Complexes of Oligoribonucleotides with D-Mannitol Inhibit Hemagglutinin–Glycan Interaction and Suppress Influenza A Virus H1N1 (A/FM/1/47) Infectivity In Vitro
Abstract
:1. Introduction
2. Results
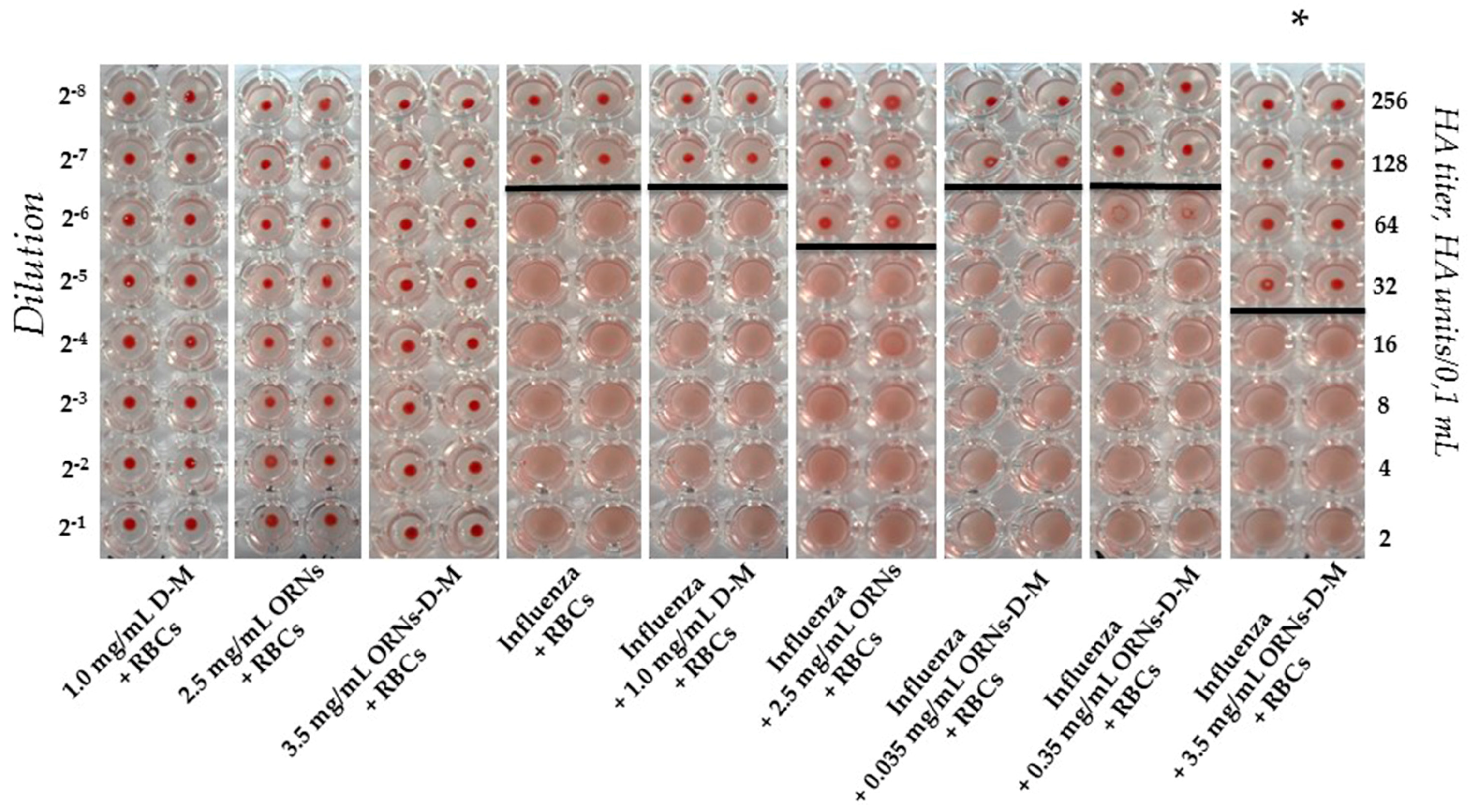
2.1. Interaction of the Influenza Virus A H1N1 HA with Glycan and Inhibition of This Interaction by the ORNs-d-M
2.2. Decrease of the Influenza Virus Infectivity after Incubation with the ORNs-d-M
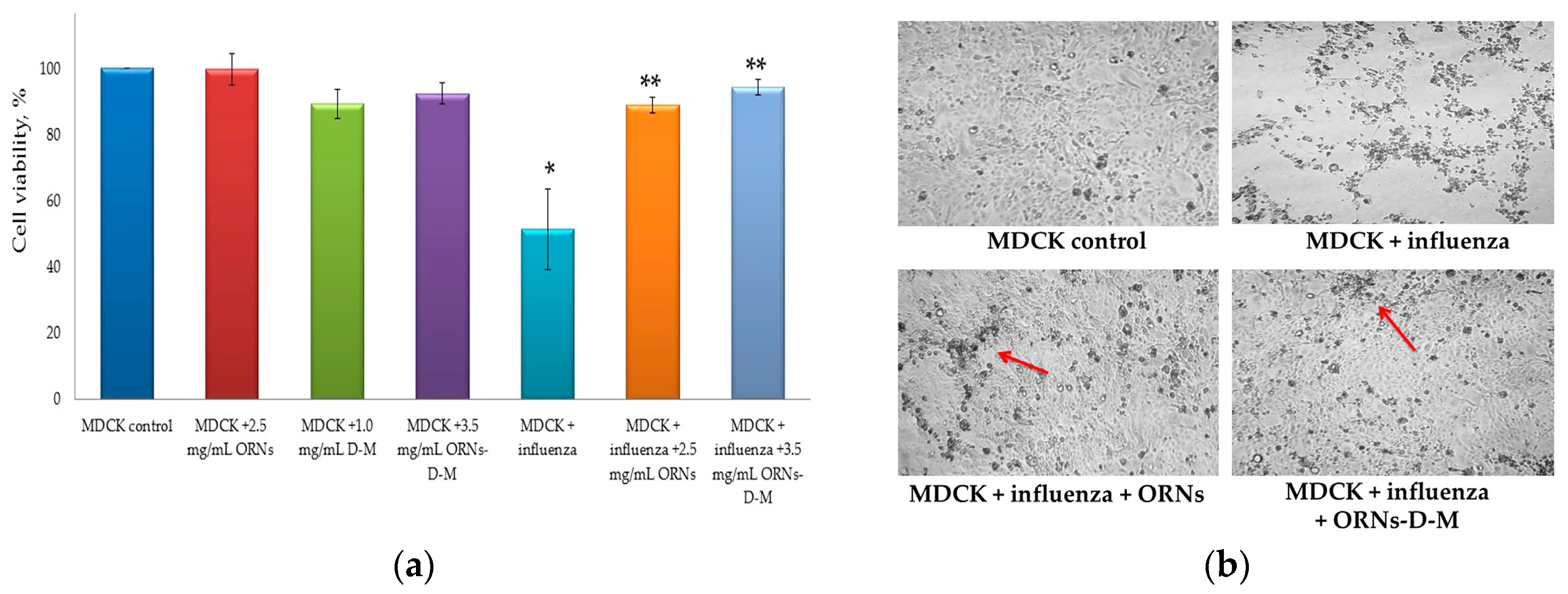
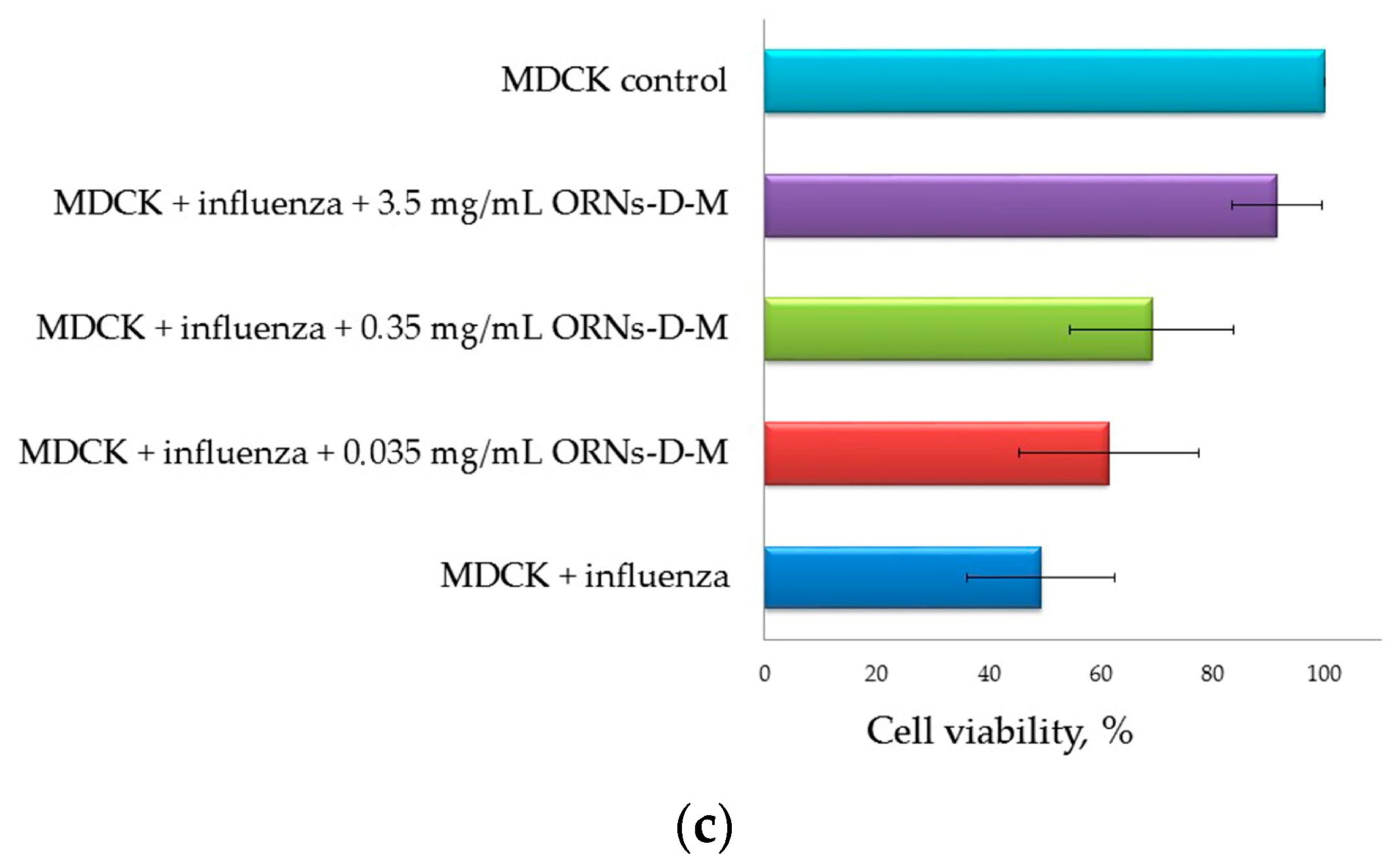
2.3. Direct Virucidal Action of the ORNs-d-M on the Influenza A Virus H1N1 (A/FM/1/47)
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Methods
4.2.1. Agglutination Assay
4.2.2. Infectious Titer of Influenza Virus (TCID50 Assay)
4.2.3. Direct Virucidal Action of the ORNs-d-M (MTT and CPE Reduction Assays)
4.2.4. Statistical Analysis
Author Contributions
Conflicts of Interest
References
- Garten, R.J.; Davis, C.T.; Russell, C.A.; Shu, B.; Lindstrom, S.; Balish, A. Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science 2009, 325, 197–201. [Google Scholar] [CrossRef] [PubMed]
- Gopinath, S.C.B.; Kumar, P.K.R. Aptamers that bind to the hemagglutinin of the recent pandemic influenza virus H1N1 and efficiently inhibit agglutination. Acta Biomater. 2013, 9, 8932–8941. [Google Scholar] [CrossRef] [PubMed]
- Brooks, G.F.; Butel, J.S.; Morse, S.A. Jawetz, Melnick, & Adelberg’s Medical Microbiology, 22nd ed.; The McGraw-Hill Companies: New York, NY, USA, 2004; PMID:3074881. [Google Scholar]
- Okazawa, A.; Maeda, H.; Fukusaki, E.; Katakura, Y.; Kobayashi, A. In vitro selection of hematoporphyrin binding DNA aptamers. Bioorg. Med. Chem. Lett. 2000, 10, 2653–2656, PMID:11128644. [Google Scholar] [CrossRef]
- Tang, J.; Xie, J.; Shao, N.; Yan, Y. The DNA aptamers that specifically recognize ricin toxin are selected by two in vitro selection methods. Electrophoresis 2006, 27, 1303–1311. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Liu, X.; Zhang, Q.; Zhang, C.; Liu, Y.; Tu, K.; Tu, J. Selection of DNA aptamers that bind to four organophosphorus pesticides. Biotechnol. Lett. 2012, 34, 869–874. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Zhao, J.; Jiang, T.; Kwon, Y.M.; Lu, H.; Jiao, P.; Liao, M.; Li, Y. Selection and characterization of DNA aptamers for use in detection of avian influenza virus H5N1. J. Virol. Methods 2013, 189, 362–369. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Swiderski, P.; Li, H.; Zhang, J.; Neff, C.P.; Akkina, R.; Rossi, J.J. Selection, characterization and application of new RNA HIV gp 120 aptamers for facile delivery of Dicer substrate siRNAs into HIV infected cells. Nucleic Acids Res. 2009, 37, 3094–3109. [Google Scholar] [CrossRef] [PubMed]
- Feng, H.; Beck, J.; Nassal, M.; Hu, K.H. A SELEXscreened aptamer of human hepatitis B virus RNA encapsidation signal suppresses viral replication. PLoS ONE 2011, 6, e27862. [Google Scholar] [CrossRef] [PubMed]
- Levchenko, S.M.; Rebriev, A.V.; Tkachuk, V.V.; Dubey, L.V.; Dubey, I.Y.; Tkachuk, Z.Y. Studies on the interaction of oligoadenylates with proteins by MALDI-TOF mass spectrometry. Biopolim. Cell 2013, 29, 42–48. [Google Scholar] [CrossRef]
- Skorobogatov, O.Y.; Lozhko, D.N.; Zhukov, I.Y.; Kozlov, O.; Tkachuk, Z.Y. Study of dephosphorylated 2′-5′-linked oligoadenylates impact on apo-S100A1 protein conformation by heteronuclear NMR and circular dichroism. Biopolym. Cell 2014, 30, 279–285. [Google Scholar] [CrossRef]
- Tkachuk, Z. Multiantivirus Compound, Composition and Method for Treatment of Virus Diseases. U.S. Patent 20,120,232,129, 16 April 2013. [Google Scholar]
- Tkachuk, Z.Y.; Zarubayev, V.V.; Semernikova, L.I. The antivirus action of Nuclex on the viruses of influenza, parainfluenza and bird flu in vitro. Ukr. Med. Alm. 2011, 6, 209–212. [Google Scholar]
- Tkachuk, Z.Y.; Rybalko, S.L.; Zharkova, L.D.; Starostyla, D.B. Antiinfluenzal activity of drug Nuclex. Rep. Natl. Acad. Sci. Ukr. 2010, 9, 191–196. [Google Scholar]
- Killian, M.L. Hemagglutination assay for the avian influenza virus. Methods Mol. Biol. 2008, 436, 47–52. [Google Scholar] [CrossRef] [PubMed]
- Reed, L.J.; Muench, H. A simple method of estimating fifty percent endpoints. Am. J. Hyg. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Takizawa, T.; Matsukawa, S.; Higuchi, Y.; Nakamura, S.; Nakanishi, Y.; Fukuda, R. Induction of programmed cell death (apoptosis) by influenza virus infection in tissue culture cells. J. Gen. Virol. 1993, 74, 2347–2355. [Google Scholar] [CrossRef]
- Tran, A.T.; Cortens, J.P.; Du, Q.; Wilkins, J.A.; Coombs, K.M. Influenza virus induces apoptosis via BAD-mediated mitochondrial dysregulation. J. Virol. 2013, 87, 1049–1060. [Google Scholar] [CrossRef] [PubMed]
- Cillóniz, C.; Shinya, K.; Peng, X.; Korth, M.J.; Proll, S.C.; Aicher, L.D.; Carter, V.S.; Chang, J.H.; Kobasa, D.; Feldmann, F.; et al. Lethal influenza virus infection in macaques is associated with early dysregulation of inflammatory related genes. PLoS Pathog. 2009, 5, e1000604. [Google Scholar] [CrossRef]
- Salzberg, S. The contents of the syringe. Nature 2008, 454, 160–161. [Google Scholar] [CrossRef] [PubMed]
- Davlin, S.L. Influenza activity—United States, 2015–2016 season and composition of the 2016–2017 influenza vaccine. MMWR Morb. Mortal. Wkly. Rep. 2016, 65, 567–575. [Google Scholar] [CrossRef] [PubMed]
- Hensley, S.E. Challenges of selecting seasonal influenza vaccine strains for humans with diverse pre-exposure histories. Curr. Opin. Virol. 2014, 8, 85–89. [Google Scholar] [CrossRef] [PubMed]
- Houser, K.; Subbarao, K. Influenza vaccines: Challenges and solutions. Cell Host Microbe 2015, 17, 295–300. [Google Scholar] [CrossRef] [PubMed]
- De Clercq, E. Antiviral agents active against influenza A viruses. Nat. Rev. Drug Discov. 2006, 5, 1015–1025. [Google Scholar] [CrossRef] [PubMed]
- Beigel, J.; Bray, M. Current and future antiviral therapy of severe seasonal and avian influenza. Antivir. Res. 2008, 78, 91–102. [Google Scholar] [CrossRef] [PubMed]
- Bright, R.A.; Shay, D.K.; Shu, B.; Cox, N.J.; Klimov, A.I. Adamantane resistance among influenza A viruses isolated early during the 2005–2006 influenza season in the United States. JAMA 2006, 295, 891–894. [Google Scholar] [CrossRef] [PubMed]
- Moscona, A. Global transmission of oseltamivir-resistant influenza. N. Engl. J. Med. 2009, 360, 953–956. [Google Scholar] [CrossRef] [PubMed]
- Sheu, T.G. Dual resistance to adamantanes and oseltamivir among seasonal influenza A (H1N1) viruses: 2008–2010. J. Infect. Dis. 2011, 203, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Matthew, P.; Hall, R.J.; Sonnberg, S. Pandemic (H1N1) 2009 and Seasonal Influenza A (H1N1) Co-infection, New Zealand, 2009. Emerg. Infect. Dis. 2010, 16, 1618–1620. [Google Scholar]
- Chertow, D.S.; Memoli, M.J. Bacterial coinfection in Influenza A. JAMA 2013, 309, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Tkachuk, Z.Y. 2′-5′-Oligoadenylates as a «tool» of innate immunity. Biopolym. Cell 2013, 29, 266–276. [Google Scholar] [CrossRef]
- Wongphatcharachai, M.; Wang, P.; Enomoto, S.; Webby, R.J.; Gramer, M.R. Neutralizing DNA aptamers against swine influenza H3N2 viruses. J. Clin. Microbiol. 2013, 51, 46–54. [Google Scholar] [CrossRef] [PubMed]

© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Melnichuk, N.; Semernikova, L.; Tkachuk, Z. Complexes of Oligoribonucleotides with D-Mannitol Inhibit Hemagglutinin–Glycan Interaction and Suppress Influenza A Virus H1N1 (A/FM/1/47) Infectivity In Vitro. Pharmaceuticals 2017, 10, 71. https://doi.org/10.3390/ph10030071
Melnichuk N, Semernikova L, Tkachuk Z. Complexes of Oligoribonucleotides with D-Mannitol Inhibit Hemagglutinin–Glycan Interaction and Suppress Influenza A Virus H1N1 (A/FM/1/47) Infectivity In Vitro. Pharmaceuticals. 2017; 10(3):71. https://doi.org/10.3390/ph10030071
Chicago/Turabian StyleMelnichuk, Nataliia, Larisa Semernikova, and Zenoviy Tkachuk. 2017. "Complexes of Oligoribonucleotides with D-Mannitol Inhibit Hemagglutinin–Glycan Interaction and Suppress Influenza A Virus H1N1 (A/FM/1/47) Infectivity In Vitro" Pharmaceuticals 10, no. 3: 71. https://doi.org/10.3390/ph10030071
APA StyleMelnichuk, N., Semernikova, L., & Tkachuk, Z. (2017). Complexes of Oligoribonucleotides with D-Mannitol Inhibit Hemagglutinin–Glycan Interaction and Suppress Influenza A Virus H1N1 (A/FM/1/47) Infectivity In Vitro. Pharmaceuticals, 10(3), 71. https://doi.org/10.3390/ph10030071




