Natural Proline-Rich Cyclopolypeptides from Marine Organisms: Chemistry, Synthetic Methodologies and Biological Status
Abstract
:1. Introduction
1.1. Natural Resources
1.2. Stability and Comparison with Linear Peptides
2. Chemistry
2.1. Structural Features
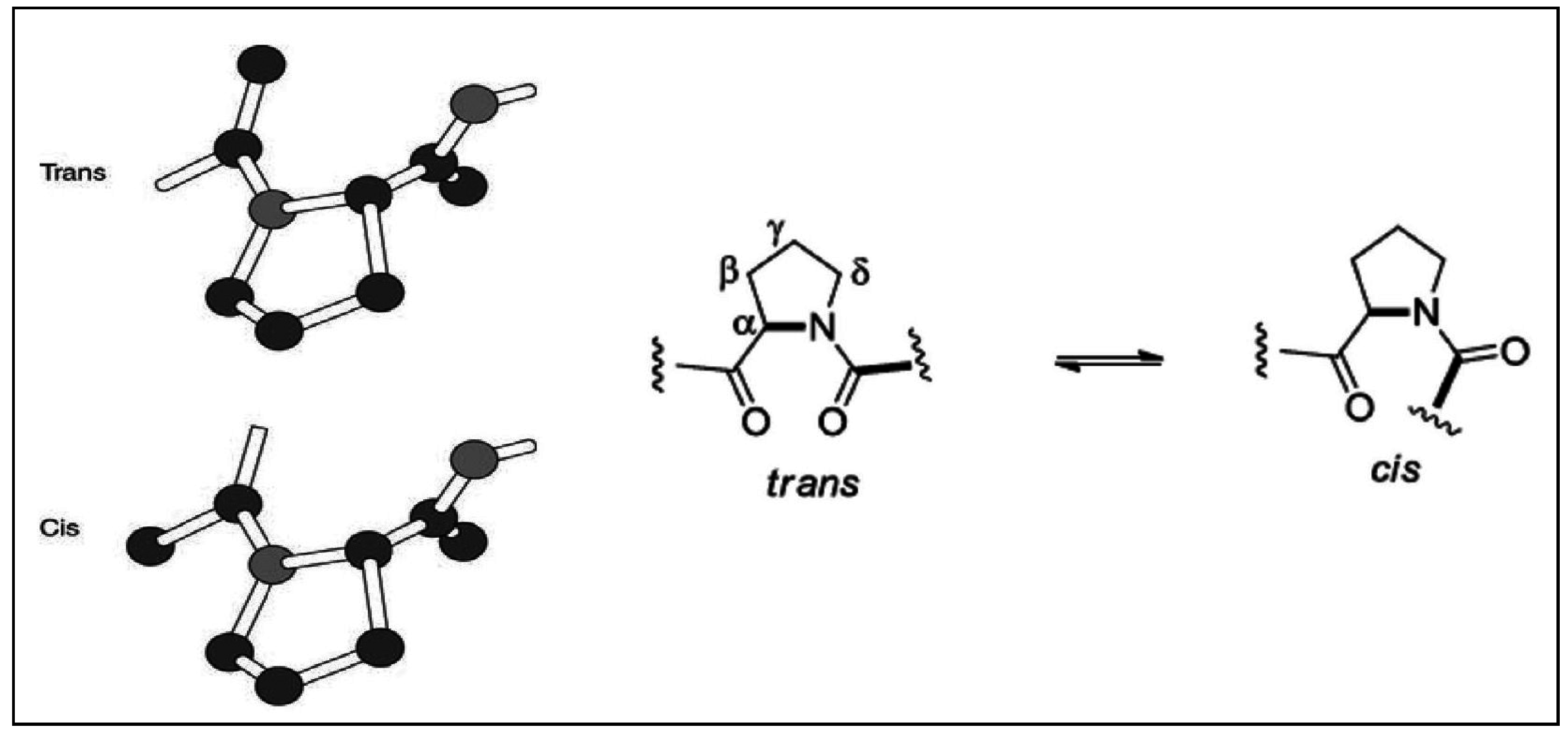
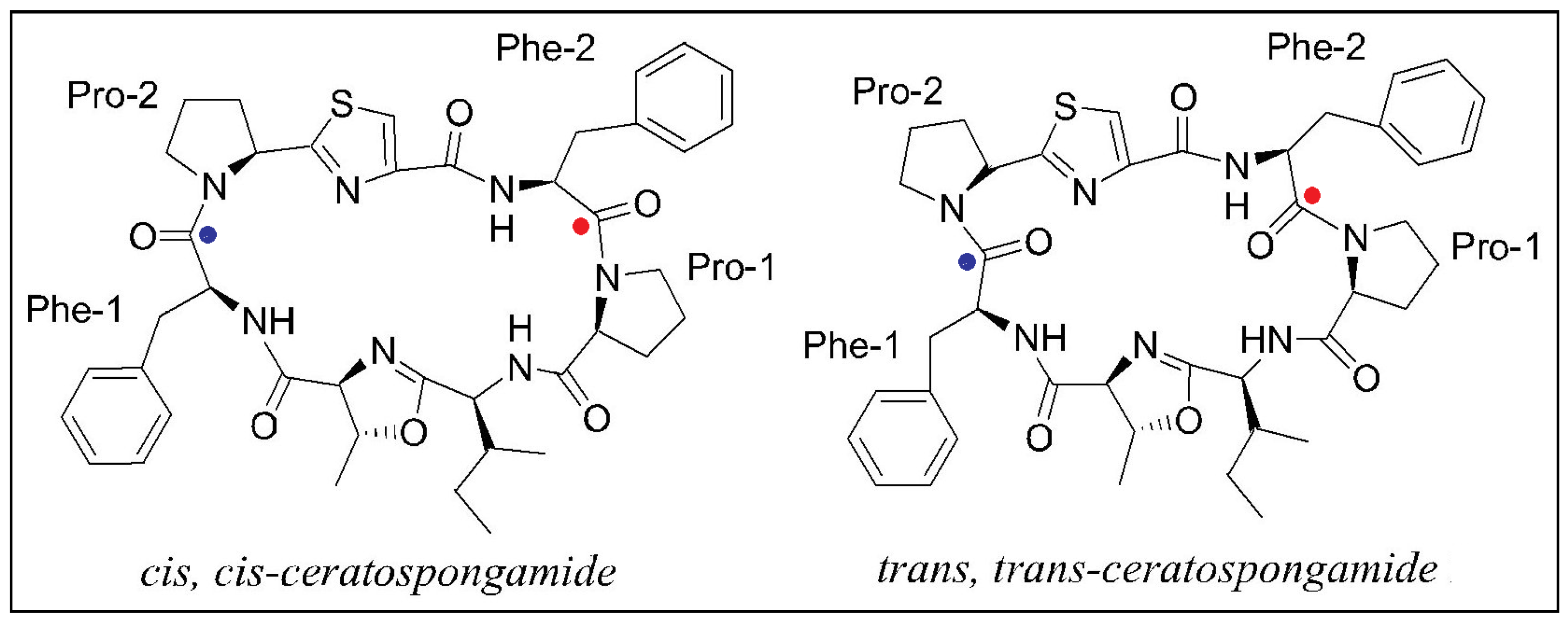
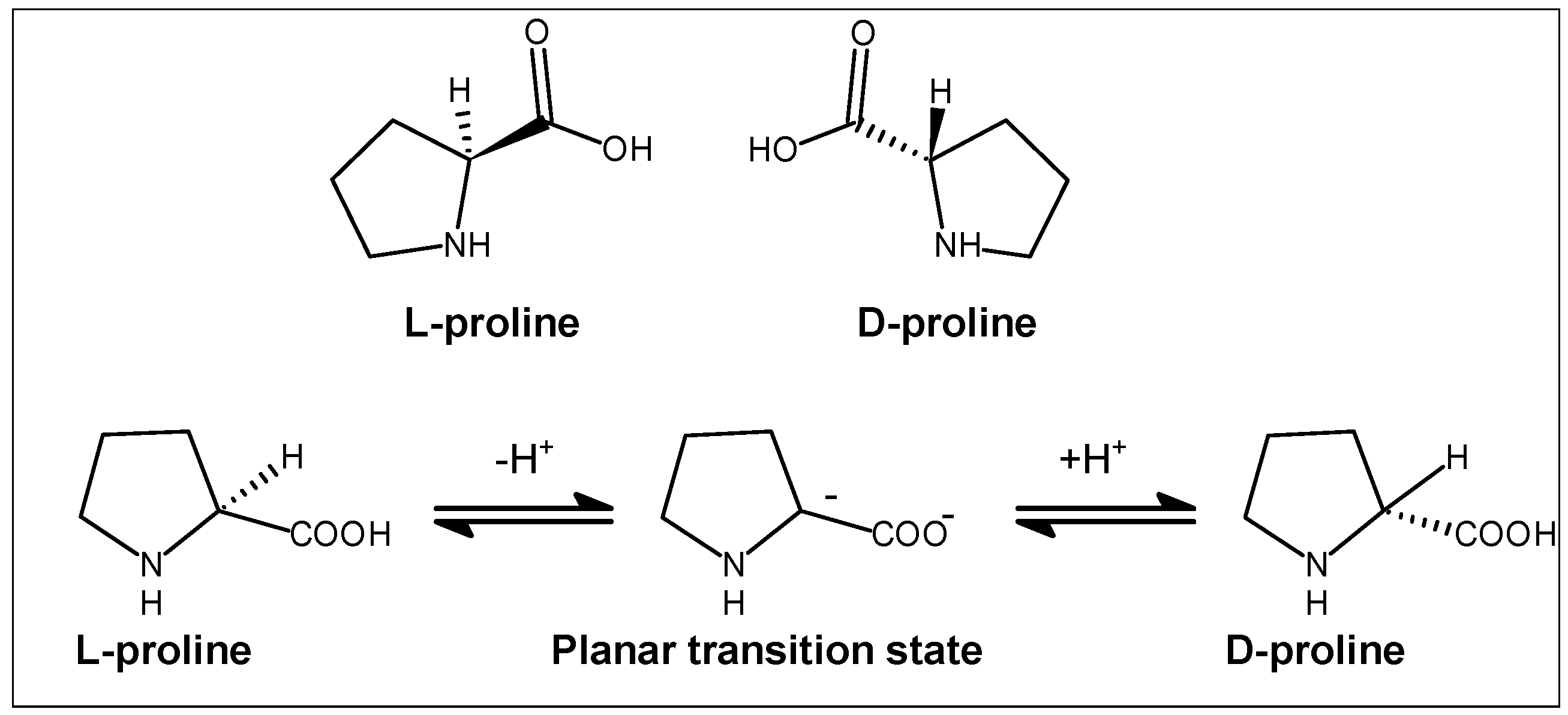
2.2. Stereochemical Aspects
2.3. Steric and Lipophilicity Parameters
2.4. Synthetic Methodologies
3. Biological Status
3.1. Mechanism of Action
3.2. Peptide Market and PRCPs in Clinical Trials
4. Conclusions and Future Prospects
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bai, R.; Friedman, S.J.; Pettit, G.R.; Hamel, E. Dolastatin 15, a potent antimitotic depsipeptide derived from Dolabella auricularia: Interaction with tubulin and effects on cellular microtubules. Biochem. Pharmacol. 1992, 43, 2637–2645. [Google Scholar] [CrossRef]
- Okamoto, S.; Iwasaki, A.; Ohno, O.; Suenaga, K. Isolation and structure of kurahyne B and total synthesis of the kurahynes. J. Nat. Prod. 2015, 78, 2719–2725. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, A.; Ohno, O.; Sumimoto, S.; Ogawa, H.; Nguyen, K.A.; Suenaga, K. Jahanyne, an apoptosis-inducing lipopeptide from the marine cyanobacterium Lyngbya sp. Org. Lett. 2015, 17, 652–655. [Google Scholar] [CrossRef] [PubMed]
- Jordan, M.A.; Walker, D.; de Arruda, M.; Barlozzari, T.; Panda, D. Suppression of microtubule dynamics by binding of cemadotin to tubulin: Possible mechanism for its antitumor action. Biochemistry 1998, 37, 17571–17578. [Google Scholar] [CrossRef] [PubMed]
- Fusetani, N.; Warabi, K.; Nogata, Y.; Nakao, Y.; Matsunaga, S.; van Soest, R.R.M. Koshikamide A1, a new cytotoxic linear peptide isolated from a marine sponge Theonella sp. Tetrahedron Lett. 1999, 40, 4687–4690. [Google Scholar] [CrossRef]
- Pettit, G.R.; Herald, C.L.; Boyd, M.R.; Leet, J.E.; Dufresne, C.; Doubek, D.L.; Schmidt, J.M.; Cerny, R.L.; Hooper, J.N.A.; Rutzler, K.C. Antineoplastic agents. 219. Isolation and structure of the cell growth inhibitory constituents from the western Pacific marine sponge Axinella sp. J. Med. Chem. 1991, 34, 3339–3340. [Google Scholar] [CrossRef] [PubMed]
- Gulavita, N.K.; Gunasekela, S.P.; Pomponi, S.A.; Robinson, E.V. Polydiscamide A: A new bioactive depsipeptide from the marine sponge Discodermia sp. J. Org. Chem. 1992, 57, 1767–1772. [Google Scholar] [CrossRef]
- Tsuda, M.; Shigemori, H.; Mikami, Y.; Kobayashi, J. Hymenamides C–E, new cyclic heptapeptides with two proline residues from the Okinawan marine sponge Hymeniacidon sp. Tetrahedron 1993, 49, 6785–6796. [Google Scholar] [CrossRef]
- Cebrat, M.; Wieczorek, Z.; Siemion, I.Z. Immunosuppressive activity of hymenistatin 1. Peptides 1996, 17, 191–196. [Google Scholar] [CrossRef]
- Tan, L.T.; Williamson, R.T.; Gerwick, W.H.; Watts, K.S.; McGough, K.; Jacobs, R. cis,cis- and trans,trans-Ceratospongamide, new bioactive cyclic heptapeptides from the indonesian red alga Ceratodictyon spongiosum and symbiotic sponge Sigmadocia symbiotica. J. Org. Chem. 2000, 65, 419–425. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Harper, M.K.; Pond, C.D.; Barrows, L.R.; Ireland, C.M.; van Wagoner, R.M. Thiazoline peptides and a tris-phenethyl urea from Didemnum molle with anti-HIV activity. J. Nat. Prod. 2012, 75, 1436–1440. [Google Scholar] [CrossRef] [PubMed]
- Sera, Y.; Adachi, K.; Fujii, K.; Shizuri, Y. A new antifouling hexapeptide from a palauan sponge, Haliclona sp. J. Nat. Prod. 2003, 66, 719–721. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, S.R.; Min, C.C.; Teuscher, F.; Ebel, R.; Kakoschke, C.; Lin, W.; Wray, V.; Edrada-Ebel, R.; Proksch, P. Callyaerins A–F and H, new cytotoxic cyclic peptides from the Indonesian marine sponge Callyspongia aerizusa. Bioorg. Med. Chem. 2010, 18, 4947–4956. [Google Scholar] [CrossRef] [PubMed]
- Vera, B.; Vicente, J.; Rodriguez, A.D. Isolation and structural elucidation of euryjanicins B–D, proline-containing cycloheptapeptides from the Caribbean marine sponge Prosuberites laughlini. J. Nat. Prod. 2009, 72, 1555–1562. [Google Scholar] [CrossRef] [PubMed]
- Berer, N.; Rudi, A.; Goldberg, I.; Benayahu, Y.; Kashman, Y. Callynormine A, a new marine cyclic peptide of a novel class. Org. Lett. 2004, 6, 2543–2545. [Google Scholar] [CrossRef] [PubMed]
- Williams, D.E.; Patrick, B.O.; Behrisch, H.W.; van soest, R.; Roberge, M.; Andersen, R.J. Dominicin, a cyclic octapeptide, and laughine, a bromopyrrole alkaloid, isolated from the Caribbean marine sponge Eurypon laughlini. J. Nat. Prod. 2005, 68, 327–330. [Google Scholar] [CrossRef] [PubMed]
- Daletos, G.; Kalscheuer, R.; Koliwer-Brandl, H.; Hartmann, R.; de Voogd, N.J.; Wray, V.; Lin, W.; Proksch, P. Callyaerins from the marine sponge Callyspongia aerizusa: Cyclic peptides with antitubercular activity. J. Nat. Prod. 2015, 78, 1910–1925. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Carroll, A.R.; Pass, D.M.; Archbold, J.K.; Avery, V.M.; Quinn, R.J. Polydiscamides B–D from a marine sponge Ircinia sp. as potent human sensory neuron-specific G protein coupled receptor agonists. J. Nat. Prod. 2008, 71, 8–11. [Google Scholar] [CrossRef] [PubMed]
- Mohammed, R.; Peng, J.; Kelly, M.; Hamann, M.T. Cyclic heptapeptides from the jamaican sponge Stylissa caribica. J. Nat. Prod. 2006, 69, 1739–1744. [Google Scholar] [CrossRef] [PubMed]
- Zhan, K.X.; Jiao, W.H.; Yang, F.; Li, J.; Wang, S.P.; Li, Y.S.; Han, B.N.; Lin, H.W. Reniochalistatins A–E, cyclic peptides from the marine sponge Reniochalina stalagmitis. J. Nat. Prod. 2014, 77, 2678–2684. [Google Scholar] [CrossRef] [PubMed]
- Randazzo, A.; Piaz, F.D.; Orrù, S.; Debitus, C.; Roussakis, C.; Pucci, P.; Gomez-Paloma, L. Axinellins A and B: New proline-containing antiproliferative cyclopeptides from the Vanuatu sponge Axinella carteri. Eur. J. Org. Chem. 1998, 11, 2659–2665. [Google Scholar] [CrossRef]
- Woo, J.K.; Jeon, J.E.; Kim, C.K.; Sim, C.J.; Oh, D.C.; Oh, K.B.; Shin, J. Gombamide A, a cyclic thiopeptide from the sponge Clathria gombawuiensis. J. Nat. Prod. 2013, 76, 1380–1383. [Google Scholar] [CrossRef] [PubMed]
- Tabudravu, J.; Morris, L.A.; Kettenes-van den Bosch, J.J.; Jaspars, M. Wainunuamide, a histidine-containing proline-rich cyclic heptapeptide isolated from the Fijian marine sponge Stylotella aurantium. Tetrahedron Lett. 2001, 42, 9273–9276. [Google Scholar] [CrossRef]
- Kita, M.; Gise, B.; Kawamura, A.; Kigoshi, H. Stylissatin A, a cyclic peptide that inhibits nitric oxide production from the marine sponge Stylissa massa. Tetrahedron Lett. 2013, 54, 6826–6828. [Google Scholar] [CrossRef]
- Zhang, H.J.; Yi, Y.H.; Yang, G.J.; Hu, M.Y.; Cao, G.D.; Yang, F.; Lin, H.W. Proline-containing cyclopeptides from the marine sponge Phakellia fusca. J. Nat. Prod. 2010, 73, 650–655. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, E.W.; Raventos-Suarez, C.; Bifano, M.; Menendez, A.T.; Fairchild, C.R.; Faulkner, D.J. Scleritodermin A, a cytotoxic cyclic peptide from the lithistid sponge Scleritoderma nodosum. J. Nat. Prod. 2004, 67, 475–478. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, R.; Omar, S.; Feliz, M.; Quinoa, E.; Riguera, R. Malaysiatin, the first cyclic heptapeptide from a marine sponge. Tetrahedron Lett. 1992, 33, 6017–6020. [Google Scholar] [CrossRef]
- Erickson, K.L.; Gustafson, K.R.; Milanowski, D.J.; Pannell, L.K.; Klose, J.R.; Boyd, M.R. Myriastramides A–C, new modified cyclic peptides from the Phillipines marine sponge Myriastra clavosa. Tetrahedron 2003, 59, 10231–10238. [Google Scholar] [CrossRef]
- Brennan, M.R.; Costello, C.E.; Maleknia, S.D.; Pettit, G.R.; Erickson, K.L. Stylopeptide 2, a proline-rich cyclodecapeptide from the sponge Stylotella sp. J. Nat. Prod. 2008, 71, 453–436. [Google Scholar] [CrossRef] [PubMed]
- Pettit, G.R.; Gao, F.; Schmidt, J.M.; Cerny, R. Isolation and structure of axinastatin 5 from a Republic of Comoros marine sponge. Bioorg. Med. Chem. Lett. 1994, 4, 2935–2940. [Google Scholar] [CrossRef]
- Kobayashi, J.; Tsuda, M.; Nakamura, T.; Mikami, Y.; Shigemori, H. Hymenamides A and B, new proline-rich cyclic heptapeptides from the okinawan marine sponge hymeniacidon sp. Tetrahedron 1993, 49, 2391–2402. [Google Scholar] [CrossRef]
- Pettit, G.R.; Cichacz, Z.; Barkoczy, J.; Dorsaz, A.C.; Herald, D.L.; Williams, M.D.; Doubek, D.L.; Schmidt, J.M.; Tackett, L.P.; Brune, D.C.; et al. Isolation and structure of the marine sponge cell growth inhibitory cyclic peptide phakellistatin 1. J. Nat. Prod. 1993, 56, 260–267. [Google Scholar] [CrossRef] [PubMed]
- Arai, M.; Yamano, Y.; Fujita, M.; Setiawan, A.; Kobayashi, M. Stylissamide X, a new proline-rich cyclic octapeptide as an inhibitor of cell migration, from an Indonesian marine sponge of Stylissa sp. Bioorg. Med. Chem. Lett. 2012, 22, 1818–1821. [Google Scholar] [CrossRef] [PubMed]
- Afifi, A.H.; El-Desoky, A.H.; Kato, H.; Mangindaan, R.E.P.; de Voogd, N.J.; Ammar, N.M.; Hifnawy, M.S.; Tsukamoto, S. Carteritins A and B, cyclic heptapeptides from the marine sponge Stylissa carteri. Tetrahedron Lett. 2016, 57, 1285–1288. [Google Scholar] [CrossRef]
- Pettit, G.R.; Clewlow, P.J.; Dufrense, C.; Doubek, D.L.; Cerny, R.L.; Rutzler, K. Antineoplastic agents. 193. Isolation and structure of the cyclic peptide hymenistatin 1. Can. J. Chem. 1990, 68, 708–711. [Google Scholar] [CrossRef]
- Vicente, J.; Vera, B.; Rodriguez, A.D.; Rodriguez-Escudero, I.; Raptis, R.G. Euryjanicin A: A new cycloheptapeptide from the Caribbean marine sponge Prosuberites laughlini. Tetrahedron Lett. 2009, 50, 4571–4574. [Google Scholar] [CrossRef] [PubMed]
- Yeung, B.K.S.; Nakao, Y.; Kinnel, R.B.; Carney, J.R.; Yoshida, W.Y.; Scheuer, P.J.; Kelly-Borges, M. The Kapakahines, cyclic peptides from the marine sponge Cribrochalina olemda. J. Org. Chem. 1996, 61, 7168–7173. [Google Scholar] [CrossRef] [PubMed]
- Rashid, M.A.; Gustafson, K.R.; Cartner, L.K.; Shigematsu, N.; Pannell, L.K.; Boyd, M.R. Microspinosamide, a new HIV-inhibitory cyclic depsipeptide from the marine sponge Sidonops microspinosa. J. Nat. Prod. 2001, 64, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Pettit, G.R.; Gao, F.; Cerny, R.L.; Doubek, D.L.; Tackett, L.P.; Schmidt, J.M.; Chapuis, J.C. Antineoplastic agents. 278. Isolation and structure of axinastatins 2 and 3 from a western Caroline Island marine sponge. J. Med. Chem. 1994, 37, 1165–1168. [Google Scholar] [CrossRef] [PubMed]
- Carroll, A.R.; Bowden, B.F.; Coll, J.C.; Hockless, D.C.R.; Skelton, B.W.; White, A.H. Studies of australian ascidians. IV. Mollamide, a cytotoxic cyclic heptapeptide from the compound ascidian Didemnum molle. Aust. J. Chem. 1994, 47, 61–69. [Google Scholar] [CrossRef]
- Tan, K.O.; Wakimoto, T.; Takada, K.; Ohtsuki, T.; Uchiyama, N.; Goda, Y.; Abe, I. Cycloforskamide, a cytotoxic macrocyclic peptide from the sea slug Pleurobranchus forskalii. J. Nat. Prod. 2013, 76, 1388–1391. [Google Scholar] [CrossRef] [PubMed]
- Whitson, E.L.; Ratnayake, A.S.; Bugni, T.S.; Harper, M.K.; Treland, C.M. Isolation, structure elucidation and synthesis of eudistomides A and B, lipopeptides from a fijian ascidian Eudistoma sp. J. Org. Chem. 2009, 74, 1156–1162. [Google Scholar] [CrossRef] [PubMed]
- Rinehart, K.L., Jr.; Gloer, J.B.; Cook, J.C., Jr.; Mizsak, S.A.; Scahill, T.A. Structures of the didemnins, antiviral and cytotoxic depsipeptides from a Caribbean tunicate. J. Am. Chem. Soc. 1981, 103, 1857–1859. [Google Scholar] [CrossRef]
- Vervoort, H.; Fenical, W. Tamandarins A and B: New cytotoxic depsipeptides from a Brazilian ascidian of the family Didemnidae. J. Org. Chem. 2000, 65, 782–792. [Google Scholar] [CrossRef] [PubMed]
- Mercader, A.G.; Duchowicz, P.R.; Sivakumar, P.M. Chemometrics Applications and Research: QSAR in Medicinal Chemistry; Apple Academic Press, Inc.: Oakville, ON, Canada, 2016; p. 278. [Google Scholar]
- Han, B.; Goeger, D.; Maier, C.S.; Gerwick, W.H. The Wewakpeptins, cyclic depsipeptides from a papua new guinea collection of the marine cyanobacterium Lyngbya semiplena. J. Org. Chem. 2005, 70, 3133–3139. [Google Scholar] [CrossRef] [PubMed]
- Lopez, J.A.V.; Al-Lihaibi, S.S.; Alarif, W.M.; Abdel-Lateff, A.; Nogata, Y.; Washio, K.; Morikawa, M.; Okino, T. Wewakazole B, a cytotoxic cyanobactin from the cyanobacterium Moorea producens collected in the red sea. J. Nat. Prod. 2016, 79, 1213–1218. [Google Scholar] [CrossRef] [PubMed]
- An, T.; Kumar, T.K.; Wang, M.; Liu, L.; Lay, J.O., Jr.; Liyanage, R.; Berry, J.; Gantar, M.; Marks, V.; Gawley, R.E.; et al. Structures of pahayokolides A and B, cyclic peptides from a Lyngbya sp. J. Nat. Prod. 2007, 70, 730–735. [Google Scholar] [CrossRef] [PubMed]
- Luo, S.; Krunic, A.; Kang, H.S.; Chen, W.L.; Woodard, J.L.; Fuchs, J.R.; Swanson, S.M.; Orjala, J. Trichormamides A and B with antiproliferative activity from the cultured freshwater cyanobacterium Trichormus sp. UIC 10339. J. Nat. Prod. 2014, 77, 1871–1880. [Google Scholar] [CrossRef] [PubMed]
- Fujii, K.; Sivonen, K.; Kashiwagi, T.; Hirayama, K.; Harada, K.I. Nostophycin, a novel cyclic peptide from the toxic cyanobacterium Nostoc sp. 152. J. Org. Chem. 1999, 64, 5777–5782. [Google Scholar] [CrossRef]
- Davies-Coleman, M.T.; Dzeha, T.M.; Gray, C.A.; Hess, S.; Pannell, L.K.; Hendricks, D.T.; Arendse, C.E. Isolation of homodolastatin 16, a new cyclic depsipeptide from a Kenyan collection of Lyngbya majuscula. J. Nat. Prod. 2003, 66, 712–715. [Google Scholar] [CrossRef] [PubMed]
- Nogle, L.M.; Gerwick, W.H. Isolation of four new cyclic depsipeptides, antanapeptins A–D, and dolastatin 16 from a madagascan collection of Lyngbya majuscula. J. Nat. Prod. 2002, 65, 21–24. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, R. Cyclopolypeptides with antifungal interest. Coll. Pharm. Commun. 2013, 1, 1–15. [Google Scholar]
- Dahiya, R.; Gautam, H. Synthesis, characterization and biological evaluation of cyclomontanin D. Afr. J. Pharm. Pharmacol. 2011, 5, 447–453. [Google Scholar] [CrossRef]
- Dahiya, R.; Gautam, H. Synthetic and pharmacological studies on a natural cyclopeptide from Gypsophila arabica. J. Med. Plant Res. 2010, 4, 1960–1966. [Google Scholar]
- Dahiya, R.; Singh, S. Synthesis, characterization and biological screening of diandrine A. Acta Pol. Pharm. 2016. submitted. [Google Scholar]
- Dahiya, R.; Gautam, H. Solution phase synthesis and bioevaluation of cordyheptapeptide B. Bull. Pharm. Res. 2011, 1, 1–10. [Google Scholar]
- Dahiya, R. Synthesis of a phenylalanine-rich peptide as potential anthelmintic and cytotoxic agent. Acta Pol. Pharm. 2007, 64, 509–516. [Google Scholar] [PubMed]
- Dahiya, R.; Gautam, H. Toward the first total synthesis of gypsin D: A natural cyclopolypeptide from Gypsophila arabica. Am. J. Sci. Res. 2010, 11, 150–158. [Google Scholar]
- Dahiya, R.; Kaur, K. Synthesis and pharmacological investigation of segetalin C as a novel antifungal and cytotoxic agent. Arzneimittelforschung 2008, 58, 29–34. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, R. Synthetic and pharmacological studies on longicalycinin A. Pak. J. Pharm. Sci. 2007, 20, 317–323. [Google Scholar] [PubMed]
- Dahiya, R.; Kumar, A. Synthetic and biological studies on a cyclopolypeptide of plant origin. J. Zhejiang Univ. Sci. B 2008, 9, 391–400. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, R.; Gautam, H. Synthesis and pharmacological studies on a cyclooligopeptide from marine bacteria. Chin. J. Chem. 2011, 29, 1911–1916. [Google Scholar]
- Dahiya, R. Synthesis, characterization and biological evaluation of a glycine-rich peptide—Cherimolacyclopeptide E. J. Chil. Chem. Soc. 2007, 52, 1224–1229. [Google Scholar] [CrossRef]
- Dahiya, R.; Gautam, H. Toward the synthesis and biological screening of a cyclotetrapeptide from marine bacteria. Mar. Drugs 2011, 9, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, R.; Maheshwari, M.; Yadav, R. Synthetic and cytotoxic and antimicrobial activity studies on annomuricatin B. Z. Naturforsch. 2009, 64, 237–244. [Google Scholar] [CrossRef]
- Aneiros, A.; Garateix, A. Bioactive peptides from marine sources: Pharmacological properties and isolation procedures. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2004, 803, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Silver, F.H. Mechanosensing and Mechanochemical Transduction in Extracellular Matrix. Biochemical, Chemical, Engineering, and Physiological Aspects. Macromolecular Structures in Tissues; Springer: Berlin/Heidelberg, Germany, 2006; Volume XVI, p. 33. [Google Scholar]
- Pandey, A.K.; Naduthambi, D.; Thomas, K.M.; Zondlo, N.J. Proline editing: A general and practical approach to the synthesis of functionally and structurally diverse peptides. Analysis of steric versus stereoelectronic effects of 4-substituted prolines on conformation within peptides. J. Am. Chem. Soc. 2013, 135, 4333–4363. [Google Scholar] [CrossRef] [PubMed]
- Roxin, A.; Zheng, G. Flexible or fixed: A comparative review of linear and cyclic cancer-targeting peptides. Future Med. Chem. 2012, 4, 1601–1618. [Google Scholar] [CrossRef] [PubMed]
- Goodwin, D.; Simerska, P.; Toth, I. Peptides as therapeutics with enhanced bioactivity. Curr. Med. Chem. 2012, 19, 4451–4461. [Google Scholar] [CrossRef] [PubMed]
- Jensen, J.E.; Mobli, M.; Brust, A.; Alewood, P.F.; King, G.F.; Rash, L.D. Cyclisation increases the stability of the sea anemone peptide APETx2 but decreases its activity at acid-sensing ion channel 3. Mar. Drugs 2012, 10, 1511–1527. [Google Scholar] [CrossRef] [PubMed]
- Roxin, A. Towards Targeted Photodynamic Therapy: Synthesis and Characterization of Aziridine Aldehyde-Cyclized Cancertargeting Peptides and Bacteriochlorin Photosensitizers. Ph.D. Thesis, Graduate Department of Pharmaceutical Sciences, University of Toronto, Toronto, ON, Canada, 2014. [Google Scholar]
- Fosgerau, K.; Hoffmann, T. Peptide therapeutics: Current status and future directions. Drug Discov. Today 2015, 20, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Shanmugam, S.; Kumar, S.T.; Selvam, K.P. Laboratory Handbook on Biochemistry, 1st ed.; Prentice-Hall of India Private Limited: New Delhi, India, 2010. [Google Scholar]
- Pettit, G.R.; Gao, F.; Cerny, R. Isolation and structure of axinastatin 4 from the western indian ocean marine sponge Axinella cf. carteri. Heterocycles 1993, 35, 711–718. [Google Scholar] [CrossRef]
- Kawagishi, H.; Somoto, A.; Kuranari, J.; Kimura, A.; Chiba, S. A novel cyclotetrapeptide produced by Lactobacillus helveticus as a tyrosinase inhibitor. Tetrahedron Lett. 1993, 34, 3439–3440. [Google Scholar] [CrossRef]
- Pergament, I.; Carmeli, S. Schizotrin A; a novel antimicrobial cyclic peptide from a cyanobacterium. Tetrahedron Lett. 1994, 35, 8473–8476. [Google Scholar] [CrossRef]
- Pettit, G.R.; Srirangam, J.K.; Herald, D.L.; Xu, J.P.; Boyd, M.R.; Cichacz, Z.; Kamano, Y.; Schmidt, J.M.; Erickson, K.L. Isolation and crystal structure of stylopeptide 1, a new marine porifera cycloheptapeptide. J. Org. Chem. 1995, 60, 8257–8261. [Google Scholar] [CrossRef]
- Carroll, A.R.; Coll, J.C.; Bourne, J.C.; MacLeod, J.K.; Zanriskie, T.M.; Ireland, C.M.; Bowden, B.F. Patellins 1-6 and Trunkamide A: Novel cyclic hexa-, hepta- and octa-peptides from colonial ascidians, Lissoclinurn sp. Aust. J. Chem. 1996, 49, 659–667. [Google Scholar]
- Kobayashi, J.; Nakamura, T.; Tsuda, M. Hymenamide F, new cyclic heptapeptide from marine sponge Hymeniacidon sp. Tetrahedron 1996, 52, 6355–6360. [Google Scholar] [CrossRef]
- Shin, H.J.; Matsuda, H.; Murakami, M.; Yamaguchi, K. Agardhipeptins A and B, two new cyclic hepta- and octapeptide, from the cyanobacterium Oscillatoria agardhii (NIES-204). Tetrahedron 1996, 52, 13129–13136. [Google Scholar] [CrossRef]
- Belofsky, G.N.; Gloer, J.B.; Wicklow, D.T.; Dowd, P.F. Shearamide A: A new cyclic peptide from the ascostromata of Eupenicillium shearii. Tetrahedron Lett. 1998, 39, 5497–5500. [Google Scholar] [CrossRef]
- Murakami, M.; Itou, Y.; Ishida, K.; Shin, H.J. Prenylagaramides A and B, new cyclic peptides from two strains of Oscillatoria agardhii. J. Nat. Prod. 1999, 62, 752–755. [Google Scholar] [CrossRef] [PubMed]
- Milanowski, D.J.; Rashid, M.A.; Gustafson, K.R.; O’Keefe, B.R.; Nawrocki, J.P.; Pannell, L.K.; Boyd, M.R. Cyclonellin, a new cyclic octapeptide from the marine sponge Axinella carteri. J. Nat. Prod. 2004, 67, 441–444. [Google Scholar] [CrossRef] [PubMed]
- Leikoski, N.; Fewer, D.P.; Jokela, J.; Wahlsten, M.; Rouhiainen, L.; Sivonen, K. Highly diverse cyanobactins in strains of the genus Anabaena. Appl. Environ. Microbiol. 2010, 76, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.X.; Zhou, L.L.; Yan, Y.M.; Chen, K.X.; Hou, F.F. Diabetic nephropathy-related active cyclic peptides from the roots of Brachystemma calycinum. Bioorg. Med. Chem. Lett. 2011, 21, 7334–7439. [Google Scholar] [CrossRef] [PubMed]
- Aviles, E.; Rodriguez, A.D. Euryjanicins E–G, poly-phenylalanine and poly-proline cyclic heptapeptides from the Caribbean sponge Prosuberites laughlini. Tetrahedron 2013, 69, 10797–10804. [Google Scholar] [CrossRef] [PubMed]
- Pettil, G.R.; Tan, R.; Williams, M.D.; Tackett, L.; Schmidt, J.M.; Cerny, R.L.; Hooper, J.N.A. Isolation and structure of phakellistatin 2 from the eastern indian ocean marine sponge phakellia carteri. Bioorg. Med. Chem. Lett. 1993, 3, 2869–2874. [Google Scholar] [CrossRef]
- Tsuda, M.; Sasaki, T.; Kobayashi, J. Hymenamides G, H, J, and K, four new cyclic octapeptides from the Okinawan marine sponge Hymeniacidon sp. Tetrahedron 1994, 50, 4667–4680. [Google Scholar] [CrossRef]
- Pettit, G.R.; Tan, R.; Ichihara, Y.; Williams, M.D.; Doubek, D.L.; Tackett, L.P.; Schmidt, J.M.; Cerny, R.L.; Boyd, M.R.; Hooper, J.N. Antineoplastic agents, 325. Isolation and structure of the human cancer cell growth inhibitory cyclic octapeptides phakellistatin 10 and 11 from Phakellia sp. J. Nat. Prod. 1995, 58, 961–965. [Google Scholar] [CrossRef] [PubMed]
- Rashid, M.A.; Gustafson, K.R.; Boswell, J.L.; Boyd, M.R. Haligramides A and B, two new cytotoxic hexapeptides from the marine sponge Haliclona nigra. J. Nat. Prod. 2000, 63, 956–959. [Google Scholar] [CrossRef] [PubMed]
- Guan, L.L.; Sera, Y.; Adachi, K.; Nishida, F.; Shizuri, Y. Isolation and evaluation of nonsiderophore cyclic peptides from marine sponges. Biochem. Biophy. Res. Commun. 2001, 283, 976–981. [Google Scholar] [CrossRef] [PubMed]
- Tabudravu, J.N.; Morris, L.A.; Kettenes-van den Bosch, J.J.; Jaspars, M. Axinellin C, a proline-rich cyclic octapeptide isolated from the Fijian marine sponge Stylotella aurantium. Tetrahedron 2002, 58, 7863–7868. [Google Scholar] [CrossRef]
- Sera, Y.; Adachi, K.; Fujii, K.; Shizuri, Y. Isolation of haliclonamides: New peptides as antifouling substances from a marine sponge species, Haliclona. Mar. Biotechnol. 2002, 4, 441–446. [Google Scholar] [CrossRef] [PubMed]
- Nogle, L.M.; Marquez, B.L.; Gerwick, W.H. Wewakazole, a novel cyclic dodecapeptide from a papua new guinea Lyngbya majuscule. Org. Lett. 2003, 5, 3–6. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Cheng, W.; de Voogd, N.J.; Proksch, P.; Lin, W. Stylissatins B–D, cycloheptapeptides from the marine sponge Stylissa massa. Tetrahedron Lett. 2016, in press. [Google Scholar]
- Wieland, T.; Luben, G.; Ottenheym, H.; Faesel, D.C.J.; de Vries, J.X.; Prox, A.; Schmid, D.C.J. The discovery, isolation, elucidation of structure, and synthesis of antamanide. Angew. Chem. Int. Ed. 1968, 7, 204–208. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, S.R.M.; Edrada-Ebel, R.A.; Mohamed, G.A.; Youssef, D.T.A.; Wray, V.; Proksch, P. Callyaerin G, a new cytotoxic cyclic peptide from the marine sponge Callyspongia aerizusa. ARKIVOC Arch. Org. Chem. 2008, 2008, 164–171. [Google Scholar]
- Pettit, G.R.; Tan, R.; Herald, D.L.; Cerny, R.L.; Williams, M.D. Antineoplastic agents. 277. Isolation and structure of phakellistatin 3 and isophakellistatin 3 from a republic of Comoros marine sponge. J. Org. Chem. 1994, 59, 1593–1595. [Google Scholar] [CrossRef]
- Martins, J.; Vasconcelos, V. Cyanobactins from cyanobacteria: Current genetic and chemical state of knowledge. Mar. Drugs 2015, 13, 6910–6946. [Google Scholar] [CrossRef] [PubMed]
- Donia, M.S.; Ravel, J.; Schmidt, E.W. A global assembly line to cyanobactins. Nat. Chem. Biol. 2008, 4, 341–343. [Google Scholar] [CrossRef] [PubMed]
- Mojsoska, B.; Jenssen, H. Peptides and peptidomimetics for antimicrobial drug design. Pharmaceuticals 2015, 8, 366–415. [Google Scholar] [CrossRef] [PubMed]
- Wedemeyer, W.J.; Welker, E.; Scheraga, H.A. Proline cis-trans isomerization and protein folding. Biochemistry 2002, 41, 14637–14644. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, P.; Reichman, C.; Saleh, T.; Birge, R.B.; Kalodimos, C.G. Proline cis-trans isomerization controls autoinhibition of a signaling protein. Mol. Cell 2007, 25, 413–426. [Google Scholar] [CrossRef] [PubMed]
- Vitagliano, L.; Berisio, R.; Mastrangelo, A.; Mazzarella, L.; Zagari, A. Preferred proline puckerings in cis and trans peptide groups: Implications for collagen stability. Protein Sci. 2001, 10, 2627–2632. [Google Scholar] [CrossRef] [PubMed]
- Bhushan, R.; Bruckner, H. Marfey’s reagent for chiral amino acid analysis: A review. Amino Acids 2004, 27, 231–247. [Google Scholar] [CrossRef] [PubMed]
- Anand, M.; Alagar, M.; Ranjitha, J.; Selvaraj, V. Total synthesis and anticancer activity of a cyclic heptapeptide from marine sponge using water soluble peptide coupling agent EDC. Arab. J. Chem. 2016, in press. [Google Scholar]
- Shinde, N.V.; Himaja, M.; Bhosale, S.K.; Ramana, M.V.; Sakarkar, D.M. Synthesis and biological evaluation of delavayin-C. Indian J. Pharm. Sci. 2008, 70, 827–831. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, R. Synthesis, spectroscopic and biological investigation of cyclic octapeptide: Cherimolacyclopeptide G. Turk. J. Chem. 2008, 32, 205–215. [Google Scholar]
- Dahiya, R. Total synthesis and biological potential of psammosilenin A. Arch. Pharm. Chem. Life Sci. 2008, 341, 502–509. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, R.; Pathak, D.; Himaja, M.; Bhatt, S. First total synthesis and biological screening of hymenamide E. Acta Pharm. 2006, 56, 399–415. [Google Scholar] [PubMed]
- Dahiya, R.; Kumar, A.; Gupta, R. Synthesis, cytotoxic and antimicrobial screening of a proline-rich cyclopolypeptide. Chem. Pharm. Bull. (Tokyo) 2009, 57, 214–217. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, R.; Gautam, H. Total synthesis and antimicrobial activity of a natural cycloheptapeptide of marine origin. Mar. Drugs 2010, 8, 2384–2394. [Google Scholar] [CrossRef] [PubMed]
- Poojary, B.; Belagali, S.L. Synthetic studies on cyclic octapeptides: Yunnanin F and hymenistatin. Eur. J. Med. Chem. 2005, 40, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Poojary, B.; Kumar, K.H.; Belagali, S.L. Synthesis and biological evaluation of pseudostellarin B. Pharmaco 2001, 56, 331–334. [Google Scholar] [CrossRef]
- Dahiya, R.; Kaur, K. Synthetic and biological studies on natural cyclic heptapeptide: Segetalin E. Arch. Pharm. Res. 2007, 30, 1380–1386. [Google Scholar] [CrossRef] [PubMed]
- El Khatib, M.; Elagawany, M.; Caliskan, E.; Davis, E.F.; Faidallah, H.M.; El-Feky, S.A.; Katritzky, A.R. Total synthesis of cyclic heptapeptide rolloamide B. Chem. Commun. (Camb.) 2013, 49, 2631–2633. [Google Scholar] [CrossRef] [PubMed]
- Poojary, B.; Kumar, K.H.; Belagali, S.L. Synthesis of a new cyclic peptide, pseudostellarin G. Z. Naturforsch. B 2004, 59, 817–820. [Google Scholar] [CrossRef]
- Zhang, C.M.; Guo, J.X.; Wang, L.; Chai, X.Y.; Hu, H.G.; Wu, Q.Y. Total synthesis of cyclic heptapeptide euryjanicin B. Chin. Chem. Lett. 2011, 22, 631–634. [Google Scholar] [CrossRef]
- McKeever, B.; Pattenden, G. Total synthesis of mollamide, a reverse prenyl substituted cytotoxic cyclic peptide from Didemnum molle. Tetrahedron Lett. 1999, 40, 9317–9320. [Google Scholar] [CrossRef]
- Dellai, A.; Maricic, I.; Kumar, V.; Arutyunyan, S.; Bouraoui, A.; Nefzi, A. Parallel synthesis and anti-inflammatory activity of cyclic peptides cyclosquamosin D and Met-cherimolacyclopeptide B and their analogs. Bioorg. Med. Chem. Lett. 2010, 20, 5653–5657. [Google Scholar] [CrossRef] [PubMed]
- Fairweather, K.A.; Sayyadi, N.; Roussakis, C.; Jolliffi, K.A. Synthesis of the cyclic heptapeptide axinellin A. Tetrahedron 2010, 66, 935–939. [Google Scholar] [CrossRef]
- Napolitano, A.; Bruno, I.; Riccio, R.; Gomez-Paloma, L. Synthesis, structure, and biological aspects of cyclopeptides related to marine phakellistatins 7–9. Tetrahedron 2005, 61, 6808–6815. [Google Scholar] [CrossRef]
- Ali, L.; Musharraf, S.G.; Shaheen, F. Solid-phase total synthesis of cyclic decapeptide phakellistatin 12. J. Nat. Prod. 2008, 71, 1059–1062. [Google Scholar] [CrossRef] [PubMed]
- Sleebs, M.M.; Scanlon, D.; Karas, J.; Maharani, R.; Hughes, A.B. Total synthesis of the antifungal depsipeptide petriellin A. J. Org. Chem. 2011, 76, 6686–6693. [Google Scholar] [CrossRef] [PubMed]
- Napolitano, A.; Bruno, I.; Rovero, P.; Lucas, R.; Peris, M.P.; Gomez-Paloma, L.; Riccio, R. Synthesis, structural aspects and bioactivity of the marine cyclopeptide hymenamide C. Tetrahedron 2001, 57, 6249–6255. [Google Scholar] [CrossRef]
- Garcia-Barrantes, P.M.; Lindsley, C.W. Total synthesis of gombamide A. Org. Lett. 2016, 18, 3810–3813. [Google Scholar] [CrossRef] [PubMed]
- Sellanes, D.; Manta, E.; Serra, G. Toward the total synthesis of scleritodermin A: Preparation of the C1–N15 fragment. Tetrahedron Lett. 2007, 48, 1827–1830. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, R.; Pathak, D. First total synthesis and biological evaluation of halolitoralin A. J. Serb. Chem. Soc. 2007, 72, 101–107. [Google Scholar] [CrossRef]
- Dahiya, R.; Maheshwari, M.; Kumar, A. Toward the synthesis and biological evaluation of hirsutide. Monatsh. Chem. 2009, 140, 121–127. [Google Scholar] [CrossRef]
- Huang, T.; Zou, Y.; Wu, M.C.; Zhao, Q.J.; Hu, H.G. Total synthesis of proline-rich cyclic octapeptide stylissamide X. Chem. Nat. Prod. 2015, 51, 523–526. [Google Scholar] [CrossRef]
- Santhakumar, G.; Payne, R.J. Total synthesis of polydiscamides B, C, and D via a convergent native chemical ligation-oxidation strategy. Org. Lett. 2014, 16, 4500–4503. [Google Scholar] [CrossRef] [PubMed]
- Pettit, G.R.; Holman, J.W.; Boland, G.M. Synthesis of the cyclic heptapeptides axinastatin 2 and axinastatin 3. J. Chem. Soc. Perkin Trans. 1 1996, 2411–2416. [Google Scholar] [CrossRef]
- Dahiya, R.; Pathak, D. Cyclic peptides: New hope for antifungal therapy. Egypt. Pharm. J. (NRC) 2006, 5, 189–199. [Google Scholar]
- Pathak, D.; Dahiya, R. Cyclic peptides as novel antineoplastic agents: A review. J. Sci. Pharm. 2003, 4, 125–131. [Google Scholar]
- Pettit, G.R.; Xu, J.P.; Dorsaz, A.C.; Williams, M.D.; Boyd, M.R.; Cerny, R.L. Isolation and structure of the human cancer cell growth inhibitory cyclic decapeptides phakellistatins 7, 8 and 9. Bioorg. Med. Chem. Lett. 1995, 5, 1339–1344. [Google Scholar] [CrossRef]
- Pettit, G.R.; Tan, R. Antineoplastic agents 390. Isolation and structure of phakellistatin 12 from a Chuuk Archipelago marine sponge. Bioorg. Med. Chem. Lett. 2003, 13, 685–688. [Google Scholar] [CrossRef]
- Li, L.H.; Timmins, L.G.; Wallace, T.L.; Krueger, W.C.; Prairie, M.D.; Im, W.B. Mechanism of action of didemnin B, a depsipeptide from the sea. Cancer Lett. 1984, 23, 279–288. [Google Scholar] [CrossRef]
- Zheng, L.H.; Wang, Y.J.; Sheng, J.; Wang, F.; Zheng, Y.; Lin, X.K.; Sun, M. Antitumor peptides from marine organisms. Mar. Drugs 2011, 9, 1840–1859. [Google Scholar] [CrossRef] [PubMed]
- Siemion, I.Z.; Cebrat, M.; Wieczorek, Z. Cyclolinopeptides and their analogs—A new family of peptide immunosuppressants affecting the calcineurin system. Arch. Immunol. Ther. Exp. 1999, 47, 143–153. [Google Scholar]
- Malaker, A.; Ahmad, S.A.I. Therapeutic potency of anticancer peptides derived from marine organism. Int. J. Eng. Appl. Sci. 2013, 2, 53–65. [Google Scholar]
- Simmons, T.L.; Andrianasolo, E.; McPhail, K.; Flatt, P.; Gerwick, W.H. Marine natural products as anticancer drugs. Mol. Cancer Ther. 2005, 4, 333–342. [Google Scholar] [PubMed]
- Proksch, P.; Ebel, R.; Edrada, R.A.; Wray, V.; Steube, K. Bioactive natural products from marine invertebrates and associated fungi. Prog. Mol. Subcell. Biol. 2003, 37, 117–142. [Google Scholar] [PubMed]
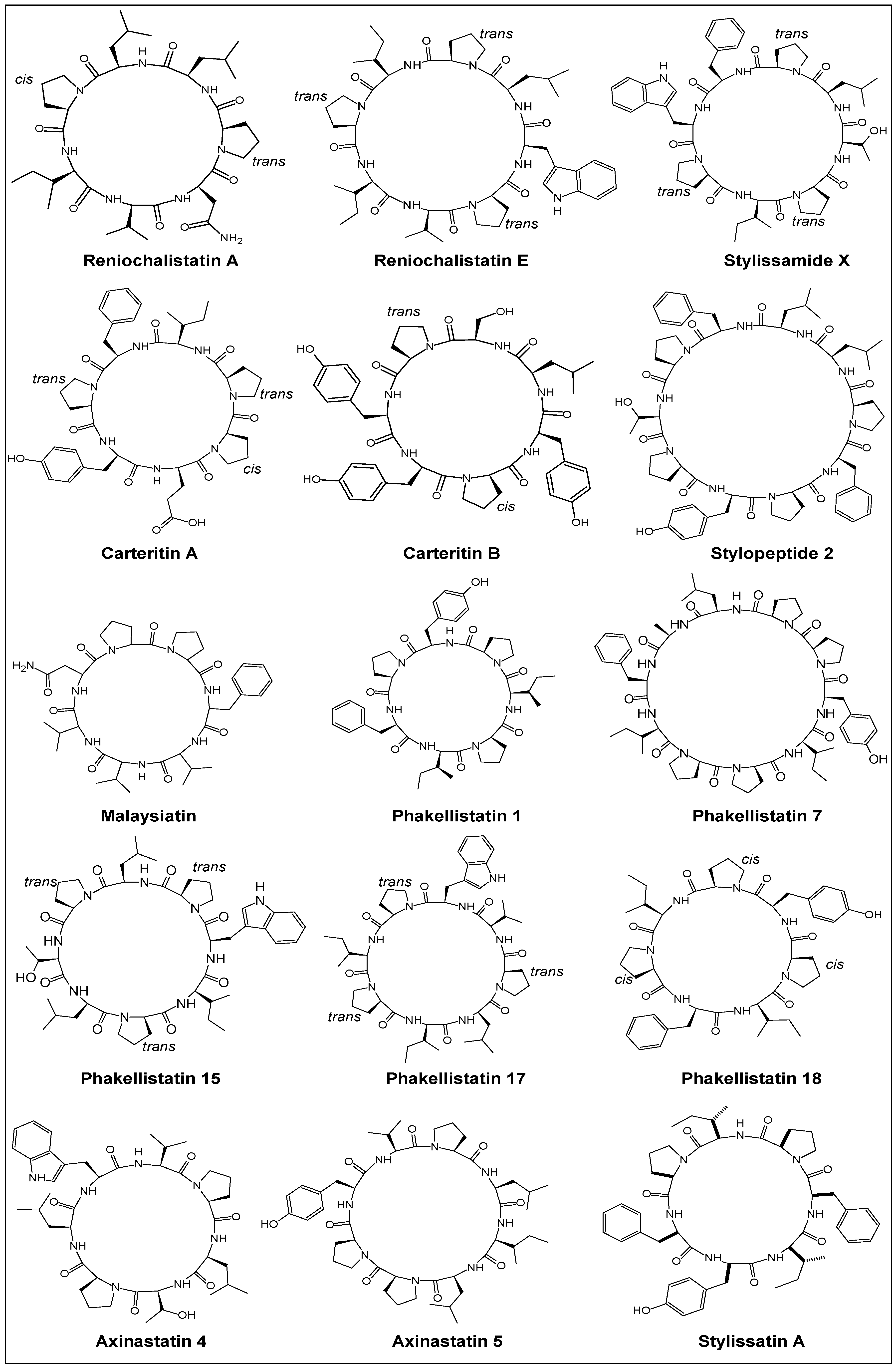
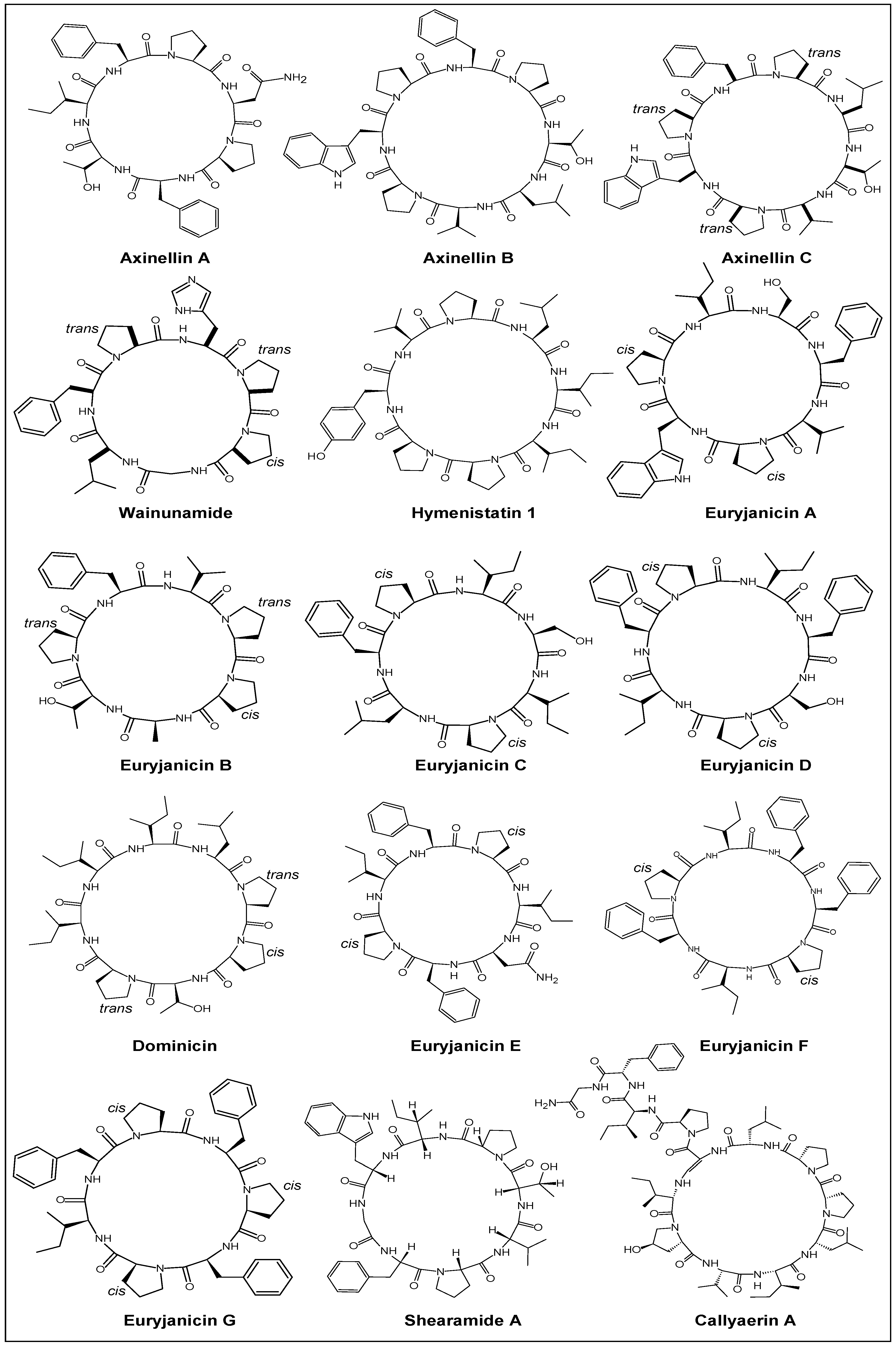
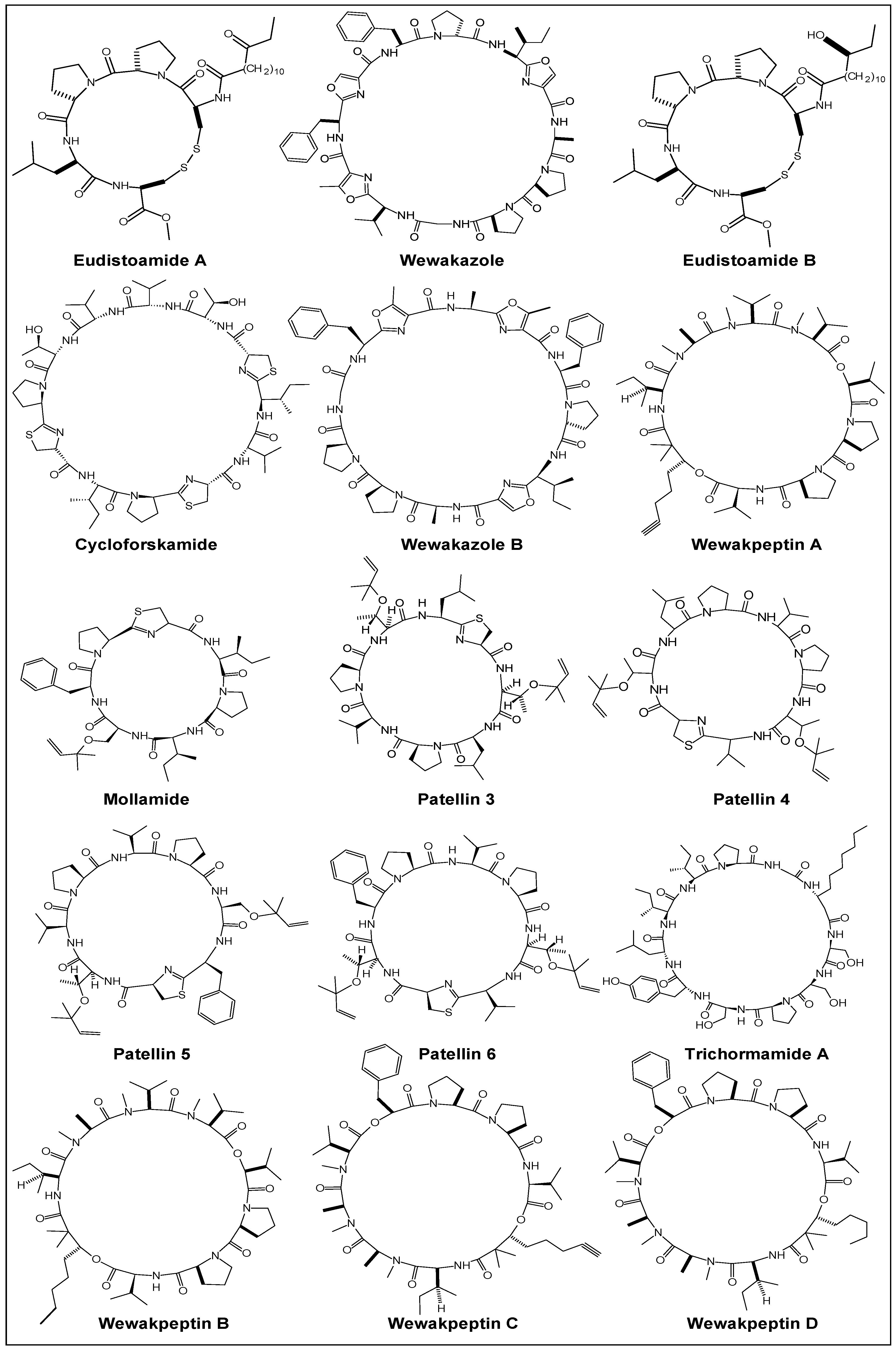
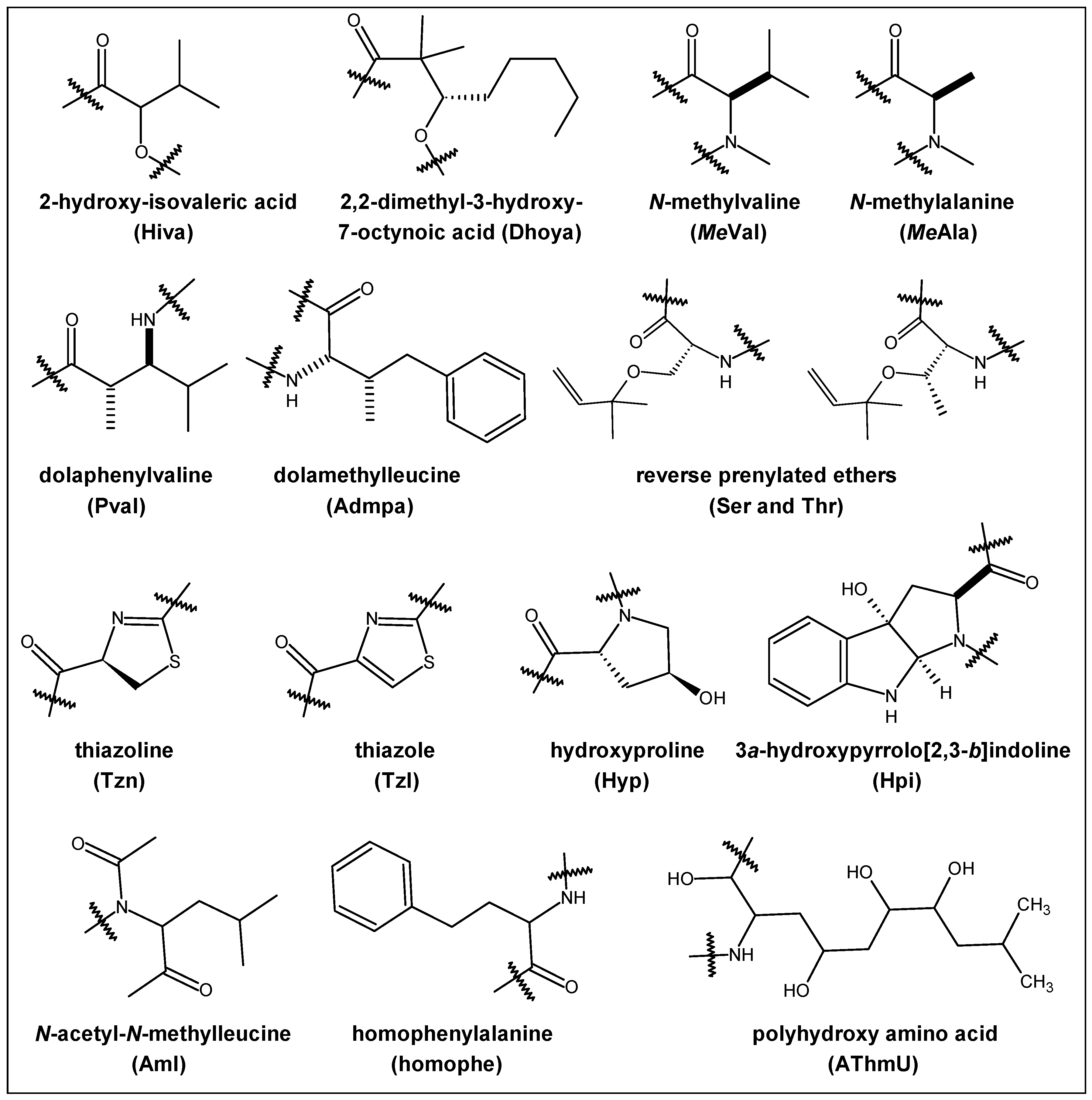
| Year | Cyclic Peptide | Molecular Formula | No. of Proline Units | Composition |
|---|---|---|---|---|
| 1981 | Didemnin B [43] | C57H89N7O15 | two | cyclodepsipeptide |
| 1988 | Aplidine [45] | C57H87N7O15 | cyclodepsipeptide | |
| 1991 | Axinastatin 1 [6] | C38H56N8O8 | cycloheptapeptide | |
| 1992 | Malaysiatin [27] | C38H56N8O8 | cycloheptapeptide | |
| 1992 | Polydiscamide A [7] | C76H109BrN19O20SNa | cyclodepsipeptide | |
| 1993 | Axinastatin 4 [76] | C42H62N8O8 | cycloheptapeptide | |
| 1993 | Cyclooligopeptide [77] | C24H32N4O5 | cyclotetrapeptide | |
| 1993 | Hymenamide B [31] | C43H56N8O10 | cycloheptapeptide | |
| 1993 | Hymenamide C [8] | C43H54N8O9 | cycloheptapeptide | |
| 1993 | Hymenamide D [8] | C38H55N7O10 | cycloheptapeptide | |
| 1993 | Hymenamide E [8] | C45H55N7O10 | cycloheptapeptide | |
| 1994 | Mollamide [40] | C42H61N7O7S | cycloheptapeptide | |
| 1994 | Schizotrin A [78] | C72H107N13O21 | cycloundecapeptide | |
| 1994 | Axinastatin 2 [39] | C39H58N8O8 | cycloheptapeptide | |
| 1994 | Axinastatin 3 [39] | C40H61N8O8 | cycloheptapeptide | |
| 1995 | Stylopeptide 1 [79] | C40H61N7O8 | cycloheptapeptide | |
| 1996 | Patellin 3 [80] | C48H78N8O9S | cyclooctapeptide | |
| 1996 | Patellin 4 [80] | C47H76N8O9S | cyclooctapeptide | |
| 1996 | Patellin 5 [80] | C49H72N8O9S | cyclooctapeptide | |
| 1996 | Patellin 6 [80] | C50H74N8O9S | cyclooctapeptide | |
| 1996 | Hymenamide F [81] | C35H60N10O7S | cycloheptapeptide | |
| 1996 | Agardhipeptin B [82] | C57H69N11O8 | cyclooctapeptide | |
| 1996 | Kapakahine A [37] | C58H72N10O9 | cyclooctapeptide | |
| 1996 | Kapakahine C [37] | C58H72N10O10 | cyclooctapeptide | |
| 1996 | Kapakahine D [37] | C58H72N10O10 | cyclooctapeptide | |
| 1998 | Axinellin A [21] | C42H56N8O9 | cycloheptapeptide | |
| 1998 | Shearamide A [83] | C47H63N9O9 | cyclooctapeptide | |
| 1999 | Prenylagaramide B [84] | C49H68N8O10 | cycloheptapeptide | |
| 1999 | Nostophycin [50] | C46H64N8O10 | cycloheptapeptide | |
| 2000 | trans,trans-ceratospongamide [10] | C41H49N7O6S | cycloheptapeptide | |
| 2000 | Tamandarine A [44] | C54H87N7O14 | cyclodepsipeptide | |
| 2000 | Tamandarine B [44] | C53H82N7O14 | cyclodepsipeptide | |
| 2001 | Microspinosamide [38] | C75H109BrN18O22S | cyclodepsipeptide | |
| 2003 | Myriastramide C [28] | C42H53N9O7S | cyclooctapeptide | |
| 2004 | Scleritodermin A [26] | C42H54N7O10SNa | cyclodepsipeptide | |
| 2004 | Cyclonellin [85] | C45H62N12O12 | cyclooctapeptide | |
| 2005 | Wewakpeptin A [46] | C52H85N7O11 | cyclodepsipeptide | |
| 2005 | Wewakpeptin B [46] | C52H89N7O11 | cyclodepsipeptide | |
| 2005 | Wewakpeptin C [46] | C54H81N7O11 | cyclodepsipeptide | |
| 2005 | Wewakpeptin D [46] | C54H85N7O11 | cyclodepsipeptide | |
| 2007 | Pahayokolide A [48] | C72H105N13O20 | cycloundecapeptide | |
| 2007 | Pahayokolide B [48] | C63H90N12O18 | cycloundecapeptide | |
| 2008 | Polydiscamide B [18] | C75H110BrN18O21S | cyclodepsipeptide | |
| 2008 | Polydiscamide C [18] | C74H107BrN18O21S | cyclodepsipeptide | |
| 2008 | Polydiscamide D [18] | C73H105BrN18O21S | cyclodepsipeptide | |
| 2009 | Euryjanicin A [36] | C44H58N8O8 | cycloheptapeptide | |
| 2009 | Euryjanicin C [14] | C40H61N7O8 | cycloheptapeptide | |
| 2009 | Euryjanicin D [14] | C44H59N7O8 | cycloheptapeptide | |
| 2009 | Eudistomide A [42] | C37H61N5O8S2 | cyclolipopeptide | |
| 2009 | Eudistomide B [42] | C37H63N5O8S2 | cyclolipopeptide | |
| 2010 | Anacyclamide A10 [86] | C49H72N12O14 | cyclodecapeptide | |
| 2011 | Duanbanhuain A [87] | C43H58N8O11 | cyclooctapeptide | |
| 2011 | Duanbanhuain B [87] | C45H57N9O10 | cyclooctapeptide | |
| 2012 | Mollamide F [12] | C33H46N6O5S | cyclohexapeptide | |
| 2013 | Stylissatin A [24] | C49H63N7O8 | cycloheptapeptide | |
| 2013 | Euryjanicin E [88] | C44H60N8O8 | cycloheptapeptide | |
| 2013 | Euryjanicin F [88] | C49H63N7O7 | cycloheptapeptide | |
| 2013 | Gombamide A [22] | C38H45N7O8S2 | cyclothiohexapeptide | |
| 2013 | Cycloforskamide [41] | C54H86N12O11S3 | cyclododecapeptide | |
| 2014 | Trichormamide A [49] | C58H93N11O15 | cycloundecapeptide | |
| 2014 | Reniochalistatin A [20] | C37H62N8O8 | cycloheptapeptide | |
| 2016 | Carteritin B [34] | C46H57N7O11 | cycloheptapeptide | |
| 1990 | Hymenistatin 1 [35] | C47H72N8O9 | three | cyclooctapeptide |
| 1993 | Phakellistatin 1 [32] | C45H61N7O8 | cycloheptapeptide | |
| 1993 | Hymenamide A [31] | C46H61N11O7 | cycloheptapeptide | |
| 1993 | Phakellistatin 2 [89] | C45H61N7O8 | cycloheptapeptide | |
| 1994 | Axinastatin 5 [30] | C47H72N8O9 | cyclooctapeptide | |
| 1994 | Hymenamide G [90] | C47H72N8O9 | cyclooctapeptide | |
| 1994 | Hymenamide H [90] | C47H69N9O9 | cyclooctapeptide | |
| 1995 | Phakellistatin 11 [91] | C53H67N9O9 | cyclooctapeptide | |
| 1996 | Waiakeamide [12] | C37H49N7O8S3 | cyclohexapeptide | |
| 1998 | Axinellin B [21] | C50H67N9O9 | cyclooctapeptide | |
| 2000 | Haligramide A [92] | C37H49N7O6S3 | cyclohexapeptide | |
| 2000 | Haligramide B [92] | C37H49N7O7S3 | cyclohexapeptide | |
| 2001 | Haliclonamide A [93] | C45H60N8O9 | cyclooctapeptide | |
| 2001 | Haliclonamide B [93] | C40H52N8O9 | cyclooctapeptide | |
| 2001 | Wainunuamide [23] | C38H51N9O7 | cycloheptapeptide | |
| 2002 | Axinellin C [94] | C50H67N9O9 | cyclooctapeptide | |
| 2002 | Dolastatin 16 [52] | C47H70N6O10 | cyclodepsipeptide | |
| 2002 | Haliclonamide C [95] | C45H60N8O10 | cyclooctapeptide | |
| 2002 | Haliclonamide D [95] | C40H54N8O10 | cyclooctapeptide | |
| 2002 | Haliclonamide E [95] | C45H62N8O10 | cyclooctapeptide | |
| 2003 | Myriastramide A [28] | C45H58N8O9 | cyclooctapeptide | |
| 2003 | Myriastramide B [28] | C45H57ClN8O9 | cyclooctapeptide | |
| 2003 | Wewakazole [96] | C59H72N12O12 | cyclododecapeptide | |
| 2005 | Dominicin [16] | C43H72N8O9 | cyclooctapeptide | |
| 2006 | Stylisin 1 [19] | C45H61N7O8 | cycloheptapeptide | |
| 2009 | Euryjanicin B [14] | C36H51N7O8 | cycloheptapeptide | |
| 2010 | Phakellistatin 15 [25] | C48H71N9O9 | cyclooctapeptide | |
| 2010 | Phakellistatin 17 [25] | C49H73N9O8 | cyclooctapeptide | |
| 2010 | Phakellistatin 18 [25] | C45H61N7O8 | cycloheptapeptide | |
| 2010 | Callyaerin B [13] | C65H108N12O14 | cyclooctapeptide b | |
| 2010 | Callyaerin C [13] | C70H105N13O16 | cycloheptapeptide c | |
| 2012 | Stylissamide X [33] | C51H69N9O9 | cyclooctapeptide | |
| 2013 | Euryjanicin G [88] | C48H59N7O7 | cyclooctapeptide | |
| 2014 | Reniochalistatins E [20] | C49H73N9O8 | cyclooctapeptide | |
| 2016 | Carteritin A [34] | C44H57N7O10 | cycloheptapeptide | |
| 2016 | Stylissatin B [97] | C38H51N9O7 | cycloheptapeptide | |
| 2016 | Stylissatin C [97] | C39H55N7O9 | cycloheptapeptide | |
| 2016 | Stylissatin D [97] | C40H57N7O9 | cycloheptapeptide | |
| 2016 | Wewakazole B [47] | C58H70N12O12 | cyclododecapeptide | |
| 1968 | Antamanide [98] | C64H78N10O10 | four | cyclodecapeptide |
| 2004 | Callynormine A [15] | C61H93N11O13 | cycloheptapeptide b | |
| 2006 | Stylisin 2 [19] | C44H57N7O8 | cycloheptapeptide | |
| 2008 | Stylopeptide 2 [29] | C63H84N10O12 | cyclodecapeptide | |
| 2010 | Callyaerin A [13] | C69H108N14O14 | cyclooctapeptide c | |
| 2010 | Callyaerin E [13] | C66H94N12O13 | cycloheptapeptide c | |
| 2010 | Callyaerin H [13] | C54H81N11O10 | cycloheptapeptide a | |
| 2008 | Callyaerin G [99] | C69H91N13O12 | five | cycloheptapeptide c |
| PRCPs | Resource | Pharmacological Activity | |
|---|---|---|---|
| Susceptibility | MIC Value | ||
| Axinastatin 1 [6] | marine sponge | Cytotoxicity against PS leukemia cell line | 0.21 μg/mL |
| Polydiscamide A [7] | marine sponge | Antiproliferative activity against human lung cancer A549 cell line; antibacterial activity against Bacillus subtilis | 0.7 μg/mL; 3.1 μg/mL |
| Hymenamide E [8] | marine sponge | Antifungal activity against pathogenic Cryptococcus neoformans | 133 μg/mL |
| trans,trans-Ceratospongamide [10] | marine red alga | Inhibition of sPLA2 expression in a cell-based model for anti-inflammation | 0.0013 μg/mL |
| Mollamide F [12] | marine tunicate | Anti-HIV activity in cytoprotective cell-based assay and HIV integrase inhibition assay | 0.0016 and 0.0031 μg/mL |
| Callyaerin A [13] | marine sponge | Anti-TB activity against M. tuberculosis, inhibitory activity toward C. albicans | 7.37 μg/mL |
| Callyaerin B [13] | marine sponge | Anti-TB activity against Mycobacterium tuberculosis | 7.8 μg/mL |
| Callyaerin E, H [13] | marine sponge | Cytotoxicity against L5178Y cell line | 7.91 and 9.59 μg/mL |
| Euryjanicin C [14] | marine sponge | Inhibitory activity against human hepatitis B virus | 49 μg/mL |
| Polydiscamides B–D [18] | marine sponge | Agonist activity against human sensory neuron-specific G protein couple receptor (SNSR) that is involved in the modulation of pain | - |
| Axinellin A, B [21] | marine sponge | Antitumor activity against human bronchopulmonary non-small-cell lung-carcinoma lines (NSCLC-N6) | 3.0 and 7.3 μg/mL |
| Wainunuamide [23] | marine sponge | Cytotoxic activity against A2780 ovarian tumor and K562 leukemia cancer cells | 19.15 and 18.36 μg/mL |
| Stylissatin A [24] | marine sponge | Inhibition of NO production in LPS-stimulated RAW264.7 cells | 0.0011 μg/mL |
| Scleritodermin A [26] | marine sponge | Inhibition of tubulin polymerization and human tumor cell lines | - |
| Axinastatin 5 [30] | marine sponge | Cytotoxic activity against human and murine cancer cells | 0.3–3.3 μg/mL |
| Phakellistatin 1 [32] | marine sponges | Cell growth inhibitory activity against P-388 murine leukemia | 7.5 μg/mL |
| Stylissamide X [33] | marine sponge | Inhibitory activity against migration of HeLa cells | 0.001–0.1 μg/mL |
| Carteritin A [34] | marine sponge | Cytotoxicity against HeLa, HCT116 and RAW264 cells | 0.0012–0.0026 μg/mL |
| Hymenistatin 1 [35] | marine sponge | Cytotoxicity against P-388 leukemia cells | 3.5 μg/mL |
| Kapakahine A, C [37] | marine sponge | Cytotoxicity against P-388 murine leukemia cells | 5.4 and 5.0 μg/mL |
| Microspinosamide [38] | marine sponge | Anti-HIV activity in CEM-SS cells | 0.2 μg/mL |
| Axinastatin 2 [39] | marine sponge | Cytotoxicity against murine leukemia P-388 cell line | 0.02 μg/mL |
| Axinastatin 3 [39] | marine sponge | Cytotoxicity against PS leukemia cell line | 0.4 μg/mL |
| Mollamide [40] | sea squirt | Cytotoxicity against P-388 (murine leukemia) and A549 (human lung carcinoma), HT29 (human colon carcinoma) cells | 1.0–2.5 μg/mL |
| Cycloforskamide [41] | sea slug | Cytotoxicity against murine leukemia P-388 cells | 8.51 μg/mL |
| Didemnin B [43] | marine tunicate | Cytotoxic activity against human L1210 lymphocytic leukemia cell lines; pancreatic carcinoma (BX-PC3) cell lines; prostatic cancer (DU-145) cell lines; head and neck carcinoma (UMSCC10b) cell lines | 0.0025 μg/mL; 0.002 μg/mL; 0.0015 μg/mL; 0.0018 μg/mL |
| Tamandarin A [44] | marine ascidian | Cytotoxic activity against human pancreatic carcinoma (BX-PC3) cell lines; prostatic cancer (DU-145) cell lines; head and neck carcinoma (UMSCC10b) cell lines | 0.0018 μg/mL; 0.0014 μg/mL; 0.0009 μg/mL |
| Wewakpeptin A [46] | marine cyanobacterium | Cytotoxicity against NCI-H460 human lung tumor and the neuro-2a mouse neuroblastoma cell lines | 0.001 μg/mL |
| Wewakazole B [47] | marine cyanobacterium | Cytotoxicity against human MCF7 breast/H460 lung cancer cells | 8.87–15.29 μg/mL |
| Pahayokolide A [48] | marine cyanobacteria | Antibacterial activity against Bacillus megaterium, Bacillus subtilis | 5 μg/mL |
| Trichormamide A [49] | marine cyanobacteria | Antiproliferative activities against the human melanoma cell line (MDA-MB-435) and the human colon cancer cell line (HT-29) | 8.45 and 8.53 μg/mL |
| Axinastatin 4 [76] | marine sponge | Cytotoxic activity against P-388 lymphocytic leukemia cell line | 0.057 μg/mL |
| Phakellistatin 2 [89] | marine sponge | Cell growth inhibitory activity against P-388 cell line | 0.34 μg/mL |
| Phakellistatin 7–9 [137] | marine sponge | Cell growth inhibitory activity against P-388 murine leukemia | 3.0, 2.9 and 4.1 μg/mL |
| Axinellin C [94] | marine sponge | Cytotoxic activity against A2780 ovarian tumor and K562 leukemia cancer cells | 13.17 and 4.46 μg/mL |
| Callyaerin G [99] | marine sponge | Cytotoxic towards the mouse lymphoma cell line (L5178Y) and HeLa cells | 0.53 and 5.4 μg/mL |
| Stylissatin B [97] | marine sponge | Inhibitory effects against human tumor cell lines including HCT-116, HepG2, BGC-823, NCI-H1650, A2780 and MCF7 | 0.0013 μg/mL |
| Phakellistatin 10, 11 [91] | marine sponge | Cell growth inhibitory activity against murine P-388 lymphocytic leukemia | 2.1, 0.20 μg/mL |
| Stylopeptide 1 [79] | marine sponge | Cell growth inhibitory activity against murine P-388 lymphocytic leukemia | 0.01 μg/mL |
| Phakellistatin 12 [138] | marine sponge | Cell growth inhibitory activity against murine P-388 lymphocytic leukemia | 2.8 μg/mL |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fang, W.-Y.; Dahiya, R.; Qin, H.-L.; Mourya, R.; Maharaj, S. Natural Proline-Rich Cyclopolypeptides from Marine Organisms: Chemistry, Synthetic Methodologies and Biological Status. Mar. Drugs 2016, 14, 194. https://doi.org/10.3390/md14110194
Fang W-Y, Dahiya R, Qin H-L, Mourya R, Maharaj S. Natural Proline-Rich Cyclopolypeptides from Marine Organisms: Chemistry, Synthetic Methodologies and Biological Status. Marine Drugs. 2016; 14(11):194. https://doi.org/10.3390/md14110194
Chicago/Turabian StyleFang, Wan-Yin, Rajiv Dahiya, Hua-Li Qin, Rita Mourya, and Sandeep Maharaj. 2016. "Natural Proline-Rich Cyclopolypeptides from Marine Organisms: Chemistry, Synthetic Methodologies and Biological Status" Marine Drugs 14, no. 11: 194. https://doi.org/10.3390/md14110194
APA StyleFang, W.-Y., Dahiya, R., Qin, H.-L., Mourya, R., & Maharaj, S. (2016). Natural Proline-Rich Cyclopolypeptides from Marine Organisms: Chemistry, Synthetic Methodologies and Biological Status. Marine Drugs, 14(11), 194. https://doi.org/10.3390/md14110194







