Binned Data Provide Better Imputation of Missing Time Series Data from Wearables
Abstract
1. Introduction
2. Materials and Methods
2.1. Data
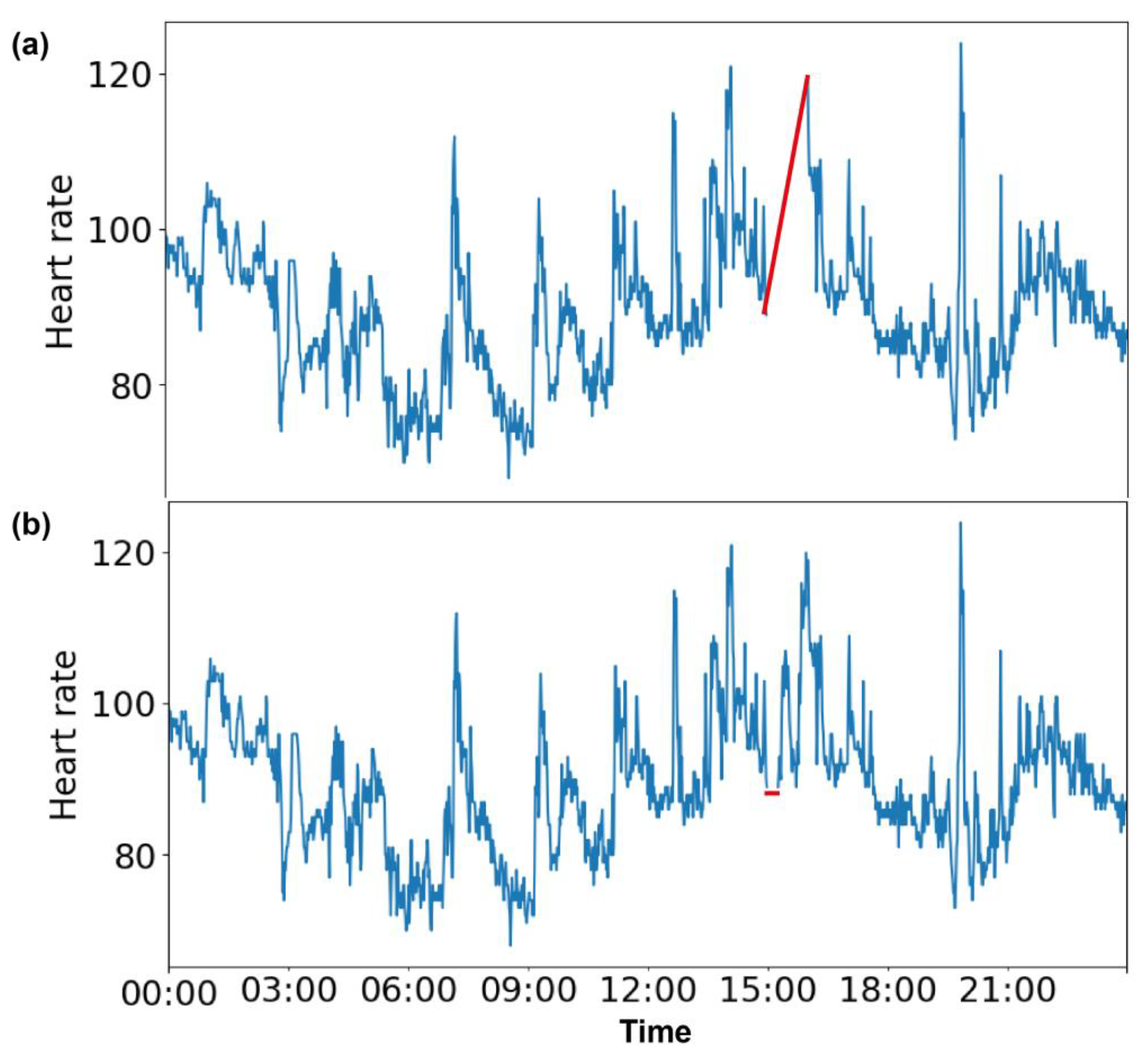
2.2. Missing Value Generation
2.3. Data Binning
2.4. Data Imputation
2.5. Performance Evaluation of Binning
3. Results
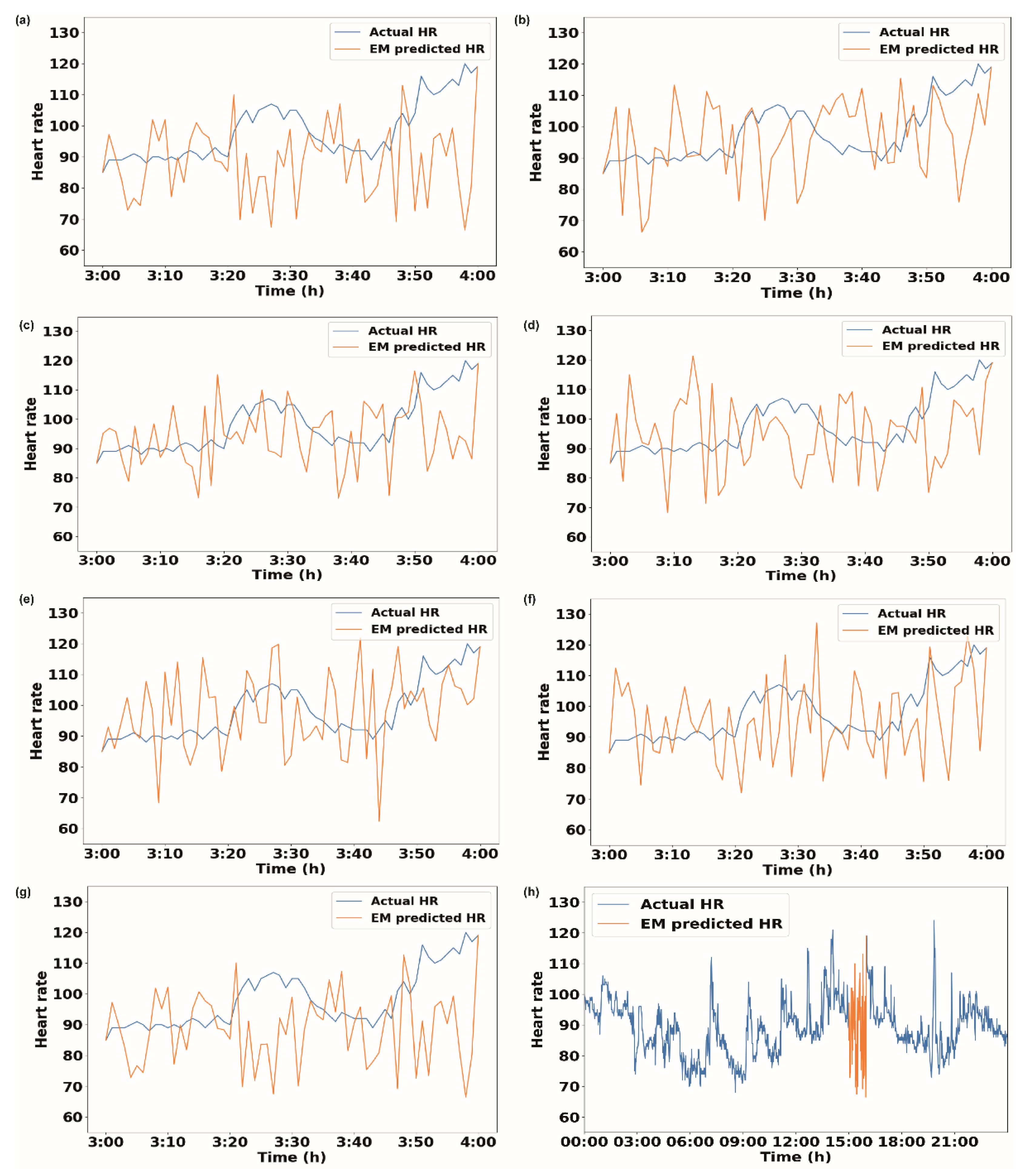
3.1. Imputation of 1 h of Missing Data
3.2. Imputation of 15 min of Missing Data
3.3. Quantitative Analysis
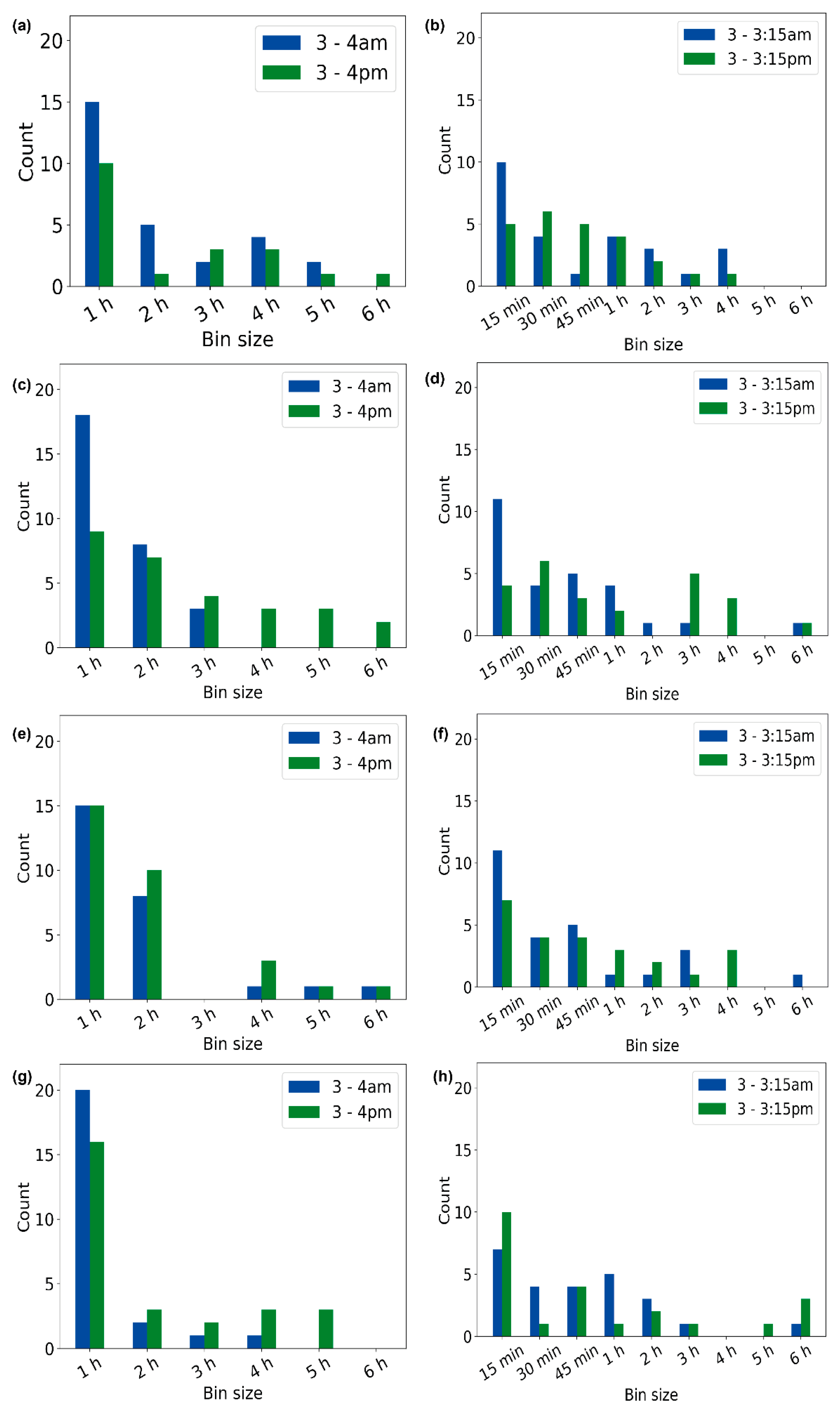
3.4. Optimal Bin Size
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Garcia-Duran, A.; West, R. Recursive Input and State Estimation: A General Framework for Learning from Time Series with Missing Data. In Proceedings of the 2021 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), ICASSP 2021, Toronto, ON, Canada, 6–11 June 2021; pp. 3535–3539. [Google Scholar] [CrossRef]
- Emmanuel, T.; Maupong, T.; Mpoeleng, D.; Semong, T.; Mphago, B.; Tabona, O. A survey on missing data in machine learning. J. Big Data 2021, 8, 140. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Mattingly, S.; Mirjafari, S.; Huang, C.; Chawla, N.V. Personalized Imputation on Wearable-Sensory Time Series via Knowledge Transfer. In Proceedings of the 29th ACM International Conference on Information & Knowledge Management, ACM: Virtual Event, Ireland, 19–23 October 2020; pp. 1625–1634. [Google Scholar]
- Bogl, M.; Filzmoser, P.; Gschwandtner, T.; Miksch, S.; Aigner, W.; Rind, A.; Lammarsch, T. Visually and Statistically Guided Imputation of Missing Values in Univariate Seasonal Time Series. In Proceedings of the 2015 IEEE Conference on Visual An-alytics Science and Technology (VAST), Chicago, IL, USA, 25–30 October 2015; IEEE: Chicago, IL, USA, 2015; pp. 189–190. [Google Scholar]
- Horton, N.J.; Lipsitz, S.R. Multiple Imputation in Practice: Comparison of Software Packages for Regression Models with Missing Variables. Am. Stat. 2001, 55, 244–254. [Google Scholar] [CrossRef]
- Jadhav, A.; Pramod, D.; Ramanathan, K. Comparison of Performance of Data Imputation Methods for Numeric Dataset. Appl. Artif. Intell. 2019, 33, 913–933. [Google Scholar] [CrossRef]
- Lakshminarayan, K.; Harp, S.A.; Goldman, R.; Samad, T. Imputation of Missing Data Using Machine Learning Techniques. In Proceedings of the 2nd International Conference on Knowledge Discovery and Data Mining (KDD-96), Portland, Oregon, 2–4 August 1996; Association for the Advancement of Artificial Intelligence (AAAI): Palo Alto, CA, USA, 1996; pp. 140–145. [Google Scholar]
- Norazian Ramli, M.N.; Yahaya, A.S.; Ramli, N.A.; Yusof, N.F.F.M.; Abdullah, M.M.A. Roles of Imputation Methods for Filling the Missing Values: A Review. Adv. Environ. Biol. 2013, 7, 3861–3869. [Google Scholar]
- Rubin, D.B.; Schenker, N. Multiple imputation in health-are databases: An overview and some applications. Stat. Med. 1991, 10, 585–598. [Google Scholar] [CrossRef]
- Koehler, E.; Brown, E.; Haneuse, S.J.-P.A. On the Assessment of Monte Carlo Error in Simulation-Based Statistical Analyses. Am. Stat. 2009, 63, 155–162. [Google Scholar] [CrossRef]
- Junger, W.; de Leon, A.P. Imputation of missing data in time series for air pollutants. Atmos. Environ. 2015, 102, 96–104. [Google Scholar] [CrossRef]
- Mir, A.A.; Kearfott, K.J.; Çelebi, F.V.; Rafique, M. Imputation by feature importance (IBFI): A methodology to envelop machine learning method for imputing missing patterns in time series data. PLoS ONE 2022, 17, e0262131. [Google Scholar] [CrossRef]
- Guk, K.; Han, G.; Lim, J.; Jeong, K.; Kang, T.; Lim, E.-K.; Jung, J. Evolution of Wearable Devices with Real-Time Disease Monitoring for Personalized Healthcare. Nanomaterials 2019, 9, 813. [Google Scholar] [CrossRef]
- Suwen, L.; Xian, W.; Gonzalo, M.; Chawla, N. Filling Missing Values on Wearable-Sensory Time Series Data. In Proceedings of the 2020 SIAM International Conference on Data Mining; Society for Industrial and Applied Mathematics: Philadelphia, PA, USA, 2020; pp. 46–54. [Google Scholar]
- Hartley, H.O.; Hocking, R.R. The Analysis of Incomplete Data. Biometrics 1971, 27, 783. [Google Scholar] [CrossRef]
- Meng, X.-L.; Rubin, D.B. Using EM to Obtain Asymptotic Variance-Covariance Matrices: The SEM Algorithm. J. Am. Stat. Assoc. 1991, 86, 899–909. [Google Scholar] [CrossRef]
- Malan, L.; Smuts, C.M.; Baumgartner, J.; Ricci, C. Missing data imputation via the expectation-maximization algorithm can improve principal component analysis aimed at deriving biomarker profiles and dietary patterns. Nutr. Res. 2020, 75, 67–76. [Google Scholar] [CrossRef]
- Feng, T.; Narayanan, S. Imputing Missing Data in Large-Scale Multivariate Biomedical Wearable Recordings Using Bidirectional Recurrent Neural Networks with Temporal Activation Regularization. In Proceedings of the 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC); IEEE: Berlin, Germany, 2019; pp. 2529–2534. [Google Scholar]
- Molenberghs, G.; Verbeke, G. Multiple Imputation and the Expectation-Maximization Algorithm. In Models for Discrete Longitudinal Data; Springer-Verlag: New York, NY, USA, 2005; pp. 511–529. ISBN 9780387251448. [Google Scholar]
- Platias, C.; Petasis, G. A Comparison of Machine Learning Methods for Data Imputation. In Proceedings of the 11th Hellenic Conference on Artificial Intelligence, Athens, Greece, 2–4 September 2020; ACM: Athens, Greece, 2020; pp. 150–159. [Google Scholar]
- Rao, A.R.; Reimherr, M. Modern multiple imputation with functional data. Stat 2021, 10, e331. [Google Scholar] [CrossRef]
- Templ, M.; Kowarik, A.; Filzmoser, P. Iterative stepwise regression imputation using standard and robust methods. Comput. Stat. Data Anal. 2011, 55, 2793–2806. [Google Scholar] [CrossRef]
- Sadhu, A.; Soni, R.; Mishra, M. Pattern-Based Comparative Analysis of Techniques for Missing Value Imputation. In Proceedings of the IEEE 5th International Conference on Computing Communication and Automation (ICCCA), Greater Noida, India, 30–31 October 2020; pp. 513–518. [Google Scholar]
- Zhang, S. Nearest neighbor selection for iteratively kNN imputation. J. Syst. Softw. 2012, 85, 2541–2552. [Google Scholar] [CrossRef]
- Tang, F.; Ishwaran, H. Random forest missing data algorithms. Stat. Anal. Data Min. ASA Data Sci. J. 2017, 10, 363–377. [Google Scholar] [CrossRef]
- Hong, S.; Lynn, H.S. Accuracy of random-forest-based imputation of missing data in the presence of non-normality, non-linearity, and interaction. BMC Med. Res. Methodol. 2020, 20, 199. [Google Scholar] [CrossRef]
- Kokla, M.; Virtanen, J.; Kolehmainen, M.; Paananen, J.; Hanhineva, K. Random forest-based imputation outperforms other methods for imputing LC-MS metabolomics data: A comparative study. BMC Bioinform. 2019, 20, 492. [Google Scholar] [CrossRef] [PubMed]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-Learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Lee, J.-H.; Lee, K.-H.; Kim, H.-J.; Youk, H.; Lee, H.-Y. Effective Prevention and Management Tools for Metabolic Syndrome Based on Digital Health-Based Lifestyle Interventions Using Healthcare Devices. Diagnostics 2022, 12, 1730. [Google Scholar] [CrossRef]
- Chakrabarti, S.; Biswas, N.; Jones, L.D.; Kesari, S.; Ashili, S. Smart Consumer Wearables as Digital Diagnostic Tools: A Review. Diagnostics 2022, 12, 2110. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, E.C.; Turley, J.P. Time series analysis as input for clinical predictive modeling: Modeling cardiac arrest in a pediatric ICU. Theor. Biol. Med. Model. 2011, 8, 40. [Google Scholar] [CrossRef]
- Lipton, Z.C.; Kale, D.; Wetzel, R. Directly Modeling Missing Data in Sequences with RNNs: Improved Classification of Clinical Time Series. In Proceedings of the 1st Machine Learning for Healthcare Conference; PMLR: Westminster, UK, 2016; pp. 253–270. [Google Scholar]
- Yozgatligil, C.; Aslan, S.; Iyigun, C.; Batmaz, I. Comparison of missing value imputation methods in time series: The case of Turkish meteorological data. Theor. Appl. Clim. 2013, 112, 143–167. [Google Scholar] [CrossRef]
- Boursalie, O.; Samavi, R.; Doyle, T.E. Evaluation Metrics for Deep Learning Imputation Models. In AI for Disease Surveillance and Pandemic Intelligence; Shaban-Nejad, A., Michalowski, M., Bianco, S., Eds.; Springer International Publishing: Cham, Switzerland, 2022; Volume 1013, pp. 309–322. ISBN 9783030930790. [Google Scholar]
- Christie, D.; Neill, S.P. Measuring and Observing the Ocean Renewable Energy Resource. In Comprehensive Renewable Energy; Elsevier: Amsterdam, The Netherlands, 2021; Volume 8, pp. 149–175. ISBN 9780128197349. [Google Scholar]
- Balasubramanian, S.; Meyyappan, T. Enhancing the Computational Intelligence of Smart Fog Gateway with Boundary-Constrained Dynamic Time Warping Based Imputation and Data Reduction. In Proceedings of the 3rd International Conference on Imaging Signal Processing and Communication (ICISPC), Singapore, 27–29 July 2019; IEEE: Singapore, 2019; pp. 15–23. [Google Scholar]
- Korkuć, P.; Arends, D.; Brockmann, G.A. Finding the Optimal Imputation Strategy for Small Cattle Populations. Front. Genet. 2019, 10, 52. [Google Scholar] [CrossRef] [PubMed]
- Støvring, H.; Kristiansen, I.S. Simple parametric survival analysis with anonymized register data: A cohort study with truncated and interval censored event and censoring times. BMC Res. Notes 2011, 4, 308. [Google Scholar] [CrossRef] [PubMed]
- Theodoridis, S. Bayesian Learning: Inference and the EM Algorithm. In Machine Learning; Academic Press: Cambridge, MA, USA, 2020; pp. 595–646. [Google Scholar] [CrossRef]
- Musil, C.M.; Warner, C.B.; Yobas, P.K.; Jones, S.L. A Comparison of Imputation Techniques for Handling Missing Data. West. J. Nurs. Res. 2002, 24, 815–829. [Google Scholar] [CrossRef]
- Ghaderpour, E.; Pagiatakis, S.D.; Hassan, Q.K. A Survey on Change Detection and Time Series Analysis with Applications. Appl. Sci. 2021, 11, 6141. [Google Scholar] [CrossRef]
- Ghaderpour, E. Multichannel antileakage least-squares spectral analysis for seismic data regularization beyond aliasing. Acta Geophys. 2019, 67, 1349–1363. [Google Scholar] [CrossRef]
- Ghaderpour, E.; Pagiatakis, S.D. Least-Squares Wavelet Analysis of Unequally Spaced and Non-stationary Time Series and Its Applications. Math. Geosci. 2017, 49, 819–844. [Google Scholar] [CrossRef]
- Rahman, S.A.; Huang, Y.; Claassen, J.; Heintzman, N.; Kleinberg, S. Combining Fourier and lagged k -nearest neighbor imputation for biomedical time series data. J. Biomed. Inform. 2015, 58, 198–207. [Google Scholar] [CrossRef] [PubMed]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chakrabarti, S.; Biswas, N.; Karnani, K.; Padul, V.; Jones, L.D.; Kesari, S.; Ashili, S. Binned Data Provide Better Imputation of Missing Time Series Data from Wearables. Sensors 2023, 23, 1454. https://doi.org/10.3390/s23031454
Chakrabarti S, Biswas N, Karnani K, Padul V, Jones LD, Kesari S, Ashili S. Binned Data Provide Better Imputation of Missing Time Series Data from Wearables. Sensors. 2023; 23(3):1454. https://doi.org/10.3390/s23031454
Chicago/Turabian StyleChakrabarti, Shweta, Nupur Biswas, Khushi Karnani, Vijay Padul, Lawrence D. Jones, Santosh Kesari, and Shashaanka Ashili. 2023. "Binned Data Provide Better Imputation of Missing Time Series Data from Wearables" Sensors 23, no. 3: 1454. https://doi.org/10.3390/s23031454
APA StyleChakrabarti, S., Biswas, N., Karnani, K., Padul, V., Jones, L. D., Kesari, S., & Ashili, S. (2023). Binned Data Provide Better Imputation of Missing Time Series Data from Wearables. Sensors, 23(3), 1454. https://doi.org/10.3390/s23031454








