1. Introduction
Functionalized magnetic nanoparticle (fMNP)-protein binding events, transduced via magnetic spectroscopy, enable biomolecule detection within a sample solution [
1,
2,
3]. Biomolecule binding increases fMNP hydrodynamic diameter (
) [
4,
5] and/or induces the formation of fMNP aggregates [
6]. The fMNP magnetic moment will have a delayed response to an external magnetic field due to
increases and aggregate formation. The magnetic moment response can be modeled via Brownian and Néel relaxation [
7,
8,
9]. fMNPs have been engineered to sense biomarkers, including SARS-CoV-2 [
10] and H1N1 nucleoprotein [
11], in vitro. Salt induced changes in fMNP aggregation pose an obstacle to the clinical translation of fMNP-based aggregation assays for in vivo applications. Additional factors impeding aggregation assay in vivo applications include, protein corona formation [
12,
13] and non-specific bio-distribution. Changes in salt concentration can alter nanoparticle (NP) stability [
4,
14]. Which can cause changes in fMNP aggregation that are not proportional to the target biomolecule concentration. To this end, we have investigated the effects of salt concentration, in the biological range, on target induced fMNP aggregation. Our aggregation assay is a model system consisting of biotinylated MNPs (biotin-MNPs) that target streptavidin. We used Lodestone Biomedical’s Nanoparticle Characterization System (an AC magnetic spectrometer, NCS) [
15] to transduce biotin-MNP-streptavidin binding into a measurable signal. The NCS emits a 1 kHz AC magnetic field with 20 mT field strength and conveys the biotin-MNP harmonic response at 3 kHz [
15]. We used the phase of the 3rd harmonic (
) to assess the level of biotin-MNP aggregation [
16,
17] and the magnitude of the 3rd harmonic (
) to determine the Fe concentration present in our samples after biotinylation [
18]. We studied changes in streptavidin induced biotin-MNP aggregation in phosphate buffered saline (PBS) solutions. 1x PBS is a water-based solution with an approximate pH of 7.4 and salt contents that match those found in the human body. 1x PBS contains 137 mM NaCl, 10 mM phosphate, and 2.7 mM KCl. To alter the ionic strength of PBS we diluted it with deionized water (DI), resulting in the following PBS solutions: 1.00x, 0.70x, 0.50x, 0.30x, 0.05x, 0.045x, 0.04x, 0.035x, 0.025x, 0.02x, 0.015x, 0.01x, 0.005, and 0.00x (DI). We determined biotin-MNP zeta potential and
in each PBS solution. Additionally, we investigated differences in biosensor response characteristics in the 1.00x, 0.50x, 0.03x, 0.015x, 0.005x, and 0.00x PBS solutions. Here, we monitored changes in biotin-MNP harmonic response when in the presence of varying amounts of streptavidin. We refer to biotin-MNP
as a function of streptavidin concentration as a streptavidin-response curve. We compared the slope, maximum
, and linear range of each streptavidin-response curve prepared in the 1.00x, 0.50x, 0.03x, 0.015x, 0.005x, and 0.00x PBS solutions.
According to the extended Derjaguin-Landau-Verwey-Overbeek (xDLVO) theory, biotin-MNP stability will be dependent on the sum of the repulsive electrostatic double layer (EDL) force, which is dependent on biotin-MNP surface charge density and the electrolyte concentration of the diluent, and the attractive van der Waals and dipolar forces acting upon similarly charged biotin-MNPs [
8,
19]. A positive sum suggests stability, whereas a negative sum suggests instability and hence aggregation. If a nanoparticle (NP) has a high surface charge density and/or is in a low salt environment, particle interactions are repulsive and dominated by the EDL force [
20]. On the other hand, if a NP has a low surface charge density and/or is in a high salt environment, particle interactions are attractive and dominated by the van der Waals force [
20]. NP zeta potential absolute value tends to decrease in high salt environments due to a counter ion layer that forms on the NP surface, decreasing its effective surface potential and leading to instability [
14]. Unstable NPs tend to aggregate, leading to an increase in NP
. Jans et al., 2009 demonstrated the effects of electrolyte concentration on gold NP (AuNP) stability [
4]. They found that the
of citrate capped AuNPs increased with increasing NaCl concentration. Interestingly, larger (∼100 nm) AuNPs were less stable in high salt environments compared to smaller (∼40 nm) AuNPs. MNPs are more apt to cluster, compared to NPs with non-magnetic cores, due to the attractive dipolar force acting amongst MNPs, in addition to the attractive van der Waals force [
21]. Nikam et al., 2014 demonstrated the effects of electrolyte concentration and zeta potential on cobalt zinc ferrite (CZF) MNP stability [
14]. Their PEGylated CZF MNPs were stable in PBS for 2 weeks, although they underwent an approximate 75 nm hydrodynamic increase in double distilled water (DDW) for the same duration, indicating instability/aggregation in DDW. Moreover, they found a linear relationship between NaCl concentration and CZF MNP zeta potential. An increase in NaCl concentration led to a decrease in zeta potential absolute value. Zeta potential absolute value decreases with increasing electrolyte concentration given an increase in EDL thickness due to surface charge screening. Given previous work investigating salt induced changes in fMNP stability, we can expect a decrease in streptavidin induced biotin-MNP aggregation with decreasing salt concentration.
In this work we also show that receptor density can be used to tune the linear region of the streptavidin-response curve for our aggregation assay. The linear region of the streptavidin-response curve is commonly referred to as the dynamic range, which is the measurable concentration range of a biosensor [
22]. Takae et al., 2005 demonstrated the tunability of AuNP-based aggregation assay dynamic range via ligand density [
23]. They investigated lactose functionalized AuNPs that target Ricinus communis agglutin (RCA120) lectin. Lactose-AuNP’s with 40% and 65% surface area lactose coverage had dynamic ranges spanning from 25 to 200
g/mL and 50 to 400
g/mL, respectively. A decrease in surface area lactose coverage led to an increase in assay sensitivity. Researchers have engineered biosensors targeting metal ions [
24], small molecules [
25], and proteins [
26] with tunable dynamic ranges. Li et al., 2013 engineered a non-cross linking AuNP-based aggregation assay with a tunable dynamic range [
27]. Their target protein, alkaline phosphatase (ALP), altered the aggregation levels of AuNPs diluted in a Tris-HCL buffer solution containing adenosine triphosphate (ATP). The dynamic range of their AuNP-based aggregation assay was tuned via the addition of metal ions to the sensing solution.
Concerning fMNP ligand density modulation, Elias et al., 2013 demonstrated a non-linear relationship between superparamagnetic MNP ligand density and cell targeting [
28]. They found that optimal cell targeting occurred when the MNPs had intermediate ligand densities. This relationship was held regardless of MNP core diameter or ligand type. Generally, MNPs with 1.7 ligands per 100 nm
2 exhibited optimal cell targeting. This topic is expanded upon in Alkilany et al., 2019 review [
29] of the relationship between a NP’s (including MNPs, AuNPs, and quantum dots) ligand density and its cellular uptake. The relationship between MNP ligand density and protein sensing dynamics, transduced via MPS, were reported by Wu et al., 2021. They functionalized MNPs with polyclonal antibodies (pAbs), to lend the MNPs and affinity for SARs-CoV-2 spike or nucleocapsid proteins and monitored odd harmonic amplitudes of magnetization as the pAb-MNPs were exposed to varying concentrations of their target [
30]. MNPs were functionalized with 1, 2, 3, or 4 pAbs per MNP. For pAb-MNPs targeting spike protein, a pAb:MNP ratio of 3:1 yielded a linear
response from 0 to 12.5 nM spike protein. Additionally, for pAb-MNPs targeting nucleocapsid protein, a pAb:MNP ratio of 4:1 yielded a linear
response from 3.13 to 400 nM nucleocapsid protein. pAb-MNPs with 1:1, 2:1, or 3:1 pAb:MNP ratios did not yield linear
responses with respect to target concentration, regardless of target protein (splike or nucleocapsid). Our work differs from that of Wu et al., 2021 in that we are studying changes in biotin-MNP biosensor response characteristics (the dynamic range, slope and
), via investigating changes in biotin-MNP
as a function of target concentration, instead of odd harmonic amplitudes. Given previous work investigating NP dynamic range tunability with ligand density, we expect an increase in biotin-MNP sensitivity with decreasing biotin density.
In this work, we verified the feasibility of using the xDLVO theory to predict fMNP target response for given salt concentrations. The results showed that biotin-MNP streptavidin targeting was independent of salt concentration for 0.005x to 1.00x PBS solutions. Additionally, we show that the dynamic range (or measurable streptavidin concentration range) for biotin-MNPs decreases with decreasing biotin density. Moreover, our biotinylation protocol and subsequent biotin-MNP physical characterization suggests that and zeta potential measurements may not be reliable metrics for functionalization confirmation protocols, given that we found a non-linear relationship amongst biotin density and as well as zeta potential.
2. Materials and Methods
2.1. Materials
Amine functionalized 25 nm magnetite (SHA 25) particles were purchased from Ocean Nanotech, San Diego, CA, USA. N-hydroxysuccinide (NHS)--biotin was purchased from Thermo Fisher Scientific, Waltham, MA, USA. Streptavidin was purchased from Rockland Immunochemicals, Pottstown, PA, USA.
2.2. Physical Characterization of SHA 25 MNPs
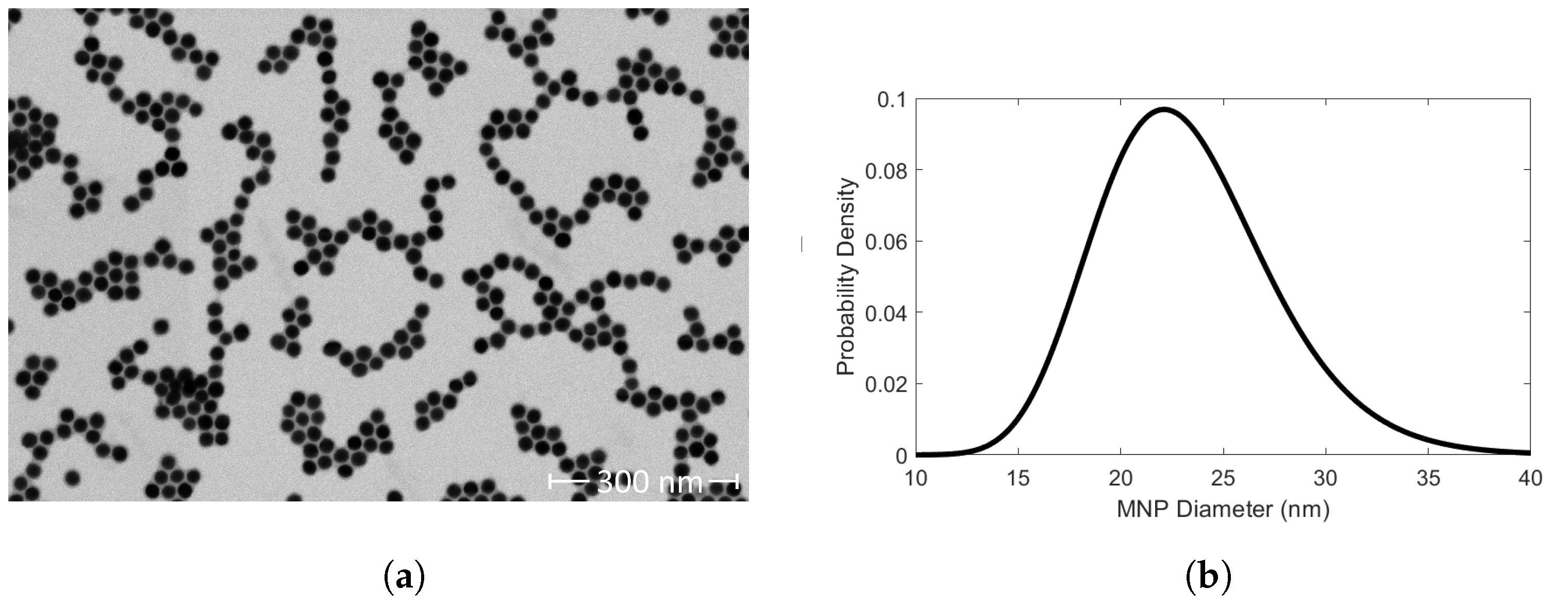
A carbon film mesh grid was dipped into a 0.5 mg Fe/mL SHA 25 solution. The SHA 25 MNPs adsorbed to the carbon film mesh grid were imaged with a combined focused ion beam scanning electron microscope (FIB-SEM, Helios 5 CX DualBeam for Materials Science, Thermo Fisher Scientific), to verify their size and shape distribution. The FIB-SEM had a current and voltage of 21 pA and 20 kV, respectively. The FIB-SEM images were analyzed with ImageJ (Fiji). ImageJ has an ‘Analyze Particles’ feature that measures the area of non-interacting particles. The ‘Analyze Particles’ feature is compatible with 8-bit binary images. We used minimum error thresholding to algorithmically threshold our image. The output of the thresholding operation was a binary image. The ‘Analyze Particles’ feature was applied to the 8-bit binary image to calculate the area of 371 SHA 25 MNPs. We algebraically manipulated the reported areas to calculate the SHA 25 MNP mean diameter and standard deviation. We conducted measurements of 10 mg Fe/L SHA 25 MNP samples. The SHA 25 MNP solution was diluted with PBS. The measurements were conducted via dynamic light scattering (DLS) with a Zetasizer Ultra Red (Malvern Panalytical). 1 mL of each SHA 25 solution was transferred to a disposable 10 × 10 plastic cell (DTS0012). The measurements were conducted in quintuplicate at 25 °C with a light scattering collection angle of 174.7° (back scatter). To verify the zeta potential of the SHA 25 MNPs in PBS, 1 mL of a 10 mg Fe/L SHA 25 MNP solution was injected into a disposable folded capillary cell (DTS1070). The zeta potential measurements were conducted in quintuplicate, with measurements before and after each zeta potential measurement to monitor MNP stability.
2.3. Biotinylation of SHA 25 MNPs
Two Eppendorf tubes containing 300 L of 5 mg Fe/mL SHA 25 MNP solutions were centrifuged at 12,000 RPM for 30 min at 25 °C. Once the centrifugation was complete, the diluent was aspirated and the SHA 25 MNPs were resuspended in 300 L of PBS. Quantities of 2 mg of NHS--biotin are stored in each vial of the Thermo ScientificTM No-WeighTM Format. To prevent moisture condensation, we allowed the NHS--biotin to equilibrate to room temperature before opening the vial. To produce biotin--SHA 25 MNPs with ∼10 available biotins for streptavidin binding per MNP (10 biotin-MNPs), the NHS--biotin was diluted to 0.50 mg/mL with DI, then 1.54 L of the 0.50 mg/mL NHS--biotin solution was added to one of the Eppendorf tubes containing the SHA 25 MNP solution. To produce biotin--SHA 25 MNPs with ∼90 available biotins for streptavidin binding per MNP (90 biotin-MNPs), the NHS--biotin was diluted to 2.86 mg/mL, then 158.80 L of the 2.86 mg/mL NHS--biotin solution was added to the SHA 25 MNP solution. Next, both mixtures were left to incubate at room temperature for 30 min. Once the SHA 25 MNPs were biotinylated, free biotin was removed from the solution via centrifugation. Each mixture was centrifuged at 12,000 RPM for 30 min at 25 °C 8 times. After each centrifugation, the diluent was aspirated and the 10 biotin-MNP and 90 biotin-MNP solutions were resuspended in 300 L of DI. The of SHA 25 serial dilutions (0, 1, 2, 3, 4 and 5 g/L Fe concentrations) were measured with the NCS. The of the 10 biotin-MNP and 90 biotin-MNP solutions were compared with those of the SHA 25 MNP serial dilutions to determine the Fe concentration of the biotin-MNP solutions. Lastly, the 10 biotin-MNP and 90 biotin-MNP solutions were diluted to 1 mg Fe/mL based on their .
2.4. Physical Characterization of Biotin-MNPs
Biotin-MNP aliquots (10 mg Fe/L concentration) were submitted to DLS, to verify their
and zeta potential in the 1.00x to 0.00x PBS series. The DLS measurements were conducted in quintuplicate. The zeta potential measurements were conducted in quintuplicate as well, with
measurements before and after each zeta potential measurement to monitor biotin-MNP stability. We measured the time evolution of biotin-MNP
in 1.00x PBS at room temperature for 8 h, with
measurements approximately every 20 min. The measurement parameters for the biotin-MNP samples were identical to those used for the physical characterization of the SHA 25 MNPs (
Section 2.2).
2.5. Magnetic Particle Spectroscopy Measurements
To obtain streptavidin-response curves for the 90 biotin-MNPs in 1.00x, 0.50x, 0.03x, 0.015x, 0.01x, 0.005x, and 0.00x PBS solutions, streptavidin serial dilutions spanning from 0.0 M to 2694.8 M streptavidin were prepared in each PBS solution. Each MPS sample contained 2.5 L of a 1 mg Fe/mL 90 biotin-MNP solution mixed with 2.0 L of a streptavidin serial dilution sample. The 1.00x PBS 0.0 M streptavidin sample was measured repeatedly at room temperature for 12 h, with measurements approximately every 20 min, to study the time evolution of biotin-MNP stability in 1.00x PBS. Additionally, streptavidin-response curves for the 10 biotin-MNPs were obtained in 1.00x PBS and DI. To obtain the streptavidin-response curve in 1.00x PBS for the 10 biotin-MNPs, a streptavidin serial dilution spanning from 0 M to 193 M streptavidin was prepared in 1.00x PBS. To obtain the streptavidin-response curve in DI for the 10 biotin-MNPs, a streptavidin serial dilution spanning from 0 M to 322 M streptavidin was prepared in DI. Each MPS sample was prepared in triplicate. The of each sample was measured in triplicate with the NCS after an approximate 40 min incubation at room temperature. At the onset of each measurement, the NCS identifies the magnetic center of each sample. Next, the sample is measured at 7 points along the z-axis of the sample (from bottom to top). Where the center point is located at the magnetic center of the sample. The 7 point measurement is repeated 4 times during each NCS measurement.
We compared the 90 biotin-MNP streptavidin-response curves in 1.00x, 0.50x, 0.03x, 0.015x, 0.01x, 0.005x, and 0.00x PBS solutions to determine the effects of salt concentration on streptavidin induced biotin-MNP aggregation. We compared the 90 biotin-MNP and 10 biotin-MNP streptavidin response curves in 1.00x PBS to determine the effects of biotin density on streptavidin induced biotin-MNP aggregation. We compared the slope, maximum , and linear range of each streptavidin-response curve. We used analysis of covariance (ANCOVA) to determine whether the slopes of each of the curves was statistically significantly different. Specifically, Matlab’s aoctool command was used to perform the ANCOVA testing.
2.6. xDLVO Interparticle Energies
We modeled the repulsive EDL energy as well as the attractive van der Waals and dipolar energies acting amongst biotin-MNPs to gain further insights on particle stability in each PBS solution. The EDL energy between 2 NPs (sphere-sphere geometry) was calculated as [
31,
32]
where
is the permittivity of free space,
is the relative permittivity of the medium,
is the biotin-MNP magnetic core radius,
is the biotin-MNP surface potential, and
is the inverse Debeye length. EDL energy will decrease with increasing electrolyte concentration due to changes in biotin MNP Debeye length and surface potential. The zeta potential of the biotin MNPs in each PBS solution was used in place of the surface potential. Walker et al., 2009 [
31] and Fiegel et al., 2019 [
32] similarly substituted NP zeta potential for surface potential in their EDL calculations. Our highest concentrated electrolyte solution, 1.00x PBS, has the following electrolyte contents: 157.00 mM Na
+, 142.00 mM Cl
−, 4.45 mM K
+, and 7.30
, and 4.60 mM H
2. We assumed electrolyte concentration decreased linearly with decreasing PBS strength. Therefore, we assumed 0.50x PBS had 50% less electrolytes than 1.00x PBS, for example. However, we handled the EDL energy calculation for DI differently to agree with reported values of the Bjerrum length
in pH 7 DI at room temperature.
is the interparticle distance at which the EDL energy is comparable in magnitude to the thermal energy of the system [
33].
in pH 7 DI water at room temperature is 1
m, which corresponded to a Debye length (
) of 4.10
m for our biotin-MNPs in DI. van der Waals and dipolar energies remain constant with respect to electrolyte concentration. The van der Waals energy between 2 NPs was calculated as [
31]
where
is the Hamaker constant and
is the characteristic wavelength of the medium, which assumed to be 100 nm [
32]. van der Waals attractive energy is proportional to the particle Hamaker constant. Bergsrtom et al., 2011 determined the Hamaker constant for magnetite NPs in water was 33 zJ [
34]. We utilized this Hamaker constant value for our van der Waals interparticle energy calculation. The dipolar energy between 2 MNPs was calculated as [
19,
31]
where
is the permeability of free space and
is the magnetic saturation. Interparticle dipolar energy is proportional to the magnetic saturation of the biotin-MNPs. Wang et al., 2021 reported that Ocean Nanotech SHA 25 MNPs had a magnetic moment per particle of 1.58 ×
emu (1.58 ×
) [
35]. We divided this value by the biotin-MNP core volume to calculate the magnetic saturation of our biotin-MNPs and ultimately the dipolar interaction energy. We modeled dipolar energy as an attarctive interaction. Dipolar energy can be either attractive or repulsive depending on the angle between the magnetic moment and the line connecting the particle centers [
36]. If the angle is
then the interaction is attractive, however if the angle is
then the interaction is repulsive. We used two-sample Kolmogorov-Smirnov (KS) tests to determine whether the total interation energy corresponding to each PBS solution had statistically significantly variations Specifically, Matlab’s kstest2 command was used to perform the KS testing.
4. Discussion
SHA 25 MNPs diluted in PBS had a
of 60.19 ± 1.90 nm (
Figure 2c). Previous work from Wu et al., 2021, reports SHA 25 MNPs had a
of 28.28 ± 10.38 nm in DI after 30 min of ultrasonication [
35]. Our SHA 25 MNP measurements were conducted without ultrasonication or other means of sample manipulation. It is important to note that the choice of diluent and Fe concentration can affect the accuracy of
measurements. DLS measurements conducted in DI typically yield
values 2-10 nm larger than the particle’s actual
[
41]. This inaccuracy is attributed to the absence of ions in solution to shield long distance interparticle interactions [
41]. Lim et al., 2013 reported an approximate 148 nm increase in MNP
due to an increase in the sample solution’s Fe concentration from 100 to 250 mg Fe/L for 18 nm superparamagnetic MNPs [
37]. Constant
data was acquired for the 18 nm superparamagnetic MNPs at Fe concentrations between 10 and 50 mg Fe/L. Therefore, we conducted our
measurements with 10 mg Fe/L MNP samples for our 23.35 ± 4.29 nm core MNPs. Moreover, MNP core size must also be considered when determining an optimal Fe concentration for
measurements. At a constant Fe concentration, larger MNPs have a lower collision frequency, hence their
would more accurately reflect the physical properties of individual MNPs, compared to smaller MNPs. When comparing literature values of NP
one must consider the diluent, NP concentration, and core size to make an informed comparative analysis.
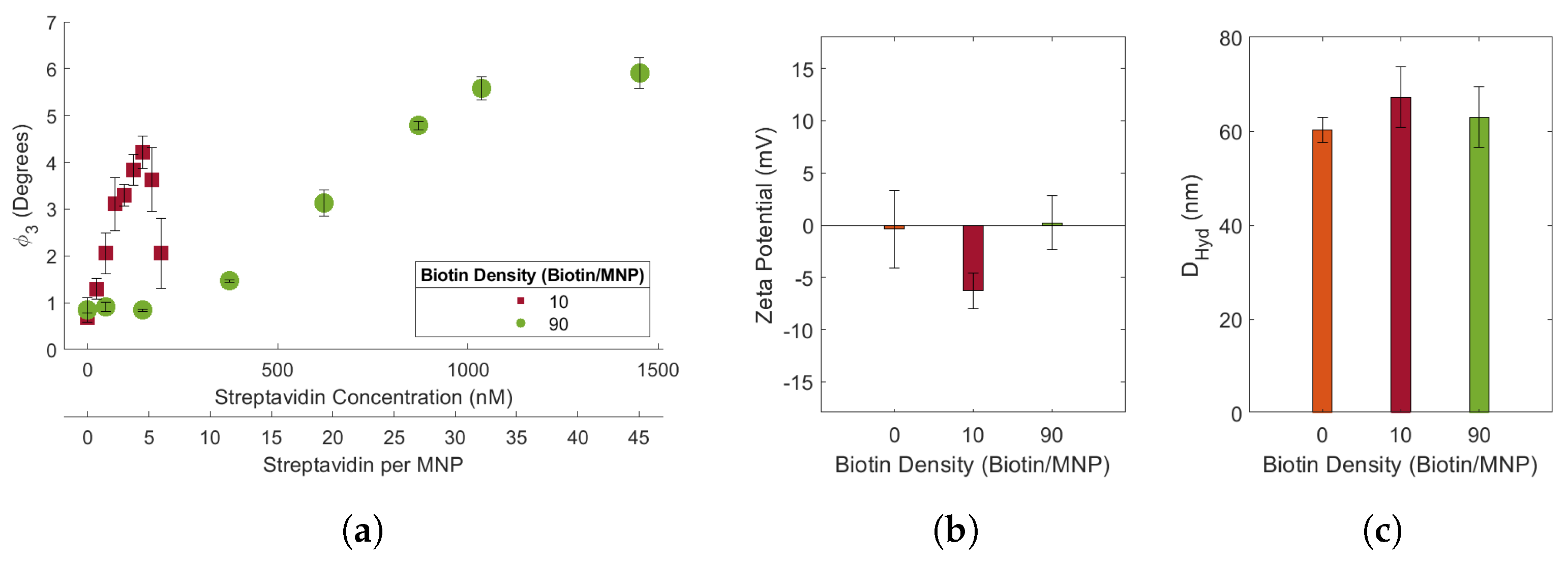
After biotinylating SHA 25 MNPs with ∼10 and 90 biotins/MNPs, we investigated the effects of ligand density on streptavidin induced biotin-MNP aggregation. We estimate that MNP biotin density is 2 times the streptavidin:biotin-MNP ratio at which the maximum
occurs. Traditional methods for NP ligand density quantification [
38] include nuclear magnetic resonance (NMR), fluorescently labelled ligands, and absorption spectroscopy. Previous works have used an absorption spectroscopy technique, wherein the ligand density of AuNPs was estimated by the shape of the absorbance versus added protein curve [
42,
43]. The protein concentration at which the absorbance curve plateaued indicated the amount of ligand bound to the AuNP surface. Our biotin-MNP ligand density quantification method is similar, however we multiply the peak value of our target response curve with 2. The multiplication factor is applied since our added protein is an aggregation inducing agent. We assume that each streptavidin is bound to 2 biotin-MNPs. Streptavidin-response curves were obtained for the 10 and 90 biotin-MNPs in 1.00x PBS. We compared the slope, maximum
, and range of the linear region of each streptavidin-response curve. Our results suggest an increase in streptavidin-response curve slope with decreasing biotin density. In other words, biotin-MNP streptavidin sensitivity increased with decreasing biotin density. Additionally, we found the streptavidin-response curve maximum
decreased with decreasing biotin-density. We suspect lower biotin density MNPs form smaller and more loosely packed clusters with streptavidin, compared to high biotin density MNPs. Our results can be leveraged for MNP-based aggregation assay development. fMNP target sensitivity and dynamic range can be tuned with ligand density. Another element of our data to consider for aggregation assay development pertains to the physical characterization of the SHA 25, 10 biotin-MNPs, and 90 biotin-MNPs. Our results suggest a non-linear relationship amongst ligand density and MNP
as well as zeta potential. NP
and zeta potential are commonly used to assess the level/success of NP functionalization. Our results suggest that
and zeta potential may not indicate the level/success of functionalization given that the SHA 25 and 90 biotin-MNPs showed no significant variation in
or zeta potential. An alternative method to assess the level of NP functionalization is to expose the bare and potentially functionalized NPs to varying amounts of target and compare transducer outputs. If the potentially functionalized NPs aggregate in the presence of their target, while the bare NPs remain fully dispersed, one can presume the NPs were successfully functionalized. This method could also be used to determine NP receptor density, wherein the target concentration corresponding to the maximum transducer output is an approximate indication of NP receptor density.
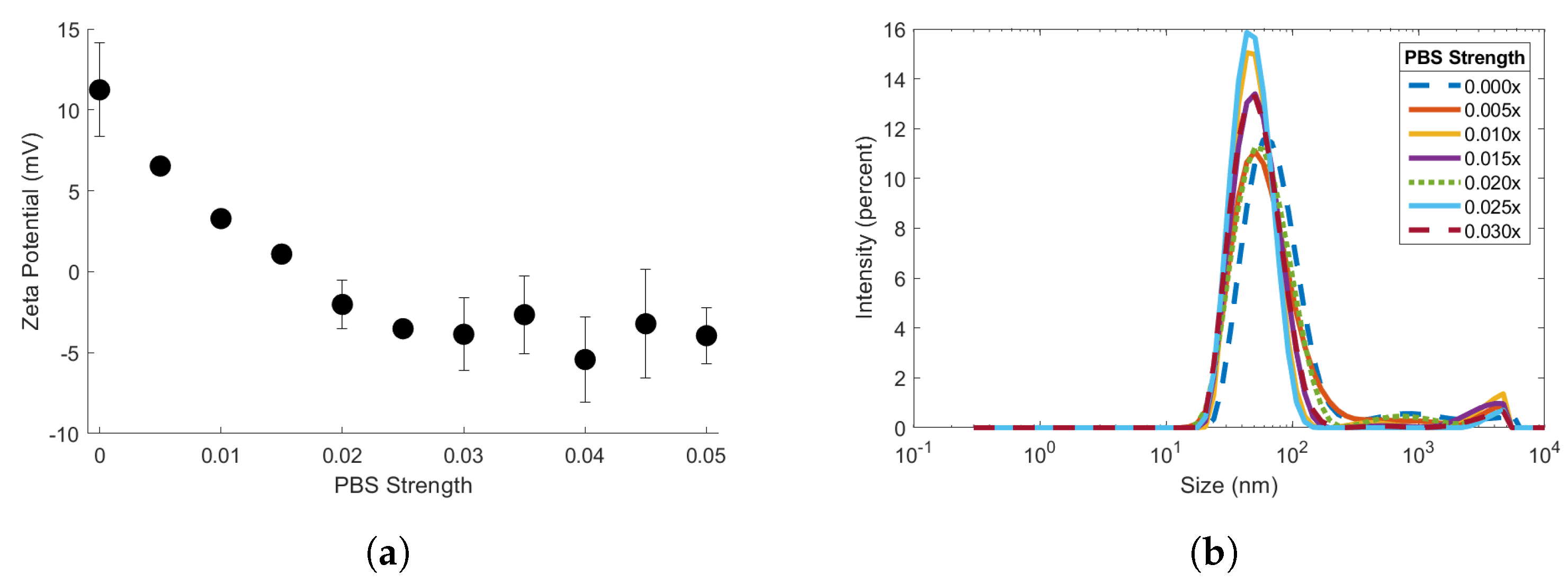
To determine the effects of electrolyte concentration on streptavidin induced biotin-MNP aggregation, streptavidin-response curves were obtained for the 90 biotin-MNPs in 1.00x, 0.50x, 0.03x, 0.015x, 0.005x, and 0.00x PBS solutions. We compared the slope, maximum , and range of the linear region of each streptavidin-response curve. There were no significant variations amongst linear region slope or range for the streptavidin-response curves prepared in 1.00x, 0.50x, 0.03x, 0.015x, and 0.005x PBS solutions. The biotin-MNPs formed aggregates with streptavidin in the 1.00x, 0.50x, 0.03x, 0.015x, and 0.005x PBS solutions, however the biotin-MNP were insusceptible to streptavidin induced aggregation in 0.00x PBS. The differences in biotin-MNP interparticle interactions were predicted with our xDLVO simulation. According to our simulation, interparticle interactions were dominated by the attractive van der Waals force in solutions with PBS strengths ranging from 0.005x to 1.00x PBS, while in 0.00x PBS interparticle interactions were dominated by the repulsive EDL force at particle separation distances between ∼23 nm and 50 nm. We have demonstrated that seemingly incremental changes in NP zeta potential and/or diluent electrolyte concentration can yield significant changes in NP-target interactions. For example, when suspended in 0.005x PBS (a relatively low electrolyte concentration diluent) the 90 biotin-MNPs had a zeta potential of 6.53 ± 0.72 mV and particle interactions were dominated by the attractive van der Waals force for separation distances (s) . However, after seemingly incremental changes in diluent to 0.00x PBS and 90 biotin-MNP zeta potential to 11.24 ± 3.88 mV, particle interactions were dominated by the repulsive EDL force for , resulting in no detectable levels of streptavidin induced biotin-MNP aggregation. Therefore, our study highlights the importance of xDLVO simulation during aggregation assay design to tune fMNP zeta potential to the diluent electrolyte concentration and pH for optimal target binding and colloidal stability. Concerning the effects of electrolyte concentration on the streptavidin-response curve maximum , no correlation was found.
and zeta potential measurements were obtained for the 90 biotin-MNPs in the 1.00x to 0.00x PBS series to assess the effects of electrolyte concentration on colloidal stability. Ninety biotin-MNP zeta potential decreased linearly in solutions with PBS strengths ranging from 0.025x to 0.00x PBS (
Figure 3a). While 90 biotin-MNP zeta potential was constant in solutions with PBS strengths ranging from 1.00x to 0.03x PBS (
Figure 3a). Ninety biotin-MNP
showed no trend with electrolyte concentration, therefore 90 biotin-MNP
is independent of electrolyte concentration in solutions with PBS strengths ≤ 1.00x. Our 90 biotin-MNP
data for solutions with PBS strengths < 1.00x is limited because it does not account for aggregate growth over time. Our
data was acquired directly after the 90 biotin-MNPs were mixed with each PBS solution, for those < 1.00x. Hence, our data reflect the initial 90 biotin-MNP size after exposure to diluents of varying PBS strength. Salt induced NP aggregates may grow in size over time depending on the salt concentration of the diluent [
40,
44,
45,
46].