Identification and Bioactivity of Compounds from the Fungus Penicillium sp. CYE-87 Isolated from a Marine Tunicate
Abstract
:1. Introduction

2. Results and Discussion
2.1. Purification of the Compounds 1–6
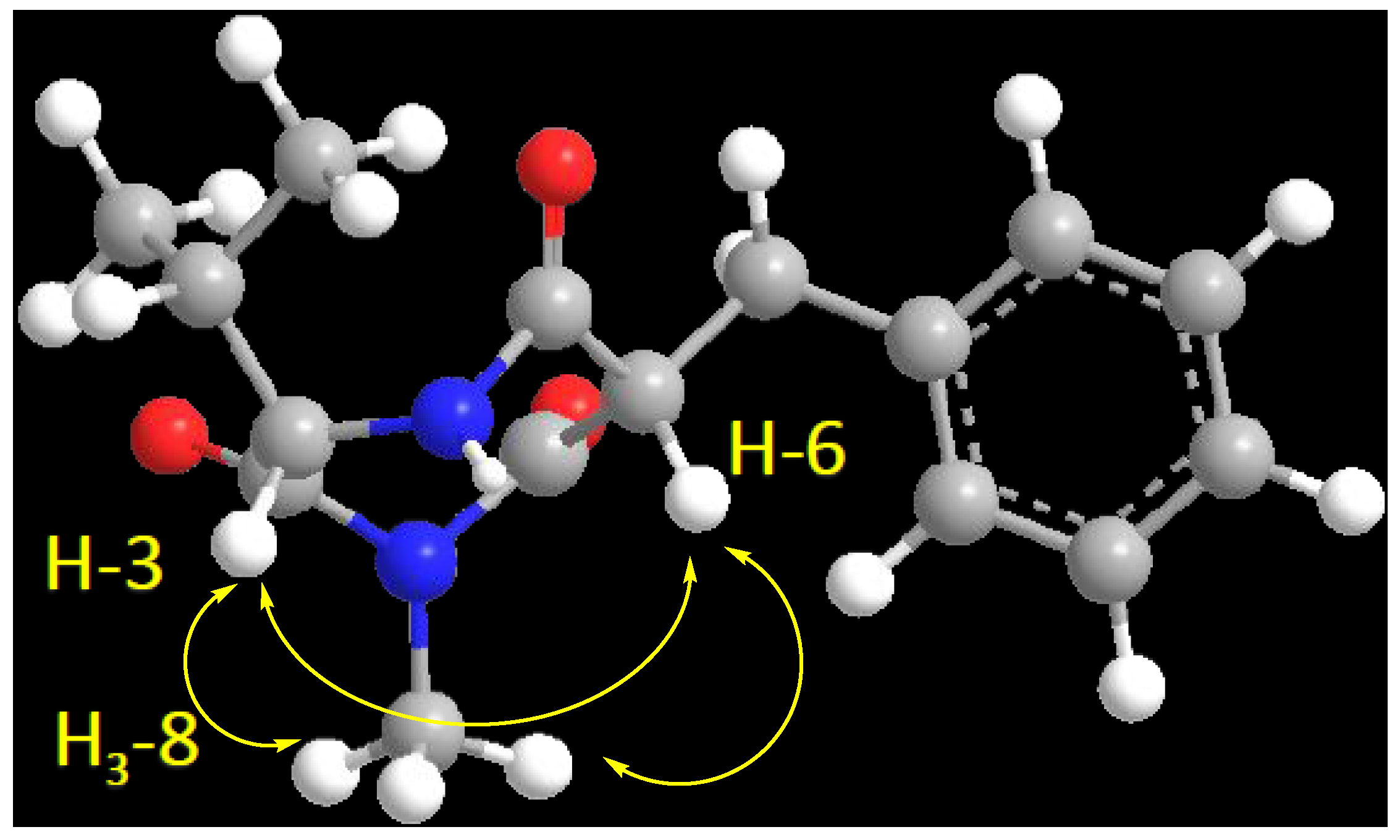
2.2. Structure Elucidation of the Compounds 1–6
| Position | δC, Type a | δH, m (J in Hz) | HMBC | NOESY |
|---|---|---|---|---|
| 2 | 173.0, C b | |||
| 3 | 60.1, CH | 4.18, d (4.8) | C-2, C-5, C-1′ | H-6, H3-8, H-1′ |
| 5 | 173.2, C | |||
| 6 | 56.4, CH | 4.68, dd (8.4, 6.6) | C-5, C-7, C-1″ | H-3, H3-8, H2-1″ |
| 7 | 173.1, C b | |||
| 8 | 22.9, CH3 | 1.90, s | C-2, C-7 | |
| 1′ | 32.2, CH | 2.07, m | C-2, C-3, C-2′, C-3′ | H-3 |
| 2′ | 19.8, CH3 | 0.81, d (7.2) | C-1′, C-3 | |
| 3′ | 18.2, CH3 | 0.78, d (7.2) | C-1′, C-3 | |
| 1″ | 39.0, CH2 | 3.11, dd (13.8, 6.6) 2.88, dd (13.8, 8.4) | C-2″, C-3″, C-6, C-7 | H-6 |
| 2″ | 138.6, C | |||
| 3″ | 130.2, CH | 7.23, m | C-2″, C-5″ | |
| 4″ | 129.4, CH | 7.24, m | C-2″ | |
| 5″ | 127.7, CH | 7.14, m | C-3″, C-7″ | |
| 6″ | 129.4, CH | 7.24, m | C-2″ | |
| 7″ | 130.2, CH | 7.23, m | C-1″, C-5″ |

2.3. Biological Activities of Compounds 1–6
| Compound | IC50 (μM) | |
|---|---|---|
| Antimigratory Activity (MDA-MB-231) | Antiproliferative Activity (HeLa Cells) | |
| 1 | >50 | >50 |
| 2 | 16.5 | >50 |
| 3 | >50 | >50 |
| 4 | >50 | >50 |
| 5 | >50 | >50 |
| 6 | 17.6 | >50 |
| S-Ethyl * | 43.4 | NT |
| Paclitaxel * | NT | 0.0017 |
3. Experimental Section
3.1. General Experimental Procedures
3.2. Biological Materials
3.2.1. Collection of the Host Tunicate, Didemnum sp.
3.2.2. Preparation of the Fungal Isolate
3.2.3. Extraction of Genome DNA from Cultured Fungal Isolate CYE-87
3.2.4. Amplification of Fungal ITS-rDNA Fragments of Isolate CYE-87
3.2.5. Sequence Fungal ITS-rDNA Regions of Isolate CYE-87
3.2.6. Characterization of the Fungal Isolate CYE-87
3.3. Culture Condition and Extraction
3.4. Isolation and Purification of Compounds 1–6
3.5. Absolute Stereochemistry of the Amino Acids in 2 and 6
3.6. Biological Activities of the Compounds
3.6.1. Evaluation of the Antimigratory of 1–6 Using the Wound Healing Assay
3.6.2. Evaluation of Antiproliferative and Cytotoxic Activities
3.6.3. Evaluation of the Antimicrobial Activity of Compounds 1–6
3.6.3.1. Determination of the Antimicrobial Activities Using the Disc Diffusion Assay
3.6.3.2. Determination of the MIC Values against C. albicans
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 2012, 75, 311–335. [Google Scholar] [CrossRef] [PubMed]
- Harvey, A.L. Natural products in drug discovery. Drug Discov. Today 2008, 13, 894–901. [Google Scholar] [CrossRef] [PubMed]
- Blunt, J.W.; Copp, B.R.; Keyzers, R.A.; Munro, M.H.G.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2012, 29, 144–222. [Google Scholar] [CrossRef] [PubMed]
- Blunt, J.W.; Copp, B.R.; Munro, M.H.G.; Northcote, P.T.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2010, 27, 165–237. [Google Scholar] [CrossRef] [PubMed]
- Imhoff, J.F.; Stöhr, R. Sponge-associated bacteria: General overview and special aspects of bacteria associated with Halichondria panicea. Prog. Mol. Subcell. Biol. 2003, 37, 35–57. [Google Scholar] [PubMed]
- Gao, Z.; Li, B.; Zheng, C.; Wang, G. Molecular detection of fungal communities in the Hawaiian marine sponges Suberites zeteki and Mycale armata. Appl. Environ. Microbiol. 2008, 74, 6091–6101. [Google Scholar] [CrossRef] [PubMed]
- Siepmann, R.; Höhnk, W. Über Hefen und einige Pilze (Fungi imp., Hyphales) aus dem Nordatlantik. Veröff. Inst. Meeresforsch. Bremerh. 1962, 8, 79–97. [Google Scholar]
- Roth, F.J., Jr.; Ahearn, D.G.; Fell, J.W.; Meyers, S.P.; Meyer, S.A. Ecology and taxonomy of yeasts isolated from various marine substrates. Limnol. Oceanogr. 1962, 7, 178–185. [Google Scholar] [CrossRef]
- Sponga, F.; Cavaletti, L.; Lazzaroni, A.; Borghi, A.; Ciciliato, I.; Losi, D.; Marinelli, F. Biodiversity and potentials of marine-derived microorganisms. J. Biotechnol. 1999, 70, 65–69. [Google Scholar] [CrossRef]
- Höller, U.; Wright, A.D.; Matthée, G.F.; König, G.M.; Draeger, S.; Aust, H.J.; Schulz, B. Fungi from marine sponges: Diversity, biological activity and secondary metabolites. Mycol. Res. 2000, 104, 1354–1365. [Google Scholar] [CrossRef]
- Namikoshi, M.; Akano, K.; Kobayashi, H.; Koike, Y.; Kitazawa, A.; Rondonuwu, A.B.; Pratasik, S.B. Distribution of marine filamentous fungi associated with marine sponges in coral reefs of Palau and Bunaken Island, Indonesia. J. Tokyo Univ. Fish. 2002, 88, 15–20. [Google Scholar]
- Morrison-Gardiner, S. Dominant fungi from Australian coral reefs. Fung. Divers. 2002, 9, 105–121. [Google Scholar]
- Proksch, P.; Ebel, R.; Edrada, R.A.; Riebe, F.; Liu, H.; Diesel, A.; Bayer, M.; Li, X.; Lin, W.H.; Grebenyuk, V.; et al. Sponge-associated fungi and their bioactive compounds: The Suberites case. Bot. Mar. 2008, 51, 209–218. [Google Scholar]
- Paz, Z.; Komon-Zelazowska, M.; Druzhinina, I.S.; Aveskamp, M.M.; Schnaiderman, A.; Aluma, A.; Carmeli, S.; Ilan, M.; Yarden, O. Diversity and potential antifungal properties of fungi associated with a Mediterranean sponge. Fung. Divers. 2010, 42, 17–26. [Google Scholar] [CrossRef]
- Bugni, T.S.; Ireland, C.M. Marine-derived fungi: A chemically and biologically diverse group of microorganisms. Nat. Prod. Rep. 2004, 21, 143–163. [Google Scholar] [CrossRef] [PubMed]
- Shang, Z.; Li, X.; Meng, L.; Li, C.; Gao, S.; Huang, C.; Wang, B. Chemical profile of the secondary metabolites produced by a deep-sea sediment-derived fungus Penicillium commune SD-118. Chin. J. Oceanol. Limnol. 2012, 30, 305–314. [Google Scholar]
- Li, H.-J.; Cai, Y.-T.; Chen, Y.-Y.; Lam, C.-K.; Lan, W.-J. Metabolites of marine fungus Aspergillus sp. collected from soft coral Sarcophyton tortuosum. Chem. Res. Chin. Univ. 2010, 26, 415–419. [Google Scholar]
- Shen, Y.; Zou, J.; Xie, D.; Ge, H.; Cao, X.; Dai, J. Butyrolactone and cycloheptanetrione from mangrove-associated fungus Aspergillus terreus. Chem. Pharm. Bull. 2012, 60, 1437–1441. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Wang, G. Diversity of fungal isolates from three Hawaiian marine sponges. Microbiol. Res. 2009, 164, 233–241. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. In PCR Protocols: A Guide to Methods and Application; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- National Center for Biotechnology Information. Available online: http://www.ncbi.nlm.nih.gov (accessed on 25 January 2015).
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The ClustalX windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A user friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Sallam, A.A.; Mohyeldin, M.M.; Foudah, A.I.; Akl, M.R.; Nazzal, S.M.; Meyer, S.A.; Liu, Y.-Y.; El Sayed, K.A. Marine natural products-inspired phenylmethylene hydantoins with potent in vitro and in vivo antitumor activities via suppression of Brk and FAK signaling. Org. Biomol. Chem. 2014, 12, 5295–5303. [Google Scholar] [CrossRef] [PubMed]
- Boyd, M.R.; Paull, K.D. Some practical considerations and applications of the National Cancer Institute in vitro anticancer discovery screen. Drug Dev. Res. 1995, 34, 91–109. [Google Scholar] [CrossRef]
- Skehan, P.; Storeng, R.; Scudiero, D.; Monks, A.; McMahon, J.; Vistica, D.; Warren, J.T.; Bokesch, H.; Kenney, S.; Boyd, M.R. New colorimetric cytotoxicity assay for anticancer-drug screening. J. Natl. Cancer Inst. 1990, 82, 1107–1112. [Google Scholar] [CrossRef] [PubMed]
- Risinger, A.L.; Jackson, E.M.; Polin, L.A.; Helms, G.L.; LeBoeuf, D.A.; Joe, P.A.; Hopper-Borge, E.; Ludueña, R.F.; Kruh, G.D.; Mooberry, S.L. The taccalonolides: Microtubule stabilizers that circumvent clinically relevant taxane resistance mechanisms. Cancer Res. 2008, 68, 8881–8888. [Google Scholar] [CrossRef]
- Kiehlbauch, J.A.; Hannett, G.E.; Salfinger, M.; Archinal, W.; Monserrat, C.; Carlyn, C. Use of the national committee for clinical laboratory standards guidelines for disk diffusion susceptibility testing in New York state laboratories. J. Clin. Microbiol. 2000, 38, 3341–3348. [Google Scholar]
- Reference Method for Broth Dilution Antifungal Susceptibility Testing of Yeast, Approved Standard, 3rd ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2008; Document M27-A3.
- Reference Method for Broth Dilution Antifungal Susceptibility Testing of Filamentous Fungi, Approved Standard, 2nd ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2008; Document M38-A2.
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shaala, L.A.; Youssef, D.T.A. Identification and Bioactivity of Compounds from the Fungus Penicillium sp. CYE-87 Isolated from a Marine Tunicate. Mar. Drugs 2015, 13, 1698-1709. https://doi.org/10.3390/md13041698
Shaala LA, Youssef DTA. Identification and Bioactivity of Compounds from the Fungus Penicillium sp. CYE-87 Isolated from a Marine Tunicate. Marine Drugs. 2015; 13(4):1698-1709. https://doi.org/10.3390/md13041698
Chicago/Turabian StyleShaala, Lamiaa A., and Diaa T. A. Youssef. 2015. "Identification and Bioactivity of Compounds from the Fungus Penicillium sp. CYE-87 Isolated from a Marine Tunicate" Marine Drugs 13, no. 4: 1698-1709. https://doi.org/10.3390/md13041698
APA StyleShaala, L. A., & Youssef, D. T. A. (2015). Identification and Bioactivity of Compounds from the Fungus Penicillium sp. CYE-87 Isolated from a Marine Tunicate. Marine Drugs, 13(4), 1698-1709. https://doi.org/10.3390/md13041698








