Oxidized Cell-Free DNA Rapidly Skews the Transcriptional Profile of Brain Cells toward Boosting Neurogenesis and Neuroplasticity
Abstract
:1. Introduction
2. Materials and Methods
2.1. Primary Cell Culture
2.2. CfDNA Purification and Oxidation
2.3. Sampling and mRNA Purification
2.4. RT PCR
2.5. Multiplex Gene Expression Analysis
2.6. Statistics
3. Results
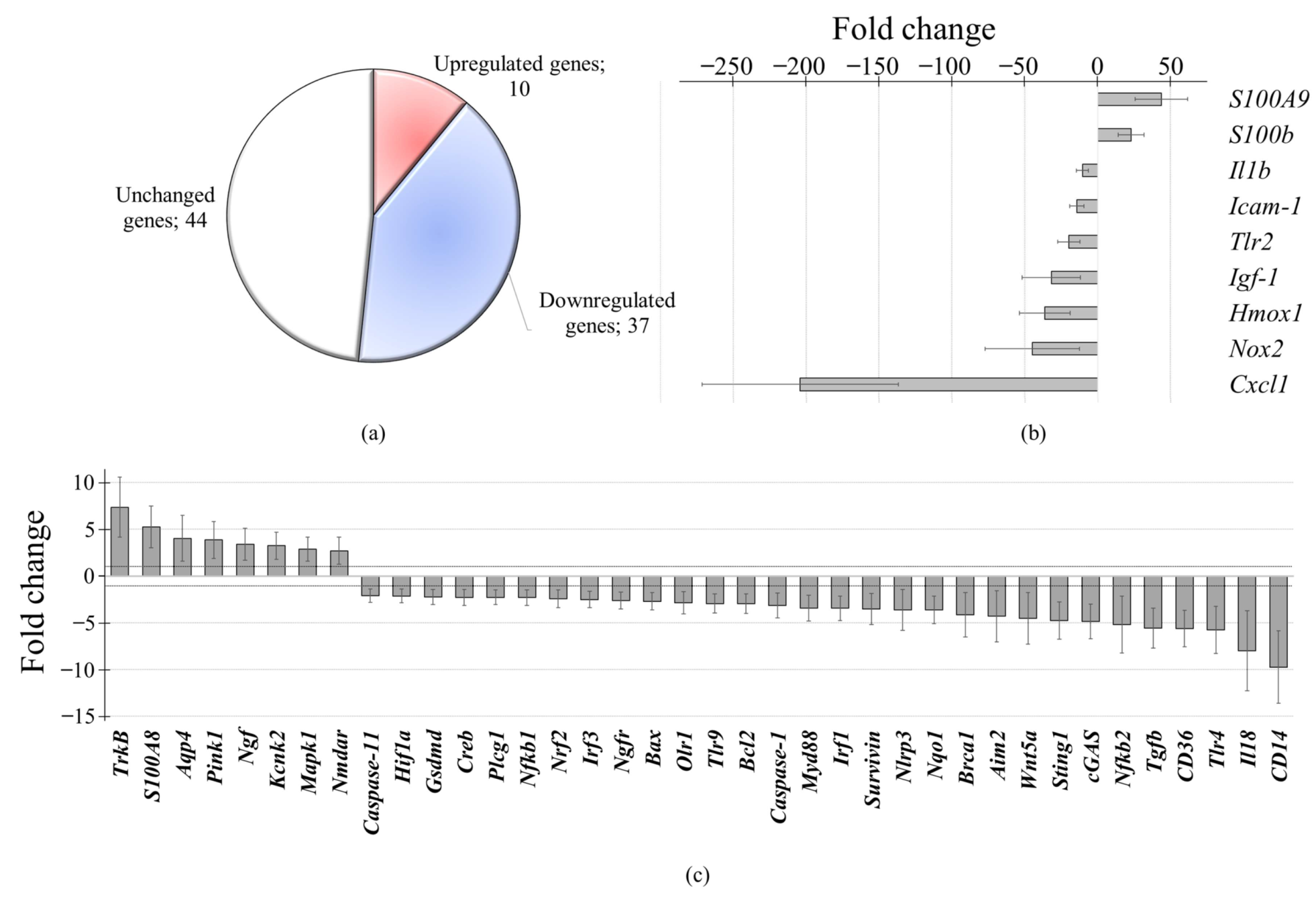
3.1. Early Effects of Oxidized and Non-Oxidized cfDNA on Gene Expression in Brain Cells
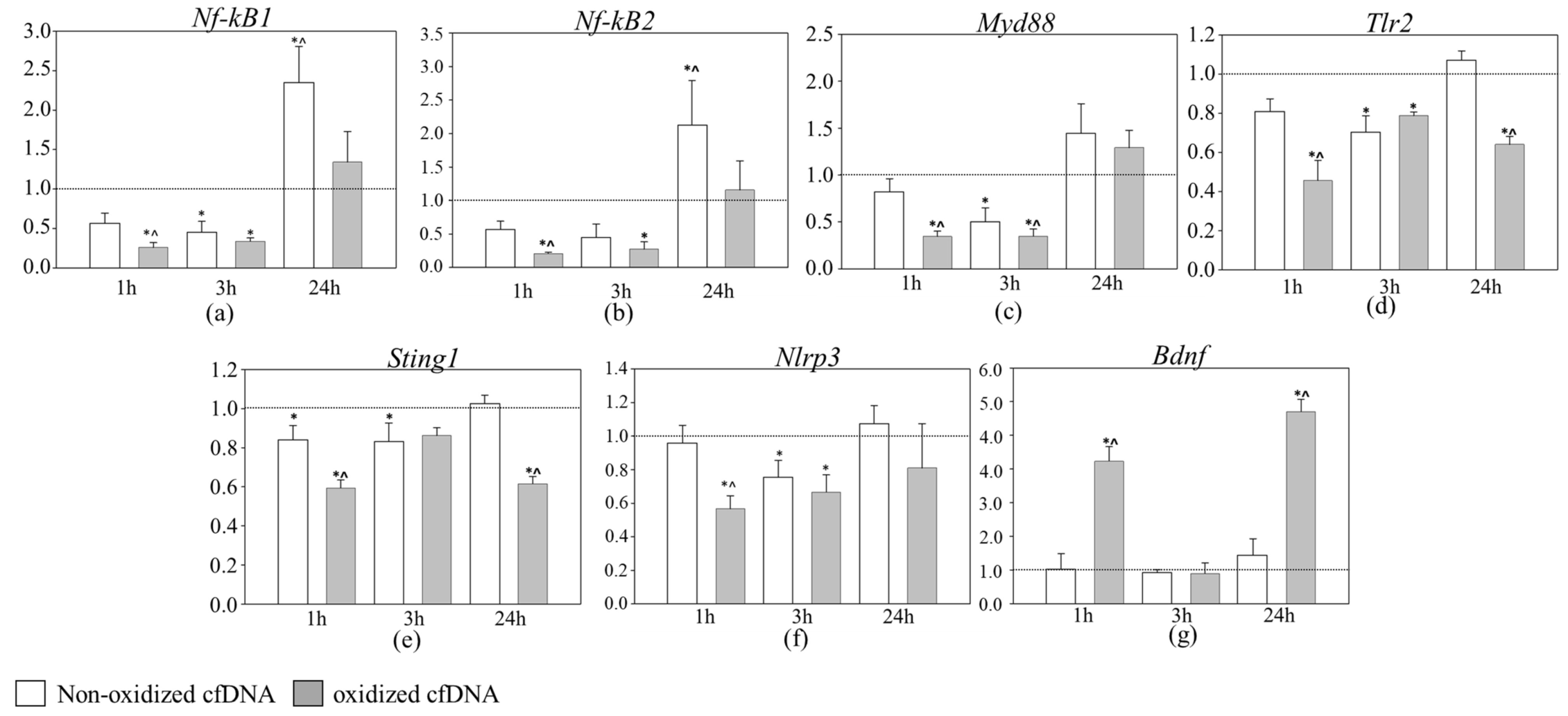
3.2. Dynamics of Inflammation-Related Gene Expression during the First 24 h after cfDNA Treatment
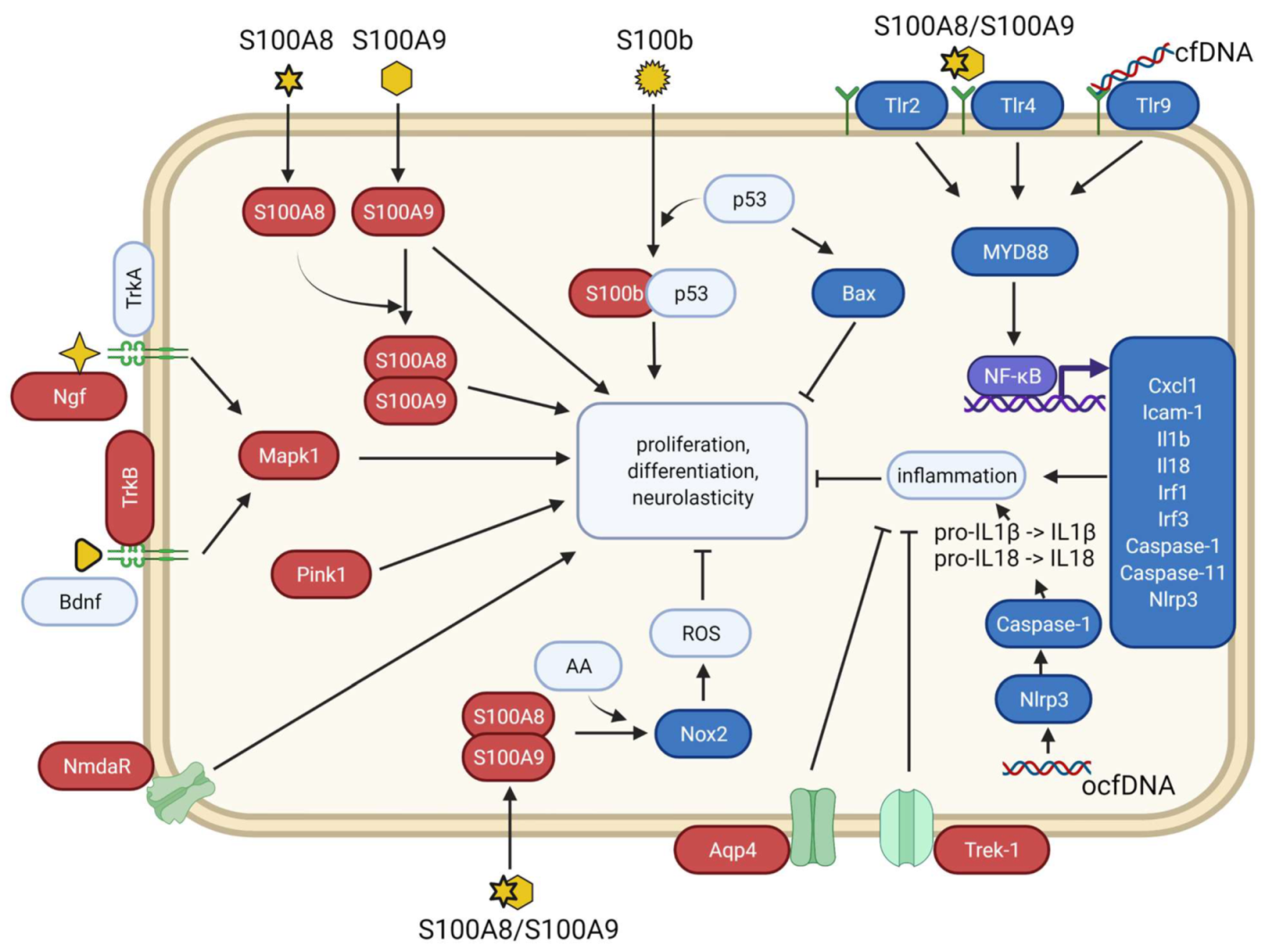
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Cameron, H.A.; McKay, R.D. Adult neurogenesis produces a large pool of new granule cells in the dentate gyrus. J. Comp. Neurol. 2001, 435, 406–417. [Google Scholar] [CrossRef]
- Spalding, K.L.; Bergmann, O.; Alkass, K.; Bernard, S.; Salehpour, M.; Huttner, H.B.; Boström, E.; Westerlund, I.; Vial, C.; Buchholz, B.A.; et al. Dynamics of hippocampal neurogenesis in adult humans. Cell 2013, 153, 1219–1227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schoenfeld, T.J.; Cameron, H.A. Adult neurogenesis and mental illness. Neuropsychopharmacology 2015, 40, 113–128. [Google Scholar] [CrossRef] [Green Version]
- Shohayeb, B.; Diab, M.; Ahmed, M.; Ng, D.C.H. Factors that influence adult neurogenesis as potential therapy. Transl. Neurodegener. 2018, 7, 4. [Google Scholar] [CrossRef]
- Fatouros, I.G.; Destouni, A.; Margonis, K.; Jamurtas, A.Z.; Vrettou, C.; Kouretas, D.; Mastorakos, G.; Mitrakou, A.; Taxildaris, K.; Kanavakis, E.; et al. Cell-free plasma DNA as a novel marker of aseptic inflammation severity related to exercise overtraining. Clin. Chem. 2006, 52, 1820–1824. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gambardella, S.; Limanaqi, F.; Ferese, R.; Biagioni, F.; Campopiano, R.; Centonze, D.; Fornai, F. ccf-mtDNA as a Potential Link Between the Brain and Immune System in Neuro-Immunological Disorders. Front. Immunol. 2019, 10, 1064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O’Connell, G.C.; Petrone, A.B.; Tennant, C.S.; Lucke-Wold, N.; Kabbani, Y.; Tarabishy, A.R.; Chantler, P.D.; Barr, T.L. Circulating extracellular DNA levels are acutely elevated in ischaemic stroke and associated with innate immune system activation. Brain Inj. 2017, 31, 1369–1375. [Google Scholar] [CrossRef]
- Schneck, E.; Samara, O.; Koch, C.; Hecker, A.; Padberg, W.; Lichtenstern, C.; Weigand, M.A.; Uhle, F. Plasma DNA and RNA differentially impact coagulation during abdominal sepsis-an explorative study. J. Surg. Res. 2017, 210, 231–243. [Google Scholar] [CrossRef]
- Khubutia, M.S.; Shabanov, A.K.; Skulachev, M.V.; Bulava, G.V.; Savchenko, I.M.; Grebenchikov, O.A.; Sergeev, A.A.; Zorov, D.B.; Zinovkin, R.A. Mitochondrial and Nuclear DNA in Patients with Severe Polytrauma. Gen. Reanimatol. 2013, 9, 24. [Google Scholar] [CrossRef]
- Li, P.; Stetler, R.A.; Leak, R.K.; Shi, Y.; Li, Y.; Yu, W.; Bennett, M.; Chen, J. Oxidative stress and DNA damage after cerebral ischemia: Potential therapeutic targets to repair the genome and improve stroke recovery. Neuropharmacology 2018, 134, 208–217. [Google Scholar] [CrossRef]
- Shmarina, G.V.; Ershova, E.S.; Simashkova, N.V.; Nikitina, S.G.; Chudakova, J.M.; Veiko, N.N.; Porokhovnik, L.N.; Basova, A.Y.; Shaposhnikova, A.F.; Pukhalskaya, D.A.; et al. Oxidized cell-free DNA as a stress-signaling factor activating the chronic inflammatory process in patients with autism spectrum disorders. J. Neuroinflamm. 2020, 17, 212. [Google Scholar] [CrossRef]
- Ershova, E.S.; Jestkova, E.M.; Martynov, A.V.; Shmarina, G.V.; Umriukhin, P.E.; Bravve, L.V.; Zakharova, N.V.; Kostyuk, G.P.; Saveliev, D.V.; Orlova, M.D.; et al. Accumulation of Circulating Cell-Free CpG-Enriched Ribosomal DNA Fragments on the Background of High Endonuclease Activity of Blood Plasma in Schizophrenic Patients. Int. J. Genom. 2019, 2019, 8390585. [Google Scholar] [CrossRef] [Green Version]
- Shock, L.S.; Thakkar, P.V.; Peterson, E.J.; Moran, R.G.; Taylor, S.M. DNA methyltransferase 1, cytosine methylation, and cytosine hydroxymethylation in mammalian mitochondria. Proc. Natl. Acad. Sci. USA 2011, 108, 3630–3635. [Google Scholar] [CrossRef] [Green Version]
- Yakes, F.M.; Van Houten, B. Mitochondrial DNA damage is more extensive and persists longer than nuclear DNA damage in human cells following oxidative stress. Proc. Natl. Acad. Sci. USA 1997, 94, 514–519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Konkova, M.S.; Kaliyanov, A.A.; Sergeeva, V.A.; Abramova, M.S.; Kostyuk, S.V. Oxidized Cell-Free DNA Is a Factor of Stress Signaling in Radiation-Induced Bystander Effects in Different Types of Human Cells. Int. J. Genom. 2019, 2019, 9467029. [Google Scholar] [CrossRef]
- Filev, A.D.; Shmarina, G.V.; Ershova, E.S.; Veiko, N.N.; Martynov, A.V.; Borzikova, M.A.; Poletkina, A.A.; Dolgikh, O.A.; Veiko, V.P.; Bekker, A.A.; et al. Oxidized Cell-Free DNA Role in the Antioxidant Defense Mechanisms under Stress. Oxid. Med. Cell Longev. 2019, 2019, 1245749. [Google Scholar] [CrossRef]
- Kostyuk, S.V.; Konkova, M.S.; Ershova, E.S.; Alekseeva, A.J.; Smirnova, T.D.; Stukalov, S.V.; Kozhina, E.A.; Shilova, N.V.; Zolotukhina, T.V.; Markova, Z.G.; et al. An exposure to the oxidized DNA enhances both instability of genome and survival in cancer cells. PLoS ONE 2013, 8, e77469. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fernando, M.R.; Jiang, C.; Krzyzanowski, G.D.; Ryan, W.L. New evidence that a large proportion of human blood plasma cell-free DNA is localized in exosomes. PLoS ONE 2017, 12, e0183915. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boyapati, R.K.; Tamborska, A.; Dorward, D.A.; Ho, G.T. Advances in the understanding of mitochondrial DNA as a pathogenic factor in inflammatory diseases. F1000Research 2017, 6, 169. [Google Scholar] [CrossRef] [Green Version]
- Zhivotosky, B.; Orrenius, S. Assessment of apoptosis and necrosis by DNA fragmentation and morphological criteria. Curr. Protoc. Cell Biol. 2001, 12, 18.3.1–18.3.23. [Google Scholar] [CrossRef]
- Ermakov, A.V.; Kostyuk, S.V.; Konkova, M.S.; Egolina, N.A.; Malinovskaya, E.M.; Veiko, N.N. Extracellular DNA fragments. Ann. N. Y. Acad. Sci. 2008, 1137, 41–46. [Google Scholar] [CrossRef]
- Van der Vaart, M.; Pretorius, P.J. The origin of circulating free DNA. Clin. Chem. 2007, 53, 2215. [Google Scholar] [CrossRef] [Green Version]
- Swarup, V.; Rajeswari, M.R. Circulating (cell-free) nucleic acids--a promising, non-invasive tool for early detection of several human diseases. FEBS Lett. 2007, 581, 795–799. [Google Scholar] [CrossRef] [Green Version]
- Zhong, Z.; Liang, S.; Sanchez-Lopez, E.; He, F.; Shalapour, S.; Lin, X.J.; Wong, J.; Ding, S.; Seki, E.; Schnabl, B.; et al. New mitochondrial DNA synthesis enables NLRP3 inflammasome activation. Nature 2018, 560, 198–203. [Google Scholar] [CrossRef]
- Javadi, A.; Shamaei, M.; Mohammadi Ziazi, L.; Pourabdollah, M.; Dorudinia, A.; Seyedmehdi, S.M.; Karimi, S. Qualification study of two genomic DNA extraction methods in different clinical samples. Tanaffos 2014, 13, 41–47. [Google Scholar]
- Filev, A.D.; Silachev, D.N.; Ryzhkov, I.A.; Lapin, K.N.; Babkina, A.S.; Grebenchikov, O.A.; Pisarev, V.M. Effect of Xenon Treatment on Gene Expression in Brain Tissue after Traumatic Brain Injury in Rats. Brain Sci. 2021, 11, 889. [Google Scholar] [CrossRef] [PubMed]
- Gubern, C.; Hurtado, O.; Rodríguez, R.; Morales, J.R.; Romera, V.G.; Moro, M.A.; Lizasoain, I.; Serena, J.; Mallolas, J. Validation of housekeeping genes for quantitative real-time PCR in in-vivo and in-vitro models of cerebral ischaemia. BMC Mol. Biol. 2009, 10, 57. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Swijsen, A.; Nelissen, K.; Janssen, D.; Rigo, J.M.; Hoogland, G. Validation of reference genes for quantitative real-time PCR studies in the dentate gyrus after experimental febrile seizures. BMC Res. Notes 2012, 5, 685. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gholami, K.; Loh, S.Y.; Salleh, N.; Lam, S.K.; Hoe, S.Z. Selection of suitable endogenous reference genes for qPCR in kidney and hypothalamus of rats under testosterone influence. PLoS ONE 2017, 12, e0176368. [Google Scholar] [CrossRef] [Green Version]
- Sun, J.; Pan, X.; Christiansen, L.I.; Yuan, X.L.; Skovgaard, K.; Chatterton, D.E.W.; Kaalund, S.S.; Gao, F.; Sangild, P.T.; Pankratova, S. Necrotizing enterocolitis is associated with acute brain responses in preterm pigs. J. Neuroinflamm. 2018, 15, 180. [Google Scholar] [CrossRef] [PubMed]
- Bresnick, A.R.; Weber, D.J.; Zimmer, D.B. S100 proteins in cancer. Nat. Rev. Cancer 2015, 15, 96–109. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lima Giacobbo, B.; Doorduin, J.; Klein, H.C.; Dierckx, R.A.J.O.; Bromberg, E.; de Vries, E.F.J. Brain-Derived Neurotrophic Factor in Brain Disorders: Focus on Neuroinflammation. Mol. Neurobiol. 2019, 56, 3295–3312. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marlin, M.C.; Li, G. Biogenesis and function of the NGF/TrkA signaling endosome. Int. Rev. Cell Mol. Biol. 2015, 314, 239–257. [Google Scholar]
- Åmellem, I.; Yovianto, G.; Chong, H.T.; Nair, R.R.; Cnops, V.; Thanawalla, A.; Tashiro, A. Role of NMDA Receptors in Adult Neurogenesis and Normal Development of the Dentate Gyrus. eNeuro 2021, 8. [Google Scholar] [CrossRef]
- Agnihotri, S.K.; Sun, L.; Yee, B.K.; Shen, R.; Akundi, R.S.; Zhi, L.; Duncan, M.J.; Cass, W.A.; Büeler, H. PINK1 deficiency is associated with increased deficits of adult hippocampal neurogenesis and lowers the threshold for stress-induced depression in mice. Behav. Brain Res. 2019, 363, 161–172. [Google Scholar] [CrossRef] [PubMed]
- Brown, S.J.; Boussaad, I.; Jarazo, J.; Fitzgerald, J.C.; Antony, P.; Keatinge, M.; Blechman, J.; Schwamborn, J.C.; Krüger, R.; Placzek, M.; et al. PINK1 deficiency impairs adult neurogenesis of dopaminergic neurons. Sci. Rep. 2021, 11, 6617. [Google Scholar] [CrossRef]
- Liang, R.; Yong, S.; Huang, X.; Kong, H.; Hu, G.; Fan, Y. Aquaporin-4 Mediates the Suppressive Effect of Lipopolysaccharide on Hippocampal Neurogenesis. Neuroimmunomodulation 2016, 23, 309–317. [Google Scholar] [CrossRef]
- Djillani, A.; Mazella, J.; Heurteaux, C.; Borsotto, M. Role of TREK-1 in Health and Disease, Focus on the Central Nervous System. Front Pharmacol. 2019, 10, 379. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, Y.; Tian, Y.; Huang, Q.; Wan, Y.; Xu, L.; Wang, W.; Pan, D.; Zhu, S.; Xie, M. Deficiency of TREK-1 potassium channel exacerbates blood-brain barrier damage and neuroinflammation after intracerebral hemorrhage in mice. J. Neuroinflamm. 2019, 16, 96. [Google Scholar] [CrossRef]
- Hummel, E.M.; Hessas, E.; Müller, S.; Beiter, T.; Fisch, M.; Eibl, A.; Wolf, O.T.; Giebel, B.; Platen, P.; Kumsta, R.; et al. Cell-free DNA release under psychosocial and physical stress conditions. Transl. Psychiatry 2018, 8, 236. [Google Scholar] [CrossRef] [Green Version]
- Seo, J.H.; Park, H.S.; Park, S.S.; Kim, C.J.; Kim, D.H.; Kim, T.W. Physical exercise ameliorates psychiatric disorders and cognitive dysfunctions by hippocampal mitochondrial function and neuroplasticity in post-traumatic stress disorder. Exp. Neurol. 2019, 322, 113043. [Google Scholar] [CrossRef]
- Małkiewicz, M.A.; Szarmach, A.; Sabisz, A.; Cubała, W.J.; Szurowska, E.; Winklewski, P.J. Blood-brain barrier permeability and physical exercise. J. Neuroinflamm. 2019, 16, 15. [Google Scholar] [CrossRef] [PubMed]
- Filev, A.D.; Kostyuk, S.V.; Pisarev, V.M.; Tabakov, V.Y.; Umriukhin, P.E. Adaptive anti-oxidative responses to chronic exposure to stress-signaling molecule, oxidized cell-free DNA, in rat neural cells. IOP Conf. Ser. Earth Environ. Sci. 2020, 548, 072039. [Google Scholar] [CrossRef]
- Zhao, X.; Rouhiainen, A.; Li, Z.; Guo, S.; Rauvala, H. Regulation of Neurogenesis in Mouse Brain by HMGB1. Cells 2020, 9, 1714. [Google Scholar] [CrossRef]
- Carletti, B.; Rossi, F. Neurogenesis in the cerebellum. Neuroscientist 2008, 14, 91–100. [Google Scholar] [CrossRef]
- Jackson Chornenki, N.L.; Coke, R.; Kwong, A.C.; Dwivedi, D.J.; Xu, M.K.; McDonald, E.; Marshall, J.C.; Fox-Robichaud, A.E.; Charbonney, E.; Liaw, P.C. Comparison of the source and prognostic utility of cfDNA in trauma and sepsis. Intensive Care Med. Exp. 2019, 7, 29. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pisarev, V.M.; Chumachenko, A.G.; Filev, A.D.; Ershova, E.S.; Kostyuk, S.V.; Veiko, N.N.; Grigoriev, E.K.; Elysina, E.V.; Cherpakov, R.A.; Tutelyan, A.V. Combination of DNA Molecular Biomarkers in the Prediction of Critical Illness Outcome. Gen. Reanimatol. 2019, 15, 31–47. (In Russian) [Google Scholar] [CrossRef]
| Gene | Sequence | Tm | Length | |
|---|---|---|---|---|
| Ppia | F | TCGCGTCTGCTTCGAGCTGT | 64.6 | 135 |
| R | TGGCACATGAATCCTGGAAT | 57.2 | ||
| Nlrp3 | F | GACCAGCCAGAGTGGAATGA | 59.38 | 118 |
| R | TACAAATCGAGATGCGGGAG | 57.49 | ||
| Nf-kB1 | F | GACCGGCAACTCACAGACAG | 60.95 | 170 |
| R | TCATAGATGGCGTCTGACAC | 57.13 | ||
| Nf-kB2 | F | CAATCACCTGCACCAGACAC | 59.12 | 187 |
| R | TCCACTGTGCAACACTGCCT | 62.27 | ||
| Myd88 | F | CTCAGCCTGTCTCCAGGTGT | 61.19 | 148 |
| R | CAAGACGGGTCCAGAACCAG | 60.32 | ||
| Tlr2 | F | GGCTGGAGGTCTCCAGGTCA | 63.10 | 157 |
| R | AGACCTGGAGCTGCCATCAC | 61.91 | ||
| Sting1 | F | GCCATGTCCAGTCCAGGTAC | 60.11 | 153 |
| R | CAAGATGCCAAGCAAGGCGC | 63.15 | ||
| Trkb | F | AAAGGTTAGAAATCATCAAT | 47.66 | 330 |
| R | CCAGAGGGGTATTCTTGCTG | 57.66 | ||
| Bdnf | F | CGTCCACGGACAAGGCAACT | 62.99 | 146 |
| R | CCAGCAGCTCTTCGATCACG | 61.14 | ||
| S100a8 | F | CCTCAGTTTGTGCAGAATAA | 53.5 | 191 |
| R | TATTCTGTAGACATATCCAA | 47.8 | ||
| S100a9 | F | GAAGAGGGAGAAAAGAAATG | 51.4 | 179 |
| R | CTTTGCCGTGGCTGTGGTCA | 63.6 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Filev, A.D.; Kostyuk, S.V.; Umriukhin, P.E.; Pisarev, V.M. Oxidized Cell-Free DNA Rapidly Skews the Transcriptional Profile of Brain Cells toward Boosting Neurogenesis and Neuroplasticity. Curr. Issues Mol. Biol. 2021, 43, 1583-1591. https://doi.org/10.3390/cimb43030112
Filev AD, Kostyuk SV, Umriukhin PE, Pisarev VM. Oxidized Cell-Free DNA Rapidly Skews the Transcriptional Profile of Brain Cells toward Boosting Neurogenesis and Neuroplasticity. Current Issues in Molecular Biology. 2021; 43(3):1583-1591. https://doi.org/10.3390/cimb43030112
Chicago/Turabian StyleFilev, Anton D., Svetlana V. Kostyuk, Pavel E. Umriukhin, and Vladimir M. Pisarev. 2021. "Oxidized Cell-Free DNA Rapidly Skews the Transcriptional Profile of Brain Cells toward Boosting Neurogenesis and Neuroplasticity" Current Issues in Molecular Biology 43, no. 3: 1583-1591. https://doi.org/10.3390/cimb43030112
APA StyleFilev, A. D., Kostyuk, S. V., Umriukhin, P. E., & Pisarev, V. M. (2021). Oxidized Cell-Free DNA Rapidly Skews the Transcriptional Profile of Brain Cells toward Boosting Neurogenesis and Neuroplasticity. Current Issues in Molecular Biology, 43(3), 1583-1591. https://doi.org/10.3390/cimb43030112







