Marine Microorganism-Invertebrate Assemblages: Perspectives to Solve the “Supply Problem” in the Initial Steps of Drug Discovery
Abstract
:1. Introduction
2. Marine Microorganism-Invertebrate Assemblages
3. The Supply Problem in Marine Drug Discovery
4. Aquaculture of Marine Invertebrates
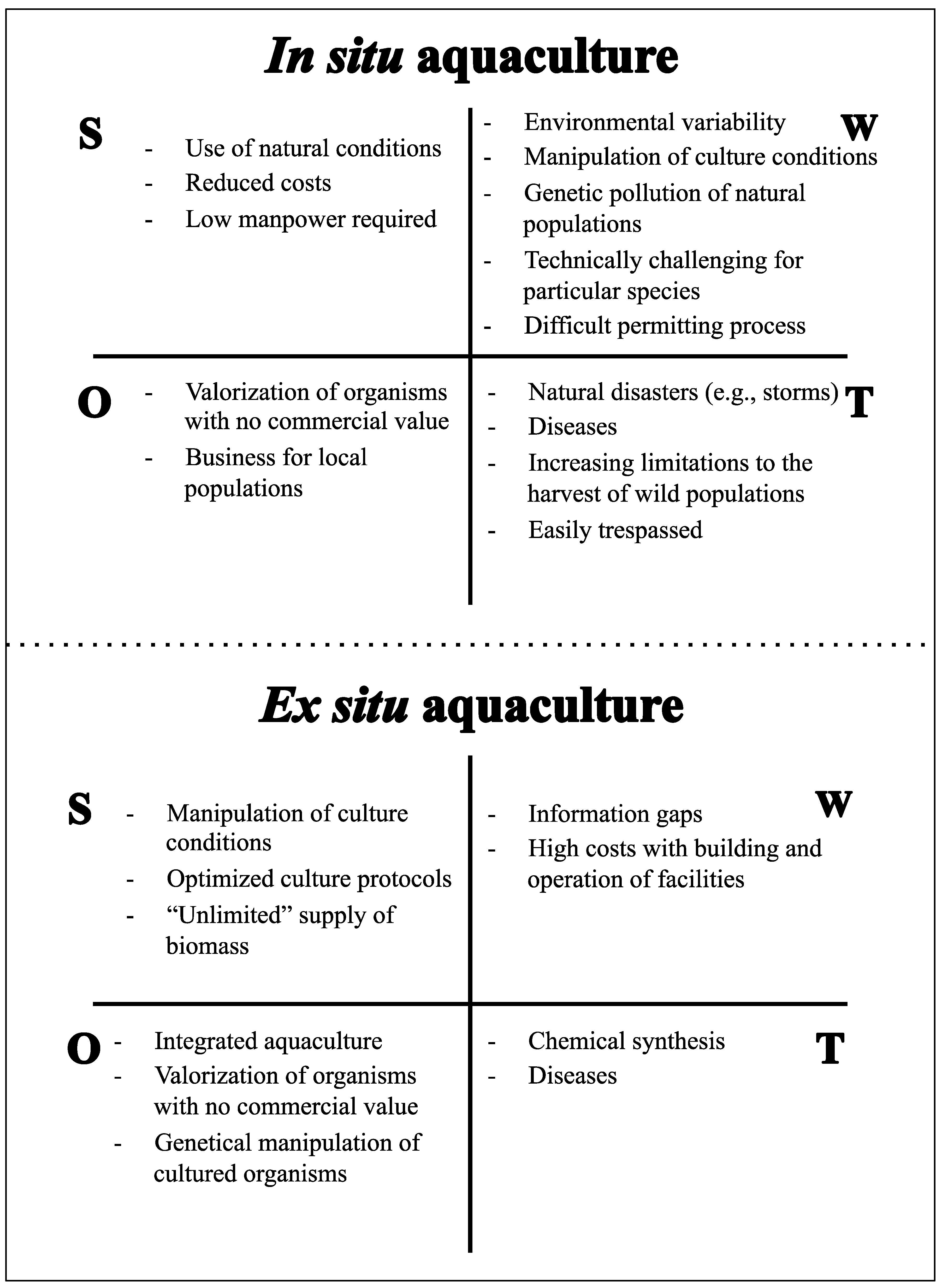
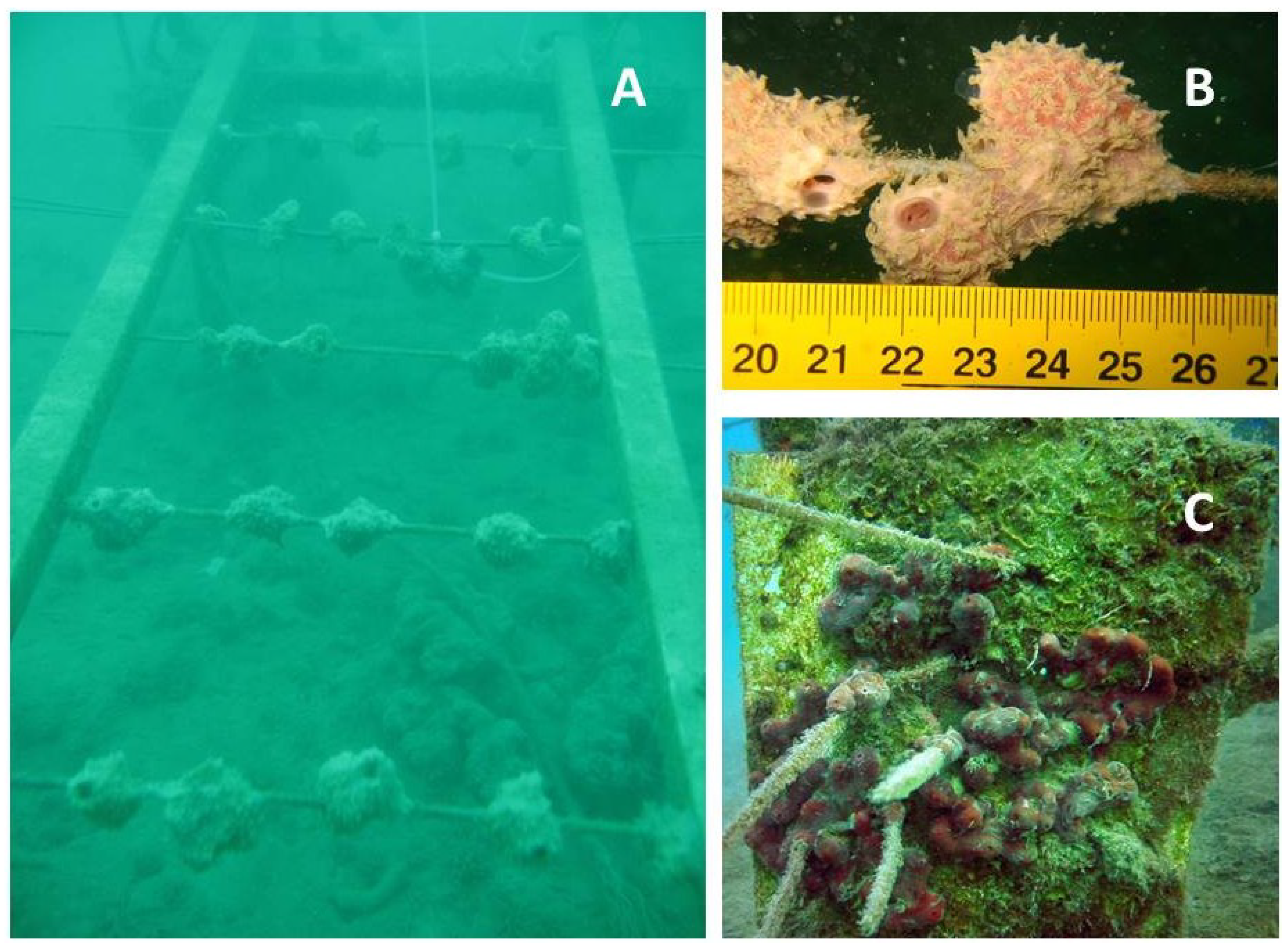
4.1. Sponges

4.2. Cnidarians
4.3. Mollusks
4.4. Other Invertebrates
5. Discovery of Bioactive Compounds from Marine Invertebrate-Associated Microorganisms and Their Production
6. Future Prospects
Acknowledgments
Conflicts of Interest
References
- Halpern, B.S.; Longo, C.; Hardy, D.; McLeod, K.L.; Samhouri, J.F.; Katona, S.K.; Kleisner, K.; Lester, S.E.; O’Leary, J.; Ranelletti, M.; et al. An index to assess the health and benefits of the global ocean. Nature 2012, 488, 615–620. [Google Scholar] [CrossRef]
- Leal, M.C.; Puga, J.; Serôdio, J.; Gomes, N.C.M.; Calado, R. Trends in the discovery of new marine natural products from invertebrates over the last two decades—Where and what are we bioprospecting? PLoS One 2012, 7, e30580. [Google Scholar]
- Leal, M.C.; Munro, M.H.G.; Blunt, J.W.; Puga, J.; Jesus, B.; Calado, R.; Rosa, R.; Madeira, C. Biogeography and biodiscovery hotspots of macroalgal marine natural products. Nat. Prod. Rep. 2013, 30, 1380–1390. [Google Scholar] [CrossRef]
- Blunt, J.W.; Munro, M.H.G. MarinLit database, 2013, Department of Chemistry, University of Canterbury. Available online: http://www.chem.canterbury.ac.nz/marinlit/marinlit.shtml (accessed on 1 August 2013).
- Appeltans, W.; Bouchet, P.; Boxshall, G.A.; de Broyer, C.; de Voogd, N.J.; Gordon, D.P.; Hoeksema, B.W.; Horton, T.; Kennedy, M.; Mees, J.; et al. World Register of Marine Species. Available online: http://www.marinespecies.org (accessed on 28 February 2014).
- Molinski, T.F.; Dalisay, D.S.; Lievens, S.L.; Saludes, J.P. Drug development from marine natural products. Nat. Rev. Drug Discov. 2009, 8, 69–85. [Google Scholar] [CrossRef]
- Rocha, J.; Peixe, L.; Gomes, N.C.M.; Calado, R. Cnidarians as a source of new marine bioactive compounds—An overview of the last decade and future steps for bioprospecting. Mar. Drugs 2011, 9, 1860–1886. [Google Scholar] [CrossRef]
- Leal, M.C.; Madeira, C.; Brandão, C.A.; Puga, J.; Calado, R. Bioprospecting of marine invertebrates for new natural products—A zoogeographical and chemical perspective. Molecules 2012, 17, 9842–9854. [Google Scholar] [CrossRef]
- Hill, R.T.; Fenical, W. Pharmaceuticals from marine natural products: Surge or ebb? Curr. Opin. Biotechnol. 2010, 21, 777–779. [Google Scholar] [CrossRef]
- Haygood, M.G.; Schmidt, E.W.; Davidson, S.K.; Faulkner, D.J. Microbial symbionts of marine invertebrates: Opportunities for microbial biotechnology. J. Mol. Microbiol. Biotechnol. 1999, 1, 33–43. [Google Scholar]
- Piel, J. Metabolites from symbiotic bacteria. Nat. Prod. Rep. 2004, 21, 519–538. [Google Scholar] [CrossRef]
- Harel, M.; Ben-Dov, E.; Rasoulouniriana, D.; Siboni, N.; Kramarsky-Winter, E.; Loya, Y.; Barak, Z.; Wiesman, Z.; Kushmaro, A. A new Thraustochytrid, strain Fng1, isolated from the surface of the hermatypic coral Fungia granulosa. FEMS Microbiol. Ecol. 2008, 64, 378–387. [Google Scholar] [CrossRef]
- Shnit-Orland, M.; Kushmaro, A. Coral mucus-associated bacteria: A possible first line of defense. FEMS Microbiol. Ecol. 2009, 67, 371–380. [Google Scholar]
- Olson, J.B.; Kellogg, C.A. Microbial ecology of corals, sponges, and algae in mesophotic coral environments. FEMS Microbiol. Ecol. 2010, 1–14. [Google Scholar]
- Sweet, M.J.; Croquer, A.; Bythell, J.C. Bacterial assemblages differ between compartments within the coral holobiont. Coral Reefs 2010, 30, 39–52. [Google Scholar]
- Hentschel, U.; Usher, K.M.; Taylor, M.W. Marine sponges as microbial fermenters. FEMS Microbiol. Ecol. 2006, 55, 167–177. [Google Scholar] [CrossRef]
- Rosenberg, E.; Koren, O.; Reshef, L.; Efrony, R.; Zilber-Rosenberg, I. The role of microorganisms in coral health, disease and evolution. Nat. Rev. Microbiol. 2007, 5, 355–362. [Google Scholar] [CrossRef]
- Rohwer, F.; Seguritan, V.; Azam, F.; Knowlton, N. Diversity and distribution of coral-associated bacteria. Mar. Ecol. Prog. Ser. 2002, 243, 1–10. [Google Scholar] [CrossRef]
- Knowlton, N.; Rohwer, F. Multispecies microbial mutualisms on coral reefs: The host as a habitat. Am. Nat. 2003, 162, S51–S62. [Google Scholar] [CrossRef]
- Agostini, S.; Suzuki, Y.; Higuchi, T.; Casareto, B.E.; Yoshinaga, K.; Nakano, Y.; Fujimura, H. Biological and chemical characteristics of the coral gastric cavity. Coral Reefs 2011, 31, 147–156. [Google Scholar]
- Leal, M.C.; Nejstgaard, J.C.; Calado, R.; Thompson, M.E.; Frischer, M.E. Molecular assessment of heterotrophy and prey digeston in zooxanthellate cnidarians. Mol. Ecol. 2013. [Google Scholar] [CrossRef]
- Webster, N.S.; Taylor, M.W. Marine sponges and their microbial symbionts: Love and other relationships. Environ. Microbiol. 2012, 14, 335–346. [Google Scholar] [CrossRef]
- Ainsworth, T.; Thurber, R.; Gates, R. The future of coral reefs: A microbial perspective. Trends Ecol. Evol. 2010, 25, 233–240. [Google Scholar] [CrossRef]
- Ainsworth, T.; Hoegh-Guldberg, O. Bacterial communities closely associated with coral tissues vary under experimental and natural reef conditions and thermal stress. Aquat. Biol. 2009, 4, 289–296. [Google Scholar] [CrossRef]
- Paul, V.J.; Puglisi, M.P. Chemical mediation of interactions among marine organisms. Nat. Prod. Rep. 2004, 21, 189–209. [Google Scholar] [CrossRef]
- Paul, V.J.; Ritson-Williams, R.; Sharp, K. Marine chemical ecology in benthic environments. Nat. Prod. Rep. 2011, 28, 345–388. [Google Scholar] [CrossRef]
- Hay, M.E. Marine Chemical Ecology: Chemical Signals and Cues Structure Marine Populations, Communities, and Ecosystems. Annu. Rev. Mar. Sci. 2009, 1, 193–212. [Google Scholar] [CrossRef]
- Kobayashi, J.; Ishibashi, M. Bioactive metabolites of symbiotic marine microorganisms. Chem. Rev. 1993, 93, 1753–1769. [Google Scholar] [CrossRef]
- Hentschel, U.; Hopke, J.; Horn, M.; Friedrich, A.B.; Wagner, M.; Hacker, J.; Moore, B.S. Molecular evidence for a uniform microbial community in sponges from different oceans. Appl. Environ. Microbiol. 2002, 68, 4431–4440. [Google Scholar] [CrossRef]
- Davies, P. Effect of daylight variations on the energy budgets of shallow-water corals. Mar. Biol. 1991, 108, 137–144. [Google Scholar] [CrossRef]
- Piel, J. Bacterial symbionts: Prospects for the sustainable production of invertebrate-derived pharmaceuticals. Curr. Med. Chem. 2006, 13, 39–50. [Google Scholar] [CrossRef]
- Li, J.W.-H.; Vederas, J.C. Drug Discovery and Natural Products: End of an Era or an Endless Frontier? Science 2009, 325, 161–165. [Google Scholar] [CrossRef]
- Radjasa, O.K.; Vaske, Y.M.; Navarro, G.; Vervoort, H.C.; Tenney, K.; Linington, R.G.; Crews, P. Highlights of marine invertebrate-derived biosynthetic products: Their biomedical potential and possible production by microbial associants. Bioorg. Med. Chem. 2011, 19, 6658–6674. [Google Scholar] [CrossRef]
- Munro, M.; Blunt, J.W.; Dumdei, E.; Hickford, S.; Lill, R.; Li, S.; Battershill, C.; Duckworth, A. The discovery and development of marine compounds with pharmaceutical potential. J. Biotechnol. 1999, 70, 15–25. [Google Scholar] [CrossRef]
- Hay, M. Marine chemical ecology: What’s known and what’s next? J. Exp. Mar. Biol. Ecol. 1996, 200, 103–134. [Google Scholar] [CrossRef]
- Aratake, S.; Tomura, T.; Saitoh, S.; Yokokura, R.; Kawanishi, Y.; Shinjo, R.; Reimer, J.D.; Tanaka, J.; Maekawa, H. Soft Coral Sarcophyton (Cnidaria: Anthozoa: Octocorallia) Species Diversity and Chemotypes. PLoS One 2012, 7, e30410. [Google Scholar] [CrossRef]
- Roberts, C.M.; McClean, C.J.; Veron, J.E.; Hawkins, J.P.; Allen, G.R.; McAllister, D.E.; Mittermeier, C.G.; Schueler, F.W.; Spalding, M.; Wells, F.; et al. Marine biodiversity hotspots and conservation priorities for tropical reefs. Science 2002, 295, 1280–1284. [Google Scholar] [CrossRef]
- Cheung, W.W.L.; Lam, V.W.Y.; Sarmiento, J.L.; Kearney, K.; Watson, R.; Pauly, D. Projecting global marine biodiversity impacts under climate change scenarios. Fish Fish. 2009, 10, 235–251. [Google Scholar] [CrossRef]
- Gerwick, W.H.; Moore, B.S. Lessons from the past and charting the future of marine natural products drug discovery and chemical biology. Chem. Biol. 2012, 19, 85–98. [Google Scholar] [CrossRef]
- Glaser, K.B.; Mayer, A. A renaissance in marine pharmacology: From preclinical curiosity to clinical reality. Biochem. Pharmacol. 2009, 78, 440–448. [Google Scholar] [CrossRef]
- Martins, A.; Vieira, H.; Gaspar, H.; Santos, S. Marketed marine natural products in the pharmaceutical and cosmoceutical industries: Tips for success. Mar. Drugs 2014, 12, 1066–1101. [Google Scholar]
- Piel, J. Approaches to capturing and designing biologically-active small molecules produced by uncultured microbes. Annu. Rev. Microbiol. 2011, 65, 431–453. [Google Scholar] [CrossRef]
- Mayer, A.M.; Glaser, K.B.; Cuevas, C.; Jacobs, R.S.; Kem, W.; Little, R.D.; McIntosh, J.M.; Newman, D.J.; Potts, B.C.; Shuster, D.E. The odyssey of marine pharmaceuticals: A current pipeline perspective. Trends Pharmacol. Sci. 2010, 31, 255–265. [Google Scholar] [CrossRef]
- Suyama, T.L.; Gerwick, W.H.; McPhail, K.L. Survey of marine natural product structure revisions: A synergy of spectroscopy and chemical synthesis. Bioorg. Med. Chem. 2011, 19, 6675–6701. [Google Scholar] [CrossRef]
- Qian, P.-Y.; Xu, Y.; Fusetani, N. Natural products as antifouling compounds: Recent progress and future perspectives. Biofouling 2010, 26, 223–234. [Google Scholar] [CrossRef]
- Wilkinson, B.; Micklefield, J. Mining and engineering natural-product biosynthetic pathways. Nat. Chem. Biol. 2007, 3, 379–386. [Google Scholar] [CrossRef]
- Leal, M.C.; Calado, R.; Sheridan, C.; Alimonti, A.; Osinga, R. Coral aquaculture to support drug discovery. Trends Biotechnol. 2013, 31, 555–561. [Google Scholar] [CrossRef]
- Ben-Dov, E.; Kramarsky-Winter, E.; Kushmaro, A. An in situ method for cultivating microorganisms using a double encapsulation technique. FEMS Microbiol. Ecol. 2009, 68, 363–371. [Google Scholar] [CrossRef]
- Page, M.J.; Handley, S.J.; Northcote, P.T.; Cairney, D.; Willan, R.C. Successes and pitfalls of the aquaculture of the sponge Mycale hentscheli. Aquaculture 2011, 312, 52–61. [Google Scholar] [CrossRef]
- Page, M.J.; Northcote, P.T.; Webb, V.L.; Mackey, S.; Handley, S.J. Aquaculture trials for the production of biologically active metabolites in the New Zealand sponge Mycale hentscheli (Demospongiae: Poecilosclerida). Aquaculture 2005, 250, 256–269. [Google Scholar] [CrossRef]
- Cognetti, G.; Maltagliati, F.; Saroglia, M. The risk of “genetic pollution” in Mediterranean fish populations related to aquaculture activities. Mar. Pollut. Bull. 2006, 52, 1321–1323. [Google Scholar] [CrossRef]
- Garren, M.; Smriga, S.; Azam, F. Gradients of coastal fish farm effluents and their effect on coral reef microbes. Environ. Microbiol. 2008, 10, 2299–2312. [Google Scholar] [CrossRef]
- Mendola, D. Aquaculture of three phyla of marine invertebrates to yield bioactive metabolites: Process developments and economics. Biomol. Eng. 2003, 20, 441–458. [Google Scholar] [CrossRef]
- Thomas, T.R.A.; Kavlekar, D.P.; LokaBharathi, P.A. Marine Drugs from Sponge-Microbe Association—A Review. Mar. Drugs 2010, 8, 1417–1468. [Google Scholar] [CrossRef]
- Kooperman, N.; Ben-Dov, E.; Kramarsky-Winter, E.; Barak, Z.; Kushmaro, A. Coral mucus-associated bacterial communities from natural and aquarium environments. FEMS Microbiol. Lett. 2007, 276, 106–113. [Google Scholar] [CrossRef]
- Isaacs, L.; Kan, J.; Nguyen, L.; Videau, P.; Anderson, M.; Wright, T.; Hill, R. Comparison of the Bacterial Communities of Wild and Captive Sponge Clathria prolifera from the Chesapeake Bay. Mar. Biotechnol. 2009, 11, 758–770. [Google Scholar] [CrossRef]
- Sweet, M.; Jones, R.; Bythell, J. Coral diseases in aquaria and in nature. J. Mar. Biol. Assoc. UK 2011, 1–11. [Google Scholar]
- Khalesi, M.K.; Beeftink, H.H.; Wijffels, R.H. Light-Dependency of Growth and Secondary Metabolite Production in the Captive Zooxanthellate Soft Coral Sinularia flexibilis. Mar. Biotechnol. 2009, 11, 488–494. [Google Scholar] [CrossRef]
- Olivotto, I.; Planas, M.; Simões, N.; Holt, G.; Avella, M.; Calado, R. Advances in breeding and rearing marine ornamentals. J. World Aquac. Soc. 2011, 42, 135–166. [Google Scholar] [CrossRef]
- Dionísio, G.; Rosa, R.; Leal, M.C.; Cruz, S.; Brandão, C.; Calado, G.; Serôdio, J.; Calado, R. Beaties and beasts: A protrait of sea slugs aquaculture. Aquaculture 2013, 408–409, 1–14. [Google Scholar]
- Sipkema, D.; Osinga, R.; Schatton, W.; Mendola, D.; Tramper, J.; Wijffels, R.H. Large-scale production of pharmaceuticals by marine sponges: Sea, cell, or synthesis? Biotechnol. Bioeng. 2005, 90, 201–222. [Google Scholar] [CrossRef]
- Sheridan, C.; Kramarsky-Winter, E.; Sweet, M.; Kushmaro, A.; Leal, M.C. Diseases in coral aquaculture: Causes, implications and preventions. Aquaculture 2013, 396–399, 124–135. [Google Scholar]
- Hooper, J.N.A.; van Soest, R.W.M. Systema Porifera: A Guide do the Classification of Sponges; Kluwer Academic/Plenum Publishers: New York, NY, USA, 2002. [Google Scholar]
- Proksch, P.; Putz, A.; Ortlepp, S.; Kjer, J.; Bayer, M. Bioactive natural products from marine sponges and fungal endophytes. Phytochem. Rev. 2010, 9, 475–489. [Google Scholar] [CrossRef]
- Proksch, P.; Ebel, R.; Edrada-Ebel, R.A.; Riebe, F.; Liu, H.; Diesel, A.; Bayer, M.; Li, X.; Li, W.H.; Grebenyuk, V.; et al. Sponge-associated fungi and their bioactive compounds: The Suberites case. Bot. Mar. 2008, 51, 209–218. [Google Scholar]
- Margassery, L.M. Biodiscovery of Natural Products from Microbes Associated with Irish Coastal Sponges. Ph.D. Thesis, University College Cork, Cork, Ireland, April 2013. [Google Scholar]
- Schippers, K.J.; Sipkema, D.; Osinga, R.; Smidt, H.; Pomponi, S.A.; Martens, D.E.; Wijffels, R.H. Cultivation of sponges, sponge cells and symbionts: Achievements and future prospects. Adv. Mar. Biol. 2012, 62, 273–337. [Google Scholar] [CrossRef]
- Duckworth, A. Farming Sponges to Supply Bioactive Metabolites and Bath Sponges: A Review. Mar. Biotechnol. 2009, 11, 669–679. [Google Scholar] [CrossRef]
- Grasela, J.J.; Pomponi, S.A.; Rinkevich, B.; Grima, J. Efforts to develop a cultured sponge cell line: Revisiting an intractable problem. In Vitro Cell. Dev. Biol.-Anim. 2012, 48, 12–20. [Google Scholar] [CrossRef]
- Dumdei, E.J.; Blunt, J.W.; Munro, M.H.G.; Battershill, C.N.; Page, M.J. The whys and whats of sponge chemistry: Why chemists extract sponges and what problems does this cause? In Sponge Sciences, Multidisciplinary Perspectives; Watanabe, Y., Fusetani, N., Eds.; Springer: Tokyo, Japan, 1998. [Google Scholar]
- Ruiz, C.; Valderrama, K.; Zea, S.; Castellanos, L. Mariculture and Natural Production of the Antitumoural (+)-Discodermolide by the Caribbean Marine Sponge Discodermia dissoluta. Mar. Biotechnol. 2013, 15, 571–583. [Google Scholar] [CrossRef]
- De Goeij, J.; Moodley, L.; Houtekamer, M.; Carballeira, N.; Duyl, F. Tracing 13C-enriched dissolved and particulate organic carbon in the bacteria-containing coral reef sponge Halisarca caerulea: Evidence for DOM feeding. Limnol. Oceanogr. 2008, 53, 1376–1386. [Google Scholar] [CrossRef]
- De Goeij, J.M.; van den Berg, H.; van Oostveen, M.M.; Epping, E.H.G.; van Duyl, F.C. Major bulk dissolved organic carbon (DOC) removal by encrusting coral reef cavity sponges. Mar. Ecol. Prog. Ser. 2008, 357, 139–151. [Google Scholar] [CrossRef]
- De Goeij, J.M.; van Oevelen, D.; Vermeij, M.J.A.; Osinga, R.; Middelburg, J.J.; Admiraal, W. Surviving in a marine desert: The sponge loop retains resources within coral reefs. Science 2013, 342, 108–110. [Google Scholar] [CrossRef]
- Yahel, G.; Sharp, J.H.; Marie, D.; Haese, C.; Genin, A. In situ feeding and element removal in the symbiont-bearing sponge Theonella swinhoei: Bulk DOC is the major source for carbon. Limnol. Oceanogr. 2003, 48, 141–149. [Google Scholar] [CrossRef]
- De Goeij, J.M.; de Kluijver, A.; van Duyl, F.C.; Vacelet, J.; Wijffels, R.H.; de Goeij, A.F.P.M.; Cleutjens, J.P.M.; Schutte, B. Cell kinetics of the marine sponge Halisarca caerulea reveal rapid cell turnover and shedding. J. Exp. Biol. 2009, 212, 3892–3900. [Google Scholar] [CrossRef]
- Muller, W.E.G.; Wimmer, W.; Schatton, W.; Bohm, M.; Batel, R.; Filic, Z. Initiation of an aquaculture of sponges for the sustainable production of bioactive metabolites in open systems: Example, Geodia cydonium. Mar. Biotechnol. 1999, 1, 569–579. [Google Scholar] [CrossRef]
- Osinga, R.; Sidri, M.; Cerig, E.; Gokalp, S.Z.; Gokalp, M. Sponge aquaculture in the East Mediterranean Sea: New approaches to earlier ideas. Open Mar. Biol. J. 2010, 4, 74–81. [Google Scholar] [CrossRef]
- Bergman, O.; Mayzel, B.; Anderson, M.A.; Shpigel, M.; Hill, R.T.; Ilan, M. Examination of Marine-Based Cultivation of Three Demosponges for Acquiring Bioactive Marine Natural Products. Mar. Drugs 2011, 9, 2201–2219. [Google Scholar] [CrossRef]
- De Caralt, S.; Sánchez-Fontenla, J.; Uriz, M.J.; Wijffels, R.H. In situ aquaculture methods of Dysidea avara (Demospongiae, Porifera) in the northwestern Mediterranean. Mar. Drugs 2010, 8, 1731–1742. [Google Scholar] [CrossRef]
- Hadas, E.; Shpigel, M.; Ilan, M. Sea ranching of the marine sponge Negombata magnifica (Demospongiae, Latruncullidae) as a first step for Latrunculin B mass production. Aquaculture 2005, 244, 159–169. [Google Scholar] [CrossRef]
- Duckworth, A.R.; Battershill, C.N.; Schiel, D.R. Effects of depth and water flow on growth, survival and bioactivity of two temperate sponges cultured in different seasons. Aquaculture 2004, 242, 237–250. [Google Scholar] [CrossRef]
- Domenicotti, C. Growth dynamics and bioactivity variation of the Mediterranean demosponges Agelas oroides (Agelasida, Agelasidae) and Petrosia ficiformis (Haplosclerida, Petrosiidae). Mar. Ecol. 2009, 30, 327–336. [Google Scholar] [CrossRef]
- Blunt, J.W.; Copp, B.R.; Munro, M.H.G.; Northcote, P.T.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2010, 27, 165–237. [Google Scholar] [CrossRef]
- Al-Zereini, W.; Yao, C.B.F.F.; Laatsch, H.; Anke, H.J. Aqabamycins A–G: Novel nitro maleimides from a marine Vibrio species. I. Taxonomy, ferentation, isolation and biological activities. J. Antibiot. 2010, 63, 297–301. [Google Scholar] [CrossRef]
- Blunt, J.W.; Copp, B.R.; Keyzers, R.A.; Munro, M.H.G.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2013, 30, 237–323. [Google Scholar] [CrossRef]
- Pomeroy, R.; Parks, J.; Balboa, C. Farming the reef: Is aquaculture a solution for reducing fishing pressure on coral reefs? Mar. Policy 2006, 30, 111–130. [Google Scholar] [CrossRef]
- Shafir, S.; van Rijn, J.; Rinkevich, B. Coral nubbins as source material for coral biological research: A prospectus. Aquaculture 2006, 259, 444–448. [Google Scholar] [CrossRef]
- Leal, M.C.; Nunes, C.; Engrola, S.; Dinis, M.; Calado, R. Optimization of monoclonal production of the glass anemone Aiptasia pallida (Agassiz in Verrill, 1864). Aquaculture 2012, 354–355, 91–96. [Google Scholar]
- Jaap, W.C. Coral reef restoration. Ecol. Eng. 2000, 15, 345–364. [Google Scholar] [CrossRef]
- Delbeek, J. Coral farming: Past, present and future trends. Aquar. Sci. Conserv. 2001, 3, 171–181. [Google Scholar]
- Ellis, S. Farming Soft Corals for the Marine Aquarium Trade; Center for Tropical and Subtropical Aquaculture: Pohnpei, Federal States of Micronesia, 1999. [Google Scholar]
- Shafir, S.; van Rijn, J.; Rinkevich, B. Steps in the construction of underwater coral nursery, an essential component in reef restoration acts. Mar. Biol. 2006, 149, 679–687. [Google Scholar] [CrossRef]
- Rocha, R.J.M.; Serôdio, J.; Leal, M.C.; Cartaxana, P.; Calado, R. Effect of light intensity on post-fragmentation photobiological performance of the soft coral Sinularia flexibilis. Aquaculture 2013, 388–391, 24–29. [Google Scholar]
- Leal, M.C.; Nunes, C.; Kempf, S.C.; Reis, A.; Silva, T.L.; Serôdio, J.; Cleary, D.F.R.; Calado, R. Effect of light, temperature and diet on the fatty acid profile of the tropical sea anemone Aiptasia pallida. Aquac. Nutr. 2013, 19, 818–826. [Google Scholar] [CrossRef]
- Rocha, R.J.M.; Pimentel, T.; Serôdio, J.; Rosa, R.; Calado, R. Comparative performance of light emitting plastma (LEP) and light emitting diode (LED) in ex situ aquaculture of scleractinian corals. Aquaculture 2013, 402–403, 38–45. [Google Scholar]
- Rocha, R.J.M.; Calado, R.; Cartaxana, P.; Furtado, J.; Serôdio, J. Photobiology and growth of leather coral Sarcophyton cf. glaucum fragments stocked under low light in a recirculated system. Aquaculture 2013, 414–415, 235–242. [Google Scholar]
- Wijgerde, T.; Henkemans, P.; Osinga, R. Effects of irradiance and light spectrum on growth of the scleractinian coral Galaxea fascicularis—Applicability of LEP and LED lighting to coral aquaculture. Aquaculture 2012, 344–349, 188–193. [Google Scholar] [CrossRef]
- Osinga, R.; Schutter, M.; Griffioen, B.; Wijffels, R.H.; Verreth, J.A.J.; Shafir, S.; Henard, S.; Taruffi, M.; Gili, C.; Lavorano, S. The Biology and Economics of Coral Growth. Mar. Biotechnol. 2011, 13, 658–671. [Google Scholar] [CrossRef]
- Vizel, M.; Loya, Y.; Downs, C.A.; Kramarsky-Winter, E. A novel method for coral explant culture and micropropagation. Mar. Biotechnol. 2011, 13, 423–432. [Google Scholar] [CrossRef]
- Petersen, D.; Laterveer, M.; van Bergen, D.; Hatta, M.; Hebbinghaus, R.; Janse, M.; Jones, R.; Richter, U.; Ziegler, T.; Visser, G.; et al. The application of sexual coral recruits for the sustainable management ofex situ populations in public aquariums to promote coral reef conservation—SECORE Project. Aquat. Conserv.: Mar. Freshw. Ecosyst. 2006, 16, 167–179. [Google Scholar] [CrossRef]
- Petersen, D.; Falcato, J.; Gilles, P.; Jones, R. Sexual coral reproduction in live coral exhibits—Current status and future perspectives. Int. Zoo Yearb. 2007, 41, 122–137. [Google Scholar] [CrossRef]
- Baums, I.B. A restoration genetics guide for coral reef conservation. Mol. Ecol. 2008, 17, 2796–2811. [Google Scholar] [CrossRef]
- Fleury, B.G.; Lages, B.G.; Barbosa, J.P.; Kaiser, C.R.; Pinto, Â.C. New hemiketal steroid from the introduced soft coral Chromonephthea braziliensis is a chemical defense against predatory fishes. J. Chem. Ecol. 2008, 34, 987–993. [Google Scholar] [CrossRef]
- Koh, E.; Sweatman, H. Chemical warfare among scleractinians: Bioactive natural products from Tubastraea faulkneri Wells kill larvae of potential competitors. J. Exp. Mar. Biol. Ecol. 2000, 251, 141–160. [Google Scholar] [CrossRef]
- Fleury, B.; Cool, J.; Sammarco, P. Complementary (secondary) metabolites in a soft coral: Sex-specific variability, inter-clonal variability, and competition. Mar. Ecol. 2006, 27, 204–218. [Google Scholar] [CrossRef]
- Houlbrèque, F.; Ferrier-Pagès, C. Heterotrophy in Tropical Scleractinian Corals. Biol. Rev. 2009, 84, 1–17. [Google Scholar] [CrossRef]
- Brown, B.; Bythell, J. Perspectives on mucus secretion in reef corals. Mar. Ecol. Prog. Ser. 2005, 296, 291–309. [Google Scholar] [CrossRef]
- Avila, C. Natural products of opisthobranch molluscs: A biological review. Oceanogr. Mar. Biol. Annu. Rev. 1995, 33, 487–559. [Google Scholar]
- Cimino, G.; Ghiselin, M.T. Chemical defense and the evolution of opisthobranch gastropods. Proc. Calif. Acad. Sci. 2009, 60, 175–422. [Google Scholar]
- Olivera, B.M.; Teichert, R.W. Diversity of the Neurotoxic Conus Peptides. Mol. Interv. 2007, 7, 251–260. [Google Scholar] [CrossRef]
- Devine, S.P. Characterization of Bacteria Associated with the Kleptoplastic Sea Slug Elysia chlorotica and Its Algal Prey Vaucheria litorea Using Metagenomics. Electronic Theses and Dissertations, University of Maine, 2012. Available online: http://digitalcommons.library.umaine.edu/etd/1757 (accessed on 1 August 2013).
- Davis, J.; Fricke, W.F.; Hamann, M.T.; Esquenazi, E.; Dorrestein, P.C.; Hill, R.T. Characterization of the Bacterial Community of the Chemically Defended Hawaiian Sacoglossan Elysia rufescens. Appl. Environ. Microbiol. 2013, 79, 7073–7081. [Google Scholar] [CrossRef]
- Hamann, M.T.; Otto, C.S.; Scheuer, P.J.; Dunbar, D.C. Kahalalides: Bioactive Peptides from a Marine Mollusk Elysia rufescens and Its Algal Diet Bryopsis sp.1. J. Org. Chem. 1996, 61, 6594–6600. [Google Scholar] [CrossRef]
- Pardo, B.; Paz-Ares, L.; Tabernero, J.; Ciruelos, E.; García, M.; Salazar, R.; López, A.; Blanco, M.; Nieto, A.; Jimeno, J. Phase I clinical and pharmacokinetic study of kahalalide F administered weekly as a 1-hour infusion to patients with advanced solid tumors. Clin. Cancer Res. 2008, 14, 1116–1123. [Google Scholar] [CrossRef]
- Luesch, H.; Moore, R.E.; Paul, V.J.; Mooberry, S.L.; Corbett, T.H. Isolation of dolastatin 10 from the marine cyanobacterium Symploca species VP642 and total stereochemistry and biological evaluation of its analogue symplostatin 1. J. Nat. Prod. 2001, 64, 907–910. [Google Scholar] [CrossRef]
- Harrigan, G.G.; Luesch, H.; Yoshida, W.Y.; Moore, R.E.; Nagle, D.G.; Paul, V.J.; Mooberry, S.L.; Corbett, T.H.; Valeriote, F.A. Symplostatin 1: A dolastatin 10 analogue from marine cyanobacterium Symploca hydnoides. J. Nat. Prod. 1998, 61, 1075–1077. [Google Scholar] [CrossRef]
- Kamiya, H.; Sakai, R.; Jimbo, M. Bioactive molecules from sea hares. Prog. Mol. Subcell. Biol. 2006, 43, 215–239. [Google Scholar] [CrossRef]
- Myers, R.A.; Cruz, L.J.; Rivier, J.E.; Olivera, B.M. Conus peptides as chemical probes for receptors and ion channels. Chem. Rev. 1993, 93, 1923–1936. [Google Scholar] [CrossRef]
- Olivera, B.M. E.E. Just Lecture, 1996: Conus Venom Peptides, Receptor and Ion Channel Targets, and Drug Design: 50 Million Years of Neuropharmacology. Mol. Biol. Cell 1997, 8, 2101–2109. [Google Scholar] [CrossRef]
- Buczek, O.; Bulaj, G.; Olivera, B.M. Conotoxins and the posttranslational modification of secreted gene products. Cell. Mol. Life Sci. 2005, 62, 3067–3079. [Google Scholar] [CrossRef]
- Prommer, E. Zinconotide: A new option for refractory pain. Drugs Today 2006, 24, 369–378. [Google Scholar] [CrossRef]
- Lin, Z.; Marett, L.; Hughen, R.W.; Flores, M.; Forteza, I.; Ammon, M.A.; Concepcion, G.P.; Espino, S.; Olivera, B.M.; Rosenberg, G.; et al. Neuroactive diol and acyloin metabolites from cone snail-associated bacteria. Bioorg. Med. Chem. Lett. 2013, 23, 4867–4869. [Google Scholar] [CrossRef]
- Peraud, O.; Biggs, J.S.; Hughen, R.W.; Light, A.R.; Concepcion, G.P.; Olivera, B.M.; Schmidt, E.W. Microhabitats within Venomous Cone Snails Contain Diverse Actinobacteria. Appl. Environ. Microbiol. 2009, 75, 6820–6826. [Google Scholar] [CrossRef]
- O’Brien, F.X.; McKindsey, C.H.; Landry, T.; Costa-Pierce, B.A. Methods of sustainable shelfish aquaculture. In Sustainable Food Production; Christou, P., Savin, R., Costa-Pierce, B.A., Misztal, I., Whitelaw, B.A., Eds.; Springer Science+Business Media: New York, NY, USA, 2013; pp. 1436–1548. [Google Scholar]
- Shawl, A.L.; Davis, M. Captive breeding behaviour of four strombidae conch. J. Shellfish Res. 2004, 23, 157–164. [Google Scholar]
- Dolorosa, R.G.; Grant, A.; Gill, J.A.; Avillanosa, A.L.; Gonzales, B.J. Indoor and Deep Sub-Tidal Intermediate Culture of Trochus niloticus for Restocking. Rev. Fish. Sci. 2013, 21, 414–423. [Google Scholar] [CrossRef]
- Capo, T.; Bardales, A.; Gillette, P.; Lara, M.; Schmale, M.; Serafy, J. Larval growth, development, and survival of laboratory-reared Aplysia californica: Effects of diet and veliger density. Comp. Biochem. Phys. C 2008, 149, 215–223. [Google Scholar] [CrossRef]
- Leal, M.C.; Nunes, C.; Alexandre, D.; Silva, T.L.; Reis, A.; Dinis, M.T.; Calado, R. Parental diets determine the embryonic fatty acid profile of the tropical nudibranch Aeolidiella stephanieae: The effect of eating bleached anemones. Mar. Biol. 2012, 159, 1745–1751. [Google Scholar] [CrossRef]
- Cruz, S.; Calado, R.; Serôdio, J.; Cartaxana, P. Crawling leaves: Photosynthesis in sacoglossan sea slugs. J. Exp. Bot. 2013, 64, 3999–4009. [Google Scholar] [CrossRef]
- Carroll, D.; Kempf, S. Laboratory culture of the aeolid nudibranch Berghia verrucicornis (Mollusca, Opisthobranchia): Some aspects of its development and life history. Biol. Bull. 1990, 179, 243–253. [Google Scholar] [CrossRef]
- Mayer, A.M.S.; Glaser, K.B. Marine pharmacology and the marine pharmaceuticals pipeline. FASEB J. 2013, 27, 1167.7. [Google Scholar]
- Haefner, B. Drugs from the deep: Marine natural products as drug candidates. Drug Discov. Today 2003, 8, 536–544. [Google Scholar] [CrossRef]
- Fusetani, N. Drugs from the Sea; Krager: Basel, Switzerland, 2000; p. 164. [Google Scholar]
- Lilly, M.; Brown, C.; Pettit, G.; Kraft, A. Bryostatin 1: A potential anti-leukemic agent for chronic myelomonocytic leukemia. Leukemia 1991, 5, 283–287. [Google Scholar]
- Mendola, D. Aquaculture production of bryostatin 1 and ecteinascidin 743. In Drugs from the Sea; Fusetani, N., Ed.; Krager: Basel, Switzerland, 2000; pp. 120–133. [Google Scholar]
- Dahms, H.-W.; Gao, Q.-F.; Hwang, J.-S. Optimized maintenance and larval production of the bryozoan Bugula neritina (Bugulidae: Gymnolaemata) in the laboratory. Aquaculture 2007, 265, 169–175. [Google Scholar] [CrossRef]
- Kahle, J.; Liebezeit, G.; Gerdes, G. Growth aspects of Flustra foliacea (Bryozoa, Cheilostomata) in laboratory. In Migrations and Dispersal of Marine Organisms; Jones, M.B., Infólfsson, A., Ólafsson, E., Helgason, G.V., Gunnarsson, K., Svavarsson, J., Eds.; Springer Science+Business Media B.V.: Dordrecht, The Netherlands, 2003; Volume 174, pp. 237–244. [Google Scholar]
- Trindade-Silva, A.E.; Lim-fong, G.E.; Sharp, K.H.; Haygood, M.G. Bryostatins: Biological context and biotechnological prospects. Curr. Opin. Biotechnol. 2010, 21, 834–842. [Google Scholar] [CrossRef]
- Cuevas, C.; Francesch, A. Development of Yondelis (R) (trabectedin, ET-743). A semisynthetic process solves the supply problem. Nat. Prod. Rep. 2009, 26, 322–337. [Google Scholar] [CrossRef]
- Zengler, K.; Toledo, G.; Rappé, M.; Elkins, J.; Mathur, E.J.; Short, J.M.; Keller, M. Cultivating the uncultured. Proc. Natl. Acad. Sci. USA 2002, 99, 15681–15686. [Google Scholar] [CrossRef]
- Amann, R.I.; Ludwig, W.; Schleifer, K.H. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 1995, 59, 143–169. [Google Scholar]
- Joint, I.; Mühling, M.; Querellou, J. Culturing marine bacteria—An essential prerequisite for biodiscovery. Microb. Biotechnol. 2010, 3, 564–575. [Google Scholar] [CrossRef]
- Pham, V.H.T.; Kim, J. Cultivation of unculturable soil bacteria. Trends Biotechnol. 2012, 30, 475–484. [Google Scholar] [CrossRef]
- Alain, K.; Querellou, J. Cultivating the uncultured: Limits, advances and future challenges. Extremophiles 2009, 13, 583–594. [Google Scholar] [CrossRef]
- Janssen, P.H.; Yates, P.S.; Grinton, B.E.; Taylor, P.M.; Sait, M. Improved Culturability of Soil Bacteria and Isolation in Pure Culture of Novel Members of the Divisions Acidobacteria, Actinobacteria, Proteobacteria, and Verrucomicrobia. Appl. Environ. Microbiol. 2002, 68, 2391–2396. [Google Scholar] [CrossRef]
- Stewart, E.J. Growing Unculturable Bacteria. J. Bacteriol. 2012, 194, 4151–4160. [Google Scholar] [CrossRef]
- Straight, P.D.; Kolter, R. Interspecies Chemical Communication in Bacterial Development. Annu. Rev. Microbiol. 2009, 63, 99–118. [Google Scholar] [CrossRef]
- Ohno, M.; Shiratori, H.; Park, M.J.; Saitoh, Y.; Kumon, Y.; Yamashita, N.; Hirata, A.; Nishida, H.; Ueda, K.; Beppu, T. Symbiobacterium thermophilum gen. nov. sp. nov. a symbiotic thermophile that depends on co-culture with a Bacillus strain for growth. Int. J. Syst. Evol. Microbiol. 2000, 50, 1829–1832. [Google Scholar]
- Pagnier, I.; Raoult, D.; La Scola, B. Isolation and identification of amoeba-resisting bacteria from water in human environment by using an Acanthamoeba polyphaga co-culture procedure. Environ. Microbiol. 2008, 10, 1135–1144. [Google Scholar] [CrossRef]
- Nichols, D.; Cahoon, N.; Trakhtenberg, E.M.; Pham, L.; Mehta, A.; Belanger, A.; Kanigan, T.; Lewis, K.; Epstein, S.S. Use of Ichip for High-Throughput In Situ Cultivation of “Uncultivable” Microbial Species. Appl. Environ. Microbiol. 2010, 76, 2445–2450. [Google Scholar] [CrossRef]
- Aoi, Y.; Kinoshita, T.; Hata, T.; Ohta, H.; Obokata, H.; Tsuneda, S. Hollow-Fiber Membrane Chamber as a Device for In Situ Environmental Cultivation. Appl. Environ. Microbiol. 2009, 75, 3826–3833. [Google Scholar] [CrossRef]
- Kaeberlein, T.; Lewis, K.; Epstein, S.S. Isolating “Uncultivable” Microorganisms in Pure Culture in a Simulated Natural Environment. Science 2002, 296, 1127–1129. [Google Scholar] [CrossRef]
- Bollmann, A.; Lewis, K.; Epstein, S.S. Incubation of Environmental Samples in a Diffusion Chamber Increases the Diversity of Recovered Isolates. Appl. Environ. Microbiol. 2007, 73, 6386–6390. [Google Scholar] [CrossRef]
- Singh, B.K.; Macdonald, C.A. Drug discovery from uncultivable microorganisms. Drug Discov. Today 2010, 15, 792–799. [Google Scholar]
- Schmidt, E.W.; Nelson, J.T.; Rasko, D.A.; Sudek, S.; Eisen, J.A.; Haygood, M.G.; Ravel, J. Patellamide A and C biosynthesis by a microcin-like pathway in Prochloron didemni, the cyanobacterial symbiont of Lissoclinum patella. Proc. Natl. Acad. Sci. USA 2005, 102, 7315–7320. [Google Scholar]
- Long, P.F.; Dunlap, W.C.; Battershill, C.N.; Jaspars, M. Shotgun Cloning and Heterologous Expression of the Patellamide Gene Cluster as a Strategy to Achieving Sustained Metabolite Production. ChemBioChem 2005, 6, 1760–1765. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Leal, M.C.; Sheridan, C.; Osinga, R.; Dionísio, G.; Rocha, R.J.M.; Silva, B.; Rosa, R.; Calado, R. Marine Microorganism-Invertebrate Assemblages: Perspectives to Solve the “Supply Problem” in the Initial Steps of Drug Discovery. Mar. Drugs 2014, 12, 3929-3952. https://doi.org/10.3390/md12073929
Leal MC, Sheridan C, Osinga R, Dionísio G, Rocha RJM, Silva B, Rosa R, Calado R. Marine Microorganism-Invertebrate Assemblages: Perspectives to Solve the “Supply Problem” in the Initial Steps of Drug Discovery. Marine Drugs. 2014; 12(7):3929-3952. https://doi.org/10.3390/md12073929
Chicago/Turabian StyleLeal, Miguel Costa, Christopher Sheridan, Ronald Osinga, Gisela Dionísio, Rui Jorge Miranda Rocha, Bruna Silva, Rui Rosa, and Ricardo Calado. 2014. "Marine Microorganism-Invertebrate Assemblages: Perspectives to Solve the “Supply Problem” in the Initial Steps of Drug Discovery" Marine Drugs 12, no. 7: 3929-3952. https://doi.org/10.3390/md12073929
APA StyleLeal, M. C., Sheridan, C., Osinga, R., Dionísio, G., Rocha, R. J. M., Silva, B., Rosa, R., & Calado, R. (2014). Marine Microorganism-Invertebrate Assemblages: Perspectives to Solve the “Supply Problem” in the Initial Steps of Drug Discovery. Marine Drugs, 12(7), 3929-3952. https://doi.org/10.3390/md12073929






