J3ExoA: A Novel Anti-HIV Immunotoxin Fusion of Anti-Gp120 J3VHH and PE38 Fragment of Pseudomonas Exotoxin A
Abstract
1. Introduction
2. Results
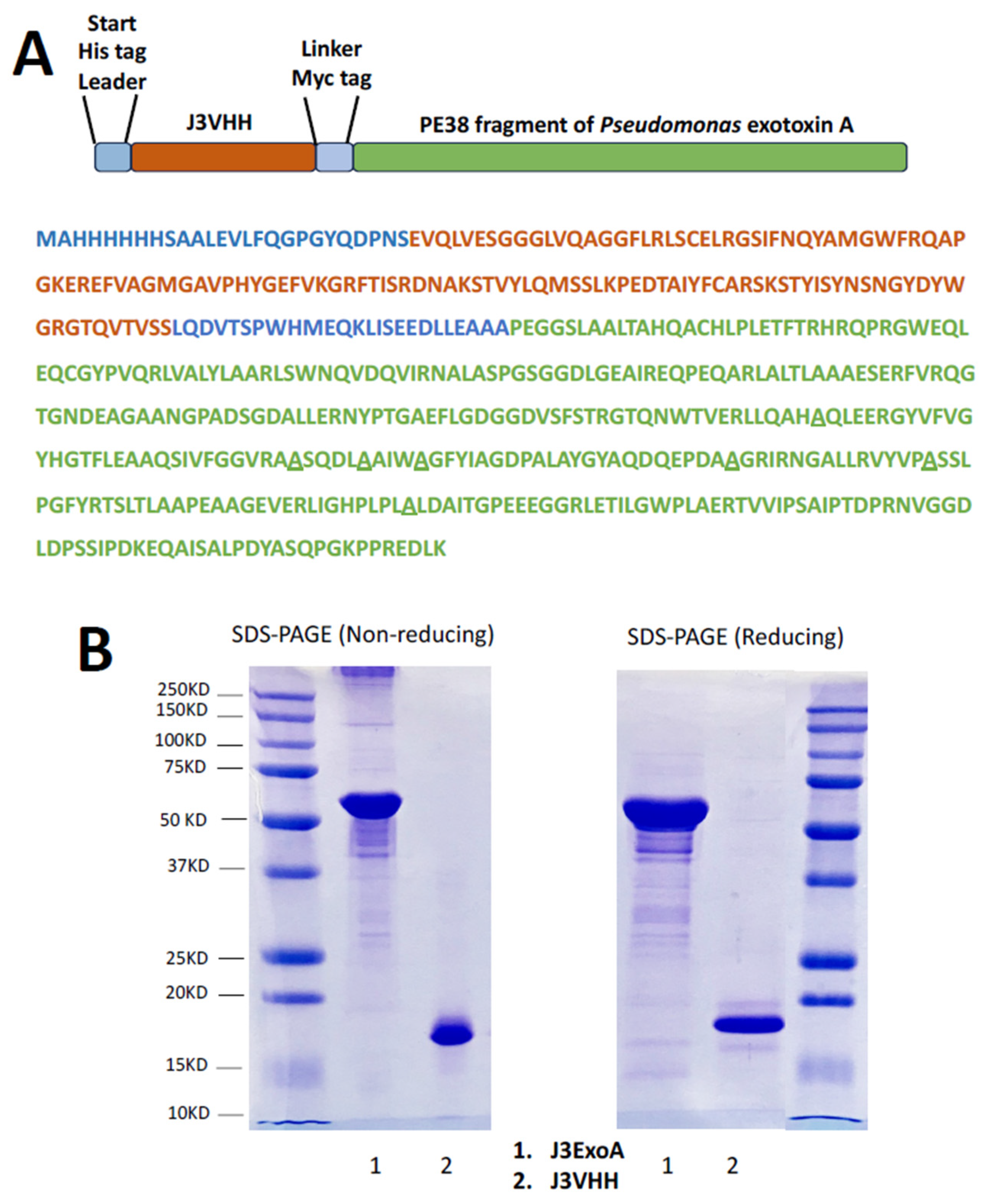
2.1. Design and Production of J3ExoA
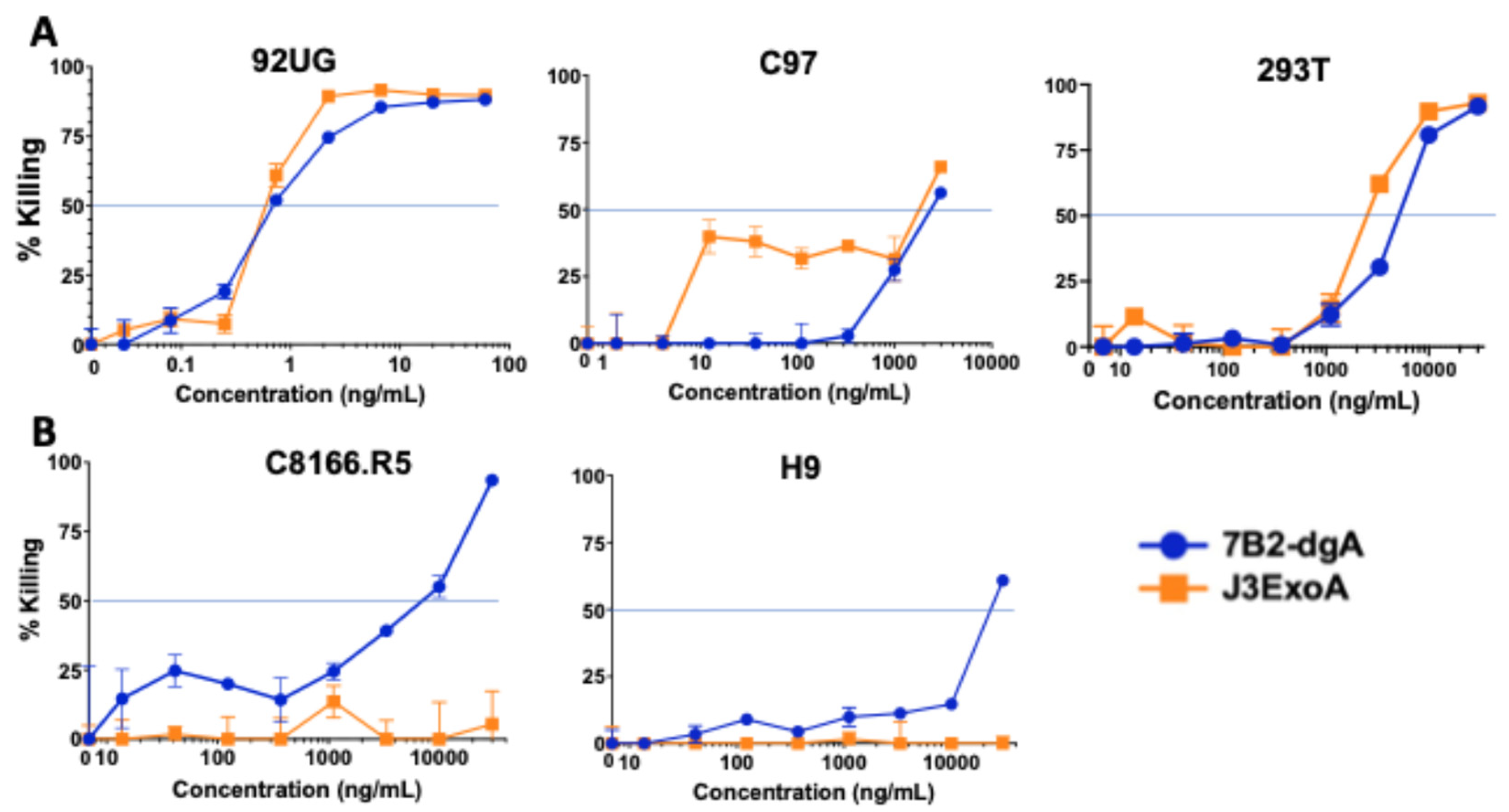
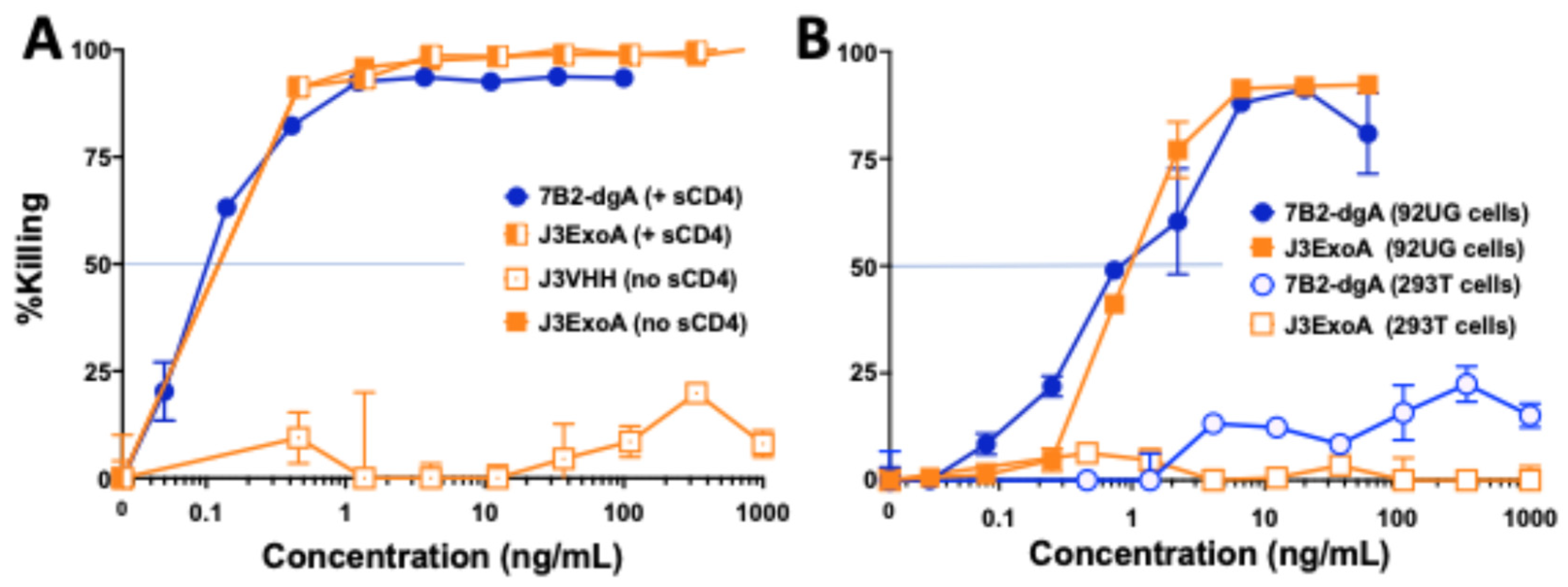
2.2. Comparative Cytotoxicity of J3ExoA and 7B2-dgA on Cell Lines
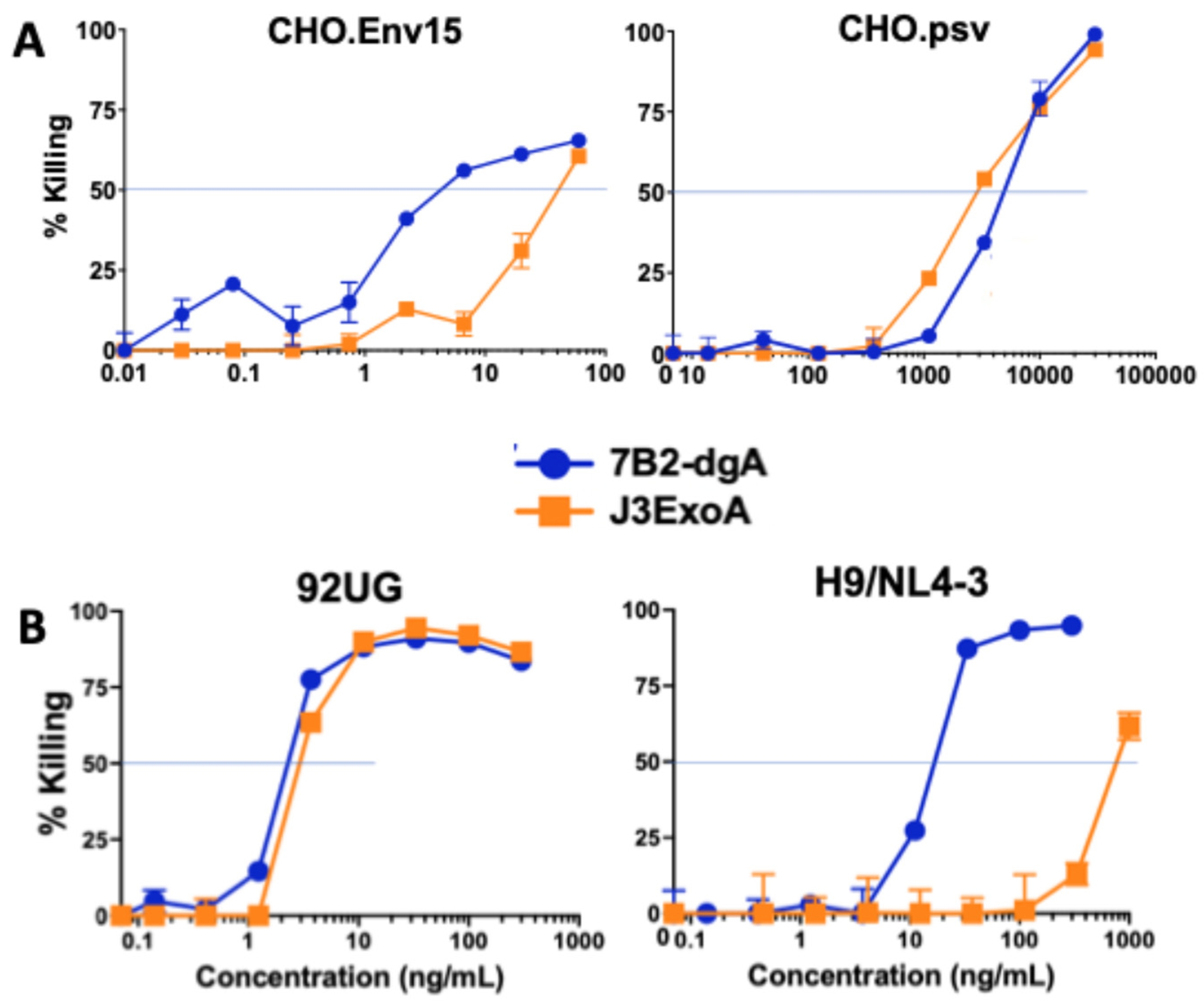
2.3. Ability of ITs to Inhibit the Spread of Tissue Culture Infection
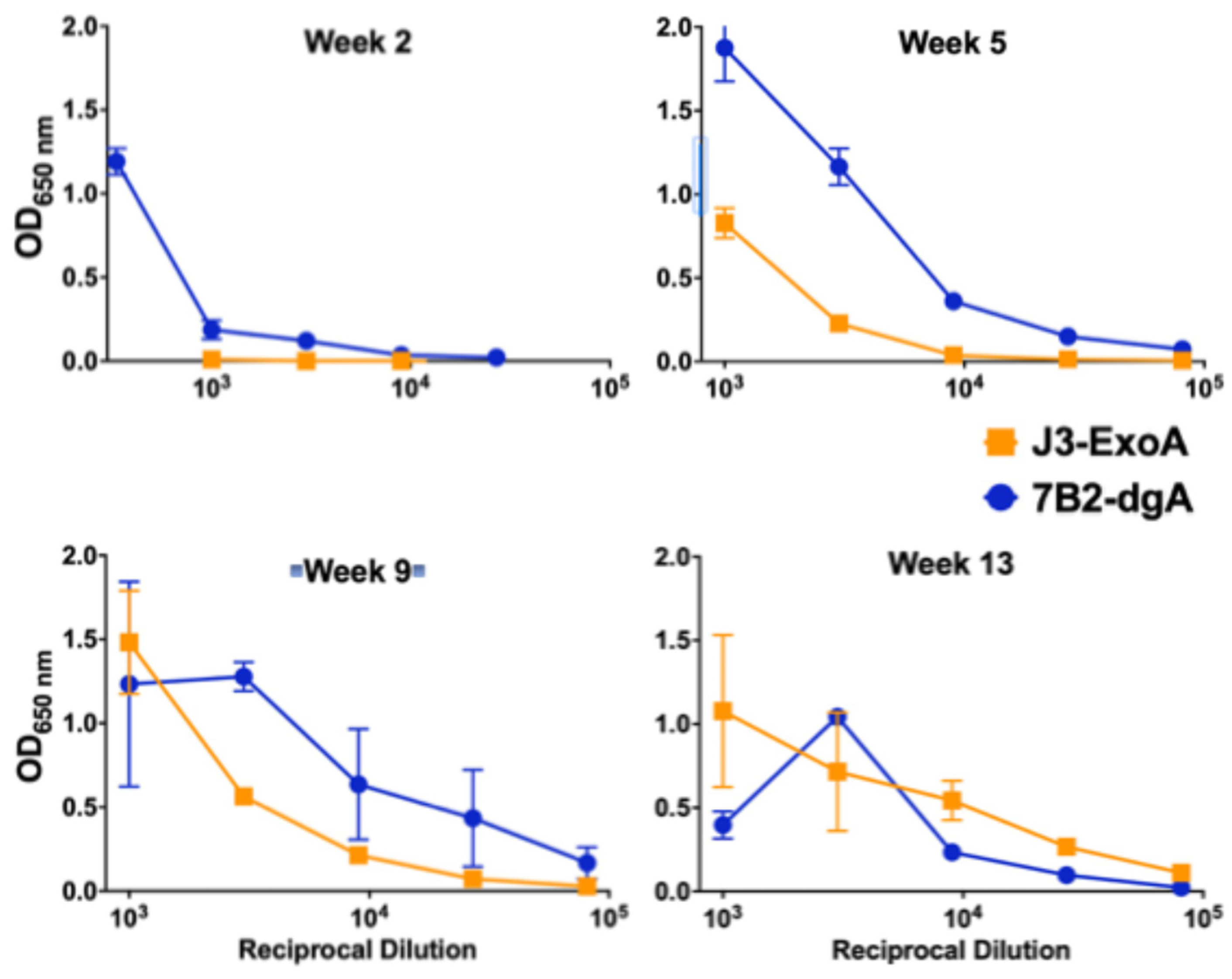
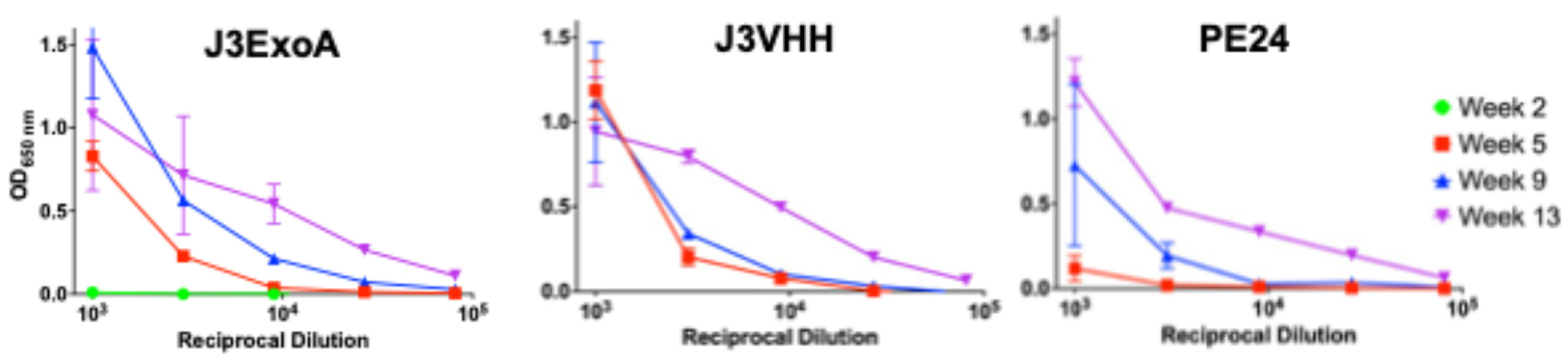
2.4. Immunogenicity and Toxicity of J3ExoA
3. Discussion
4. Materials and Methods
4.1. Production of J3ExoA and J3VHH
4.2. Reagents and Cell Lines
4.3. Immunofluorescence and Flow Cytometry
4.4. Cytotoxicity Assays
4.5. Infection Assays
4.6. Mice and Immunizations
4.7. Enzyme Linked Immunoassays
4.8. Data Display and Statistical Analyses
5. Conclusions
6. Patents
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| A | Absorbance |
| ADC | antibody-drug conjugate |
| ART | antiretroviral therapy |
| CIC | cytotoxic immunoconjugate |
| dgA | deglycosylated ricin A chain |
| Env | HIV envelope protein |
| Gp120 | surface domain of Env |
| Gp41 | transmembrane domain of Env |
| IC50 | concentration providing 50% inhibition |
| IT | immunotoxin |
| LDA | latency disrupting agent |
| mAb | monoclonal antibody |
| PBA | PBS, 1% bovine serum albumin, 0.01% Na azide |
| PHA | Phytohemagglutinin, an activator of T cells |
| PBMC | peripheral blood mononuclear cells |
| R10 | RPMI1640 medium plus 10% fetal bovine serum |
| sCD4 | soluble CD4 |
| VHH | single domain camelid antibody |
References
- Siliciano, J.D.; Siliciano, R.F. HIV cure: The daunting scale of the problem. Science 2024, 383, 703–705. [Google Scholar] [CrossRef] [PubMed]
- Margolis, D.M.; Archin, N.M.; Cohen, M.S.; Eron, J.J.; Ferrari, G.; Garcia, J.V.; Gay, C.L.; Goonetilleke, N.; Joseph, S.B.; Swanstrom, R.; et al. Curing HIV: Seeking to Target and Clear Persistent Infection. Cell 2020, 181, 189–206. [Google Scholar] [CrossRef]
- Siliciano, J.D.; Siliciano, R.F. In Vivo Dynamics of the Latent Reservoir for HIV-1: New Insights and Implications for Cure. Annu. Rev. Pathol. 2022, 17, 271–294. [Google Scholar] [CrossRef]
- Wei, D.G.; Chiang, V.; Fyne, E.; Balakrishnan, M.; Barnes, T.; Graupe, M.; Hesselgesser, J.; Irrinki, A.; Murry, J.P.; Stepan, G.; et al. Histone Deacetylase Inhibitor Romidepsin Induces HIV Expression in CD4 T Cells from Patients on Suppressive Antiretroviral Therapy at Concentrations Achieved by Clinical Dosing. PLoS Pathog. 2014, 10, e1004071. [Google Scholar] [CrossRef]
- Elliott, J.H.; Wightman, F.; Solomon, A.; Ghneim, K.; Ahlers, J.; Cameron, M.J.; Smith, M.Z.; Spelman, T.; McMahon, J.; Velayudham, P.; et al. Activation of HIV transcription with short-course vorinostat in HIV-infected patients on suppressive antiretroviral therapy. PLoS Pathog. 2014, 10, e1004473. [Google Scholar] [CrossRef]
- Hsu, D.C.; Schuetz, A.; Imerbsin, R.; Silsorn, D.; Pegu, A.; Inthawong, D.; Sopanaporn, J.; Visudhiphan, P.; Chuenarom, W.; Keawboon, B.; et al. TLR7 agonist, N6-LS and PGT121 delayed viral rebound in SHIV-infected macaques after antiretroviral therapy interruption. PLoS Pathog. 2021, 17, e1009339. [Google Scholar] [CrossRef]
- Borducchi, E.N.; Liu, J.; Nkolola, J.P.; Cadena, A.M.; Yu, W.H.; Fischinger, S.; Broge, T.; Abbink, P.; Mercado, N.B.; Chandrashekar, A.; et al. Antibody and TLR7 agonist delay viral rebound in SHIV-infected monkeys. Nature 2018, 563, 360–364. [Google Scholar] [CrossRef] [PubMed]
- Molyer, B.; Kumar, A.; Angel, J.B. SMAC Mimetics as Therapeutic Agents in HIV Infection. Front. Immunol. 2021, 12, 780400. [Google Scholar] [CrossRef] [PubMed]
- Pincus, S.H.; Cole, F.M.; Ober, K.; Tokmina-Lukaszewska, M.; Marcotte, T.; Kovacs, E.W.; Copié, V.; Peters, T. Conjugation of Anti-HIV gp41 Monoclonal Antibody to a Drug Capable of Targeting Resting Lymphocytes Produces an Effective Cytotoxic Anti-HIV Immunoconjugate. J. Virol. 2024, 98, e00647-24. [Google Scholar] [CrossRef]
- Pincus, S.H.; Stackhouse, M.; Watt, C.; Ober, K.; Cole, F.M.; Chen, H.-C.; Smith III, A.B.; Peters, T. Soluble CD4 and low molecular weight CD4-mimetic compounds sensitize cells to be killed by anti-HIV cytotoxic immunoconjugates. J. Virol. 2023, 97, e1128. [Google Scholar] [CrossRef]
- Klug, G.; Cole, F.M.; Hicar, M.D.; Watt, C.; Peters, T.; Pincus, S.H. Identification of Anti-gp41 Monoclonal Antibodies That Effectively Target Cytotoxic Immunoconjugates to Cells Infected with Human Immunodeficiency Virus, Type 1. Vaccines 2023, 11, 829. [Google Scholar] [CrossRef]
- Pincus, S.H.; Song, K.; Maresh, G.A.; Hamer, D.H.; Dimitrov, D.S.; Chen, W.; Zhang, M.-Y.; Ghetie, V.F.; Chan-Hui, P.-Y.; Robinson, J.E.; et al. Identification of Human Anti-HIV gp160 Monoclonal Antibodies That Make Effective Immunotoxins. J. Virol. 2017, 91, e01955-16. [Google Scholar] [CrossRef] [PubMed]
- Pincus, S.H.; Song, K.; Maresh, G.A.; Frank, A.; Worthylake, D.; Chung, H.K.; Polacino, P.; Hamer, D.H.; Coyne, C.P.; Rosenblum, M.G.; et al. Design and In Vivo Characterization of Immunoconjugates Targeting HIV gp160. J. Virol. 2017, 91, e01360-16. [Google Scholar] [CrossRef]
- Pincus, S.H.; Fang, H.; Wilkinson, R.A.; Marcotte, T.K.; Robinson, J.E.; Olson, W.C. In vivo efficacy of anti-gp41, but not anti-gp120, immunotoxins in a mouse model of HIV infection. J. Immunol. 2003, 170, 2236–2241. [Google Scholar] [CrossRef] [PubMed]
- Pincus, S.H.; Wehrly, K.; Chesebro, B. Treatment of HIV tissue culture infection with monoclonal antibody-ricin A chain conjugates. J. Immunol. 1989, 142, 3070–3075. [Google Scholar] [CrossRef]
- Dey, B.; Berger, E.A. Towards an HIV cure based on targeted killing of infected cells: Different approaches against acute versus chronic infection. Curr. Opin. HIV AIDS 2015, 10, 207–213. [Google Scholar] [CrossRef]
- Denton, P.W.; Long, J.M.; Wietgrefe, S.W.; Sykes, C.; Spagnuolo, R.A.; Snyder, O.D.; Perkey, K.; Archin, N.M.; Choudhary, S.K.; Yang, K.; et al. Targeted Cytotoxic Therapy Kills Persisting HIV Infected Cells During ART. PLoS Pathog. 2014, 10, e1003872. [Google Scholar] [CrossRef] [PubMed]
- Berger, E.A.; Pastan, I. Immunotoxin complementation of HAART to deplete persisting HIV-infected cell reservoirs. PLoS Pathog. 2010, 6, e1000803. [Google Scholar] [CrossRef]
- Kennedy, P.E.; Bera, T.K.; Wang, Q.C.; Gallo, M.; Wagner, W.; Lewis, M.G.; Berger, E.A.; Pastan, I. Anti-HIV-1 immunotoxin 3B3(Fv)-PE38: Enhanced potency against clinical isolates in human PBMCs and macrophages, and negligible hepatotoxicity in macaques. J. Leukoc. Biol. 2006, 80, 1175–1182. [Google Scholar] [CrossRef]
- Till, M.A.; Zolla-Pazner, S.; Gorny, M.K.; Patton, J.S.; Uhr, J.W.; Vitetta, E.S. Human immunodeficiency virus-infected T cells and monocytes are killed by monoclonal human anti-gp41 antibodies coupled to ricin A chain. Proc. Natl. Acad. Sci. USA 1989, 86, 1987–1991. [Google Scholar] [CrossRef]
- Till, M.A.; Ghetie, V.; Gregory, T.; Patzer, E.J.; Porter, J.P.; Uhr, J.W.; Capon, D.J.; Vitetta, E.S. HIV-infected cells are killed by rCD4-ricin A chain. Science 1988, 242, 1166–1168. [Google Scholar] [CrossRef]
- Dadachova, E.; Kitchen, S.G.; Bristol, G.; Baldwin, G.C.; Revskaya, E.; Empig, C.; Thornton, G.B.; Gorny, M.K.; Zolla-Pazner, S.; Casadevall, A. Pre-Clinical Evaluation of a 213Bi-Labeled 2556 Antibody to HIV-1 gp41 Glycoprotein in HIV-1 Mouse Models as a Reagent for HIV Eradication. PLoS ONE 2012, 7, e31866. [Google Scholar] [CrossRef]
- Dadachova, E.; Patel, M.C.; Toussi, S.; Apostolidis, C.; Morgenstern, A.; Brechbiel, M.W.; Gorny, M.K.; Zolla-Pazner, S.; Casadevall, A.; Goldstein, H. Targeted killing of virally infected cells by radiolabeled antibodies to viral proteins. PLoS Med. 2006, 3, e427. [Google Scholar] [CrossRef]
- Goldstein, H.; Pettoello-Mantovani, M.; Bera, T.K.; Pastan, I.; Berger, E.A. Chimeric toxins targeted to the human immunodeficiency virus type 1 envelope glycoprotein augment the in vivo activity of combination antiretroviral therapy in thy/liv-SCID-Hu mice. J. Inf. Dis. 2000, 181, 921–926. [Google Scholar] [CrossRef] [PubMed]
- Pincus, S.H.; Wehrly, K.; Cole, R.; Fang, H.; Lewis, G.K.; McClure, J.; Conley, A.J.; Wahren, B.; Posner, M.R.; Notkins, A.L.; et al. In vitro effects of anti-HIV immunotoxins directed against multiple epitopes on the HIV-1 envelope glycoprotein gp160. AIDS Res. Hum. Retrovir. 1996, 12, 1041–1051. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Zheng, F.; He, W.; Muyldermans, S.; Wen, Y. Single domain antibody: Development and application in biotechnology and biopharma. Immunol. Rev. 2024, 328, 98–112. [Google Scholar] [CrossRef]
- McCoy, L.E.; Quigley, A.F.; Strokappe, N.M.; Bulmer-Thomas, B.; Seaman, M.S.; Mortier, D.; Rutten, L.; Chander, N.; Edwards, C.J.; Ketteler, R.; et al. Potent and broad neutralization of HIV-1 by a llama antibody elicited by immunization. J. Exp. Med. 2012, 209, 1091–1103. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Chen, L.; Gorman, J.; Wang, S.; Kwon, Y.D.; Lin, B.C.; Louder, M.K.; Rawi, R.; Stancofski, E.D.; Yang, Y.; et al. Structural basis for llama nanobody recognition and neutralization of HIV-1 at the CD4-binding site. Structure 2022, 30, 862–875.E4. [Google Scholar] [CrossRef]
- Mazor, R.; Pastan, I. Immunogenicity of Immunotoxins Containing Pseudomonas Exotoxin A: Causes, Consequences, and Mitigation. Front. Immunol. 2020, 11, 1261. [Google Scholar] [CrossRef]
- Mazor, R.; Eberle, J.A.; Hu, X.; Vassall, A.N.; Onda, M.; Beers, R.; Lee, E.C.; Kreitman, R.J.; Lee, B.; Baker, D.; et al. Recombinant immunotoxin for cancer treatment with low immunogenicity by identification and silencing of human T-cell epitopes. Proc. Natl. Acad. Sci. 2014, 111, 8571–8576. [Google Scholar] [CrossRef]
- Pincus, S.H.; McClure, J. Soluble CD4 enhances the efficacy of immunotoxins directed against gp41 of the human immunodeficiency virus. Proc. Natl. Acad. Sci. USA 1993, 90, 332–336. [Google Scholar] [CrossRef] [PubMed]
- Krowicka, H.; Robinson, J.E.; Clark, R.; Hager, S.; Broyles, S.; Pincus, S.H. Use of tissue culture cell lines to evaluate HIV antiviral resistance. AIDS Res. Hum. Retroviruses 2008, 24, 957–967. [Google Scholar] [CrossRef]
- Vitetta, E.S.; Thorpe, P.E.; Uhr, J.W. Immunotoxins: Magic bullets or misguided missiles? Immunol. Today 1993, 14, 252–258. [Google Scholar] [CrossRef] [PubMed]
- Chapman, A.P. PEGylated antibodies and antibody fragments for improved therapy: A review. Adv. Drug Deliv. Rev. 2002, 54, 531–545. [Google Scholar] [CrossRef]
- Gelber, E.E.; Vitetta, E.S. Effect of immunosuppressive agents on the immunogenicity and efficacy of an immunotoxin in mice. Clin. Cancer Res. An. Off. J. Am. Assoc. Cancer Res. 1998, 4, 1297–1304. [Google Scholar]
- King, E.M.; Mazor, R.; Cuburu, N.; Pastan, I. Low-Dose Methotrexate Prevents Primary and Secondary Humoral Immune Responses and Induces Immune Tolerance to a Recombinant Immunotoxin. J. Immunol. (Baltim. Md. 1950) 2018, 200, 2038–2045. [Google Scholar] [CrossRef]
- Skorupan, N.; Peer, C.J.; Zhang, X.; Choo-Wosoba, H.; Ahmad, M.I.; Lee, M.J.; Rastogi, S.; Sato, N.; Yu, Y.; Pegna, G.J.; et al. Tofacitinib to prevent anti-drug antibody formation against LMB-100 immunotoxin in patients with advanced mesothelin-expressing cancers. Front. Oncol. 2024, 14, 1386190. [Google Scholar] [CrossRef]
- Margolin, K.; Gordon, M.S.; Holmgren, E.; Gaudreault, J.; Novotny, W.; Fyfe, G.; Adelman, D.; Stalter, S.; Breed, J. Phase Ib trial of intravenous recombinant humanized monoclonal antibody to vascular endothelial growth factor in combination with chemotherapy in patients with advanced cancer: Pharmacologic and long-term safety data. J. Clin. Oncol. 2001, 19, 851–856. [Google Scholar] [CrossRef]
- Nakakido, M.; Kinoshita, S.; Tsumoto, K. Development of novel humanized VHH synthetic libraries based on physicochemical analyses. Sci. Rep. 2024, 14, 19533. [Google Scholar] [CrossRef]
- Safarzadeh Kozani, P.; Safarzadeh Kozani, P.; Rahbarizadeh, F. Humanization of the antigen-recognition domain does not impinge on the antigen-binding, cytokine secretion, and antitumor reactivity of humanized nanobody-based CD19-redirected CAR-T cells. J. Transl. Med. 2024, 22, 679. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Quintero, M.L.; Guarnera, E.; Musil, D.; Pekar, L.; Sellmann, C.; Freire, F.; Sousa, R.L.; Santos, S.P.; Freitas, M.C.; Bandeiras, T.M.; et al. On the humanization of VHHs: Prospective case studies, experimental and computational characterization of structural determinants for functionality. Protein Sci. 2024, 33, e5176. [Google Scholar] [CrossRef]
- Martins, C.; Diharce, J.; Nadaradjane, A.A.; de Brevern, A.G. Evaluation of the Potential Impact of In Silico Humanization on V(H)H Dynamics. Int. J. Mol. Sci. 2023, 24, 14586. [Google Scholar] [CrossRef]
- Kashmiri, S.V.; Iwahashi, M.; Tamura, M.; Padlan, E.A.; Milenic, D.E.; Schlom, J. Development of a minimally immunogenic variant of humanized anti-carcinoma monoclonal antibody CC49. Crit. Rev. Oncol. Hematol. 2001, 38, 3–16. [Google Scholar] [CrossRef]
- Brock, H.P.M.; Tekoppele, j.; Hakim, J.; Kerwin, J.A.; Nijenhuis, E.M.; De Groot, C.W.; Bontrop, R.E.; T’Hart, B.A. Prophylactic andtherapeutic effects of a humanized monoclonal antibody against the IL-2 receptor DACLIZUMAB) oncollagen-induced arthritis CIA) n rhesus monkeys. Clin. Exp. Immunol. 2001, 134, 134–141. [Google Scholar]
- Wu, S.; Luo, Q.; Li, F.; Zhang, S.; Zhang, C.; Liu, J.; Shao, B.; Hong, Y.; Tan, T.; Dong, X.; et al. Development of novel humanized CD19/BAFFR bicistronic chimeric antigen receptor T cells with potent antitumor activity against B-cell lineage neoplasms. Br. J. Haematol. 2024, 205, 1361–1373. [Google Scholar] [CrossRef]
- Tanaka, Y.; Miyazaki, Y.; Kawanishi, M.; Yamasaki, H.; Takeuchi, T. Long-term safety and efficacy of anti-TNF multivalent VHH antibodies ozoralizumab in patients with rheumatoid arthritis. RMD Open 2024, 10, e004480. [Google Scholar] [CrossRef] [PubMed]
- Kyuuma, M.; Kaku, A.; Mishima-Tsumagari, C.; Ogawa, B.; Endo, M.; Tamura, Y.; Ishikura, K.I.; Mima, M.; Nakanishi, Y.; Fujii, Y. Unique structure of ozoralizumab, a trivalent anti-TNFalpha NANOBODY((R)) compound, offers the potential advantage of mitigating the risk of immune complex-induced inflammation. Front. Immunol. 2023, 14, 1149874. [Google Scholar] [CrossRef] [PubMed]
- King, C.; Garza, E.N.; Mazor, R.; Linehan, J.L.; Pastan, I.; Pepper, M.; Baker, D. Removing T-cell epitopes with computational protein design. Proc. Natl. Acad. Sci. USA 2014, 111, 8577–8582. [Google Scholar] [CrossRef]
- Frankel, A.E.; Woo, J.H.; Ahn, C.; Pemmaraju, N.; Medeiros, B.C.; Carraway, H.E.; Frankfurt, O.; Forman, S.J.; Yang, X.A.; Konopleva, M.; et al. Activity of SL-401, a targeted therapy directed to interleukin-3 receptor, in blastic plasmacytoid dendritic cell neoplasm patients. Blood 2014, 124, 385–392. [Google Scholar] [CrossRef]
- Candolfi, M.; Kroeger, K.M.; Xiong, W.; Liu, C.; Puntel, M.; Yagiz, K.; Muhammad, A.G.; Mineharu, Y.; Foulad, D.; Wibowo, M.; et al. Targeted toxins for glioblastoma multiforme: Pre-clinical studies and clinical implementation. Anticancer. Agents Med. Chem. 2011, 11, 729–738. [Google Scholar] [CrossRef]
- Geoghegan, E.M.; Zhang, H.; Desai, P.J.; Biragyn, A.; Markham, R.B. Antiviral activity of a single-domain antibody immunotoxin binding to glycoprotein D of herpes simplex virus 2. Antimicrob. Agents Chemother. 2015, 59, 527–535. [Google Scholar] [CrossRef]
- Brinkmann, U.; Buchner, J.; Pastan, I. Independent domain folding of Pseudomonas exotoxin and single-chain immunotoxins: Influence of interdomain connections. Proc. Natl. Acad. Sci. USA 1992, 89, 3075–3079. [Google Scholar] [CrossRef]
- Chen, J.; Kovacs, J.M.; Peng, H.; Rits-Volloch, S.; Lu, J.; Park, D.; Zablowsky, E.; Seaman, M.S.; Chen, B. Effect of the cytoplasmic domain on antigenic characteristics of HIV-1 envelope glycoprotein. Science 2015, 349, 191–195. [Google Scholar] [CrossRef] [PubMed]
- Pitts, T.W.; Bohanon, M.J.; Leach, M.F.; McQuade, T.J.; Marschke, C.K.; Merritt, J.A.; Wierenga, W.; Nicholas, J.A. Soluble CD4-PE40 Is Cytotoxic for a Transfected MammalianmCell Line Stably Expressing the Envelope Protein of Human Immunodeficiency Virus (HIV-1), and Cytotoxicity Is Variably Inhibited by the Sera of HIV-1-Infected Patients. AIDS Res. Hum. Retroviruses 1991, 7, 742–749. [Google Scholar] [CrossRef] [PubMed]
- Adachi, A.; Gendelman, H.E.; Koenig, S.; Folks, T.; Willey, R.; Rabson, A.; Martin, M.A. Production of acquired immunodeficiency syndrome-associated retrovirus in human and nonhuman cells transfected with an infectious molecular clone. J. Virol. 1986, 59, 284–291. [Google Scholar] [CrossRef]
- Santra, S.; Tomaras, G.D.; Warrier, R.; Nicely, N.I.; Liao, H.X.; Pollara, J.; Liu, P.; Alam, S.M.; Zhang, R.; Cocklin, S.L.; et al. Human Non-neutralizing HIV-1 Envelope Monoclonal Antibodies Limit the Number of Founder Viruses during SHIV Mucosal Infection in Rhesus Macaques. PLoS Pathog. 2015, 11, e1005042. [Google Scholar] [CrossRef] [PubMed]
- Walker, L.M.; Huber, M.; Doores, K.J.; Falkowska, E.; Pejchal, R.; Julien, J.-P.; Wang, S.-K.; Ramos, A.; Chan-Hui, P.-Y.; Moyle, M.; et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 2011, 477, 466–470. [Google Scholar] [CrossRef]
- Gauduin, M.-C.; Allaway, G.P.; Olson, W.C.; Weir, R.; Maddon, P.J.; Koup, R.A. CD4-IgG2 protects Hu-PBL-SCID mice against challenge by primary HIV-1 isolates. J. Virol. 1998, 72, 3475–3478. [Google Scholar] [CrossRef]
- Allaway, G.P.; Davis-Bruno, K.L.; Beaudry, G.A.; Garcia, E.B.; Wong, E.L.; Ryder, A.M.; Hasel, K.W.; Gauduin, M.-C.; Koup, R.A.; McDougal, S.; et al. Expression and characterization of CD4-IgG2, a novel heterotetramer that neutralizes primary HIV-1 isolates. AIDS Res. Hum. Retroviruses 1995, 11, 533–539. [Google Scholar] [CrossRef]
- Pincus, S.H.; Marcotte, T.K.; Forsyth, B.M.; Fang, H. In vivo testing of anti-HIV immunotoxins. In Immunotoxin Methods and Protocols; Pincus, S.H.; Marcotte, T.K.; Forsyth, B.M.; Fang, H. Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2001; Volume 166, pp. 277–294. [Google Scholar]
- Chackerian, B.; Long, E.M.; Luciw, P.A.; Overbaugh, J. HIV-1 co-receptors participates in post-entry stages of the virus replication cycle and function in SIV infection. J. Virol. 1997, 71, 3932–3939. [Google Scholar] [CrossRef]
- Maddaloni, M.; Cooke, C.; Wilkinson, R.; Stout, A.V.; Eng, L.; Pincus, S.H. Immunological characteristics associated with protective efficacy of antibodies to ricin. J. Immunol. 2004, 172, 6221–6228. [Google Scholar] [CrossRef] [PubMed]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pincus, S.H.; Luo, K.; Peters, T.; Gordy, J.T.; Cole, F.M.; Klug, G.; Ober, K.; Marcotte, T.K.; Markham, R.B. J3ExoA: A Novel Anti-HIV Immunotoxin Fusion of Anti-Gp120 J3VHH and PE38 Fragment of Pseudomonas Exotoxin A. Pharmaceuticals 2025, 18, 1305. https://doi.org/10.3390/ph18091305
Pincus SH, Luo K, Peters T, Gordy JT, Cole FM, Klug G, Ober K, Marcotte TK, Markham RB. J3ExoA: A Novel Anti-HIV Immunotoxin Fusion of Anti-Gp120 J3VHH and PE38 Fragment of Pseudomonas Exotoxin A. Pharmaceuticals. 2025; 18(9):1305. https://doi.org/10.3390/ph18091305
Chicago/Turabian StylePincus, Seth H., Kun Luo, Tami Peters, James T. Gordy, Frances M. Cole, Grant Klug, Kelli Ober, Tamera K. Marcotte, and Richard B. Markham. 2025. "J3ExoA: A Novel Anti-HIV Immunotoxin Fusion of Anti-Gp120 J3VHH and PE38 Fragment of Pseudomonas Exotoxin A" Pharmaceuticals 18, no. 9: 1305. https://doi.org/10.3390/ph18091305
APA StylePincus, S. H., Luo, K., Peters, T., Gordy, J. T., Cole, F. M., Klug, G., Ober, K., Marcotte, T. K., & Markham, R. B. (2025). J3ExoA: A Novel Anti-HIV Immunotoxin Fusion of Anti-Gp120 J3VHH and PE38 Fragment of Pseudomonas Exotoxin A. Pharmaceuticals, 18(9), 1305. https://doi.org/10.3390/ph18091305








