Additional New Cytotoxic Triquinane-Type Sesquiterpenoids Chondrosterins K–M from the Marine Fungus Chondrostereum sp.
Abstract
:1. Introduction
2. Results and Discussion
2.1. Structure Elucidation
2.2. Biological Evaluation
3. Materials and Methods
3.1. General Experimental Procedures
3.2. Fungal Material
3.3. Fermentation, Extraction, and Isolation
3.4. Cytotoxic Assay
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Liang, L.F.; Guo, Y.W. Terpenes from the soft corals of the genus Sarcophyton: Chemistry and biological activities. Chem. Biodiver. 2013, 10, 2161–2196. [Google Scholar] [CrossRef] [PubMed]
- Hasan, S.; Ansari, M.I.; Mishra, M.; Ahmad, A. Major bioactive metabolites from marine fungi: A Review. Bioinformation 2015, 11, 176–181. [Google Scholar] [CrossRef] [PubMed]
- Rateb, M.E.; Ebel, R. Secondary metabolites of fungi from marine habitats. Nat. Prod. Rep. 2011, 28, 290–344. [Google Scholar] [CrossRef] [PubMed]
- Stadler, M.; Anke, T.; Dasenbrock, J. Phellodonic acid, a new biologically active hirsutane derivative from Phellodon melaleucus (Thelephoraceae, Basidiomycetes). Z. Naturforschung C 1993, 48, 545–549. [Google Scholar]
- Wang, G.Y.S.; Abrell, L.M.; Avelar, A. New hirsutane based sesquiterpenes from salt water cultures of a marine sponge-derived fungus and the terrestrial fungus Coriolus consors. Tetrahedron 1998, 54, 7335–7342. [Google Scholar] [CrossRef]
- Kupka, J.; Anke, T.; Giannetti, B.M.; Steglich, W. Isolation and biological characterization of hypnophilin, pleurotellol, and pleurotellic acid from Pleurotellus hypnophilus (Berk.) Sacc. Arch. Microbiol. 1981, 130, 223–227. [Google Scholar] [CrossRef]
- Birnbacher, J.; Schüffler, A.; Deininger, F. Isolation and biological activity of new norhirsutanes from Creolophus cirrhatus. Z. Naturforschung C 2008, 63, 203–206. [Google Scholar] [CrossRef]
- Liermann, J.C.; Schüffler, A.; Wollinsky, B. Hirsutane-type sesquiterpenes with uncommon modifications from three basidiomycetes. J. Org. Chem. 2010, 75, 2955–2961. [Google Scholar] [CrossRef] [PubMed]
- Rukachaisirikul, V.; Tansakul, C.; Saithong, S.; Pakawatchai, C.; Isaka, M.; Suvannakad, R. Hirsutane sesquiterpenes from the fungus Lentinus connatus BCC8996. J. Nat. Prod. 2005, 68, 1674–1676. [Google Scholar] [CrossRef] [PubMed]
- Li, H.J.; Xie, Y.L.; Xie, Z.L.; Chen, Y.; Lam, C.K.; Lan, W.J. Chondrosterins A–E, triquinane-type sesquiterpenoids from soft coral-associated fungus Chondrostereum sp. Mar. Drugs 2012, 10, 627–638. [Google Scholar] [CrossRef] [PubMed]
- Li, H.J.; Chen, T.; Xie, Y.L.; Chen, W.D.; Zhu, X.F.; Lan, W.J. Isolation and structure elucidation of Chondrosterins F–H from the marine fungus Chondrostereum sp. Mar. Drugs 2013, 11, 551–558. [Google Scholar] [CrossRef] [PubMed]
- Li, H.J.; Jiang, W.H.; Liang, W.L.; Huang, J.X.; Mo, Y.F.; Ding, Y.Q.; Lam, C.K.; Qian, X.J.; Zhu, X.F.; Lan, W.J. Induced marine fungus Chondrostereum sp. as a means of producing new sesquiterpenoids Chondrosterins I and J by using glycerol as the carbon source. Mar. Drugs 2014, 12, 167–175. [Google Scholar] [CrossRef] [PubMed]
- Li, H.J.; Lan, W.J.; Lam, C.K.; Yang, F.; Zhu, X.F. Hirsutane sesquiterpenoids from the marine-derived fungus Chondrostereum sp. Chem. Biodiver. 2011, 8, 317–324. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Chen, W.D.; Deng, R.; Li, D.D.; Wu, K.W.; Feng, G.K.; Li, H.J.; Zhu, X.F. Hirsutanol A induces apoptosis and autophagy via reactive oxygen species accumulation in breast cancer MCF-7 cells. J. Pharm. Sci. 2012, 119, 214–220. [Google Scholar] [CrossRef]
- Yang, F.; Chen, W.D.; Deng, R.; Zhang, H.; Tang, J.; Wu, K.W.; Li, D.D.; Feng, G.K.; Lan, W.J.; Li, H.J.; et al. Hirsutanol A, a novel sesquiterpene compound from fungus Chondrostereum sp., induces apoptosis and inhibits tumor growth through mitochondrial-independent ROS production: Hirsutanol A inhibits tumor growth through ROS production. J. Transl. Med. 2013, 11. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Gao, Y.H.; Wu, K.W.; Deng, R.; Li, D.D.; Wei, Z.X.; Jiang, S.; Wu, X.Q.; Feng, G.K.; Li, H.J.; et al. A novel sesquiterpene Hirsutanol A induces autophagical cell death in human hepatocellular carcinoma cells by increasing reactive oxygen species. Chin. J. Cancer 2010, 29, 655–660. [Google Scholar] [CrossRef] [PubMed]
- Etchri, A.; William, A.A.; Lois, M.B. Antifungal sesquiterpenoids from an arthroconidial fungus. J. Nat. Prod. 1989, 52, 1042–1054. [Google Scholar]
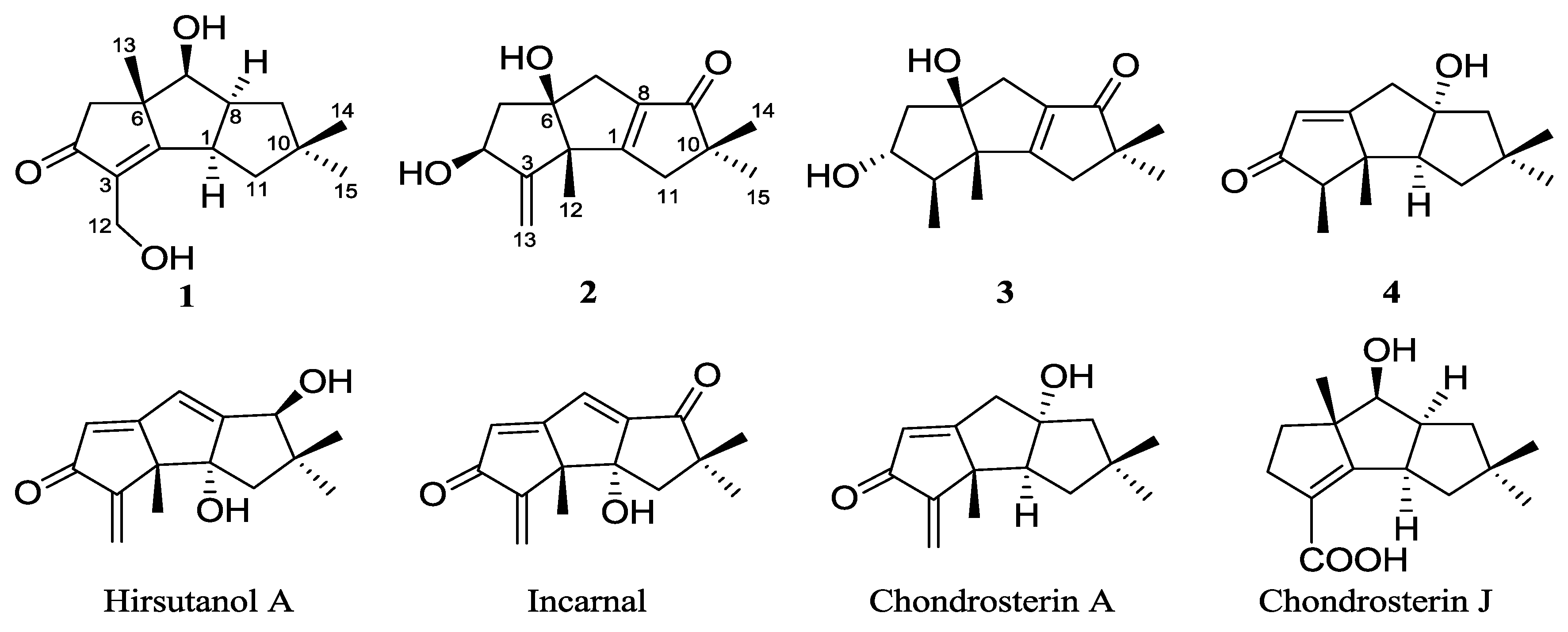
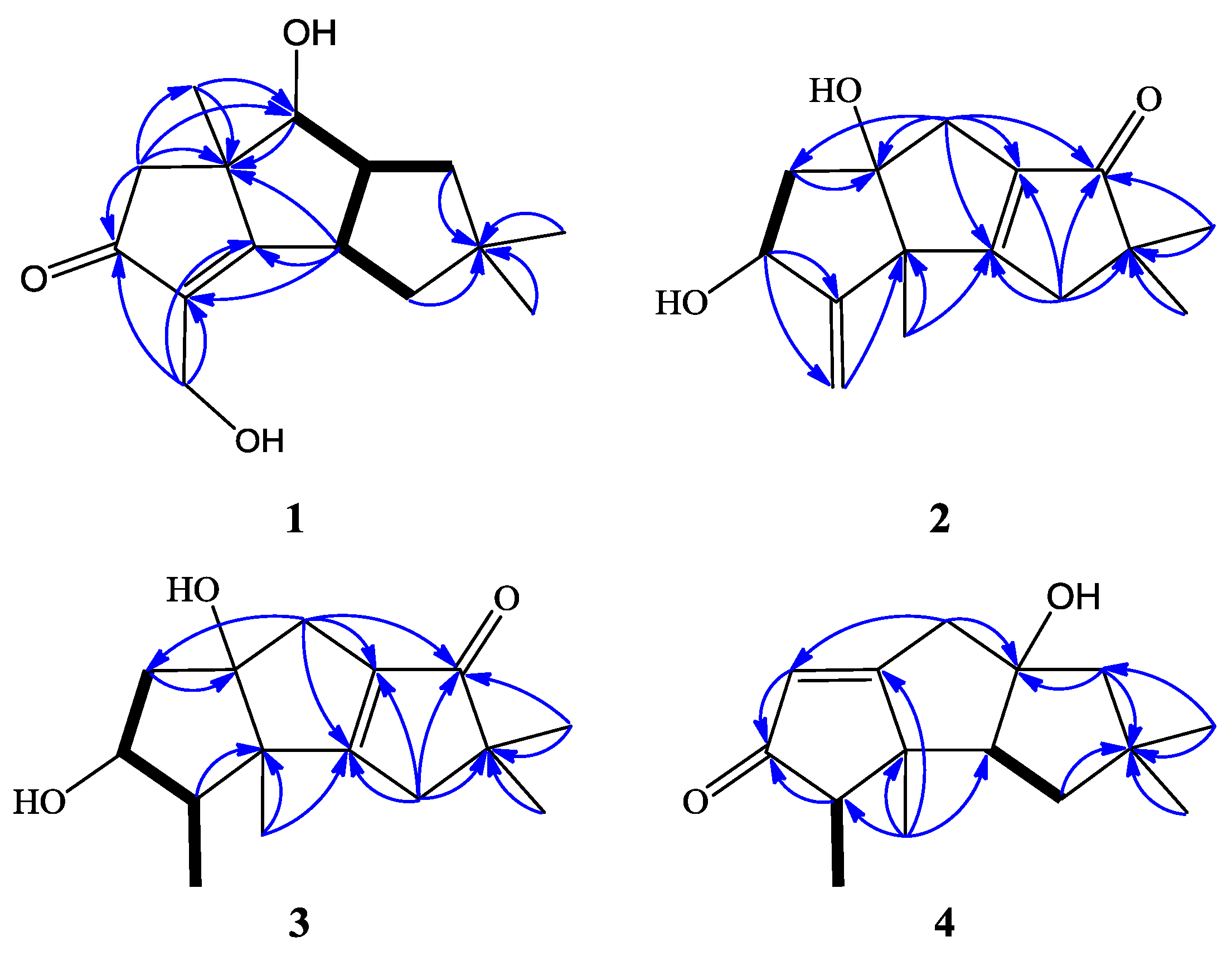
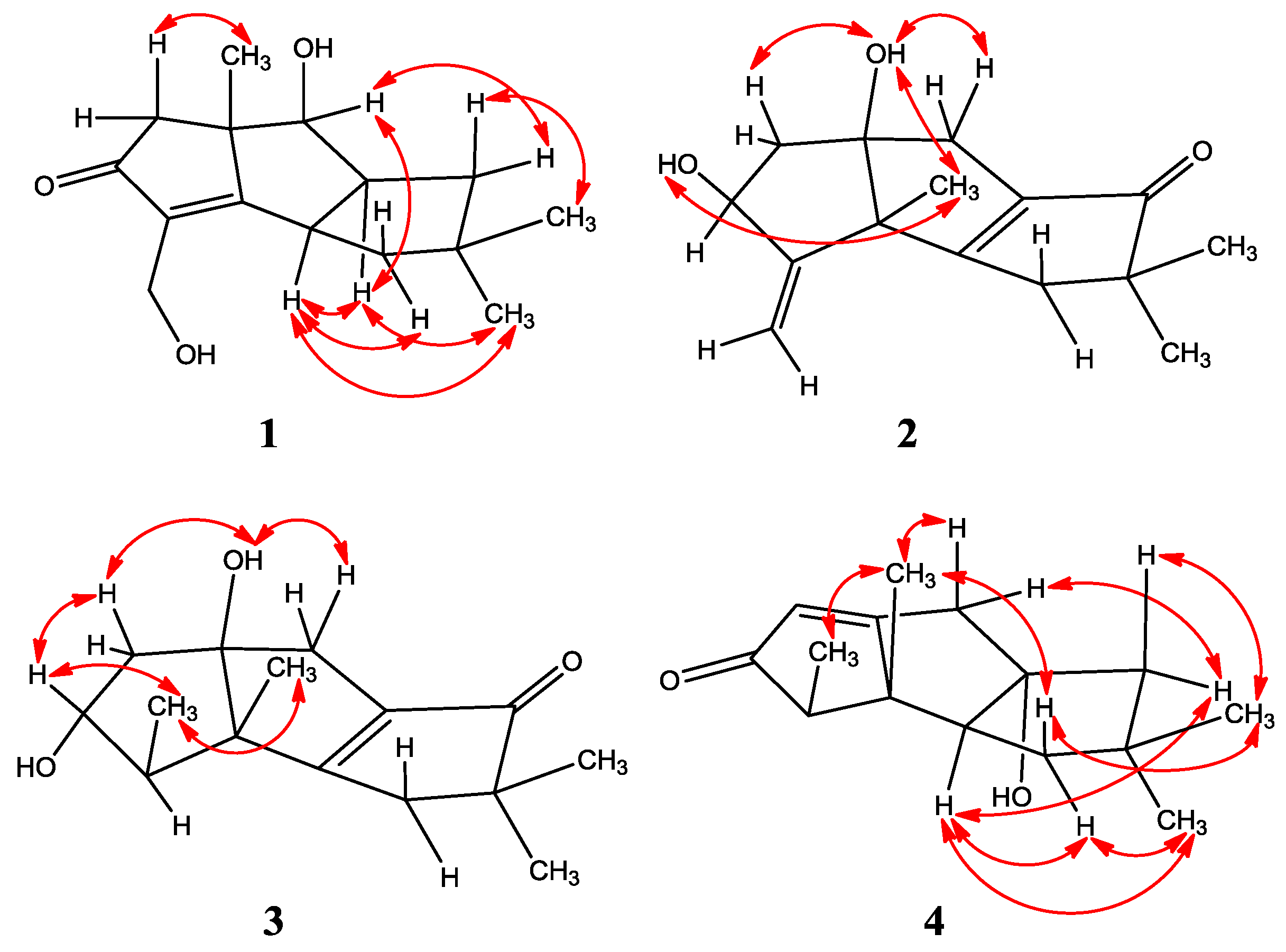
| Position | 1 a | 2 | 3 | 4 | |||
|---|---|---|---|---|---|---|---|
| In CDCl3 | In CDCl3 a | In Acetone-d6 b | In CDCl3 b | In Acetone-d6 b | In CDCl3 a | In Acetone-d6 b | |
| 1 | 42.6, CH | 184.8, C | 184.6 | 183.7, C | 184.7 | 63.4, CH | 63.6 |
| 2 | 186.4, C | 59.3, C | 59.4 | 59.1, C | 59.0 | 53.4, C | 53.9 |
| 3 | 135.2, C | 156.8, C | 158.8 | 52.1, CH | 52.8 | 57.7, CH | 58.1 |
| 4 | 210.1, C | 75.9, CH | 73.4 | 77.7, CH | 76.1 | 211.6, C | 210.8 |
| 5 | 53.3, CH2 | 46.1, CH2 | 48.7 | 48.5, CH2 | 50.6 | 122.9, CH | 122.8 |
| 6 | 52.6, C | 93.2, C | 91.4 | 91.2, C | 90.1 | 190.8, C | 192.7 |
| 7 | 77.8, CH | 36.4, CH2 | 40.2 | 40.1, CH2 | 42.0 | 43.9, CH2 | 44.5 |
| 8 | 50.2, CH | 139.1, C | 140.2 | 141.2, C | 141.9 | 92.7, C | 92.7 |
| 9 | 41.6, CH2 | 209.7, C | 208.3 | 209.1, C | 208.4 | 55.9, CH2 | 56.2 |
| 10 | 43.2, C | 48.7, C | 48.9 | 48.8, CH | 49.0 | 43.3, C | 43.7 |
| 11 | 46.1, CH2 | 39.2, CH2 | 39.4 | 42.5, CH3 | 42.8 | 41.8, CH2 | 42.3 |
| 12 | 56.1, CH2 | 19.9, CH3 | 18.4 | 19.4, CH3 | 19.0 | 20.8, CH3 | 21.1 |
| 13 | 22.6, CH3 | 110.9, CH2 | 107.4 | 14.5, CH3 | 14.0 | 9.5, CH3 | 9.7 |
| 14 | 28.5, CH3 | 25.224, CH3 | 25.5 | 25.6, CH3 | 25.8 | 30.2, CH3 | 30.5 |
| 15 | 26.8, CH3 | 25.211, CH3 | 25.4 | 25.2, CH3 | 25.4 | 28.1, CH3 | 28.4 |
| Position | 1 a | 2 | |
|---|---|---|---|
| In CDCl3 a | In Acetone-d6 b | ||
| 1 | 3.47, ddd (10.4, 9.6, 9.2) | ||
| 4 | 4.45, dd (4.4, 2.0) | 4.32, dddd (7.8, 6.0, 1.8, 1.8) | |
| 5 | 2.35, s | β: 1.87, dd (14.0, 4.4) α: 2.08, dd (14.4, 2.0) | β: 1.85, dd (12.6, 7.8) α: 2.17, dd (12.6, 6.0) |
| 7 | 4.05, d (9.2) | β: 2.43, dt (16.0, 3.2) α: 2.61, dt (16.0, 2.0) | β: 2.45, ddd (16.8, 3.6, 3.0) α: 2.54, ddd (16.8, 3.6, 2.4) |
| 8 | 3.16, dddd (9.6, 9.6, 9.2, 9.2) | ||
| 9 | β: 1.55, ddd (12.8, 9.2, 2.0) α: 1.92, dd (12.8, 9.6) | ||
| 11 | β: 1.63, dd (12.8, 10.4) α: 1.87, ddd (12.8, 9.2, 2.0) | 2.31, dd (3.2, 2.0) | β: 2.27, ddd (18.0, 3.6, 3.0) α: 2.36, ddd (18.0, 3.6, 2.4) |
| 12 | β: 4.29, d (13.6) α: 4.34, d (13.6) | 1.31, s | 1.28, s |
| 13 | 1.33, s | 5.35, s 5.10, s | 5.23, d (1.8) 5.07, d (1.8) |
| 14 | 1.16, s | 1.12, s | 1.00, s |
| 15 | 1.01, s | 1.06, s | 1.07, s |
| 4-OH | 2.97, brs | 4.16, brs | |
| 6-OH | 2.97, brs | 2.98, brs | |
| 7-OH | 2.15, brs | ||
| 12-OH | 2.15, brs | ||
| Position | 3 | 4 | ||
|---|---|---|---|---|
| In CDCl3 b | In Acetone-d6 b | In CDCl3 a | In Acetone-d6 b | |
| 1 | 2.38, dd (10.5, 8.4) | 2.42, dd (10.2, 9.0) | ||
| 3 | 1.93, dq (6.4, 7.2) | 1.76, dq (9.6, 7.2) | 2.32, q (7.2) | 2.27, q (7.2) |
| 4 | 3.63, dd (6.4, 5.6, 5.6) | 3.40, ddd (10.2, 9.6, 6.6) | ||
| 5 | β: 1.91, dd (13.2, 5.6) α: 2.28, dd (13.2, 5.6) | β: 1.82, dd (12.6, 10.2) α: 2.24, dd (12.6, 6.6) | 5.82, d (1.8) | 5.69, d (1.8) |
| 7 | β: 2.45, d (10.0) α: 2.66, d (10.0) | β: 2.45, d (10.0) α: 2.66, d (10.0) | β: 2.71, dd (15.6, 1.8) α: 2.79, d (15.6) | β: 2.75, d (15.6) α: 2.81, dd (15.6, 1.8) |
| 9 | β: 1.66, d (13.8) α: 1.86, dd (13.8, 1.2) | β: 1.70, d (13.8) α: 1.85, dd (13.8, 1.2) | ||
| 11 | β: 2.38, d (10.0) α: 2.44, d (10.0) | β: 2.39, d (10.0) α: 2.41, d (10.0) | β: 1.46, dd (12.9, 10.5) α: 1.70, ddd (12.9, 8.4, 1.2) | β: 1.51, dd (12.6, 10.2) α: 1.67, ddd (12.6, 9.0, 1.2) |
| 12 | 1.21, s | 1.16, s | 0.91, s | 0.92, s |
| 13 | 1.06, d (7.2) | 1.06, d (7.2) | 1.08, d (7.2) | 1.00, d (7.2) |
| 14 | 1.15, s | 1.08, s | 1.11, s | 1.08, s |
| 15 | 1.12, s | 1.03, s | 1.20, s | 1.19, s |
| 4-OH | 2.15, brs | 4.17, brs | ||
| 6-OH | 2.53, brs | 4.07, brs | ||
| 8-OH | 1.98, brs | 3.93, brs | ||
| Cancer Cell Lines | 1 | 2 | 3 | Hirsutanol A |
|---|---|---|---|---|
| CNE1 | 17.66 | 33.55 | 42.00 | 10.08 |
| CNE2 | 12.03 | 22.50 | 44.08 | 12.72 |
| HONE1 | 22.06 | 34.60 | 46.11 | 17.40 |
| SUNE1 | 16.44 | 30.40 | 58.83 | 3.50 |
| A549 | 23.51 | 29.67 | 49.58 | 11.96 |
| GLC82 | 18.08 | 37.47 | 55.90 | 10.11 |
| HL7702 | 22.14 | 34.26 | 56.40 | 9.76 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, L.; Lan, W.-J.; Deng, R.; Feng, G.-K.; Xu, Q.-Y.; Hu, Z.-Y.; Zhu, X.-F.; Li, H.-J. Additional New Cytotoxic Triquinane-Type Sesquiterpenoids Chondrosterins K–M from the Marine Fungus Chondrostereum sp. Mar. Drugs 2016, 14, 157. https://doi.org/10.3390/md14090157
Huang L, Lan W-J, Deng R, Feng G-K, Xu Q-Y, Hu Z-Y, Zhu X-F, Li H-J. Additional New Cytotoxic Triquinane-Type Sesquiterpenoids Chondrosterins K–M from the Marine Fungus Chondrostereum sp. Marine Drugs. 2016; 14(9):157. https://doi.org/10.3390/md14090157
Chicago/Turabian StyleHuang, Lei, Wen-Jian Lan, Rong Deng, Gong-Kan Feng, Qing-Yan Xu, Zhi-Yu Hu, Xiao-Feng Zhu, and Hou-Jin Li. 2016. "Additional New Cytotoxic Triquinane-Type Sesquiterpenoids Chondrosterins K–M from the Marine Fungus Chondrostereum sp." Marine Drugs 14, no. 9: 157. https://doi.org/10.3390/md14090157
APA StyleHuang, L., Lan, W.-J., Deng, R., Feng, G.-K., Xu, Q.-Y., Hu, Z.-Y., Zhu, X.-F., & Li, H.-J. (2016). Additional New Cytotoxic Triquinane-Type Sesquiterpenoids Chondrosterins K–M from the Marine Fungus Chondrostereum sp. Marine Drugs, 14(9), 157. https://doi.org/10.3390/md14090157







