1. Introduction
Vascular diseases, characterized by either blocked or excessive blood supply to organs and tissues, are among the most serious health challenges globally [
1], leading to life-threatening conditions including stroke or heart attack. They are associated with abnormalities in blood vessel lumen geometry, such as stenoses caused by pathological deposition of atherosterotic plaque inside the vessel [
2]. The diagnosis and treatment of vascular diseases requires accurate lumen geometry quantification, in a non-invasive way where possible [
3]. Three-dimensional imaging is the main technique used to acquire quantitative information about the vasculature [
4]. Example modalities include magnetic resonance angiography (MRA) which can be either blood-flow-dependent (time-of-flight (TOF) or phase contrast angiography (PCA)) or flow-independent [
5,
6]. An invasive alternative is computed tomography angiography (CTA) [
7]. Vasculature images can be nonenhanced or contrast-enhanced (CE), which involves injecting a contrastive substance into the blood circulatory system [
8]. Vascular ultrasonography (US) is another imaging technique with good potential [
9]. However, modeling the US image formation requires different theoretical treatment [
10] to that applicable to MR and CT volumes. Further discussion of US imaging lies beyond the scope of this paper.
Parametric evaluation of the shape of the blood vessel lumen requires identification and quantification of 3D image regions representing the relevant part of the circulatory system. This is not a trivial task, for several reasons. Firstly, the vasculature exhibits a complex tree-like structure of highly curved branches of different diameters [
11,
12] (p. 9). These diameters range broadly, from 30 mm in the aorta and vena cava to as small as 30 µm in venules and arterioles [
13]. At the same time, the spatial resolution of imaging systems is limited. For instance, in clinical scanners, the spatial resolution is limited to cuboids (voxels) with a minimum sidelength of 0.2 mm. Thus, the thin regions of vessels are heavily blurred in the image, and the thinnest of them are not visible. To enhance the resolution, various technical solutions are employed to reduce the voxel volume [
14,
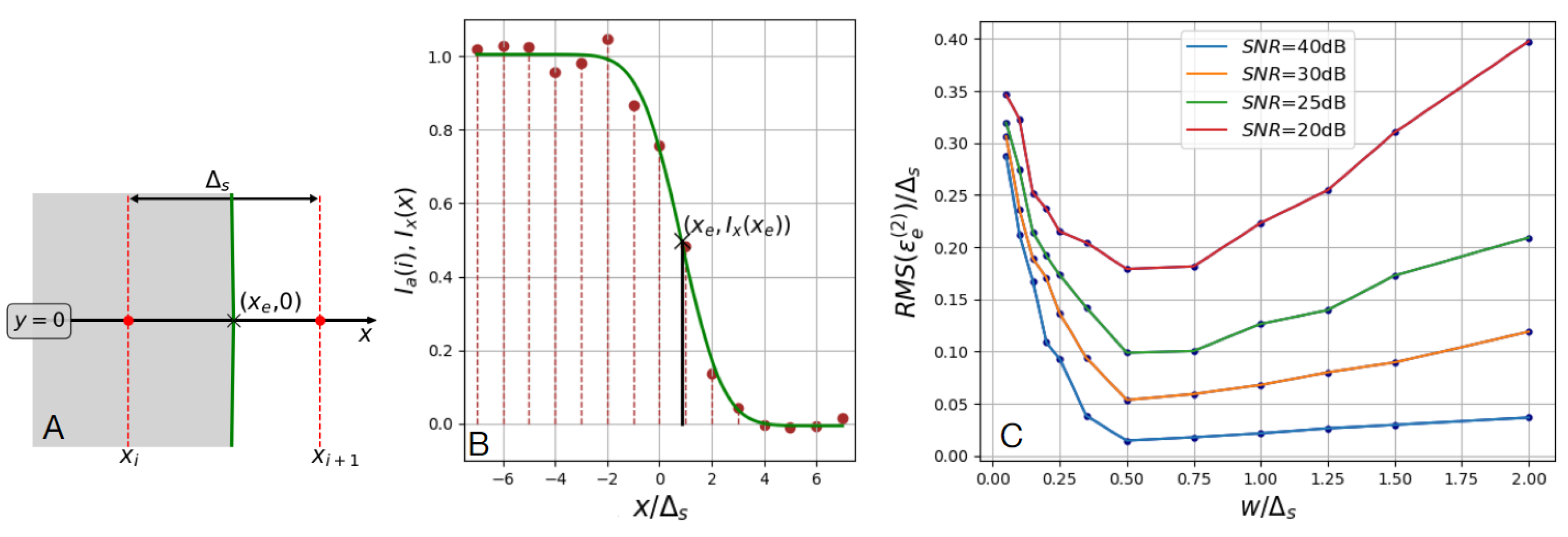
15]. However, this causes a decrease in the signal collected from each voxel, and as a consequence increases the impact of random noise as a component of the image intensity. The presence of random noise increases the uncertainty of assessing intensity levels inside the lumen, the vessel wall, and the surrounding background, adversely affecting the lumen geometry measurements (Appendix A). Image artifacts further complicate vessel image analysis.
Typically, the vessels of interest are closely surrounded by other arteries, veins and tissues. Thus, the background region features uneven image intensity and the spatially blurred signals from different regions overlap (
Figure 1). Moreover, the intensity inside the vessel walls is not constant. The plaque depositions, especially those that are calcified, exhibit different physical properties to the blood or to the contrastive medium [
16]. Under these conditions, classical, numerically efficient segmentation through thresholding becomes highly inaccurate. These factors make the task of vessel quantification in 3D images especially challenging for radiologists. Manual vessel delineation in 3D images is tedious, time-consuming, and error-prone [
17]. Its adequacy depends much on the reader’s experience and their level of fatigue. There is a strong need to develop automated techniques for accurate, fast, and objective vascularity evaluation from volumetric image data [
18].
Numerous methods of lumen segmentation and quantification have been proposed over the last few decades [
17,
18,
19]. Research in this area has further intensified with the development of deep learning as an approach to image segmentation [
20,
21]. There are two main approaches to vascular structure quantification in 3D images [
22,
23]:
The result of direct 3D image segmentation is another image. In its basic form, this image is a grid with the same spatial resolution as the original volume, where the nodes (e.g., the centers of the voxels) are attributed binary intensity values. Some of these voxels are marked by a label indicating that they represent the lumen. All the others are labeled as background. The boundary of the lumen region, which is smooth and continuous in the physical space, is discretized during image acquisition and so it remains after segmentation. For random boundary location in space, the variance of the discretization error can only be reduced by increasing imaging resolution (
Appendix A) usually at the price of increased noise.
Thus, when binary segmented, the lumen surface exhibits a stair-like or voxelized form. It requires additional spatial smoothing for visualization or to build a geometric model for blood flow simulation. However, part of the information about the actual course of this continuous surface, which is needed for faithful reconstruction, is lost in the binary segmentation step, when the continuous intensities are replaced by two-level values. From this point of view, image segmentation, although intuitive, is an unnecessary processing step. Moreover, some segmentation algorithms involve time-consuming iterative calculations [
24,
25].
Here, we focus on the second approach to estimating lumen geometry parameters, from 3D image cross-sections. These cross-sections are computed as 2D images on planes perpendicular to the vessel centerline, which is approximated by a smooth curve in 3D space [
26]. Various algorithms for the centerline extraction are available [
27,
28]. Normal vectors to the centerline define the cross-section planes. The cross-section images are obtained through 3D discrete image interpolation and resampling. The center of the cross-section image grid is usually set to lie on the centerline. The contours of the lumen are delineated, either semi-manually [
29] or automatically [
26]. For automated contour delineation, a 2D image formation model is defined. This model accounts for image smoothing by an equivalent scanner impulse response, either one-dimensional along radial lines [
26,
30] or two-dimensional over the image plane [
31], as well as for the random noise. The effect of smoothing, naturally featured by imaging sensors, plays an important role in the chosen approach. Namely, smoothing converts distances between image points and the lumen edge to image intensity variations. Thus, more relevant information is available (in the transition region around the edge) compared to voxelwise thresholding (
Appendix A). The model is fitted with the use of the least-squares (LS) algorithm. This involves long-lasting iterative minimization of a nonlinear error function and is likely to become stuck in its local minima, depending on the initial parameter values.
We use LS fitting as a reference method and apply the convolutional neural network (CNN) driven by 2D cross-section images for fast estimation of the model parameters [
31,
32]. The contours found at predefined increments along the centerline arc are lofted to build the geometric models, e.g., for visualization (
Figure 2). Lofting is a technique used in computer-assisted design programs to create 3D surfaces or solid objects [
29]. They are formed from sequences of cross-section curves—the contours of the lumen, in our case.
Past works that followed the modeling approach incorporate circular or elliptical cylinders as abstract geometric objects that represent the vessel lumen in the image space. In fact, the actual shape of the lumen contours significantly deviates from these idealized figures (
Figure 2). This happens in particular in the vessels narrowed by atherosclerotic plaque [
33]. The novelty of our method lies in using B-splines—parameterized curves that accurately represent the natural lumen shapes in normal and pathological vasculature—as part of the image formation model. Although B-splines have been used previously for blood vessel contouring [
34,
35], they were fitted to the approximate, spatially discretized surface of the lumen region in binary segmented 3D image. In our approach, the fitting process takes place in the image at full bit depth intensity instead of in the discretized spatial domain. Innovatively, the lumen contour parameters are estimated by a neural network, providing increased speed and robustness compared to LS fitting. The use of B-splines makes our method compatible with the isogeometric approach [
36] to image analysis, test object (physical phantoms) design and computational blood flow simulation.
2. Materials and Methods
To illustrate the properties and capabilities of the proposed lumen modeling method, the following real-life and synthetic images were used:
2.1. PAVES MR Dataset
Steady-state MR contrast-enhanced volumes, available as part of the PAVES dataset, are high spatial resolution images which, in principle, allow evaluation of the shape of the artery lumen, e.g., for the purpose of identifying stenosis. However, these images take several minutes to acquire. During this rather long time, both the arteries and veins become filled with the gadolinium-based contrast medium. As a result, the intensities of these regions show similar values, making the arterial regions difficult to identify and distinguish from the neighboring veins. The arteries are smaller in diameter, and their cross-sections are close to circular, except for pathological stenosed vessels featuring wall-thickening. On the other hand, veins typically have oval shapes and are located in pairs next to an artery (
Figure S1). Thus, the cross-sections of artery–vein complexes do not exhibit circular shapes. They are non-circular, although their boundaries are smooth. As highlighted in [
39], veins can be used by surgeons to “bypass” a blocked artery, provided they are of the correct diameter and length. Therefore, delineating veins and arteries in high-resolution GdCE (gadolinium contrast-enhanced) MR images is a valid medical image processing task, formulated as a Grand Challenge [
37].
The research problem defined in this example is to evaluate the suitability of the proposed method to delineate boundaries of the veins and arteries, given their steady-state GdCE MR volume. The PAVES dataset 5 was selected for this purpose, as the clinical significance of its properties was demonstrated in [
39]. To obtain the cross-sections, the centerlines of the blood vessel regions were extracted and smoothly approximated by application of the following 3D image preprocessing steps [
26]:
Multiscale “vesselness” filtering [
27,
40] to enhance the blood vessels regions [
41];
Vesselness map thresholding to obtain binary lumen representation;
Binary region thinning to produce a skeleton;
Skeleton parsing [
42] to identify the blood vessel tree branches between bifurcations;
Approximating the skeleton branches by a differentiable function in 3D (to initialize their centerline).
Figure 3A shows a maximum intensity projection on the axial plane of the PAVES dataset 5 TWIST (subtracted time-resolved acquisition) volume, for left volunteer extremity. The arrow indicates a stenosis in the anterior tibial artery. The above preprocessing algorithm was applied for the TWIST 3D image. The parsed skeleton is illustrated in
Figure 3B. The tibial artery segment b14 was selected for further analysis, among other vessel-tree branches enumerated by the skeleton parsing algorithm.
Tangent vectors to the smooth vessel centerline curve were computed at a number of consecutive points at distances of about 0.5 mm from each other. Together with the binormal vectors, these tangent vectors define the Frenet-Serret frame at each point. This frame often becomes rotated around the tangent vector, causing an undesirable twist in the surface of an elongated object. A rotation minimizing frame algorithm [
43] was then used to compute the vessel local coordinate systems 0
xy, in which the 2D lumen cross-sections were calculated by resampling the MR volume. These image preprocessing and analysis algorithms were implemented in both
(version R2022B) and
(version 3.10.13). The codes are available on request.
Examples of the cross-sections are presented in
Figure 3C, in the form of a mosaic of 15 × 15-pixel images, where the pixel size is 1.0 mm × 1.0 mm. The consecutive lumen cross-sections show minimal twist. Basically, a cross-section contains a dark background where there is no tissue filled with blood, a medium-intensity vein region, and a bright artery blob with the highest concentration of the contrastive medium. As shown in the upper five rows of
Figure 3C, the healthy arteries feature circle-like cross-sections. However, the atherosclerotic occlusions cause narrowing of the vessel lumen (row six, right half) and reduce the intensity of the artery region. As an effect of collateral blood circulation, the width of the artery lumen may be restored down the vessel, as can be seen in rows nine to fifteen (
Figure 3C).
2.2. CAT08 Coronary CTA Dataset
We applied the proposed lumen modeling method to 3D images of a training set made available by the Rotterdam Coronary Algorithm Evaluation Framework [
38,
44,
45]. It was designed for the development of algorithms for lumen segmentation, as well as detection and quantification of coronary artery stenoses, and provided in the MICCAI 2008 Coronary Artery Tracking Challenge (CAT08). There are 48 datasets available in the Rotterdam Challenge data repository. In 18 of the datasets, anatomical segments of the arteries (specified in [
44]) have been manually annotated by three experts. These 18 sets constitute the CAT08 training set. For 78 of the training set segments, expertly annotated contours of the arteries cross-sections are available. We used this information, together with the corresponding 3D images, to train the CNN estimator and as a reference for testing. It is worth noting the three observers needed approximately 300 h to complete their annotation of the data [
38].
The CT images of the Rotterdam dataset were acquired with different scanners over a period of one year, for a number of patients under observation for cardiac diseases. The average voxel size for the whole dataset is 0.32 mm × 0.32 mm × 0.4 mm. We clipped the CT volume intensity to HU (Hounsfield units) and normalized it to . The volumes were resampled on the planes of marked contours, to obtain lumen cross-sections centered at the centerline points.
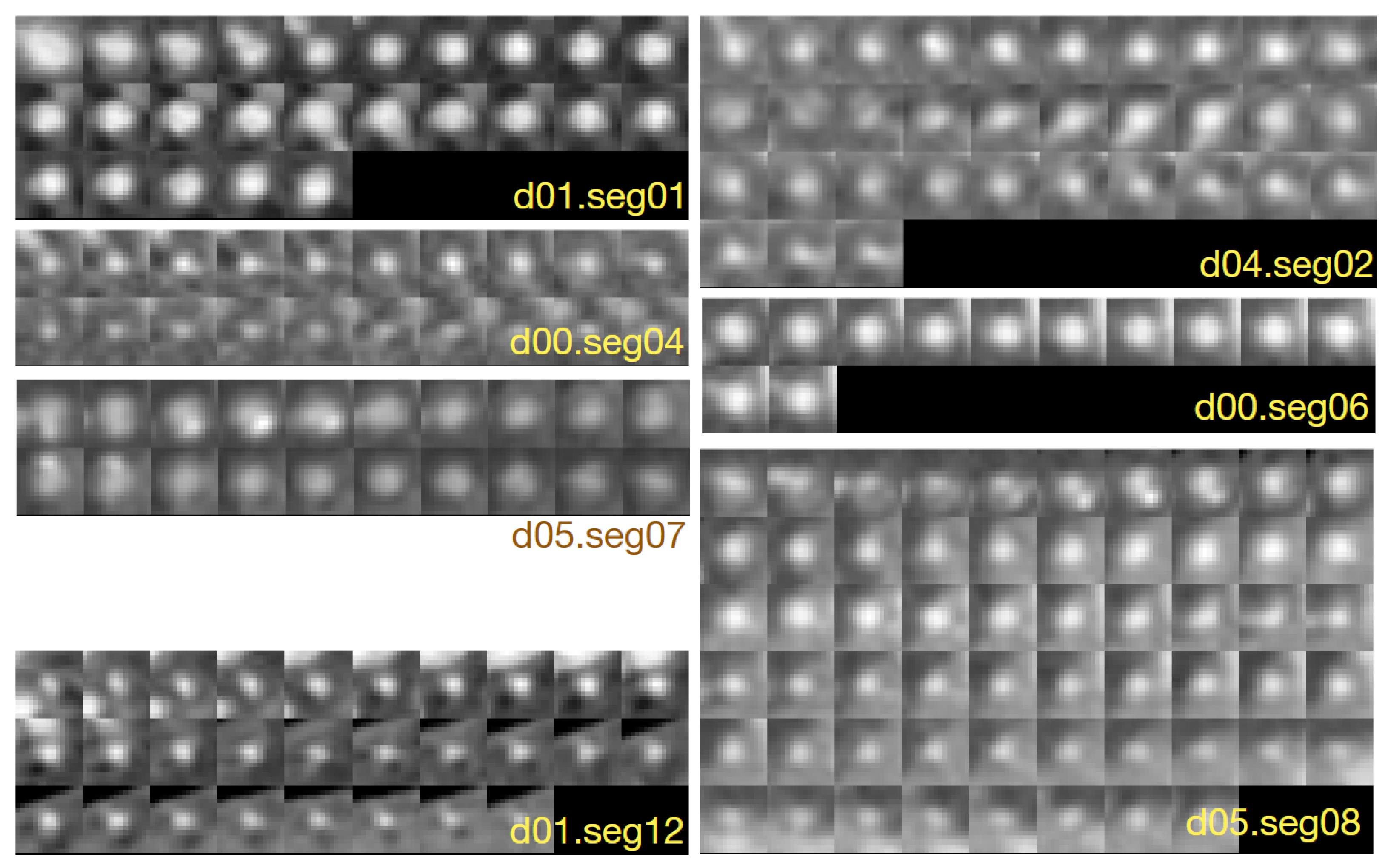
Figure 1 shows example cross-sections of a few segments in the CAT08 dataset, computed for a sampling interval of 0.45 mm. Segments
01 and
02 are consecutive anatomical parts of the right coronary artery (RCA). However, the sections of these segments presented in
Figure 1 belong to different datasets and differ in appearance quite significantly. The contrast for the 25 images of
01 is much higher and the area of its lumen region does not change as much as the lumen region for the 33 sections of
02.
Thresholding would not be useful for lumen region segmentation of 02. The lumen diameter noticeably decreases along the 20 sections of 04 (from left to right, row by row) and approaches subpixel values in the bottom row. At the same time, bright regions in the background dominate the image content. The contrast of images in 06 is good. The lumen region is well defined and its shape is close to circular. However, it shows a significant blur, similar to all other images. This property, originating in the limited spatial resolution of the CT scanners, leads to high uncertain localization of lumen boundaries, especially in noisy images.
Segments
07 and
08 form consecutive parts of the left anterior descending artery (LAD). The first section (the leftmost in the upper row) of
07 is closest to the heart, the last section (rightmost in the lower row) is the immediate neighbor of the first section in
08. The sections in the upmost row of
08 exhibit a reduced lumen area and bright image spots in the sixth and seventh sections, apparently caused by atherosclerotic plaque. If they become part of the lumen model as a result of image analysis, postprocessing will be needed to detect and quantify them.
12 is the first obtuse marginal artery (OM1). Its lumen is well distinguished from the immediate background. However, a significant area is occupied by black triangular-shaped regions where the normalized intensity is close to zero (−300 HU or less in the scanner-acquired CT volumes). The typical cross-section shape of the vessel lumen marked by the observers is neither circular nor elliptical (
Figure 4).
All the image nonidealities pointed out in the previous paragraph make lumen modeling a difficult task. In our numerous experiments, the LS model fitting method failed in the case of too many CAT08 arterial branches, as background regions were of locally higher intensity than the lumen. On the other hand, the proposed parameterized B-spline lumen contour model can be robustly identified by a feedforward CNN, which will be demonstrated in the Results section.
2.3. Image Formation Model and the Synthetic Dataset Generation
We assumed the blood vessel cross-section image centered at any point
in the 3D space is a convolution of the function
, which represents the lumen and its background, with
− the imaging system effective impulse response (point spread function PSF):
where
are image coordinates on the cross-section plane. The function
in (
1) combines the effects of the 3D image scanner impulse response and interpolation necessary for computing the cross-section intensity from the 3D image via resampling. This assumption, though idealized, is relevant to most practical situations [
46].
We further assume the function is constant within the lumen region, surrounded by a background of different but also constant intensity. These assumptions are often not met, as mentioned before with regard to the coronary artery CE CTA images. We will demonstrate in the Results section that CNN-based lumen geometry restoration is robust to image intensity variations in the regions of interest.
We considered equidistant sampling on a Cartesian grid of points (although other sampling strategies are possible to tune the estimator properties):
where
denotes the sampling interval. The cross-section image size is
. The image intensity at
is the sampled convolution (
1) multiplied by lumen intensity step
b and added to the background intensity
a:
The lumen section boundary at each centerline point
,
is described in this paper by a closed (periodic) B-spline curve which encloses the higher-intensity lumen region
in
:
The in-plane blur in our model is assumed to be an isotropic Gaussian [
46,
47,
48], where
w is a parameter:
The lumen contour is approximated by a parameterized curve lying on the plane of the vessel cross-section. For this purpose, we selected a B-spline curve [
49,
50], which is a smooth, piecewise polynomial allowing stable, locally controlled approximation of real-world data. To define a curve of this kind, it is necessary to specify a sequence of
,
, real numbers, the knots
which define the curve segment boundaries,
, and
n as the B-spline degree. At the boundaries (internal knots), the segments are joined together to satisfy continuity constraints. We use third-degree polynomials, meaning that the first and second derivatives are continuous along the curve arclength.
The closed B-spline of degree
n is controlled by
points
, such that the first control point
overlaps with the last one in the sequence. For any
t within the B-spline segments, the curve points can be computed as [
51]
The B-spline (
7) is a linear combination of degree
n basis B-splines
. These can be defined recursively using the de Boor algorithm [
49]
To compute the cross-section model intensity (
3), one needs to specify the boundary of the lumen region
in (
4). This is described by a B-spline curve controlled by a sequence of points that have to be defined. In our approach, the control points for a cross-section centered at the vessel centerline are located on the radial lines extruding from the image center at
D angles:
The distance in physical space from the image center to the
ith control point
is
,
.
R is a scale factor (fixed for the image of given size) and
is a vector of adjustable B-spline parameters. The coordinates of the control points are computed as
Based on our experience with blood vessel images of various modalities, the
w parameter of the PSF in (
5) does not change significantly within an image. Therefore, a constant value of
w is assumed and measured separately, e.g., via analysis of appropriate edge blur. Then, there are altogether
adjustable parameters
. Combining expressions from (
1)–(
11), one obtains
where
given by (
1) is multiplied by the parameter
b (the intensity pulse corresponding to the lumen region) and added to the background intensity
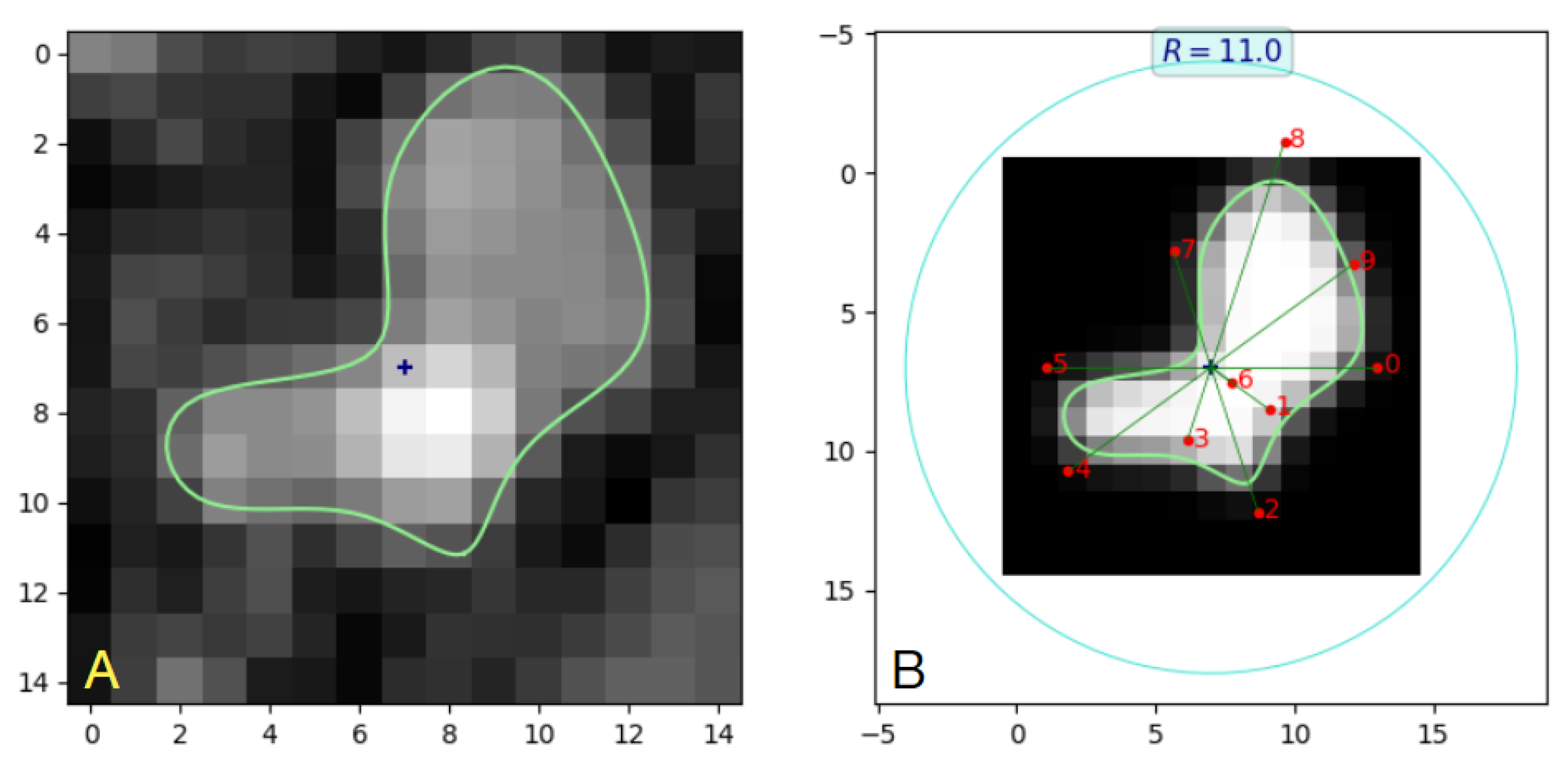
a. Geometrical interpretation of the elements of vector
is provided in
Figure 5A, for
. The lumen is represented in (
4) by a set of points of unit intensity (the shaded area in
Figure 5B), surrounded by the background points with intensity equal to zero (the white area in
Figure 5B).
Figure 5C shows a low-resolution image obtained as a result of sampling a blurred version of the ideal image depicted in
Figure 5B.
Considering the complex shape of the region
, no closed-form expression for the integral (
1) appears to be available. To compute 2D images for the training and testing datasets, we evaluated the right-hand side in (
1) numerically, with a
implementation of the
(version 4.6.0) library. A synthetic dataset comprising 10,000 images was created. The synthetic dataset was considered in two versions: noiseless and noisy. The noise addition procedure is described in the Results subsection devoted to the PAVES dataset.
2.4. Least-Squares Model Fitting for the PAVES Dataset
The aim is to estimate the parameter vector
, given an acquired 2D digital image
of the lumen cross-section of known
R,
w, and
. This image is the input to the CNN, which approximates the mapping from the image space to the parameter space. We will compare the performance of two approaches to parameter vector estimation: least-squares (LS) model fitting and prediction by a convolutional neural network [
31]. Using the former method, the lumen contour parameters can be estimated by minimizing the sum of squared differences between the acquired and modeled image samples:
The
(version 1.11.4)
module was used to perform minimization of the sum-of-squares in (
13).
2.5. Quantitative Evaluation of Contours Similarity
The lumen contour is defined by a closed continuous curve on the cross-section plane in a physical space. For numerical analysis, it is represented by a finite set of discrete point—vertices of a polygon chosen to approximate the curve. We will use the term “contour” to refer to its discrete version. We applied two measures to evaluate the discrepancy between contours. Consider the contours
A and
B, comprised of
and
points, respectively, resampled to
and
. The mean distance between them is defined as
where
is a set of Euclidean distances between a given point
x and each of the points in set
Y. In our experiments, we interpolated the contours being compared such that their cardinality was
=
= 100 and
=
= 500.
The second measure was the Dice Similarity Coefficient (DSC), computed for the sets
and
. These sets denote all the points of a common contour plane that lie inside the contour
A and
B, respectively. The DSC is defined as
In our numerical experiments, the sets and were approximated by all the points of a high-resolution image (e.g., 800 × 800 points, coplanar with the contours), which were within a polygon defined by the points A and B, respectively.
2.6. CNN-Based B-Spline Model Parameter Estimation
In the proposed method, the CNN plays a role of a nonlinear regressor. Its input is a lumen cross-section image. It predicts the values of the image model parameters at the output. A feedforward network was trained to predict the B-spline parameter vector
d. The architecture is shown in
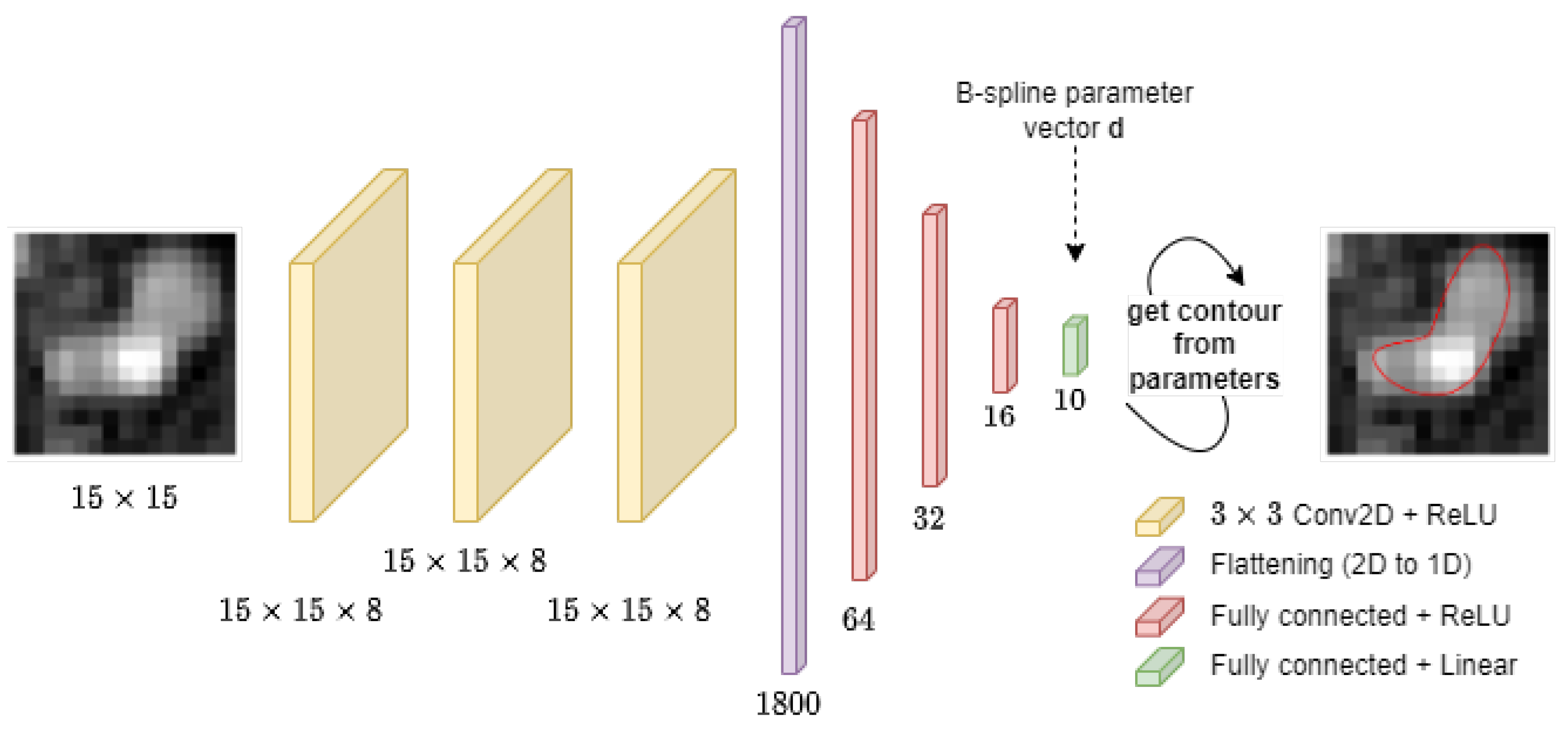
Figure 6. Three 2D convolution layers are followed by a flattening layer and by four fully connected layers. The three convolutional layers feature 3 × 3 kernels, with a kernel number equal to 8 for each layer, “same” input padding type, and ReLU nonlinear activation function [
52]. The flattening layer transforms the multichannel 2D feature maps into vectors. The vectors are then processed by three fully connected layers with 64, 32, and 16 neurons, respectively, and ReLU activation. The output layer estimates the 10 elements of vector
d and features a linear activation function. In total, the network has 119,290 trainable parameters and was implemented in
keras (version 2.13.1).
Network training was performed with the mean squared error as the cost function, optimized using the Adam algorithm [
53]. Mini-batches of 64 examples were used during training. The maximum number of training epochs was set to 500, but an early stopping algorithm was employed to stop training before overfitting occurred. The early stopping patience parameter was set to 15 epochs.
The synthetic dataset was used to train the network for prediction on the PAVES dataset. For the test set, 10% of the data was excluded, which yielded 9000 examples. The validation set used in the early stopping algorithm was extracted from the training set as its 20% fraction.
The CAT08 dataset used for neural network training, validation, and testing has 652 examples. Of these, 66 (10%) were excluded for the test set, and the rest were split into training and validation sets in the same proportion of 90:10. Targets in the form of B-spline-approximated expert contour annotations were separated into three sets—one for each expert. The training procedure was repeated for each expert’s target set, yielding three trained models.
In summary, five models were trained: two for the PAVES datasets (noiseless and noisy) and three for the CAT08 datasets. These models are evaluated in the Results.
2.7. Methods and Tools for Statistical Analysis of the Results
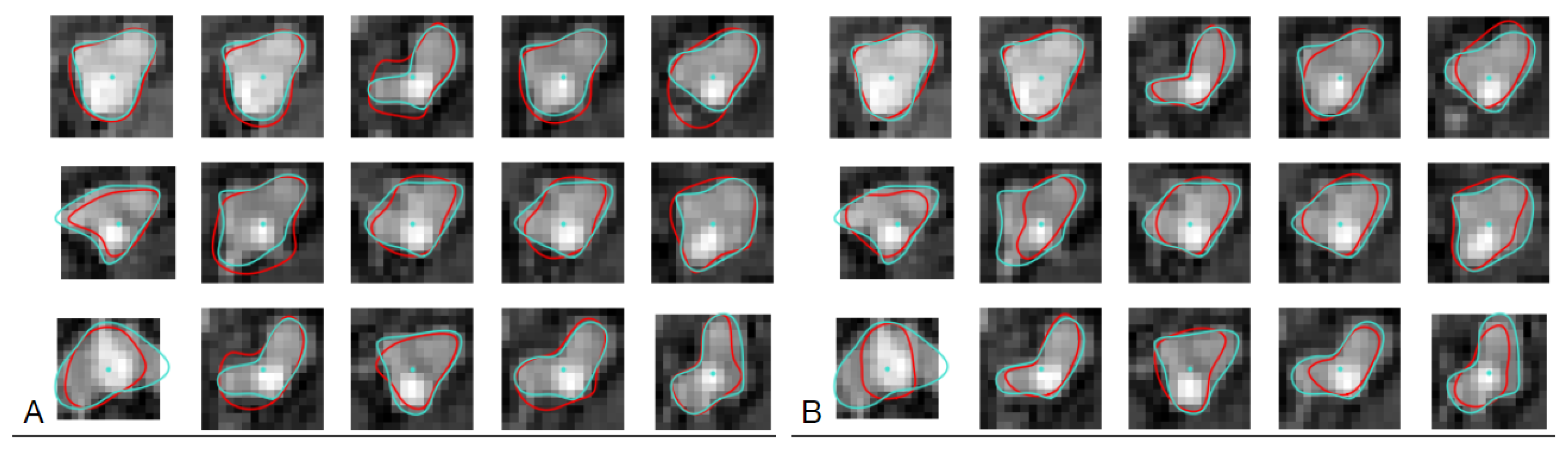
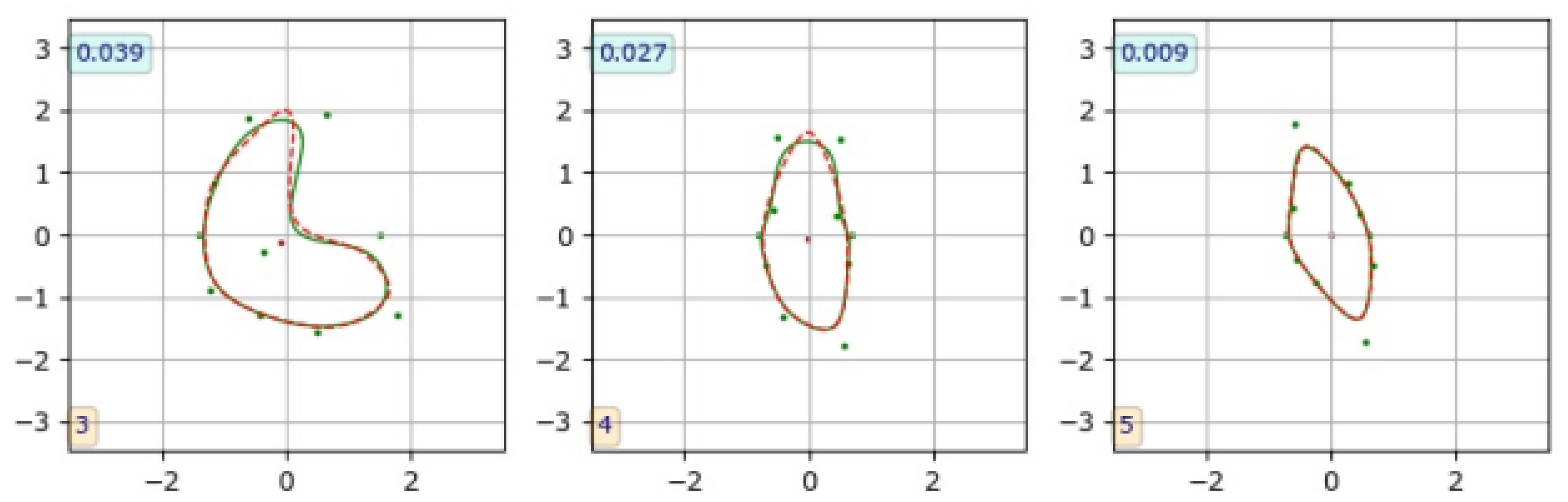
The cross-sections of coronary arteries visualized in the CAT08 dataset were annotated by three observers. Each of them delineated the lumen boundaries, presented as polygons on cross-section planes. The polygon vertices are available in the dataset. The three trained CNNs were applied to the same images to generate corresponding lumen contours. For each test image from the test set of
images, two contours were then available, one marked by an observer and the other predicted by the corresponding CNN instance (
Figure 7).
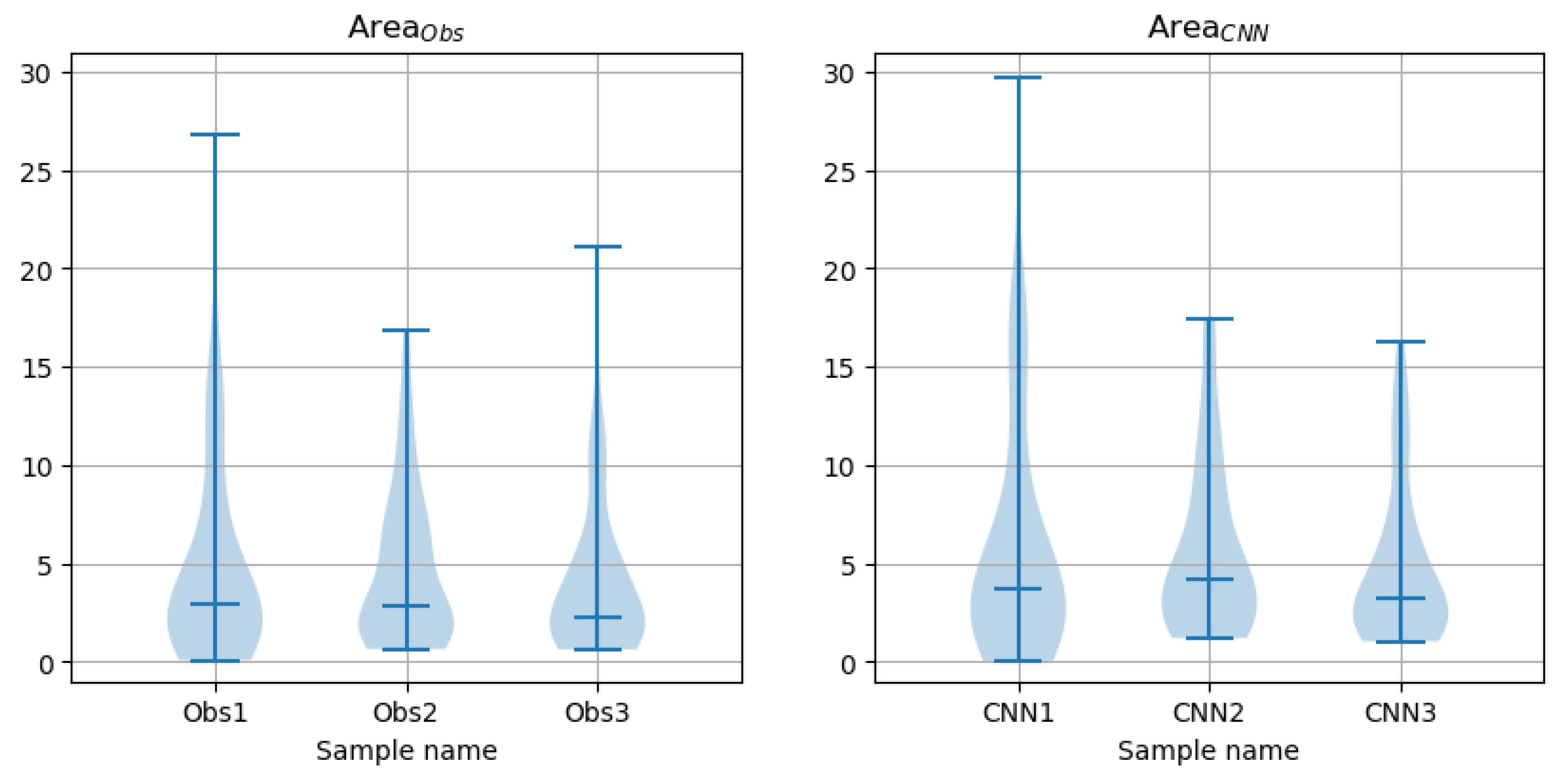
The image area covered by each contour was computed to give
= 198 pairs of values—
and
, respectively, for the observer and for the CNN. These are the data points. Since they relate to the same image space, they are paired and match each other. In the context of this study, the question is whether the two methods of marking the lumen contour (by the observer and by the CNN) differ [
54]. This question can be answered by a paired statistical test. The choice of a particular test has to be preceded by a test for the normality of the data populations. D’Agostino and Pearson’s normality test, available as
scipy.stats.normaltest() function in the
Scipy (version 1.11.4) library, was applied for this study. The area measurements failed the normality test. Based on this result, the paired samples Wilcoxon test was applied to the data,
scipy.stats.wilcoxon(). It is a nonparametric test of the null hypothesis that two samples come from the same population against an alternative hypothesis, as described and discussed in the Results.
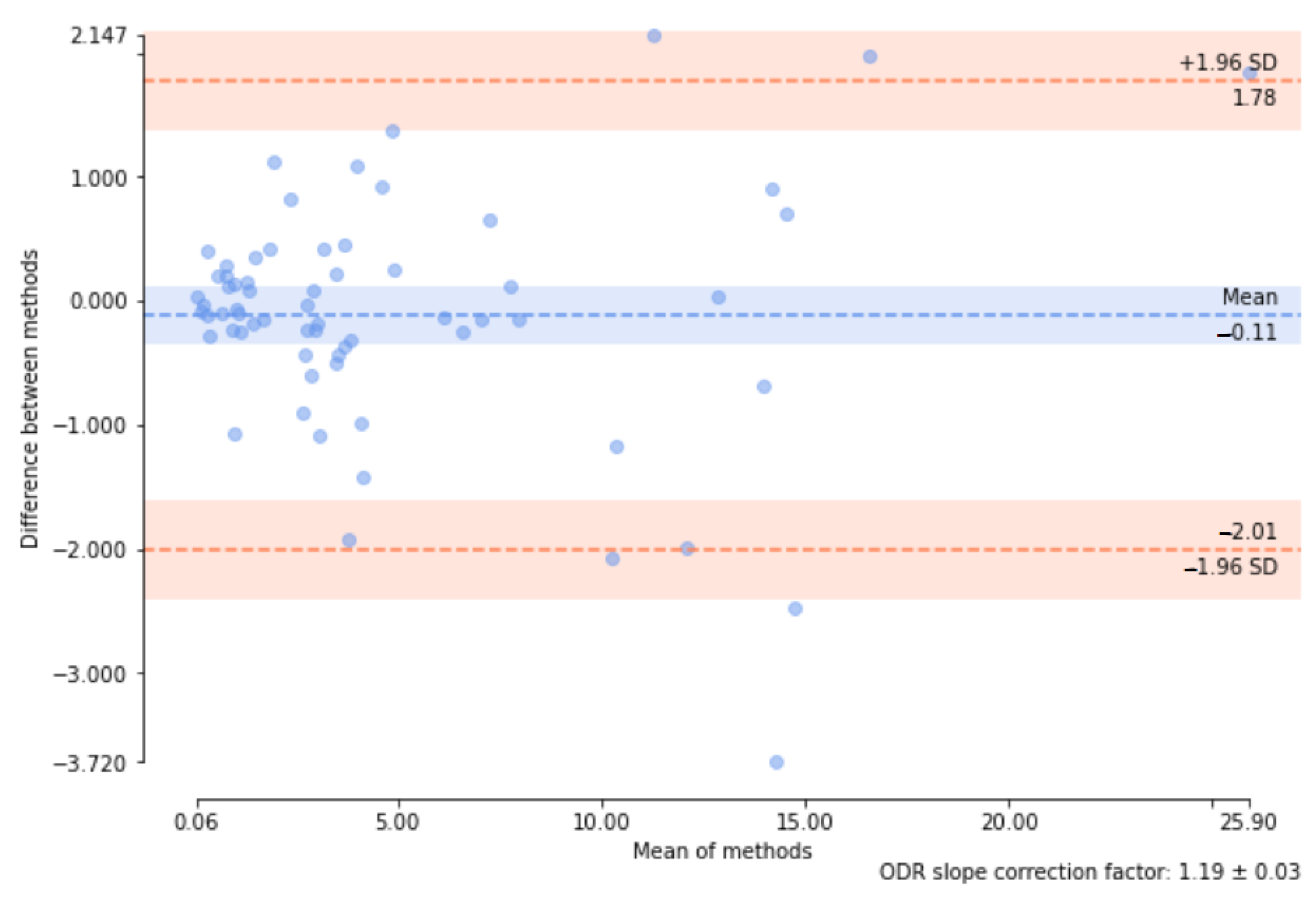
More quantitative information about statistical differences between the methods is available from the Bland–Altman plot which visualizes both bias and confidence intervals of the sample differences [
55].
Figure 8 obtained with
pyCompare library (version 1.5.4) is an example of such a plot.
All the computations in this work including statistical analyses were performed on a standard PC computer (Intel Core i5-8300 H, 2300 GHz, 16 GB RAM, NVIDIA GeForce 1050) under Windows 10 Home operating system in miniconda3 Python (version 3.10.13) programming environment. Occasionally, a laptop or an iPad Pro machines, of similar performance were used.
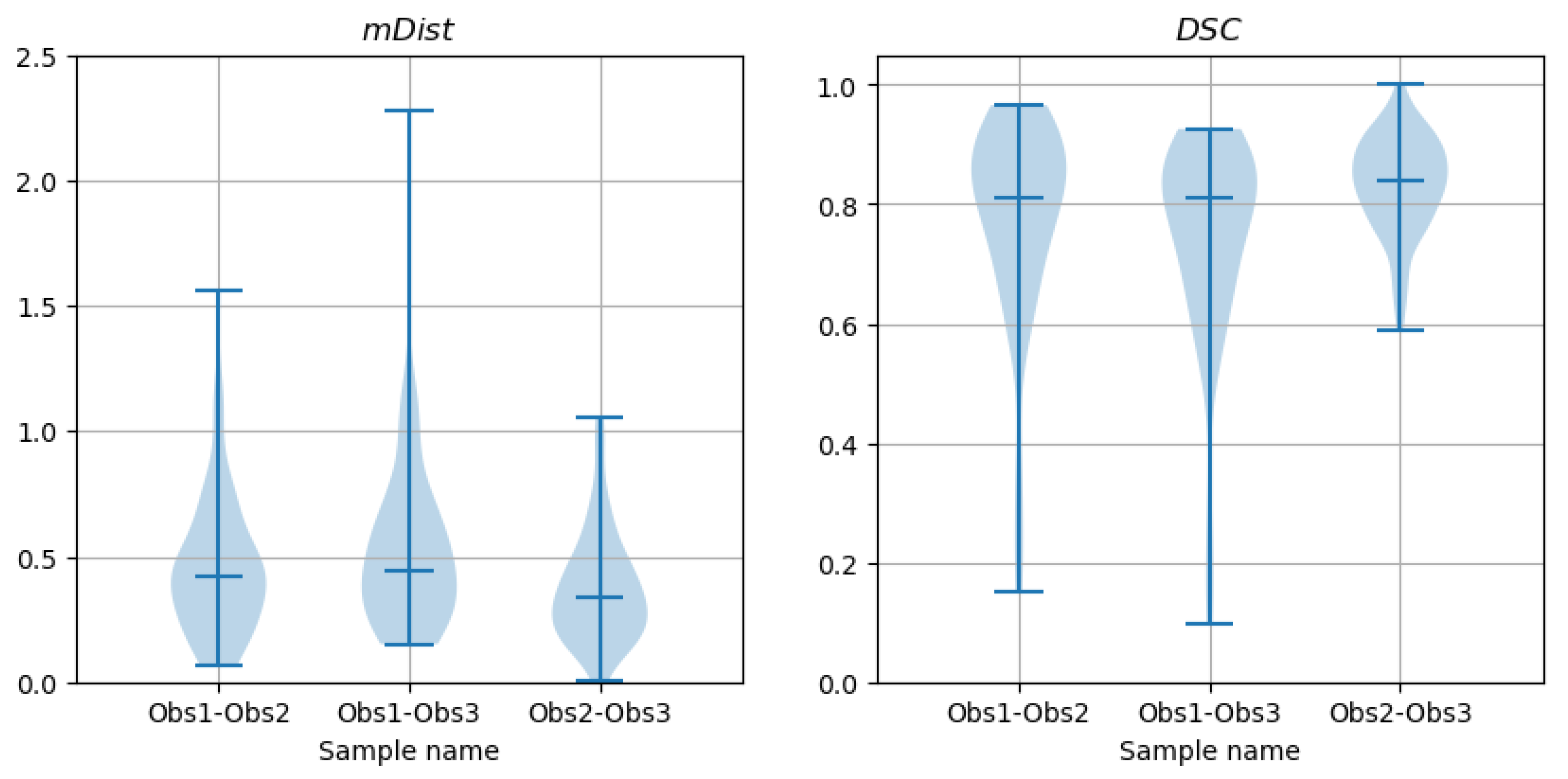
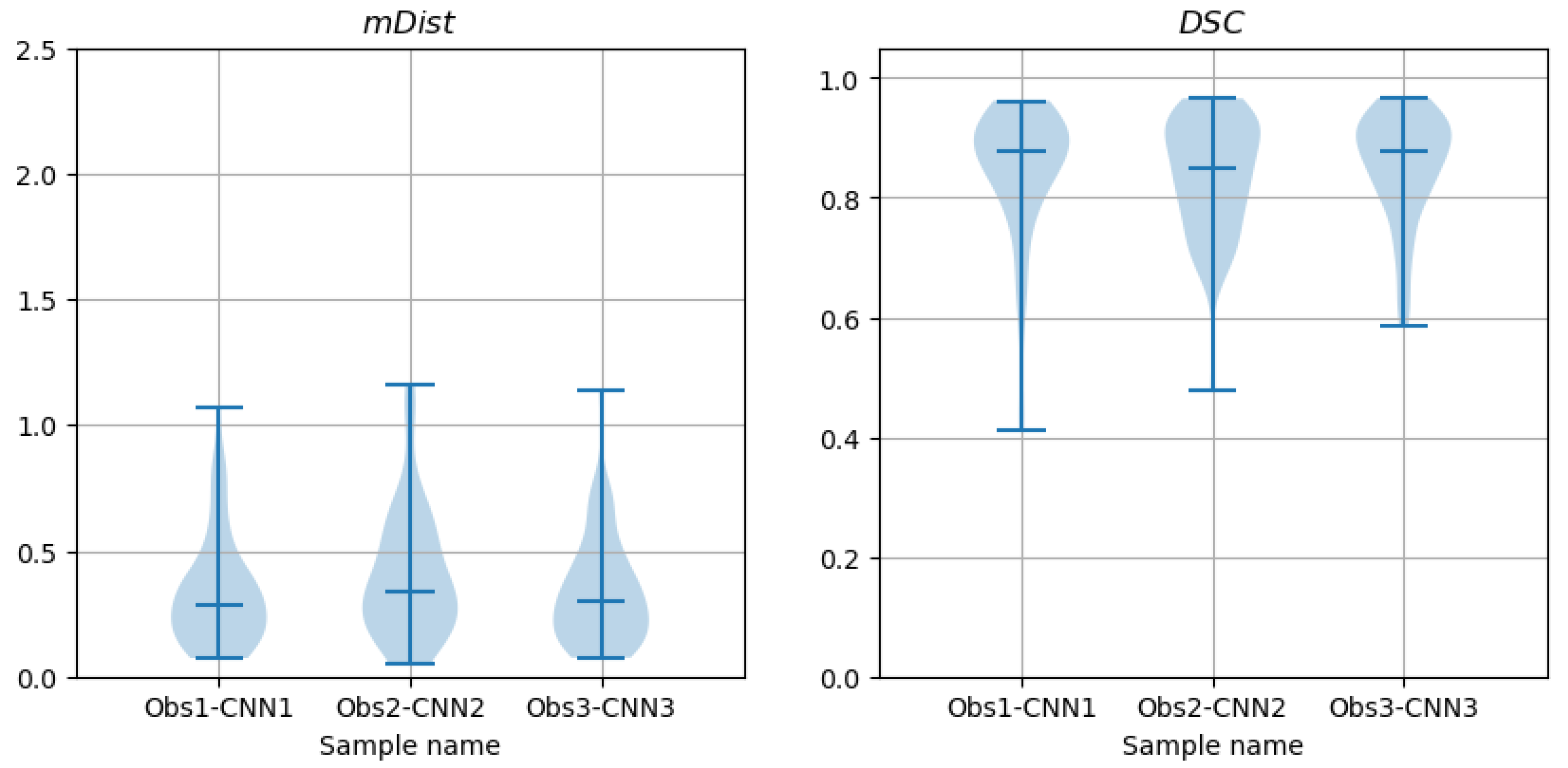
The above procedure was applied to the
(
14) and
(
15) coefficients which quantify the distances between contours marked by different methods, e.g., to test whether the populations of
data computed for contours marked by Observer 1 and Observer 2 and the
data for contours outlined by Observer 1 and Observer 3 are the same or different. Those tests are described and discussed in Results.
4. Discussion
This study has demonstrated that the integration of B-splines with advanced deep learning techniques significantly enhances the precision and reliability of blood vessel lumen modeling from 3D images. Based on two publicly available datasets, PAVES and CAT08, B-spline-approximated lumen contours were used in the image formation model to quantify the non-circular shape of blood vessels with subvoxel accuracy. Such accuracy cannot be obtained using existing circular or elliptical approximations.
The experimental results obtained for the PAVES dataset demonstrated good agreement between the B-spline parameters estimated by the CNN and those computed with the use of LS model fitting. It was shown that the differences can be reduced by proper augmentation of the synthesized images used for the CNN transfer training. Unfortunately, the PAVES dataset does not contain any reference information. No radiologists’ annotations are available, so there are no ground-truth contours to compare with those related to B-spline parameters found in the course of LS model fitting or CNN-based prediction. Still, the dataset was useful to demonstrate that the proposed lumen quantification approach, based on the image formation model, can be successfully applied to highly non-circular blood vessel cross-sections. Moreover, we demonstrated the possibility of both obtaining reasonable contours for real-life MR images with the use of a transfer-trained CNN and making the CNN predictions robust to spatial intensity variations by adding fractal texture patterns to the training images. Optimizing the augmentation strategy will be the topic of future research.
A ground-truth reference is available within the CAT08 dataset quantified with our method. Due to the complexity of CT images of coronary artery cross-sections, LS model fitting does not provide the expected stability. Even with the use of troublesome constrained optimization, it finds local minima too often and the fitting results depend heavily on the initialization points. In contrast, the CNN can be trained to ignore image nonidealities (e.g., bright objects in the background) and produce lumen contours which are consistent with the observers’ delineations. Of course, it is also much faster than human experts. Its application to image annotation would relieve the radiologist of this task.
Our experiments show that the CNN can be trained to mimic an individual observer’s style of contour marking. There was good agreement between the shapes of CNN- and observer-produced contours. The CNN reproduced the interobserver variability very well. This raises the question of the ground truth. Future work is planned to design and implement realistic digital and physical phantoms of known tubular geometry and blood distributions. These phantoms will be used to simulate and acquire images, which will then be evaluated by both radiologists and CNNs. The aim is to conduct a quantitative comparison of the respective contour delineations with the phantom properties.
There is a significant difference between the CNN and LS algorithms in terms of the computation time needed. When implemented on a moderate-performance PC (Intel Core i5 plus NVIDIA GeForce 1050), the CNN predicts the contour parameters in microseconds per image, compared to about 100 s for the iterative nonlinear LS program.
The CNN trained to predict the lumen model parameters has a simple architecture (
Figure 6) and a relatively small number of adjustable weights. Its training on a dataset of 500 images requires only about 2 min on a standard laptop computer (MS Windows 11 Pro, Intel Core i5, 2400 GHz) with no GPU acceleration. Implementing the neural network is straightforward and does not present any major technical challenges. The proposed solution is much less computationally demanding than the majority of popular CNN applications.
Presumably, the efficiency of the proposed approach stems from the CNN being guided by the centerline while processing image samples. The CNN receives direct information about the distance of the sampling points to the lumen boundary. This information is clear-cut and encoded in the sampled intensity. Such a priori knowledge is built into our method. The CNN has only to approximate the inverse mapping from the intensity space to the point coordinates space, which is expected to be smooth (
Appendix A).
5. Conclusions
Accurate geometric modeling of blood vessel lumen from 3D images is crucial for vessel quantification, as part of the diagnosis, treatment, and monitoring of cardiovascular diseases. It is a challenging task, given the complex geometry and internal structure of normal and pathological blood vessels, their widely varying diameters, and the limited capabilities of imaging sensors and scanners. The arteries and veins are composed of highly curved branches, interleaved in a space filled with various organs and tissues.
Manual vessel delineation by radiologists is time-consuming, tedious, and error-prone. The development of automated techniques for lumen quantification has been the subject of intensive research over the last few decades. Research has taken two main directions: 3D segmentation-based lumen quantification and model-fitting-based 2D lumen cross-section quantification along an approximate centerline. Binary segmentation introduces discretization errors, which may be critical for lumens with small diameters. Some segmentation algorithms require time-consuming iterative calculations. Therefore, we have followed the second approach.
Unlike other centerline-based methods, which assume a circular or elliptical vessel cross-section, the proposed method employs parametric B-splines combined with image formation system equations to accurately localize highly curved lumen boundaries. The parameters of this model are estimated using a feedforward convolutional neural network driven by full bit depth image samples on planes orthogonal to the centerline. This is in contrast to known B-spline-based lumen models which are identified from coarsely outlined binary regions in segmented images. The need for segmentation is avoided. Superior modeling accuracy of highly curved lumen boundaries in datasets comprising real-life MR and CT images is demonstrated.
The CNN, which has a rather straightforward architecture, can be trained on either computer-synthesized or expert-annotated images within just minutes using a standard desktop or laptop computer equipped with a standard graphics processor. The trained network predicts the parameter values in microseconds, compared to the minutes per image needed for alternative LS model fitting methods. High speed is thus one more advantage of the proposed estimator.
Due to its high speed, high accuracy and robustness, the proposed method can have an impact on and appear significant to medical diagnosis and research. It could be of great help to radiologists in their routine work. Also, the trained CNNs can be used for automated annotation of images in large datasets needed for the development of data-driven artificial intelligence solutions for healthcare. As the high accuracy of the vessel wall reconstruction is crucial for faithful blood flow simulation, this research area can benefit from using CNNs and B-splines for lumen modeling.