Research Progress on Rolling Circle Amplification (RCA)-Based Biomedical Sensing
Abstract
1. Introduction
2. Enzyme-Aided RCA Biosensor
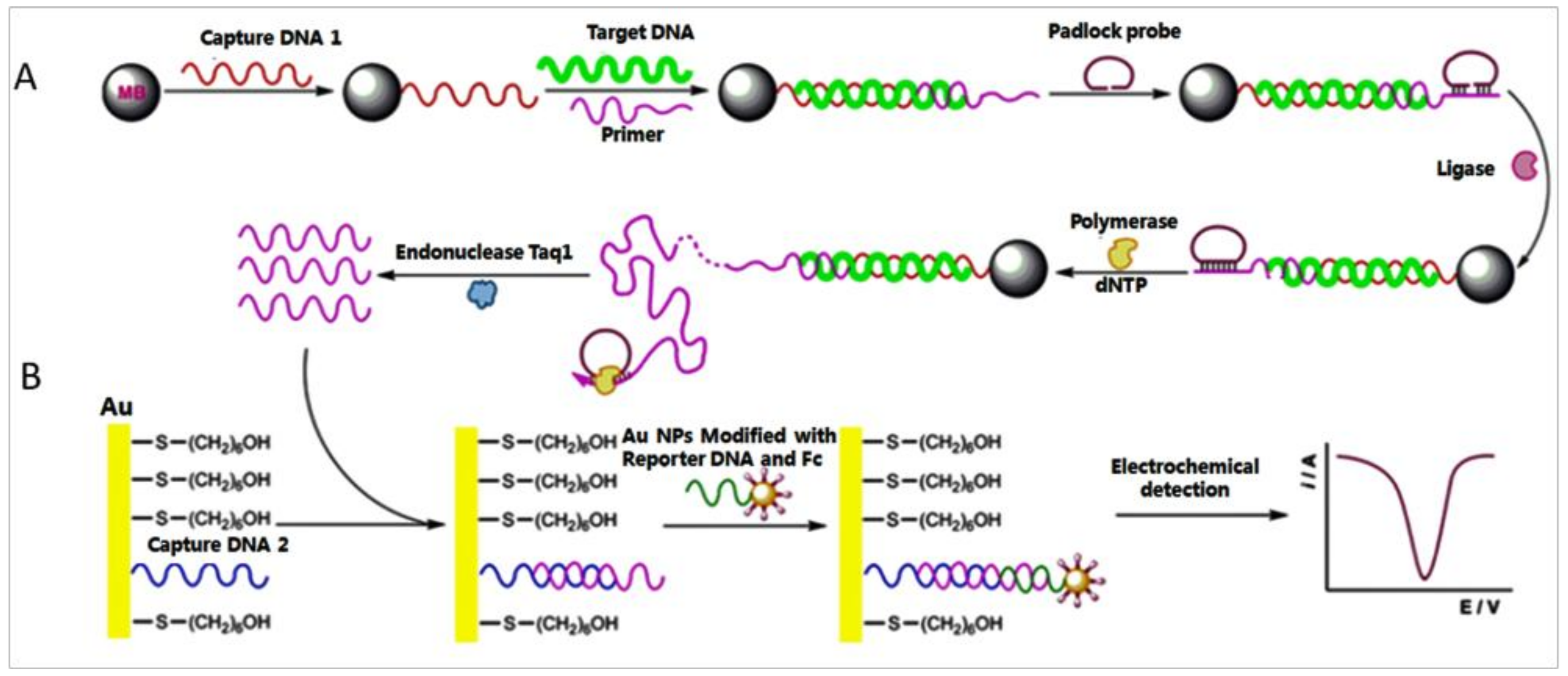
3. AuNP-RCA Biosensor for Multiple Pathogens Detection
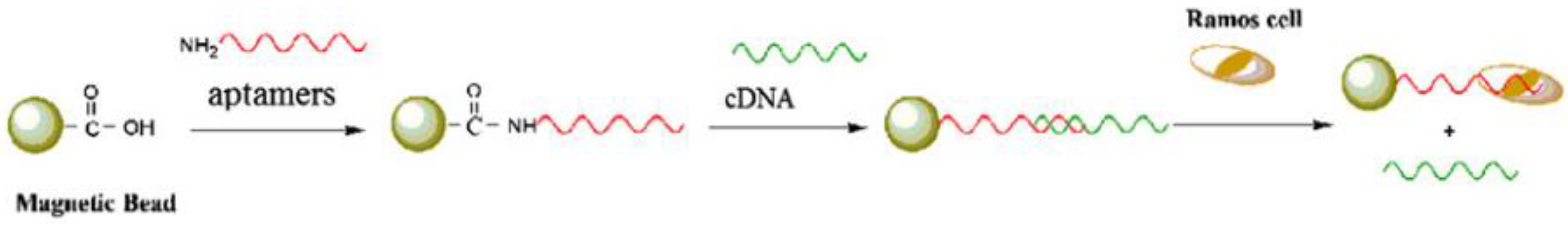
4. Aptamer Biosensor Based on RCA
5. Micromolecular Biosensor Based on Electrochemistry Coupled to RCA
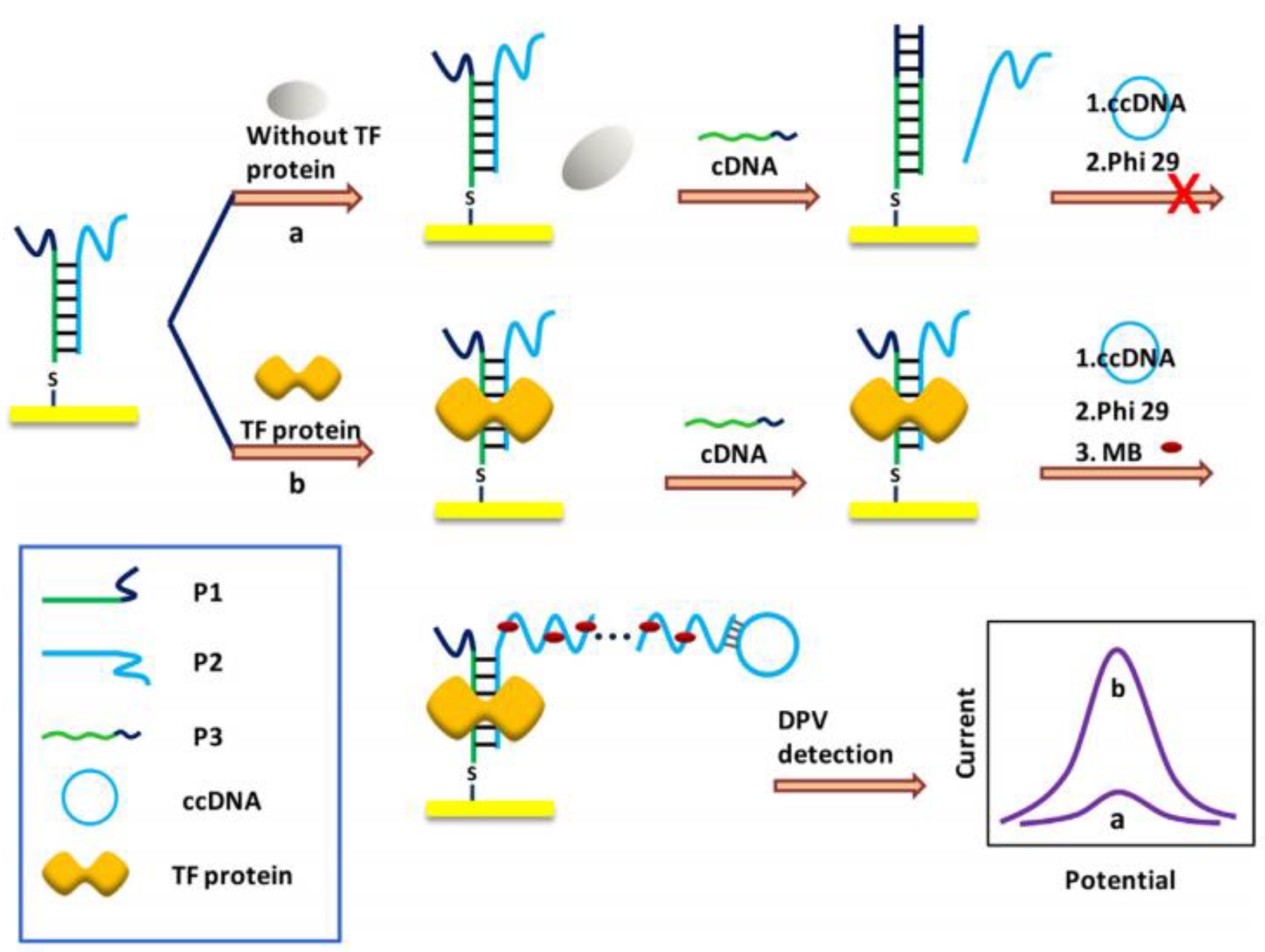
6. Transcription Factor (TF) Biosensor Based on RCA
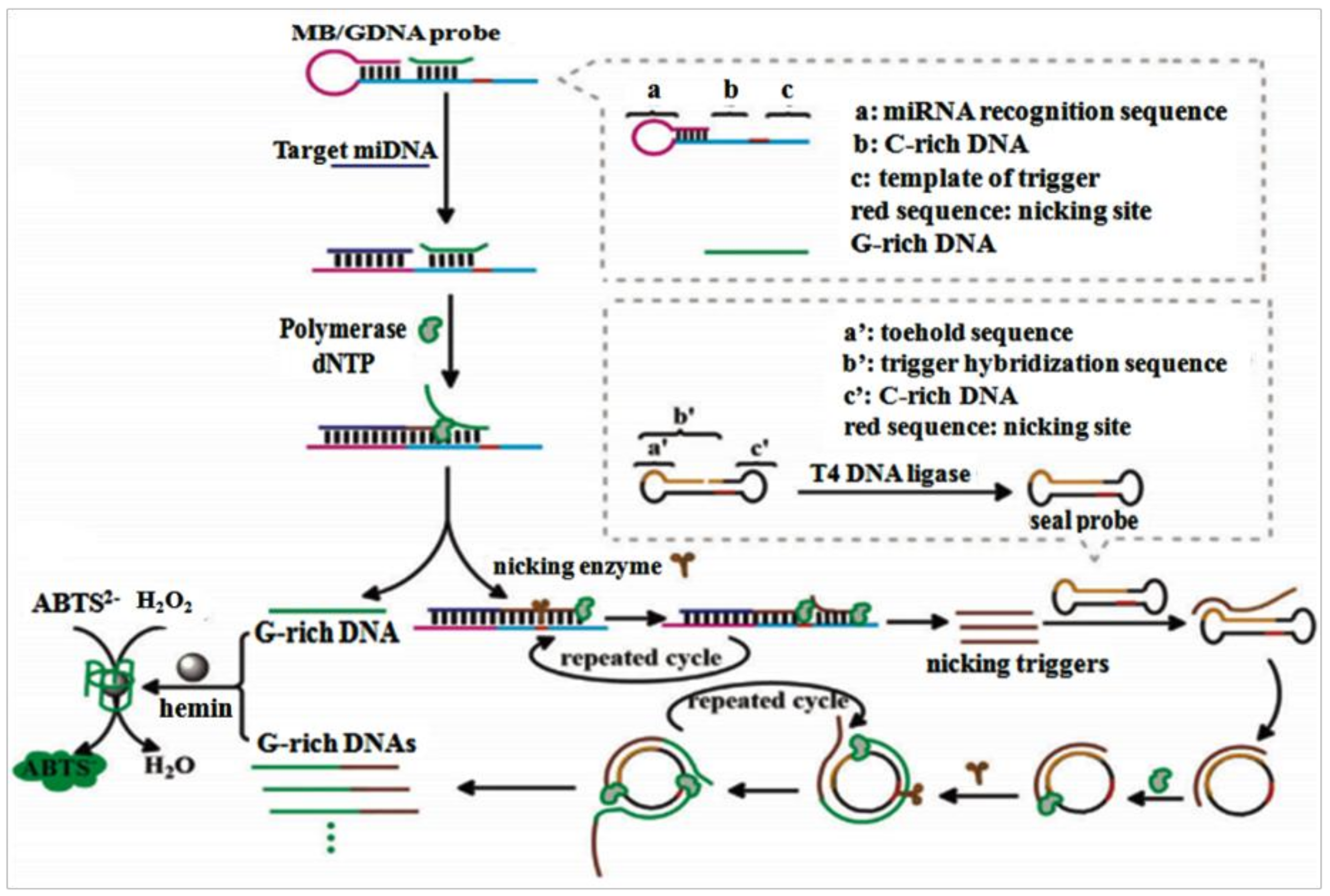
7. MiRNAs Biosensor Based on RCA
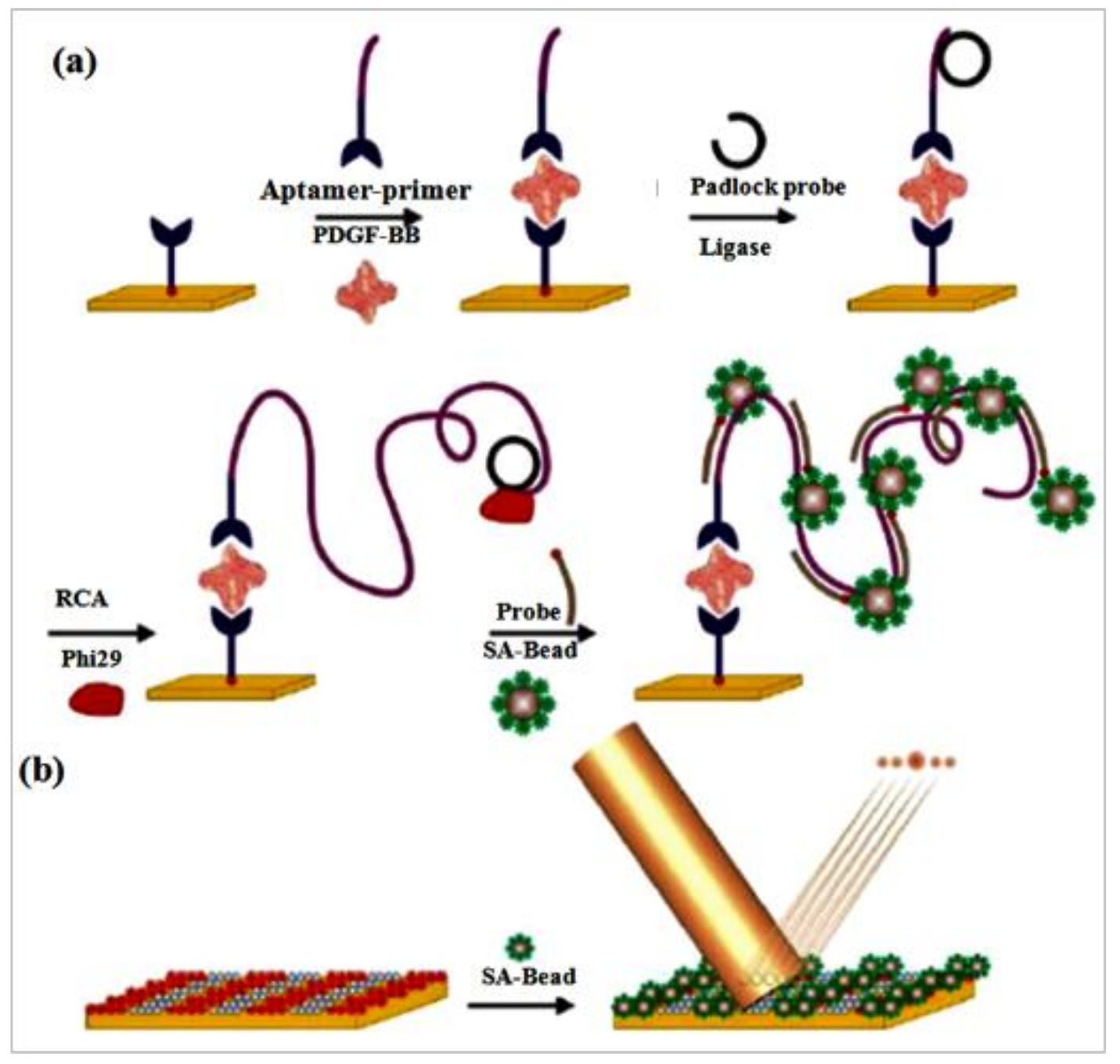
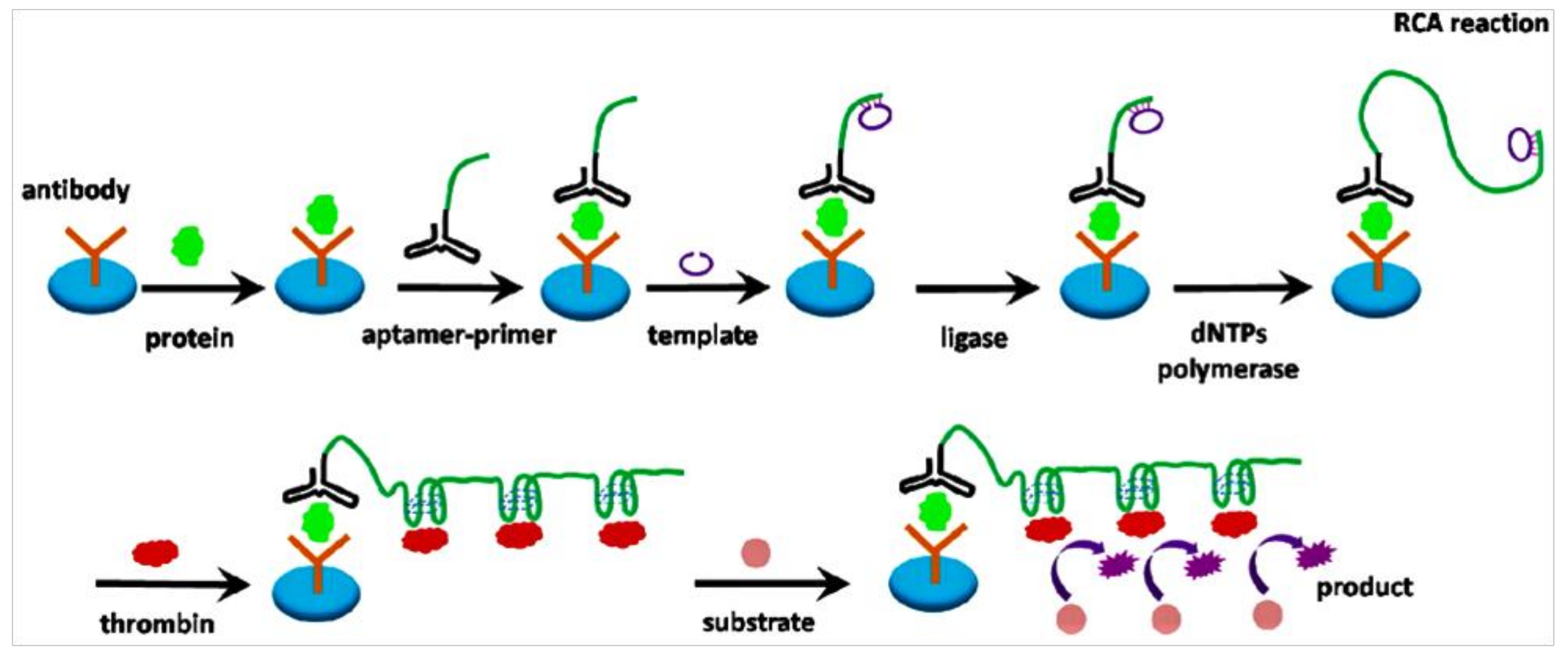
8. Protein Biosensor Based on RCA
9. Other Novel Biosensors Based on RCA
10. Outlook
Acknowledgments
Conflicts of Interest
References
- Nilsson, M.; Gullberg, M.; Dahl, F.; Szuhai, K.; Raap, A.K. Real-time monitoring of rolling-circle amplification using a modified molecular beacon design. Nucleic Acids Res. 2002, 30, e66. [Google Scholar] [CrossRef]
- Liu, X.; Xue, Q.; Ding, Y.; Zhu, J.; Wang, L.; Jiang, W. Cascade Signal Amplification Strategy for Sensitive and Label-free DNA Detection Based on Exo III-catalyzed Recycling Coupled with Rolling Circle Amplification. Analyst 2014, 139, 2884–2889. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zeng, Y.; Ji, X.; Li, X.; Ren, R. Cascade signal amplification for sensitive detection of cancer cell based on self-assembly of DNA scaffold and rolling circle amplification. Sens. Actuators B Chem. 2012, 171–172, 361–366. [Google Scholar] [CrossRef]
- Khan, S.A. Plasmid rolling-circle replication: Highlights of two decades of research. Plasmid 2005, 53, 126–136. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Cui, C.H.; Bose, S.; Guo, D.; Shen, C.; Wong, W.P.; Halvorsen, K.; Farokhzad, O.C.; Teo, G.S.L.; Phillips, J.A.; et al. Bioinspired multivalent DNA network for capture and release of cells. Proc. Natl. Acad. Sci. USA 2012, 109, 19626–19631. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Ali, M.M.; Eckert, M.A.; Kang, D.K.; Chen, Y.Y.; Sender, L.S.; Fruman, D.A.; Zhao, W. A polyvalent aptamer system for targeted drug delivery. Biomaterials 2013, 34, 9728–9735. [Google Scholar] [CrossRef] [PubMed]
- Ali, M.M.; Li, Y. Colorimetric Sensing by Using Allosteric-DNAzyme-Coupled Rolling Circle Amplification and a Peptide Nucleic Acid–Organic Dye Probe. Angew. Chem. 2009, 121, 3564–3567. [Google Scholar] [CrossRef]
- Cheglakov, Z.; Weizmann, Y.; Basnar, B.; Willner, I. Diagnosing viruses by the rolling circle amplified synthesis of DNAzymes. Org. Biomol. Chem. 2007, 5, 223–225. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Wang, C.; Xiong, Y.; Lu, H.; Ju, H.; Zhang, X. Highly sensitive and selective chemiluminescent imaging for DNA detection by ligation-mediated rolling circle amplified synthesis of DNAzyme. Biosens. Bioelectron. 2013, 41, 348–353. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.; Liu, Y.; Ali, M.M.; Kang, D.K.; Zhao, W.; Li, J. Colorimetric and ultrasensitive bioassay based on a dual-amplification system using aptamer and DNAzyme. Anal. Chem. 2012, 84, 4711–4717. [Google Scholar] [CrossRef] [PubMed]
- Dahl, F.; Banér, J.; Gullberg, M.; Mendel-Hartvig, M.; Landegren, U.; Nilsson, M. Circle-to-circle amplification for precise and sensitive DNA analysis. Proc. Natl. Acad. Sci. USA 2004, 101, 4548–4553. [Google Scholar] [CrossRef] [PubMed]
- Linck, L.; Reiß, E.; Bier, F.; Resch-Genger, U. Direct labeling rolling circle amplification as a straightforward signal amplification technique for biodetection formats. Anal. Methods 2012, 4, 1215–1220. [Google Scholar] [CrossRef]
- Zhao, W.; Gao, Y.; Kandadai, S.A.; Brook, M.A.; Li, Y. DNA Polymerization on Gold Nanoparticles through Rolling Circle Amplification: Towards Novel Scaffolds for Three-Dimensional Periodic Nanoassemblies. Angew. Chem. Int. Ed. 2006, 45, 2409–2413. [Google Scholar] [CrossRef] [PubMed]
- Ali, M.M.; Aguirre, S.D.; Xu, Y.; Filipe, C.D.M.; Pelton, R.; Li, Y. Detection of DNA using bioactive paper strips. Chem. Commun. 2009, 45, 6640–6642. [Google Scholar] [CrossRef] [PubMed]
- Berr, A.; Schubert, I. Interphase chromosome arrangement in Arabidopsis thaliana is similar in differentiated and meristematic tissues and shows a transient mirror symmetry after nuclear division. Genetics 2007, 176, 853–863. [Google Scholar] [CrossRef] [PubMed]
- Beyer, S.; Nickels, P.; Simmel, F.C. Periodic DNA nanotemplates synthesized by rolling circle amplification. Nano Lett. 2005, 5, 719–722. [Google Scholar] [CrossRef] [PubMed]
- Su, H.; Yuan, R.; Chai, Y.; Mao, L.; Zhuo, Y. Ferrocenemonocarboxylic–HRP@ Pt nanoparticles labeled RCA for multiple amplification of electro-immunosensing. Biosens. Bioelectron. 2011, 26, 4601–4604. [Google Scholar] [CrossRef] [PubMed]
- Thomas, D.; Lizardi, P.; Nardone, G.; Winndeen, E. Cascade rolling circle amplification, a homogeneous fluorescence detection system for DNA diagnostics. Clin. Chem. 1997, 11, 38. [Google Scholar]
- Smolina, I.; Lee, C.; Frank-Kamenetskii, M. Detection of low-copy-number genomic DNA sequences in individual bacterial cells by using peptide nucleic acid-assisted rolling-circle amplification and fluorescence in situ hybridization. Appl. Environ. Microbiol. 2007, 73, 2324–2328. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Xie, X.; Li, D.; Yang, Z.; Li, T.; Liu, X. Ultrasensitive Colorimetric DNA Detection using a Combination of Rolling Circle Amplification and Nicking Endonuclease-Assisted Nanoparticle Amplification (NEANA). Small 2012, 8, 1846–1850. [Google Scholar] [CrossRef] [PubMed]
- Johne, R.; Müller, H.; Rector, A.; Van Ranst, M.; Stevens, H. Rolling-circle amplification of viral DNA genomes using phi29 polymerase. Trends Microbiol. 2009, 17, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Schopf, E.; Fischer, O.N.; Chen, Y.; Tok, J.B.H. Sensitive and selective viral DNA detection assay via microbead-based rolling circle amplification. Bioorg. Med. Chem. Lett. 2008, 18, 5871–5874. [Google Scholar] [CrossRef] [PubMed]
- Schopf, E.; Chen, Y. Attomole DNA detection assay via rolling circle amplification and single molecule detection. Anal. Biochem. 2010, 397, 115–117. [Google Scholar] [CrossRef] [PubMed]
- Thomas, D.C.; Nardone, G.A.; Randall, S.K. Amplification of padlock probes for DNA diagnostics by cascade rolling circle amplification or the polymerase chain reaction. Arch. Pathol. Lab. Med. 1999, 123, 1170–1176. [Google Scholar] [PubMed]
- Christian, A.T.; Pattee, M.S.; Attix, C.M.; Reed, B.E.; Sorensen, K.J.; Tucker, J.D. Detection of DNA point mutations and mRNA expression levels by rolling circle amplification in individual cells. Proc. Natl. Acad. Sci. USA 2001, 98, 14238–14243. [Google Scholar] [CrossRef] [PubMed]
- Lagunavicius, A.; Merkiene, E.; Kiveryte, Z.; Savaneviciute, A.; Zimbaite-Ruskuliene, V.; Radzvilavicius, T.; Janulaitis, A. Novel application of Phi29 DNA polymerase: RNA detection and analysis in vitro and in situ by target RNA-primed RCA. RNA 2009, 15, 765–771. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Calciano, M.; Hamann, S.; Leamon, J.H.; Strugnell, T.; Christian, M.W.; Lizardi, P.M. In situ detection of messenger RNA using digoxigenin-labeled oligonucleotides and rolling circle amplification. Exp. Mol. Pathol. 2001, 70, 281–288. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Ma, X.; Li, M.; Zhou, D.; Xiao, P.; Lu, Z. Analysis of CpG island methylation using rolling circle amplification (RCA) product microarray. J. Biomed. Nanotechnol. 2011, 7, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Qi, X.; Bakht, S.; Devos, K.M.; Gale, M.D.; Osbourn, A. L-RCA (ligation-rolling circle amplification): A general method for genotyping of single nucleotide polymorphisms (SNPs). Nucleic Acids Res. 2001, 29, e116. [Google Scholar] [CrossRef] [PubMed]
- Pickering, J.; Bamford, A.; Godbole, V.; Briggs, J.; Scozzafava, G.; Roe, P.; Wheeler, C.; Ghouze, F.; Cuss, S. Integration of DNA ligation and rolling circle amplification for the homogeneous, end-point detection of single nucleotide polymorphisms. Nucleic Acids Res. 2002, 30, e60. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.Y.; Cheng, Y.Q.; Du, Q.; Zhang, H.X.; Li, Z.P. Integration of rolling circle amplification and cationic conjugated polymer for the homogeneous detection of single nucleotide polymorphisms. Chin. Sci. Bull. 2011, 56, 3247–3252. [Google Scholar] [CrossRef]
- Zhang, S.; Wu, Z.; Shen, G.; Yu, R. A label-free strategy for SNP detection with high fidelity and sensitivity based on ligation-rolling circle amplification and intercalating of methylene blue. Biosens. Bioelectron. 2009, 24, 3201–3207. [Google Scholar] [CrossRef] [PubMed]
- Deng, K.; Li, C.; Huang, H.; Li, X. Rolling circle amplification based on signal-enhanced electrochemical DNA sensor for ultrasensitive transcription factor detection. Sens. Actuators B Chem. 2017, 238, 1302–1308. [Google Scholar] [CrossRef]
- Meng, F.; Miao, P.; Wang, B.; Tang, Y.; Yin, J. Identification of glutathione by voltammetric analysis with rolling circle amplification. Anal. Chim. Acta 2016, 943, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Mao, X.; Chen, X.; Wang, Z.; Wang, K.; Zhu, X. The analysis of proteins and small molecules based on sterically tunable nucleic acid hyperbranched rolling circle amplification. Biosens. Bioelectron. 2017, 91, 136–142. [Google Scholar] [CrossRef] [PubMed]
- Ding, C.; Liu, H.; Wang, N.; Wang, Z. Cascade signal amplification strategy for the detection of cancer cells by rolling circle amplification and nanoparticles tagging. Chem. Commun. 2012, 48, 5019–5021. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, F.; Kenzaka, T.; Yamaguchi, N.; Tani, K.; Nasu, M. Visualization and enumeration of bacteria carrying a specific gene sequence by in situ rolling circle amplification. Appl. Environ. Microbiol. 2005, 71, 7933–7940. [Google Scholar] [CrossRef] [PubMed]
- Ali, M.M.; Li, F.; Zhang, Z.; Zhang, K.; Kang, D.K.; Ankrum, J.A.; Chris Le, X.; Zhao, W. Rolling circle amplification: A versatile tool for chemical biology, materials science and medicine. Chem. Soc. Rev. 2014, 43, 3324–3341. [Google Scholar] [CrossRef] [PubMed]
- Doherty, A.J.; Dafforn, T.R. Nick Recognition by DNA Ligases. J. Mol. Biol. 2000, 296, 43–56. [Google Scholar] [CrossRef] [PubMed]
- Doherty, A.J.; Suh, S.W. Structural and mechanistic conservation in DNA ligases. Nucleic Acids Res. 2000, 28, 4051–4058. [Google Scholar] [CrossRef] [PubMed]
- Lehman, I. DNA Iigase: Structure mechanism, function. Science 1974, 186, 790–797. [Google Scholar] [CrossRef] [PubMed]
- Blundred, R.M.; Stewart, G.S. DNA double-strand break repair, immunodeficiency and the RIDDLE syndrome. Expert Rev. Clin. Immunol. 2011, 7, 169–185. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.-C.; Ding, S.; Chang, C.H.; Kuo, S.H.; Chan, S.T.; Hsu, G.C.; Hsu, H.M.; Hou, M.F.; Jung, L.Y.; Cheng, C.W.; et al. Genetic susceptibility to the development and progression of breast cancer associated with polymorphism of cell cycle and ubiquitin ligase genes. Carcinogenesis 2009, 30, 1562–1570. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Siribal, S.; Weinfeld, M.; Karimi-Busheri, F.; Mark Glover, J.N.; Bernstein, N.K.; Aceytuno, D.; Chavalitshewinkoon-Petmitr, P. Molecular characterization of Plasmodium falciparum putative polynucleotide kinase/phosphatase. Mol. Biochem. Parasitol. 2011, 180, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Allinson, S.L. DNA end-processing enzyme polynucleotide kinase as a potential target in the treatment of cancer. Future Oncol. 2010, 6, 1031–1042. [Google Scholar] [CrossRef] [PubMed]
- Freschauf, G.K.; Karimi-Busheri, F.; Ulaczyk-Lesanko, A.; Mereniuk, T.R.; Ahrens, A.; Koshy, J.M.; Rasouli-Nia, A.; Pasarj, P.; Holmes, C.F.B.; Rininsland, F.; et al. Identification of a small molecule inhibitor of the human DNA repair enzyme polynucleotide kinase/phosphatase. Cancer Res. 2009, 69, 7739–7746. [Google Scholar] [CrossRef] [PubMed]
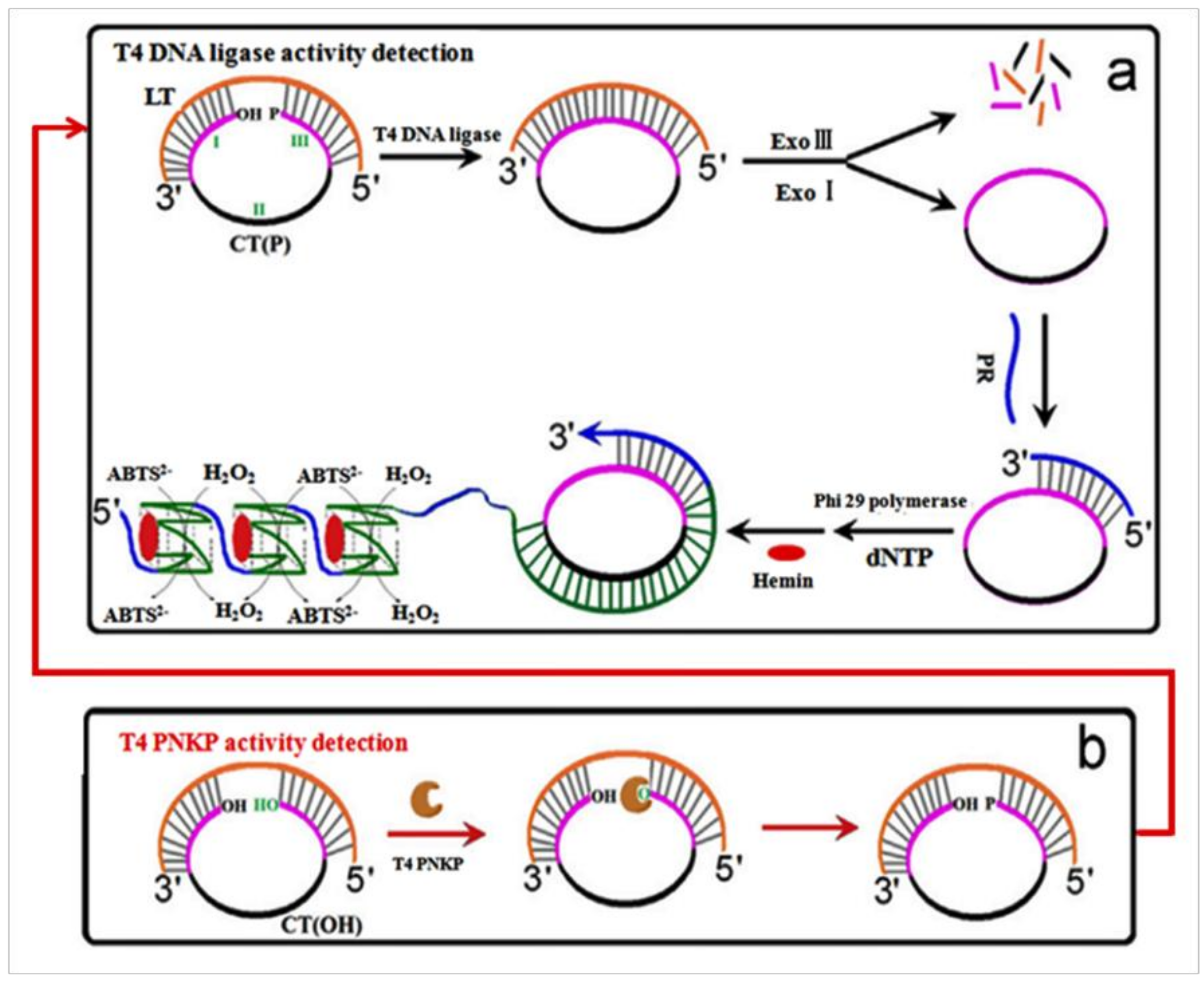
- Jiang, H.-X.; Kong, D.-M.; Shen, H.-X. Amplified detection of DNA ligase and polynucleotide kinase/phosphatase on the basis of enrichment of catalytic G-quadruplex DNAzyme by rolling circle amplification. Biosens. Bioelectron. 2014, 55, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Bodrossy, L.; Sessitsch, A. Oligonucleotide microarrays in microbial diagnostics. Curr. Opin. Microbiol. 2004, 7, 245–254. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, R.; Jobs, M.; Ekstrand, C.; Ullberg, M.; Herrmann, B.; Landegren, U.; Nilsson, M.; Blomberg, J. Multiplex and quantifiable detection of nucleic acid from pathogenic fungi using padlock probes, generic real time PCR and specific suspension array readout. J. Microbiol. Methods 2009, 78, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Atkins, S.D.; Clark, I.M. Fungal molecular diagnostics: A mini review. J. Appl. Genet. 2004, 45, 3–15. [Google Scholar] [PubMed]
- Cho, S.-N.; Brennan, P.J. Tuberculosis: Diagnostics. Tuberculosis 2007, 87, S14–S17. [Google Scholar] [CrossRef] [PubMed]
- López, M.M.; Bertolini, E.; Olmos, A.; Caruso, P.; Gorris, M.T.; Llop, P.; Penyalver, R.; Cambra, M. Innovative tools for detection of plant pathogenic viruses and bacteria. Int. Microbiol. 2003, 6, 233–243. [Google Scholar] [CrossRef] [PubMed]
- Manso, J.; Mena, M.L.; Yanez-Sedeno, P.; Pingarrón, J.M. Bienzyme amperometric biosensor using gold nanoparticle-modified electrodes for the determination of inulin in foods. Anal. Biochem. 2008, 375, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Homola, J. Surface plasmon resonance sensors for detection of chemical and biological species. Chem. Rev. 2008, 108, 462–493. [Google Scholar] [CrossRef] [PubMed]
- Cosnier, S.; Mailley, P. Recent advances in DNA sensors. Analyst 2008, 133, 984–991. [Google Scholar] [CrossRef] [PubMed]
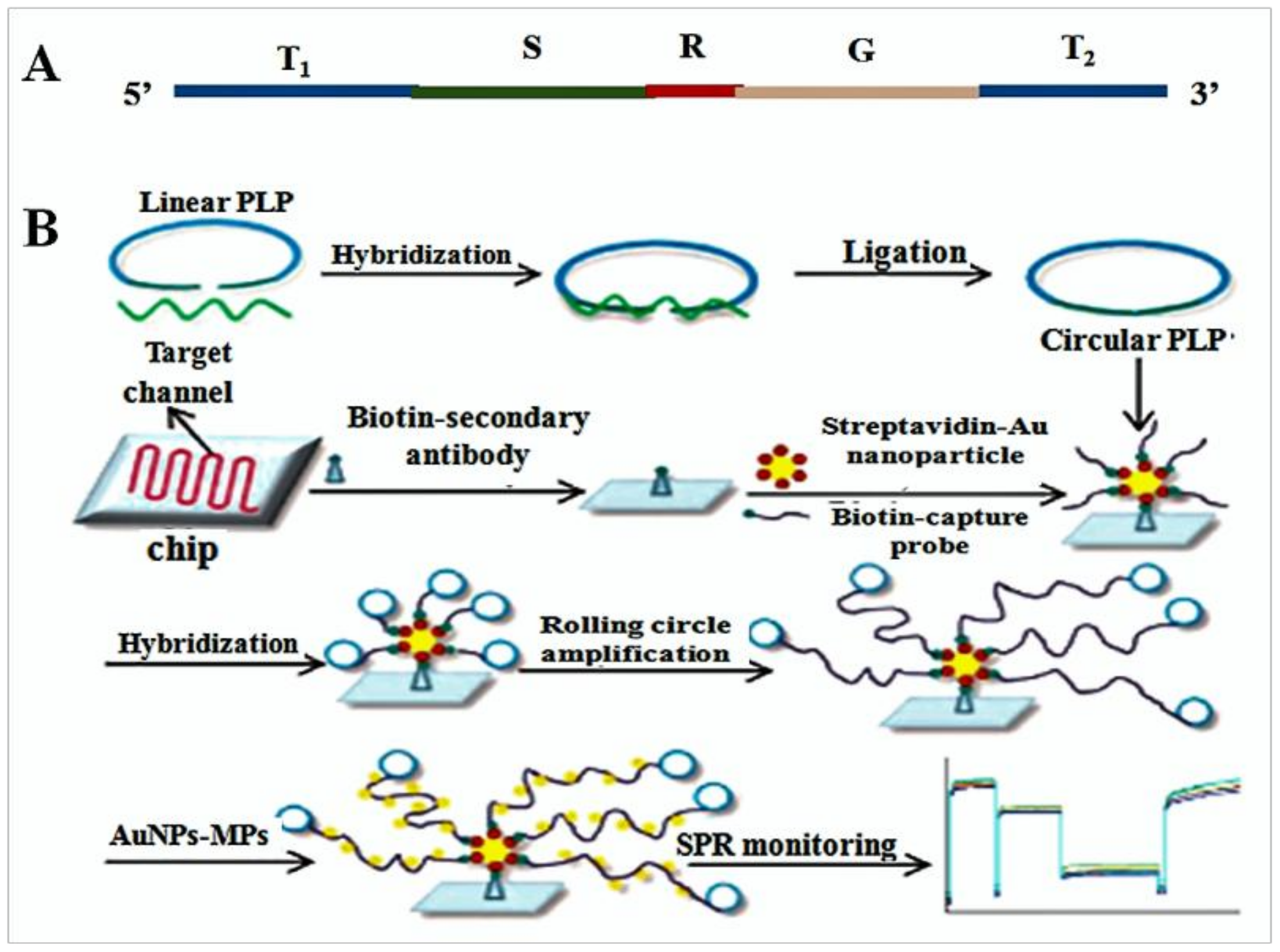
- Shi, D.; Huang, J.; Chuai, Z.; Chen, D.; Zhu, X.; Wang, H.; Peng, J.; Wu, H.; Huang, Q.; Fu, W. Isothermal and rapid detection of pathogenic microorganisms using a nano-rolling circle amplification-surface plasmon resonance biosensor. Biosens. Bioelectron. 2014, 62, 280–287. [Google Scholar] [CrossRef] [PubMed]
- Daniel, D.; Lalitha, R. Tumor markers—A bird’s eye view. J. Oral Maxillofac. Surg. Med. Pathol. 2016, 28, 475–480. [Google Scholar] [CrossRef]
- De Rancher, M.-A.R.; Oudart, J.B.; Maquart, F.X.; Monboisse, J.C.; Ramont, L. Evaluation of Lumipulse® G1200 for the measurement of six tumor markers: Comparison with AIA® 2000. Clin. Biochem. 2016, 49, 1302–1306. [Google Scholar] [CrossRef] [PubMed]
- Chung, S.; Moon, J.M.; Ban, C.; Shim, Y.B. A Simple and Fast SELEX Using an Alternating Current Potential Modulated Microfluidic Channel and an Evaluation of Sensing Ability of Aptamers. In Meeting Abstracts; The Electrochemical Society: Pennington, NJ, USA, 2016. [Google Scholar]
- Mu, Q.; Annapragada, A.; Srivastava, M.; Thiviyanathan, V.; Li, X.; Gorenstein, D.; Annapragada, A.; Vigneswaran, N. Conjugate-SELEX, a novel screening method, identifies aptamers that deliver payload to the cytosol of target cells. Cancer Res. 2016, 76, 3913. [Google Scholar] [CrossRef]
- Sanghavi, B.J.; Moore, J.A.; Chávez, J.L.; Hagen, J.A.; Kelley-Loughnane, N.; Chou, C.F.; Swami, N.S. Aptamer-functionalized nanoparticles for surface immobilization-free electrochemical detection of cortisol in a microfluidic device. Biosens. Bioelectron. 2016, 78, 244–252. [Google Scholar] [CrossRef] [PubMed]
- Théodorou, I.; Quang, N.N.; Gombert, K.; Thézé, B.; Lelandais, B.; Ducongé, F. In Vitro and In Vivo Imaging of Fluorescent Aptamers. In Nucleic Acid Aptamers: Selection, Characterization, and Application; Springer: New York, NY, USA, 2016; pp. 135–150. [Google Scholar]
- Wu, L.; Qi, P.; Fu, X.; Liu, H.; Li, J.; Wang, Q.; Fan, H. A novel electrochemical PCB77-binding DNA aptamer biosensor for selective detection of PCB77. J. Electroanal. Chem. 2016, 771, 45–49. [Google Scholar] [CrossRef]
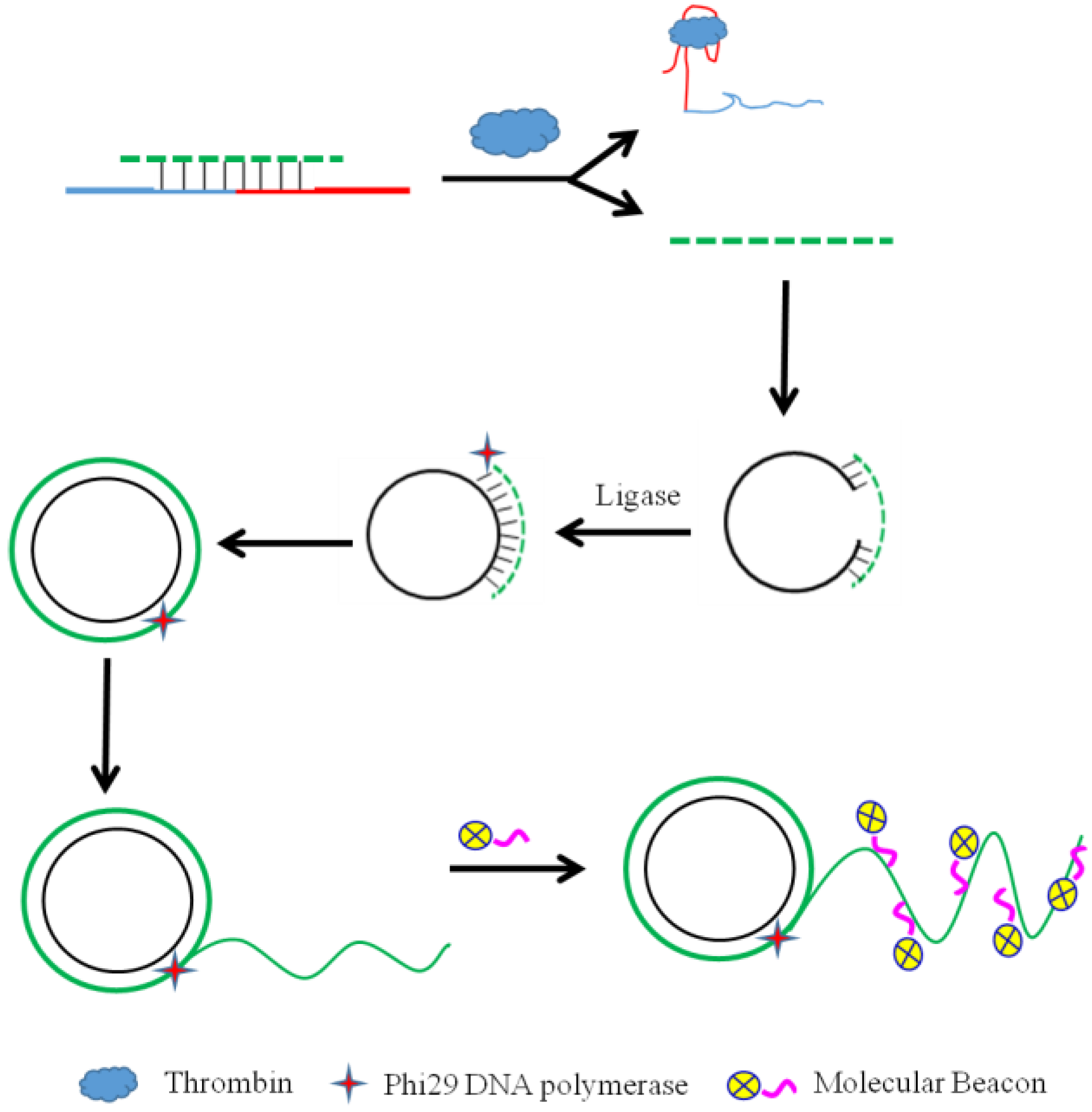
- Zhang, S.-B.; Zheng, L.Y.; Hu, X.; Shen, G.Y.; Liu, X.W.; Shen, G.L.; Yu, R.Q. Highly Sensitive Fluorescent Aptasensor for Thrombin Detection Based on Competition Triggered Rolling Circle Amplification. Chin. J. Anal. Chem. 2015, 43, 1688–1694. [Google Scholar] [CrossRef]
- Kerman, K.; Ozkan, D.; Kara, P.; Meric, B.; Gooding, J.J.; Ozsoz, M. Voltammetric determination of DNA hybridization using methylene blue and self-assembled alkanethiol monolayer on gold electrodes. Anal. Chim. Acta 2002, 462, 39–47. [Google Scholar] [CrossRef]
- Wang, Q.; Zheng, H.; Gao, X.; Lin, Z.; Chen, G. A label-free ultrasensitive electrochemical aptameric recognition system for protein assay based on hyperbranched rolling circle amplification. Chem. Commun. 2013, 49, 11418–11420. [Google Scholar] [CrossRef] [PubMed]
- Alves-Balvedi, R.P.; Caetano, L.P.; Madurro, J.M.; Brito-Madurro, A.G. Use of 3,3′,5,5′ tetramethylbenzidine as new electrochemical indicator of DNA hybridization and its application in genossensor. Biosens. Bioelectron. 2016, 85, 226–231. [Google Scholar] [CrossRef] [PubMed]
- Berg, H.; Horn, G.; Luthardt, U.; Ihn, W. Interaction of anthracycline antibiotics with biopolymers: Part V. Polarographic behavior and complexes with DNA. Bioelectrochem. Bioenerget. 1981, 8, 537–553. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, K.; Ma, H. Electrochemical DNA biosensor based on silver nanoparticles/poly (3-(3-pyridyl) acrylic acid)/carbon nanotubes modified electrode. Anal. Biochem. 2009, 387, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Liu, B.; Zhao, Z.; Ma, C.; Luo, P.; Liu, C.; Xie, G. Ultrasensitive electrochemical immunosensor for HE4 based on rolling circle amplification. Biosens. Bioelectron. 2012, 33, 216–221. [Google Scholar] [CrossRef] [PubMed]
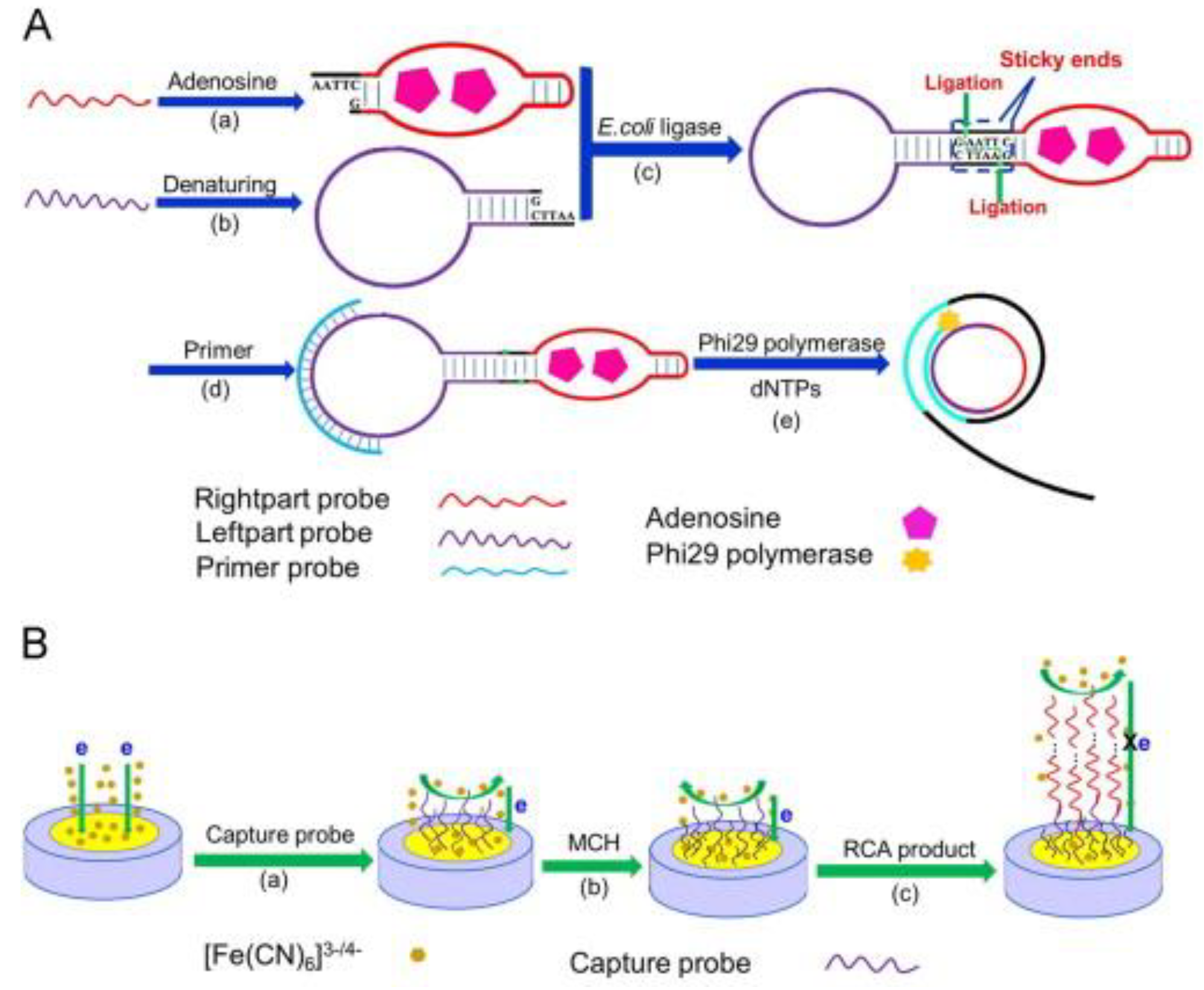
- Yi, X.; Li, L.; Peng, Y.; Guo, L. A universal electrochemical sensing system for small biomolecules using target-mediated sticky ends-based ligation-rolling circle amplification. Biosens. Bioelectron. 2014, 57, 103–109. [Google Scholar] [CrossRef] [PubMed]
- Latchman, D.S. Transcription factors: An overview. Int. J. Biochem. Cell Biol. 1997, 29, 1305–1312. [Google Scholar] [CrossRef]
- Rosenbauer, F.; Tenen, D.G. Transcription factors in myeloid development: Balancing differentiation with transformation. Nat. Rev. Immunol. 2007, 7, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Engelkamp, D.; van Heyningen, V. Transcription factors in disease. Curr. Opin. Genet. Dev. 1996, 6, 334–342. [Google Scholar] [CrossRef]
- Zhang, Y.; Ma, F.; Tang, B.; Zhang, C. Recent advances in transcription factor assays in vitro. Chem. Commun. 2016, 52, 4739–4748. [Google Scholar] [CrossRef] [PubMed]
- Garner, M.M.; Revzin, A. A gel electrophoresis method for quantifying the binding of proteins to specific DNA regions: Application to components of the Escherichia coli lactose operon regulatory system. Nucleic Acids Res. 1981, 9, 3047–3060. [Google Scholar] [CrossRef] [PubMed]
- Galas, D.J.; Schmitz, A. DNAase footprinting a simple method for the detection of protein-DNA binding specificity. Nucleic Acids Res. 1978, 5, 3157–3170. [Google Scholar] [CrossRef] [PubMed]
- Burnette, W.N. “Western blotting”: Electrophoretic transfer of proteins from sodium dodecyl sulfate-polyacrylamide gels to unmodified nitrocellulose and radiographic detection with antibody and radioiodinated protein A. Anal. Biochem. 1981, 112, 195–203. [Google Scholar] [CrossRef]
- Li, C.; Qiu, X.; Hou, Z.; Deng, K. A dumbell probe-mediated rolling circle amplification strategy for highly sensitive transcription factor detection. Biosens. Bioelectron. 2015, 64, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Ridzon, D.; Wong, L.; Chen, C. Characterization of microRNA expression profiles in normal human tissues. BMC Genom. 2007, 8. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef] [PubMed]
- Heneghan, H.M.; Miller, N.; Lowery, A.J.; Sweeney, K.J.; Newell, J.; Kerin, M.J. Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Ann. Surg. 2010, 251, 499–505. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Feng, Y.; Coukos, G.; Zhang, L. Therapeutic microRNA strategies in human cancer. AAPS J. 2009, 11, 747–757. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Qin, W.; Atasoy, U.; Sauter, E.R. Circulating microRNAs in breast cancer and healthy subjects. BMC Res. Notes 2009, 2. [Google Scholar] [CrossRef] [PubMed]
- Stenvang, J.; Silahtaroglu, A.N.; Lindow, M.; Elmen, J.; Kauppinen, S. The utility of LNA in microRNA-based cancer diagnostics and therapeutics. In Seminars in Cancer Biology; Elsevier: Amsterdam, The Netherlands, 2008. [Google Scholar]
- Cho, W.C. MicroRNAs: Potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int. J. Biochem. Cell Biol. 2010, 42, 1273–1281. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Tan, S.; Kooger, R.; Zhang, C.; Zhang, Y. MicroRNAs as novel biological targets for detection and regulation. Chem. Soc. Rev. 2014, 43, 506–517. [Google Scholar] [CrossRef] [PubMed]
- Volinia, S.; Calin, G.A.; Liu, C.G.; Ambs, S.; Cimmino, A.; Petrocca, F.; Visone, R.; Iorio, M.; Roldo, C.; Ferracin, M.; et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. USA 2006, 103, 2257–2261. [Google Scholar] [CrossRef] [PubMed]
- Válóczi, A.; Hornyik, C.; Varga, N.; Burgyán, J.; Kauppinen, S.; Havelda, Z. Sensitive and specific detection of microRNAs by northern blot analysis using LNA-modified oligonucleotide probes. Nucleic Acids Res. 2004, 32, e175. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Real-time quantification of microRNAs by stem–loop RT–PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Ruan, K. MicroRNA detection by microarray. Anal. Bioanal. Chem. 2009, 394, 1117–1124. [Google Scholar] [CrossRef] [PubMed]
- Bi, S.; Li, L.; Cui, Y. Exonuclease-assisted cascaded recycling amplification for label-free detection of DNA. Chem. Commun. 2012, 48, 1018–1020. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Fu, Z.; Li, B.; Zhou, Y.; Yin, H.; Ai, S. One-step, ultrasensitive, and electrochemical assay of microRNAs based on T7 exonuclease assisted cyclic enzymatic amplification. Anal. Chem. 2014, 86, 5606–5610. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.Q.; Zhang, M.; Yin, B.C.; Ye, B.C. Attomolar ultrasensitive microRNA detection by DNA-scaffolded silver-nanocluster probe based on isothermal amplification. Anal. Chem. 2012, 84, 5165–5169. [Google Scholar] [CrossRef] [PubMed]
- Bi, S.; Cui, Y.; Dong, Y.; Zhang, N. Target-induced self-assembly of DNA nanomachine on magnetic particle for multi-amplified biosensing of nucleic acid, protein, and cancer cell. Biosens. Bioelectron. 2014, 53, 207–213. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Li, L.; Duan, L.; Wang, X.; Xie, Y.; Tong, L.; Wang, Q.; Tang, B. High specific and ultrasensitive isothermal detection of microRNA by padlock probe-based exponential rolling circle amplification. Anal. Chem. 2013, 85, 7941–7947. [Google Scholar] [CrossRef] [PubMed]
- Deng, R.; Tang, L.; Tian, Q.; Wang, Y.; Lin, L.; Li, J. Toehold-initiated Rolling Circle Amplification for Visualizing Individual MicroRNAs In Situ in Single Cells. Angew. Chem. Int. Ed. 2014, 53, 2389–2393. [Google Scholar] [CrossRef] [PubMed]
- JamesáYang, C. A T7 exonuclease-assisted cyclic enzymatic amplification method coupled with rolling circle amplification: A dual-amplification strategy for sensitive and selective microRNA detection. Chem. Commun. 2014, 50, 1576–1578. [Google Scholar]
- Li, D.; Cheng, W.; Yan, Y.; Zhang, Y.; Yin, Y.; Ju, H.; Ding, S. A colorimetric biosensor for detection of attomolar microRNA with a functional nucleic acid-based amplification machine. Talanta 2016, 146, 470–476. [Google Scholar] [CrossRef] [PubMed]
- Gao, X. The Progress Study of Gastric Cancer and Tumor Associated Protein. Asian Case Rep. Oncol. 2016, 5, 61–71. [Google Scholar] [CrossRef]
- Olmsted, J. Affinity purification of antibodies from diazotized paper blots of heterogeneous protein samples. J. Biol. Chem. 1981, 256, 11955–11957. [Google Scholar] [PubMed]
- Stavitsky, A.B.; Jarchow, C. Micromethods for the study of proteins and antibodies I. Procedure and general applications of hemagglutination and hemagglutination-inhibition reactions with tannic acid and protein-treated red blood cells. J. Immunol. 1954, 72, 360–367. [Google Scholar] [PubMed]
- Savran, C.A.; Knudsen, S.M.; Ellington, A.D.; Manalis, S.R. Micromechanical detection of proteins using aptamer-based receptor molecules. Anal. Chem. 2004, 76, 3194–3198. [Google Scholar] [CrossRef] [PubMed]
- Eissa, S.; Ng, A.; Siaj, M.; Zourob, M. Selection, Characterization, and Application of High Affinity Microcystin-Targeting Aptamers in a Graphene-Based Biosensing Platform. In Meeting Abstracts; The Electrochemical Society: Pennington, NJ, USA, 2015. [Google Scholar]
- Chen, I.H.; Horikawa, S.; Du, S.; Liu, Y.; Wikle, H.C.; Barbaree, J.M.; Chin, B.A. Thermal Stability of Phage Peptide Probes vs. Aptamer for Salmonella Detection on Magnetoelastic Biosensors Platform. ECS Trans. 2016, 75, 165–173. [Google Scholar] [CrossRef]
- Acharya, G.; Chang, C.L.; Doorneweerd, D.D.; Vlashi, E.; Henne, W.A.; Hartmann, L.C.; Low, P.S.; Savran, C.A. Immunomagnetic diffractometry for detection of diagnostic serum markers. J. Am. Chem. Soc. 2007, 129, 15824–15829. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Icoz, K.; Roberts, A.; Ellington, A.D.; Savran, C.A. Diffractometric detection of proteins using microbead-based rolling circle amplification. Anal. Chem. 2010, 82, 197–202. [Google Scholar] [CrossRef] [PubMed]
- Noskovicova, N.; Petřek, M.; Eickelberg, O.; Heinzelmann, K. Platelet-derived growth factor signaling in the lung. From lung development and disease to clinical studies. Am. J. Respir. Cell Mol. Biol. 2015, 52, 263–284. [Google Scholar] [CrossRef] [PubMed]
- Kuai, J.; Mosyak, L.; Brooks, J.; Cain, M.; Carven, G.J.; Ogawa, S.; Ishino, T.; Tam, M.; Lavallie, E.R.; Yang, Z.; et al. Characterization of binding mode of action of a blocking anti-platelet-derived growth factor (PDGF)-B monoclonal antibody, MOR8457, reveals conformational flexibility and avidity needed for PDGF-BB to bind PDGF receptor-β. Biochemistry 2015, 54, 1918–1929. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Hao, L.; Zhao, Q. An aptamer assay using rolling circle amplification coupled with thrombin catalysis for protein detection. Anal. Bioanal. Chem. 2016, 408, 4715–4722. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Liu, L.; Zhang, L.; Guo, Q.; Cui, Y.; Yang, M. Sensitive immunosensing of the carcinoembryonic antigen utilizing aptamer-based in-situ formation of a redox-active heteropolyacid and rolling circle amplification. Microchim. Acta 2017. [Google Scholar] [CrossRef]
- Risch, N.; Merikangas, K. The future of genetic studies of complex human diseases. Science 1996, 273, 1516–1517. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Orner, B.P. Induction of pluripotency in fibroblasts through the expression of only four nuclear proteins. ACS Chem. Biol. 2006, 1, 557–558. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, J.; Chua, L.S.; Lynn, D.M. Multilayered thin films that sustain the release of functional DNA under physiological conditions. Langmuir 2004, 20, 8015–8021. [Google Scholar] [CrossRef] [PubMed]
- Miao, W.; Bard, A.J. Electrogenerated chemiluminescence. 77. DNA hybridization detection at high amplification with [Ru(bpy)3]2+-containing microspheres. Anal. Chem. 2004, 76, 5379–5386. [Google Scholar] [CrossRef] [PubMed]
- Ding, C.; Wang, N.; Zhang, J.; Wang, Z. Rolling circle amplification combined with nanoparticle aggregates for highly sensitive identification of DNA and cancercells. Biosens. Bioelectron. 2013, 42, 486–491. [Google Scholar] [CrossRef] [PubMed]
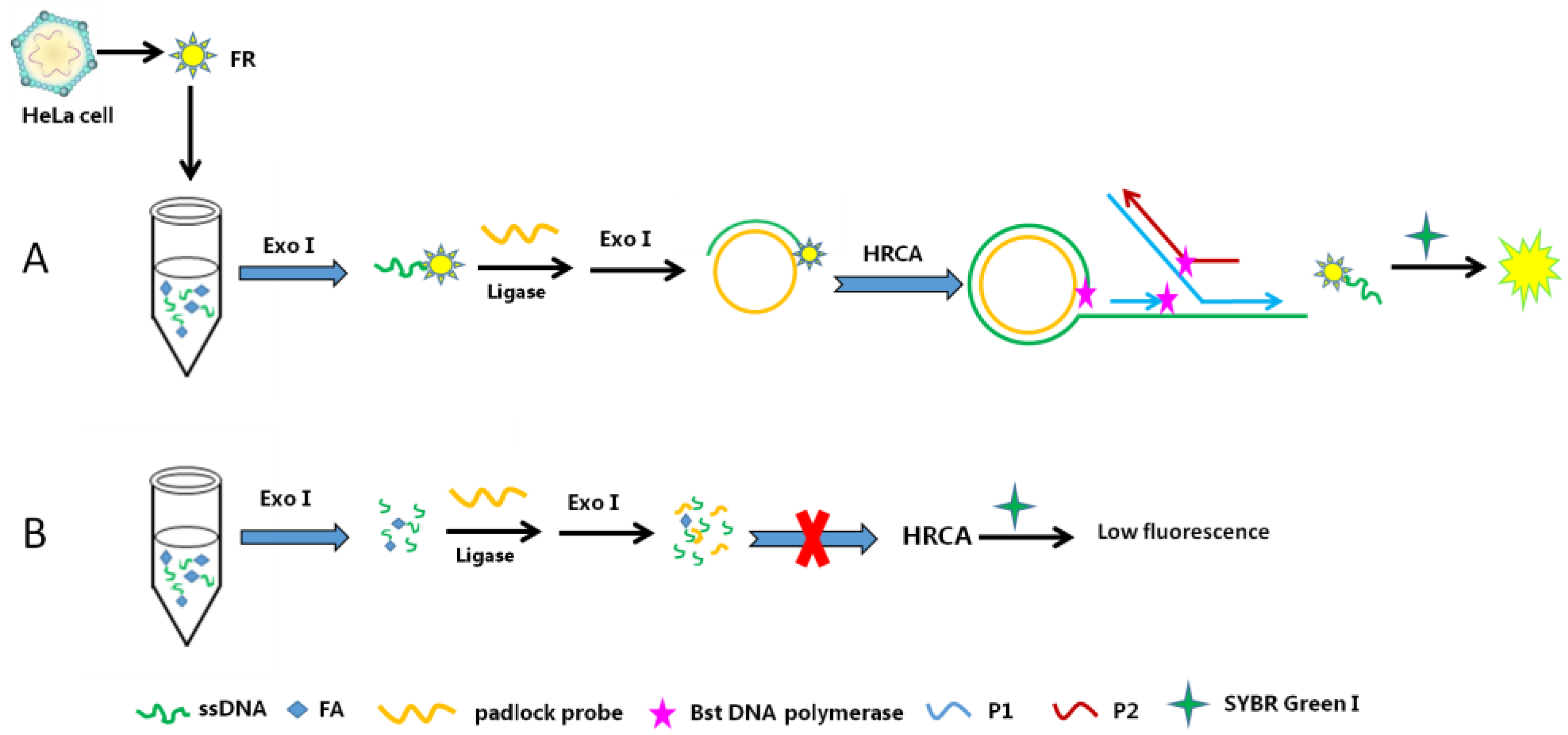
- Akbar, S.; Anwar, A.; Kanwal, Q. Electrochemical determination of folic acid: A short review. Anal. Biochem. 2016, 510, 98–105. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Ma, Y.; Zhang, Y.; Lin, Z. Fluorescence biosensor for folate receptors in cancer cells based on terminal protection and hyperbranched rolling circle amplification. Anal. Methods 2016, 8, 6231–6235. [Google Scholar] [CrossRef]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gu, L.; Yan, W.; Liu, L.; Wang, S.; Zhang, X.; Lyu, M. Research Progress on Rolling Circle Amplification (RCA)-Based Biomedical Sensing. Pharmaceuticals 2018, 11, 35. https://doi.org/10.3390/ph11020035
Gu L, Yan W, Liu L, Wang S, Zhang X, Lyu M. Research Progress on Rolling Circle Amplification (RCA)-Based Biomedical Sensing. Pharmaceuticals. 2018; 11(2):35. https://doi.org/10.3390/ph11020035
Chicago/Turabian StyleGu, Lide, Wanli Yan, Le Liu, Shujun Wang, Xu Zhang, and Mingsheng Lyu. 2018. "Research Progress on Rolling Circle Amplification (RCA)-Based Biomedical Sensing" Pharmaceuticals 11, no. 2: 35. https://doi.org/10.3390/ph11020035
APA StyleGu, L., Yan, W., Liu, L., Wang, S., Zhang, X., & Lyu, M. (2018). Research Progress on Rolling Circle Amplification (RCA)-Based Biomedical Sensing. Pharmaceuticals, 11(2), 35. https://doi.org/10.3390/ph11020035




