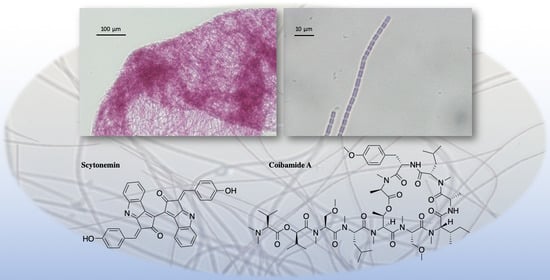
The Chemistry, Biochemistry and Pharmacology of Marine Natural Products from Leptolyngbya, a Chemically Endowed Genus of Cyanobacteria
Abstract
1. Introduction
2. Chemical Diversity of the Secondary Metabolites Isolated from Leptolyngbya
2.1. Polypeptides
2.2. Simple Esters
2.3. Macrolides
2.4. Pyrones
2.5. Polyaromatics
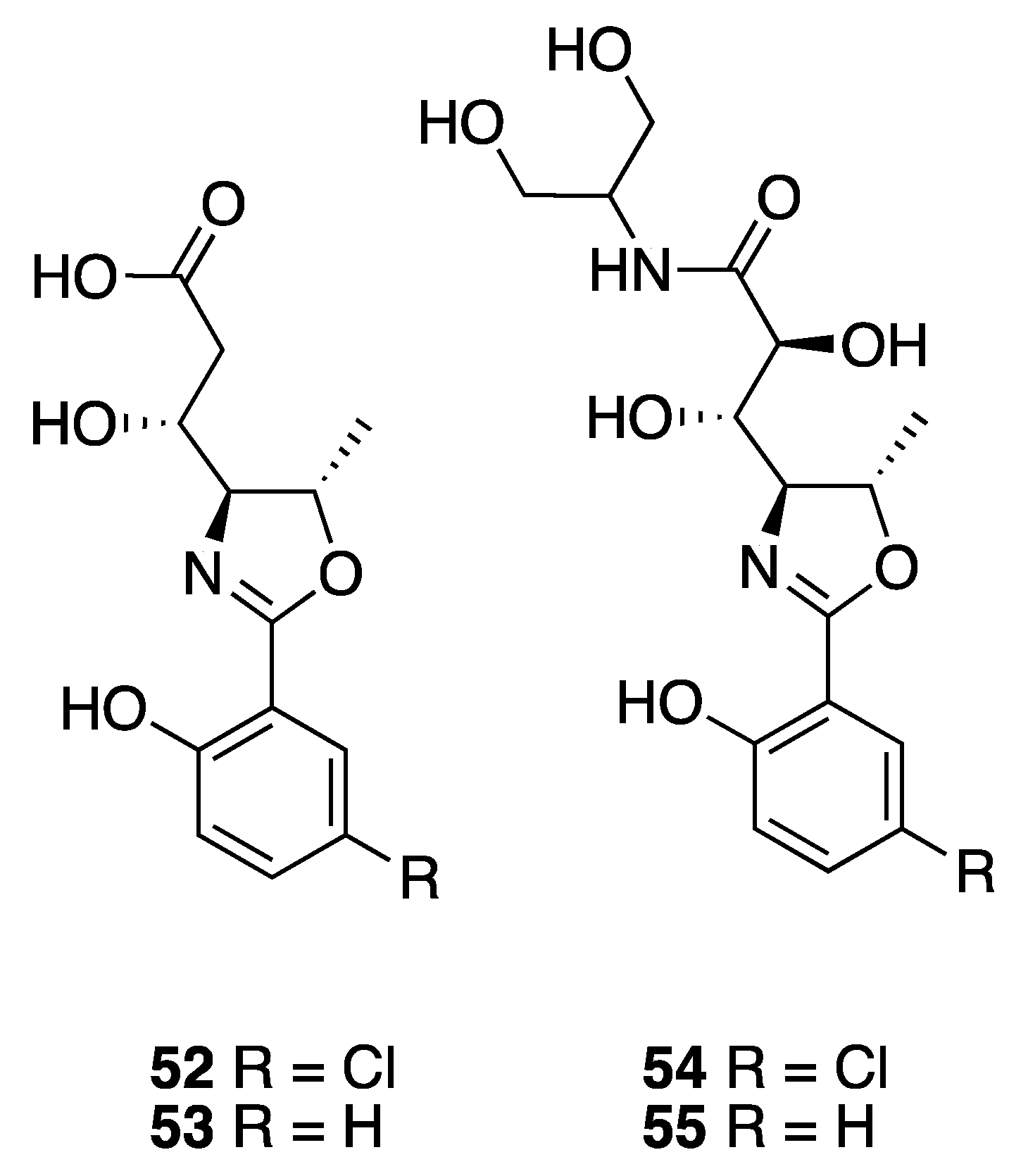
2.6. Oxazolines
2.7. Other
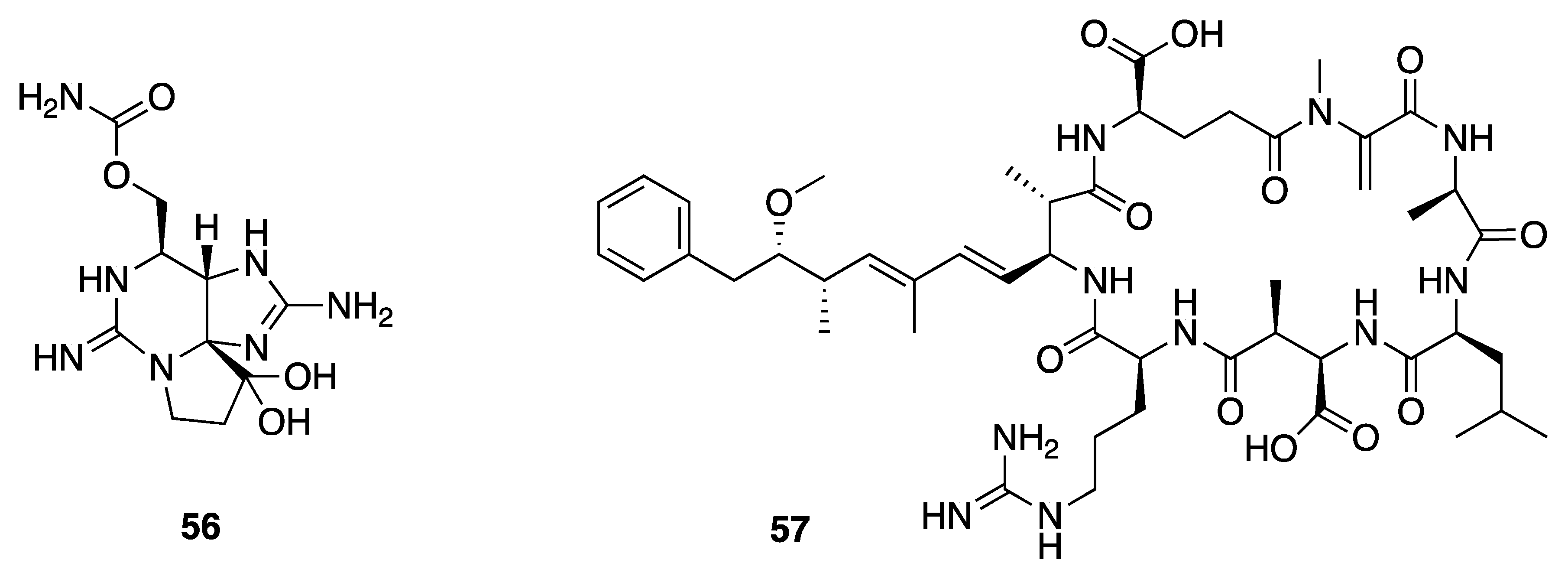
2.7.1. Toxins
2.7.2. Non-Toxic Metabolites
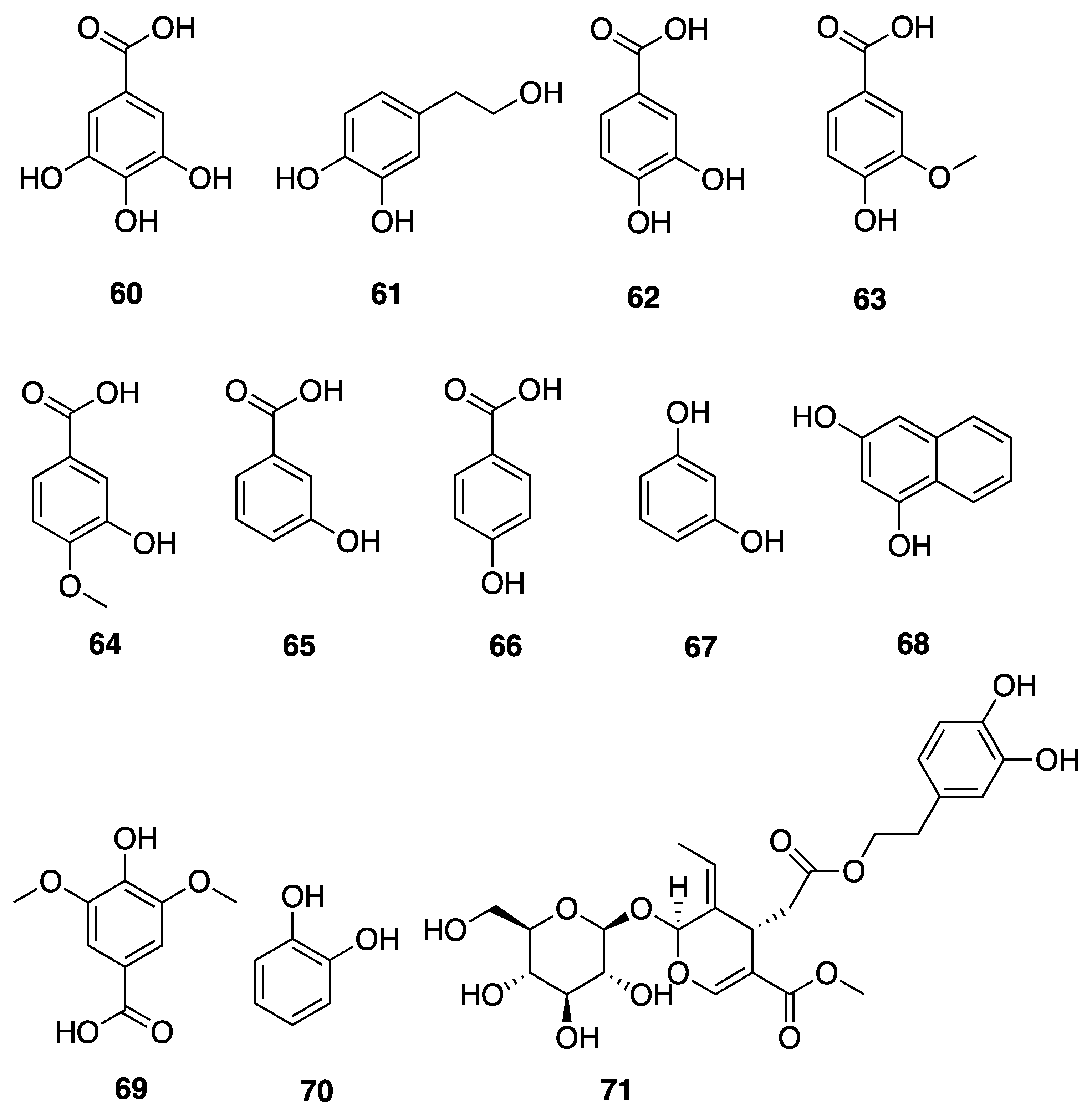
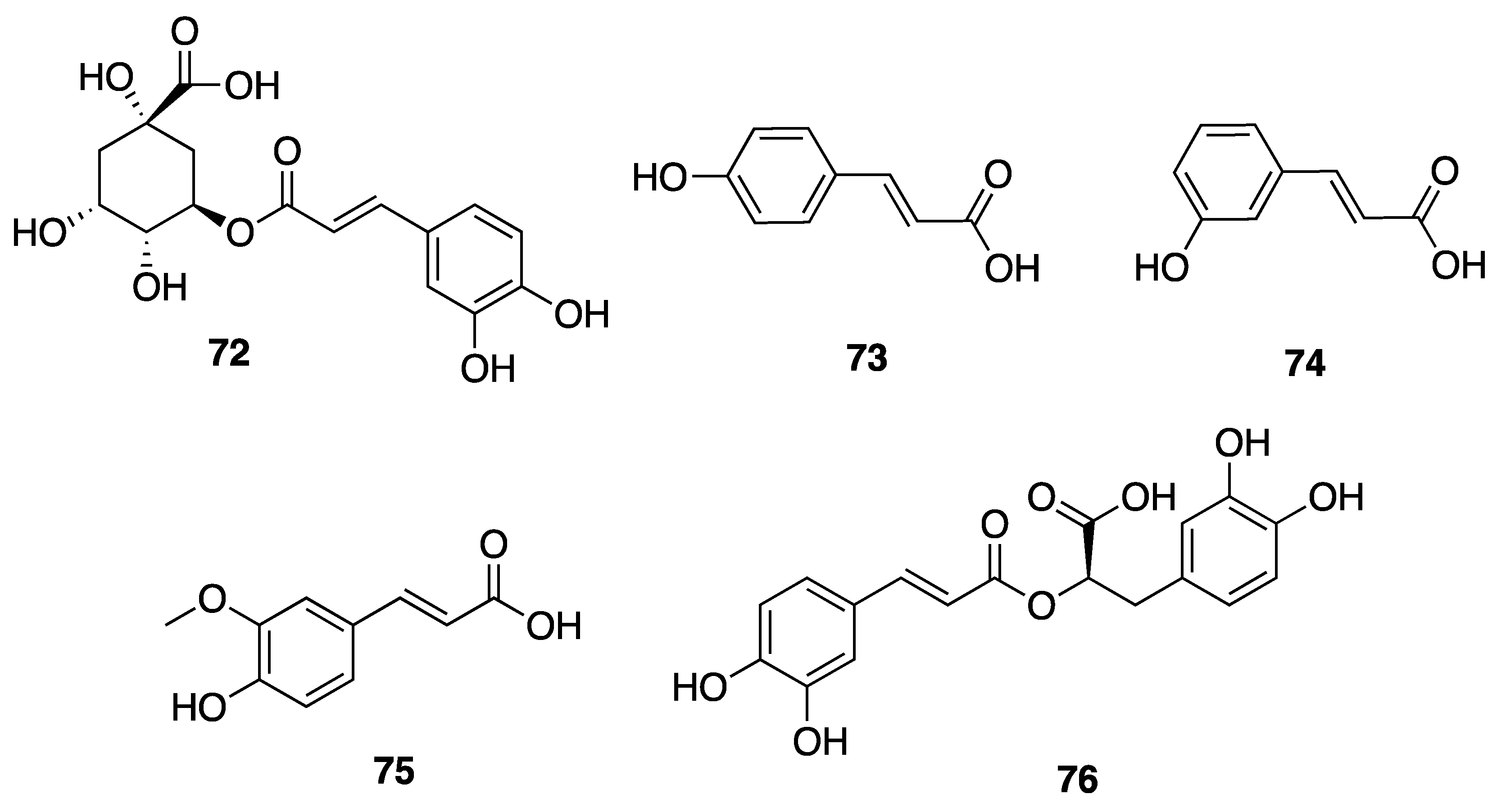
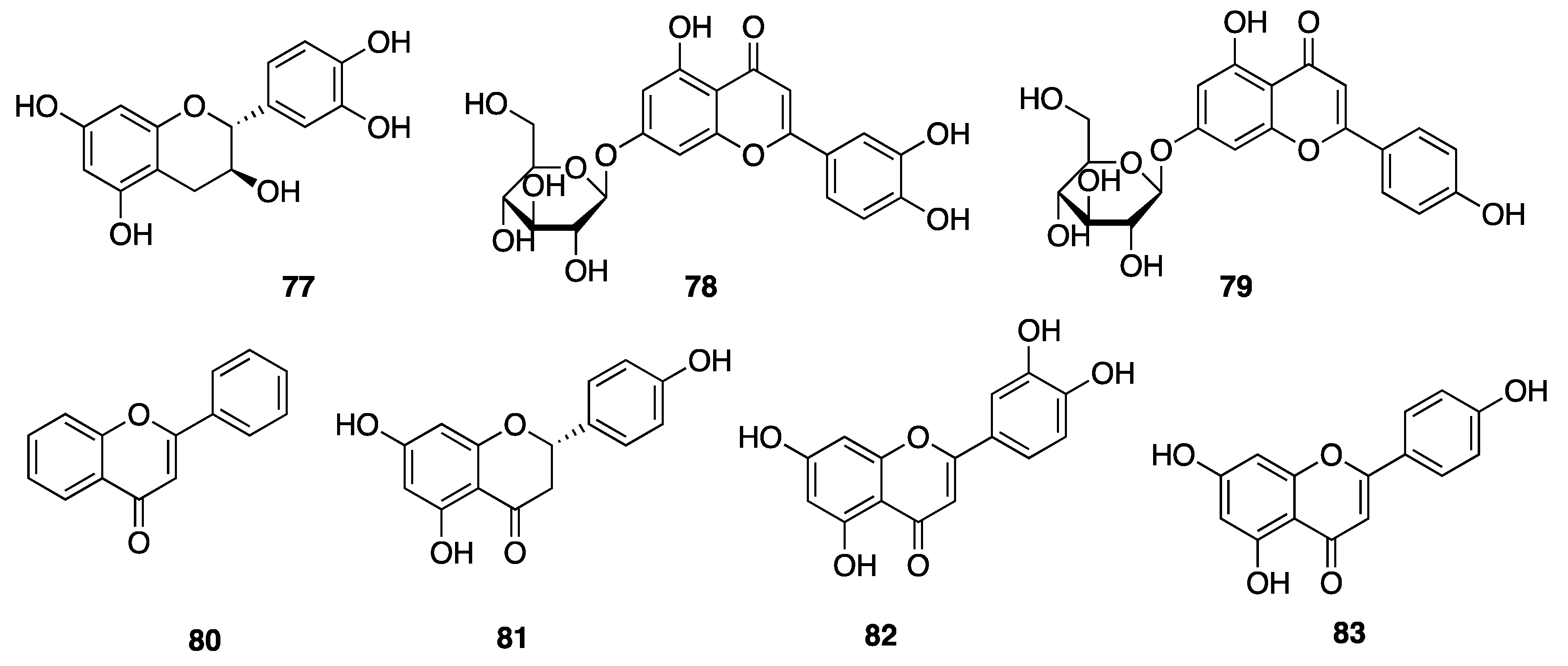
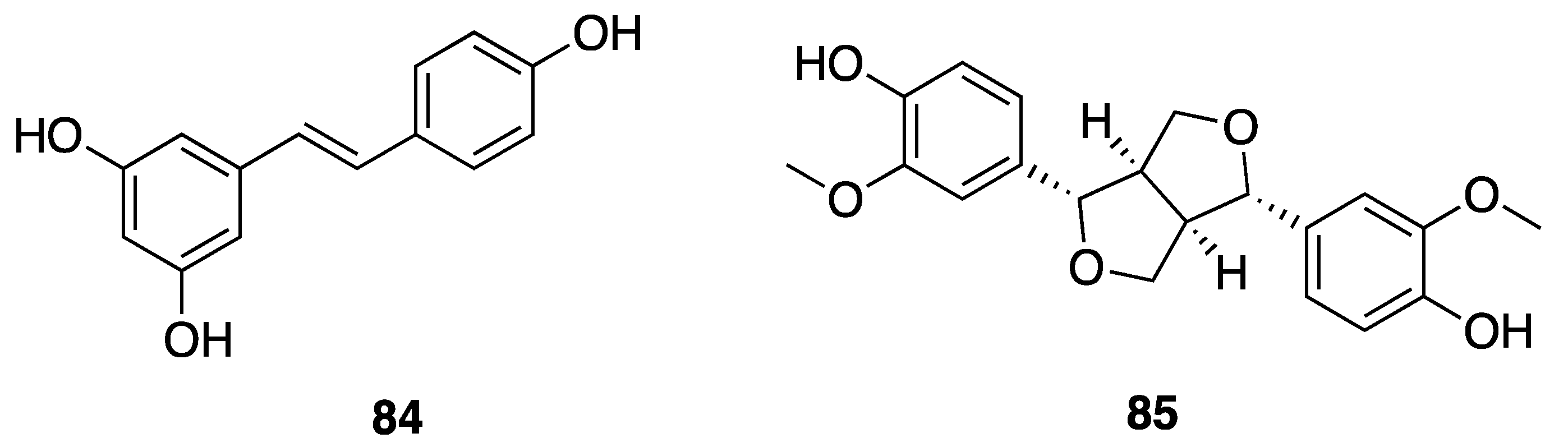
2.7.3. Phenolic Compounds
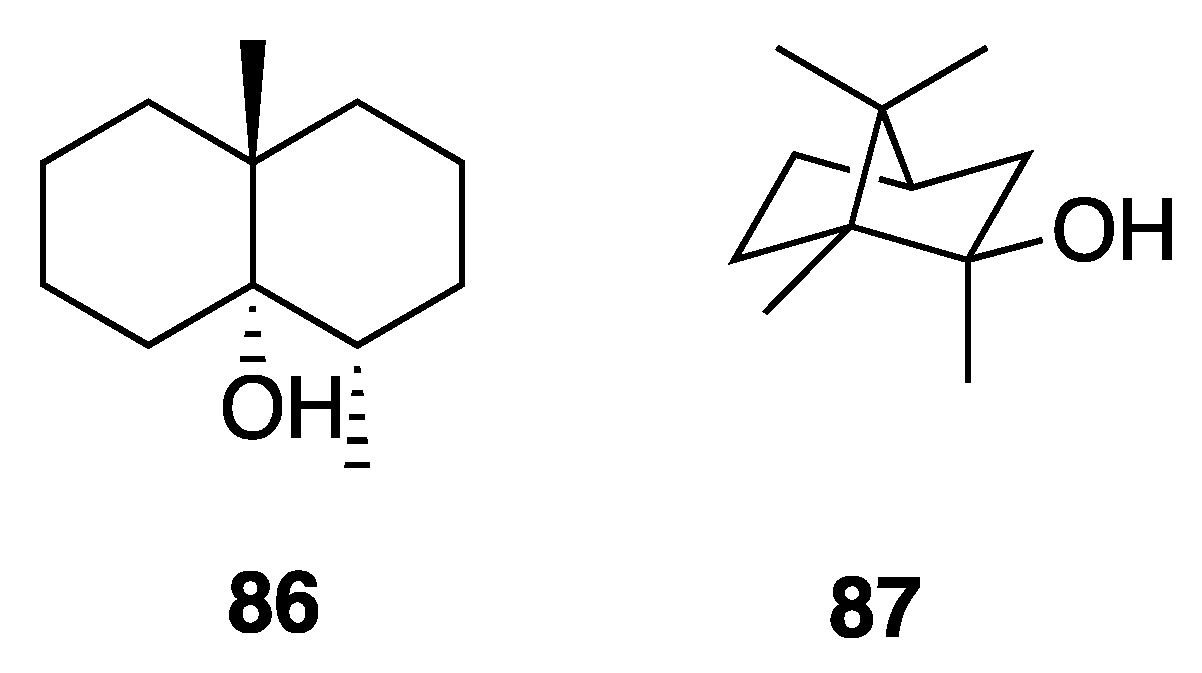
2.7.4. Odorous Metabolites
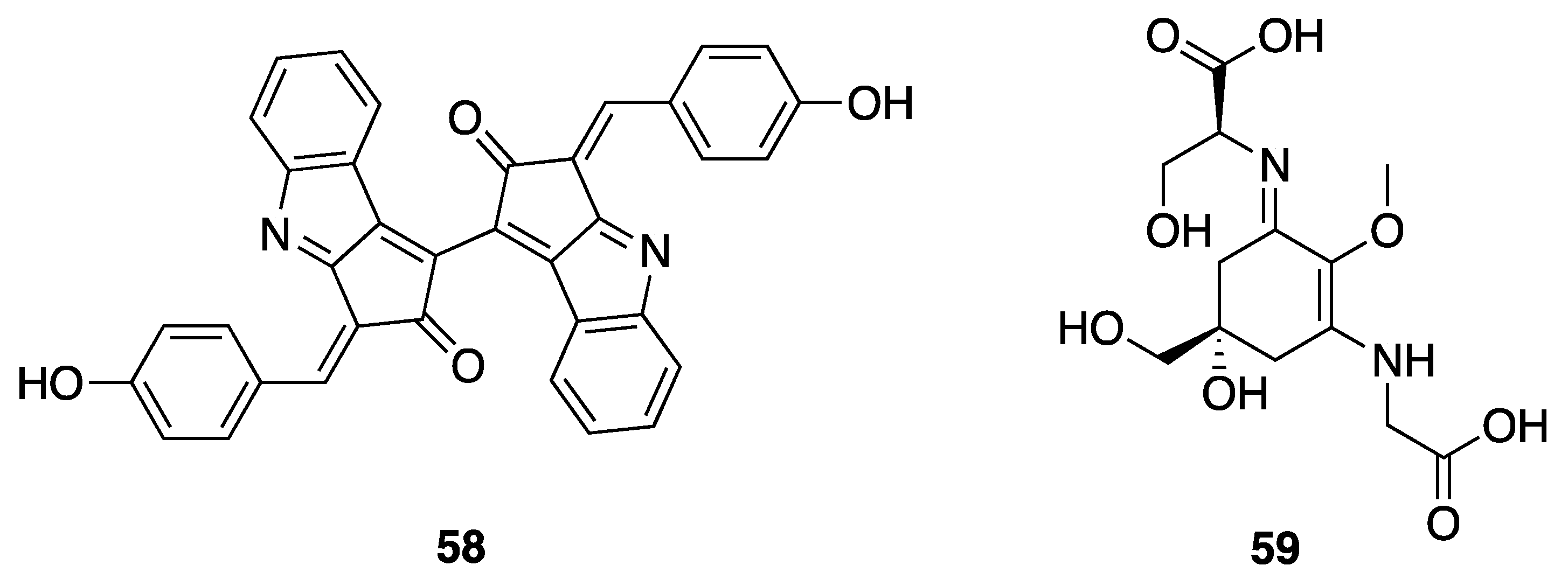
2.7.5. Pigments
3. Laboratory Cultivation
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gademann, K.; Portmann, C. Secondary metabolites from cyanobacteria: Complex structures and powerful bioactivities. Curr. Org. Chem. 2008, 12, 326–341. [Google Scholar] [CrossRef]
- Summons, R.E.; Jahnke, L.L.; Hope, J.M.; Logan, G.A. 2-Methylhopanoids as biomarkers forcyanobacterial oxygenic photosynthesis. Nature 1999, 400, 554–557. [Google Scholar] [CrossRef] [PubMed]
- Blankenship, R.E.; Hartman, H. The origin and evolution of oxygenic photosynthesis. Trends Biochem. Sci. 1998, 23, 94–97. [Google Scholar] [CrossRef]
- Berman-Frank, I.; Lundgren, P.; Falkowski, P. Nitrogen fixation and photosynthetic oxygen evolution in cyanobacteria. Res. Microbiol. 2003, 154, 157–164. [Google Scholar] [CrossRef]
- Kim, J.H.; Choi, W.; Jeon, S.M.; Kim, T.; Park, A.; Kim, J.; Heo, S.J.; Oh, C.; Shim, W.B.; Kang, D.H. Isolation and characterization of Leptolyngbya sp. KIOST-1, a basophilic and euryhaline filamentous cyanobacterium from an open paddle-wheel raceway Arthrospira culture pond in Korea. J. Appl. Microbiol. 2015, 119, 1597–1612. [Google Scholar] [CrossRef]
- Mai, T.; Johansen, J.R.; Pietrasiak, N.; Bohunická, M.; Martin, M.P. Revision of the Synechococcales (Cyanobacteria) through recognition of four families including Oculatellaceae fam. nov. and Trichocoleaceae fam. nov. and six new genera containing 14 species. Phytotaxa 2018, 365, 1–59. [Google Scholar] [CrossRef]
- Bruno, L.; Billi, D.; Bellezza, S.; Albertano, P. Cytomorphological and genetic characterization of troglobitic Leptolyngbya strains isolated from roman hypogea. Appl. Environ. Microbiol. 2009, 75, 608–617. [Google Scholar] [CrossRef]
- Anagnostidis, K.; Komárek, J. Modern approach to the classification system of cyanophytes. 3—Oscillatoriales. Algol. Stud. /Arch. Hydrobiol. Suppl. Vol. 1988, 50–53, 327–472. [Google Scholar]
- Komárek, J.; Kaštovský, J.; Mareš, J.; Johansen, J.R. Taxonomic classification of cyanoprokaryotes (cyanobacterial genera) 2014, using a polyphasic approach. Preslia 2014, 86, 295–335. [Google Scholar]
- Cirés, S.; Casero, M.C.; Quesada, A. Toxicity at the edge of life: A review on cyanobacterial toxins from extreme environments. Mar. Drugs 2017, 15, 233. [Google Scholar] [CrossRef]
- Ahmed, M.; Stal, L.J.; Hasnain, S. The morphology and bioactivity of the rice field cyanobacterium Leptolyngbya. Rev. Biol. Trop. 2014, 62, 1251–1260. [Google Scholar] [PubMed]
- Veerabadhran, M.; Chakraborty, S.; Mitra, S.; Karmakar, S.; Mukherjee, J. Effects of flask configuration on biofilm growth and metabolites of intertidal cyanobacteria isolated from a mangrove forest. J. Appl. Microbiol. 2018, 125, 190–202. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, Z.A.; Al-Shehri, A.M. Biodiversity and toxin production of cyanobacteria in mangrove swamps in the Red Sea off the southern coast of Saudi Arabia. Bot. Mar. 2015, 58, 23–34. [Google Scholar] [CrossRef]
- Yuu, H.; Fujisawa, T.; Ohtsubo, Y.; Katayama, M.; Misawa, N.; Wakazuki, S.; Shimura, Y.; Nakamura, Y.; Kawachi, M.; Yoshikawa, H.; et al. Complete genome sequence of cyanobacterium Leptolyngbya sp. NIES-3755. Am. Soc. Microbiol. 2016, 4, e0009-16. [Google Scholar]
- Brito, Â.; Ramos, V.; Seabra, R.; Santos, A.; Santos, C.L.; Lopo, M.; Ferreira, S.; Martins, A.; Mota, R.; Frazão, B.; et al. Culture-dependent characterization of cyanobacterial diversity in the intertidal zones of the Portuguese coast: A polyphasic study. Syst. Appl. Microbiol. 2012, 35, 110–119. [Google Scholar] [CrossRef]
- Costa, M.; Garcia, M.; Costa-Rpdrigues, J.; Costa, M.S.; Ribeiro, M.J.; Fernandes, M.H.; Barros, P.; Barreiro, A.; Vasconcelos, V.; Martins, R. Exploring bioactive properties of marine cyanobacteria isolated from the Portuguese Coast: High potential as a source of anticancer compounds. Mar. Drugs 2014, 12, 98–114. [Google Scholar] [CrossRef]
- Semary, N.A. El Microscopic, molecular, and biochemical investigations to characterize a benthic cyanoprokaryote Leptolyngbya strain from Egypt. Microsc. Res. Tech. 2013, 76, 249–257. [Google Scholar] [CrossRef]
- Guiry, M.D.; Guiry, G.M. AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. Available online: https://www.algaebase.org (accessed on 3 October 2020).
- Komárek, J.; Hauer, T. The On-line Database of Cyanobacterial Genera. Available online: http://www.cyanodb.cz (accessed on 3 October 2020).
- Singh, D.P.; Khattar, J.I.S.; Gupta, M.; Kaur, G. Evaluation of toxicological impact of cartap hydrochloride on some physiological activities of a non-heterocystous cyanobacterium Leptolyngbya foveolarum. Pestic. Biochem. Physiol. 2014, 110, 63–70. [Google Scholar] [CrossRef]
- Pramanik, A.; Sundararaman, M.; Das, S.; Ghosh, U.; Mukherjee, J. Isolation and characterization of cyanobacteria possessing antimicrobial activity from the sundarbans, the world’s largest tidal mangrove forest. J. Phycol. 2011, 47, 731–743. [Google Scholar] [CrossRef]
- Xue, Y.; Zhao, P.; Quan, C.; Zhao, Z.; Gao, W.; Li, J.; Zu, X.; Fu, D.; Feng, S.; Bai, X.; et al. Cyanobacteria-derived peptide antibiotics discovered since 2000. Peptides 2018, 107, 17–24. [Google Scholar] [CrossRef]
- Haque, F.; Banayan, S.; Yee, J.; Chiang, Y.W. Extraction and applications of cyanotoxins and other cyanobacterial secondary metabolites. Chemosphere 2017, 183, 164–175. [Google Scholar] [CrossRef]
- Vijayakumar, S.; Menakha, M. Pharmaceutical applications of cyanobacteria-A review. J. Acute Med. 2015, 5, 15–23. [Google Scholar] [CrossRef]
- Engene, N.; Gunasekera, S.P.; Gerwick, W.H.; Paul, V.J. Phylogenetic inferences reveal a large extent of novel biodiversity in chemically rich tropical marine cyanobacteria. Appl. Environ. Microbiol. 2013, 79, 1882–1888. [Google Scholar] [CrossRef]
- Bertin, M.J.; Vulpanovici, A.; Monroe, E.A.; Korobeynikov, A.; Sherman, D.H.; Gerwick, L.; Gerwick, W.H. The Phormidolide biosynthetic gene cluster: A trans-AT PKS pathway encoding a toxic macrocyclic polyketide. ChemBioChem 2016, 17, 164–173. [Google Scholar] [CrossRef] [PubMed]
- Castenholz, R.W. Culturing methods for cyanobacteria. Methods Enzymol. 1988, 167, 68–93. [Google Scholar]
- Moss, N.A.; Leao, T.; Glukhov, E.; Gerwick, L.; Gerwick, W.H. Collection, Culturing, and Genome Analyses of Tropical Marine Filamentous Benthic Cyanobacteria, 1st ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2018; Volume 604, ISBN 9780128139592. [Google Scholar]
- Leão, P.N.; Ramos, V.; Goncalves, P.B.; Viana, F.; Lage, O.M.; Gerwick, W.H.; Vasconcelos, V.M. Chemoecological screening reveals high bioactivity in diverse culturable portuguese marine cyanobacteria. Mar. Drugs 2013, 11, 1316–1335. [Google Scholar] [CrossRef] [PubMed]
- Yao, G.; Pan, Z.; Wu, C.; Wang, W.; Fang, L.; Su, W. Efficient synthesis and stereochemical revision of Coibamide, A. J. Am. Chem. Soc. 2015, 137, 13488–13491. [Google Scholar] [CrossRef]
- Yao, G.; Wang, W.; Ao, L.; Cheng, Z.; Wu, C.; Pan, Z.; Liu, K.; Li, H.; Su, W.; Fang, L. Improved total synthesis and biological evaluation of coibamide A analogues. J. Med. Chem. 2018, 61, 8908–8916. [Google Scholar] [CrossRef]
- Medina, R.A.; Goeger, D.E.; Hills, P.; Mooberry, S.L.; Huang, N.; Romero, L.I.; Ortega-Barría, E.; Gerwick, W.H.; McPhail, K.L. Coibamide A, a potent antiproliferative cyclic depsipeptide from the panamanian marine cyanobacterium Leptolyngbya sp. J. Am. Chem. Soc. 2008, 130, 6324–6325. [Google Scholar] [CrossRef]
- Sable, G.A.; Park, J.; Kim, H.; Lim, S.J.; Jang, S.; Lim, D. Solid-Phase Total synthesis of the proposed structure of coibamide A and its derivative: Highly methylated cyclic depsipeptides. Eur. J. Org. Chem. 2015, 2015, 7043–7052. [Google Scholar] [CrossRef]
- He, W.; Qiu, H.B.; Chen, Y.J.; Xi, J.; Yao, Z.J. Total synthesis of proposed structure of coibamide A, a highly N- and O-methylated cytotoxic marine cyclodepsipeptide. Tetrahedron Lett. 2014, 55, 6109–6112. [Google Scholar] [CrossRef]
- Nabika, R.; Suyama, T.L.; Hau, A.M.; Misu, R.; Ohno, H.; Ishmael, J.E.; McPhail, K.L.; Oishi, S.; Fujii, N. Synthesis and biological evaluation of the [d-MeAla11]-epimer of coibamide A. Bioorg. Med. Chem. Lett. 2015, 25, 302–306. [Google Scholar] [CrossRef] [PubMed]
- Thornburg, C.C.; Thimmaiah, M.; Shaala, L.A.; Hau, A.M.; Malmo, J.M.; Ishmael, J.E.; Youssef, D.T.A.; McPhail, K.L. Cyclic depsipeptides, grassypeptolides D and E and Ibu- epidemethoxylyngbyastatin 3, from a Red Sea Leptolyngbya cyanobacterium. J. Nat. Prod. 2011, 74, 1677–1685. [Google Scholar] [CrossRef]
- Harrigan, G.G.; Yoshida, W.Y.; Moore, R.E.; Nagle, D.G.; Park, P.U.; Biggs, J.; Paul, V.J.; Mooberry, S.L.; Corbett, T.H.; Valeriote, F.A. Isolation, structure determination, and biological activity of dolastatin 12 and lyngbyastatin 1 from Lyngbya majuscula/Schizothrix calcicola cyanobacterial assemblages. J. Nat. Prod. 1998, 61, 1221–1225. [Google Scholar] [CrossRef] [PubMed]
- Pettit, G.R.; Kamano, Y.; Kizu, H.; Dufresne, C.; Herald, C.L.; Bontems, R.J.; Schmidt, J.M.; Boettner, F.E.; Nieman, R.A. Isolation and structure of the cell growth inhibitory depsipeptides dolastatins 11 and 12. Heterocycles 1989, 28, 3–8. [Google Scholar] [CrossRef]
- Bates, R.B.; Brusoe, K.G.; Burns, J.J.; Caldera, S.; Cui, W.; Gangwar, S.; Gramme, M.R.; McClure, K.J.; Rouen, G.P.; Schadow, H.; et al. Dolastatins. 26. synthesis and stereochemistry of dolastatin 11. J. Am. Chem. Soc. 1997, 119, 2111–2113. [Google Scholar] [CrossRef]
- Al-Awadhi, F.H.; Paul, V.J.; Luesch, H. Structural diversity and anticancer activity of marine-derived elastase inhibitors: Key features and mechanisms mediating the antimetastatic effects in invasive breast cancer. ChemBioChem 2018, 19, 815–825. [Google Scholar] [CrossRef]
- Maneechote, N.; Yingyongnarongkul, B.E.; Suksamran, A.; Lumyong, S. Inhibition of Vibrio spp. by 2-Hydroxyethyl-11-hydroxyhexadec-9-enoate of marine cyanobacterium Leptolyngbya sp. LT19. Aquac. Res. 2017, 48, 2088–2095. [Google Scholar] [CrossRef]
- Choi, H.; Mascuch, S.J.; Villa, F.A.; Byrum, T.; Teasdale, M.E.; Smith, J.E.; Preskitt, L.B.; Rowley, D.C.; Gerwick, L.; Gerwick, W.H. Honaucins A-C, potent inhibitors of inflammation and bacterial quorum sensing: Synthetic derivatives and structure-activity relationships. Chem. Biol. 2012, 19, 589–598. [Google Scholar] [CrossRef]
- Sapkota, M.; Li, L.; Choi, H.; Gerwick, W.H.; Soh, Y. Bromo-honaucin A inhibits osteoclastogenic differentiation in RAW 264.7 cells via Akt and ERK signaling pathways. Eur. J. Pharmacol. 2015, 769, 100–109. [Google Scholar] [CrossRef]
- Cui, J.; Morita, M.; Ohno, O.; Kimura, T.; Teruya, T.; Watanabe, T.; Suenaga, K.; Shibasaki, M. Leptolyngbyolides, cytotoxic macrolides from the marine cyanobacterium Leptolyngbya sp.: Isolation, biological activity, and catalytic asymmetric total synthesis. Chem. A Eur. J. 2017, 23, 8500–8509. [Google Scholar] [CrossRef] [PubMed]
- Pereira, A.R.; Cao, Z.; Engene, N.; Soria-Mercado, I.E.; Murray, T.F.; Gerwick, W.H. Palmyrolide A, an unusually stabilized neuroactive macrolide from palmyra atoll cyanobacteria. Org. Lett. 2010, 12, 4490–4493. [Google Scholar] [CrossRef] [PubMed]
- Williamson, R.T.; Boulanger, A.; Vulpanovici, A.; Roberts, M.A.; Gerwick, W.H. Structure and absolute stereochemistry of phormidolide, a new toxic metabolite from the marine cyanobacterium Phormidium sp. J. Org. Chem. 2002, 67, 7927–7936. [Google Scholar] [CrossRef] [PubMed]
- Bertin, M.J.; Demirkiran, O.; Navarro, G.; Moss, N.A.; Lee, J.; Goldgof, G.M.; Vigil, E.; Winzeler, E.A.; Valeriote, F.A.; Gerwick, W.H. Kalkipyrone B, a marine cyanobacterial γ-pyrone possessing cytotoxic and anti-fungal activities. Phytochemistry 2016, 122, 113–118. [Google Scholar] [CrossRef]
- Graber, M.A.; Gerwick, W.H. Kalkipyrone, a toxic γ-pyrone from an assemblage of the marine cyanobacteria Lyngbya majuscula and Tolypothrix sp. J. Nat. Prod. 1998, 61, 677–680. [Google Scholar] [CrossRef] [PubMed]
- Inuzuka, T.; Yamamoto, K.; Iwasaki, A.; Ohno, O.; Suenaga, K.; Kawazoe, Y.; Uemura, D. An inhibitor of the adipogenic differentiation of 3T3-L1 cells, yoshinone A, and its analogs, isolated from the marine cyanobacterium Leptolyngbya sp. Tetrahedron Lett. 2014, 55, 6711–6714. [Google Scholar] [CrossRef]
- Choi, H.; Engene, N.; Smith, J.E.; Preskitt, L.B.; Gerwick, W.H. Crossbyanols A-D, toxic brominated polyphenyl ethers from the hawai’ian bloom-forming cyanobacterium Leptolyngbya crossbyana. J. Nat. Prod. 2010, 73, 517–522. [Google Scholar] [CrossRef]
- Bhandari Neupane, J.; Neupane, R.P.; Luo, Y.; Yoshida, W.Y.; Sun, R.; Williams, P.G. Characterization of leptazolines A-D, polar oxazolines from the cyanobacterium Leptolyngbya sp., reveals a glitch with the “willoughby-hoye” scripts for calculating NMR chemical shifts. Org. Lett. 2019, 21, 8449–8453. [Google Scholar] [CrossRef]
- Hau, A.M.; Greenwood, J.A.; Löhr, C.V.; Serrill, J.D.; Proteau, P.J.; Ganley, I.G.; McPhail, K.L.; Ishmael, J.E. Coibamide A induces mTOR-independent autophagy and cell death in human glioblastoma cells. PLoS ONE 2013, 8, e65250. [Google Scholar] [CrossRef]
- Snyder, K.M.; Sikorska, J.; Ye, T.; Fang, L.; Su, W.; Carter, R.G.; McPhail, K.L.; Cheong, P.H.Y. Towards theory driven structure elucidation of complex natural products: Mandelalides and coibamide A. Org. Biomol. Chem. 2016, 14, 5826–5831. [Google Scholar] [CrossRef]
- Pettit, G.R.; Kamano, Y.; Fujii, Y.; Herald, C.L.; Inoue, M.; Brown, P.; Gust, D.; Kitahara, K.; Schmidt, J.M.; Michael, C.; et al. Marine animal biosynthetic constituents for cancer chemotherapy. J. Nat. Prod. 1981, 44, 482–485. [Google Scholar] [CrossRef] [PubMed]
- Adams, B.; Pörzgen, P.; Pittman, E.; Yoshida, W.Y.; Westenburg, H.E.; Horgen, F.D. Isolation and structure determination of malevamide E, a dolastatin 14 analogue, from the marine cyanobacterium Symploca laete-viridis. J. Nat. Prod. 2008, 71, 750–754. [Google Scholar] [CrossRef] [PubMed]
- Thornburg, C.C.; Mcphail, K.L. Investigation of Unique Marine Environments for Microbial Natural Products; Oregon State University: Corvallis, OR, USA, 2013. [Google Scholar]
- Williams, P.G.; Moore, R.E.; Paul, V.J. Isolation and structure determination of lyngbyastatin 3, a lyngbyastatin 1 homologue from the marine cyanobacterium Lyngbya majuscula. Determination of the configuration of the 4-Amino-2,2-dimethyl-3-oxopentanoic acid unit in majusculamide C, dolastatin 12, lyngbyastatin 1, and lyngbyastatin 3 from cyanobacteria. J. Nat. Prod. 2003, 66, 1356–1363. [Google Scholar] [PubMed]
- Gerwick, W.H.; Tan, L.T.; Sitachitta, N. Nitrogen-Containing Metabolites Marine Cyanobacteria; Elsevier: Amsterdam, The Netherlands, 2001; Volume 57. [Google Scholar]
- Kwan, J.C.; Ratnayake, R.; Abboud, K.A.; Paul, V.J.; Luesch, H. Grassypeptolides A-C, cytotoxic bis-thiazoline containing marine cyclodepsipeptides. J. Org. Chem. 2010, 75, 8012–8023. [Google Scholar] [CrossRef] [PubMed]
- Kwan, J.C.; Rocca, J.R.; Abboud, K.A.; Paul, V.J.; Luesch, H. Total structure determination of grassypeptolide, a new marine cyanobacterial cytotoxin. Org. Lett. 2008, 10, 789–792. [Google Scholar] [CrossRef]
- Gogineni, V.; Hamann, M.T. Marine natural product peptides with therapeutic potential: Chemistry, biosynthesis, and pharmacology. Biochim. Biophys. Acta Gen. Subj. 2018, 1862, 81–196. [Google Scholar] [CrossRef]
- Popplewell, W.L.; Ratnayake, R.; Wilson, J.A.; Beutler, J.A.; Colburn, N.H.; Henrich, C.J.; McMahon, J.B.; McKee, T.C. Grassypeptolides F and G, cyanobacterial peptides from Lyngbya majuscula. J. Nat. Prod. 2011, 74, 1686–1691. [Google Scholar] [CrossRef]
- Kwan, J.C.; Liu, Y.; Ratnayake, R.; Hatano, R.; Kuribara, A.; Morimoto, C.; Ohnuma, K.; Paul, V.J.; Ye, T.; Luesch, H. Grassypeptolides as natural inhibitors of dipeptidyl peptidase 8 and T-cell activation. ChemBioChem 2014, 15, 799–804. [Google Scholar] [CrossRef]
- Mascuch, S.J.; Boudreau, P.D.; Carland, T.M.; Pierce, N.T.; Olson, J.; Hensler, M.E.; Choi, H.; Campanale, J.; Hamdoun, A.; Nizet, V.; et al. Marine natural product honaucin A attenuates inflammation by activating the Nrf2-ARE pathway. J. Nat. Prod. 2018, 81, 506–514. [Google Scholar] [CrossRef]
- Skiba, M.A.; Sikkema, A.P.; Moss, N.A.; Lowell, A.N.; Su, M.; Sturgis, R.M.; Gerwick, L.; Gerwick, W.H.; Sherman, D.H.; Smith, J.L. Biosynthesis of t-butyl in apratoxin A: Functional analysis and architecture of a PKS loading module. ACS Chem. Biol. 2018, 13, 1640–1650. [Google Scholar] [CrossRef]
- Wadsworth, A.D.; Furkert, D.P.; Brimble, M.A. Total synthesis of the macrocyclic N-Methyl enamides palmyrolide A and 2S-sanctolide A. J. Org. Chem. 2014, 79, 11179–11193. [Google Scholar] [CrossRef]
- Tello-Aburto, R.; Johnson, E.M.; Valdez, C.K.; Maio, W.A. Asymmetric total synthesis and absolute stereochemistry of the neuroactive marine macrolide palmyrolide A. Org. Lett. 2012, 14, 2150–2153. [Google Scholar] [CrossRef]
- Tello-Aburto, R.; Newar, T.D.; Maio, W.A. Evolution of a protecting-group-free total synthesis: Studies en route to the neuroactive marine macrolide (-)-palmyrolide A. J. Org. Chem. 2012, 77, 6271–6289. [Google Scholar] [CrossRef]
- Lam, N.Y.S.; Muir, G.; Challa, V.R.; Britton, R.; Paterson, I. A counterintuitive stereochemical outcome from a chelation-controlled vinylmetal aldehyde addition leads to the configurational reassignment of phormidolide A. Chem. Commun. 2019, 55, 9717–9720. [Google Scholar] [CrossRef]
- Helfrich, E.J.N.; Ueoka, R.; Dolev, A.; Rust, M.; Meoded, R.A.; Bhushan, A.; Califano, G.; Costa, R.; Gugger, M.; Steinbeck, C.; et al. Automated structure prediction of trans-acyltransferase polyketide synthase products. Nat. Chem. Biol. 2019, 15, 813–821. [Google Scholar] [CrossRef]
- Ndukwe, I.E.; Wang, X.; Lam, N.Y.S.; Ermanis, K.; Alexander, K.L.; Bertin, M.J.; Martin, G.E.; Muir, G.; Paterson, I.; Britton, R.; et al. Synergism of anisotropic and computational NMR methods reveals the likely configuration of phormidolide A. Chem. Commun. 2020, 56, 7565–7568. [Google Scholar] [CrossRef]
- Shinomiya, S.; Iwasaki, A.; Ohno, O.; Suenaga, K. Total synthesis and stereochemical determination of yoshinone A. Phytochemistry 2016, 132, 109–114. [Google Scholar] [CrossRef]
- Koyama, T.; Kawazoe, Y.; Iwasaki, A.; Ohno, O.; Suenaga, K.; Uemura, D. Anti-obesity activities of the yoshinone A and the related marine γ-pyrone compounds. J. Antibiot. 2016, 69, 348–351. [Google Scholar] [CrossRef]
- Mohamed, Z. Harmful cyanobacteria and their cyanotoxins in Egyptian fresh waters–state of knowledge and research needs. Afr. J. Aquat. Sci. 2016, 41, 361–368. [Google Scholar] [CrossRef]
- Joshi, D.; Mohandass, C.; Dhale, M. Effect of UV-B radiation and desiccation stress on photoprotective compounds accumulation in marine Leptolyngbya sp. Appl. Biochem. Biotechnol. 2018, 184, 35–47. [Google Scholar] [CrossRef]
- Babu, S.V.; Ashokkumar, B.; Sivakumar, N.; Sudhakarsamy, P.; Varalakshmi, P. Indole-3-acetic acid from filamentous cyanobacteria: Screening, strain identification and production. J. Sci. Ind. Res. 2013, 72, 581–584. [Google Scholar]
- Stevenson, C.S.; Capper, E.A.; Roshak, A.K.; Marquez, B.; Eichman, C.; Jackson, J.R.; Mattern, M.; Gerwick, W.H.; Jacobs, R.S.; Marshall, L.A. The identification and characterization of the marine natural product scytonemin as a novel antiproliferative pharmacophore. J. Pharmacol. Exp. Ther. 2002, 303, 858–866. [Google Scholar] [CrossRef] [PubMed]
- Soule, T.; Stout, V.; Swingley, W.D.; Meeks, J.C.; Garcia-Pichel, F. Molecular genetics and genomic analysis of scytonemin biosynthesis in Nostoc punctiforme ATCC 29133. J. Bacteriol. 2007, 189, 4465–4472. [Google Scholar] [CrossRef] [PubMed]
- Balskus, E.P.; Walsh, C.T. Investigating the initial steps in the biosynthesis of cyanobacterial sunscreen scytonemin. J. Am. Chem. Soc. 2008, 130, 15260–15261. [Google Scholar] [CrossRef]
- Sorrels, C.M.; Proteau, P.J.; Gerwick, W.H. Organization, evolution, and expression analysis of the biosynthetic gene cluster for scytonemin, a cyanobacterial UV-absorbing pigment. Appl. Environ. Microbiol. 2009, 75, 4861–4869. [Google Scholar] [CrossRef]
- Jones, C.S.; Esquenazi, E.; Dorrestein, P.C.; Gerwick, W.H. Probing the in vivo biosynthesis of scytonemin, a cyanobacterial ultraviolet radiation sunscreen, through small scale stable isotope incubation studies and MALDI-TOF mass spectrometry. Bioorg. Med. Chem. 2011, 19, 6620–6627. [Google Scholar] [CrossRef]
- Martins, S.; Mussatto, S.I.; Martínez-Avila, G.; Montañez-Saenz, J.; Aguilar, C.N.; Teixeira, J.A. Bioactive phenolic compounds: Production and extraction by solid-state fermentation. A review. Biotechnol. Adv. 2011, 29, 365–373. [Google Scholar] [CrossRef]
- Ljaz, S.; Hasnain, S. Antioxidant potential of indigenous cyanobacterial strains in relation with their phenolic and flavonoid contents. Nat. Prod. Res. 2016, 30, 1297–1300. [Google Scholar]
- Trabelsi, L.; Mnari, A.; Abdel-Daim, M.M.; Abid-Essafi, S.; Aleya, L. Therapeutic properties in Tunisian hot springs: First evidence of phenolic compounds in the cyanobacterium Leptolyngbya sp. biomass, capsular polysaccharides and releasing polysaccharides. BMC Complement. Altern. Med. 2016, 16, 515. [Google Scholar] [CrossRef]
- Smith, J.L.; Boyer, G.L.; Zimba, P.V. A review of cyanobacterial odorous and bioactive metabolites: Impacts and management alternatives in aquaculture. Aquaculture 2008, 280, 5–20. [Google Scholar] [CrossRef]
- Wang, Z.; Xiao, P.; Song, G.; Li, Y.; Li, R. Isolation and characterization of a new reported cyanobacterium Leptolyngbya bijugata coproducing odorous geosmin and 2-methylisoborneol. Environ. Sci. Pollut. Res. 2015, 22, 12133–12140. [Google Scholar] [CrossRef] [PubMed]
- Schipper, K.; Fortunati, F.; Oostlander, P.C.; Al Muraikhi, M.; Al Jabri, H.M.S.J.; Wijffels, R.H.; Barbosa, M.J. Production of phycocyanin by Leptolyngbya sp. in desert environments. Algal Res. 2020, 47, 101875. [Google Scholar] [CrossRef]
- Singh, N.K.; Sonani, R.R.; Awasthi, A.; Prasad, B.; Patel, A.R.; Kumar, J.; Madamwar, D. Phycocyanin moderates aging and proteotoxicity in caenorhabditis elegans. J. Appl. Phycol. 2016, 28, 2407–2417. [Google Scholar] [CrossRef]
- Kim, S.-K. Handbook of Anticancer Drugs from Marine Origin; Springer: Berlin/Heidelberg, Germany, 2015; ISBN 9783319071442. [Google Scholar]
- Dittmann, E.; Gugger, M.; Sivonen, K.; Fewer, D.P. Natural product biosynthetic diversity and comparative genomics of the cyanobacteria. Trends Microbiol. 2015, 23, 642–652. [Google Scholar] [CrossRef]
- Videau, P.; Wells, K.N.; Singh, A.J.; Eiting, J.; Proteau, P.J.; Philmus, B. Expanding the natural products heterologous expression repertoire in the model cyanobacterium Anabaena sp. strain PCC 7120: Production of pendolmycin and teleocidin B-4. ACS Synth. Biol. 2020, 9, 63–75. [Google Scholar] [CrossRef]
- Taton, A.; Ecker, A.; Diaz, B.; Moss, N.A.; Anderson, B.; Leão, T.F.; Simkovsky, R.; Dorrestein, P.C.; Gerwick, L.; William, H. Heterologous expression of cryptomaldamide in a cyanobacterial host. BioRxiv 2020. [Google Scholar] [CrossRef]
| Name | Geographic Location | Culture | Total Synthesis | Bioactivity | Cell Line | Activity | Reference |
|---|---|---|---|---|---|---|---|
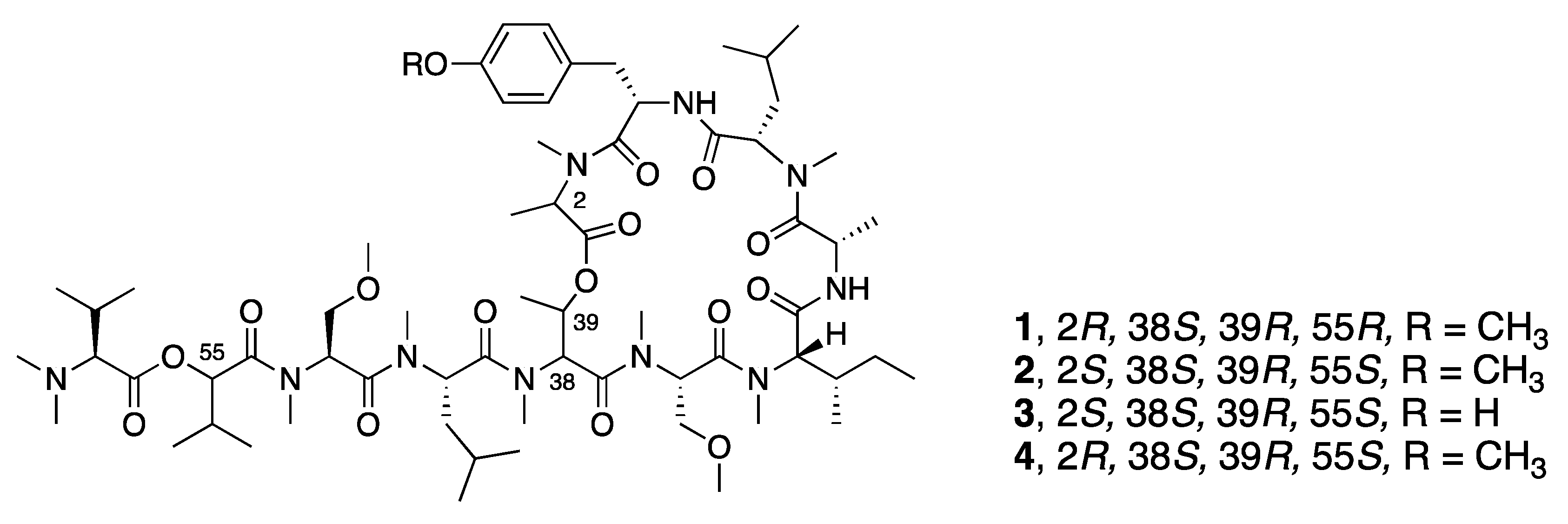
| Coibamide A (1) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | IC50 3.9 nM | [30] |
| Cytotoxicity | A549 | IC50 3.6 nM | |||||
| Cytotoxicity | MCF-7 | IC50 35.7 nM | |||||
| N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 5.0 nM | [31] | |
| Cytotoxicity | A549 | GI50 5.4 nM | |||||
| Cytotoxicity | PANC-1 | GI50 3.1 nM | |||||
| Coiba National Park, Panama | N a | N a | Cytotoxicity | H460 | LC50 < 23 nM | [32] | |
| Cytotoxicity | Mouse neuro2a | LC50 < 23 nM | |||||
| Cytotoxicity | MDA-MB-231 | GI50 2.8 nM | |||||
| Cytotoxicity | LOX IMVI | GI50 7.4 nM | |||||
| Cytotoxicity | HL-60(TB) | GI50 7.4 nM | |||||
| Cytotoxicity | SNB-75 | GI50 7.6 nM | |||||
| Histological Selectivity | Breast, CNS, colon, and ovarian cancer cells | Good | |||||
| Coiba National Park, Panama | N a | N a | Cytotoxicity | U87-MG | EC50 28.8 nM | [33] | |
| Cytotoxicity | SF-295 glioblastoma cell | EC50 96.2 nM | |||||
| Cytotoxicity | MEFs | EC50 96.2 nM | |||||
| Synthetic l-HVA, l-MeAla-Coibamide A (2) | N a | N a | Y b | Cytotoxicity | COLO205 | IC50 11.5 µM | [33] |
| Cytotoxicity | H460 | 45% inhibition at 20 µM | |||||
| Cytotoxicity | MDA-MB-231 | IC50 17.98 µM | [34] | ||||
| Cytotoxicity | MCF-7 | IC50 11.77 µM | |||||
| Cytotoxicity | A549 | IC50 22.80 µM | |||||
| Cytotoxicity | MDA-MB-231 | GI50 > 16000 nM | [31] | ||||
| Cytotoxicity | A549 | GI50 22800 nM | |||||
| Cytotoxicity | PANC-1 | ND c | |||||
| N a | N a | N a | Cytotoxicity | H292 | IC50 124 nM | [35] | |
| Cytotoxicity | MDA-MB-231 | IC50 66 nM | |||||
| Cytotoxicity | PC-3 | IC50 80 nM | |||||
| Cytotoxicity | SF-295 | IC50 219 nM | |||||
| Synthetic O-Desmethyl, l-HVA, l-MeAla-Coibamide A (3) | N a | N a | Y b | Cytotoxicity | COLO205 | IC50 13 µM | [33] |
| Cytotoxicity | H460 | 36% inhibition at 20 µM | |||||
| Synthetic l-HVA, d-MeAla-Coibamide A (4) | N a | N a | Y b | Cytotoxicity | A549 | IC50 19.0 nM | [35] |
| Cytotoxicity | HCT116 | IC50 44.6 nM | |||||
| Cytotoxicity | MCF-7 | IC50 48.6 nM | |||||
| Cytotoxicity | B16 | IC50 54.4 nM | |||||
| Cytotoxicity | H292 | IC50 610 nM | |||||
| Cytotoxicity | MDA-MB-231 | IC50 545 nM | |||||
| Cytotoxicity | PC-3 | IC50 424 nM | |||||
| Cytotoxicity | SF-295 | IC50 816 nM | |||||
| Cytotoxicity | MDA-MB-231 | GI50 545 nM | [31] | ||||
| Cytotoxicity | A549 | GI50 19 nM | |||||
| Cytotoxicity | PANC-1 | ND c | |||||
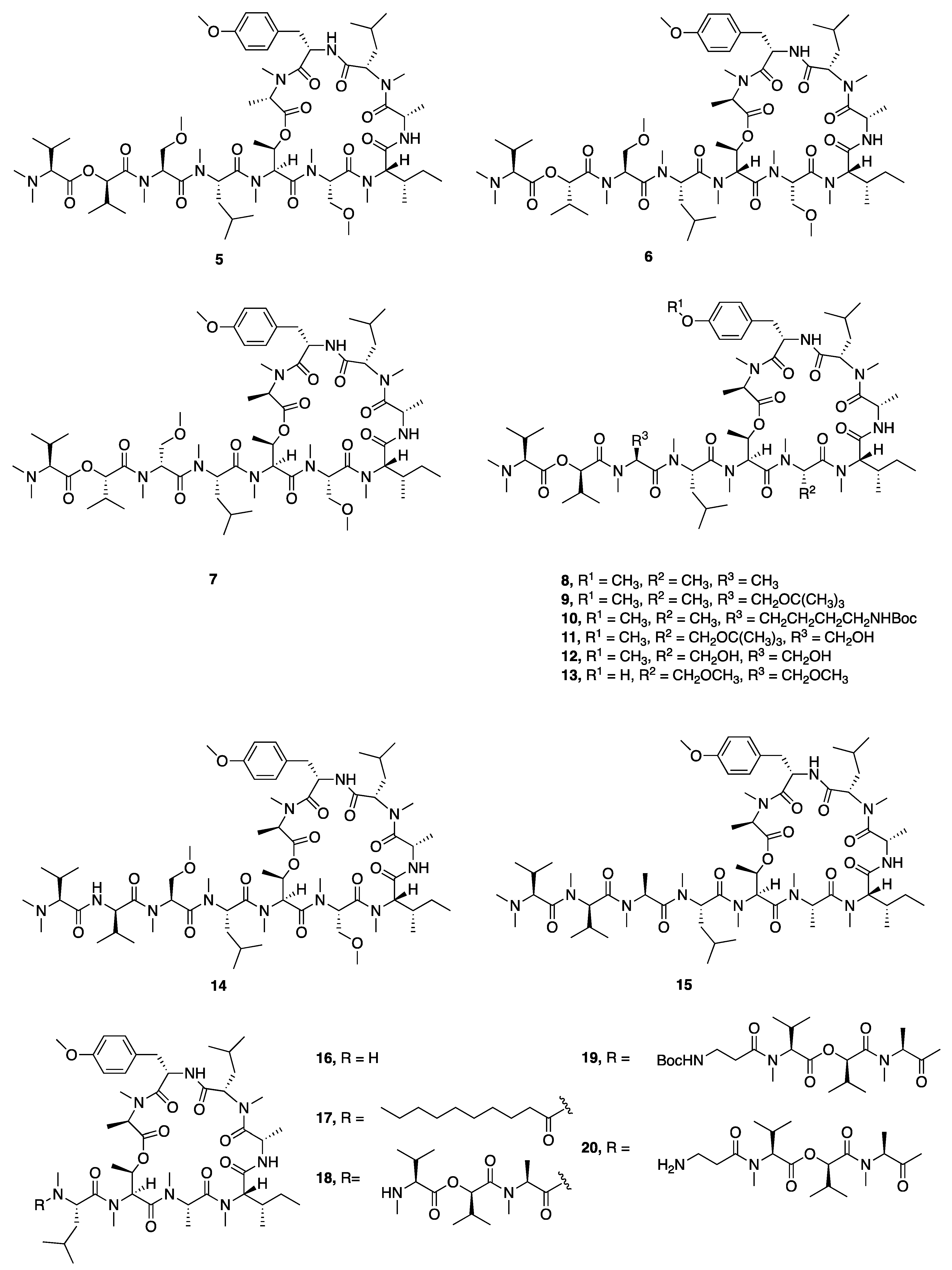
| Synthetic Coibamide A-1c (5) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 7518 nM | [31] |
| Cytotoxicity | A549 | GI50 20091 nM | |||||
| Cytotoxicity | PANC-1 | GI50 12417 nM | |||||
| Synthetic Coibamide A-1d (6) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 10809 nM | [31] |
| Cytotoxicity | A549 | ND c | |||||
| Cytotoxicity | PANC-1 | ND c | |||||
| Synthetic Coibamide A-1e (7) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 2662 nM | [31] |
| Cytotoxicity | A549 | GI50 1995 nM | |||||
| Cytotoxicity | PANC-1 | GI50 1906 nM | |||||
| Synthetic MeAla3-MeAla6-Coibamide A-1f (8) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 5.1 nM | [31] |
| Cytotoxicity | A549 | GI50 7.3 nM | |||||
| Cytotoxicity | PANC-1 | GI50 7.0 nM | |||||
| Synthetic Coibamide A-1g (9) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 5.3 nM | [31] |
| Cytotoxicity | A549 | GI50 12.4 nM | |||||
| Cytotoxicity | PANC-1 | GI50 32.9 nM | |||||
| Synthetic Coibamide A-1h (10) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 61.6 nM | [31] |
| Cytotoxicity | A549 | GI50 81.7 nM | |||||
| Cytotoxicity | PANC-1 | GI50 124 nM | |||||
| Synthetic Coibamide A-1i (11) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 20.8 nM | [31] |
| Cytotoxicity | A549 | GI50 194 nM | |||||
| Cytotoxicity | PANC-1 | GI50 46.3 nM | |||||
| Synthetic Coibamide A-1j (12) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 2056 nM | [31] |
| Cytotoxicity | A549 | ND c | |||||
| Cytotoxicity | PANC-1 | GI50 2178 nM | |||||
| Synthetic Coibamide A-1k (13) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 183 nM | [31] |
| Cytotoxicity | A549 | GI50 222 nM | |||||
| Cytotoxicity | PANC-1 | GI50 277 nM | |||||
| Synthetic Coibamide A-1l (14) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 450 nM | [31] |
| Cytotoxicity | A549 | GI50 473 nM | |||||
| Cytotoxicity | PANC-1 | GI50 601 nM | |||||
| Synthetic Coibamide A-1m (15) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 415 nM | [31] |
| Cytotoxicity | A549 | GI50 511 nM | |||||
| Cytotoxicity | PANC-1 | GI50 723 nM | |||||
| Synthetic Coibamide A-1n (16) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 >16000nM | [31] |
| Cytotoxicity | A549 | ND c | |||||
| Cytotoxicity | PANC-1 | ND c | |||||
| Synthetic Coibamide A-1o (17) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 470 nM | [31] |
| Cytotoxicity | A549 | GI50 733 nM | |||||
| Cytotoxicity | PANC-1 | GI50 828 nM | |||||
| Synthetic Coibamide A-1p (18) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 236 nM | [31] |
| Cytotoxicity | A549 | GI50 360 nM | |||||
| Cytotoxicity | PANC-1 | GI50 204 nM | |||||
| Synthetic Coibamide A-1q (19) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 239 nM | [31] |
| Cytotoxicity | A549 | GI50 443 nM | |||||
| Cytotoxicity | PANC-1 | GI50 415 nM | |||||
| Synthetic Coibamide A-1r (20) | N a | N a | Y b | Cytotoxicity | MDA-MB-231 | GI50 >16000 nM | [31] |
| Cytotoxicity | A549 | ND c | |||||
| Cytotoxicity | PANC-1 | ND c | |||||
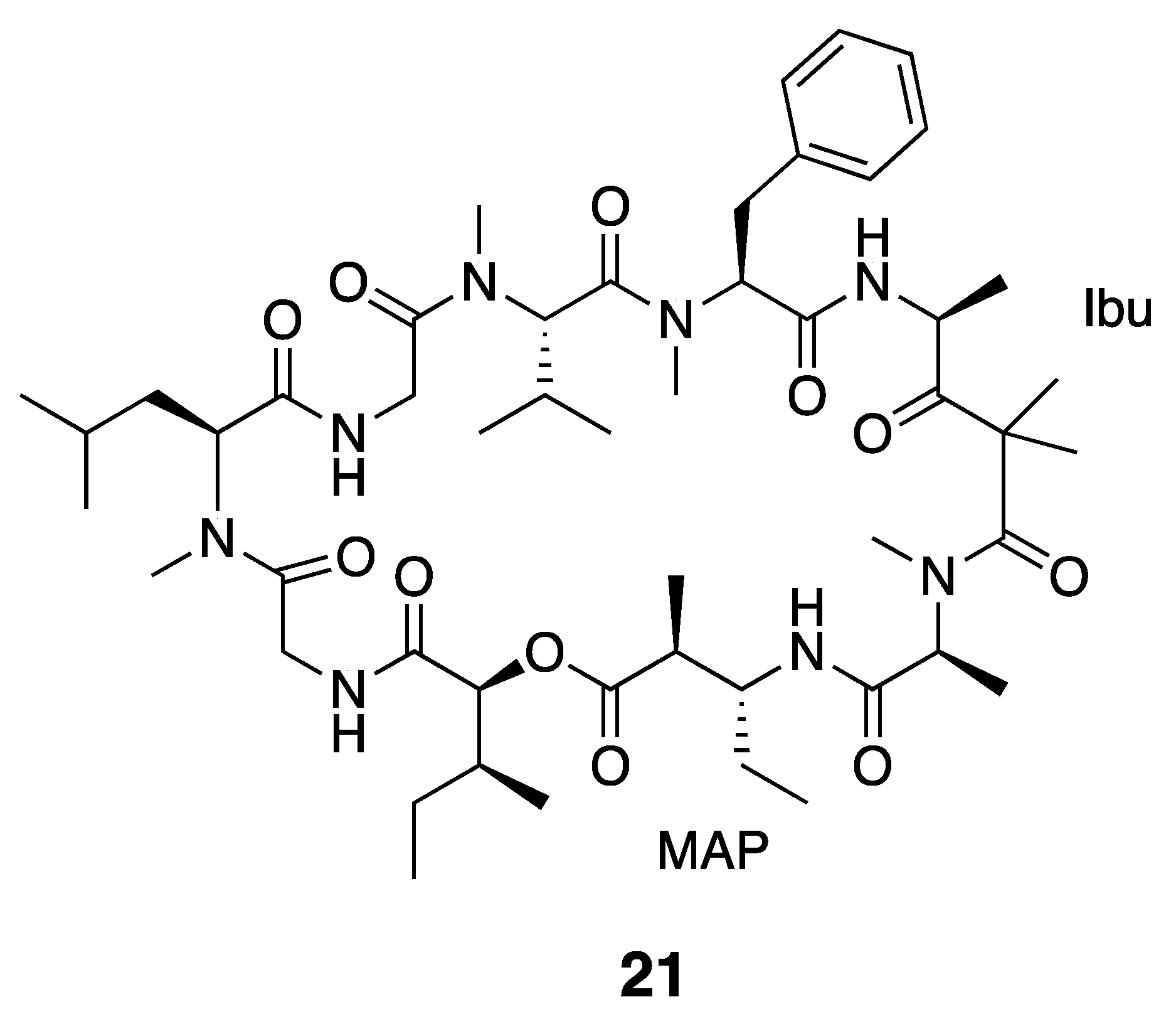
| Dolastatin 12 (21) | The Red Sea | Y b | N a | Cytotoxicity | HeLa cells | IC50 > 1 μM | [36] |
| KB (human nasopharyngeal carcinoma cell line) | MICs <0.05 µg/mL | [37] | |||||
| LoVo (a human colon adenocarcinoma cell line) | 0.08 µg/mL | ||||||
| Murine P388 lymphocytic leukemia | ED50 7.5 × 10−2 µg/mL | [38,39] | |||||
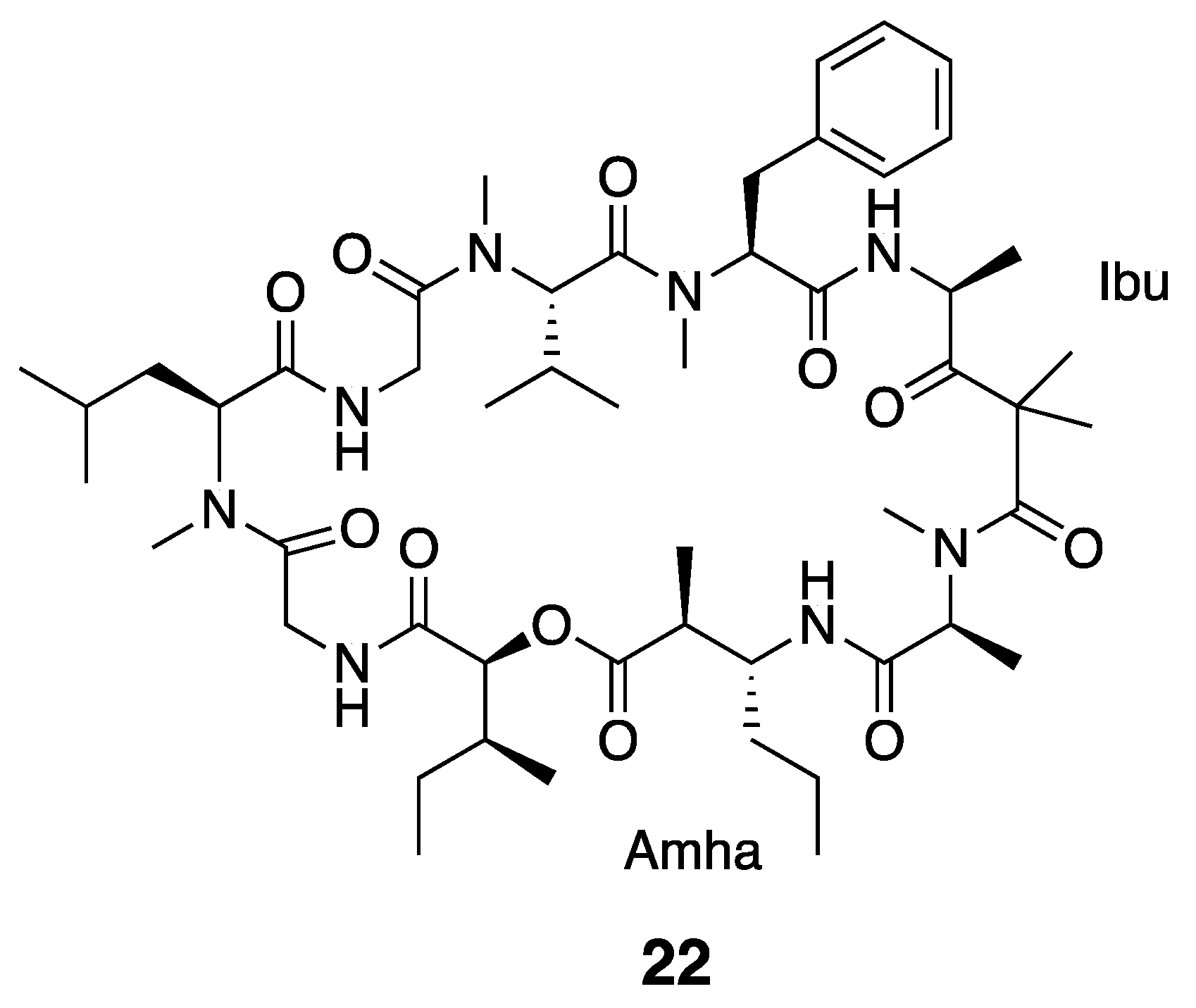
| Ibu-Epidemethoxylyngbyastatin 3 (22) | The Red Sea | Y b | N a | Cytotoxicity | HeLa cells | IC50 > 10 μM | [36] |
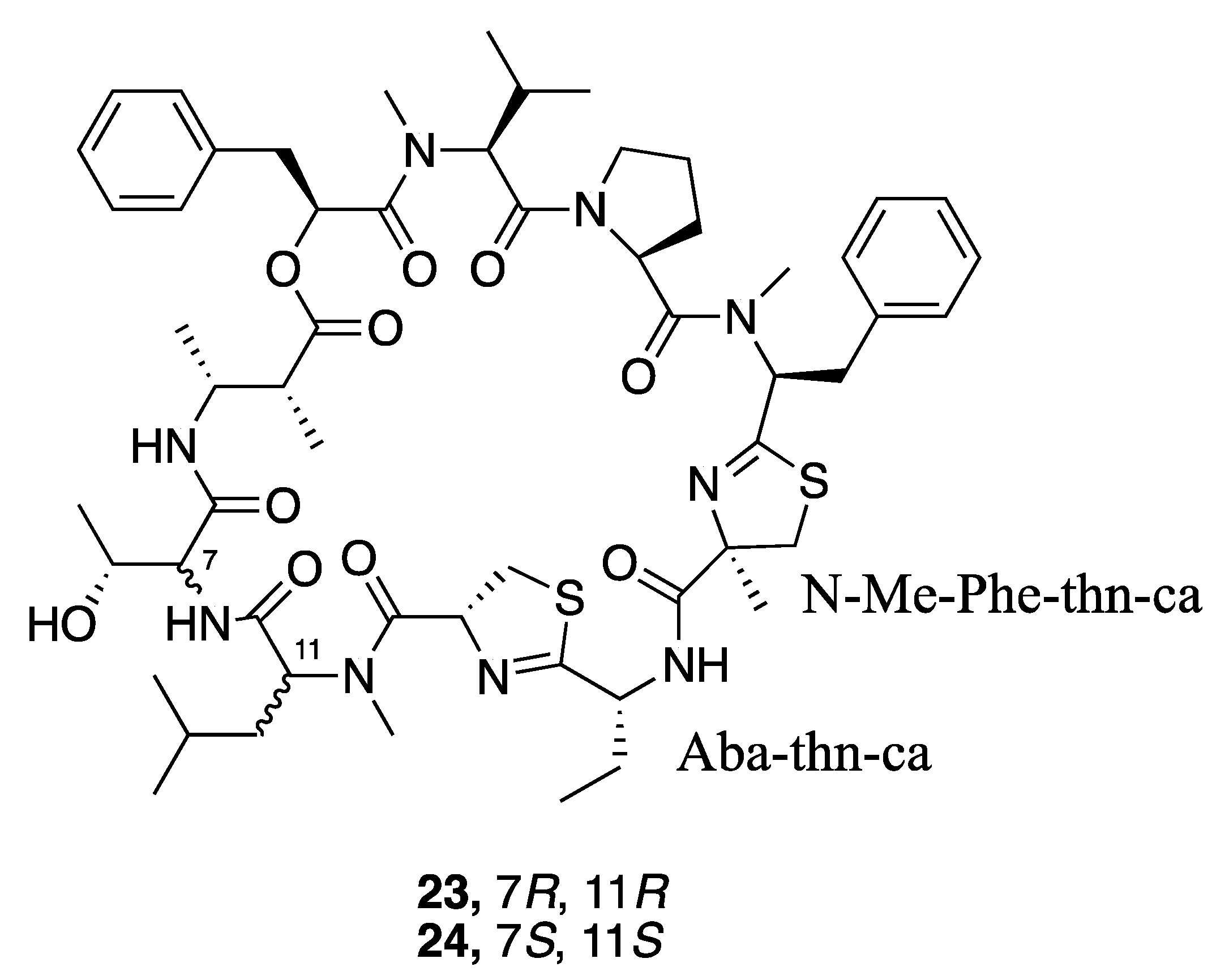
| Grassypeptolide D (23) | The Red Sea | Y b | N a | Cytotoxicity | HeLa cells | IC50 335 nM | [36] |
| Cytotoxicity | Mouse neuro2a blastoma cells | IC50 599 nM | |||||
| Grassypeptolide E (24) | Cytotoxicity | HeLa cells | IC50 192 nM | [36] | |||
| Cytotoxicity | Mouse neuro2a blastoma cells | IC50 407 nM | |||||
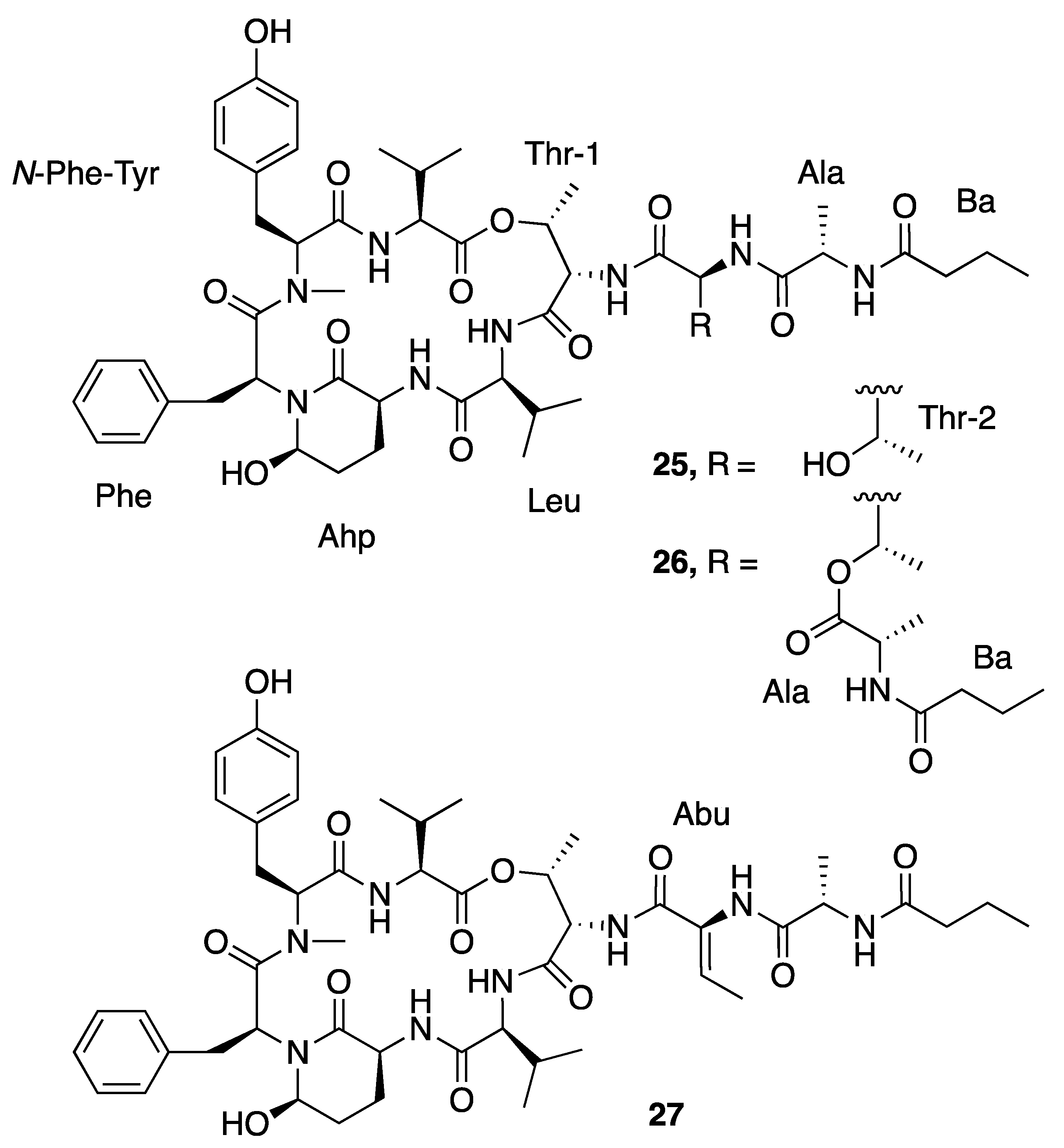
| Loggerpeptin A (25) | Florida, USA | N a | N a | Antiproteolytic Activity | Bovine pancreatic chymotrypsin | IC50 0.24 μM | [40] |
| Porcine pancreatic elastase | IC50 0.24 μM | ||||||
| Human neutrophil elastase | IC50 0.29 μM | ||||||
| Loggerpeptin B (26) | Florida, USA | N a | N a | Antiproteolytic Activity | Bovine pancreatic chymotrypsin | IC50 0.22 μM | [40] |
| Porcine pancreatic elastase | IC50 0.28 μM | ||||||
| Human neutrophil elastase | IC50 0.89 μM | ||||||
| Loggerpeptin C (27) | Florida, USA | N a | N a | Antiproteolytic Activity | Bovine pancreatic chymotrypsin | IC50 0.35 μM | [40] |
| Porcine pancreatic elastase | IC50 0.54 μM | ||||||
| Human neutrophil elastase | IC50 0.62 μM | ||||||
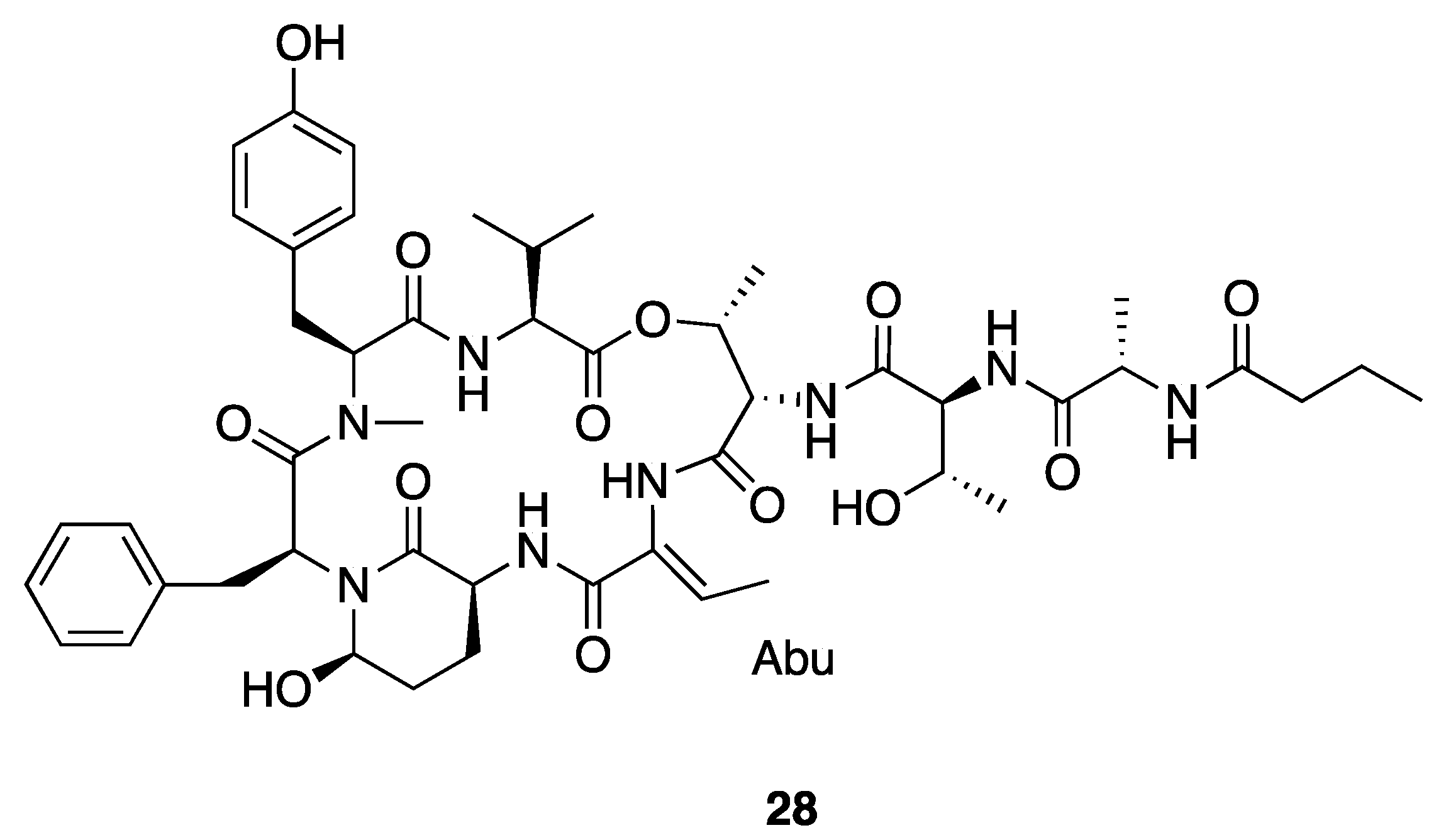
| Molassamide (28) | Florida, USA | N a | N a | Antiproteolytic Activity | Bovine pancreatic chymotrypsin | IC50 0.24 μM | [40] |
| Porcine pancreatic elastase | IC50 0.05 μM | ||||||
| Human neutrophil elastase | IC50 0.11 μM | ||||||
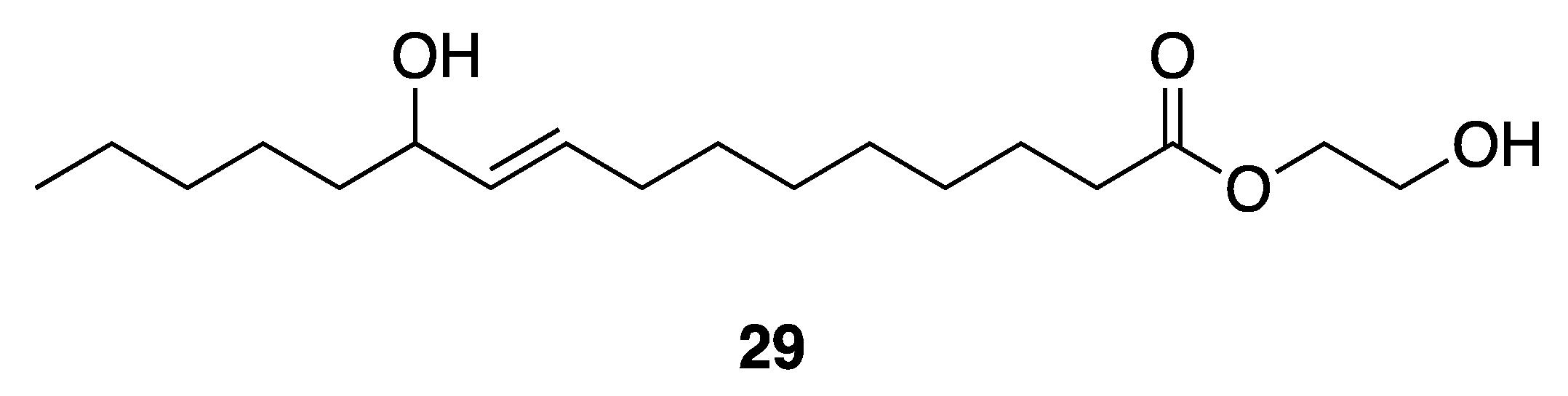
| 2-Hydroxyethyl-11-Hydroxyhexadec-9-Enoate (29) | Gulf of Thailand | Y b | N a | Antibacterial Activities | Vibrio harveyi | MIC 250–1000 µg/mL | [41] |
| Antibacterial Activities | Vibrio parahaemolyticus | MIC 350–1000 µg/mL | |||||
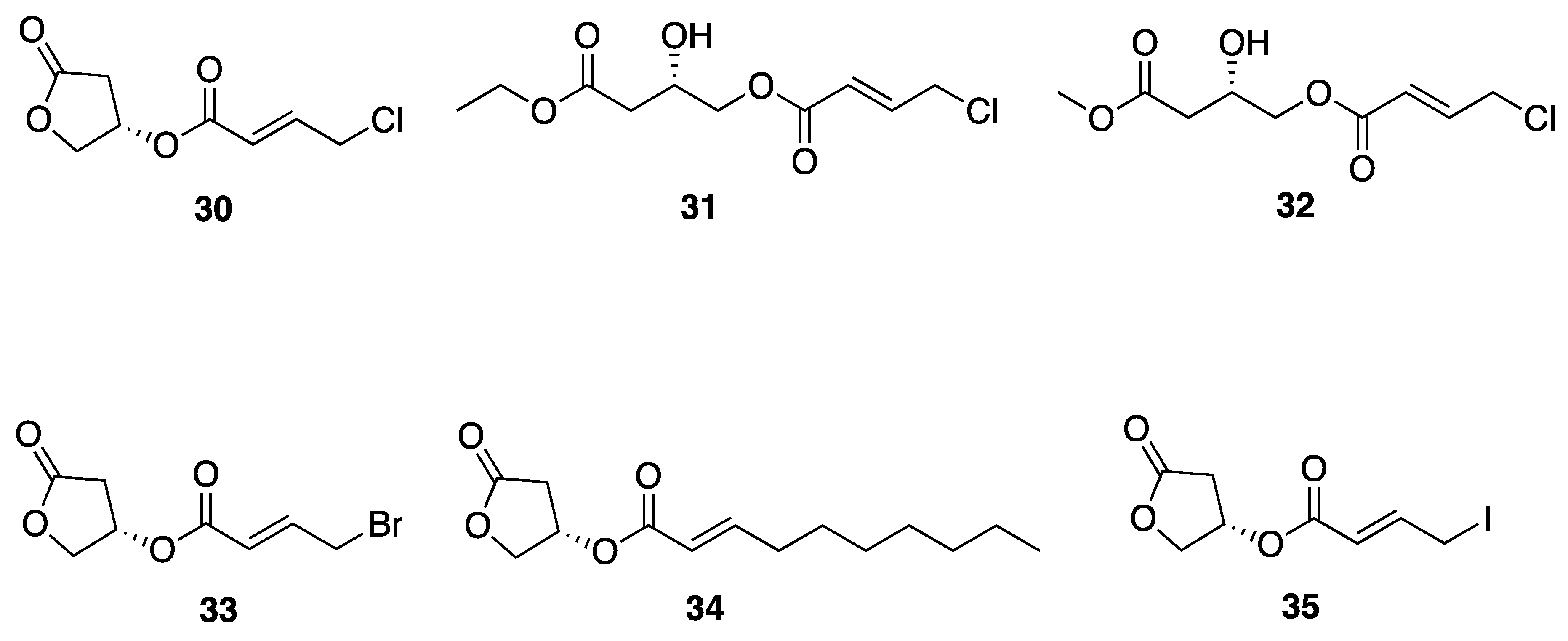
| Honaucin A (30) | Hawaii, USA | N a | N a | Anti-Inflammatory Activity | LPS-stimulated RAW264.7 murine macrophages | IC50 4.0 µM | [42] |
| Antioxidant Activity | Radical Oxygen Scavenging | No activity at 146 µM | |||||
| QS-Inhibitory activities | V. harveyi BB120 | IC50 5.6 µM | |||||
| QS-Inhibitory activities | E. coli JB525 | IC50 38.5 µM | |||||
| Cytotoxicity | RAW264.7 cells | No activity at 1 µg/mL | [43] | ||||
| Cellular TRAP Activity | RANKL-induced osteoclastogenesis in RAW264.7 cells | IC50 0.63 μg/mL | |||||
| Honaucin B (31) | Hawaii, USA | N a | N a | Anti-Inflammatory Activity | LPS-stimulated RAW264.7 murine macrophages | IC50 4.5 µM | [42] |
| QS-Inhibitory activities | V. harveyi BB120 | IC50 17.6 µM | |||||
| QS-Inhibitory activities | E. coli JB525 | IC50 > 500 µM | |||||
| Honaucin C (32) | Hawaii, USA | N a | N a | Anti-Inflammatory Activity | LPS-stimulated RAW264.7 murine macrophages | IC50 7.8 µM | [42] |
| QS-Inhibitory activities | V. harveyi BB120 | IC50 14.6 µM | |||||
| QS-Inhibitory activities | E. coli JB525 | IC50 > 500 µM | |||||
| Synthetic Br-Honaucin A (33) | N a | N a | Y b | Cytotoxicity | RAW264.7 cells | No activity at 1 µg/mL | [43] |
| Cellular TRAP Activity | RANKL-induced osteoclastogenesis in RAW264.7 cells | IC50 0.54 μg/mL | |||||
| Synthetic Hex-Honaucin A (34) | N a | N a | Y b | Cytotoxicity | RAW264.7 cells | 71.6% cell viability at 1 µg/mL | [43] |
| Cellular TRAP Activity | RANKL-induced osteoclastogenesis in RAW264.7 cells | IC50 0.68 μg/mL | |||||
| Synthetic I-Honaucin A (35) | N a | N a | Y b | Cytotoxicity | RAW264.7 cells | No activity at 1 µg/mL | [43] |
| Cellular TRAP Activity | RANKL-induced osteoclastogenesis in RAW264.7 cells | IC50 0.61 μg/mL | |||||
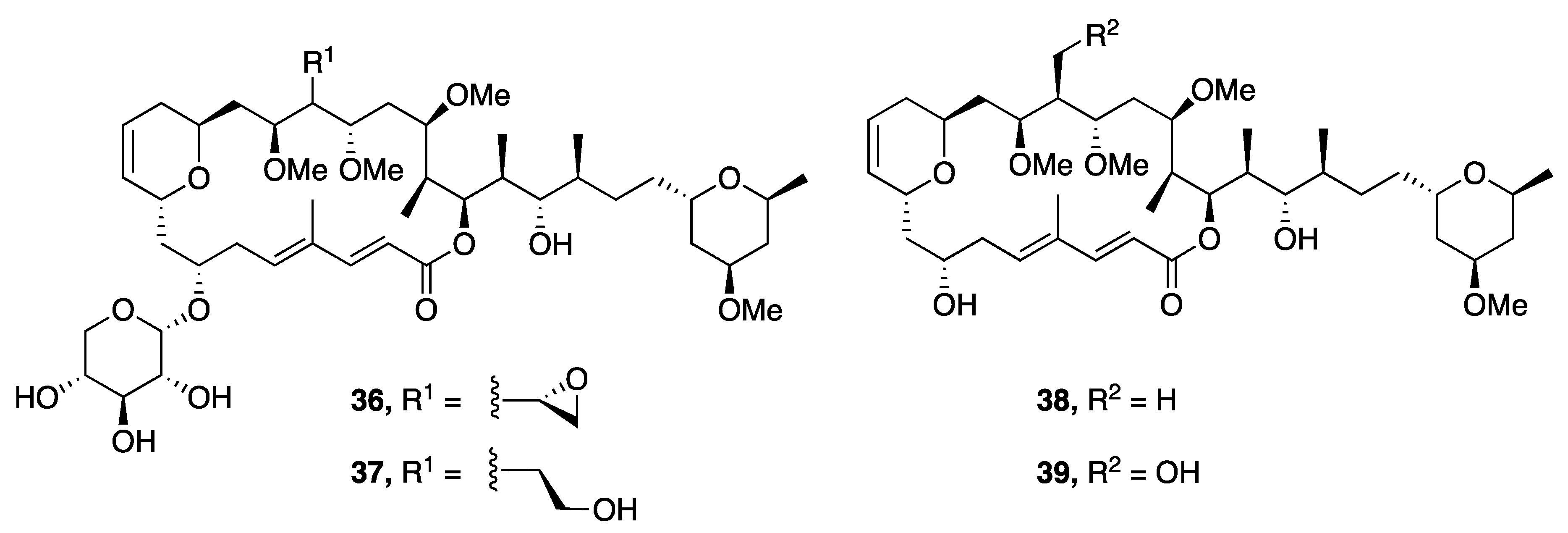
| Leptolyngbyolide A (36) | Okinawa, Japan | N a | Y b | Cytotoxicity | HeLa S3 cell | IC50 0.099 µM | [44] |
| Actin-Depolymerizing activity | F-actin | EC50 12.6 µM | |||||
| Leptolyngbyolide B (37) | Okinawa, Japan | N a | Y b | Cytotoxicity | HeLa S3 cell | IC50 0.16 µM | [44] |
| Actin-Depolymerizing activity | F-actin | EC50 11.6 µM | |||||
| Leptolyngbyolide C (38) | Okinawa, Japan | N a | Y b | Cytotoxicity | HeLa S3 cell | IC50 0.64 µM | [44] |
| Actin-Depolymerizing activity | F-actin | EC50 26.9 µM | |||||
| Leptolyngbyolide D (39) | Okinawa, Japan | N a | Y b | Cytotoxicity | HeLa S3 cell | IC50 0.15 µM | [44] |
| Actin-Depolymerizing activity | F-actin | EC50 21.5 µM | |||||
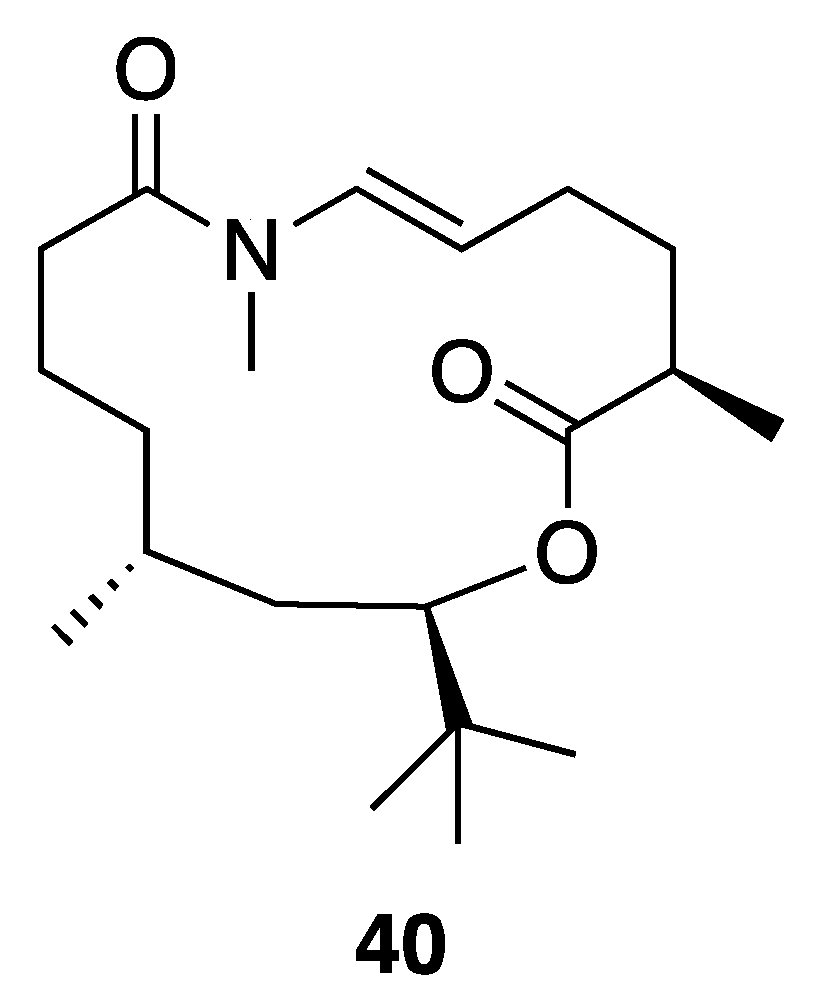
| Palmyrolide A (40) | Palmyra Atoll | N a | Y b | Ca2+ Influx (Inhibition) | Murine cerebrocortical neurons | IC50 3.70 µM (2.29–5.98 µM, 95% CI) | [45] |
| Na+ Channel Blocking Activity | Mouse neuroblastoma (neuro2a) | IC50 5.2 µM | |||||
| Cytotoxicity | H460 | No activity at 20 µM | |||||
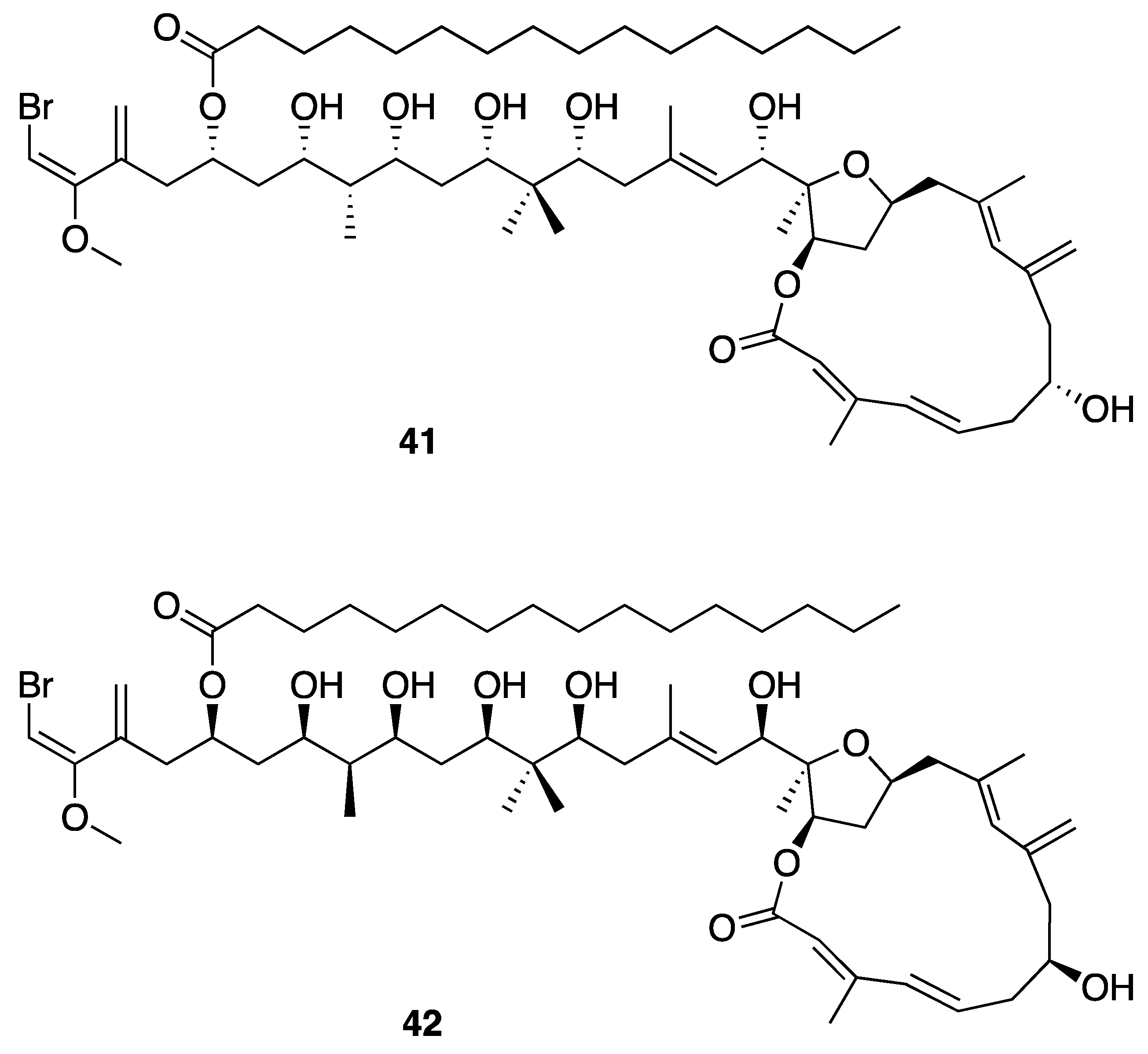
| Phormidolide (42) | The Red Sea | Y b | N a | Brine Shrimp Toxicity | LC50 1.5 µM | [46] | |
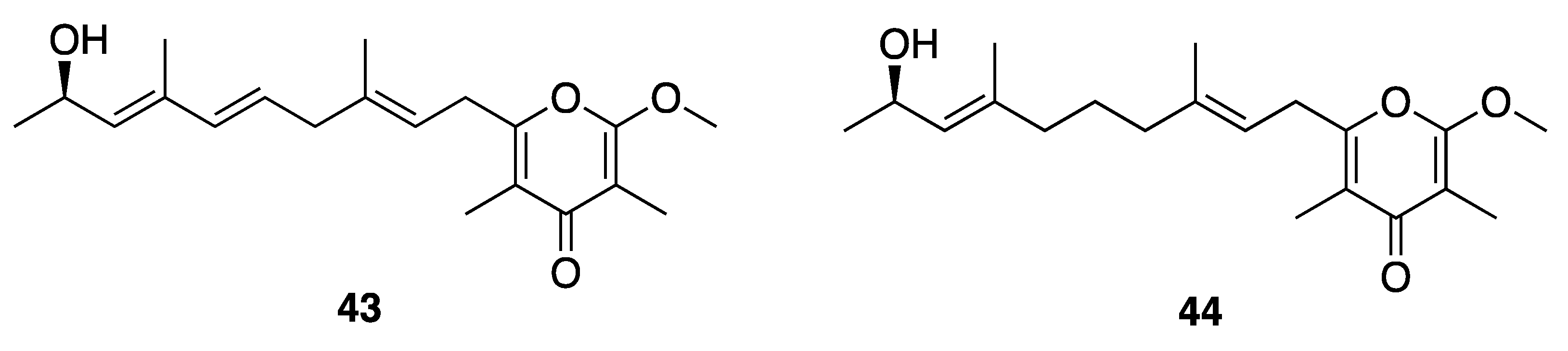
| Kalkipyrone A (43) | America Samoa | N a | N a | Cytotoxicity | H460 cells | EC50 0.9 µM | [47] |
| Cytotoxicity | Saccharomyces cerevisiae ABC16-monster | IC50 14.6 µM | |||||
| Brine Shrimp Toxicity | Brine shrimp (Artemia salina) | LD50 1 µg/mL | [48] | ||||
| Ichthyotoxicity | Goldfish Carassius auratus | LD50 2 µg/mL | |||||
| Cytotoxicity | NCI’s 60 human tumor cell line | Modestly inhibitory to several renal and melanoma cell lines | |||||
| Kalkipyrone B (44) | America Samoa | N a | N a | Cytotoxicity | H460 cells | EC50 9.0 µM | [47] |
| Cytotoxicity | Saccharomyces cerevisiae ABC16-monster | IC50 13.4 µM | |||||
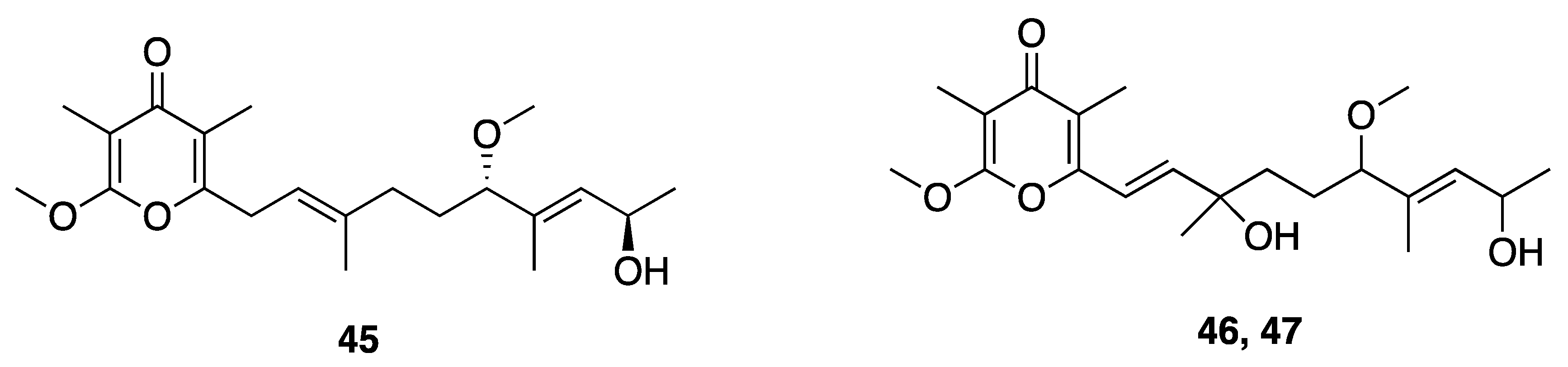
| Yoshinone A (45) | Ishigaki island, Japan | N a | Y b | Adipogenic Differentiation | 3T3-L1 cells | EC50 420 nM | [49] |
| Cytotoxicity | 3T3-L1 cells | IC50 > 50 µM | |||||
| Cytotoxicity | HeLa | IC50 > 50 µM | |||||
| Cytotoxicity | Saccharomyces cerevisiae ABC16-monster | IC50 63.8 µM | [47] | ||||
| Cytotoxicity | H460 cells | EC50 > 10 µM | |||||
| Yoshinone B1 (46) | Ishigaki island, Japan | N a | N a | Adipogenic Differentiation | 3T3-L1 cells | <50% inhibition at 5 µM | [49] |
| Yoshinone B2 (47) | Ishigaki island, Japan | N a | N a | Adipogenic Differentiation | 3T3-L1 cells | <50% inhibition at 5 µM | [49] |
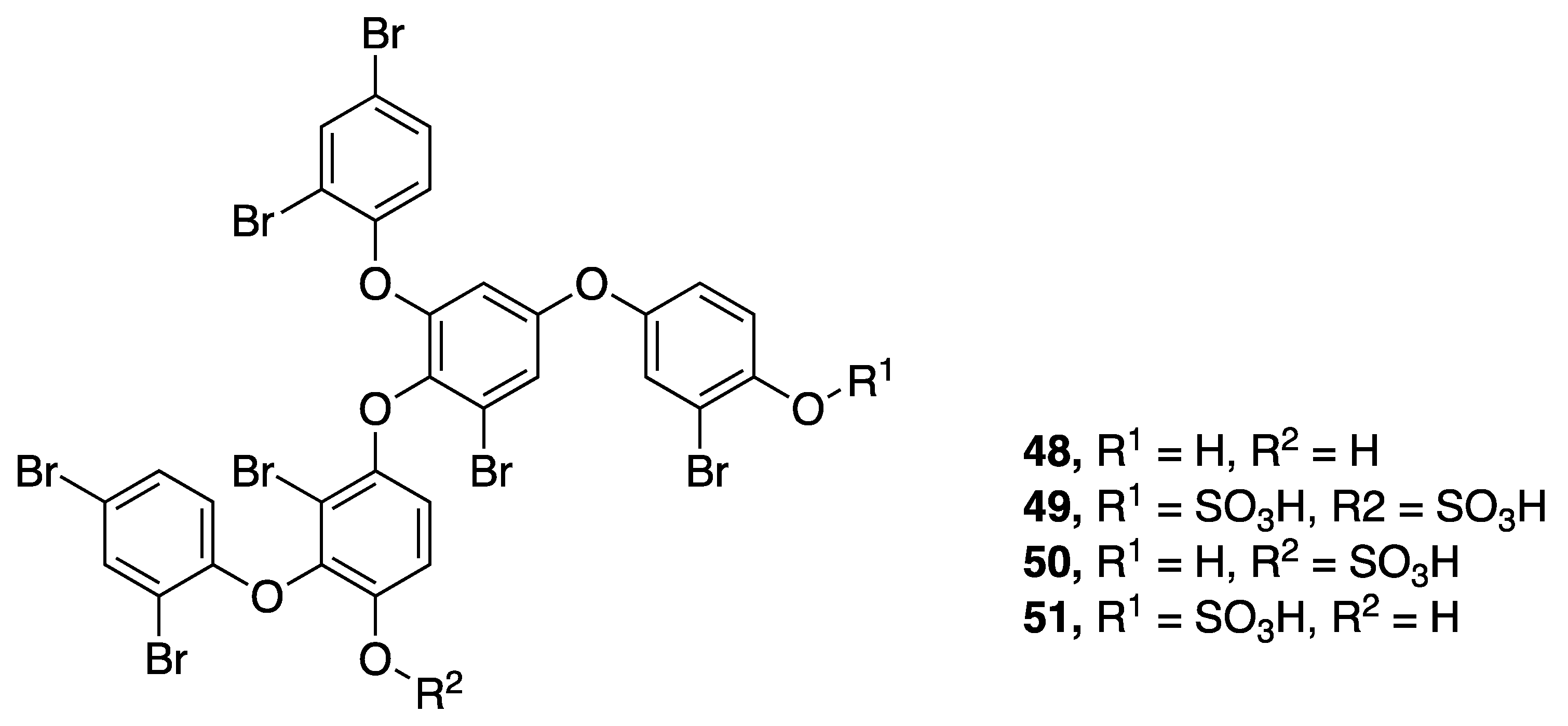
| Crossbyanol A (48) | Hawaii, USA | N a | N a | Cytotoxicity | H460 human lung cancer cells | IC50 30 µg/ mL | [50] |
| Na+ Influx (Activation and Inhibition) | Mouse neuroblastoma (neuro2a) | IC50 20 µg/mL(Activation) | |||||
| Antibacterial Activity | Methicillin-resistant Staphylococcus aureus (MRSA) | No activity at 125 µg/mL | |||||
| Brine Shrimp Toxicity | Brine shrimp (Artemia salina) | No activity at 25 µg/mL | |||||
| Crossbyanol B (49) | Hawaii, USA | N a | N a | Cytotoxicity | H460 human lung cancer cells | No activity at 20 µg/mL | [50] |
| Na+ Influx (Activation and Inhibition) | Mouse neuroblastoma (neuro2a) | No activity at 20 µg/mL | |||||
| Antibacterial activity | Methicillin-resistant Staphylococcus aureus (MRSA) | MIC 2.0–3.9 µg/mL | |||||
| Brine Shrimp Toxicity | Brine shrimp (Artemia salina) | IC50 2.8 µg/mL | |||||
| Crossbyanol C (50) | Hawaii, USA | N a | N a | Cytotoxicity | H460 human lung cancer cells | No activity at 20 µg/mL | [50] |
| Na+ Influx (Activation and Inhibition) | Mouse neuroblastoma (neuro2a) | No activity at 20 µg/mL | |||||
| Crossbyanol D (51) | Hawaii, USA | N a | N a | Cytotoxicity | H460 human lung cancer cells | No activity at 20 µg/mL | [50] |
| Na+ Influx (Activation and Inhibition) | Mouse neuroblastoma (neuro2a) | No activity at 20 µg/mL | |||||
| Leptazoline A (52) | Honolulu | Y b | N a | Cytotoxicity | PANC-1 | No significant activity | [51] |
| Leptazoline B (53) | Honolulu | Y b | N a | Cytotoxicity | PANC-1 | GI50 10 µM |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Naman, C.B.; Alexander, K.L.; Guan, H.; Gerwick, W.H. The Chemistry, Biochemistry and Pharmacology of Marine Natural Products from Leptolyngbya, a Chemically Endowed Genus of Cyanobacteria. Mar. Drugs 2020, 18, 508. https://doi.org/10.3390/md18100508
Li Y, Naman CB, Alexander KL, Guan H, Gerwick WH. The Chemistry, Biochemistry and Pharmacology of Marine Natural Products from Leptolyngbya, a Chemically Endowed Genus of Cyanobacteria. Marine Drugs. 2020; 18(10):508. https://doi.org/10.3390/md18100508
Chicago/Turabian StyleLi, Yueying, C. Benjamin Naman, Kelsey L. Alexander, Huashi Guan, and William H. Gerwick. 2020. "The Chemistry, Biochemistry and Pharmacology of Marine Natural Products from Leptolyngbya, a Chemically Endowed Genus of Cyanobacteria" Marine Drugs 18, no. 10: 508. https://doi.org/10.3390/md18100508
APA StyleLi, Y., Naman, C. B., Alexander, K. L., Guan, H., & Gerwick, W. H. (2020). The Chemistry, Biochemistry and Pharmacology of Marine Natural Products from Leptolyngbya, a Chemically Endowed Genus of Cyanobacteria. Marine Drugs, 18(10), 508. https://doi.org/10.3390/md18100508