Marine Microbiological Enzymes: Studies with Multiple Strategies and Prospects
Abstract
:1. Introduction
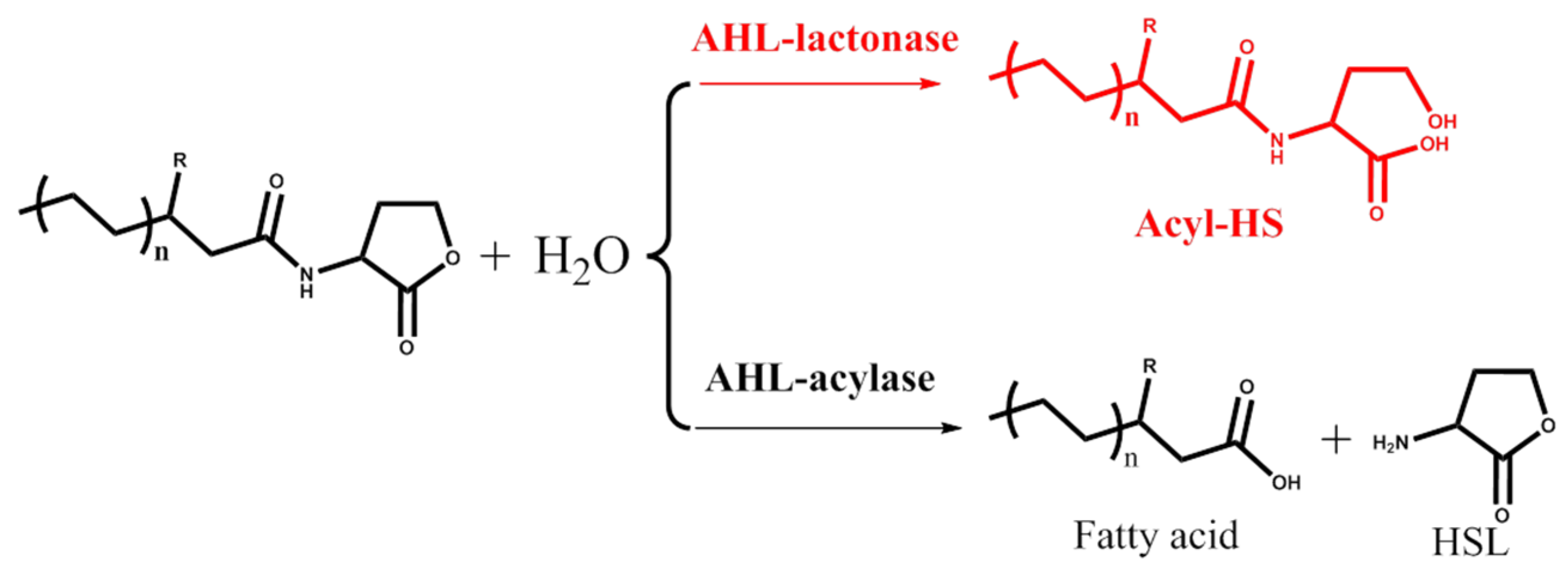
2. AHL Lactonase
2.1. Introduction
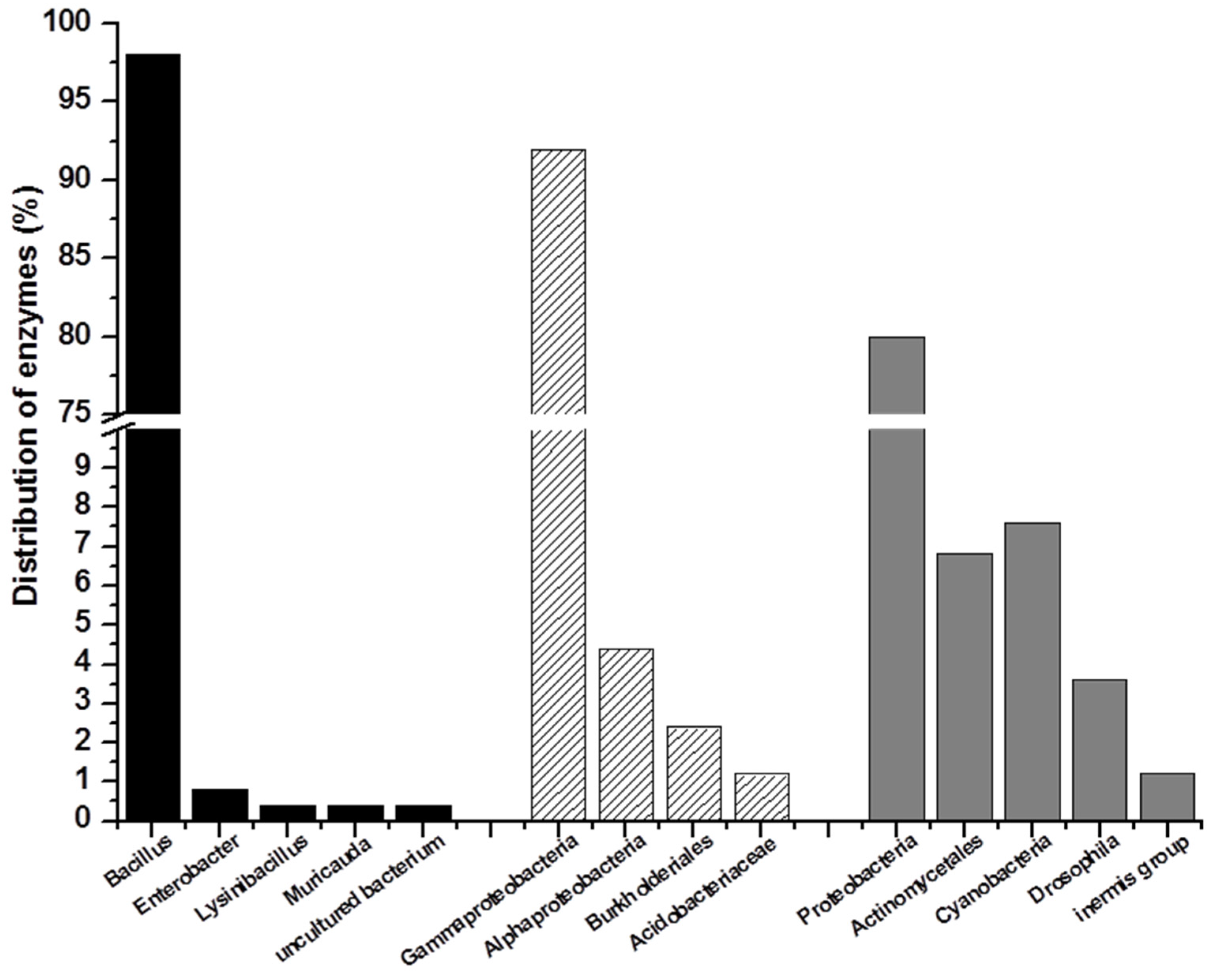
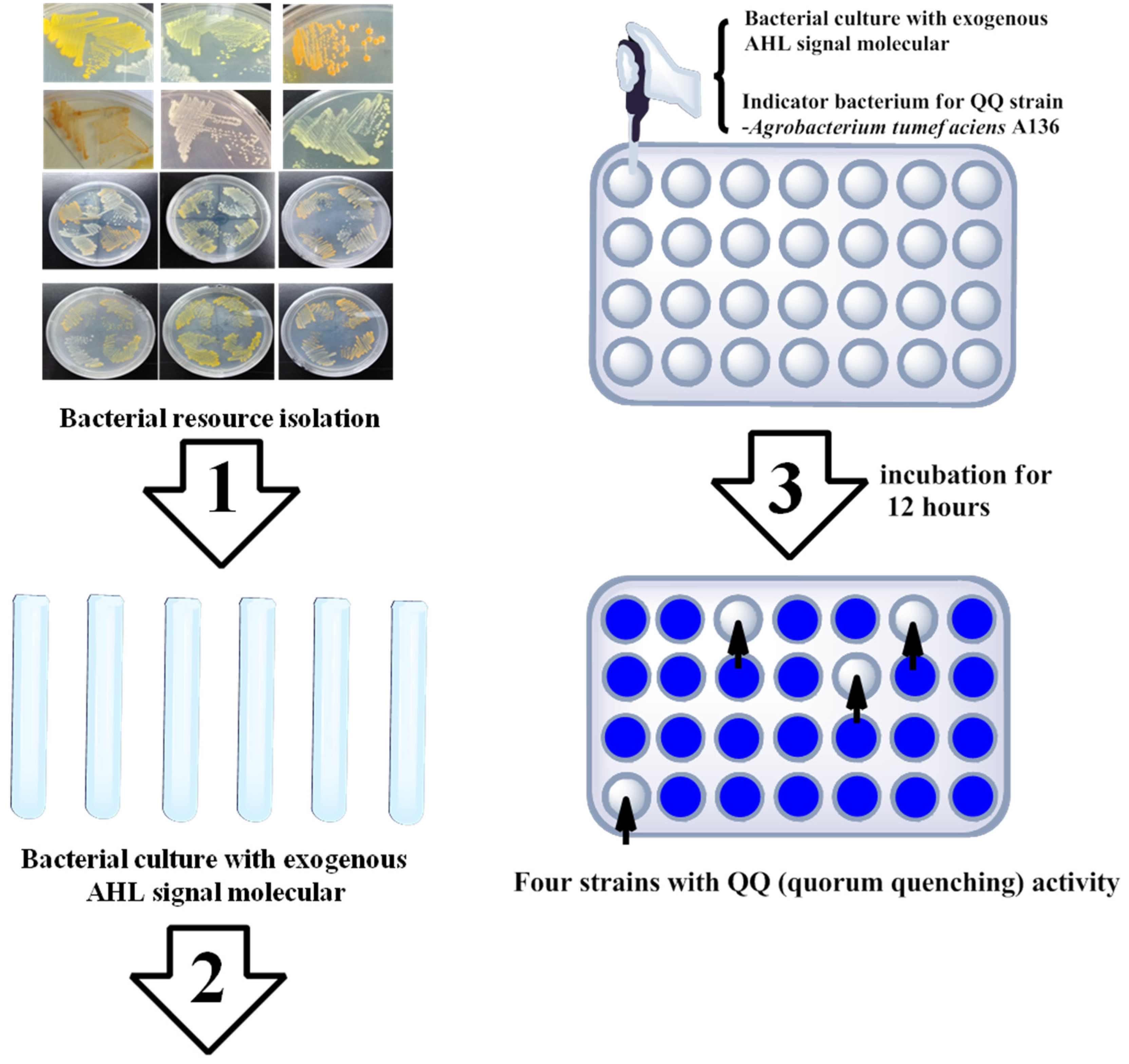
2.2. Marine Resources of AHL Lactonase and Research Methods
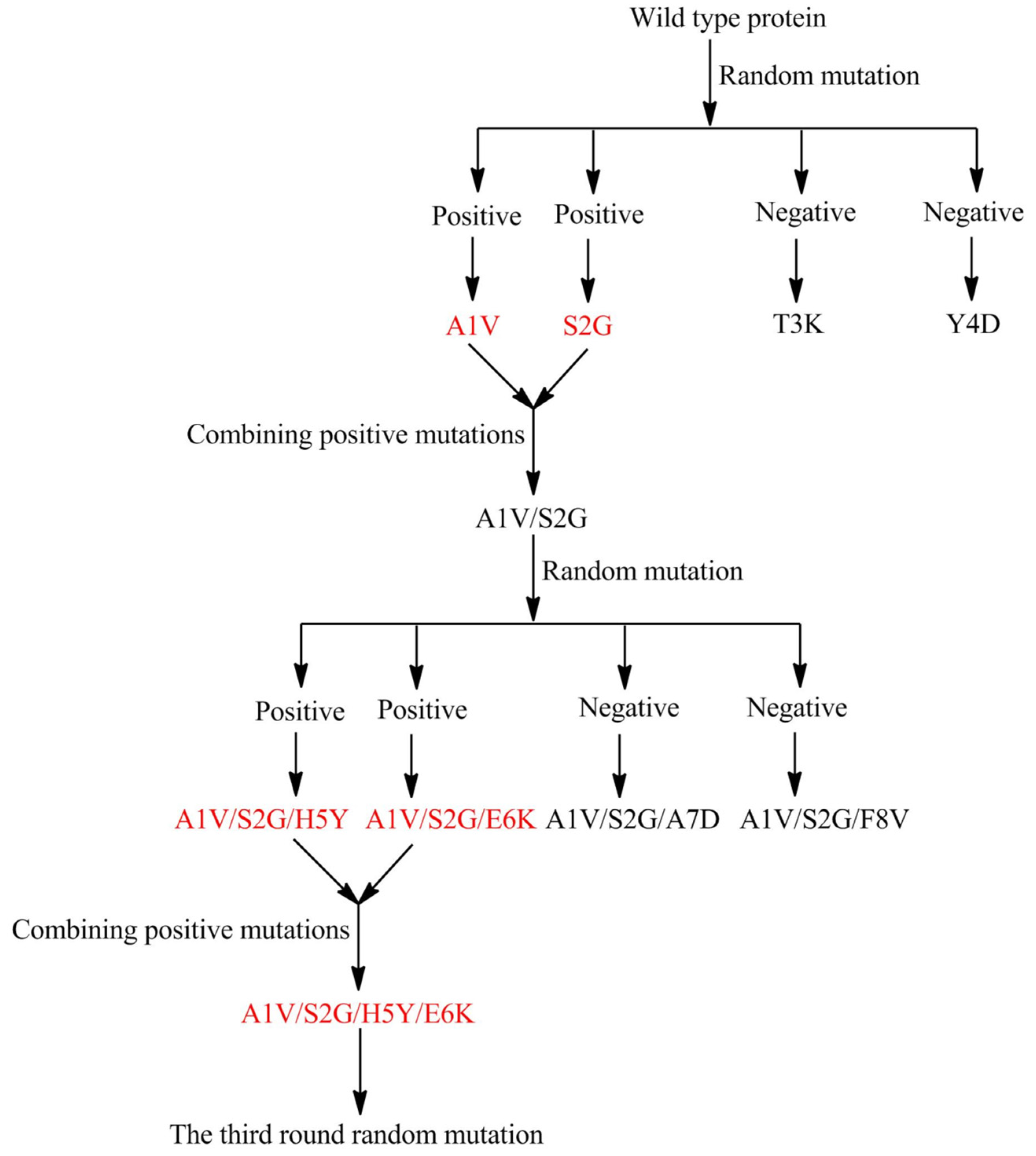
2.3. Prospects for AHL Lactonase
3. Amylase
3.1. Introduction
3.2. Marine Resources of Amylase and Related Catalytic Mechanisms
3.3. Prospects for Amylase
4. Alginate Lyase
4.1. Introduction
4.2. Marine Resources of Alginate Lyase and Catalytic Mechanisms
4.3. Prospects for Alginate Lyase
5. Chitinase
5.1. Introduction
5.2. Marine Resources of Chitinase and Catalytic Mechanisms
5.3. Prospects for Chitinase
6. Cellulase
6.1. Introduction
6.2. Marine Resources of Cellulase and Catalytic Mechanisms
6.3. Prospects for Cellulase
7. Conclusions
Acknowledgments
Conflicts of Interest
References
- Antranikian, G.; Vorgias, C.E.; Bertoldo, C. Extreme environments as a resource for microorganisms and novel biocatalysts. Adv. Biochem. Eng. Biotechnol. 2005, 96, 219–262. [Google Scholar] [PubMed]
- Bhattacharya, A.; Pletschke, B.I. Review of the enzymatic machinery of Halothermothrix orenii with special reference to industrial applications. Enzyme Microb. Technol. 2014, 55, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Huang, Z.; Zhang, X.; Shao, Z.; Liu, Z. Cloning, expression and characterization of a novel cold-active and halophilic xylanase from Zunongwangia profunda. Extremophiles 2014, 18, 441–450. [Google Scholar] [CrossRef] [PubMed]
- Jaiganesh, R.; Sampath Kumar, N.S. Marine bacterial sources of bioactive compounds. Adv. Food Nutr. Res. 2012, 65, 389–408. [Google Scholar] [PubMed]
- Zotchev, S.B. Marine actinomycetes as an emerging resource for the drug development pipelines. J. Biotechnol. 2012, 158, 168–175. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.S.; Kim, Y.S.; Park, S.; Kim, J.; Kang, S.J.; Lee, M.H.; Ryu, S.; Choi, J.M.; Oh, T.K.; Yoon, J.H. Exceptional production of both prodigiosin and cycloprodigiosin as major metabolic constituents by a novel marine bacterium, Zooshikella rubidus S1-1. Appl. Environ. Microbiol. 2011, 77, 4967–4973. [Google Scholar] [CrossRef] [PubMed]
- Martins, A.; Vieira, H.; Gaspar, H.; Santos, S. Marketed marine natural products in the pharmaceutical and cosmeceutical industries: Tips for success. Mar. Drugs 2014, 12, 1066–1101. [Google Scholar] [CrossRef] [PubMed]
- Rahman, H.; Austin, B.; Mitchell, W.J.; Morris, P.C.; Jamieson, D.J.; Adams, D.R.; Spragg, A.M.; Schweizer, M. Novel anti-infective compounds from marine bacteria. Mar. Drugs 2010, 8, 498–518. [Google Scholar] [CrossRef] [PubMed]
- Abdelmohsen, U.R.; Yang, C.; Horn, H.; Hajjar, D.; Ravasi, T.; Hentschel, U. Actinomycetes from Red Sea sponges: Sources for chemical and phylogenetic diversity. Mar. Drugs 2014, 12, 2771–2789. [Google Scholar] [CrossRef] [PubMed]
- Komaki, H.; Ichikawa, N.; Hosoyama, A.; Fujita, N.; Igarashi, Y. Draft genome sequence of marine-derived streptomyces sp. TP-A0873, a producer of a pyrrolizidine alkaloid bohemamine. Genome Announc. 2015, 3. [Google Scholar] [CrossRef] [PubMed]
- Zhou, K.; Zhang, X.; Zhang, F.; Li, Z. Phylogenetically diverse cultivable fungal community and polyketide synthase (PKS), non-ribosomal peptide synthase (NRPS) genes associated with the South China Sea sponges. Microb. Ecol. 2011, 62, 644–654. [Google Scholar] [CrossRef] [PubMed]
- Zhu, P.; Zheng, Y.; You, Y.; Yan, X.; Shao, J. Molecular phylogeny and modular structure of hybrid NRPS/PKS gene fragment of Pseudoalteromonas sp. NJ6-3-2 isolated from marine sponge Hymeniacidon perleve. J. Microbiol. Biotechnol. 2009, 19, 229–237. [Google Scholar] [PubMed]
- Bassler, B.L. Cell-to-cell communication in bacteria: A chemical discourse. Harvey Lect. 2004, 100, 123–142. [Google Scholar] [PubMed]
- Miyamoto, C.M.; Meighen, E.A. Involvement of LuxR, a quorum sensing regulator in Vibrio harveyi, in the promotion of metabolic genes: argA, purM, lysE and rluA. Biochim. Biophys. Acta 2006, 1759, 296–307. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, S.; Schmidt-Dannert, C. Applications of quorum sensing in biotechnology. Appl. Microbiol. Biotechnol. 2010, 86, 1267–1279. [Google Scholar] [CrossRef] [PubMed]
- Mangwani, N.; Dash, H.R.; Chauhan, A.; Das, S. Bacterial quorum sensing: Functional features and potential applications in biotechnology. J. Mol. Microbiol. Biotechnol. 2012, 22, 215–227. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Momb, J.; Thomas, P.W.; Moulin, A.; Petsko, G.A.; Fast, W.; Ringe, D. Mechanism of the quorum-quenching lactonase (AiiA) from Bacillus thuringiensis. 1. Product-bound structures. Biochemistry 2008, 47, 7706–7714. [Google Scholar] [CrossRef] [PubMed]
- Momb, J.; Wang, C.; Liu, D.; Thomas, P.W.; Petsko, G.A.; Guo, H.; Ringe, D.; Fast, W. Mechanism of the quorum-quenching lactonase (AiiA) from Bacillus thuringiensis. 2. Substrate modeling and active site mutations. Biochemistry 2008, 47, 7715–7725. [Google Scholar] [CrossRef] [PubMed]
- Thomas, P.W.; Stone, E.M.; Costello, A.L.; Tierney, D.L.; Fast, W. The quorum-quenching lactonase from Bacillus thuringiensis is a metalloprotein. Biochemistry 2005, 44, 7559–7569. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; He, S.; Zhou, Z.; Zhang, M.; Mao, W.; Zhang, H.; Yao, B. Orally administered thermostable N-acyl homoserine lactonase from Bacillus sp. strain AI96 attenuates Aeromonas hydrophila infection in zebrafish. Appl. Environ. Microbiol. 2012, 78, 1899–1908. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Zhou, Z.; Cao, Y.; Bai, Y.; Yao, B. High yield expression of an AHL-lactonase from Bacillus sp. B546 in Pichia pastoris and its application to reduce Aeromonas hydrophila mortality in aquaculture. Microb. Cell Fact. 2010, 9, 39. [Google Scholar] [CrossRef] [PubMed]
- Carlier, A.; Uroz, S.; Smadja, B.; Fray, R.; Latour, X.; Dessaux, Y.; Faure, D. The Ti plasmid of Agrobacterium tumefaciens harbors an attM-paralogous gene, aiiB, also encoding N-Acyl homoserine lactonase activity. Appl. Environ. Microbiol. 2003, 69, 4989–4893. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Lee, S.J.; Oh, T.K.; Oh, J.W.; Koo, B.T.; Yum, D.Y.; Lee, J.K. AhlD, an N-acylhomoserine lactonase in Arthrobacter sp., and predicted homologues in other bacteria. Microbiology 2003, 149, 1541–1550. [Google Scholar] [CrossRef] [PubMed]
- Uroz, S.; Oger, P.M.; Chapelle, E.; Adeline, M.T.; Faure, D.; Dessaux, Y. A Rhodococcus qsdA-encoded enzyme defines a novel class of large-spectrum quorum-quenching lactonases. Appl. Environ. Microbiol. 2008, 74, 1357–1366. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.Z.; Morohoshi, T.; Ikenoya, M.; Someya, N.; Ikeda, T. AiiM, a novel class of N-acylhomoserine lactonase from the leaf-associated bacterium Microbacterium testaceum. Appl. Environ. Microbiol. 2010, 76, 2524–2530. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.B.; Wang, L.H.; Zhang, L.H. Genetic control of quorum-sensing signal turnover in Agrobacterium tumefaciens. Proc. Natl. Acad. Sci. USA 2002, 99, 4638–4643. [Google Scholar] [CrossRef] [PubMed]
- Gao, A.; Mei, G.Y.; Liu, S.; Wang, P.; Tang, Q.; Liu, Y.P.; Wen, H.; An, X.M.; Zhang, L.Q.; Yan, X.X.; et al. High-resolution structures of AidH complexes provide insights into a novel catalytic mechanism for N-acyl homoserine lactonase. Acta Crystallogr. D Biol. Crystallogr. 2013, 69, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Mei, G.Y.; Yan, X.X.; Turak, A.; Luo, Z.Q.; Zhang, L.Q. AidH, an alpha/beta-hydrolase fold family member from an Ochrobactrum sp. strain, is a novel N-acylhomoserine lactonase. Appl. Environ. Microbiol. 2010, 76, 4933–4942. [Google Scholar] [CrossRef] [PubMed]
- Tang, K.; Su, Y.; Brackman, G.; Cui, F.; Zhang, Y.; Shi, X.; Coenye, T.; Zhang, X.H. MomL, a novel marine-derived N-acyl homoserine lactonase from Muricauda olearia. Appl. Environ. Microbiol. 2015, 81, 774–782. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Lin, Y.; Yi, S.; Liu, P.; Shen, J.; Shao, Z.; Liu, Z. QsdH, a novel AHL lactonase in the RND-type inner membrane of marine Pseudoalteromonas byunsanensis strain 1A01261. PLoS ONE 2012, 7, e46587. [Google Scholar] [CrossRef] [PubMed]
- Tang, K.; Zhang, Y.; Yu, M.; Shi, X.; Coenye, T.; Bossier, P.; Zhang, X.H. Evaluation of a new high-throughput method for identifying quorum quenching bacteria. Sci. Rep. 2013, 3, 2935. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Xie, X.; Pashkov, I.; Sawaya, M.R.; Laidman, J.; Zhang, W.; Cacho, R.; Yeates, T.O.; Tang, Y. Directed evolution and structural characterization of a simvastatin synthase. Chem. Biol. 2009, 16, 1064–1074. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, R.F.; Holtman, M.A.; Zylstra, G.J.; White, J.F.; Kobayashi, D.Y. Taxonomic positioning of two biological control agents for plant diseases as Lysobacter enzymogenes based on phylogenetic analysis of 16S rDNA, fatty acid composition and phenotypic characteristics. J. Appl. Microbiol. 2003, 94, 1079–1086. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Qian, G.; Li, Y.; Wright, S.; Shen, Y.; Liu, F.; Du, L. Biosynthetic mechanism for sunscreens of the biocontrol agent Lysobacter enzymogenes. PLoS ONE 2013, 8, e66633. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Wright, S.; Shen, Y.; Du, L. Bioactive natural products from Lysobacter. Nat. Prod. Rep. 2012, 29, 1277–1287. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Qian, G.; Liu, F.; Li, Y.Z.; Shen, Y.; Du, L. Facile method for site-specific gene integration in Lysobacter enzymogenes for yield improvement of the anti-MRSA antibiotics WAP-8294A and the antifungal antibiotic HSAF. ACS Synth. Biol. 2013, 2, 670–678. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Li, Y.; Qian, G.; Wang, Y.; Chen, H.; Li, Y.Z.; Liu, F.; Shen, Y.; Du, L. Identification and characterization of the anti-methicillin-resistant Staphylococcus aureus WAP-8294A2 biosynthetic gene cluster from Lysobacter enzymogenes OH11. Antimicrob. Agents Chemother. 2011, 55, 5581–5589. [Google Scholar] [CrossRef] [PubMed]
- Lou, L.; Qian, G.; Xie, Y.; Hang, J.; Chen, H.; Zaleta-Rivera, K.; Li, Y.; Shen, Y.; Dussault, P.H.; Liu, F.; et al. Biosynthesis of HSAF, a tetramic acid-containing macrolactam from Lysobacter enzymogenes. J. Am. Chem. Soc. 2011, 133, 643–645. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Zaleta-Rivera, K.; Zhu, X.; Huffman, J.; Millet, J.C.; Harris, S.D.; Yuen, G.; Li, X.C.; Du, L. Structure and biosynthesis of heat-stable antifungal factor (HSAF), a broad-spectrum antimycotic with a novel mode of action. Antimicrob. Agents Chemother. 2007, 51, 64–72. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Wang, Y.; Tombosa, S.; Wright, S.; Huffman, J.; Yuen, G.; Qian, G.; Liu, F.; Shen, Y.; Du, L. Identification of a small molecule signaling factor that regulates the biosynthesis of the antifungal polycyclic tetramate macrolactam HSAF in Lysobacter enzymogenes. Appl. Microbiol. Biotechnol. 2015, 99, 801–811. [Google Scholar] [CrossRef] [PubMed]
- Di Cagno, R.; De Angelis, M.; Calasso, M.; Gobbetti, M. Proteomics of the bacterial cross-talk by quorum sensing. J. Proteom. 2011, 74, 19–34. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.H.; Zhang, L.H. Quorum sensing and quorum-quenching enzymes. J. Microbiol. 2005, 43, 101–109. [Google Scholar] [PubMed]
- Martino, P.D.; Fursy, R.; Bret, L.; Sundararaju, B.; Phillips, R.S. Indole can act as an extracellular signal to regulate biofilm formation of Escherichia coli and other indole-producing bacteria. Can. J. Microbiol. 2003, 49, 443–449. [Google Scholar] [CrossRef] [PubMed]
- Meiser, P.; Bode, H.B.; Muller, R. The unique DKxanthene secondary metabolite family from the myxobacterium Myxococcus xanthus is required for developmental sporulation. Proc. Natl. Acad. Sci. USA 2006, 103, 19128–19133. [Google Scholar] [CrossRef] [PubMed]
- Poplawsky, A.R.; Walters, D.M.; Rouviere, P.E.; Chun, W. A gene for a dioxygenase-like protein determines the production of the DF signal in Xanthomonas campestris pv. campestris. Mol. Plant. Pathol. 2005, 6, 653–657. [Google Scholar] [CrossRef] [PubMed]
- Stevens, A.M.; Queneau, Y.; Soulere, L.; von Bodman, S.; Doutheau, A. Mechanisms and synthetic modulators of AHL-dependent gene regulation. Chem. Rev. 2011, 111, 4–27. [Google Scholar] [CrossRef] [PubMed]
- Guzman-Maldonado, H.; Paredes-Lopez, O. Amylolytic enzymes and products derived from starch: A review. Crit. Rev. Food Sci. Nutr. 1995, 35, 373–403. [Google Scholar] [CrossRef] [PubMed]
- Song, Q.; Wang, Y.; Yin, C.; Zhang, X.H. LaaA, a novel high-active alkalophilic alpha-amylase from deep-sea bacterium Luteimonas abyssi XH031(T). Enzyme Microb. Technol. 2016, 90, 83–92. [Google Scholar] [CrossRef] [PubMed]
- Sibakov, M.; Palva, I. Isolation and the 5′-end nucleotide sequence of Bacillus licheniformis alpha-amylase gene. Eur. J. Biochem. 1984, 145, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Takkinen, K.; Pettersson, R.F.; Kalkkinen, N.; Palva, I.; Soderlund, H.; Kaariainen, L. Amino acid sequence of alpha-amylase from Bacillus amyloliquefaciens deduced from the nucleotide sequence of the cloned gene. J. Biol. Chem. 1983, 258, 1007–1013. [Google Scholar] [PubMed]
- Feller, G.; Payan, F.; Theys, F.; Qian, M.; Haser, R.; Gerday, C. Stability and structural analysis of alpha-amylase from the antarctic psychrophile Alteromonas haloplanctis A23. Eur. J. Biochem. 1994, 222, 441–447. [Google Scholar] [CrossRef] [PubMed]
- Hagihara, H.; Igarashi, K.; Hayashi, Y.; Endo, K.; Ikawa-Kitayama, K.; Ozaki, K.; Kawai, S.; Ito, S. Novel alpha-amylase that is highly resistant to chelating reagents and chemical oxidants from the alkaliphilic Bacillus isolate KSM-K38. Appl. Environ. Microb. 2001, 67, 1744–1750. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Zhang, Y.; Wang, Q.; Liu, J.; Wang, H.; Xue, Y.; Ma, Y. Gene cloning and characterization of a novel alpha-amylase from alkaliphilic Alkalimonas amylolytica. Biotechnol. J. 2006, 1, 1258–1265. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.P.; Morikawa, M.; Takagi, M.; Imanaka, T. Cloning of the aapT gene and characterization of its product, alpha-amylase-pullulanase (AapT), from thermophilic and alkaliphilic Bacillus sp. strain XAL601. Appl. Environ. Microbiol. 1994, 60, 3764–3773. [Google Scholar] [PubMed]
- Igarashi, K.; Hatada, Y.; Hagihara, H.; Saeki, K.; Takaiwa, M.; Uemura, T.; Ara, K.; Ozaki, K.; Kawai, S.; Kobayashi, T.; et al. Enzymatic properties of a novel liquefying alpha-amylase from an alkaliphilic Bacillus isolate and entire nucleotide and amino acid sequences. Appl. Environ. Microbiol. 1998, 64, 3282–3289. [Google Scholar] [PubMed]
- Zhang, J.W.; Zeng, R.Y. Purification and characterization of a cold-adapted alpha-amylase produced by Nocardiopsis sp. 7326 isolated from Prydz Bay, Antarctic. Mar. Biotechnol. 2008, 10, 75–82. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.U.; Gu, B.G.; Jeong, J.Y.; Byun, S.M.; Shin, Y.C. Purification and characterization of a maltotetraose-forming alkaline (alpha)-amylase from an alkalophilic Bacillus Strain, GM8901. Appl. Environ. Microbiol. 1995, 61, 3105–3112. [Google Scholar] [PubMed]
- Boyer, E.W.; Ingle, M.B. Extracellular alkaline amylase from a Bacillus species. J. Bacteriol. 1972, 110, 992–1000. [Google Scholar] [PubMed]
- Sharma, A.; Satyanarayana, T. Cloning and expression of acidstable, high maltose-forming, Ca2+-independent alpha-amylase from an acidophile Bacillus acidicola and its applicability in starch hydrolysis. Extremophiles 2012, 16, 515–522. [Google Scholar] [CrossRef] [PubMed]
- Prieto, J.A.; Bort, B.R.; Martinez, J.; Randezgil, F.; Buesa, C.; Sanz, P. Purification and characterization of a new alpha-amylase of intermediate thermal-stability from the yeast Lipomyces kononenkoae. Biochem. Cell Biol. 1995, 73, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Schwermann, B.; Pfau, K.; Liliensiek, B.; Schleyer, M.; Fischer, T.; Bakker, E.P. Purification, properties and structural aspects of a thermoacidophilic alpha-amylase from Alicyclobacillus acidocaldarius atcc 27009. Insight into acidostability of proteins. Eur. J. Biochem. 1994, 226, 981–991. [Google Scholar] [CrossRef] [PubMed]
- Asoodeh, A.; Chamani, J.; Lagzian, M. A novel thermostable, acidophilic alpha-amylase from a new thermophilic “Bacillus sp. Ferdowsicous” isolated from Ferdows hot mineral spring in Iran: Purification and biochemical characterization. Int. J. Biol. Macromol. 2010, 46, 289–297. [Google Scholar] [PubMed]
- Buonocore, V.; Caporale, C.; De Rosa, M.; Gambacorta, A. Stable, inducible thermoacidophilic alpha-amylase from Bacillus acidocaldarius. J. Bacteriol. 1976, 128, 515–521. [Google Scholar] [PubMed]
- Ali, I.; Akbar, A.; Anwar, M.; Prasongsuk, S.; Lotrakul, P.; Punnapayak, H. Purification and characterization of a polyextremophilic alpha-amylase from an obligate halophilic Aspergillus penicillioides isolate and its potential for souse with detergents. Biomed. Res. Int. 2015. [Google Scholar] [CrossRef] [PubMed]
- Xian, L.; Wang, F.; Luo, X.; Feng, Y.L.; Feng, J.X. Purification and characterization of a highly efficient calcium-independent alpha-amylase from Talaromyces pinophilus 1-95. PLoS ONE 2015, 10, e0121531. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Lu, Z.; Lu, M.; Qin, S.; Liu, H.; Deng, X.; Lin, Q.; Chen, J. Identification of archaeon-producing hyperthermophilic alpha-amylase and characterization of the alpha-amylase. Appl. Microbiol. Biotechnol. 2008, 80, 605–614. [Google Scholar] [CrossRef] [PubMed]
- Han, P.; Zhou, P.; Hu, S.Q.; Yang, S.Q.; Yan, Q.J.; Jiang, Z.Q. A novel multifunctional alpha-amylase from the thermophilic fungus Malbranchea cinnamomea: Biochemical characterization and three-dimensional structure. Appl. Biochem. Biotechnol. 2013, 170, 420–435. [Google Scholar] [CrossRef] [PubMed]
- Silva, T.M.; Damasio, A.R.; Maller, A.; Michelin, M.; Squina, F.M.; Jorge, J.A.; Polizeli Mde, L. Purification, partial characterization, and covalent immobilization-stabilization of an extracellular alpha-amylase from Aspergillus niveus. Folia Microbiol. 2013, 58, 495–502. [Google Scholar] [CrossRef] [PubMed]
- El-Sayed, A.K.; Abou Dobara, M.I.; El-Fallal, A.A.; Omar, N.F. Purification, sequencing, and biochemical characterization of a novel calcium-independent alpha-amylase AmyTVE from Thermoactinomyces vulgaris. Appl. Biochem. Biotechnol. 2013, 170, 483–497. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Morimoto, N.; Saburi, W.; Mukai, A.; Imoto, K.; Takehana, T.; Koike, S.; Mori, H.; Matsui, H. Purification and characterization of a liquefying alpha-amylase from alkalophilic thermophilic Bacillus sp. AAH-31. Biosci. Biotechnol. Biochem. 2012, 76, 1378–1383. [Google Scholar] [CrossRef] [PubMed]
- Michelin, M.; Silva, T.M.; Benassi, V.M.; Peixoto-Nogueira, S.C.; Moraes, L.A.; Leao, J.M.; Jorge, J.A.; Terenzi, H.F.; Polizeli Mde, L. Purification and characterization of a thermostable alpha-amylase produced by the fungus Paecilomyces variotii. Carbohydr. Res. 2010, 345, 2348–2353. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Wang, S.; Fang, Y.; Li, H.; Liu, S.; Liu, H. Cloning, expression, purification, and characterization of cold-adapted alpha-amylase from Pseudoalteromonas arctica GS230. Protein J. 2010, 29, 591–597. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.D.; Xu, Y. A novel raw starch digesting alpha-amylase from a newly isolated Bacillus sp. YX-1: Purification and characterization. Bioresour. Technol. 2008, 99, 4315–4320. [Google Scholar] [CrossRef] [PubMed]
- Uma Maheswar Rao, J.L.; Satyanarayana, T. Purification and characterization of a hyperthermostable and high maltogenic alpha-amylase of an extreme thermophile Geobacillus thermoleovorans. Appl. Biochem. Biotechnol. 2007, 142, 179–193. [Google Scholar] [CrossRef] [PubMed]
- Champreda, V.; Kanokratana, P.; Sriprang, R.; Tanapongpipat, S.; Eurwilaichitr, L. Purification, biochemical characterization, and gene cloning of a new extracellular thermotolerant and glucose tolerant maltooligosaccharide-forming alpha-amylase from an endophytic ascomycete Fusicoccum sp. BCC4124. Biosci. Biotechnol. Biochem. 2007, 71, 2010–2020. [Google Scholar] [CrossRef] [PubMed]
- Nagarajan, D.R.; Rajagopalan, G.; Krishnan, C. Purification and characterization of a maltooligosaccharide-forming alpha-amylase from a new Bacillus subtilis KCC103. Appl. Microbiol. Biotechnol. 2006, 73, 591–597. [Google Scholar] [CrossRef] [PubMed]
- Najafi, M.F.; Deobagkar, D.; Deobagkar, D. Purification and characterization of an extracellular alpha-amylase from Bacillus subtilis AX20. Protein Expr. Purif. 2005, 41, 349–354. [Google Scholar] [CrossRef] [PubMed]
- Mijts, B.N.; Patel, B.K. Cloning, sequencing and expression of an alpha-amylase gene, amyA, from the thermophilic halophile Halothermothrix orenii and purification and biochemical characterization of the recombinant enzyme. Microbiology 2002, 148, 2343–2349. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, K.; Bhattacharyya, B.K.; Sen, S.K. Purification and characterization of a thermostable alpha-amylase from Bacillus stearothermophilus. Folia Microbiol. 2000, 45, 207–210. [Google Scholar] [CrossRef]
- Egas, M.C.; da Costa, M.S.; Cowan, D.A.; Pires, E.M. Extracellular alpha-amylase from Thermus filiformis Ork A2: Purification and biochemical characterization. Extremophiles 1998, 2, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Marco, J.L.; Bataus, L.A.; Valencia, F.F.; Ulhoa, C.J.; Astolfi-Filho, S.; Felix, C.R. Purification and characterization of a truncated Bacillus subtilis alpha-amylase produced by Escherichia coli. Appl. Microbiol. Biotechnol. 1996, 44, 746–752. [Google Scholar] [PubMed]
- Shih, N.J.; Labbe, R.G. Purification and characterization of an extracellular alpha-amylase from Clostridium perfringens type A. Appl. Environ. Microbiol. 1995, 61, 1776–1779. [Google Scholar] [PubMed]
- Spiess, C.; Happersberger, H.P.; Glocker, M.O.; Spiess, E.; Rippe, K.; Ehrmann, M. Biochemical characterization and mass spectrometric disulfide bond mapping of periplasmic alpha-amylase MalS of Escherichia coli. J. Biol. Chem. 1997, 272, 22125–22133. [Google Scholar] [CrossRef] [PubMed]
- Mantsala, P.; Zalkin, H. Membrane-bound and soluble extracellular alpha-amylase from Bacillus subtilis. J. Biol. Chem. 1979, 254, 8540–8547. [Google Scholar] [PubMed]
- Buisson, G.; Duee, E.; Haser, R.; Payan, F. Three dimensional structure of porcine pancreatic alpha-amylase at 2.9 Å resolution. Role of calcium in structure and activity. EMBO J. 1987, 6, 3909–3916. [Google Scholar] [PubMed]
- Nielsen, J.E.; Borchert, T.V. Protein engineering of bacterial α-amylases. Biochim. Biophys. Acta 2000, 1543, 253–274. [Google Scholar] [CrossRef]
- Tang, S.Y.; Le, Q.T.; Shim, J.H.; Yang, S.J.; Auh, J.H.; Park, C.; Park, K.H. Enhancing thermostability of maltogenic amylase from Bacillus thermoalkalophilus ET2 by DNA shuffling. FEBS J. 2006, 273, 3335–3345. [Google Scholar] [CrossRef] [PubMed]
- Tomazic, S.J.; Klibanov, A.M. Mechanisms of irreversible thermal inactivation of Bacillus alpha-amylases. J. Biol. Chem. 1988, 263, 3086–3091. [Google Scholar] [PubMed]
- Vallee, B.L.; Stein, E.A.; Sumerwell, W.N.; Fischer, E.H. Metal content of alpha-amylases of various origins. J. Biol. Chem. 1959, 234, 2901–2905. [Google Scholar] [PubMed]
- Machius, M.; Declerck, N.; Huber, R.; Wiegand, G. Activation of Bacillus licheniformis alpha-amylase through a disorder-order transition of the substrate-binding site mediated by a calcium–sodium–calcium metal triad. Structure 1998, 6, 281–292. [Google Scholar] [CrossRef]
- Machius, M.; Wiegand, G.; Huber, R. Crystal structure of calcium-depleted Bacillus licheniformis alpha-amylase at 2.2 Å resolution. J. Mol. Biol. 1995, 246, 545–559. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Yang, J.; Li, J.; Long, L.; Xiao, Y.; Tian, X.; Wang, F.; Zhang, S. Role of two amino acid residues’ insertion on thermal stability of thermophilic alpha-amylase AMY121 from a deep sea bacterium Bacillus sp. SCSIO 15121. Bioprocess Biosyst. Eng. 2014, 38, 871–879. [Google Scholar] [CrossRef] [PubMed]
- Ghollasi, M.; Khajeh, K.; Naderi-Manesh, H.; Ghasemi, A. Engineering of a Bacillus alpha-amylase with improved thermostability and calcium independency. Appl. Biochem. Biotechnol. 2010, 162, 444–459. [Google Scholar] [CrossRef] [PubMed]
- Ben Ali, M.; Khemakhem, B.; Robert, X.; Haser, R.; Bejar, S. Thermostability enhancement and change in starch hydrolysis profile of the maltohexaose-forming amylase of Bacillus stearothermophilus US100 strain. Biochem. J. 2006, 394, 51–56. [Google Scholar] [CrossRef] [PubMed]
- Khemakhem, B.; Ben Ali, M.; Aghajari, N.; Juy, M.; Haser, R.; Bejar, S. The importance of an extra loop in the B-domain of an alpha-amylase from B. stearothermophilus US100. Biochem. Biophys. Res. Commun. 2009, 385, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Binter, A.; Staunig, N.; Jelesarov, I.; Lohner, K.; Palfey, B.A.; Deller, S.; Gruber, K.; Macheroux, P. A single intersubunit salt bridge affects oligomerization and catalytic activity in a bacterial quinone reductase. FEBS J. 2009, 276, 5263–5274. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.H.; Lu, F.P.; Li, Y.; Wang, J.L.; Gao, C. Acid stabilization of Bacillus licheniformis alpha amylase through introduction of mutations. Appl. Microbiol. Biotechnol. 2008, 80, 795–803. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Bai, A.; Gao, L.; Zhang, Z.; Zheng, B.; Feng, Y. Glu88 in the non-catalytic domain of acylpeptide hydrolase plays dual roles: Charge neutralization for enzymatic activity and formation of salt bridge for thermodynamic stability. Biochim. Biophys. Acta 2009, 1794, 94–102. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Liu, L.; Shin, H.D.; Chen, R.R.; Li, J.; Du, G.; Chen, J. Structure-based engineering of histidine residues in the catalytic domain of alpha-amylase from Bacillus subtilis for improved protein stability and catalytic efficiency under acidic conditions. J. Biotechnol. 2013, 164, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Haghani, K.; Khajeh, K.; Naderi-Manesh, H.; Ranjbar, B. Evidence regarding the hypothesis that the histidine-histidine contact pairs may affect protein stability. Int. J. Biol. Macromol. 2012, 50, 1040–1047. [Google Scholar] [CrossRef] [PubMed]
- Wong, T.Y.; Preston, L.A.; Schiller, N.L. ALGINATE LYASE: Review of major sources and enzyme characteristics, structure-function analysis, biological roles, and applications. Annu. Rev. Microbiol. 2000, 54, 289–340. [Google Scholar] [CrossRef] [PubMed]
- Tseng, C.-H.; Yamaguchi, K.; Kitamikado, M. Two types of alginate lyase from a marine bacterium Vibrio sp. Al-9. Nippon Suisan Gakkaishi 1992, 58, 743–749. [Google Scholar] [CrossRef]
- Tseng, C.-H.; Yamaguchi, K.; Kitamikado, M. Isolation and some properties of alginate lyase from a marine bacterium Vibrio sp. Al-128. Nippon Suisan Gakkaishi 1992, 58, 533–538. [Google Scholar] [CrossRef]
- Song, Y.; Yu, W.G.; Han, F.; Han, W.J.; Li, J.B. Purification and characterization of aginate lyase from marine bacterium Vibrio sp. QY101. Acta Biochim. Biophys. Sin. 2003, 35, 473–477. [Google Scholar] [PubMed]
- Han, F.; Gong, Q.H.; Song, K.; Li, J.B.; Yu, W.G. Cloning, sequence analysis and expression of gene alyVI encoding alginate lyase from marine bacterium Vibrio sp. QY101. DNA Seq. 2004, 15, 344–350. [Google Scholar] [CrossRef] [PubMed]
- Takeshita, S.; Sato, N.; Igarashi, M.; Muramatsu, T. A highly denaturant-durable alginate Lyase from a marine bacterium: Purification and properties. Biosci. Biotechnol. Biochem. 1993, 57, 1125–1128. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Yue, L.; Chi, Z.; Yu, W.; Madzak, C. The surface display of the alginate lyase on the cells of Yarrowia lipolytica for hydrolysis of alginate. Mar. Biotechnol. 2009, 11, 619–626. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Jiang, X.; Guan, H.; Wang, P.; Guo, H. Three alginate lyases from marine bacterium Pseudomonas fluorescens HZJ216: Purification and characterization. Appl. Biochem. Biotechnol. 2011, 164, 305–317. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.; Yang, J.; Zhang, X.Y.; Shi, M.; Song, X.Y.; Chen, X.L.; Zhang, Y.Z. Cultivable alginate lyase-excreting bacteria associated with the Arctic brown alga Laminaria. Mar. Drugs 2012, 10, 2481–2491. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; Inoue, A.; Tanaka, H.; Ojima, T. cDNA cloning of an alginate lyase from a marine gastropod Aplysia kurodai and assessment of catalytically important residues of this enzyme. Biochimie 2011, 93, 1720–1730. [Google Scholar] [CrossRef] [PubMed]
- MacDonald, L.C.; Berger, B.W. A polysaccharide lyase from Stenotrophomonas maltophilia with a unique, pH-regulated substrate specificity. J. Biol. Chem. 2014, 289, 312–325. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Uchimura, K.; Miyazaki, M.; Nogi, Y.; Horikoshi, K. A new high-alkaline alginate lyase from a deep-sea bacterium Agarivorans sp. Extremophiles 2009, 13, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Miyake, O.; Ochiai, A.; Hashimoto, W.; Murata, K. Origin and diversity of alginate lyases of families PL-5 and -7 in Sphingomonas sp. strain A1. J. Bacteriol. 2004, 186, 2891–2896. [Google Scholar] [CrossRef] [PubMed]
- Osawa, T.; Matsubara, Y.; Muramatsu, T.; Kimura, M.; Kakuta, Y. Crystal structure of the alginate (poly alpha-l-guluronate) lyase from Corynebacterium sp. at 1.2 Å resolution. J. Mol. Biol. 2005, 345, 1111–1118. [Google Scholar] [CrossRef] [PubMed]
- Duan, G.; Han, F.; Yu, W. Cloning, sequence analysis, and expression of gene alyPI encoding an alginate lyase from marine bacterium Pseudoalteromonas sp. CY24. Can. J. Microbiol. 2009, 55, 1113–1118. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, S.; Sahara, T.; Sato, D.; Kawasaki, K.; Ohgiya, S.; Inoue, A.; Ojima, T. Catalytically important amino-acid residues of abalone alginate lyase HdAly assessed by site-directed mutagenesis. Enzyme Microb. Technol. 2008, 43, 396–402. [Google Scholar] [CrossRef]
- Zhu, B.; Yin, H. Alginate lyase: Review of major sources and classification, properties, structure-function analysis and applications. Bioengineered 2015, 6, 125–131. [Google Scholar] [CrossRef] [PubMed]
- Matsushima, R.; Danno, H.; Uchida, M.; Ishihara, K.; Suzuki, T.; Kaneniwa, M.; Ohtsubo, Y.; Nagata, Y.; Tsuda, M. Analysis of extracellular alginate lyase and its gene from a marine bacterial strain, Pseudoalteromonas atlantica AR06. Appl. Microbiol. Biotechnol. 2010, 86, 567–576. [Google Scholar] [CrossRef] [PubMed]
- Matsushima, R.; Watanabe, R.; Tsuda, M.; Suzuki, T. Analysis of extracellular alginate lyase (alyA) expression and its regulatory region in a marine bacterial strain, Pseudoalteromonas atlantica AR06, using a gfp gene reporter system. Mar. Biotechnol. 2013, 15, 349–356. [Google Scholar] [CrossRef] [PubMed]
- Ashton, R.S.; Banerjee, A.; Punyani, S.; Schaffer, D.V.; Kane, R.S. Scaffolds based on degradable alginate hydrogels and poly(lactide-co-glycolide) microspheres for stem cell culture. Biomaterials 2007, 28, 5518–5525. [Google Scholar] [CrossRef] [PubMed]
- Hoiby, N.; Bjarnsholt, T.; Givskov, M.; Molin, S.; Ciofu, O. Antibiotic resistance of bacterial biofilms. Int. J. Antimicrob. Agents 2010, 35, 322–332. [Google Scholar] [CrossRef] [PubMed]
- Iehata, S.; Inagaki, T.; Okunishi, S.; Nakano, M.; Tanaka, R.; Maeda, H. Improved gut environment of abalone Haliotis gigantea through Pediococcus sp Ab1 treatment. Aquaculture 2010, 305, 59–65. [Google Scholar] [CrossRef]
- Younes, I.; Rinaudo, M. Chitin and chitosan preparation from marine sources. Structure, properties and applications. Mar. Drugs 2015, 13, 1133–1174. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Shao, B.; Fu, H.; Rao, P. Isolation of a thermostable legume chitinase and study on the antifungal activity. Appl. Microbiol. Biotechnol. 2009, 85, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Hirono, I.; Yamashita, M.; Aoki, T. Note: Molecular cloning of chitinase genes from Vibrio anguillarum and V. parahaemolyticus. J. Appl. Microbiol. 1998, 84, 1175–1178. [Google Scholar] [CrossRef] [PubMed]
- Aunpad, R.; Panbangred, W. Cloning and characterization of the constitutively expressed chitinase C gene from a marine bacterium, Salinivibrio costicola strain 5SM-1. J. Biosci. Bioeng. 2003, 96, 529–536. [Google Scholar] [CrossRef]
- Howard, M.B.; Ekborg, N.A.; Taylor, L.E.; Weiner, R.M.; Hutcheson, S.W. Genomic analysis and initial characterization of the chitinolytic system of Microbulbifer degradans strain 2-40. J. Bacteriol. 2003, 185, 3352–3360. [Google Scholar] [CrossRef] [PubMed]
- Tsujibo, H.; Kubota, T.; Yamamoto, M.; Miyamoto, K.; Inamori, Y. Characterization of chitinase genes from an alkaliphilic actinomycete, Nocardiopsis prasina OPC-131. Appl. Environ. Microbiol. 2003, 69, 894–900. [Google Scholar] [CrossRef] [PubMed]
- Fukamizo, T. Chitinolytic enzymes: Catalysis, substrate binding, and their application. Curr. Protein Pept. Sci. 2000, 1, 105–124. [Google Scholar] [CrossRef] [PubMed]
- Horn, S.J.; Sorbotten, A.; Synstad, B.; Sikorski, P.; Sorlie, M.; Varum, K.M.; Eijsink, V.G. Endo/exo mechanism and processivity of family 18 chitinases produced by Serratia marcescens. FEBS J. 2006, 273, 491–503. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Yang, B.; Zhang, F.; Miao, X.; Li, Z. Characterization of antifungal chitinase from marine Streptomyces sp. DA11 associated with South China Sea sponge Craniella australiensis. Mar. Biotechnol. 2009, 11, 132–140. [Google Scholar] [CrossRef] [PubMed]
- Stefanidi, E.; Vorgias, C.E. Molecular analysis of the gene encoding a new chitinase from the marine psychrophilic bacterium Moritella marina and biochemical characterization of the recombinant enzyme. Extremophiles 2008, 12, 541–552. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.L.; Liang, T.W.; Lin, B.S.; Wang, C.L.; Wu, P.C.; Liu, J.R. Purification and characterization of chitinase from a new species strain Pseudomonas sp. TKU008. J. Microbiol. Biotechnol. 2010, 20, 1001–1005. [Google Scholar] [PubMed]
- Suginta, W.; Songsiriritthigul, C.; Kobdaj, A.; Opassiri, R.; Svasti, J. Mutations of Trp275 and Trp397 altered the binding selectivity of Vibrio carchariae chitinase A. Biochim. Biophys. Acta 2007, 1770, 1151–1160. [Google Scholar] [CrossRef] [PubMed]
- De Assis, C.F.; Costa, L.S.; Melo-Silveira, R.F.; Oliveira, R.M.; Pagnoncelli, M.G.; Rocha, H.A.; De Macedo, G.R.; Santos, E.S. Chitooligosaccharides antagonize the cytotoxic effect of glucosamine. World J. Microbiol. Biotechnol. 2012, 28, 1097–1105. [Google Scholar] [CrossRef] [PubMed]
- Chi, Z.; Zhang, T.; Liu, G.; Li, J.; Wang, X. Production, characterization and gene cloning of the extracellular enzymes from the marine-derived yeasts and their potential applications. Biotechnol. Adv. 2009, 27, 236–255. [Google Scholar] [CrossRef] [PubMed]
- Mba Medie, F.; Davies, G.J.; Drancourt, M.; Henrissat, B. Genome analyses highlight the different biological roles of cellulases. Nat. Rev. Microbiol. 2012, 10, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, M.D.; Reeves, R.A.; Farrington, G.K.; Anderson, P.; Williams, D.P.; Bergquist, P.L. Multidomain and multifunctional glycosyl hydrolases from the extreme thermophile Caldicellulosiruptor isolate Tok7B.1. Curr. Microbiol. 2000, 40, 333–340. [Google Scholar] [CrossRef] [PubMed]
- Bronnenmeier, K.; Kern, A.; Liebl, W.; Staudenbauer, W.L. Purification of Thermotoga maritima enzymes for the degradation of cellulosic materials. Appl. Environ. Microbiol. 1995, 61, 1399–1407. [Google Scholar] [PubMed]
- Hakamada, Y.; Koike, K.; Yoshimatsu, T.; Mori, H.; Kobayashi, T.; Ito, S. Thermostable alkaline cellulase from an alkaliphilic isolate, Bacillus sp. KSM-S237. Extremophiles 1997, 1, 151–156. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.-H.; Kim, B.-K.; Lee, Y.-J.; Chung, C.-H.; Lee, J.-W. Industrial scale of optimization for the production of carboxymethyl cellulase from rice bran by a marine bacterium, Bacillus subtilis subsp. subtilis A-53. Enzyme Microb. Technol. 2010, 46, 38–42. [Google Scholar] [CrossRef]
- Alfredsson, G.A.; Kristjansson, J.K.; Hjorleifsdottir, S.; Stetter, K.O. Rhodothermus marinus, gen. nov., sp. nov., a thermophilic, halophilic bacterium from submarine hot springs in iceland. J. Gen. Microbiol. 1988, 134, 299–306. [Google Scholar] [CrossRef]
- Trivedi, N.; Gupta, V.; Kumar, M.; Kumari, P.; Reddy, C.R.K.; Jha, B. An alkali-halotolerant cellulase from Bacillus flexus isolated from green seaweed Ulva lactuca. Carbohydr. Polym. 2011, 83, 891–897. [Google Scholar] [CrossRef]
- Fang, Z.; Fang, W.; Liu, J.; Hong, Y.; Peng, H.; Zhang, X.; Sun, B.; Xiao, Y. Cloning and characterization of a beta-glucosidase from marine microbial metagenome with excellent glucose tolerance. J. Microbiol. Biotechnol. 2010, 20, 1351–1358. [Google Scholar] [CrossRef] [PubMed]
| QQ Enzyme | Length (aa) | Predictable Domains | Signal Peptide | Host Organisms | Origin | Structure | Reference |
|---|---|---|---|---|---|---|---|
| AiiA | 231aa | Beta-lactamase family (15–216) | No signal | Bacillus | terrestrial | 3DHB | [18] |
| AiiB | 276aa | Beta-lactamase family (42–259) | No signal | Agrobacterium | terrestrial | unknown | [22] |
| AttM | 295aa | Beta-lactamase family (78–282) | 1–17 | Agrobacterium | terrestrial | unknown | [25] |
| QsdA | 323aa | Phosphotriesterase family (11–322) | No signal | Rhodococcus | terrestrial | unknown | [24] |
| AidH | 279aa | Alpha/beta hydrolase (25–147) | No signal | Ochrobactrum | terrestrial | unknown | [27,28] |
| GKL | 330aa | Phosphotriesterase family (16–329) | No signal | Geobacillus | terrestrial | unknown | [26] |
| MomL | 293aa | Beta-lactamase family (72–277) | 1–21 | Muricauda | oceanic | unknown | [29] |
| QsdH | 968aa | AcrB/AcrD/AcrF family (182–964) | 1–23 | Pseudoalteromonas | oceanic | unknown | [30] |
| Stain | UniProtKB | Molecular Mass (kDa) | Signal Peptide (aa) | Temperature Optimum (°C) | Thermostabiliy | pH Optimum | pH Stability | Specific Activity with Soluble Starch (U/mg) | References |
|---|---|---|---|---|---|---|---|---|---|
| Luteimonas abyssi | NM | 49 | 35 | 50 | 34%, 50 °C, 20 min | 9 | >50%, 6–11, 50 °C, 1 h | 8881 a | [48] |
| Bacillus licheniformis | Q208A7 | 55 | 29 | 90 | Clear halos, 100 °C, 120 min | NM | NM | NM | [49] |
| Bacillus amyloliquefaciens | P00692 | 54.8 | 31 | 60 | NM | NM | NM | NM | [50] |
| Alteromonas haloplanctis A23 | P29957 | 50 | 24 | 25 | 6%, 25 °C | 7 | NM | NM | [51] |
| Bacillus sp. strain KSM-K38 | Q93I48 | 55 | 21 | 55–60 | 20%, 50 °C, 30 min | 8.0–9.5 | >80%, 6–11, 40 °C, 30 min | 4221 a | [52] |
| alkaliphilic bacterium N10 | Q6WUB6 | 61 | 31 | 50 | 71%, 50 °C, 30 min | 9.5 | >80%, 8.5–11, 50 °C, 10 min | 7826 a | [53] |
| Bacillus sp. XAL601 | Q45643 | 225 | 31 | 70 | NM | 9.0 | NM | 57.3 a | [54] |
| Bacillus sp. | O82839 | 53 | 31 | 55 | 25%, 80 °C, 10 min | 8.0–8.5 | >50%, 6–9, 40 °C, 30 min | 5009 a | [55] |
| Nocardiopsis sp. 7326 | NM | 55 | NM | 35 | 18%, 55 °C, 30 min | 8.0 | >60%, 7–9, 4 °C, 24 h | 548 a | [56] |
| Bacillus sp. strain GM8901 | NM | 97 | NM | 60 | 37%, 60 °C, 2 h (−Ca), 78%, 60 °C, 2 h (+Ca) | 11–12 | >85%, 6–13, 50 °C, 1 h | 157.5 a | [57] |
| Bacillus sp. NRRL B-3881 | NM | NM | NM | 50 | 50%, 55 °C | 9.2 | >50%, 7.0–10.5 | 3485 a | [58] |
| Bacillus acidicola | J9PQD2 | 62 | no signal | 60 | 50%, 90 °C, 10 min | 4 | 100%, 4, 12 h, 100%, 3, 1 h | 1166 a | [59] |
| Lipomyces kononenkoae | Q01117 | 76 | 28 | 70 | 0, 70 °C, 10 min | 4.5–5.0 | >70%, 3–8, 1 h | 258 a | [60] |
| Alicyclobacillus acidocaldarius | C8WUR2 | 160 | 23 | 75 | NM | 3 | NM | 16.9 b | [61] |
| Bacillus sp. Ferdowsicous | P86331 | 53 | NM | 70 | 75%, 75 °C, 45 min | 4.5 | >75%, 3.5–6, 60 min | 267 a | [62] |
| Bacillus acidocaldarius | NM | 68 | NM | 75 | 50%, 60 °C, 5 days | 3.5 | Stable below 4.5 | 257 b | [63] |
| Aspergillus penicillioides | NM | 42 | NM | 80 | 60%, 100 °C | 9 | >80%, 7–10 | 118.42 a | [64] |
| Talaromyces pinophilus 1–95 | NM | 58 | NM | 55 | <45 °C, 1 h | 4–5 | 5–9.5, 24 h | 673.08 a | [65] |
| Thermococcus sp. HJ21 | B4X9V8 | 51.4 | NM | 95 | 50%, 90 °C, 5 h, 40%, 30%; 100 °C; 2 h, 3 h | 5 | 5–9 | 8.3 a | [66] |
| Malbranchea cinnamomea | K9L8F3 | 60.3 | 21 | 65 | 50%, 60 °C, 41.1 min | 6.5 | >90%, 5–10, 30 min | 514.6 a | [67] |
| Aspergillus niveus | NM | 60 | NM | 65 | 50%, 70 °C, 20 m | 6 | 4–7, 24 h | 168 a | [68] |
| Thermoactinomyces vulgaris | G8ZE61 | 40.6 | NM | 50 | 50%, 50 °C, 2 h | 6–7 | 4–9 | 127,100.33 b | [69] |
| Bacillus sp. AAH-31 | S6BGD1 | 91 | 28 | 70 | <60 °C | 8.5 | 6.4–10.3 | 16.7 a | [70] |
| Paecilomyces variotii | NM | 75 | NM | 60 | 50%, 60 °C, 53 min | 4 | >70%, 5–8, 1 h | 612.5 a | [71] |
| Pseudoalteromonas arctica GS230 | NM | 55 | 24 | 30 | 49%, 30 °C, 150 min | 7.5 | >60%, 7–8.5, 1 h | 25.5 a | [72] |
| Bacillus sp. YX-1 | A9YDD9 | 56 | 31 | 40–50 | 60%, 60 °C, 1 h | 5 | >80%, 4.5–11, 1 h | 607 b | [73] |
| Geobacillus thermoleovorans | NM | 26 | NM | 100 | 50%, 100 °C, 3.6 h | 8 | 50%, 6, 4.5 h, 50%, 7, 7.5 h | 450 a | [74] |
| Fusicoccum sp. BCC4124 | Q0Z8K1 | 50 | no signal | 70 | 95%, 50 °C, 1 h | 7 | NM | 90 a | [75] |
| Bacillus subtilis KCC103 | A8VWC5 | 53 | 33 | 65–70 | 50%, 70 °C, 7 min | 6–7 | >98%, 5–9.5 | 483 a | [76] |
| Bacillus subtilis AX20 | NM | 149 | NM | 55 | 50 °C, 30 min | 6 | 5–9, 24 h | 4133 a | [77] |
| Halothermothrix orenii | Q8GPL8 | 60 | 23 | 65 | 37–75 °C | 7.5 | 6–9.5 | 22.32 a | [78] |
| Bacillus stearothermophilus | NM | 64 | NM | 50 | 92%, 100 °C, 1 h | 7 | 23%, 3, 1 h, 26%, 10, 1 h | 77.2 b | [79] |
| Thermus filiformis Ork A2 | NM | 60 | NM | 95 | 50%, 95 °C, 19 min | 5.5–6 | >80%, 4.5–8, 1 h | 6352 b | [80] |
| Bacillus subtilis | NM | 48 | NM | 50 | 70%, 60 °C, 1 h | 6.5 | 5–6.5 | 772.7 a | [81] |
| Clostridium perfringens NCTC 8679 | NM | 76 | NM | 30 | NM | 6.5 | NM | NM | [82] |
| Escherichia coli (strain K12) | P25718 | 75.7 | 17 | NM | NM | 8 | NM | NM | [83] |
| Bacillus subtilis | P00691 | 67 | 27 | NM | NM | 8.5 | NM | NM | [84] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Y.; Song, Q.; Zhang, X.-H. Marine Microbiological Enzymes: Studies with Multiple Strategies and Prospects. Mar. Drugs 2016, 14, 171. https://doi.org/10.3390/md14100171
Wang Y, Song Q, Zhang X-H. Marine Microbiological Enzymes: Studies with Multiple Strategies and Prospects. Marine Drugs. 2016; 14(10):171. https://doi.org/10.3390/md14100171
Chicago/Turabian StyleWang, Yan, Qinghao Song, and Xiao-Hua Zhang. 2016. "Marine Microbiological Enzymes: Studies with Multiple Strategies and Prospects" Marine Drugs 14, no. 10: 171. https://doi.org/10.3390/md14100171
APA StyleWang, Y., Song, Q., & Zhang, X.-H. (2016). Marine Microbiological Enzymes: Studies with Multiple Strategies and Prospects. Marine Drugs, 14(10), 171. https://doi.org/10.3390/md14100171





