Antimicrobial Peptides in 2014
Abstract
:1. Introduction
2. New Host Defense Peptides Reported in 2014
| APD ID | Name | Source | Peptide amino acid sequence | Unique features 1 |
|---|---|---|---|---|
| 2381 | Gageotetrin A | Bacteria | LE | The shortest lipopeptide |
| 2397 | Sonorensin | Bacteria | CWSCMGHSCWSCMGHSCWSCAGHSCWSCMGHSCWSCMGHSCWSCAGHCCGSCWHGGM | Repeating CWSCXGHS motif |
| 2372 | Baceridin | Bacteria | WAIVLL | The shortest circular peptide consisting entirely of hydrophobic amino acids |
| 2440 | Copsin | Fungi | QNCPTRRGLCVTSGLTACRNHCRSCHRGDVGCVRCSNAQCTGFLGTTCTCINPCPRC | The first fungal defensin with six disulfide bonds |
| 2407 | Hispidalin | Plants | SDYLNNNPLFPRYDIGNVELSTAYRSFANQKAPGRLNQNWALTADYTYR | A unique peptide with 31% similarity to known sequences. Not predicted by existing programs |
| 2477 | EcAMP3 | Plants | GADRCRERCERRHRGDWQGKQRCLMECRRREQEED | The first disulfide-stabilized hairpin-like helical peptide that inhibits phytopathogenic bacteria |
| 2424 | Crotalicidin | Animals | KRFKKFFKKVKKSVKKRLKKIFKKPMVIGVTIPF | Rich in lysine (38%) |
3. New Light on Known Human Antimicrobial Peptides
4. Mechanisms of Action of Antimicrobial Peptides and Genetic Basis of Bacterial Resistance
4.1. Peptide at Work
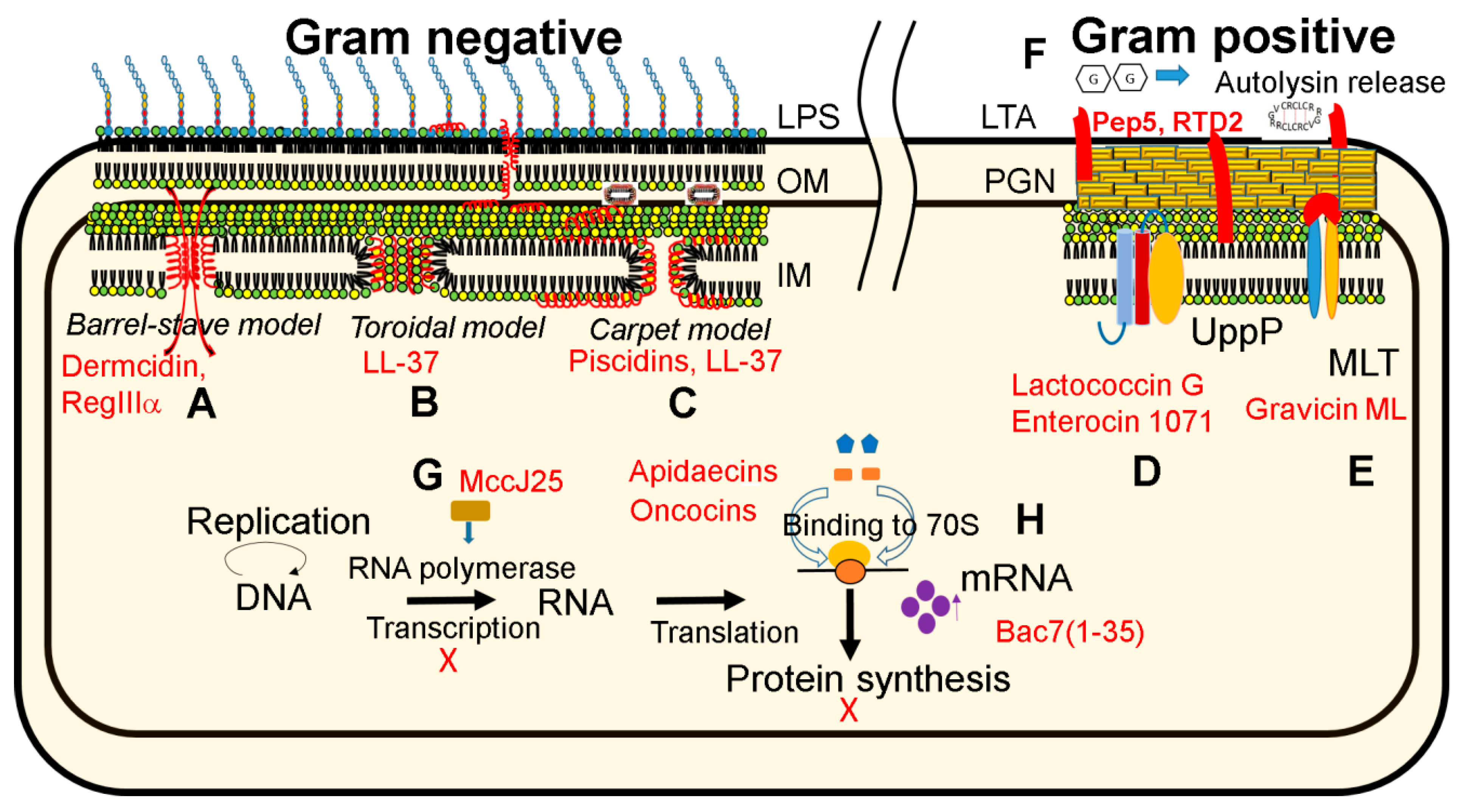
| Mechanism | Lantibiotic Examples | Disulfide-Linked Examples |
|---|---|---|
| Inhibition of cell wall synthesis 1 | Nisin A, lacticin 3147, mersacidin, bovicin HJ50 | HNP1, hBD-3, plectasin, lucifensin, eurocin, copsin |
| Membrane and autolysin release | Pep5 | θ-defensins such as RTD-2 |
| Binding to lipid PE | Duramycins, cinnamycin | Kalata B1, cycloviolacin O2 |
4.2. Resistance Genes for Pathogens and Survival Skills for Commensal Bacteria
4.2.1. Gram-Positive Bacteria
4.2.2. Gram-Negative Bacteria
5. Potential Applications of Antimicrobial Peptides
5.1. Toward Therapeutic Uses
5.2. Peptide Surface Coating
5.3. Nanoparticle-Based Drug Delivery Systems
5.4. Biosensors and Detection
6. Perspectives
Acknowledgements
Author Contributions
Conflicts of Interest
References
- Zasloff, M. Antimicrobial peptides of multicellullar organisms. Nature 2002, 415, 359–365. [Google Scholar] [CrossRef]
- Lai, Y.; Gallo, R.L. AMPed up immunity: How antimicrobial peptides have multiple roles in immune defense. Trends Immunol. 2009, 30, 131–141. [Google Scholar] [CrossRef] [PubMed]
- Hancock, R.E.; Lehrer, R. Cationic peptides: A new source of antibiotics. Trends Biotechnol. 1998, 16, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Yeaman, M.R.; Yount, N.Y. Mechanisms of Antimicrobial Peptide Action and Resistance. Pharmacol. Rev. 2003, 55, 27–55. [Google Scholar] [CrossRef] [PubMed]
- Brogden, K.A. Antimicrobial peptides: Pore formers or metabolic inhibitors in bacteria? Natl. Rev. Microbiol. 2005, 3, 238–250. [Google Scholar] [CrossRef]
- NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2015, 43, D6–D17. [Google Scholar]
- Wang, Z.; Wang, G. APD: The antimicrobial peptide database. Nucleic Acids Res. 2004, 32, D590–D592. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Li, X.; Wang, Z. The updated antimicrobial peptide database and its application in peptide design. Nucleic Acids Res. 2009, 37, D933–D937. [Google Scholar] [CrossRef] [PubMed]
- Wang, G. Improved methods for classification, prediction, and design of antimicrobial peptides. Methods Mol. Biol. 2015, 1268, 43–66. [Google Scholar] [PubMed]
- Timeline of Antimicrobial Peptide Discovery. http://aps.unmc.edu/AP/timeline.php. Accessed on 16 March 2015.
- Chakchouk-Mtibaa, A.; Elleuch, L.; Smaoui, S.; Najah, S.; Sellem, I.; Abdelkafi, S.; Mellouli, L. An antilisterial bacteriocin BacFL31 produced by Enterococcus faecium FL31 with a novel structure containing hydroxyproline residues. Anaerobe 2014, 27C, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Niggemann, J.; Bozko, P.; Bruns, N.; Wodtke, A.; Gieseler, M.T.; Thomas, K.; Jahns, C.; Nimtz, M.; Reupke, I.; Brüser, T.; et al. Baceridin, a cyclic hexapeptide from an epiphytic Bacillus strain, inhibits the proteasome. Chembiochem 2014, 15, 1021–1029. [Google Scholar] [CrossRef] [PubMed]
- Gavrish, E.; Sit, C.S.; Cao, S.; Kandror, O.; Spoering, A.; Peoples, A.; Ling, L.; Fetterman, A.; Hughes, D.; Bissell, A.; et al. Lassomycin, a ribosomally synthesized cyclic peptide, kills mycobacterium tuberculosis by targeting the ATP-dependent protease ClpC1P1P2. Chem. Biol. 2014, 21, 509–518. [Google Scholar] [CrossRef] [PubMed]
- Yamanaka, K.; Reynolds, K.A.; Kersten, R.D.; Ryan, K.S.; Gonzalez, D.J.; Nizet, V.; Dorrestein, P.C.; Moore, B.S. Direct cloning and refactoring of a silent lipopeptide biosynthetic gene cluster yields the antibiotic taromycin A. Proc. Natl. Acad. Sci. U.S.A. 2014, 111, 1957–1962. [Google Scholar] [CrossRef] [PubMed]
- Scholz, R.; Vater, J.; Budiharjo, A.; Wang, Z.; He, Y.; Dietel, K.; Schwecke, T.; Herfort, S.; Lasch, P.; Borriss, R. Amylocyclicin, a Novel Circular Bacteriocin Produced by Bacillus amyloliquefaciens FZB42. J. Bacteriol. 2014, 196, 1842–1852. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.Q.; Yuan, J.; Osapay, G.; Osapay, K.; Tran, D.; Miller, C.J.; Ouellette, A.J.; Selsted, M.E. A cyclic antimicrobial peptide produced in primate leukocytes by the ligation of two truncated alpha-defensins. Science 1999, 286, 498–502. [Google Scholar] [CrossRef] [PubMed]
- Falcao, C.B.; de La Torre, B.G.; Pérez-Peinado, C.; Barron, A.E.; Andreu, D.; Rádis-Baptista, G. Vipericidins: a novel family of cathelicidin-related peptides from the venom gland of South American pit vipers. Amino Acids 2014, 46, 2561–2571. [Google Scholar] [CrossRef] [PubMed]
- Conlon, J.M.; Mechkarska, M. Host-defense peptides with therapeutic potential from skin secretions of frogs from the family pipidae. Pharmaceuticals 2014, 7, 58–77. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Hu, Y.; Li, J.; Liu, Y.; Li, S.; Yan, K.; Wang, X.; Liu, J.; Wang, H. Identification of multiple peptides with antioxidant and antimicrobial activities from skin and its secretions of Hylarana taipehensis, Amolops lifanensis, and Amolops granulosus. Biochimie. 2014, 105, 192–201. [Google Scholar] [CrossRef] [PubMed]
- Ryazantsev, D.Y.; Rogozhin, E.A.; Dimitrieva, T.V.; Drobyazina, P.E.; Khadeeva, N.V.; Egorov, T.A.; Grishin, E.V.; Zavriev, S.K. A novel hairpin-like antimicrobial peptide from barnyard grass (Echinochloa crusgalli L.) seeds: Structure-functional and molecular-genetics characterization. Biochimie 2014, 99, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Li, M.F.; Zhang, B.C.; Li, J.; Sun, L. Sil: A Streptococcus iniae bacteriocin with dual role as an antimicrobial and an immunomodulator that inhibits innate immune response and promotes S. iniae infection. PLoS One. 2014, 9, e96222. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Weisshoff, H.; Hentschel, S.; Zaspel, I.; Jarling, R.; Krause, E.; Pham, T.L. PPZPMs—A novel group of cyclic lipodepsipeptides produced by the Phytophthora alni associated strain Pseudomonas sp. JX090307—The missing link between the viscosin and amphisin group. Nat. Prod. Commun. 2014, 9, 989–996. [Google Scholar] [PubMed]
- Trindade, F.; Amado, F.; Pinto da Costa, J.; Ferreira, R.; Maia, C.; Henriques, I.; Colaco, B.; Vitorino, R. Salivary peptidomic as a tool to disclose new potential antimicrobial peptides. J. Proteomics 2014, 115C, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Bouzid, W.; Verdenaud, M.; Klopp, C.; Ducancel, F.; Noirot, C.; Vetillard, A. de novo sequencing and transcriptome analysis for tetramorium bicarinatum: A comprehensive venom gland transcriptome analysis from an ant species. BMC Genomics 2014, 15, 987. [Google Scholar] [CrossRef] [PubMed]
- Capriotti, A.L.; Cavaliere, C.; Foglia, P.; Piovesana, S.; Samperi, R.; Zenezini Chiozzi, R.; Lagana, A. Development of an analytical strategy for the identification of potential bioactive peptides generated by in vitro tryptic digestion of fish muscle proteins. Anal. Bioanal. Chem. 2015, 407, 845–854. [Google Scholar] [CrossRef] [PubMed]
- Wang, G. Database-guided discovery of potent peptides to combat HIV-1 or superbugs. Pharmaceuticals 2013, 6, 728–758. [Google Scholar] [CrossRef] [PubMed]
- Tareq, F.S.; Lee, M.A.; Lee, H.S.; Lee, Y.J.; Lee, J.S.; Hasan, C.M.; Islam, M.T.; Shin, H.J. Gageotetrins A-C, Noncytotoxic antimicrobial linear lipopeptides from a marine bacterium Bacillus subtilis. Org. Lett. 2014, 16, 928–931. [Google Scholar] [CrossRef] [PubMed]
- Makovitzki, A.; Avrahami, D.; Shai, Y. Ultrashort antibacterial and antifungal lipopeptides. Proc. Natl. Acad. Sci. U.S.A. 2006, 103, 15997–16002. [Google Scholar] [CrossRef] [PubMed]
- First in a New Class of Antibiotics. FDA Consum 2003, 37, 4.
- Chopra, L.; Singh, G.; Choudhary, V.; Sahoo, D.K. Sonorensin: an antimicrobial peptide, belonging to the heterocycloanthracin subfamily of bacteriocins, from a new marine isolate, Bacillus sonorensis MT93. Appl. Environ. Microbiol. 2014, 80, 2981–2990. [Google Scholar] [CrossRef] [PubMed]
- Mygind, P.H.; Fischer, R.L.; Schnorr, K.M.; Hansen, M.T.; Sonksen, C.P.; Ludvigsen, S.; Raventos, D.; Buskov, S.; Christensen, B.; De Maria, L.; et al. Plectasin is a peptide antibiotic with therapeutic potential from a saprophytic fungus. Nature 2005, 437, 975–980. [Google Scholar] [CrossRef] [PubMed]
- Oeemig, J.S.; Lynggaard, C.; Knudsen, D.H.; Hansen, F.T.; Norgaard, K.D.; Schneider, T.; Vad, B.S.; Sandvang, D.H.; Nielsen, L.A.; Neve, S.; et al. Eurocin, a new fungal defensin: structure, lipid binding, and its mode of action. J. Biol. Chem. 2012, 287, 42361–42372. [Google Scholar] [CrossRef] [PubMed]
- Essig, A.; Hofmann, D.; Munch, D.; Gayathri, S.; Kunzler, M.; Kallio, P.T.; Sahl, H.G.; Wider, G.; Schneider, T.; Aebi, M. Copsin, a novel peptide-based fungal antibiotic interfering with the peptidoglycan synthesis. J. Biol. Chem. 2014, 289, 34953–34964. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Verma, H.N.; Sharma, N.K. Cationic bioactive peptide from the seeds of Benincasa hispida. Int. J. Pept. 2014, 2014, 156060. [Google Scholar] [CrossRef] [PubMed]
- Kawabata, S.; Nagayama, R.; Hirata, M.; Shigenaga, T.; Agarwala, K.L.; Saito, T.; Cho, J.; Nakajima, H.; Takagi, T.; Iwanaga, S. Tachycitin, a small granular component in horseshoe crab hemocytes, is an antimicrobial protein with chitin-binding activity. J. Biochem. 1996, 120, 1253–1260. [Google Scholar] [CrossRef] [PubMed]
- Conlon, J.M.; Mechkarska, M.; Emanahmed Coquet, L.; Jouenne, T.; Jérômeleprince Vaudry, H.; Hayes, M.P.; Padgett-Flohr, G. Host defense peptides in skin secretions of the Oregon spotted frog Ranapretiosa: Implications for species resistance to chytridiomycosis. Dev. Comp. Immunol. 2011, 35, 644–649. [Google Scholar] [CrossRef] [PubMed]
- Thomas, S.; Karnik, S.; Barai, R.S.; Jayaraman, V.K.; Idicula-Thomas, S. CAMP: A useful resource for research on antimicrobial peptides. Nucleic Acids Res. 2010, 38, D774–D780. [Google Scholar] [CrossRef] [PubMed]
- Lata, S.; Mishra, N.K.; Raghava, G.P. AntiBP2: Improved version of antibacterial peptide prediction. BMC Bioinformatics 2010, 11, S19. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.; Wang, P.; Lin, W.Z.; Jia, J.H.; Chou, K.C. iAMP-2L: A two-level multi-label classifier for identifying antimicrobial peptides and their functional types. Anal. Biochem. 2013, 436, 168–177. [Google Scholar] [CrossRef] [PubMed]
- Zanetti, M. The role of cathelicidins in the innate host defenses of mammals. Curr. Issues Mol. Biol. 2005, 7, 179–196. [Google Scholar] [PubMed]
- Hao, X.; Yang, H.; Wei, L.; Yang, S.; Zhu, W.; Ma, D.; Yu, H.; Lai, R. Amphibian cathelicidin fills the evolutionary gap of cathelicidin in vertebrate. Amino acids 2012, 43, 677–685. [Google Scholar] [CrossRef] [PubMed]
- Ling, G.; Gao, J.; Zhang, S.; Xie, Z.; Wei, L.; Yu, H.; Wang, Y. Cathelicidins from the bullfrog Rana catesbeiana provides novel template for peptide antibiotic design. PLoS One 2014, 9, e93216. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, B.O.; Ehmann, D.; Precht, J.C.; Castillo, P.A.; Kuchler, R.; Berger, J.; Schaller, M.; Stange, E.F.; Wehkamp, J. Paneth Cell Alpha-defensin 6 (HD-6) is an antimicrobial peptide. Mucosal Immunol. 2014. [Google Scholar] [CrossRef]
- Liu, H.; Yu, H.; Xin, A.; Shi, H.; Gu, Y.; Zhang, Y.; Diao, H.; Lin, D. Production and characterization of recombinant human beta-defensin DEFB120. J. Pept. Sci. 2014, 20, 251–257. [Google Scholar] [CrossRef] [PubMed]
- Gela, A.; Kasetty, G.; Jovic, S.; Ekoff, M.; Nilsson, G.; Morgelin, M.; Kjellstrom, S.; Pease, J.E.; Schmidtchen, A.; Egesten, A. Eotaxin-3 (CCL26) Exerts innate host defense activities that are modulated by mast cell proteases. Allergy 2015, 70, 161–170. [Google Scholar] [CrossRef] [PubMed]
- Becknell, B.; Eichler, T.E.; Beceiro, S.; Li, B.; Easterling, R.S.; Carpenter, A.R.; James, C.L.; McHugh, K.M.; Hains, D.S.; Partida-Sanchez, S.; et al. Ribonucleases 6 and 7 have antimicrobial function in the human and murine urinary tract. Kidney Int. 2015, 87, 151–161. [Google Scholar] [CrossRef] [PubMed]
- Wang, G. Human antimicrobial peptides and proteins. Pharmaceuticals 2014, 7, 545–594. [Google Scholar] [CrossRef] [PubMed]
- Boman, H.G. Antibacterial peptides: Basic facts and emerging concepts. J. Inter. Med. 2003, 254, 197–215. [Google Scholar] [CrossRef]
- Gschwandtner, M.; Zhong, S.; Tschachler, A.; Mlitz, V.; Karner, S.; Elbe-Bürger, A.; Mildner, M. Fetal human keratinocytes produce large amounts of antimicrobial peptides: Involvement of histone-methylation processes. J. Invest Dermatol. 2014, 134, 2192–2201. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.J.; Guerrero-Juarez, C.F.; Hata, T.; Bapat, S.P.; Ramos, R.; Plikus, M.V.; Gallo, R.L. Innate immunity. Dermal adipocytes protect against invasive Staphylococcus aureus skin infection. Science 2015, 347, 67–71. [Google Scholar]
- Lee, P.H.; Ohtake, T.; Zaiou, M.; Murakami, M.; Rudisill, J.A.; Lin, K.H.; Gallo, R.L. Expression of an additional cathelicidin antimicrobial peptide protects against bacterial skin infection. Proc. Natl. Acad. Sci. U.S.A. 2005, 102, 3750–3755. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Wang, I.; Lehrer, R.I. Widespread expression of beta-defensin hBD-1 in human secretory glands and epithelial cells. FEBS Lett. 1996, 396, 319–322. [Google Scholar] [CrossRef] [PubMed]
- Furci, L.; Baldan, R.; Bianchini, V.; Trovato, A.; Ossi, C.; Cichero, P.; Cirillo, D.M. A new role for human alpha-defensin 5 in the fight against Clostridium difficile hypervirulent strains. Infect. Immun. 2015, 83, 986–995. [Google Scholar] [CrossRef] [PubMed]
- Hubert, P.; Herman, L.; Roncarati, P.; Maillard, C.; Renoux, V.; Demoulin, S.; Erpicum, C.; Foidart, J.M.; Boniver, J.; Noel, A.; et al. Altered alpha-defensin 5 expression in cervical squamocolumnar junction: implication in the formation of a viral/tumour-permissive microenvironment. J. Pathol. 2014, 234, 464–477. [Google Scholar] [CrossRef] [PubMed]
- Wiens, M.E.; Smith, J.G. Alpha-Defensin HD5 Inhibits Furin Cleavage of HPV16 L2 to Block Infection. J. Virol. 2014, 89, 2866–2874. [Google Scholar]
- Wommack, A.J.; Ziarek, J.J.; Tomaras, J.; Chileveru, H.R.; Zhang, Y.; Wagner, G.; Nolan, E.M. Discovery and characterization of a disulfide-locked C(2)-symmetric defensin peptide. J. Am. Chem. Soc. 2014, 136, 13494–13497. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, B.O.; Wu, Z.; Nuding, S.; Groscurth, S.; Marcinowski, M.; Beisner, J.; Buchner, J.; Schaller, M.; Stange, E.F.; Wehkamp, J. Reduction of disulphide bonds unmasks potent antimicrobial activity of human beta-defensin 1. Nature 2011, 469, 419–423. [Google Scholar] [CrossRef] [PubMed]
- Porter, E.M.; Poles, M.A.; Lee, J.S.; Naitoh, J.; Bevins, C.L.; Ganz, T. Isolation of human intestinal defensins from ileal neobladder urine. FEBS Lett. 1998, 434, 272–276. [Google Scholar] [CrossRef] [PubMed]
- Chu, H.; Pazgier, M.; Jung, G.; Nuccio, S.P.; Castillo, P.A.; de Jong, M.F.; Winter, M.G.; Winter, S.E.; Wehkamp, J.; Shen, B.; et al. Human Alpha-Defensin 6 Promotes mucosal innate immunity through self-assembled peptide nanonets. Science 2012, 337, 477–481. [Google Scholar] [CrossRef] [PubMed]
- Abou Alaiwa, M.H.; Reznikov, L.R.; Gansemer, N.D.; Sheets, K.A.; Horswill, A.R.; Stoltz, D.A.; Zabner, J.; Welsh, M.J. PH Modulates the activity and synergism of the airway surface liquid antimicrobials beta-defensin-3 and LL-37. Proc. Natl. Acad. Sci. U.S.A. 2014, 111, 18703–18708. [Google Scholar] [CrossRef] [PubMed]
- Johansson, J.; Gudmundsson, G.H.; Rottenberg, M.E.; Berndt, K.D.; Agerberth, B. Conformation-dependent antibacterial activity of the naturally occurring human peptide LL-37. J. Biol. Chem. 1998, 273, 3718–3724. [Google Scholar] [CrossRef] [PubMed]
- Xhindoli, D.; Pacor, S.; Guida, F.; Antcheva, N.; Tossi, A. Native oligomerization determines the mode of action and biological activities of human cathelicidin LL-37. Biochem. J. 2014, 457, 263–275. [Google Scholar] [CrossRef] [PubMed]
- Wang, G. Antimicrobial Peptides: Discovery, Design and Novel Therapeutic Strategies; CABI: Wallingford, UK, 2010. [Google Scholar]
- Singh, D.; Vaughan, R.; Kao, C.C. LL-37 peptide enhancement of signal transduction by toll-like receptor 3 is regulated by pH: identification of a peptide antagonist of LL-37. J. Biol. Chem. 2014, 289, 27614–27624. [Google Scholar] [CrossRef] [PubMed]
- Yamasaki, K.; Schauber, J.; Coda, A.; Lin, H.; Dorschner, R.A.; Schechter, N.M.; Bonnart, C.; Descargues, P.; Hovnanian, A.; Gallo, R.L. Kallikrein-mediated proteolysis regulates the antimicrobial effects of cathelicidins in skin. FASEB J. 2006, 20, 2068–2080. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Mishra, B.; Epand, R.F.; Epand, R.M. High-quality 3D structures shine light on antibacterial, anti-biofilm and antiviral activities of human cathelicidin LL-37 and its fragments. Biochim. Biophys. Acta 2014, 1838, 2160–2172. [Google Scholar] [CrossRef] [PubMed]
- Koziel, J.; Bryzek, D.; Sroka, A.; Maresz, K.; Glowczyk, I.; Bielecka, E.; Kantyka, T.; Pyrc, K.; Svoboda, P.; Pohl, J.; et al. Citrullination alters immunomodulatory function of ll-37 essential for prevention of endotoxin-induced sepsis. J. Immunol. 2014, 192, 5363–5372. [Google Scholar] [CrossRef] [PubMed]
- Picchianti, M.; Russo, C.; Castagnini, M.; Biagini, M.; Soldaini, E.; Balducci, E. NAD-Dependent ADP-ribosylation of the human antimicrobial and immune-modulatory peptide LL-37 by ADP-ribosyltransferase-1. Innate Immun. 2015, 21, 314–321. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, O.E.; Gram, L.; Johnsen, A.H.; Andersson, E.; Bangsbøll, S.; Tjabringa, G.S.; Hiemstra, P.S.; Malm, J.; Egesten, A.; Borregaard, N. Processing of seminal plasma hCAP-18 to ALL-38 by gastricsin: A novel mechanism of generating antimicrobial peptides in vagina. J. Biol. Chem. 2003, 278, 28540–28546. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.T.; Haney, E.F.; Vogel, H.J. The expanding scope of antimicrobial peptide structures and their modes of action. Trends Biotechnol. 2011, 29, 464–472. [Google Scholar] [CrossRef] [PubMed]
- Gazit, E.; Miller, I.R.; Biggin, P.C.; Sansom, M.S.; Shai, Y. Structure and orientation of the mammalian antibacterial peptide cecropin p1 within phospholipid membranes. J. Mol. Biol. 1996, 258, 860–870. [Google Scholar] [CrossRef] [PubMed]
- Perrin, B.S., Jr.; Tian, Y.; Fu, R.; Grant, C.V.; Chekmenev, E.Y.; Wieczorek, W.E.; Dao, A.E.; Hayden, R.M.; Burzynski, C.M.; Venable, R.M.; et al. High-resolution structures and orientations of antimicrobial peptides piscidin 1 and piscidin 3 in fluid bilayers reveal tilting, kinking, and bilayer immersion. J. Am. Chem. Soc. 2014, 136, 3491–3504. [Google Scholar] [CrossRef] [PubMed]
- Fox, R.O., Jr.; Richards, F.M. A voltage-gated ion channel model inferred from the crystal structure of alamethicin at 1.5-Å resolution. Nature 1982, 300, 325–330. [Google Scholar] [CrossRef] [PubMed]
- Kovacs, F.; Quine, J.; Cross, T.A. Validation of the single-stranded channel conformation of gramicidin A by solid-state NMR. Proc. Natl. Acad. Sci. U.S.A. 1999, 96, 7910–7915. [Google Scholar] [CrossRef] [PubMed]
- Burian, M.; Schittek, B. The secrets of dermcidin action. Int. J. Med. Microbiol. 2015, 305, 283–286. [Google Scholar] [CrossRef] [PubMed]
- Song, C.; Weichbrodt, C.; Salnikov, E.S.; Dynowski, M.; Forsberg, B.O.; Bechinger, B.; Steinem, C.; de Groot, B.L.; Zachariae, U.; Zeth, K. Crystal structure and functional mechanism of a human antimicrobial membrane channel. Proc. Natl. Acad. Sci. U.S.A. 2013, 110, 4586–4591. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, S.; Zheng, H.; Derebe, M.G.; Callenberg, K.M.; Partch, C.L.; Rollins, D.; Propheter, D.C.; Rizo, J.; Grabe, M.; Jiang, Q.X.; et al. Antibacterial membrane attack by a pore-forming intestinal c-type lectin. Nature 2014, 505, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Henzler Wildman, K.A.; Lee, D.K.; Ramamoorthy, A. Mechanism of lipid bilayer disruption by the human antimicrobial peptide, LL-37. Biochemistry 2003, 42, 6545–6558. [Google Scholar] [CrossRef] [PubMed]
- Oren, Z.; Lerman, J.C.; Gudmundsson, G.H.; Agerberth, B.; Shai, Y. Structure and organization of the human antimicrobial peptide LL-37 in phospholipid membranes: Relevance to the molecular basis for its non-cell-selective activity. Biochem. J. 1999, 341 (Pt 3), 501–513. [Google Scholar] [CrossRef] [PubMed]
- Kjos, M.; Oppegard, C.; Diep, D.B.; Nes, I.F.; Veening, J.W.; Nissen-Meyer, J.; Kristensen, T. Sensitivity to the two-peptide bacteriocin lactococcin G is dependent on UppP, an enzyme involved in cell-wall synthesis. Mol. Microbiol. 2014, 92, 1177–1187. [Google Scholar] [CrossRef] [PubMed]
- Cotter, P.D. An 'Upp'-turn in bacteriocin receptor identification. Mol. Microbiol. 2014, 92, 1159–1163. [Google Scholar] [CrossRef] [PubMed]
- Wilmes, M.; Stockem, M.; Bierbaum, G.; Schlag, M.; Gotz, F.; Tran, D.Q.; Schaal, J.B.; Ouellette, A.J.; Selsted, M.E.; Sahl, H.G. Killing of staphylococci by theta-defensins involves membrane impairment and activation of autolytic enzymes. Antibiotics (Basel) 2014, 3, 617–631. [Google Scholar] [CrossRef]
- Mukhopadhyay, J.; Sineva, E.; Knight, J.; Levy, R.M.; Ebright, R.H. Antibacterial peptide microcin J25 inhibits transcription by binding within and obstructing the RNA polymerase secondary channel. Mol. Cell 2004, 14, 739–751. [Google Scholar] [CrossRef] [PubMed]
- Krizsan, A.; Volke, D.; Weinert, S.; Strater, N.; Knappe, D.; Hoffmann, R. Insect-derived proline-rich antimicrobial peptides kill bacteria by inhibiting bacterial protein translation at the 70S ribosome. Angew. Chem. Int. Ed. Engl. 2014, 53, 12236–12239. [Google Scholar] [CrossRef] [PubMed]
- Mardirossian, M.; Grzela, R.; Giglione, C.; Meinnel, T.; Gennaro, R.; Mergaert, P.; Scocchi, M. The Host antimicrobial peptide Bac7(1–35) binds to bacterial ribosomal proteins and inhibits protein synthesis. Chem. Biol. 2014, 21, 1639–1647. [Google Scholar] [CrossRef] [PubMed]
- De Leeuw, E.; Li, C.; Zeng, P.; Li, C.; Diepeveen-de Buin, M.; Lu, W.Y.; Breukink, E.; Lu, W. Functional interaction of human neutrophil peptide-1 with the cell wall precursor lipid II. FEBS Lett. 2010, 584, 1543–1548. [Google Scholar] [CrossRef] [PubMed]
- Sass, V.; Schneider, T.; Wilmes, M.; Körner, C.; Tossi, A.; Novikova, N.; Shamova, O.; Sahl, H.G. Human beta-defensin 3 inhibits cell wall biosynthesis in Staphylococci. Infect. Immun. 2010, 78, 2793–2800. [Google Scholar] [CrossRef] [PubMed]
- Bierbaum, G.; Sahl, H.G. Lantibiotics: Mode of action, biosynthesis and bioengineering. Curr. Pharm. Biotechnol. 2009, 10, 2–18. [Google Scholar] [CrossRef] [PubMed]
- Iwamoto, K.; Hayakawa, T.; Murate, M.; Makino, A.; Ito, K.; Fujisawa, T.; Kobayashi, T. Curvature-dependent recognition of ethanolamine phospholipids by duramycin and cinnamycin. Biophys. J. 2007, 93, 1608–1619. [Google Scholar] [CrossRef] [PubMed]
- Henriques, S.T.; Huang, Y.H.; Castanho, M.A.; Bagatolli, L.A.; Sonza, S.; Tachedjian, G.; Daly, N.L.; Craik, D.J. phosphatidylethanolamine binding is a conserved feature of cyclotide-membrane interactions. J. Biol. Chem. 2012, 287, 33629–33643. [Google Scholar] [CrossRef] [PubMed]
- Troeira Henriques, S.; Huang, Y.H.; Chaousis, S.; Wang, C.K.; Craik, D.J. Anticancer and toxic properties of cyclotides are dependent on phosphatidylethanolamine phospholipid targeting. Chembiochem 2014, 15, 1956–1965. [Google Scholar] [CrossRef] [PubMed]
- Otvos, L., Jr.; Insug, O.; Rogers, M.E.; Consolvo, P.J.; Condie, B.A.; Lovas, S.; Bulet, P.; Blaszczyk-Thurin, M. Interaction between heat shock proteins and antimicrobial peptides. Biochemistry 2000, 39, 14150–14159. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.; Kar, R.K.; Jana, J.; Saha, A.; Jana, B.; Krishnamoorthy, J.; Kumar, D.; Ghosh, S.; Chatterjee, S.; Bhunia, A. Indolicidin targets duplex DNA: Structural and mechanistic insight through a combination of spectroscopy and microscopy. ChemMedChem 2014, 9, 2052–2058. [Google Scholar] [CrossRef] [PubMed]
- Wenzel, M.; Chiriac, A.I.; Otto, A.; Zweytick, D.; May, C.; Schumacher, C.; Gust, R.; Albada, H.B.; Penkova, M.; Kramer, U.; et al. Small cationic antimicrobial peptides delocalize peripheral membrane proteins. Proc. Natl. Acad. Sci. U.S.A. 2014, 111, E1409–E1418. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Peterkofsky, A.; Clore, G.M. A novel membrane anchor function for the N-terminal amphipathic sequence of the signal-transducing protein IIAglucose of the Escherichia coli phosphotransferase system. J. Biol. Chem. 2000, 275, 39811–39814. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Li, Y.; Li, X. Correlation of three-dimensional structures with the antibacterial activity of a group of peptides designed based on a nontoxic bacterial membrane anchor. J. Biol. Chem. 2005, 280, 5803–5811. [Google Scholar] [CrossRef] [PubMed]
- Wang, G. Structures of human host defense cathelicidin LL-37 and its smallest antimicrobial peptide KR-12 in lipid micelles. J. Biol. Chem. 2008, 283, 32637–32643. [Google Scholar] [CrossRef] [PubMed]
- Epand, RF; Wang, G.; Berno, B.; Epand, R.M. Lipid segregation explains selective toxicity of a series of fragments derived from the human cathelicidin LL-37. Antimicrob. Agents Chemother. 2009, 53, 3705–3714. [Google Scholar] [CrossRef] [PubMed]
- Bals, R.; Wilson, J.M. Cathelicidins—A family of multifunctional antimicrobial peptides. Cell. Mol. Life Sci. 2003, 60, 711–720. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Epand, R.F.; Mishra, B.; Lushnikova, T.; Thomas, V.C.; Bayles, K.W.; Epand, R.M. Decoding the functional roles of cationic side chains of the major antimicrobial region of human cathelicidin LL-37. Antimicrob. Agents Chemother. 2012, 56, 845–856. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, Y.; Gallo, R.L. Endogenous intracellular cathelicidin enhances TLR9 activation in dendritic cells and macrophages. J. Immunol. 2015, 194, 1274–1284. [Google Scholar] [CrossRef] [PubMed]
- Neumann, A.; Berends, E.T.; Nerlich, A.; Molhoek, E.M.; Gallo, R.L.; Meerloo, T.; Nizet, V.; Naim, H.Y.; von Kockritz-Blickwede, M. The antimicrobial peptide LL-37 facilitates the formation of neutrophil extracellular traps. Biochem. J. 2014, 464, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Chen, Q.; Schmidt, A.P.; Anderson, G.M.; Wang, J.M.; Wooters, J.; Oppenheim, J.J.; Chertov, O. LL-37, the neutrophil granule- and epithelial cell-derived cathelicidin, utilizes formyl peptide receptor-like 1 (FPRL1) as a receptor to chemoattract human peripheral blood neutrophils, monocytes, and T cells. J. Exp. Med. 2000, 192, 1069–1074. [Google Scholar] [CrossRef] [PubMed]
- Elssner, A.; Duncan, M.; Gavrilin, M.; Wewers, M.D. A novel P2X7 receptor activator, the human cathelicidin-derived peptide LL37, induces IL-1 beta processing and release. J. Immunol. 2004, 172, 4987–4994. [Google Scholar] [CrossRef] [PubMed]
- Wan, M.; van der Does, A.M.; Tang, X.; Lindbom, L.; Agerberth, B.; Haeggstrom, J.Z. Antimicrobial peptide LL-37 promotes bacterial phagocytosis by human macrophages. J. Leukoc. Biol. 2014, 95, 971–981. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, S.; Verma, A.; Kim, E.J.; White, M.R.; Hartshorn, K.L. LL-37 modulates human neutrophil responses to influenza a virus. J. Leukoc. Biol. 2014, 96, 931–938. [Google Scholar] [CrossRef] [PubMed]
- Lande, R.; Botti, E.; Jandus, C.; Dojcinovic, D.; Fanelli, G.; Conrad, C.; Chamilos, G.; Feldmeyer, L.; Marinari, B.; Chon, S.; et al. The antimicrobial peptide LL37 is a T-Cell autoantigen in psoriasis. Nat. Commun. 2014, 5, 5621. [Google Scholar] [CrossRef]
- Peschel, A. How do bacteria resist human antimicrobial peptides? Trends Microbiol. 2002, 10, 179–186. [Google Scholar] [CrossRef] [PubMed]
- Gebhard, S.; Fang, C.; Shaaly, A.; Leslie, D.J.; Weimar, M.R.; Kalamorz, F.; Carne, A.; Cook, G.M. Identification and characterization of a bacitracin resistance network in Enterococcus faecalis. Antimicrob. Agents Chemother. 2014, 58, 1425–1433. [Google Scholar] [CrossRef] [PubMed]
- Dintner, S.; Staron, A.; Berchtold, E.; Petri, T.; Mascher, T.; Gebhard, S. Coevolution of ABC transporters and two-component regulatory systems as resistance modules against antimicrobial peptides in Firmicutes Bacteria. J. Bacteriol. 2011, 193, 3851–3862. [Google Scholar] [CrossRef] [PubMed]
- Falord, M.; Karimova, G.; Hiron, A.; Msadek, T. GraXSR proteins interact with the VraFG ABC transporter to form a five-component system required for cationic antimicrobial peptide sensing and resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 2012, 56, 1047–1058. [Google Scholar] [CrossRef] [PubMed]
- Weidenmaier, C.; Peschel, A.; Kempf, V.A.; Lucindo, N.; Yeaman, M.R.; Bayer, A.S. DltABCD- and mprF-mediated cell envelope modifications of Staphylococcus aureus confer resistance to platelet microbicidal proteins and contribute to virulence in a rabbit endocarditis model. Infect. Immun. 2005, 73, 8033–8038. [Google Scholar] [CrossRef] [PubMed]
- Li, M; Cha, D.J.; Lai, Y.; Villaruz, A.E.; Sturdevant, D.E.; Otto, M. The antimicrobial peptide-sensing system aps of Staphylococcus aureus. Mol. Microbiol. 2007, 66, 1136–1147. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.-J.; Bayer, A.S.; Mishra, N.N.; Meehl, M.; Ledala, N.; Yeaman, M.R.; Xiong, Y.Q.; Cheung, A.L. The Staphylococcus aureus two-component regulatory system, GraRS, senses and confers resistance to selected cationic antimicrobial peptides. Infect. Immun. 2012, 80, 74–81. [Google Scholar] [CrossRef] [PubMed]
- Cheung, A.L.; Bayer, A.S.; Yeaman, M.R.; Xiong, Y.Q.; Waring, A.J.; Memmi, G.; Donegan, N.; Chaili, S.; Yang, S.J. Site-specific mutation of the sensor kinase GraS in Staphylococcus aureus alters the adaptive response to distinct cationic antimicrobial peptides. Infect. Immun. 2014, 82, 5336–5345. [Google Scholar] [CrossRef] [PubMed]
- Velarde, J.J.; Ashbaugh, M.; Wessels, M.R. The human antimicrobial peptide LL-37 binds directly to CsrS, a sensor histidine kinase of group A Streptococcus, to activate expression of virulence factors. J. Biol. Chem. 2014, 289, 36315–36324. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.D.; Groisman, E.A. The biology of the PmrA/PmrB two-component system: The major regulator of lipopolysaccharide modifications. Annu. Rev. Microbiol. 2013, 67, 83–112. [Google Scholar] [CrossRef] [PubMed]
- Henderson, J.C.; Fage, C.D.; Cannon, J.R.; Brodbelt, J.S.; Keatinge-Clay, A.T.; Trent, M.S. Antimicrobial peptide resistance of Vibrio cholerae results from an lps modification pathway related to nonribosomal peptide synthetases. ACS Chem. Biol. 2014, 9, 2382–2392. [Google Scholar] [CrossRef] [PubMed]
- Tzeng, Y.L.; Ambrose, K.D.; Zughaier, S.; Zhou, X.; Miller, Y.K.; Shafer, W.M.; Stephens, D.S. Cationic antimicrobial peptide resistance in Neisseria meningitidis. J. Bacteriol. 2005, 187, 5387–5396. [Google Scholar] [CrossRef] [PubMed]
- Balthazar, J.T.; Gusa, A.; Martin, L.E.; Choudhury, B.; Carlson, R.; Shafer, W.M. Lipooligosaccharide structure is an important determinant in the resistance of Neisseria gonorrhoeae to antimicrobial agents of innate host defense. Front. Microbiol. 2011, 2, 30. [Google Scholar] [CrossRef] [PubMed]
- Handing, J.W.; Criss, A.K. The lipooligosaccharide-modifying enzyme LptA enhances gonococcal defence against human neutrophils. Cell. Microbiol. 2014. [Google Scholar] [CrossRef]
- Kandler, J.L.; Joseph, S.J.; Balthazar, J.T.; Dhulipala, V.; Read, T.D.; Jerse, A.E.; Shafer, W.M. Phase-variable expression of lpta modulates the resistance of Neisseria gonorrhoeae to cationic antimicrobial peptides. Antimicrob. Agents Chemother. 2014, 58, 4230–4233. [Google Scholar] [CrossRef] [PubMed]
- Packiam, M.; Yedery, R.D.; Begum, A.A.; Carlson, R.W.; Ganguly, J.; Sempowski, G.D.; Ventevogel, M.S.; Shafer, W.M.; Jerse, A.E. Phosphoethanolamine decoration of Neisseria gonorrhoeae lipid a plays a dual immunostimulatory and protective role during experimental genital tract infection. Infect. Immun. 2014, 82, 2170–2179. [Google Scholar] [CrossRef] [PubMed]
- Groisman, E.A.; Parra-Lopez, C.; Salcedo, M.; Lipps, C.J.; Heffron, F. Resistance to host antimicrobial peptides is necessary for Salmonella virulence. Proc. Natl. Acad. Sci. U.S.A. 1992, 89, 11939–11943. [Google Scholar] [CrossRef] [PubMed]
- Dalebroux, Z.D.; Matamouros, S.; Whittington, D.; Bishop, R.E.; Miller, S.I. PhoPQ regulates acidic glycerophospholipid content of the Salmonella typhimurium outer membrane. Proc. Natl. Acad. Sci. U.S.A. 2014, 111, 1963–1968. [Google Scholar] [CrossRef] [PubMed]
- Matamouros, S.; Miller, S.I. S. Typhimurium strategies to resist killing by cationic antimicrobial peptides. Biochim. Biophys. Acta 2015. [Google Scholar] [CrossRef]
- Dalebroux, Z.D.; Miller, S.I. Salmonellae PhoPQ regulation of the outer membrane to resist innate immunity. Curr Opin Microbiol. 2014, 17, 106–113. [Google Scholar] [CrossRef] [PubMed]
- Cullen, T.W.; Schofield, W.B.; Barry, N.A.; Putnam, E.E.; Rundell, E.A.; Trent, M.S.; Degnan, P.H.; Booth, C.J.; Yu, H.; Goodman, A.L. Gut Microbiota. Antimicrobial peptide resistance mediates resilience of prominent gut commensals during inflammation. Science 2015, 347, 170–175. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Hanke, M.L.; Mishra, B.; Lushnikova, T.; Heim, C.E.; Chittezham Thomas, V.; Bayles, K.W.; Kielian, T. Transformation of human cathelicidin LL-37 into selective, stable, and potent antimicrobial compounds. ACS Chem. Biol. 2014, 9, 1997–2002. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Li, Y.; Han, H.; Miller, D.W.; Wang, G. Solution structures of human LL-37 fragments and NMR-based identification of a minimal membrane-targeting antimicrobial and anticancer region. J. Am. Chem. Soc. 2006, 128, 5776–5785. [Google Scholar] [CrossRef] [PubMed]
- Leake, I. IBD: cathelicidin can reverse intestinal fibrosis in models of colitis. Nat. Rev. Gastroenterol. Hepatol. 2015, 12, 3. [Google Scholar] [PubMed]
- Steinstraesser, L.; Lam, M.C.; Jacobsen, F.; Porporato, P.E.; Chereddy, K.K.; Becerikli, M.; Stricker, I.; Hancock, R.E.; Lehnhardt, M.; Sonveaux, P.; et al. Skin electroporation of a plasmid encoding hCAP-18/LL-37 host defense peptide promotes wound healing. Mol. Ther. 2014, 22, 734–742. [Google Scholar] [CrossRef] [PubMed]
- Woo, J.I.; Kil, S.H.; Brough, D.E.; Lee, Y.J.; Lim, D.J.; Moon, S.K. Therapeutic potential of adenovirus-mediated delivery of beta-defensin 2 for experimental otitis media. Innate Immun. 2015, 21, 215–224. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Curiel, I.; Trujillo, V.; Montoya-Rosales, A.; Rincon, K.; Rivas-Calderon, B.; deHaro-Acosta, J.; Marin-Luevano, P.; Lozano-Lopez, D.; Enciso-Moreno, J.A.; Rivas-Santiago, B. 1,25-Dihydroxyvitamin D3 induces LL-37 and HBD-2 production in keratinocytes from diabetic foot ulcers promoting wound healing: An in vitro model. PLoS One 2014, 9, e111355. [Google Scholar] [CrossRef] [PubMed]
- Mallbris, L.; Edstrom, D.W.; Sundblad, L.; Granath, F.; Stahle, M. UVB upregulates the antimicrobial protein hCAP18 mRNA in human skin. J. Invest. Dermatol. 2005, 125, 1072–1074. [Google Scholar] [CrossRef] [PubMed]
- Eda, N.; Shimizu, K.; Suzuki, S.; Tanabe, Y.; Lee, E.; Akama, T. Effects of yoga exercise on salivary beta-defensin 2. Eur. J. Appl. Physiol. 2013, 113, 2621–2627. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.P.; Li, C.Y.; Suzuki, K.; Chang, C.K.; Chou, K.M.; Fang, S.H. Green tea consumption after intense taekwondo training enhances salivary defense factors and antibacterial capacity. PLoS One 2014, 9, e87580. [Google Scholar] [CrossRef] [PubMed]
- Aboye, T.L.; Ha, H.; Majumder, S.; Christ, F.; Debyser, Z.; Shekhtman, A.; Neamati, N.; Camarero, J.A. Design of a novel cyclotide-based CXCR4 antagonist with anti-human immunodeficiency virus (HIV)-1 activity. J. Med. Chem. 2012, 55, 10729–10734. [Google Scholar] [CrossRef] [PubMed]
- Gunasekera, S.; Foley, F.M.; Clark, R.J.; Sando, L.; Fabri, L.J.; Craik, D.J.; Daly, N.L. Engineering stabilized vascular endothelial growth factor-a antagonists: synthesis, structural characterization, and bioactivity of grafted analogues of cyclotides. J. Med. Chem. 2008, 51, 7697–7704. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.K.; Gruber, C.W.; Cemazar, M.; Siatskas, C.; Tagore, P.; Payne, N.; Sun, G.; Wang, S.; Bernard, C.C.; Craik, D.J. Molecular grafting onto a stable framework yields novel cyclic peptides for the treatment of multiple sclerosis. ACS Chem. Biol. 2014, 9, 156–163. [Google Scholar] [CrossRef] [PubMed]
- Craik, D.J.; Henriques, S.T.; Mylne, J.S.; Wang, C.K. Cyclotide isolation and characterization. Methods Enzymol. 2012, 516, 37–62. [Google Scholar] [PubMed]
- Gunasekera, S.; Daly, N.L.; Anderson, M.A.; Craik, D.J. Chemical synthesis and biosynthesis of the cyclotide family of circular proteins. IUBMB Life 2006, 58, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Jia, X.; Kwon, S.; Wang, C.I.; Huang, Y.H.; Chan, L.Y.; Tan, C.C.; Rosengren, K.J.; Mulvenna, J.P.; Schroeder, C.I.; Craik, D.J. Semienzymatic cyclization of disulfide-rich peptides using sortase A. J. Biol. Chem. 2014, 289, 6627–6638. [Google Scholar] [CrossRef] [PubMed]
- Stanger, K.; Maurer, T.; Kaluarachchi, H.; Coons, M.; Franke, Y.; Hannoush, R.N. Backbone cyclization of a recombinant cystine-knot peptide by engineered sortase A. FEBS Lett. 2014, 588, 4487–4496. [Google Scholar] [CrossRef] [PubMed]
- Lim, K.; Chua, R.R.; Ho, B.; Tambyah, P.A.; Hadinoto, K.; Leong, S.S. Development of a catheter functionalized by a polydopamine peptide coating with antimicrobial and antibiofilm properties. Acta Biomater. 2014, 15, 127–138. [Google Scholar] [CrossRef] [PubMed]
- Godoy-Gallardo, M.; Mas-Moruno, C.; Fernandez-Calderon, M.C.; Perez-Giraldo, C.; Manero, J.M.; Albericio, F.; Gil, F.J.; Rodriguez, D. Covalent immobilization of hLf1-11 peptide on a titanium surface reduces bacterial adhesion and biofilm formation. Acta Biomater. 2014, 10, 3522–3534. [Google Scholar] [CrossRef] [PubMed]
- Lombana, A.; Raja, Z.; Casale, S.; Pradier, C.M.; Foulon, T.; Ladram, A.; Humblot, V. Temporin-SHa peptides grafted on gold surfaces display antibacterial activity. J. Pept. Sci. 2014, 20, 563–569. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Han, X.; He, N.; Chen, Z.; Brooks, C.L., 3rd. Molecular structures of C- and N-terminus cysteine modified cecropin P1 chemically immobilized onto maleimide-terminated self-assembled monolayers investigated by molecular dynamics simulation. J. Phys. Chem. B 2014, 118, 5670–5680. [Google Scholar] [CrossRef] [PubMed]
- Mishra, B.; Basu, A.; Chua, R.R.Y.; Saravanan, R.; Tambyah, P.P.; Ho, B.; Chang, M.W.; Leong, S.S.J. Site specific immobilization of a potent antimicrobial peptide onto silicone catheters: evaluation against urinary tract infection pathogens. J. Mater. Chem. B 2014, 2, 1706–1716. [Google Scholar] [CrossRef]
- Salwiczek, M.; Qu, Y.; Gardiner, J.; Strugnell, R.A.; Lithgow, T.; McLean, K.M.; Thissen, H. Emerging rules for effective antimicrobial coatings. Trends Biotechnol. 2014, 32, 82–90. [Google Scholar] [CrossRef] [PubMed]
- Dutta, D.; Ozkan, J.; Willcox, M.D. Biocompatibility of antimicrobial melimine lenses: Rabbit and human studies. Optom. Vis. Sci. 2014, 91, 570–581. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.W.; Goh, T.W.; Saraswathi, P.; Nyein, C.L.; Setiawan, M.; Riau, A.; Lakshminarayanan, R.; Liu, S.; Tan, D.; Beuerman, R.W.; et al. Effectiveness of antimicrobial peptide immobilization for preventing perioperative cornea implant-associated bacterial infection. Antimicrob. Agents Chemother. 2014, 58, 5229–5238. [Google Scholar] [CrossRef] [PubMed]
- Medeiros, K.A.; Joanitti, G.A.; Silva, L.P. Chitosan nanoparticles for dermaseptin peptide delivery toward tumor cells in vitro. Anticancer Drugs 2014, 25, 323–331. [Google Scholar] [CrossRef] [PubMed]
- Miao, D.; Jiang, M.; Liu, Z.; Gu, G.; Hu, Q.; Kang, T.; Song, Q.; Yao, L.; Li, W.; Gao, X.; et al. Co-administration of dual-targeting nanoparticles with penetration enhancement peptide for antiglioblastoma therapy. Mol. Pharm. 2014, 11, 90–101. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, A.; Ranjan, S.; Zhang, W.; Zou, J.; Pyykko, I.; Kinnunen, P.K. Novel endosomolytic peptides for enhancing gene delivery in nanoparticles. Biochim. Biophys. Acta 2015, 1848, 544–553. [Google Scholar] [CrossRef] [PubMed]
- Keohane, K.; Brennan, D.; Galvin, P.; Griffin, B.T. Silicon microfluidic flow focusing devices for the production of size-controlled PLGA based drug loaded microparticles. Int. J. Pharm. 2014, 467, 60–69. [Google Scholar] [CrossRef] [PubMed]
- Chereddy, K.K.; Her, C.H.; Comune, M.; Moia, C.; Lopes, A.; Porporato, P.E.; Vanacker, J.; Lam, M.C.; Steinstraesser, L.; Sonveaux, P.; et al. PLGA Nanoparticles loaded with host defense peptide ll37 promote wound healing. J. Control. Release 2014, 194, 138–147. [Google Scholar] [CrossRef] [PubMed]
- Laverty, G.; McCloskey, A.P.; Gilmore, B.F.; Jones, D.S.; Zhou, J.; Xu, B. Ultrashort cationic naphthalene-derived self-assembled peptides as antimicrobial nanomaterials. Biomacromolecules 2014, 15, 3429–3439. [Google Scholar] [CrossRef] [PubMed]
- Lillehoj, P.B.; Kaplan, C.W.; He, J.; Shi, W.; Ho, C.M. Rapid, electrical impedance detection of bacterial pathogens using immobilized antimicrobial peptides. J. Lab. Autom. 2014, 19, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Etayash, H.; Jiang, K.; Thundat, T.; Kaur, K. Impedimetric detection of pathogenic gram-positive bacteria using an antimicrobial peptide from class IIa bacteriocins. Anal. Chem. 2014, 86, 1693–1700. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Brehm-Stecher, B.F. Design and evaluation of peptide nucleic acid probes for specific identification of Candida albicans. J. Clin. Microbiol. 2015, 53, 511–521. [Google Scholar] [CrossRef] [PubMed]
- Strauss-Grabo, M.; Atiyem, S.; Le, T.; Kretschmar, M. Decade-long use of the antimicrobial peptide combination tyrothricin does not pose a major risk of acquired resistance with gram-positive bacteria and Candida spp. Pharmazie 2014, 69, 838–841. [Google Scholar]
- Shenkarev, Z.O.; Gizatullina, A.K.; Finkina, E.I.; Alekseeva, E.A.; Balandin, S.V.; Mineev, K.S.; Arseniev, A.S.; Ovchinnikova, T.V. Heterologous expression and solution structure of defensin from lentil Lens culinaris. Biochem. Biophys. Res. Commun. 2014, 451, 252–257. [Google Scholar] [CrossRef] [PubMed]
- De Medeiros, L.N.; Angeli, R.; Sarzedas, C.G.; Barreto-Bergter, E.; Valente, A.P.; Kurtenbach, E.; Almeida, F.C. Backbone dynamics of the antifungal Psd1 pea defensin and its correlation with membrane interaction by NMR spectroscopy. Biochim. Biophys. Acta 2010, 1798, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Neves de Medeiros, L.; Domitrovic, T.; Cavalcante de Andrade, P.; Faria, J.; Barreto Bergter, E.; Weissmüller, G.; Kurtenbach, E. Psd1 binding affinity toward fungal membrane components as assessed by SPR: The role of glucosylceramide in fungal recognition and entry. Biopolymers 2014, 102, 456–464. [Google Scholar] [CrossRef] [PubMed]
- De Paula, V.S.; Pomin, V.H.; Valente, A.P. Unique properties of human β-defensin 6 (hBD6) and glycosaminoglycan complex: sandwich-like dimerization and competition with the chemokine receptor 2 (CCR2) binding site. J. Biol. Chem. 2014, 289, 22969–22979. [Google Scholar] [CrossRef] [PubMed]
- Forde, E.; Humphreys, H.; Greene, C.M.; Fitzgerald-Hughes, D.; Devocelle, M. Potential of host defense peptide prodrugs as neutrophil elastase-dependent anti-infective agents for cystic fibrosis. Antimicrob. Agents Chemother. 2014, 58, 978–985. [Google Scholar] [CrossRef] [PubMed]
- Forde, E.; Devocelle, M. Pro-moieties of antimicrobial peptide prodrugs. Molecules 2015, 20, 1210–1227. [Google Scholar] [CrossRef] [PubMed]
- Neumann, A.; Völlger, L.; Berends, E.T.; Molhoek, E.M.; Stapels, D.A.; Midon, M.; Friães, A.; Pingoud, A.; Rooijakkers, S.H.; Gallo, R.L.; et al. Novel role of the antimicrobial peptide LL-37 in the protection of neutrophil extracellular traps against degradation by bacterial nucleases. J. Innate Immun. 2014, 6, 860–868. [Google Scholar] [CrossRef] [PubMed]
- Sol, A.; Skvirsky, Y.; Nashef, R.; Zelentsova, K.; Burstyn-Cohen, T.; Blotnick, E.; Muhlrad, A.; Bachrach, G. Actin enables the antimicrobial action of LL-37 peptide in the presence of microbial proteases. J. Biol. Chem. 2014, 289, 22926–22941. [Google Scholar] [CrossRef] [PubMed]
- Sol, A.; Wang, G.; Blotnick, E.; Golla, R.; Bachrach, G. Interaction of the core fragments of the LL-37 host defense peptide with actin. RSC Adv. 2015, 5, 9361–9367. [Google Scholar] [CrossRef]
- Svensson, D.; Westman, J.; Wickstrom, C.; Jonsson, D.; Herwald, H.; Nilsson, B.O. Human Endogenous Peptide p33 Inhibits Detrimental Effects of LL-37 on Osteoblast Viability. J. Periodontal. Res. 2015, 50, 80–88. [Google Scholar] [CrossRef] [PubMed]
- Hiemstra, T.F.; Charles, P.D.; Gracia, T.; Hester, S.S.; Gatto, L.; Al-Lamki, R.; Floto, R.A.; Su, Y.; Skepper, J.N.; Lilley, K.S.; et al. Human Urinary Exosomes as Innate Immune Effectors. J. Am. Soc. Nephrol. 2014, 50, 80–88. [Google Scholar]
- Carmona-Ribeiro, A.M.; de Melo Carrasco, L.D. Novel formulations for antimicrobial peptides. Int. J. Mol. Sci. 2014, 15, 18040–18083. [Google Scholar] [CrossRef] [PubMed]
- Pina, A.S.; Batalha, I.L.; Fernandes, C.S.; Aoki, M.A.; Roque, A.C. Exploring the potential of magnetic antimicrobial agents for water disinfection. Water Res. 2014, 66, 160–168. [Google Scholar] [CrossRef] [PubMed]
- Silva, R.R.; Avelino, K.Y.; Ribeiro, K.L.; Franco, O.L.; Oliveira, M.D.; Andrade, C.A. Optical and dielectric sensors based on antimicrobial peptides for microorganism diagnosis. Front. Microbiol. 2014, 5, 443. [Google Scholar] [PubMed]
- Otter, J.A.; Vickery, K.; Walker, J.T.; deLancey Pulcini, E.; Stoodley, P.; Goldenberg, S.D.; Salkeld, J.A.; Chewins, J.; Yezli, S.; Edgeworth, J.D. Surface-attached cells, biofilms and biocide susceptibility: implications for hospital cleaning and disinfection. J. Hosp. Infect. 2015, 89, 16–27. [Google Scholar] [CrossRef] [PubMed]
- Pimentel-Filho Nde, J.; Martins, M.C.; Nogueira, G.B.; Mantovani, H.C.; Vanetti, M.C. Bovicin HC5 and nisin reduce Staphylococcus aureus adhesion to polystyrene and change the hydrophobicity profile and gibbs free energy of adhesion. Int. J. Food Microbiol. 2014, 190, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; McCoy, E.; Zhang, M.; Yang, L. Inhibitory effects of nisin-coated multi-walled carbon nanotube sheet on biofilm formation from Bacillus anthracis spores. J. Environ. Sci. (China) 2014, 26, 2526–2534. [Google Scholar] [CrossRef]
- Singh, A.P.; Preet, S.; Rishi, P. Nisin/β-Lactam adjunct therapy against Salmonella enterica serovar Typhimurium: a mechanistic approach. J. Antimicrob. Chemother. 2014, 69, 1877–1887. [Google Scholar] [CrossRef] [PubMed]
- Dosler, S.; Karaaslan, E. Inhibition and destruction of Pseudomonas aeruginosa biofilms by antibiotics and antimicrobial peptides. Peptides 2014, 62, 32–37. [Google Scholar] [CrossRef] [PubMed]
- Maiti, S.; Patro, S.; Purohit, S.; Jain, S.; Senapati, S.; Dey, N. Effective control of Salmonella infections by employing combinations of recombinant antimicrobial human β-defensins hBD-1 and hBD-2. Antimicrob Agents Chemother. 2014, 58, 6896–6903. [Google Scholar] [CrossRef] [PubMed]
- Donelli, G.; Francolini, I.; Romoli, D.; Guaglianone, E.; Piozzi, A.; Ragunath, C.; Kaplan, J.B. Synergistic activity of dispersin B and cefamandole nafate in inhibition of staphylococcal biofilm growth on polyurethanes. Antimicrob Agents Chemother. 2007, 51, 2733–2740. [Google Scholar] [CrossRef] [PubMed]
- Tong, Z.; Zhang, L.; Ling, J.; Jian, Y.; Huang, L.; Deng, D. An in vitro study on the effect of free amino acids alone or in combination with nisin on biofilms as well as on planktonic bacteria of Streptococcus mutans. PLoS One 2014, 9, e99513. [Google Scholar] [CrossRef] [PubMed]
- De la Fuente-Nunez, C.; Reffuveille, F.; Haney, E.F.; Straus, S.K.; Hancock, R.E. Broad-Spectrum anti-biofilm peptide that targets a cellular stress response. PLoS Pathog. 2014, 10, e1004152. [Google Scholar] [CrossRef] [PubMed]
- Bommarius, B.; Anyanful, A.; Izrayelit, Y.; Bhatt, S.; Cartwright, E.; Wang, W.; Swimm, A.I.; Benian, G.M.; Schroeder, F.C.; Kalman, D. A family of indoles regulate virulence and Shiga toxin production in pathogenic E. coli. PLoS One. 2013, 8, e54456. [Google Scholar] [CrossRef] [PubMed]
- Scopel, M.; Abraham, W.R.; Antunes, A.L.; Henriques, A.T.; Macedo, A.J. Mevalonolactone: an inhibitor of Staphylococcus epidermidis adherence and biofilm formation. Med. Chem. 2014, 10, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Liaqat, I.; Bachmann, R.T.; Edyvean, R.G. Type 2 quorum sensing monitoring, inhibition and biofilm formation in marine microrganisms. Curr. Microbiol. 2014, 68, 342–351. [Google Scholar] [CrossRef] [PubMed]
- Pereira, U.A.; Barbosa, L.C.; Maltha, C.R.; Demuner, A.J.; Masood, M.A.; Pimenta, A.L. γ-Alkylidene-γ-lactones and isobutylpyrrol-2(5H)-ones analogues to rubrolides as inhibitors of biofilm formation by gram-positive and gram-negative bacteria. Bioorg. Med. Chem. Lett. 2014, 24, 1052–1056. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, G.; Mishra, B.; Lau, K.; Lushnikova, T.; Golla, R.; Wang, X. Antimicrobial Peptides in 2014. Pharmaceuticals 2015, 8, 123-150. https://doi.org/10.3390/ph8010123
Wang G, Mishra B, Lau K, Lushnikova T, Golla R, Wang X. Antimicrobial Peptides in 2014. Pharmaceuticals. 2015; 8(1):123-150. https://doi.org/10.3390/ph8010123
Chicago/Turabian StyleWang, Guangshun, Biswajit Mishra, Kyle Lau, Tamara Lushnikova, Radha Golla, and Xiuqing Wang. 2015. "Antimicrobial Peptides in 2014" Pharmaceuticals 8, no. 1: 123-150. https://doi.org/10.3390/ph8010123
APA StyleWang, G., Mishra, B., Lau, K., Lushnikova, T., Golla, R., & Wang, X. (2015). Antimicrobial Peptides in 2014. Pharmaceuticals, 8(1), 123-150. https://doi.org/10.3390/ph8010123







