Application-Oriented Marine Isomerases in Biocatalysis
Abstract
1. Introduction
2. Literature Search
3. Application-Oriented Biocatalysts
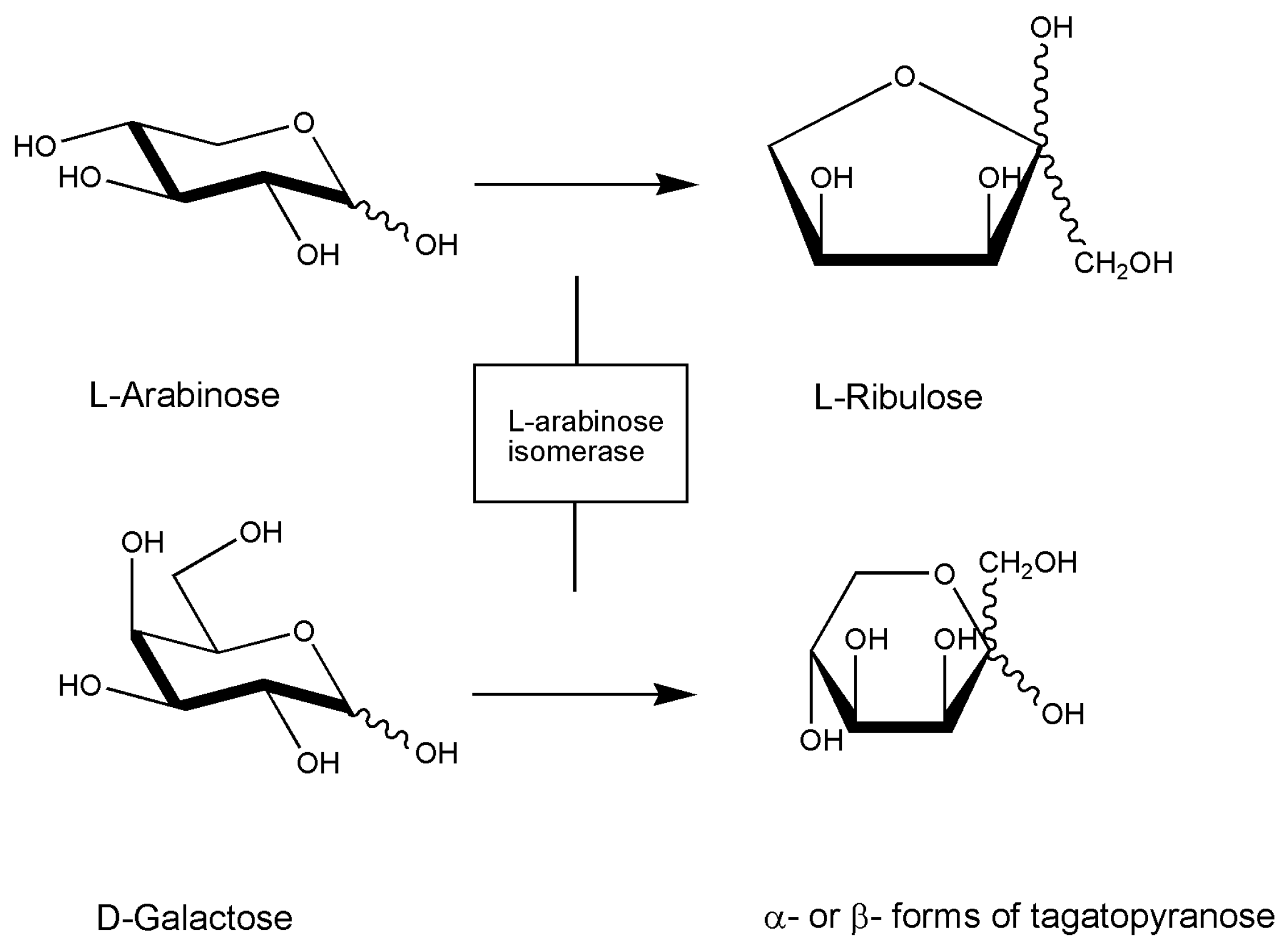
4. Marine Isomerases Acting on Sugar Molecules
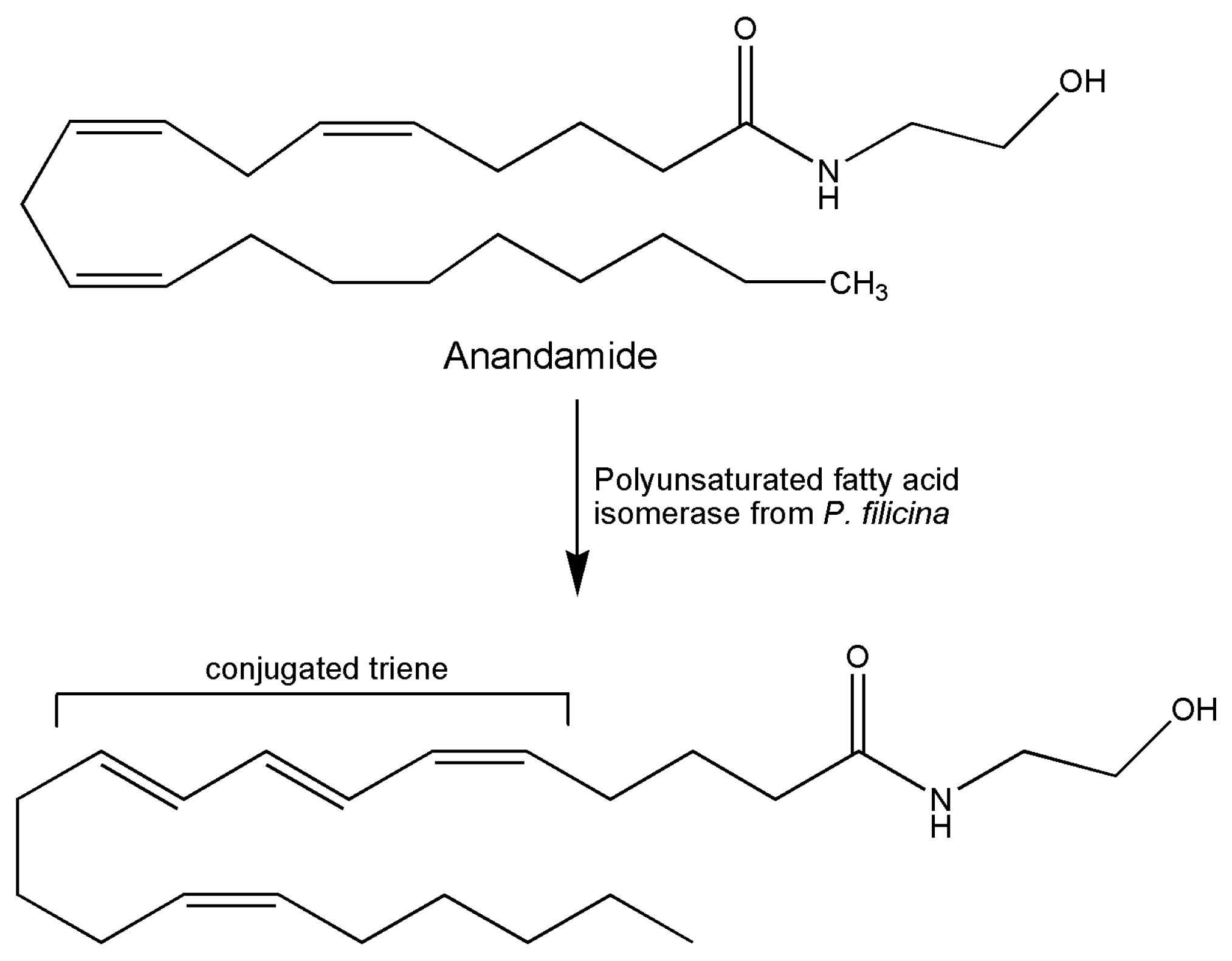
5. Marine Isomerases Acting on Lipid Molecules
6. Marine Isomerases Acting on Amino Acids and Peptides
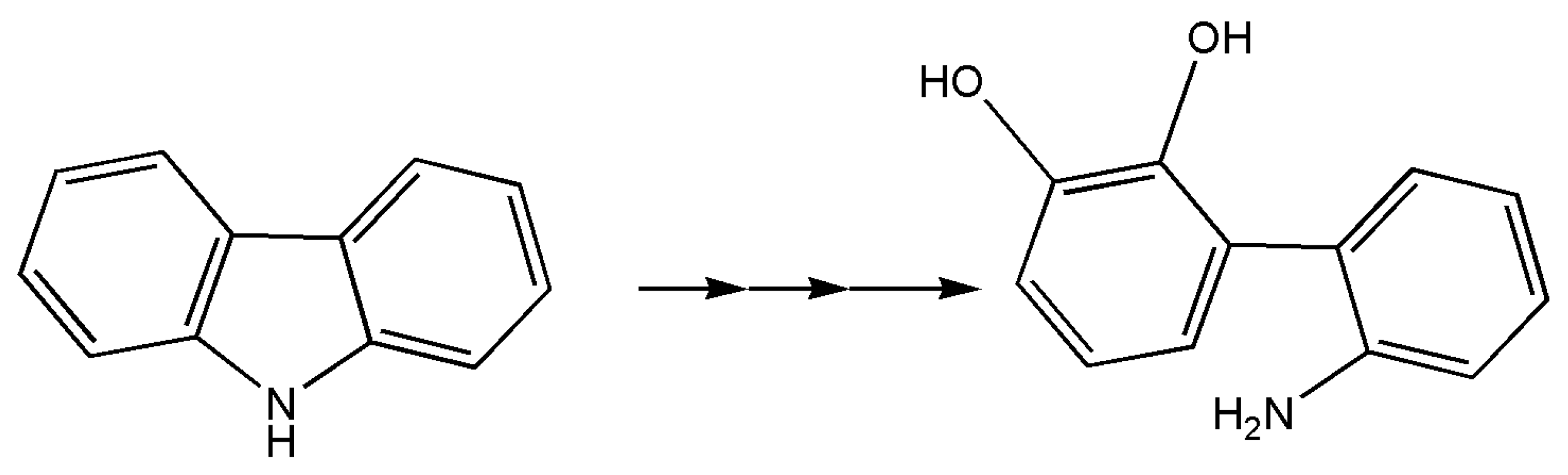
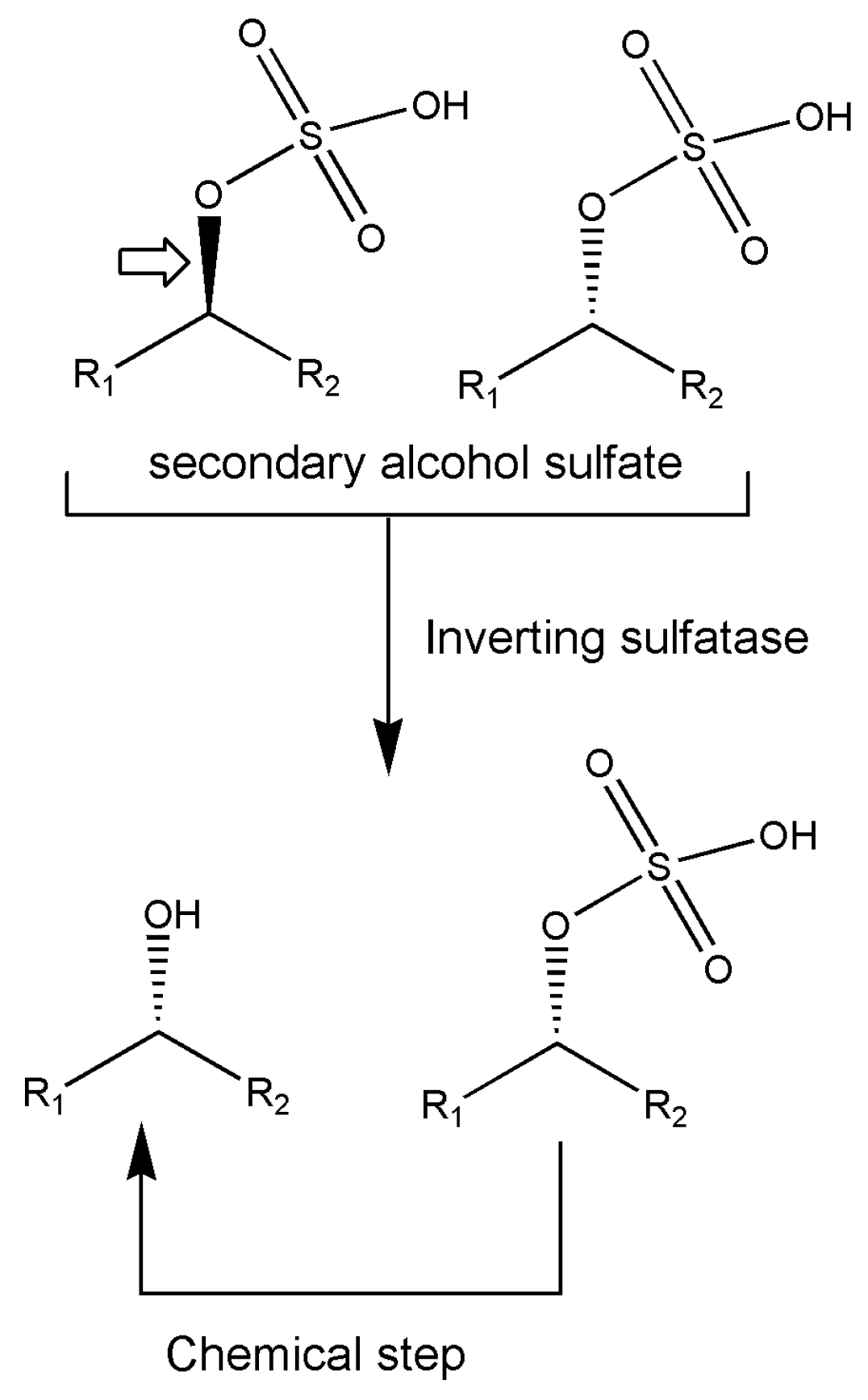
7. Other Enzymes
8. Conclusions
Funding
Conflicts of Interest
References
- Trincone, A. Potential biocatalysts originating from sea environments. J. Mol. Catal. B Enzym. 2010, 66, 241–256. [Google Scholar] [CrossRef]
- Uo, T.; Ueda, M.; Nishiyama, T.; Yoshimura, T.; Esaki, N. Purification and characterization of alanine racemase from hepatopancreas of black-tiger prawn, Penaeus monodon. J. Mol. Catal. B Enzym. 2001, 12, 137–144. [Google Scholar] [CrossRef]
- Zheng, W.; Wise, M.L.; Wyrick, A.; Metz, J.G.; Yuan, L.; Gerwick, W.H. Polyenoic fatty acid isomerase from the marine alga Ptilota filicina: Protein characterization and functional expression of the cloned cDNA. Arch. Biochem. Biophys. 2002, 401, 11–20. [Google Scholar] [CrossRef]
- Lima, R.N.; Porto, A.L.M. Recent advances in marine enzymes for biotechnological processes. In Marine Enzymes Biotechnology: Production and Industrial Applications, Part I—Production of Enzymes; Kim, S.-K., Toldra, F., Eds.; Academic Press Elsevier: New York, NY, USA, 2016; pp. 153–192. [Google Scholar] [CrossRef]
- Sun, H.; Zhang, H.; Ang, E.L.; Zhao, H. Biocatalysis for the synthesis of pharmaceuticals and pharmaceutical intermediates. Bioorg. Med. Chem. 2018, 26, 1275–1284. [Google Scholar] [CrossRef]
- Femmer, C.; Bechtold, M.; Roberts, T.M.; Panke, S. Exploiting racemases. Appl. Microbiol. Biotechnol. 2016, 100, 7423–7436. [Google Scholar] [CrossRef] [PubMed]
- Ju, X.; Xu, X.; Shen, M.; Mo, X.; Fan, H.; Liangzhi Li, L. Biochemical and structural insights into an Ochrobactrum sp. CSL1 ribose-5-phosphate isomerase A and its roles in isomerization of rare sugars. Enz. Microb. Technol. 2020, 140, 109604. [Google Scholar] [CrossRef]
- Noel, P.; Wilkins, P. Phosphoglucose isomerase in marine molluscs. In Isozymes; Markert, C.L., Ed.; Academic Press: Cambridge, MA, USA, 1975; pp. 931–943. [Google Scholar] [CrossRef]
- Madgwick, J.; Haug, A.; Larsen, B. Polymannuronic acid 5-epimerase from the marine alga Pelvetia canaliculata (L.) Dcne. et Thur. Acta Chem. Scand. 1973, 27, 3592–3594. [Google Scholar] [CrossRef]
- Sawyer, M.H.; Baumann, P.; Baumann, L. Pathways of D-fructose and D-glucose catabolism in marine species of Alcaligenes, Pseudomonas marina, and Alteromonas communis. Arch. Microbiol. 1977, 112, 169–172. [Google Scholar] [CrossRef]
- Lavie, B.; Nevo, E.; Zoller, U. Differential viability of phosphoglucose isomerase allozyme genotypes of marine snails in nonionic detergent and crude oil-surfactant mixtures. Environ. Res. 1984, 35, 270–276. [Google Scholar] [CrossRef]
- Hall, J.G. The adaptation of enzymes to temperature: Catalytic characterization of glucosephosphate isomerase homologues isolated from Mytilus edulis and Isognomon alatus, bivalve molluscs inhabiting different thermal environments. Mol. Biol. Evol. 1985, 2, 251–269. [Google Scholar] [CrossRef][Green Version]
- Adler, E.; Knowles, J. A Thermolabile triosephosphate isomerase from the psychrophile Vibrio sp. strain ANT-300. Arch. Biochem. Biophys. 1995, 321, 137–139. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Ragan, M.A. Cloning and characterization of the nuclear gene and cDNAs for triosephosphate isomerase of the marine red alga Gracilaria verrucosa. Curr. Genet. 1995, 28, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Manchenko, G.P. Unusual isozyme patterns of glucose-6-phosphate isomerase in Polydora brevipalpa (Polychaeta: Spionidae). Biochem. Genet. 2001, 39, 285–288. [Google Scholar] [CrossRef] [PubMed]
- Iwahara, K.; Takahashi, R.; Naomi, T.; Kida, M.; Miyamoto, R.; Tokuyama, T. Purification and comparison of triosephosphate isomerases from ammonia-oxidizing bacteria isolated from terrestrial and marine environments. J. Biosci. Bioeng. 2001, 91, 603–606. [Google Scholar] [CrossRef]
- Goulard, F.; Pondaven, P.; Diouris, M.; Deslandes, E.; Floch, J.Y. Partial Purification and Characterization of UDP-Glucose-4-Epimerase from Solieria chordalis (Rhodophyceae). Bot. Mar. 2003, 46, 107–111. [Google Scholar] [CrossRef]
- Caponera, J.A.; Rawson, P.D. Highly divergent duplicate mannose-6-phosphate isomerase (Mpi) genes in the blue mussel, Mytilus edulis. Mar. Genom. 2008, 1, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Fitriani, D.; Saksono, B. Cloning of araA Gene Encoding L-Arabinose Isomerase from Marine Geobacillus stearothermophilus Isolated from Tanjung Api, Poso, Indonesia. HAYATI J. Biosci. 2010, 17, 58–62. [Google Scholar] [CrossRef]
- Schoville, S.D.; Flowers, J.M.; Burton, R.S. Diversifying Selection Underlies the Origin of Allozyme Polymorphism at the Phosphoglucose Isomerase Locus in Tigriopus californicus. PLoS ONE 2012, 7, e40035. [Google Scholar] [CrossRef][Green Version]
- Rodionova, I.A.; Scott, D.A.; Grishin, N.V.; Osterman, A.L.; Rodionov, D.A. Tagaturonate-fructuronate epimerase UxaE, a novel enzyme in the hexuronate catabolic network in Thermotoga maritima. Environ. Microbiol. 2012, 14, 2920–2934. [Google Scholar] [CrossRef]
- Chung, S.-K.; Ryu, S.-I.; Lee, S.-B. Characterization of UDP-glucose 4-epimerase from Pyrococcus horikoshii: Regeneration of UDP to produce UDP-galactose using two-enzyme system with trehalose. Bioresour. Technol. 2012, 110, 423–429. [Google Scholar] [CrossRef]
- Raedts, J.; Lundgren, M.; Kengen, S.W.M.; Li, J.-P.; Van Der Oost, J. A Novel Bacterial Enzyme with d-Glucuronyl C5-epimerase Activity. J. Biol. Chem. 2013, 288, 24332–24339. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Wang, X.; Zhang, Y.; Yu, J.; Liu, T.; Chen, S.; Chi, S. Origin and evolution of alginate-c5-mannuronan-epimerase gene based on transcriptomic analysis of brown algae. Acta Oceanol. Sin. 2014, 33, 73–85. [Google Scholar] [CrossRef]
- Yun, E.J.; Lee, S.; Kim, H.T.; Pelton, J.G.; Kim, S.; Ko, H.-J.; Choi, I.-G.; Kim, K.H. The novel catabolic pathway of 3,6-anhydro-L-galactose, the main component of red macroalgae, in a marine bacterium. Environ. Microbiol. 2014, 17, 1677–1688. [Google Scholar] [CrossRef] [PubMed]
- Akutsu, J.-I.; Zhang, Z.; Morita, R.; Kawarabayasi, Y. Identification and characterization of a thermostable bifunctional enzyme with phosphomannose isomerase and sugar-1-phosphate nucleotidylyltransferase activities from a hyperthermophilic archaeon, Pyrococcus horikoshii OT3. Extremophiles 2015, 19, 1077–1085. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Zavala, A.A.; Carrasco-Miranda, J.S.; Ramirez-Aguirre, C.D.; López-Hidalgo, M.; Benítez-Cardoza, C.G.; Ochoa-Leyva, A.; Cardona-Felix, C.S.; Diaz-Quezada, C.; Rudiño-Piñera, E.; Sotelo-Mundo, R.R.; et al. Structural insights from a novel invertebrate triosephosphate isomerase from Litopenaeus vannamei. Biochim. et Biophys. Acta (BBA) Bioenerg. 2016, 1864, 1696–1706. [Google Scholar] [CrossRef] [PubMed]
- Lajoie, C.A.; Kitner, J.B.; Potochnik, S.J.; Townsend, J.M.; Beatty, C.C.; Kelly, C.J. Cloning, expression and characterization of xylose isomerase from the marine bacterium Fulvimarina pelagiin in Escherichia coli. Biotechnol. Prog. 2016, 32, 1230–1237. [Google Scholar] [CrossRef]
- Fischl, R.; Bertelsen, K.; Gaillard, F.; Coelho, S.; Michel, G.; Klinger, M.; Boyen, C.; Czjzek, M.; Hervé, C. The cell-wall active mannuronan C5-epimerases in the model brown alga Ectocarpus: From gene context to recombinant protein. Glycobiology 2016, 26, 973–983. [Google Scholar] [CrossRef]
- Inoue, A.; Satoh, A.; Morishita, M.; Tokunaga, Y.; Miyakawa, T.; Tanokura, M.; Ojima, T. Functional heterologous expression and characterization of mannuronan C5-epimerase from the brown alga Saccharina japonica. Algal Res. 2016, 16, 282–291. [Google Scholar] [CrossRef]
- Lee, S.; Yun, E.J.; Kim, K.H.; Kim, H.-Y.; Choi, I.-G. 3,6-Anhydro-L-galactonate cycloisomerase from Vibrio sp. strain EJY3: Crystallization and X-ray crystallographic analysis. Acta Crystallogr. Sect. F Struct. Biol. Commun. 2017, 73, 511–514. [Google Scholar] [CrossRef]
- Yang, Y.; Chen, Z.-W.; Hurlburt, B.K.; Li, G.-L.; Zhang, Y.-X.; Fei, D.-X.; Shen, H.-W.; Cao, M.-J.; Liu, G.-M. Identification of triosephosphate isomerase as a novel allergen in Octopus fangsiao. Mol. Immunol. 2017, 85, 35–46. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, Y.-X.; Liu, M.; Maleki, S.J.; Zhang, M.-L.; Liu, Q.-M.; Cao, M.-J.; Su, W.-J.; Liu, G.-M. Triosephosphate isomerase and filamin C share common epitopes as novel allergens of Procambarus clarkii. J. Agric. Food Chem. 2017, 65, 950–963. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Chen, Q.; Song, Y.; Huang, H.; Li, J.; Ma, J.; Li, Q.; Ju, J. Deciphering the sugar biosynthetic pathway and tailoring steps of nucleoside antibiotic A201A biosynthesis unveils GDP-l-galactose mutase. Proc. Natl. Acad. Sci. USA 2017, 114, 4948–4953. [Google Scholar] [CrossRef] [PubMed]
- Merkx-Jacques, A.; Rasmussen, H.; Muise, D.M.; Benjamin, J.J.R.; Kottwitz, H.; Tanner, K.; Milway, M.T.; Purdue, L.M.; Scaife, M.A.; Armenta, R.E.; et al. Engineering xylose metabolism in thraustochytrid T18. Biotechnol. Biofuels 2018, 11, 248. [Google Scholar] [CrossRef] [PubMed]
- Xia, F.; Li, M.-S.; Liu, Q.-M.; Liu, M.; Yang, Y.; Cao, M.-J.; Chen, G.-X.; Jin, T.; Liu, G.-M. Crystal Structure Analysis and Conformational Epitope Mutation of Triosephosphate Isomerase, a Mud Crab Allergen. J. Agric. Food Chem. 2019, 67, 12918–12926. [Google Scholar] [CrossRef]
- Hu, Y.; Du, Q.; Mi, P.; Shang, E.; Sui, Z. Gene cloning and expression regulation in the pathway of agar and floridean starch synthesis of Gracilariopsis lemaneiformis (Rhodophyta). Environ. Boil. Fishes 2019, 31, 1889–1896. [Google Scholar] [CrossRef]
- Rengasamy, S.; Subramanian, M.R.; Perumal, V.; Ganeshan, S.; Al Khulaifi, M.M.; Al-Shwaiman, H.A.; Elgorban, A.M.; Syed, A.; Thangaprakasam, U. Purification and kinetic behavior of glucose isomerase from Streptomyces lividans RSU26. Saudi J. Biol. Sci. 2020, 27, 1117–1123. [Google Scholar] [CrossRef]
- Sakai, N.; Tanaka, M.; Takahashi, M.; Adachi, S.; Nagahama, Y. Isolation and expression of rainbow trout (Oncorhynchus mykiss) ovarian cDNA encoding 3β-hydroxysteroid dehydrogenase/Δ5−4-isomerase. Fish Physiol. Biochem. 1993, 11, 273–279. [Google Scholar] [CrossRef]
- Misawa, N.; Shimada, H. Metabolic engineering for the production of carotenoids in non-carotenogenic bacteria and yeasts. J. Biotechnol. 1998, 59, 169–181. [Google Scholar] [CrossRef]
- Wise, M.L.; Rossi, J.; Gerwick, W.H. Characterization of the substrate binding site of polyenoic fatty acid isomerase, a novel enzyme from the marine alga Ptilota filicina. Biochemistry 1997, 36, 2985–2992. [Google Scholar] [CrossRef]
- Wang, C.W.; Oh, M.K.; Liao, J.C. Engineered isoprenoid pathway enhances astaxanthin production in Escherichia coli. Biotechnol. Bioeng. 1999, 62, 235–241. [Google Scholar] [CrossRef]
- Huang, J.; Jiang, X.; Zhang, X.; Chen, W.; Tian, B.; Shu, Z.; Hu, S. Expressed sequence tag analysis of marine fungus Schizochytrium producing docosahexaenoic acid. J. Biotechnol. 2008, 138, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Leblond, J.D.; Dodson, J.; Khadka, M.; Holder, S.; Seipelt, R.L. Sterol composition and biosynthetic genes of the recently discovered photosynthetic alveolate, Chromera velia (chromerida), a close relative of apicomplexans. J. Eukaryot. Microbiol. 2012, 59, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Liu, M.; He, M.; Ye, Y.; Huang, J. Illustrating and Enhancing the Biosynthesis of Astaxanthin and Docosahexaenoic Acid in Aurantiochytrium sp. SK4. Mar. Drugs 2019, 17, 45. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.-L.; Zheng, J.-L.; Liu, J. Transcription activation of β-carotene biosynthetic genes at the initial stage of stresses as an indicator of the increased β-carotene accumulation in isolated Dunaliella salina strain GY-H13. Aquat. Toxicol. 2020, 222, 105472. [Google Scholar] [CrossRef] [PubMed]
- Omura, Y.; Hayashi, Y.; Matsushima, O.; Katayama, H.; Yamada, K. Partial purification and characterization of alanine racemase from the brackish-water bivalve Corbicula japonica. J. Exp. Mar. Biol. Ecol. 1985, 94, 281–289. [Google Scholar] [CrossRef]
- Yamada, A.; Matsushima, O. The relation of d-alanine and alanine racemase activity in molluscs. Comp. Biochem. Physiol. Part B 1992, 103, 617–621. [Google Scholar] [CrossRef]
- Oren, A.; Gurevich, P. Diversity of lactate metabolism in halophilic archaea. Can. J. Microbiol. 1995, 41, 302–307. [Google Scholar] [CrossRef]
- Iida, T.; Furutani, M.; Iwabuchi, T.; Maruyama, T. Gene for a cyclophilin-type peptidyl–prolyl cis–trans isomerase from a halophilic archaeum, Halobacterium cutirubrum. Gene 1997, 204, 139–144. [Google Scholar] [CrossRef]
- Shibata, K.; Shirasuna, K.; Motegi, K.; Kera, Y.; Abe, H.; Yamada, R.-H. Purification and properties of alanine racemase from crayfish Procambarus clarkii. Comp. Biochem. Physiol. Part B: Biochem. Mol. Biol. 2000, 126, 599–608. [Google Scholar] [CrossRef]
- Yokoyama, T.; Tanaka, Y.; Sato, M.; Kan-No, N.; Nakano, T.; Yamaguchi, T.; Nagahisa, E. Alanine racemase activity in the microalga Thalassiosira sp. Fish. Sci. 2005, 71, 924–930. [Google Scholar] [CrossRef]
- Yokoyama, T.; Mimura, Y.; Sato, M.; Nagahisa, E. Purification and some properties of alanine racemase from the marine gastropod Cellana grata. Fish. Sci. 2006, 72, 195–201. [Google Scholar] [CrossRef]
- Safavi-Hemami, H.; Siero, W.A.; Gorasia, D.G.; Young, N.D.; Macmillan, D.; Williamson, N.A.; Purcell, A.W. Specialisation of the Venom Gland Proteome in Predatory Cone Snails Reveals Functional Diversification of the Conotoxin Biosynthetic Pathway. J. Proteome Res. 2011, 10, 3904–3919. [Google Scholar] [CrossRef]
- Sha, Z.-X.; Liu, H.; Wang, Q.-L.; Liu, Y.; Lu, Y.; Li, M.; Chen, S.-L. Channel catfish (Ictalurus punctatus) protein disulphide isomerase, PDIA6: Molecular characterization and expression regulated by bacteria and virus inoculation. Fish Shellfish. Immunol. 2012, 33, 220–228. [Google Scholar] [CrossRef] [PubMed]
- Li, L.-C.; Hsu, Y.-T.; Chang, H.-L.; Wu, T.-M.; Sung, M.-S.; Cho, C.-L.; Lee, T.-M. Polyamine effects on protein disulfide isomerase expression and implications for hypersalinity stress in the marine algaUlva lactucaLinnaeus1. J. Phycol. 2013, 49, 1181–1191. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Zhang, D.; Wang, L.; Gai, Y.; Zhou, Z.; Zhang, H.; Song, L. The molecular characterization of a cyclophilin A from Chinese mitten crab Eriocheir sinensis and the antifungal activity of its recombinant protein. Electron. J. Biotechnol. 2013, 16. [Google Scholar] [CrossRef]
- Kim, S.-H.; Lee, J.M.; Kang, D.S.; Kim, D.-G.; Ahn, S.-H.; Kong, I.-S. Expression, purification and characterization of soluble recombinant peptidyl-prolyl cis/trans isomerase from Vibrio anguillarum. Protein Expr. Purif. 2014, 101, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Tang, W.; Wang, X.; Chen, Y.; Wu, Y.; Qiang, Y.; Feng, Y.; Ren, Z.; Chen, S.; Xu, A. PPIase is associated with the diversity of conotoxins from cone snail venom glands. Biochimie 2015, 112, 129–138. [Google Scholar] [CrossRef]
- Jakob, R.P.; Schmidpeter, P.A.M.; Koch, J.R.; Schmid, F.X.; Maier, T. Structural and Functional Characterization of a Novel Family of Cyclophilins, the AquaCyps. PLoS ONE 2016, 11, e0157070. [Google Scholar] [CrossRef]
- Ekubota, T.; Shimamura, S.; Kobayashi, T.; Nunoura, T.; Deguchi, S. Distribution of eukaryotic serine racemases in the bacterial domain and characterization of a representative protein in Roseobacter litoralis Och 149. Microbiology 2016, 162, 53–61. [Google Scholar] [CrossRef]
- Subin, C.S.; Pradeep, M.A.; Vijayan, K.K. FKBP-type peptidyl-prolyl cis-trans isomerase from thermophilic microalga, Scenedesmus sp.: Molecular characterisation and demonstration of acquired salinity and thermotolerance in E. coli by recombinant expression. Environ. Boil. Fishes 2016, 28, 3307–3315. [Google Scholar] [CrossRef]
- Hoppstock, L.; Trusch, F.; Lederer, C.; Van West, P.; Könneke, M.; Bayer, P. NmPin from the marine thaumarchaeote Nitrosopumilus maritimus is an active membrane associated prolyl isomerase. BMC Biol. 2016, 14, 53. [Google Scholar] [CrossRef] [PubMed]
- Safavi-Hemami, H.; Li, Q.; Jackson, R.L.; Song, A.S.; Boomsma, W.; Bandyopadhyay, P.K.; Gruber, C.W.; Purcell, A.W.; Yandell, M.; Olivera, B.M.; et al. Rapid expansion of the protein disulfide isomerase gene family facilitates the folding of venom peptides. Proc. Natl. Acad. Sci. USA 2016, 113, 3227–3232. [Google Scholar] [CrossRef] [PubMed]
- Figueroa-Montiel, A.; Ramos, M.A.; Mares, R.E.; Dueñas, S.; Pimienta, G.; Ortiz, E.; Possani, L.D.; Licea-Navarro, A.F. In Silico Identification of Protein Disulfide Isomerase Gene Families in the De Novo Assembled Transcriptomes of Four Different Species of the Genus Conus. PLoS ONE 2016, 11, e0148390. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Xie, J.; Yang, S.; Ye, S.; Luo, M.; Wu, X. First characterization of three cyclophilin family proteins in the oyster, Crassostrea ariakensis Gould. Fish Shellfish. Immunol. 2016, 55, 257–266. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, X.; Ren, Z.; Tang, W.; Zou, Q.; Wang, J.; Chen, S.; Zhang, H.; Xu, A. Oxidative Folding of Conopeptides Modified by Conus Protein Disulfide Isomerase. Protein J. 2017, 36, 407–416. [Google Scholar] [CrossRef]
- Muhammad, F.; Zhi-Feng, Z.; Ming-Yu, S.; Shafi, M. cDNA Cloning and expression of cyclophilin A (LvCypA) in white leg shrimp, Litopenaeus vannamei. Pak. J. Zoöl. 2017, 49, 935–941. [Google Scholar] [CrossRef]
- Lee, H.; Kim, S.H.; Han, Y.-J.; Im, S. PsCYP1 of marine red alga Pyropia seriata (Bangiales, Rhodophyta) confers salt and heat tolerance in Chlamydomonas. J. Appl. Phycol. 2017, 29, 617–625. [Google Scholar] [CrossRef]
- Morinaka, B.I.; Vagstad, A.L.; Piel, J. Radical S-Adenosylmethionine Peptide Epimerases: Detection of Activity and Characterization of d-Amino Acid Products. Methods Enzymol. 2018, 604, 237–257. [Google Scholar] [CrossRef]
- Qiu, X.; Zhu, W.; Wang, W.; Jin, H.; Zhu, P.; Zhuang, R.; Yan, X. Structural and functional insights into the role of a cupin superfamily isomerase in the biosynthesis of Choi moiety of aeruginosin. J. Struct. Biol. 2019, 205, 44–52. [Google Scholar] [CrossRef]
- Shang, Z.; Winter, J.M.; Kauffman, C.A.; Yang, I.; Fenical, W. Salinipeptins: Integrated genomic and chemical approaches reveal unusual D-amino acid-containing ribosomally synthesized and post-translationally modified peptides (RiPPs) from a Great Salt Lake Streptomyces sp. ACS Chem. Biol. 2019, 14, 415–425. [Google Scholar] [CrossRef]
- Company, R.; Antúnez, O.; Cosson, R.P.; Serafim, A.; Shillito, B.; Cajaraville, M.; Bebianno, M.J.; Torreblanca, A. Protein expression profiles in Bathymodiolus azoricus exposed to cadmium. Ecotoxicol. Environ. Saf. 2019, 171, 621–630. [Google Scholar] [CrossRef] [PubMed]
- Ulagesan, S.; Choi, J.-W.; Nam, T.-J.; Choi, Y.H. Peptidyl-prolyl isomerase and the biological activities of recombinant protein cyclophilin from Pyropia yezoensis (PyCyp). Protein Expr. Purif. 2020, 172, 105636. [Google Scholar] [CrossRef] [PubMed]
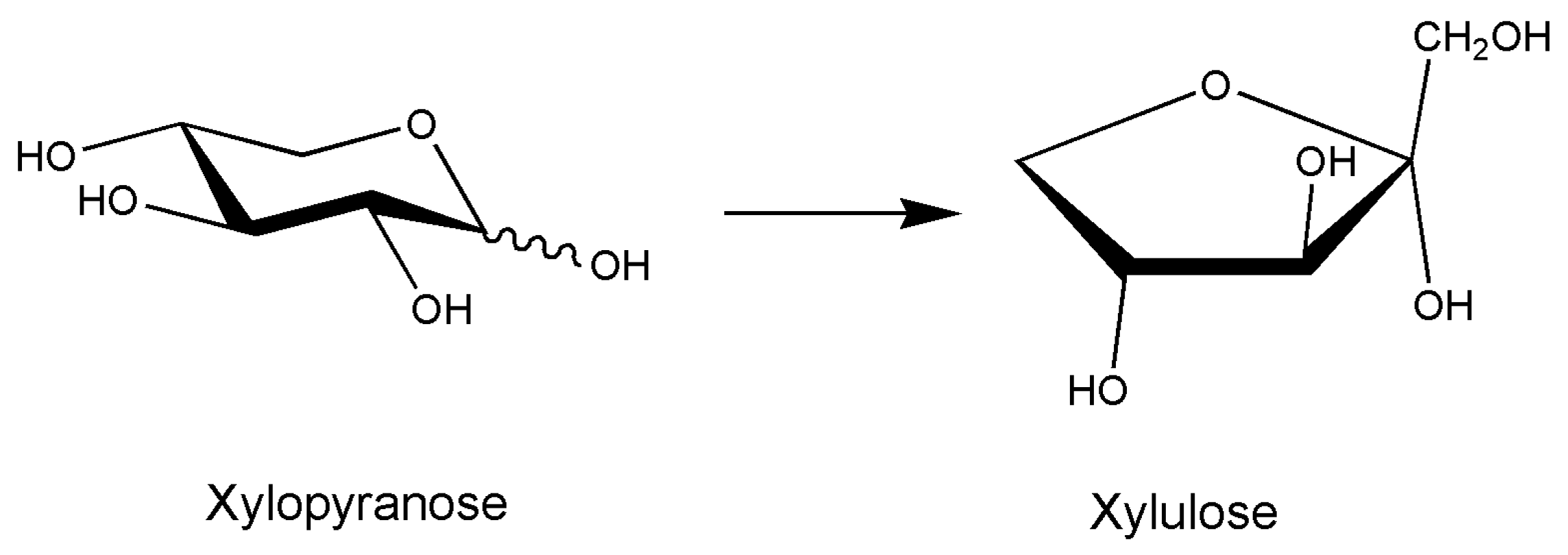
- Umemoto, Y.; Shibata, T.; Araki, T. d-Xylose Isomerase from a Marine Bacterium, Vibrio sp. Strain XY-214, and d-Xylulose Production from β-1,3-Xylan. Mar. Biotechnol. 2011, 14, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Wang, Z.-P.; Sheng, J.; Zheng, Y.; Ji, X.-F.; Zhou, H.-X.; Liu, X.-Y.; Chi, Z.-M. High and efficient isomaltulose production using an engineered Yarrowia lipolytica strain. Bioresour. Technol. 2018, 265, 577–580. [Google Scholar] [CrossRef]
- Mu, W.; Li, W.; Wang, X.; Zhang, T.; Jiang, B. Current studies on sucrose isomerase and biological isomaltulose production using sucrose isomerase. Appl. Microbiol. Biotechnol. 2014, 98, 6569–6582. [Google Scholar] [CrossRef]
- Jeong, D.W.; Hyeon, J.E.; Shin, S.K.; Han, S.O. Trienzymatic Complex System for Isomerization of Agar-Derived d-Galactose into d-Tagatose as a Low-Calorie Sweetener. J. Agric. Food Chem. 2020, 68, 3195–3202. [Google Scholar] [CrossRef]
- Wise, M.L.; Hamberg, M.; Gerwick, W.H. Biosynthesis of conjugated triene-containing fatty acids by a novel isomerase from the red marine alga Ptilota filicina. Biochemistry 1994, 33, 15223–15232. [Google Scholar] [CrossRef]
- Wise, M.L.; Söderström, K.; Murray, T.F.; Gerwick, W.H. Synthesis and cannabinoid receptor binding activity of conjugated triene anandamide, a novel eicosanoid. Cell. Mol. Life Sci. 1996, 52, 88–92. [Google Scholar] [CrossRef]
- Gerwick, W.H.; Moghaddam, M.; Hamberg, M. Oxylipin metabolism in the red alga Gracilariopsis lemaneiformis: Mechanism of formation of vicinal dihydroxy fatty acids. Arch. Biochem. Biophys. 1991, 290, 436–444. [Google Scholar] [CrossRef]
- Ghioni, C.; Porter, A.E.A.; Sadler, I.H.; Tocher, D.R.; Sargent, J.R. Cultured fish cells metabolize octadecapentaenoic acid (all-cis Δ3,6,9,12,15–18:5) to octadecatetraenoic acid (all-cis Δ 6,9,12,15–18:4) via its 2-trans intermediate (trans Δ2, all-cis δ6,9,12,15–18:5). Lipids 2001, 36, 145–153. [Google Scholar] [CrossRef]
- Rontani, J.-F.; Bonin, P.; Jameson, I.; Volkman, J.K. Degradation of alkenones and related compounds during oxic and anoxic incubation of the marine haptophyte Emiliania huxleyi with bacterial consortia isolated from microbial mats from the Camargue, France. Org. Geochem. 2005, 36, 603–618. [Google Scholar] [CrossRef]
- Nagashima, H.; Zulkharnain, A.; Maeda, R.; Fuse, H.; Iwata, K.; Omori, T. Cloning and nucleotide sequences of carbazole degradation genes from marine bacterium Neptuniibacter sp. strain CAR-SF. Curr. Microbiol. 2009, 61, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Glueck, S.; Gadler, P.; Kroutil, W.; Nestl, B.M.; Larissegger-Schnell, B.; Ueberbacher, B.; Wallner, S.; Faber, K. Biocatalytic approaches for the quantitative production of single stereoisomers from racemates. Biochem. Soc. Trans. 2006, 34, 296–300. [Google Scholar] [CrossRef] [PubMed]
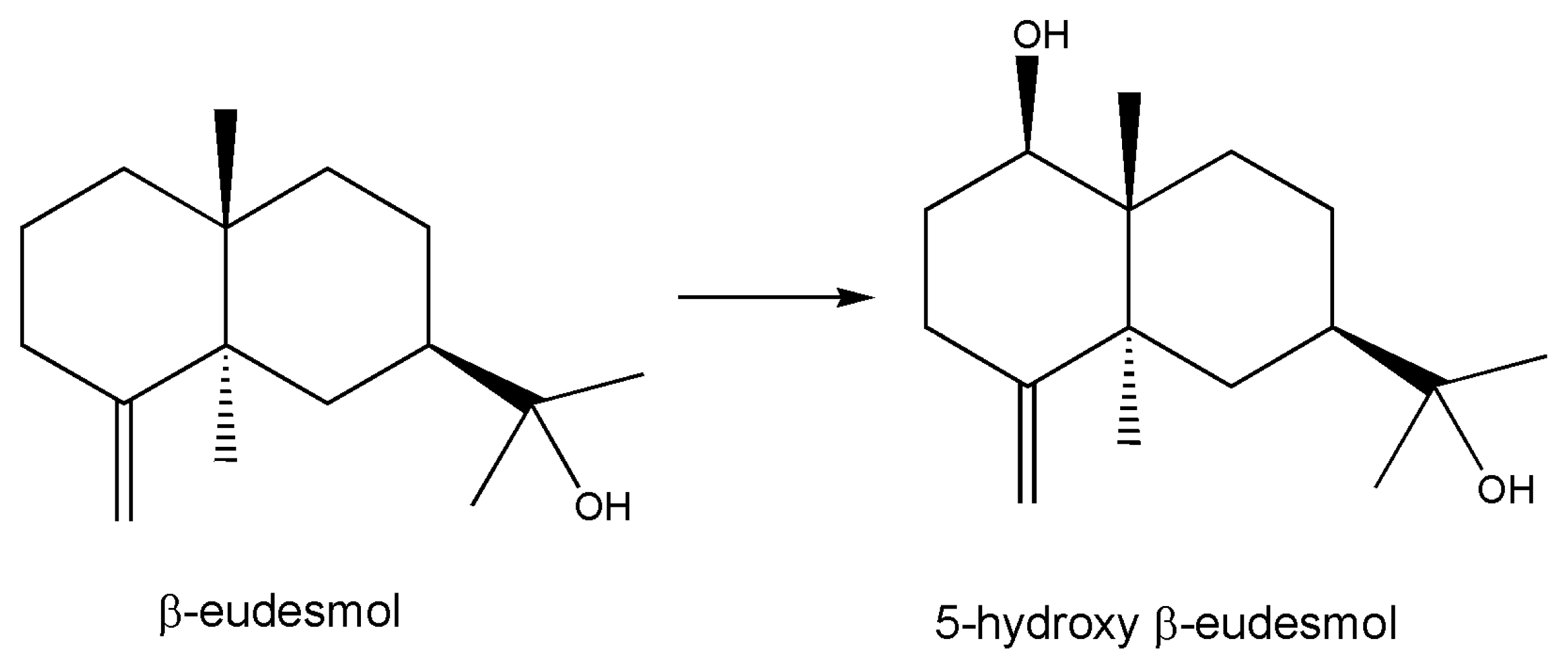
- Misawa, N.; Nodate, M.; Otomatsu, T.; Shimizu, K.; Kaido, C.; Kikuta, M.; Ideno, A.; Ikenaga, H.; Ogawa, J.; Shimizu, S.; et al. Bioconversion of substituted naphthalenes and β-eudesmol with the cytochrome P450 BM3 variant F87V. Appl. Microbiol. Biotechnol. 2010, 90, 147–157. [Google Scholar] [CrossRef]
- Jung, B.-O.; Roseman, S.; Park, J.K. The central concept for chitin catabolic cascade in marine bacterium, Vibrios. Macromol. Res. 2008, 16, 1–5. [Google Scholar] [CrossRef]
- Rosani, U.; Domeneghetti, S.; Maso, L.; Wegner, K.M.; Venier, P. An Evolutionary Perspective of Dopachrome Tautomerase Enzymes in Metazoans. Genes 2019, 10, 495. [Google Scholar] [CrossRef]
- Waisiel, A.A.; Rozeboom, H.J.; Hauke, D.; Baas, B.-J.; Zandvoort, E.; Quax, W.J.; Thunnissen, A.-M.W.H.; Poelarends, G.J. Structural and functional characterization of macrophage migration inhibiotry factor homologue from the marine cyanobacterium Prochlorocuccus marinus. Biochemitry 2010, 49, 7572–7581. [Google Scholar] [CrossRef][Green Version]
- Zuasti, A.; Martínez-Liarte, J.H.; Solano, F.; Ferrer, C. Melanization stimulating factors in the integument of the Mugil cephalus and Dicertranchus labrax. Histol. Histopathol. 2000, 15, 1145–1150. [Google Scholar]
- Xu, F.; Li, M.-Y.; Chen, J.; Feng, X.; Ming-Yun, L.; Chen, J. D-dopachrome tautomerase from Japanese sea bass (Lateolabrax japonicus) is a chemokine-like cytokine and functional homolog of macrophage migration inhibitory factor. Zool Res. 2020, 41, 39–50. [Google Scholar] [CrossRef]
- Wang, D.; Yang, D.; Wang, Q.; Zhao, Y.; Li, C.; Wei, Q.; Han, Y.; Zhao, J. Two macrophage migration inhibitory factors (MIFs) from the clam Ruditapes philippinarum: Molecular characterization, localization and enzymatic activities. Fish Shellfish. Immunol. 2018, 78, 158–168. [Google Scholar] [CrossRef]
- Trincone, A. Enzymatic Processes in Marine Biotechnology. Mar. Drugs 2017, 15, 93. [Google Scholar] [CrossRef] [PubMed]
| Databases | Search Statement | Hits |
|---|---|---|
| Science Direct | Isomerase * and marine in titles, abstracts and keywords | 31 |
| PubMed | Isomerase * and marine | 172 |
| WoS 1 | Marine epimerase * in All fields | 53 |
| WoS | Marine racemase * in All fields | 44 |
| WoS | Marine cis–trans isomerase * in All fields | 56 |
| WoS | Marine cycloisomerase * in All fields | 3 |
| WoS | Marine tautomerase * in All fields | 26 |
| WoS | Marine mutase * in All fields | 45 |
| Reference/Year | Organism | Enzyme | Reaction | Note |
|---|---|---|---|---|
| [9] 1973 | Alga Pelvetia canaliculata | Polymannuronic-5-epimerase | Conversion of polymannuronic acid to a mixed polymer containing guluronic acid | Preparation of ammonium sulfate fraction of the enzyme |
| [10] 1977 | Marine species of Alcaligenes, Pseudomonas marina, and Alteromonas communis | P-hexose isomerase | Glycolytic pathway | Entner–Doudoroff pathway |
| [11] 1984 | Marine snails | Phosphoglucose isomerase | Glycolytic enzyme | Tolerance to detergents as monitoring tool |
| [12] 1985 | Bivalve mollusks: Mytilus edulis and Isognomon alatus | Glucose phosphate isomerase | Glycolytic enzyme | Biochemical-based study of adaptation of enzyme to temperature |
| [13] 1995 | Psychrophilic marine eubacterium Vibrio sp. strain ANT-300 | Triosephosphate isomerase | Interconversion dihydroxyacetone phosphate and d-glyceraldehyde-3-phosphate | Thermolability study |
| [14] 1995 | Marine red alga Gracilaria verrucosa | Triosephosphate isomerase | Interconversion dihydroxyacetone phosphate and d-glyceraldehyde-3-phosphate | Genetic study |
| [15] 2001 | Polychaeta Polydora brevipalpa | Glucose-6-phosphate isomerase | Glycolytic enzyme | Study of isozyme pattern |
| [16] 2001 | Marine ammonia-oxidizing bacteria Nitrosomonas | Triosephosphate isomerases | Interconversion dihydroxyacetone phosphate and d-glyceraldehyde-3-phosphate | Purification and characterization |
| [17] 2003 | Macroalga Solieria chordalis | UDP-glucose-4-epimerase | Catalyzing both the synthesis of UDP-Gal and UDP-Glc | Characterization of the enzyme |
| [18] 2008 | Blue mussel Mytilus edulis | Mannose-6-phosphate isomerase | Glycolytic enzyme | Genetic study |
| [19] 2010 | Marine Geobacillus stearothermophilus | l-Arabinose Isomerase | Converting d-galactose to d-tagatose | Clone and sequence araA gene |
| [20] 2012 | Marine copepod Tigriopus californicus | Phosphoglucose isomerase | Glycolytic enzyme | Genetic variability study |
| [21] 2012 | Thermotoga maritima | Tagaturonate-fructuronate epimerase UxaE | Epimerization of tagaturonate to fructuronate | Study of metabolism of galacturonate and glucuronate from pectin and xylan |
| [22] 2012 | Pyrococcus horikoshii | UDP-glucose 4-epimerase | Catalyzing both the synthesis of UDP-Gal and UDP-Glc | Characterization study of the enzyme that could be coupled with trehalose synthase |
| [23] 2013 | Marine bacterium Bermanella marisrubri sp. RED65 | d-glucuronyl C5-epimerase | Epimerization of d-glucuronic acid to its C5-epimer l-iduronic acid | Recombinant protein expressed in Escherichia coli showed epimerization activity |
| [24] 2014 | Brown algae | Alginate-C5-mannuronan-epimerase | Catalyze the conversion of mannuronate to guluronate and determine the M/G ratio of alginate | Genetic study: predicted 94 algG genes open reading frame (ORF) sequences of brown algae |
| [25] 2015 | Marine bacterium Vibrio sp. | 3,6-Anhydro-l-galactonate cycloisomerase | Converts 3,6-anhydro-l-galactonate into 2-keto-3-deoxygalactonate | Identification of intermediate products of 3,6-anhydro-l-galactose |
| [26] 2015 | Pyrococcus horikoshii | Phosphomannose isomerase | Mannosylglycerate biosynthetic pathway | Recombinant protein expressed in E. coli with double activity (Man-1-P GTase activity) |
| [27] 2016 | Marine Pacific whiteleg shrimp Litopenaeus vannamei | Triosephosphate isomerase | Interconversion dihydroxyacetone phosphate and d-glyceraldehyde-3-phosphate | Structural and mechanistic study and insights into glycolysis regulation in crustaceans |
| [28] 2016 | Marine bacterium Fulvimarina pelagi | Xylose isomerase | Interconversion of d-xylose and d-xylulose | Cloning, expression, and characterization for use in biofuels’ production |
| [29] 2016 | Brown alga Ectocarpus | Mannuronan C5-epimerase | Control the distribution pattern of (1-4) linked β-d-mannuronic acid (M) and alpha-l-guluronic acid (G) residues in alginates | Transcript expression |
| [30] 2016 | Alga Saccharina japonica | Mannuronan C5-epimerase | Control of the distribution pattern of (1-4) linked β-d-mannuronic acid (M) and alpha-l-guluronic acid (G) residues in alginates | Functional recombinant expression of protein in insect-cell system revealing alternate epimerization of beta-d-mannuronic acid to alpha-l-guluronic acid |
| [31] 2017 | Vibrio sp. strain EJY3 | 3,6-Anhydro-l-galactonate cycloisomerase | Converts 3,6-anhydro-l-galactonate into 2-keto-3-deoxygalactonate | Crystallization and X-ray analysis of recombinant protein |
| [32] 2017 | Octopus fangsiao | Triosephosphate isomerase | Interconversion dihydroxyacetone phosphate and d-glyceraldehyde-3-phosphate | Study of allergen function |
| [33] 2017 | Freshwater crayfish Procambarus clarkii | Triosephosphate isomerase | Interconversion dihydroxyacetone phosphate and d-glyceraldehyde-3-phosphate | Study of allergen function |
| [34] 2017 | Marinactinospora thermotolerans | GDP-l-galactose mutase | Conversion of pyranose form to furanose structure | Study of the sugar biosynthetic pathway |
| [35] 2018 | Marine fungus-like thraustochytrids | Xylose isomerase | Interconversion of d-xylose and d-xylulose | Identification and characterization of xylose metabolism |
| [36] 2019 | Scylla paramamosain | Triosephosphate isomerase | Interconversion dihydroxyacetone phosphate and d-glyceraldehyde-3-phosphate | Crystal structure |
| [37] 2019 | Gracilariopsis lemaneiformis | Mannose-6-phosphate isomerase, GDP-mannose-3,5-epimerase | Pathways of floridean starch | Transcriptomic study for the study of the mechanism of substrate competition of synthesis pathways of floridean starch |
| [38] 2020 | Marine Streptomyces lividans RSU26 | Glucose isomerase | Fructose to glucose conversion | Characterization study and optimization of enzyme production |
| Reference/Year | Organism | Enzyme | Reaction | Note |
|---|---|---|---|---|
| [39] 1993 | Rainbow trout Oncorhynchus mykiss | 3β-hydroxysteroid dehydrogenase/Δ(5-4)-isomerase | Steroidogenic enzymes involved in the production of 17α-hydroxyprogesterone | Genetic study |
| [40] 1997 | Marine bacterium Agrobacterium aurantiacum | Carotenoid gene cluster | β-carotene biosynthesis | Metabolic engineering study |
| [41] 1997 | Alga Ptilota filicina | Polyenoic fatty acid isomerase | Assay by conversion of arachidonic acid to a conjugated triene | Biochemical study of binding site characteristics |
| [42] 1999 | Marine bacterium Agrobacterium aurantiacum | Isopentenyl diphosphate (IPP) isomerase and gene cluster (crtBIYZW) | Isoprenoid pathway | Study to enhance astaxanthin production by engineering isoprenoid pathway |
| [3] 2002 | Marine alga Ptilota filicina | Polyenoic fatty acid isomerase | Assay by conversion of arachidonic acid to a conjugated triene | Study of protein characterization and functional expression |
| [43] 2008 | Marine fungus Schizochytrium | Enzymes involved in biosynthesis of fatty acid via polyketide synthases | Confirmation PKS pathway | Genetic study of docohexanoic acid biosynthesis |
| [44] 2012 | Marine alveolate Chromera velia | Isopentenyl diphosphate Δ-isomerase | Sterol biosynthesis | Study of sterol composition of Chromera velia for chemotaxonomic relationships |
| [45] 2019 | Marine thraustochytrid Aurantiochytrium | Isopentenyl pyrophosphate isomerase | Biosynthetic pathways of docosahexaenoic acid (DHA) and ketocarotenoid astaxanthin | Analyses of the genome, transcriptome, key enzymes, and pathway products |
| [46] 2020 | Dunaliella salina | 15-cis-Z-carotene isomerase, prolycopene isomerase | β-carotene biosynthesis | Study of β-carotene biosynthesis: seven full length cDNA sequences cloned |
| Reference/Year | Organism | Enzyme | Reaction | Note |
|---|---|---|---|---|
| [47] 1985 | Bivalve Corbicula japonica | Alanine racemase | l to d alanine | Partial purification and characterization |
| [48] 1992 | Eighteen molluscan species | Alanine racemase | l to d alanine | Comparative study and distribution |
| [49] 1995 | Haloferax volcanii and Haloarcula species | Lactate racemase | l to d lactate | Study of enzymatic diversity among species |
| [50] 1997 | Halobacterium cutirubrum | Peptidyl-prolyl cis/trans isomerase | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Genetic study and expression in E. coli |
| [51] 2000 | Crayfish Procambarus clarkii | Alanine racemase | l to d alanine | Isolation, kinetic properties, substrate specificity, structural characteristics |
| [2] 2001 | Black-tiger prawn, Penaeus monodon | Alanine racemase | l to d alanine | Kinetic properties and substrate specificity |
| [52] 2005 | Microalga Thalassiosira sp. | Alanine racemase | l to d alanine | Kinetic properties and substrate specificity |
| [53] 2006 | Marine gastropod Cellana grata | Alanine racemase | l to d alanine | First purification study and kinetic assessment in gastropod |
| [54] 2011 | Marine cone snails | Disulfide isomerase | Oxidation, isomerization, and reduction of S–S bonds | Proteomic study showing presence of multitude of isoform of the enzyme |
| [55] 2012 | Channel catfish Ictalurus punctatus | Disulfide isomerase | Oxidation, isomerization and reduction of S–S bonds | Genetic study |
| [56] 2013 | Marine alga Ulva lactuca | Disulfide isomerase | Oxidation, Isomerization, and reduction of S–S bonds | Study of cloning and expression |
| [57] 2013 | Crab Eriocheir sinensis | Peptidyl-prolyl cis/trans isomerase | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Purification of recombinant protein and assessment of antifungal properties |
| [58] 2014 | Marine bacterium Vibrio anguillarum | Peptidyl-prolyl cis/trans isomerase | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Changes in protein expression of V. anguillarum, gene expression in E. coli and biochemical characterization |
| [59] 2015 | Core snails | Disulfide isomerase | Oxidation, Isomerization, and reduction of S–S bonds | Proteomic study |
| [60] 2016 | Marine Alphaproteobacteria | A novel family of peptidyl-prolyl isomerase | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Structural and functional characterization |
| [61] 2016 | Marine heterotrophic bacterium Roseobacter litoralis | Serine racemase | Racemization and minor dehydration of serine | Genomic analysis |
| [62] 2016 | Thermophilic chlorophycean microalga, Scenedesmus sp. | Peptidyl-prolyl cis/trans isomerase | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Cloning and expression of the enzyme in E. coli and indication of role in stress-tolerance mechanisms |
| [63] 2016 | Marine thaumarchaeote Nitrosopumilus maritimus | Peptidyl-prolyl cis/trans isomerase | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | A protein structure study |
| [64] 2016 | Superfamily Conoidea | Disulfide isomerase | Oxidation, Isomerization, and reduction of S–S bonds | Study of diversification of enzymatic protein folding correlated with diversity of conotoxins |
| [65] 2016 | Marine snails belonging to Conus | Disulfide isomerase | Oxidation, Isomerization, and reduction of S–S bonds | Transcriptomic and in silico analysis and characterization of the group of PDI protein sequences |
| [66] 2016 | Oyster Crassostrea ariakensis Gould | Peptidyl-prolyl isomerase (cyclophilins) | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Enzymatic tissue distribution and role of the three enzymes identified and involvement in in oyster immune response |
| [67] 2017 | Cone snail species | Disulfide isomerase | Oxidation, Isomerization, and reduction of S–S bonds | Cloned 12 disulfide isomerase genes and study of reaction on conopeptides |
| [68] 2017 | Shrimp, Litopenaeus vannamei | Peptidyl-prolyl isomerase (cyclophilins) | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Cloning and tissue distribution of the enzyme |
| [69] 2017 | Red alga Pyropia seriata | Peptidyl-prolyl isomerase (cyclophilins) | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Transcriptomic study |
| [70] 2018 | Cyanobacterial genomes | PoyD, a member of the radical S-adenosylmethionine superfamily | Introducing d-amino acids into a ribosomally synthesized peptide | Heterologous expression in E. coli, detection of epimerase activity, and localization of epimerization sites |
| [71] 2019 | Cyanobacteria | AerE, a cupin superfamily enzyme | 1,3-allylic isomerization | Study of the biosynthesis of aeruginosins trapeptides possessing antithrombotic activity |
| [72] 2019 | Halotolerant Streptomyces sp. strain GSL-6C | Inferring new epimerases | Conversion of l- to d-amino acids | Genome analysis integrating a study on salinipeptins |
| [73] 2019 | Hydrothermal vent mussel Bathymodiolus azoricus | Peptidyl-prolyl cis/trans isomerase | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Study of proteome changes upon Cd exposure for bioindicator identification |
| [74] 2020 | Marine red algae Pyropia yezoensis | Peptidyl-prolyl isomerase (cyclophilins) | Isomerization of peptide bonds (trans-cis) at Pro residues; facilitates protein folding | Analysis of the biological activity of recombinant cyclophilin |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trincone, A. Application-Oriented Marine Isomerases in Biocatalysis. Mar. Drugs 2020, 18, 580. https://doi.org/10.3390/md18110580
Trincone A. Application-Oriented Marine Isomerases in Biocatalysis. Marine Drugs. 2020; 18(11):580. https://doi.org/10.3390/md18110580
Chicago/Turabian StyleTrincone, Antonio. 2020. "Application-Oriented Marine Isomerases in Biocatalysis" Marine Drugs 18, no. 11: 580. https://doi.org/10.3390/md18110580
APA StyleTrincone, A. (2020). Application-Oriented Marine Isomerases in Biocatalysis. Marine Drugs, 18(11), 580. https://doi.org/10.3390/md18110580





