Bacteriocin as Weapons in the Marine Animal-Associated Bacteria Warfare: Inventory and Potential Applications as an Aquaculture Probiotic
Abstract
:1. Introduction
2. Probiotics for Aquaculture
3. Bacteriocins
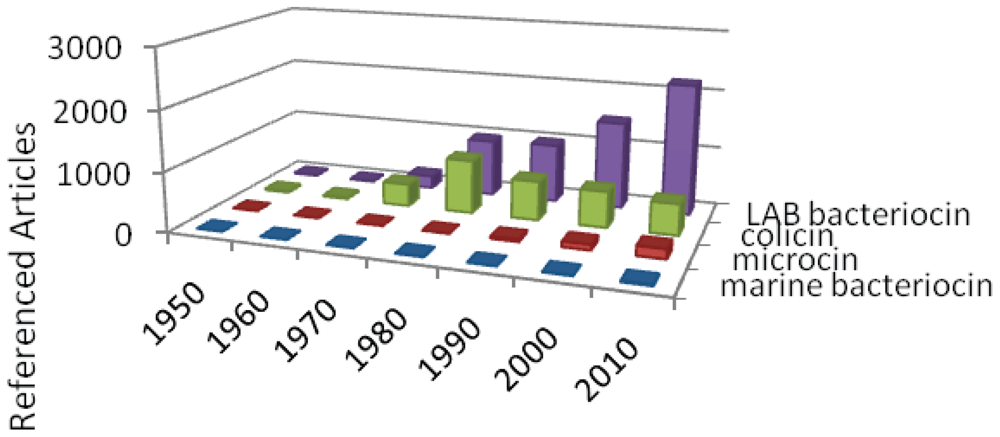
3.1. Bacteriocin story
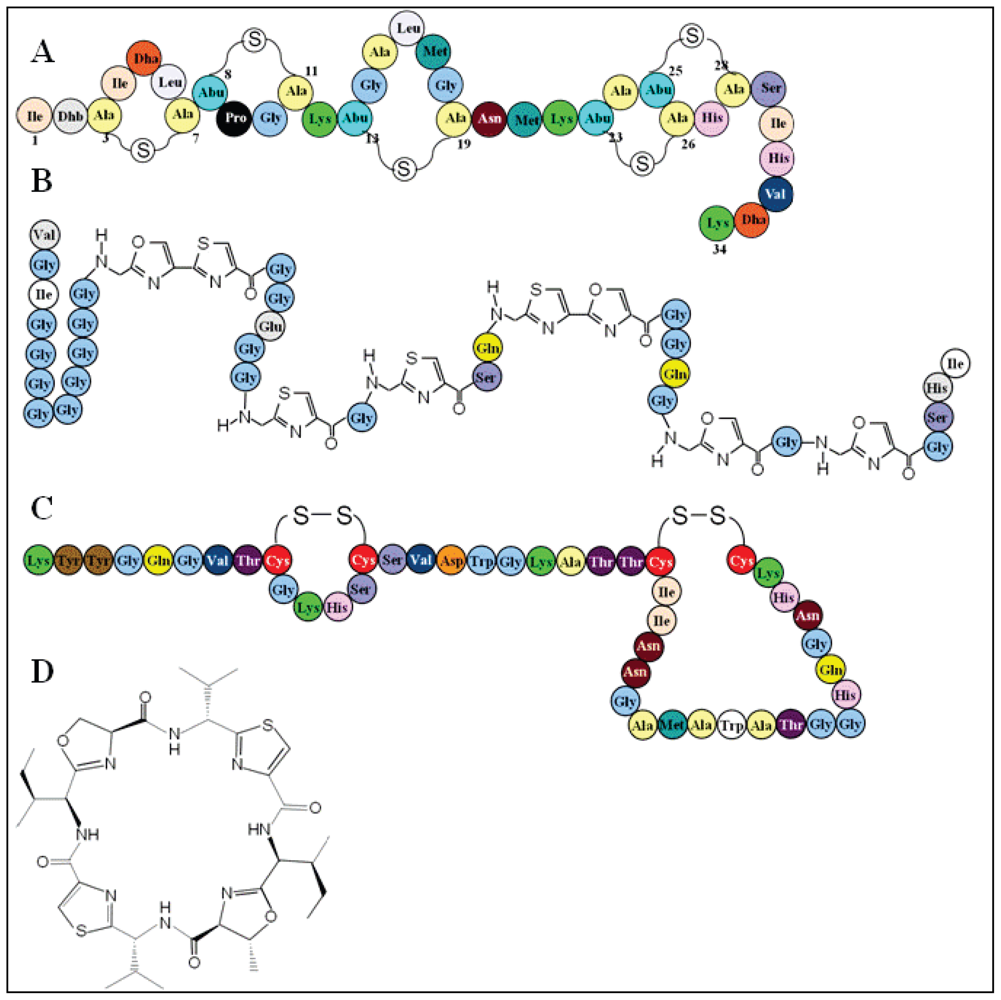
3.2. Bacteriocin classification
3.3. Bacteriocin specificity
4. Marine Animal-Associated Microorganisms as Bacteriocin Producers
4.1. BLIS from Vibrio sp
4.2. BLIS from marine Aeromonas sp
4.3. BLIS from marine Pseudoalteromonas sp
4.4. Bacteriocin from Firmicutes and LAB associated to marine animals
4.5. Bacteriocin from marine cyanobacteria
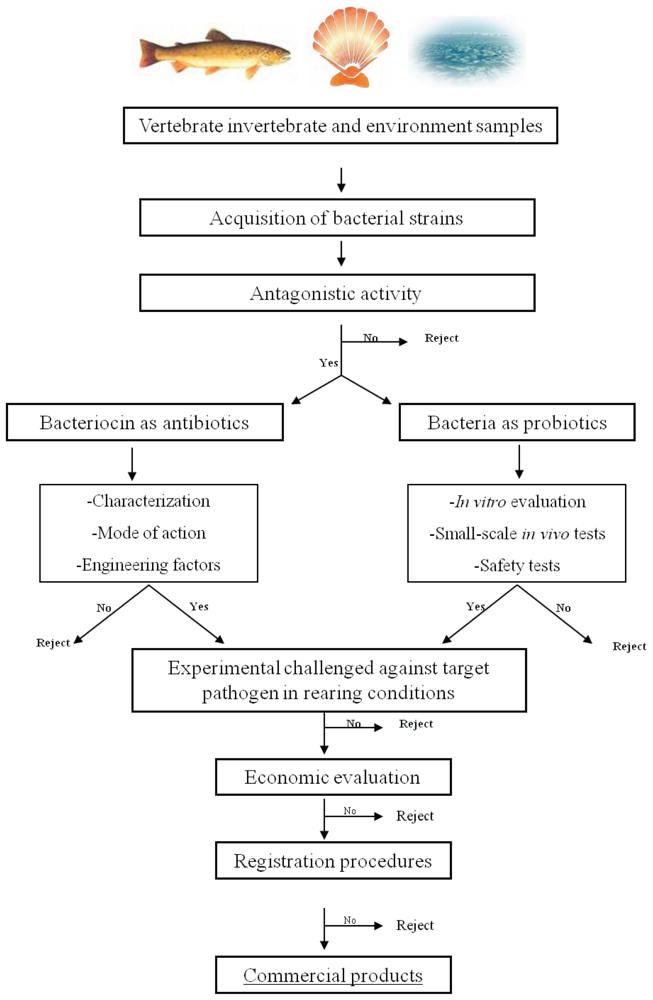
5. Bacteriocin-Based Strategy to Select a Probiotic for Aquaculture
6. Conclusions
Acknowledgements
- Samples Availability: Available from the authors.
References and Notes
- Kurath, G. Biotechnology and DNA vaccines for aquatic animals. Rev Sci Tech Off Int Epiz 2008, 27, 175–196. [Google Scholar]
- Toranzo, AE; Magariños, B; Romalde, JL. A review of the main bacterial fish diseases in mariculture systems. Aquaculture 2005, 246, 37–61. [Google Scholar]
- Austin, B; Zhang, XH. Vibrio harveyi: a significant pathogen of marine vertebrates and invertebrates. Lett Appl Microbiol 2006, 43, 119–124. [Google Scholar]
- Paillard, C; Le Roux, F; Borrego, JJ. Bacterial disease in marine bivalves, a review of recent studies: Trends and evolution. Aquat Living Res 2004, 17, 477–498. [Google Scholar]
- Marcogliese, D. The impact of climate change on the parasites and infectious diseases of aquatic animals. Rev Sci Tech Off Int Epiz 2008, 27, 467–484. [Google Scholar]
- Cabello, FC. Heavy use of prophylactic antibiotics in aquaculture: a growing problem for human and animal health and for the environment. Environ Microbiol 2006, 8, 1137–1144. [Google Scholar]
- Zhou, Q; Li, K; Jun, X; Bo, L. Role and functions of beneficial microorganisms in sustainable aquaculture. Bioresour Technol 2009, 100, 3780–3786. [Google Scholar]
- Dorrington, T; Gomez-Chiarri, M. Antimicrobial Peptides for Use in Oyster Aquaculture: Effect on Pathogens, Commensals, and Eukaryotic Expression Systems. J Shellfish Res 2008, 27, 365–374. [Google Scholar]
- Kesarcodi-Watson, A; Kaspar, H; Lategan, MJ; Gibson, L. Probiotics in aquaculture: The need, principles and mechanisms of action and screening processes. Aquaculture 2008, 274, 1–14. [Google Scholar]
- Joerger, RD. Alternatives to antibiotics: bacteriocins, antimicrobial peptides and bacteriophages. Poult Sci 2003, 82, 640–647. [Google Scholar]
- Gillor, O; Etzion, A; Riley, MA. The dual role of bacteriocins as anti- and probiotics. Appl Microbiol Biotechnol 2008, 81, 591–606. [Google Scholar]
- Gillor, O; Nigro, LM; Riley, LM. Genetically Engineered Bacteriocins and their Potential as the Next Generation of Antimicrobials. Curr Pharm Des 2005, 11, 1067–1075. [Google Scholar]
- Lenski, RE; Riley, MA. Chemical warfare from an ecological perspective. Proc Natl Acad Sci USA 2002, 99, 556–558. [Google Scholar]
- Riley, MA; Gordon, DM. The ecological role of bacteriocins in bacterial competition. Trends Microbiol 1999, 7, 129–133. [Google Scholar]
- Riley, MA; Wertz, JE. BACTERIOCINS: Evolution, Ecology, and Application. Annu Rev Microbiol 2002, 56, 117–137. [Google Scholar]
- Kollath, W. Nutrition and the tooth system; general review with special reference to vitamins. 1953, 8, 7–16. [Google Scholar]
- Parker, RB. Probiotics, the other half of the antibiotic story. Anim Nutr Health 1974, 29, 4–8. [Google Scholar]
- Fuller, R. Probiotics in man and animals. J Appl Microbiol 1989, 66, 365–378. [Google Scholar]
- Salminen, S; Ouwehand, A; Benno, Y; Lee, YK. Probiotics: how should they be defined. Trends Food Sci Technol 1999, 10, 107–110. [Google Scholar]
- Ouwehand, AC; Salminen, SJ. The health effects of cultured milk products with viable and non-viable bacteria. Int Dairy J 1998, 8, 749–758. [Google Scholar]
- Reid, G; Sanders, ME; Gaskins, HR; Gibson, GR; Mercenier, A; Rastall, R; Roberfroid, M; Rowland, I; Cherbut, C; Klaenhammer, TR. New Scientific Paradigms for Probiotics and Prebiotics. J Clin Gastroenterol 2003, 37, 105–118. [Google Scholar]
- Moriarty, DJW. Control of luminous Vibrio species in penaeid aquaculture ponds. Aquaculture 1998, 164, 351–358. [Google Scholar]
- Cahill, MM. Bacterial flora of fishes: A review. Microb Ecol 1990, 19, 21–41. [Google Scholar]
- Jorquera, MA; Silva, FR; Riquelme, CE. Bacteria in the culture of the scallop Argopecten purpuratus (Lamarck, 1819). Aquaculture Int 2001, 9, 285–303. [Google Scholar]
- Verschuere, L; Rombaut, G; Sorgeloos, P; Verstraete, W. Probiotic Bacteria as Biological Control Agents in Aquaculture. Microbiol Mol Biol Rev 2000, 64, 655–671. [Google Scholar]
- Tinh, N; Dierckens, K; Sorgeloos, P; Bossier, P. A review of the functionality of probiotics in the larviculture food chain. Mar Biotechnol 2008, 10, 1–12. [Google Scholar]
- Balcázar, JL; Blas, Id; Ruiz-Zarzuela, I; Cunningham, D; Vendrell, D; Múzquiz, JL. The role of probiotics in aquaculture. Vet Microbiol 2006, 114, 173–186. [Google Scholar]
- Bomba, A; Nemcová, R; Mudronová, D; Guba, P. The possibilities of potentiating the efficacy of probiotics. Trends Food Sci Technol 2002, 13, 121–126. [Google Scholar]
- Musa, HH; Wu, SL; Zhu, CH; Seri, HI; Zhu, GQ. The Potential Benefits of Probiotics in Animal Production and Health. J Anim Vet Adv 2009, 8, 313–321. [Google Scholar]
- Isolauri, E; Sutas, Y; Kankaanpaa, P; Arvilommi, H; Salminen, S. Probiotics: effects on immunity. Am J Clin Nutr 2001, 73, 444S–450S. [Google Scholar]
- Kelly, D; Conway, S; Aminov, R. Commensal gut bacteria: mechanisms of immune modulation. Trends Immunol 2005, 26, 326–333. [Google Scholar]
- Cornil, V; Babes, V. Les bactéries et leur rôle dans l’anatomie et l’histologie pathologiques des maladies infectieuses: ouvrage contenant les méthodes spéciales de la bactériologie; F. Alcan: Paris, France, 1885. [Google Scholar]
- Gratia, A. Sur un remarquable exemple d’antagonisme entre deux souches de colibacille. CR Soc Biol 1925, 93, 1040–1041. [Google Scholar]
- Fredericq, P; Joiris, E; Betz-Barreau, M; Gratia, A. Recherche des germes producteurs de colicines dans les selles de malades atteints de fièvre paratyphoide. C R Soc Biol 1949, 143, 556–559. [Google Scholar]
- Duquesne, S; Destoumieux-Garzon, D; Peduzzi, J; Rebuffat, S. Microcins, gene-encoded antibacterial peptides from enterobacteria. Nat Prod Rep 2007, 24, 708–734. [Google Scholar]
- Jacob, F; Lwoff, A; Siminovitch, A; Wollman, E. Définition de quelques termes relatifs à la lysogenie. Ann Inst Pasteur 1953, 84, 222–224. [Google Scholar]
- Bradley, DE. Ultrastructure of phages and bacteriocins. Bacteriol Rev 1967, 31, 230–314. [Google Scholar]
- Reeves, P. The bacteriocins. Bacteriol Rev 1965, 29, 24–45. [Google Scholar]
- Rogers, LA. The inhibitory effect of Streptococcus lactis on Lactobacillus bulgaricus. J Bacteriol 1928, 16, 321–325. [Google Scholar]
- Jansen, EF; Hirschmann, DJ. Subtilin, an antibacterial product of Bacillus subtilis: culturing conditions and properties. Arch Biochem 1944, 4, 297–304. [Google Scholar]
- Gross, E; Kiltz, HH; Nebelin, E. Subtilin, VI: the structure of subtilin (author’s transl). Hoppe Seylers Z Physiol Chem 1973, 354, 810–812. [Google Scholar]
- Gross, E; Morell, JL. Nisin. The assignment of sulfide bridges of beta-methyllanthionine to a novel bicyclic structure of identical ring size. J Am Chem Soc 1970, 92, 2919–2920. [Google Scholar]
- Anonymous. Nisin preparation: Affirmation of GRAS status as a direct human food ingredient. Fed Regist 1988, Part 184 53, 11247–11251. [Google Scholar]
- Abee, T; Krockel, L; Hill, C. Bacteriocins: modes of action and potentials in food preservation and control of food poisoning. Int J Food Microbiol 1995, 28, 169–185. [Google Scholar]
- Deegan, LH; Cotter, PD; Hill, C; Ross, PR. Bacteriocins: Biological tools for bio-preservation and shelf-life extension. Int Dairy J 2006, 16, 1058–1071. [Google Scholar]
- Gálvez, A; Abriouel, H; López, RL; Omar, NB. Bacteriocin-based strategies for food biopreservation. Int J Food Microbiol 2007, 120, 51–70. [Google Scholar]
- Nes, IF; Johnsborg, O. Exploration of antimicrobial potential in LAB by genomics. Curr Opin Biotechnol 2004, 15, 100–104. [Google Scholar]
- Papagianni, M; Anastasiadou, S. Pediocins: The bacteriocins of Pediococci. Sources, production, properties and applications. Microb Cell Fact 2009, 8, 3. [Google Scholar]
- Juncioni de Arauza, L; Jozalaa, AF; Mazzolab, PG; Vessoni Pennaa, TC. Nisin biotechnological production and application: a review. Trends Food Sci Technol 2009, 20, 146–154. [Google Scholar]
- Papagianni, M. Ribosomally synthesized peptides with antimicrobial properties: biosynthesis, structure, function and applications. Biotechnol Adv 2003, 21, 465–499. [Google Scholar]
- Holtsmark, I; Eijsink, VG; Brurberg, MB. Bacteriocins from plant pathogenic bacteria. FEMS Microbiol Lett 2008, 280, 1–7. [Google Scholar]
- Vidaver, AK. Bacteriocins: the lure and the reality. Plant Dis 1983, 67, 471–474. [Google Scholar]
- Tagg, JR; Dierksen, KP. Bacterial replacement therapy: adapting ‘germ warfare’ to infection prevention. Trends Biotechnol 2003, 21, 217–223. [Google Scholar]
- Hammami, R; Zouhir, A; Ben Hamida, J; Fliss, I. BACTIBASE: a new web-accessible database for bacteriocin characterization. BMC Microbiol 2007, 7, 89. [Google Scholar]
- de Jong, A; van Hijum, SA; Bijlsma, JJ; Kok, J; Kuipers, OP. BAGEL: a web-based bacteriocin genome mining tool. Nucleic Acids Res 2006, 34, W273–279. [Google Scholar]
- Wang, G; Li, X; Wang, Z. APD2: the updated antimicrobial peptide database and its application in peptide design. Nucleic Acids Res 2009, 37, D933–937. [Google Scholar]
- Wang, Z; Wang, G. APD: the Antimicrobial Peptide Database. Nucleic Acids Res 2004, 32, D590–592. [Google Scholar]
- Wang, CK; Kaas, Q; Chiche, L; Craik, DJ. CyBase: a database of cyclic protein sequences and structures, with applications in protein discovery and engineering. Nucleic Acids Res 2008, 36, D206–210. [Google Scholar]
- Duquesne, S; Petit, V; Peduzzi, J; Rebuffat, S. Structural and functional diversity of microcins, gene-encoded antibacterial peptides from enterobacteria. J Mol Microbiol Biotechnol 2007, 13, 200–209. [Google Scholar]
- Severinov, K; Semenova, E; Kazakov, A; Kazakov, T; Gelfand, MS. Low-molecular-weight post-translationally modified microcins. Mol Microbiol 2007, 65, 1380–1394. [Google Scholar]
- Jack, RW; Jung, G. Lantibiotics and microcins: polypeptides with unusual chemical diversity. Curr Opin Chem Biol 2000, 4, 310–317. [Google Scholar]
- Klaenhammer, TR. Genetics of bacteriocins produced by lactic acid bacteria. FEMS Microbiol Rev 1993, 12, 39–85. [Google Scholar]
- Cotter, PD; Hill, C; Ross, RP. Bacteriocins: developing innate immunity for food. Nat Rev Microbiol 2005, 3, 777–788. [Google Scholar]
- Cotter, PD; Hill, C; Ross, PR. What’s in a name? Class distinction for bacteriocins. Nat Rev Microbiol 2006, 4. [Google Scholar] [CrossRef]
- Heng, NCK; Tagg, JR. What’s in a name? Class distinction for bacteriocins. Nat Rev Microbiol 2006, 4. [Google Scholar] [CrossRef]
- Riley, MA. Molecular mechanisms of bacteriocin evolution. Annu Rev Genet 1998, 32, 255–278. [Google Scholar]
- Cascales, E; Buchanan, SK; Duche, D; Kleanthous, C; Lloubes, R; Postle, K; Riley, M; Slatin, S; Cavard, D. Colicin biology. Microbiol Mol Biol Rev 2007, 71, 158–229. [Google Scholar]
- Davies, JK; Reeves, P. Genetics of resistance to colicins in Escherichia coli K12: cross-resistance among resistance of group A. J Bacteriol 1975, 123, 102–117. [Google Scholar]
- Riley, MA; Wertz, JE. Bacteriocin diversity: ecological and evolutionary perspectives. Biochimie 2002, 84, 357–364. [Google Scholar]
- Duport, C; Baysse, C; Michel-Briand, Y. Molecular characterization of pyocin S3, a novel S-type pyocin from Pseudomonas aeruginosa. J Biol Chem 1995, 270, 8920–8927. [Google Scholar]
- Wertz, JE; Riley, MA. Chimeric nature of two plasmids of Hafnia alvei encoding the bacteriocins alveicins A and B. J Bacteriol 2004, 186, 1598–1605. [Google Scholar]
- James, R. Molecular Cloning and Purification of Klebicin B. J Gen Microbiol 1988, 134, 2525–2533. [Google Scholar]
- Riley, MA; Pinou, T; Wertz, JE; Tan, Y; Valletta, CM. Molecular characterization of the klebicin B plasmid of Klebsiella pneumoniae. Plasmid 2001, 45, 209–221. [Google Scholar]
- Jabrane, A; Sabri, A; Compere, P; Jacques, P; Vandenberghe, I; Van Beeumen, J; Thonart, P. Characterization of serracin P, a phage-tail-like bacteriocin, and its activity against Erwinia amylovora, the fire blight pathogen. Appl Environ Microbiol 2002, 68, 5704–5710. [Google Scholar]
- Heu, S; Oh, J; Kang, Y; Ryu, S; Cho, SK; Cho, Y; Cho, M. gly gene cloning and expression and purification of glycinecin A, a bacteriocin produced by Xanthomonas campestris pv. glycines 8ra. Appl Environ Microbiol 2001, 67, 4105–4110. [Google Scholar]
- Pham, HT; Riu, KZ; Jang, KM; Cho, SK; Cho, M. Bactericidal activity of glycinecin A, a bacteriocin derived from Xanthomonas campestris pv. glycines, on phytopathogenic Xanthomonas campestris pv. vesicatoria cells. Appl Environ Microbiol 2004, 70, 4486–4490. [Google Scholar]
- Strauch, E; Kaspar, H; Schaudinn, C; Dersch, P; Madela, K; Gewinner, C; Hertwig, S; Wecke, J; Appel, B. Characterization of enterocoliticin, a phage tail-like bacteriocin, and its effect on pathogenic Yersinia enterocolitica strains. Appl Environ Microbiol 2001, 67, 5634–5642. [Google Scholar]
- Nguyen, HA; Kaneko, J; Kamio, Y. Temperature-dependent production of carotovoricin Er and pectin lyase in phytopathogenic Erwinia carotovora subsp. carotovora Er. Biosci Biotech Biochem 2002, 66, 444–447. [Google Scholar]
- Joerger, MC; Klaenhammer, TR. Cloning, expression, and nucleotide sequence of the Lactobacillus helveticus 481 gene encoding the bacteriocin helveticin J. J Bacteriol 1990, 172, 6339–6347. [Google Scholar]
- Beukes, M; Bierbaum, G; Sahl, HG; Hastings, JW. Purification and partial characterization of a murein hydrolase, millericin B, produced by Streptococcus milleri NMSCC 061. Appl Environ Microbiol 2000, 66, 23–28. [Google Scholar]
- Nilsen, T; Nes, IF; Holo, H. Enterolysin A, a cell wall-degrading bacteriocin from enterococcus faecalis LMG 2333. Appl. Environ Microbiol 2003, 69, 2975–2984. [Google Scholar]
- Kumar, JK. Lysostaphin: an antistaphylococcal agent. Appl Environ Microbiol 2008, 80, 555–561. [Google Scholar]
- Trayer, HR; Buckley, CE. Molecular properties of Lysostaphin, a bacteriolytic agent specific for staphylococcus aureus. J Biol Chem 1970, 245, 4842–4846. [Google Scholar]
- Brotz, H; Sahl, HG. New insights into the mechanism of action of lantibiotics--diverse biological effects by binding to the same molecular target. J Antimicrob Chemother 2000, 46, 1–6. [Google Scholar]
- Nagao, J; Asaduzzaman, SM; Aso, Y; Okuda, K; Nakayama, J; Sonomoto, K. Lantibiotics: insight and foresight for new paradigm. J Biosci Bioeng 2006, 102, 139–149. [Google Scholar]
- Dufour, A; Hindré, T; Haras, D; Le Pennec, J-P. The biology of lantibiotics from the lacticin481 group is coming of age. FEMS Microbiol Rev 2007, 31, 134–167. [Google Scholar]
- Drider, D; Fimland, G; Hechard, Y; McMullen, LM; Prevost, H. The continuing story of class IIa bacteriocins. Microbiol Mol Biol Rev 2006, 70, 564–582. [Google Scholar]
- Oppegård, C; Rogne, P; Emanuelsen, L; Kristiansen, PE; Fimland, G; Nissen-Meyer, J. The Two-Peptide Class II bacteriocins: structure, production, and mode of action. J Mol Microbiol Biotechnol 2007, 13, 210–219. [Google Scholar]
- Maqueda, M; Sánchez-Hidalgo, M; Fernández, M; Montalbán-López, M; Valdivia, E; Martínez-Bueno, M. Genetic features of circular bacteriocins produced by Gram-positive bacteria. FEMS Microbiol Rev 2008, 32, 2–22. [Google Scholar]
- Martin-Visscher, LA; Gong, X; Duszyk, M; Vederas, JC. The three-dimensional structure of carnocyclin A reveals that many circular bacteriocins share a common structural motif. J Biol Chem 2009, 284, 28674–28681. [Google Scholar]
- Diep, DB; Skaugen, M; Salehian, Z; Holo, H; Nes, IF. Common mechanisms of target cell recognition and immunity for class II bacteriocins. Proc Natl Acad Sci USA 2007, 104, 2384–2389. [Google Scholar]
- Schmidt, EW; Nelson, JT; Rasko, DA; Sudek, S; Eisen, JA; Haygood, MG; Ravel, J. Patellamide A and C biosynthesis by a microcin-like pathway in Prochloron didemni, the cyanobacterial symbiont of Lissoclinum patella. Proc Natl Acad Sci USA 2005, 102, 7315–7320. [Google Scholar]
- Kiss, A; Baliko, G; Csorba, A; Chuluunbaatar, T; Medzihradszky, KF; Alfoldi, L. Cloning and characterization of the DNA region responsible for Megacin A-216 production in Bacillus megaterium 216. J Bacteriol 2008, 190, 6448–6457. [Google Scholar]
- Pons, AM; Lanneluc, I; Cottenceau, G; Sable, S. New developments in non-post translationally modified microcins. Biochimie 2002, 84, 531–537. [Google Scholar]
- Parks, WM; Bottrill, AR; Pierrat, OA; Durrant, MC; Maxwell, A. The action of the bacterial toxin, microcin B17, on DNA gyrase. Biochimie 2007, 89, 500–507. [Google Scholar]
- Bieler, S; Silva, F; Soto, C; Belin, D. Bactericidal activity of both secreted and nonsecreted microcin E492 requires the mannose permease. J Bacteriol 2006, 188, 7049–7061. [Google Scholar]
- Bastos, MC; Ceotto, H; Coelho, ML; Nascimento, JS. Staphylococcal antimicrobial peptides: relevant properties and potential biotechnological applications. Curr Pharm Biotechnol 2009, 10, 38–61. [Google Scholar]
- Bierbaum, G; Sahl, HG. Lantibiotics: mode of action, biosynthesis and bioengineering. Curr Pharm Biotechnol 2009, 10, 2–18. [Google Scholar]
- Breukink, E. A lesson in efficient killing from two-component lantibiotics. Mol Microbiol 2006, 61, 271–273. [Google Scholar]
- Cooper, LE; McClerren, AL; Chary, A; van der Donk, WA. Structure-activity relationship studies of the two-component lantibiotic haloduracin. Chem Biol 2008, 15, 1035–1045. [Google Scholar]
- Lawton, EM; Ross, RP; Hill, C; Cotter, PD. Two-peptide lantibiotics: a medical perspective. Mini-Rev Med Chem 2007, 7, 1236–1247. [Google Scholar]
- Nissen-Meyer, J; Rogne, P; Oppegard, C; Haugen, HS; Kristiansen, PE. Structure-Function Relationships of the Non-Lanthionine-Containing Peptide (class II) Bacteriocins Produced by Gram-Positive Bacteria. Curr Pharm Biotechnol 2009, 10, 19–37. [Google Scholar]
- Draper, LA; Ross, RP; Hill, C; Cotter, PD. Lantibiotic immunity. Curr Protein Pept Sci 2008, 9, 39–49. [Google Scholar]
- Lubelski, J; Rink, R; Khusainov, R; Moll, GN; Kuipers, OP. Biosynthesis, immunity, regulation, mode of action and engineering of the model lantibiotic nisin. Cell Mol Life Sci 2008, 65, 455–476. [Google Scholar]
- Kjos, M; Nes, IF; Diep, DB. Class II one-peptide bacteriocins target a phylogenetically defined subgroup of mannose phosphotransferase systems on sensitive cells. Microbiology 2009, 155, 2949–2961. [Google Scholar]
- Romanenko, LA; Uchino, M; Kalinovskaya, NI; Mikhailov, VV. Isolation, phylogenetic analysis and screening of marine mollusc-associated bacteria for antimicrobial, hemolytic and surface activities. Microbiol Res 2008, 163, 633–644. [Google Scholar]
- Wilson, GS; Raftos, DA; Corrigan, SL; Nair, SV. Diversity and antimicrobial activities of surface-attached marine bacteria from Sydney Harbour, Australia. Microbiol Res 2009. in Press. [Google Scholar]
- Selvin, J; Joseph, S; Asha, KR; Manjusha, WA; Sangeetha, VS; Jayaseema, DM; Antony, MC; Denslin Vinitha, AJ. Antibacterial potential of antagonistic Streptomyces sp. isolated from marine sponge Dendrilla nigra. FEMS Microbiol Ecol 2004, 50, 117–122. [Google Scholar]
- Morris, JJÂG. Cholera and Other Types of Vibriosis: A Story of Human Pandemics and Oysters on the Half Shell. Clin Infect Dis 2003, 37, 272–280. [Google Scholar]
- Zai, AS; Ahmad, S; Rasool, SA. Bacteriocin production by indigenous marine catfish associated Vibrio spp. Pak J Pharm Sci 2009, 22, 162–167. [Google Scholar]
- Carraturo, A; Raieta, K; Ottaviani, D; Russo, GL. Inhibition of Vibrio parahaemolyticus by a bacteriocin-like inhibitory substance (BLIS) produced by Vibrio mediterranei 1. J Appl Microbiol 2006, 101, 234–241. [Google Scholar]
- Prasad, S; Morris, PC; Hansen, R; Meaden, PG; Austin, B. A novel bacteriocin-like substance (BLIS) from a pathogenic strain of Vibrio harveyi. Microbiology 2005, 151, 3051–3058. [Google Scholar]
- Zhang, X-H; Austin, B. Pathogenicity of Vibrio harveyi to salmonids. J Fish Dis 2000, 23, 93–102. [Google Scholar]
- McCall, JO; Sizemore, RK. Description of a bacteriocinogenic plasmid in Beneckea harveyi. Appl Environ Microbiol 1979, 38, 974–979. [Google Scholar]
- Hoyt, PR; Sizemore, RK. Competitive Dominance by a Bacteriocin-Producing Vibrio harveyi Strain. Appl Environ Microbiol 1982, 44, 653–658. [Google Scholar]
- Shehane, SD; Sizemore, RK. Isolation and preliminary characterization of bacteriocins produced by Vibrio vulnificus. J Appl Microbiol 2002, 92, 322–328. [Google Scholar]
- Sugita, H; Matsuo, N; Hirose, Y; Iwato, M; Deguchi, Y. Vibrio sp. strain NM 10, isolated from the intestine of a Japanese coastal fish, has an inhibitory effect against Pasteurella piscicida. Appl Environ Microbiol 1997, 63, 4986–4989. [Google Scholar]
- Pedron Moro, EM; Niederauer Weiss, RD; Salete Friedrich, R; Paiva Nunes, M. Bacteriocin-like Substance of Aeromonas hydrophila. Mem Inst Oswaldo Cruz 1997, 92, 115–116. [Google Scholar]
- Messi, P; Guerrieri, E; Bondi, M. Bacteriocin-like substance (BLS) production in Aeromonas hydrophila water isolates. FEMS Microbiol Lett 2003, 220, 121–125. [Google Scholar]
- Pirzada, ZA; Ali, SA; Khan, BM; Rasool, SA. Production And Physico-Chemical Characterization Of Bacteriocins-Like Inhibitory Substances From Marine Bacterium ZM81. Pak J Biol Sci 2004, 7, 2026–2030. [Google Scholar]
- Longeon, A; Peduzzi, J; Barthelemy, M; Corre, S; Nicolas, J-L; Guyot, M. Purification and Partial Identification of Novel Antimicrobial Protein from Marine Bacterium Pseudoalteromonas Species Strain X153. Mar Biotechnol 2004, 6, 633–641. [Google Scholar]
- Ringø, E; Gatesoupe, FJ. Lactic acid bacteria in fish: a review. Aquaculture 1998, 160, 177–203. [Google Scholar]
- Rihakova, J; Belguesmia, Y; Petit, VW; Pilet, MF; Prevost, H; Dousset, X; Drider, D. Divercin V41 from gene characterization to food applications: 1998–2008, a decade of solved and unsolved questions. Lett Appl Microbiol 2009, 48, 1–7. [Google Scholar]
- Hosseini, SV; Arlindo, S; Böhme, K; Fernández-No, C; Calo-Mata, P; Barros-Velázquez, J. Molecular and probiotic characterization of bacteriocin-producing Enterococcus faecium strains isolated from non fermented animal foods. J Appl Microbiol 2009, 107, 1392–1403. [Google Scholar]
- Pinto, AL; Fernandes, M; Pinto, C; Albano, H; Castilho, F; Teixeira, P; Gibbs, PA. Characterization of anti-Listeria bacteriocins isolated from shellfish: Potential antimicrobials to control non-fermented seafood. Int J Food Microbiol 2009, 129, 50–58. [Google Scholar]
- Duffes, F; Leroi, F; Boyaval, P; Dousset, X. Inhibition of Listeria monocytogenes by Carnobacterium spp. strains in a simulated cold smoked fish system stored at 4 °C. Int J Food Microbiol 1999, 47, 33–42. [Google Scholar]
- Métivier, A; Pilet, M-F; Dousset, X; Sorokine, O; Anglade, P; Zagorec, M; Piard, J-C; Marion, D; Cenatiempo, Y; Fremaux, C. Divercin V41, a new bacteriocin with two disulphide bonds produced by Carnobacterium divergens V41: primary structure and genomic organization. Microbiology 1998, 144, 2837–2844. [Google Scholar]
- Pilet, M-F; Dousser, X; Barre, R; Novel, G; Dezmazeaud, M; Piard, J-C. Evidence for two bacteriocins produced by Carnobacterium piscicola and Carnobacterium divergens isolated from fish and active against Listeria monocytogenes. J Food Prot 1995, 58, 256–262. [Google Scholar]
- Richard, C; Drider, D; Elmorjani, K; Marion, D; Prevost, H. Heterologous Expression and Purification of Active Divercin V41, a Class IIa Bacteriocin Encoded by a Synthetic Gene in Escherichia coli. J Bacteriol 2004, 186, 4276–4284. [Google Scholar]
- Bhugaloo-Vial, P; Dousset, X; Metivier, A; Sorokine, O; Anglade, P; Boyaval, P; Marion, D. Purification and amino acid sequences of piscicocins V1a and V1b, two class IIa bacteriocins secreted by Carnobacterium piscicola V1 that display significantly different levels of specific inhibitory activity. Appl Environ Microbiol 1996, 62, 4410–4416. [Google Scholar]
- Moore, BS. Biosynthesis of marine natural products: microorganisms (Part A). Nat Prod Rep 2005, 22, 580–593. [Google Scholar]
- Sudek, S; Haygood, MG; Youssef, DTA; Schmidt, EW. Structure of Trichamide, a Cyclic Peptide from the Bloom-Forming Cyanobacterium Trichodesmium erythraeum, Predicted from the Genome Sequence. Appl Environ Microbiol 2006, 72, 4382–4387. [Google Scholar]
- Ziemert, N; Ishida, K; Quillardet, P; Bouchier, C; Hertweck, C; de Marsac, NT; Dittmann, E. Microcyclamide Biosynthesis in Two Strains of Microcystis aeruginosa: from Structure to Genes and Vice Versa. Appl Environ Microbiol 2008, 74, 1791–1797. [Google Scholar]
- Long, PF; Dunlap, WC; Battershill, CN; Jaspars, M. Shotgun Cloning and Heterologous Expression of the Patellamide Gene Cluster as a Strategy to Achieving Sustained Metabolite Production13. Chem Bio Chem 2005, 6, 1760–1765. [Google Scholar]
- Jüttner, F; Todorova, AK; Walch, N; von Philipsborn, W. Nostocyclamide M: a cyanobacterial cyclic peptide with allelopathic activity from Nostoc 31. Phytochemistry 2001, 57, 613–619. [Google Scholar]
- Banker, R; Carmeli, S. Tenuecyclamides A-D, Cyclic Hexapeptides from the Cyanobacterium Nostoc spongiaeforme var. tenue. J Nat Prod 1998, 61, 1248–1251. [Google Scholar]
- Linington, RG; Gonzalez, J; Urena, L-D; Romero, LI; Ortega-Barria, E; Gerwick, WH. Venturamides A and B: Antimalarial Constituents of the Panamanian Marine Cyanobacterium Oscillatoria sp. J Nat Prod 2007, 70, 397–401. [Google Scholar]
- Ogino, J; Moore, RE; Patterson, GML; Smith, CD. Dendroamides, New Cyclic Hexapeptides from a Blue-Green Alga. Multidrug-Resistance Reversing Activity of Dendroamide A. J Nat Prod 1996, 59, 581–586. [Google Scholar]
- Ishida, K; Nakagawa, H; Murakami, M. Microcyclamide, a Cytotoxic Cyclic Hexapeptide from the Cyanobacterium Microcystis aeruginosa. J Nat Prod 2000, 63, 1315–1317. [Google Scholar]
- Gatesoupe, FJ. Updating the importance of lactic acid bacteria in fish farming: natural occurrence and probiotic treatments. J Mol Microbiol Biotechnol 2008, 14, 107–114. [Google Scholar]
- Wang, Y-B; Li, J-R; Lin, J. Probiotics in aquaculture: Challenges and outlook. Aquaculture 2008, 281, 1–4. [Google Scholar]
- Hong, HA; Duc, LH; Cutting, SM. The use of bacterial spore formers as probiotics. FEMS Microbiol Rev 2005, 29, 813–835. [Google Scholar]
- Vine, NG; Leukes, WD; Kaiser, H. Probiotics in marine larviculture. FEMS Microbiol Rev 2006, 30, 404–427. [Google Scholar]
- Das, S; Ward, L; Burke, C. Prospects of using marine actinobacteria as probiotics in aquaculture. Appl Microbiol Biotechnol 2008, 81, 419–429. [Google Scholar]
- von Wright, A. Regulating the Safety of Probiotics - The European Approach. Curr Pharm Des 2005, 11, 17–23. [Google Scholar]
- 101.70, C. Subpart E-Specific Requirements for Health Claims. Code Fed Regul 2005, 21, 126–129.
- Sahu, M; Swarnakumar, N; Sivakumar, K; Thangaradjou, T; Kannan, L. Probiotics in aquaculture: importance and future perspectives. Indian J Microbiol 2008, 48, 299–308. [Google Scholar]
- Guo, J-J; Liu, K-F; Cheng, S-H; Chang, CI; Lay, J-J; Hsu, Y-O; Yang, J-Y; Chen, T-I. Selection of probiotic bacteria for use in shrimp larviculture. Aquaculture Res 2009, 40, 609–618. [Google Scholar]
- Ruiz-Ponte, C; Samain, JF; Sánchez, JL; Nicolas, JL. The Benefit of a Roseobacter Species on the Survival of Scallop Larvae. Mar Biotechnol 1999, 1, 52–59. [Google Scholar]
| (A) | ||||||
|---|---|---|---|---|---|---|
| Protein-Bacteriocins | Class | Sub-Class | Name | MM (kDa) | Mode of action | Ref. |
| Gracilicutes | ||||||
| Escherichia coli | Colicins | Groupe A | 40 to 80 | Nuclease/Pore-forming | [69] | |
| Groupe B | 40 to 80 | Nuclease/Pore-forming | [69] | |||
| Pseudomonas aeruginosa | Pyocins | R-type | Pyocin R2 | 270 (AA) | Pore-forming | |
| S-type | Pyocin S1,S2,AP41 | 75/84/94 | Phage-tail like | [70] | ||
| F-type | Pyocin F | Phage-tail like | ||||
| Hafnia alvei | Alveicins | Colicin like | Alveicin A, B | 408/358 (AA) | Pore forming | [71] |
| Klebsiella pneumonia | Klebicin | Colicin-like | Klebicin C, D | 96 | Nuclease | [72,73] |
| Serratia plymithicum | Serracin | Serracin P | 66 | Phage-tail like | [74] | |
| Xanthomonas campestris | Glynericin | Glynericin A | 50 | Phage tail like | [75,76] | |
| Yersinia enterocolitica | Enterocoliticin | 669 | Phage tail like | [77] | ||
| Erwinia carotovora | Carotovoricin | Carotovoricin Er | 68/76 | Phage tail like | [78] | |
| Firmicutes | ||||||
| Lactobacillus helveticus | Helveticin J | Class III | 37,5 | to be defined | [79] | |
| Streptococcus milleri | Millericin | Class III | 30 | Peptidoglycan hydrolysis | [80] | |
| Enterococcus faecalis | Enterolysin | Class III | 34,5 | Peptidoglycan hydrolysis | [81] | |
| Staphylococcus aureus | Lysostaphin | Class III | 25 | Peptidoglycan hydrolysis | [82,83] | |
| (B) | ||||||||
|---|---|---|---|---|---|---|---|---|
| Peptide-Bacteriocin | Class | Sub-Class | Name | MM (kDa) | PTM | Mode of action | Ref. | |
| Gracilicutes | ||||||||
| Escherichia coli | Microcin | Class I | Microcin B17 | 3.1 | drastic | intracellular enzymes | ||
| Class II | IIa | Microcin V | 8.8 | light | pore-forming | [35,59,61] | ||
| IIb | Microcin E492 | 7.9 | drastic | pore forming | ||||
| Firmicutes | ||||||||
| Lactic acid bacteria | Class I | A-type | A1 | Nisin | 3.5 | drastic | pore-forming | [84,85] |
| (mainly) | or Lantibiotic | A2 | Lacticin 481 | 3 | drastic | pore forming | [86] | |
| B-type | Mersacidin | 2 | [61] | |||||
| Class II | class IIa | Pediocin | 4.6 | light | pore forming | [48,87] | ||
| class IIb | Plantaricin E/F | 3.5/3.7 | light | pore forming | [88] | |||
| Class IIc | carnocyclin A | 5.8 | cyclic | pore forming | [89,90] | |||
| Class IId | Lactococcin A | 5.8 | none | pore forming | [91] | |||
| Cyanobacteria | ||||||||
| Prochloron didemni | microcin –like | - | Patellamides | 0.7 | drastic | [92] | ||
| Producing strain | Bacteriocin | Inhibited strain(s) | Isolated from | MM (kDa) | Ref. |
|---|---|---|---|---|---|
| Listonella anguillarum AVP10 | Vibriocin AVP10 | Escherichia coli Listonella anguillarum AVS91 | Healthy and infected catfishes (Arius thalassimus) | ? | [110] |
| Vibrio mediterranei | BLIS | V. parahaemolyticus V. mediterranei 5 | Fresh & frozen seafood | 63–65a | [111] |
| Vibrio harveyi VIB 571 | BLIS | Vibrio harveyi1 V. fischeri V. gazogenes V. parahaemolyticus | - | ~32a,b | [112] |
| Vibrio harveyi (Beneckea harveyi SY) | Harveyicin SY | V. harveyi1 | area of Galveston Island | 24 | [114,115] |
| Vibrio vulnificus | IW1 | V. vulnificus V. cholera | Water samples from Wilmington (NC, USA) | 9 | [116] |
| BC1 | V. parahaemolyticus | 7,5 | |||
| BC2 | Vibrio spp. Plesiomonas shigelloides E. coli | 1,35 | |||
| Vibrio sp. Strain NM 10 | BLIS | Pasteurella piscicida K-III; E. coli; V. vulnificus Enterococcus seriolicida | Leiognathus nuchalis intestine | < 5d | [117] |
| Bacteriocinogenic strain marine strain ZM81 (Gram positif pleomorphic strain) | Bacteriocins/BLIS | Marine bacterial strain ZM19 | Open sea region of Karachi coast | >10 | [120] |
| Aeromonas hydrophila | BLIS | Staphylococcus aureus strains | Water tank containing alligators | ? | [118] [119] |
| Pseudoalteromonas Species Strain X153 | Antibiotic protein P-153 | Ichthyopathogenic Vibrio1 Staphylococcus epidermidis Propionibacterium acnes Propionibacterium granulosum | Substrates on the littoral of Brittany | 280a,b | [121] |
| Producing strain | Bacteriocin | Inhibited strain(s) | Isolated from | MM (kDa) | Ref. |
|---|---|---|---|---|---|
| Enterococcus faecium LHICA 28.4, 34.5, 40.4, 46 | Enterocin P | Carnobacterium maltaromaticum Listeria monocytogenes Staphylococcus aureus | Turbot muscle | [124] | |
| Enterococcus faecium ALP7 | bac ALP7 | Listeria monocytogenes | Non-fermented shellfish including oysters, mussels and clams | <10 | [125] |
| Pediococcus pentosaceus ALP57 | bac ALP57 | Bacillus subtilis Enterococcus faecalis Lactobacillus brevis gravensis; Lactobacillus curvatus Listeria innocua | |||
| Carnobacterium divergens V41 | Divercin V41 | Listeria monocytogenes | Salmon intestine | 4,509 | [126–129] |
| Carnobacterium piscicola V1 | Piscicocin V1a Piscicocin V1b | Listeria monocytogenes | Trout intestine | 4,416 4,526 | [128,130] |
Abbreviations
| APD2 | Antimicrobial peptide database 2 |
| BLIS | Bacteriocin-like inhibitory substance |
| FDA | Food and Drug Administration |
| GRAS | Generally recognize as safe |
| LAB | Lactic acid bacteria |
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Desriac, F.; Defer, D.; Bourgougnon, N.; Brillet, B.; Le Chevalier, P.; Fleury, Y. Bacteriocin as Weapons in the Marine Animal-Associated Bacteria Warfare: Inventory and Potential Applications as an Aquaculture Probiotic. Mar. Drugs 2010, 8, 1153-1177. https://doi.org/10.3390/md8041153
Desriac F, Defer D, Bourgougnon N, Brillet B, Le Chevalier P, Fleury Y. Bacteriocin as Weapons in the Marine Animal-Associated Bacteria Warfare: Inventory and Potential Applications as an Aquaculture Probiotic. Marine Drugs. 2010; 8(4):1153-1177. https://doi.org/10.3390/md8041153
Chicago/Turabian StyleDesriac, Florie, Diane Defer, Nathalie Bourgougnon, Benjamin Brillet, Patrick Le Chevalier, and Yannick Fleury. 2010. "Bacteriocin as Weapons in the Marine Animal-Associated Bacteria Warfare: Inventory and Potential Applications as an Aquaculture Probiotic" Marine Drugs 8, no. 4: 1153-1177. https://doi.org/10.3390/md8041153
APA StyleDesriac, F., Defer, D., Bourgougnon, N., Brillet, B., Le Chevalier, P., & Fleury, Y. (2010). Bacteriocin as Weapons in the Marine Animal-Associated Bacteria Warfare: Inventory and Potential Applications as an Aquaculture Probiotic. Marine Drugs, 8(4), 1153-1177. https://doi.org/10.3390/md8041153






