Datasets for Automated Affect and Emotion Recognition from Cardiovascular Signals Using Artificial Intelligence— A Systematic Review
Abstract
:Simple Summary
Abstract
1. Introduction
Research Questions
- What are the datasets used for AAER from CV signals with AI techniques?
- What are the CV signals most often gathered in datasets for AAER?
- What were other signals are collected in analysed papers?
- What are the characteristics of the population in included studies?
- What instruments were used to assess emotion and affect in included papers?
- What confounders were taken into account in analysed papers?
- What devices were used to collect the signals in included studies?
- What stimuli are most often used for preparing datasets for AAER from CV signals?
- What are the characteristics of investigated stimuli?
- What is the credibility of included studies?
2. Methods
2.1. Eligibility Criteria, Protocol
2.2. Search Methods
2.3. Definitions
2.4. Data Collection
2.5. Quality Assessment
- Was the sample size pre-specified?
- Were eligibility criteria for the experiment provided?
- Were all inclusions and exclusions of the study participants appropriate?
- Was the measurement of the exposition clearly stated?
- Was the measurement of the outcome clearly stated?
- Did all participants receive a reference standard?
- Did participants receive the same reference standard?
- Were the confounders measured?
2.6. Analyses
3. Results
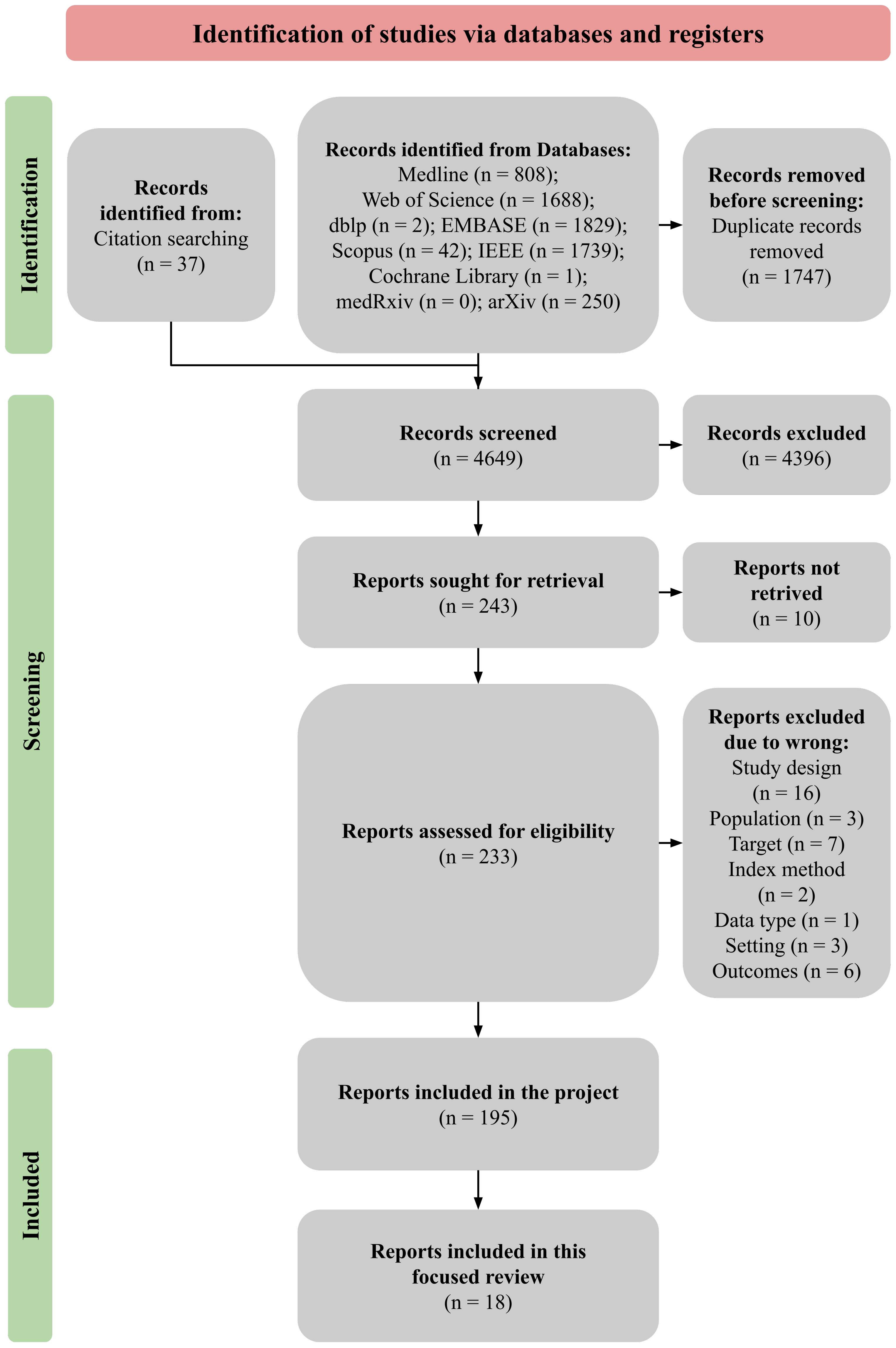
3.1. Included Studies
3.2. Experiments
3.3. Signals and Devices
3.4. Validation
3.5. Population
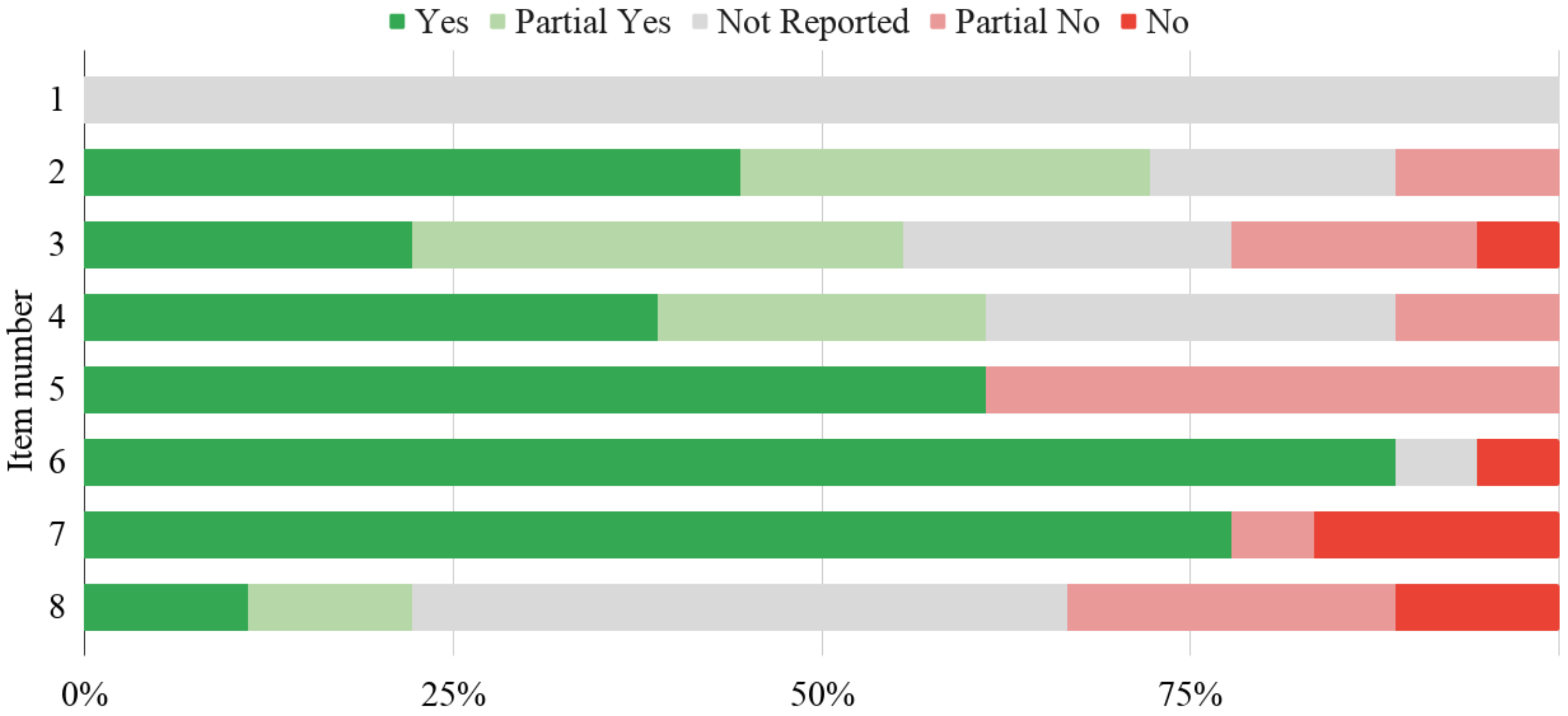
3.6. Credibility
3.7. Additional Analyses
4. Discussion
Strengths and Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAER | automated affect and/or emotion recognition |
| ABP | arterial blood pressure |
| AI | artificial intelligence |
| AMIGOS | a dataset for Affect, personality and Mood research on Individuals and GrOupS |
| ASCERTAIN | a multimodal databaASe for impliCit pERsonaliTy and Affect recognitIoN using commercial physiological sensors |
| AuDB-4 | AUgsburg Database of Biosignal 4 |
| BVP | blood volume pressure |
| CV | cardiovascular |
| DAG | Database for Affective Gaming |
| DECAF | MEG-based multimodal database for DECoding AFfective physiological responses |
| DEAP | Database for Emotion Analysis using Physiological signals |
| DL | deep learning |
| DTA | diagnostic test accuracy |
| ECG | electrocardiogram |
| EDA | electrodermal activity |
| EEG | electroencephalography |
| EMG | electromyography |
| emoFBVP | database of multimodal (Face, Body gesture, Voice and Physiological signals) recordings |
| EPQ | Eysenck Personality Questionnaire |
| ERS | Emotion Recognition Smartwatch |
| HCI | human–computer interaction |
| HR | heart rate |
| HRV | heart rate variability |
| ICP | intracranial pressure |
| ITMDER | IT Multimodal Dataset for Emotion Recognition |
| MAHNOB-HCI | Multimodal Analysis of Human NOnverbal Behaviour in real-world settings-Human-Computer Interaction |
| MAPD | Multi-subject Affective Physiological Database |
| MD | Mazeball Dataset |
| MEG | magnetoencephalography |
| ML | machine learning |
| MPED | Multi-modal Physiological Emotion Database for discrete emotion recognition |
| NB | naive Bayes |
| OSF | Open Science Framework |
| PANAS | Positive and Negative Affect Schedule |
| POX | pulse oximetry |
| PPG | photoplethysmogram |
| PPV | pulse pressure variation |
| PRISMA | Preferred Reporting Items for Systematic reviews and Meta-Analyses |
| PROBAST | Prediction model Risk Of Bias ASsessment Tool |
| QUADAS | QUality Assessment of Diagnostic Accuracy Studies |
| RF | random forest |
| RoB | Risk of Bias |
| SAM | Self-Assessment Manikin |
| TSST | Trier Social Stress Test |
| VRAD | Virtual Reality Affective Dataset |
| WESAD | WEarable Stress and Affect Detection |
Appendix A. Papers Included in This Focused Review
Appendix B. Papers Excluded
| Study ID | Reason of Excluding |
|---|---|
| [120] | Wrong study design |
| [121] | Wrong study design |
| [122] | Wrong study design |
| [123] | Wrong study design |
| [124] | Wrong study design |
| [125] | Wrong study design |
| [126] | Wrong study design |
| [127] | Wrong study design |
| [128] | Wrong study design |
| [129] | Wrong study design |
| [130] | Wrong study design |
| [131] | Wrong study design |
| [132] | Wrong study design |
| [133] | Wrong study design |
| [134] | Wrong study design |
| [135] | Wrong study design |
| [136] | Wrong population |
| [137] | Wrong population |
| [138] | Wrong population |
| [139] | Wrong target |
| [140] | Wrong target |
| [141] | Wrong target |
| [142] | Wrong target |
| [143] | Wrong target |
| [144] | Wrong target |
| [145] | Wrong target |
| [146] | Wrong index method |
| [147] | Wrong index method |
| [148] | Wrong type of data |
| [149] | Wrong setting |
| [150] | Wrong setting |
| [151] | Wrong setting |
| [152] | Wrong outcomes |
| [153] | Wrong outcomes |
| [154] | Wrong outcomes |
| [155] | Wrong outcomes |
| [156] | Wrong outcomes |
| [157] | Wrong outcomes |
Appendix C. Risk of Bias Tool
| Domain 1 | Review Authors’s Judgement | Criteria for Judgement | |
|---|---|---|---|
| Sample [74] | 1. Was the sample size prespecified? | Yes/partial yes | The experiment was preceded by calculating the minimum sample size, and the method used was adequate and well-described. |
| No/partial no | It is stated that the minimum sample size has not been calculated, or it has been calculated, but no details of the method used are provided. | ||
| Not reported | No sufficient information is provided in this regard. | ||
| Sample [74] | 2. Were eligibility criteria for the experiment provided? | Yes/partial yes | The criteria for inclusion in the experiment are specified. |
| No/partial no | The criteria for inclusion in the experiment were used, however not specified in the article. | ||
| Not reported | No sufficient information is provided in this regard. | ||
| Participants [73] | 3. Were all inclusions and exclusions of participants appropriate? | Yes/partial yes | The criteria for inclusion and exclusion are relevant to the aim of the study. Conditions that may affect the participant’s state or collected physiological signals and ability to recognise emotions were considered, including cardiovascular and mental disorders. |
| No/partial no | The established criteria for inclusion and exclusion are irrelevant to the aim of the study. | ||
| Not reported | No sufficient information is provided in this regard. | ||
| Measurement [74] | 4. Was the measurement of exposition clearly stated? | Yes/partial yes | The selection of stimuli is adequately justified in the context of eliciting emotions, e.g., selection from a standardised database, pilot studies. |
| No/partial no | The selection of stimuli was carried out based on inadequate criteria. | ||
| Not reported | No sufficient information is provided in this regard. | ||
| Measurement [74] | 5. Was the measurement of outcome clearly stated? | Yes/partial yes | The assessment tool used for emotions measurement is described in detail, adequate, and validated. |
| No/partial no | The assessment tool used for emotions measurement is not described, or the measurement method is inadequate, or not validated. | ||
| Not reported | No sufficient information is provided in this regard. | ||
| Flow and Timing [72] | 6. Did all participants receive a reference standard? | Yes/partial yes | Emotions were measured in all participants, and the measurement was performed after each stimulus. |
| No/partial no | Not all participants had their emotions measured. | ||
| Not reported | No sufficient information is provided in this regard. | ||
| Flow and Timing [72] | 7. Did participants receive the same reference standard? | Yes/partial yes | The same assessment standard was used in all participants who had their emotions measured |
| No/partial no | A different assessment standard was used in some of the participants to measure their emotions. | ||
| Not reported | No sufficient information is provided in this regard. | ||
| Control of confounders [74] | 8. Were the confounders measured? | Yes/partial yes | Adequate confounding factors were measured, and relevant justification is provided. |
| No/partial no | The control of confounding factors is not justified, or the measured factors are inadequate. | ||
| Not reported | No sufficient information is provided in regard to confounding factors. | ||
| Scenario 1: Overall quality (elicitation) Scenario 2: Overall quality (without judgement of 1. item) | High | All judgements are yes or partial yes. | |
| Low | At least one judgement is no or partial no. | ||
| Unclear | All judgements are yes or partial yes with at least one not reported. | ||
References
- Hacker, P. Teaching fairness to artificial intelligence: Existing and novel strategies against algorithmic discrimination under EU law. Common Mark. Law Rev. 2018, 55, 1143–1185. [Google Scholar]
- Butterworth, M. The ICO and artificial intelligence: The role of fairness in the GDPR framework. Comput. Law Secur. Rev. 2018, 34, 257–268. [Google Scholar] [CrossRef]
- Fan, X.; Yan, Y.; Wang, X.; Yan, H.; Li, Y.; Xie, L.; Yin, E. Emotion Recognition Measurement based on Physiological Signals. In Proceedings of the 2020 13th International Symposium on Computational Intelligence and Design (ISCID), Hangzhou, China, 12–13 December 2020; pp. 81–86. [Google Scholar]
- Xia, H.; Wu, J.; Shen, X.; Yang, F. The Application of Artificial Intelligence in Emotion Recognition. In Proceedings of the 2020 International Conference on Intelligent Computing and Human-Computer Interaction (ICHCI), Sanya, China, 4–6 December 2020; pp. 62–65. [Google Scholar]
- Jemioło, P.; Storman, D.; Giżycka, B.; Ligęza, A. Emotion elicitation with stimuli datasets in automatic affect recognition studies—Umbrella review. In Proceedings of the IFIP Conference on Human-Computer Interaction, Bari, Italy, 30 August–3 September 2021; Springer: Berlin/Heidelberg, Germany, 2021; pp. 248–269. [Google Scholar] [CrossRef]
- Ekman, P.; Friesen, W.V.; O’sullivan, M.; Chan, A.; Diacoyanni-Tarlatzis, I.; Heider, K.; Krause, R.; LeCompte, W.A.; Pitcairn, T.; Ricci-Bitti, P.E.; et al. Universals and cultural differences in the judgments of facial expressions of emotion. J. Personal. Soc. Psychol. 1987, 53, 712. [Google Scholar] [CrossRef]
- Barrett, L.F. How Emotions Are Made: The Secret Life of the Brain; Houghton Mifflin Harcourt: Boston, MA, USA, 2017. [Google Scholar]
- Plutchik, R. A general psychoevolutionary theory of emotion. In Theories of Emotion; Elsevier: Amsterdam, The Netherlands, 1980; pp. 3–33. [Google Scholar]
- Sarma, P.; Barma, S. Review on Stimuli Presentation for Affect Analysis Based on EEG. IEEE Access 2020, 8, 51991–52009. [Google Scholar] [CrossRef]
- Bradley, M.M.; Lang, P.J. Measuring emotion: The self-assessment manikin and the semantic differential. J. Behav. Ther. Exp. Psychiatry 1994, 25, 49–59. [Google Scholar] [CrossRef]
- Bandara, D.; Song, S.; Hirshfield, L.; Velipasalar, S. A more complete picture of emotion using electrocardiogram and electrodermal activity to complement cognitive data. In Proceedings of the International Conference on Augmented Cognition, Toronto, ON, Canada, 17–22 July 2016; Springer: Berlin/Heidelberg, Germany, 2016; pp. 287–298. [Google Scholar]
- Nardelli, M.; Greco, A.; Valenza, G.; Lanata, A.; Bailón, R.; Scilingo, E.P. A novel heart rate variability analysis using lagged poincaré plot: A study on hedonic visual elicitation. In Proceedings of the 2017 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Jeju, Korea, 11–15 July 2017; pp. 2300–2303. [Google Scholar]
- Jang, E.H.; Park, B.J.; Kim, S.H.; Chung, M.A.; Park, M.S.; Sohn, J.H. Emotion classification based on bio-signals emotion recognition using machine learning algorithms. In Proceedings of the 2014 International Conference on Information Science, Electronics and Electrical Engineering, Sapporo, Japan, 26–28 April 2014; Volume 3, pp. 1373–1376. [Google Scholar]
- Kołakowska, A.; Szwoch, W.; Szwoch, M. A review of emotion recognition methods based on data acquired via smartphone sensors. Sensors 2020, 20, 6367. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.; Wang, Z.; Yu, Z.; Guo, B. EmotionSense: Emotion recognition based on wearable wristband. In Proceedings of the 2018 IEEE SmartWorld, Ubiquitous Intelligence & Computing, Advanced & Trusted Computing, Scalable Computing & Communications, Cloud & Big Data Computing, Internet of People and Smart City Innovation (SmartWorld/SCALCOM/UIC/ATC/CBDCom/IOP/SCI), Guangzhou, China, 8–12 October 2018; pp. 346–355. [Google Scholar]
- Akalin, N.; Köse, H. Emotion recognition in valence-arousal scale by using physiological signals. In Proceedings of the 2018 26th Signal Processing and Communications Applications Conference (SIU), Izmir, Turkey, 2–5 May 2018; pp. 1–4. [Google Scholar]
- Nasoz, F.; Lisetti, C.L.; Vasilakos, A.V. Affectively intelligent and adaptive car interfaces. Inf. Sci. 2010, 180, 3817–3836. [Google Scholar] [CrossRef]
- Hsiao, P.W.; Chen, C.P. Effective attention mechanism in dynamic models for speech emotion recognition. In Proceedings of the 2018 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Calgary, AB, Canada, 15–20 April 2018; pp. 2526–2530. [Google Scholar]
- Shu, L.; Yu, Y.; Chen, W.; Hua, H.; Li, Q.; Jin, J.; Xu, X. Wearable emotion recognition using heart rate data from a smart bracelet. Sensors 2020, 20, 718. [Google Scholar] [CrossRef] [Green Version]
- Ragot, M.; Martin, N.; Em, S.; Pallamin, N.; Diverrez, J.M. Emotion recognition using physiological signals: Laboratory vs. wearable sensors. In In Proceedings of the International Conference on Applied Human Factors and Ergonomics, Los Angeles, CA, USA, 17–21 July 2017; Springer: Berlin/Heidelberg, Germany, 2017; pp. 15–22. [Google Scholar]
- Ali, H.; Hariharan, M.; Yaacob, S.; Adom, A.H. Facial emotion recognition using empirical mode decomposition. Expert Syst. Appl. 2015, 42, 1261–1277. [Google Scholar] [CrossRef]
- Liu, Z.T.; Wu, M.; Cao, W.H.; Mao, J.W.; Xu, J.P.; Tan, G.Z. Speech emotion recognition based on feature selection and extreme learning machine decision tree. Neurocomputing 2018, 273, 271–280. [Google Scholar] [CrossRef]
- Gómez-Zaragozá, L.; Marín-Morales, J.; Parra, E.; Guixeres, J.; Alcañiz, M. Speech Emotion Recognition from Social Media Voice Messages Recorded in the Wild. In Proceedings of the International Conference on Human-Computer Interaction, Copenhagen, Denmark, 19–24 July 2020; Springer: Berlin/Heidelberg, Germany, 2020; pp. 330–336. [Google Scholar]
- Abdullah, S.M.S.A.; Ameen, S.Y.A.; Sadeeq, M.A.; Zeebaree, S. Multimodal emotion recognition using deep learning. J. Appl. Sci. Technol. Trends 2021, 2, 52–58. [Google Scholar] [CrossRef]
- Harper, R.; Southern, J. A bayesian deep learning framework for end-to-end prediction of emotion from heartbeat. IEEE Trans. Affect. Comput. 2020. [Google Scholar] [CrossRef] [Green Version]
- Oh, S.; Lee, J.Y.; Kim, D.K. The design of CNN architectures for optimal six basic emotion classification using multiple physiological signals. Sensors 2020, 20, 866. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ravindran, A.S.; Nakagome, S.; Wickramasuriya, D.S.; Contreras-Vidal, J.L.; Faghih, R.T. Emotion recognition by point process characterization of heartbeat dynamics. In Proceedings of the 2019 IEEE Healthcare Innovations and Point of Care Technologies, (HI-POCT), Bethesda, MD, USA, 20–22 November 2019; pp. 13–16. [Google Scholar]
- Gadea, G.H.; Kreuder, A.; Stahlschmidt, C.; Schnieder, S.; Krajewski, J. Brute Force ECG Feature Extraction Applied on Discomfort Detection. In Proceedings of the International Conference on Information Technologies in Biomedicine, Kamień Śląski, Poland, 18–20 June 2018; Springer: Berlin/Heidelberg, Germany, 2018; pp. 365–376. [Google Scholar]
- Moharreri, S.; Dabanloo, N.J.; Maghooli, K. Detection of emotions induced by colors in compare of two nonlinear mapping of heart rate variability signal: Triangle and parabolic phase space (TPSM, PPSM). J. Med. Biol. Eng. 2019, 39, 665–681. [Google Scholar] [CrossRef]
- Basu, A.; Routray, A.; Shit, S.; Deb, A.K. Human emotion recognition from facial thermal image based on fused statistical feature and multi-class SVM. In Proceedings of the 2015 Annual IEEE India Conference (INDICON), New Delhi, India, 17–20 December 2015; pp. 1–5. [Google Scholar]
- Ferdinando, H.; Seppänen, T.; Alasaarela, E. Emotion recognition using neighborhood components analysis and ecg/hrv-based features. In Proceedings of the International Conference on Pattern Recognition Applications and Methods, Porto, Portugal, 24–26 February 2017; Springer: Berlin/Heidelberg, Germany, 2017; pp. 99–113. [Google Scholar]
- Higgins, J.P.; Thomas, J.; Chandler, J.; Cumpston, M.; Li, T.; Page, M.J.; Welch, V.A. Cochrane Handbook for Systematic Reviews of Interventions; John Wiley & Sons: Hoboken, NJ, USA, 2019. [Google Scholar]
- Mamica, M.; Kapłon, P.; Jemioło, P. EEG-Based Emotion Recognition Using Convolutional Neural Networks. In Proceedings of the International Conference on Conceptual Structures, ICCS. Krakow, Poland, 16–18 June 2021; Springer: Berlin/Heidelberg, Germany, 2021. [Google Scholar] [CrossRef]
- Saxena, A.; Khanna, A.; Gupta, D. Emotion recognition and detection methods: A comprehensive survey. J. Artif. Intell. Syst. 2020, 2, 53–79. [Google Scholar] [CrossRef]
- Resnick, B. More Social Science Studies just Failed to Replicate. Here’s Why This Is Good. Available online: Https://www.vox.com/science-and-health/2018/8/27/17761466/psychology-replication-crisis-nature-social-science (accessed on 15 February 2022).
- Maxwell, S.E.; Lau, M.Y.; Howard, G.S. Is psychology suffering from a replication crisis? What does “failure to replicate” really mean? Am. Psychol. 2015, 70, 487. [Google Scholar] [CrossRef]
- Kilkenny, M.F.; Robinson, K.M. Data Quality: “Garbage In—Garbage Out”. Health Inf. Manag. J. 2018, 47, 103–105. [Google Scholar] [CrossRef] [Green Version]
- Vidgen, B.; Derczynski, L. Directions in abusive language training data, a systematic review: Garbage in, garbage out. PLoS ONE 2020, 15, e0243300. [Google Scholar] [CrossRef]
- Stodden, V.; Seiler, J.; Ma, Z. An empirical analysis of journal policy effectiveness for computational reproducibility. Proc. Natl. Acad. Sci. USA 2018, 115, 2584–2589. [Google Scholar] [CrossRef] [Green Version]
- Fehr, J.; Heiland, J.; Himpe, C.; Saak, J. Best practices for replicability, reproducibility and reusability of computer-based experiments exemplified by model reduction software. arXiv 2016, arXiv:1607.01191. [Google Scholar]
- Mann, F.; von Walter, B.; Hess, T.; Wigand, R.T. Open access publishing in science. Commun. ACM 2009, 52, 135–139. [Google Scholar] [CrossRef]
- Kluyver, T.; Ragan-Kelley, B.; Pérez, F.; Granger, B.E.; Bussonnier, M.; Frederic, J.; Kelley, K.; Hamrick, J.B.; Grout, J.; Corlay, S.; et al. Jupyter Notebooks—A Publishing Format for Reproducible Computational Workflows; IOS Press: Amsterdam, The Netherlands, 2016. [Google Scholar]
- ReScicenceX. Available online: Http://rescience.org/x (accessed on 12 February 2022).
- ReScicence C. Available online: Https://rescience.github.io/ (accessed on 12 February 2022).
- Simmons, J.P.; Nelson, L.D.; Simonsohn, U. Pre-registration: Why and how. J. Consum. Psychol. 2021, 31, 151–162. [Google Scholar] [CrossRef]
- Nosek, B.A.; Ebersole, C.R.; DeHaven, A.C.; Mellor, D.T. The preregistration revolution. Proc. Natl. Acad. Sci. USA 2018, 115, 2600–2606. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Joffe, M.M.; Ten Have, T.R.; Feldman, H.I.; Kimmel, S.E. Model selection, confounder control, and marginal structural models: Review and new applications. Am. Stat. 2004, 58, 272–279. [Google Scholar] [CrossRef]
- Pourhoseingholi, M.A.; Baghestani, A.R.; Vahedi, M. How to control confounding effects by statistical analysis. Gastroenterol. Hepatol. Bed Bench 2012, 5, 79. [Google Scholar]
- Salminen, J.K.; Saarijärvi, S.; Äärelä, E.; Toikka, T.; Kauhanen, J. Prevalence of alexithymia and its association with sociodemographic variables in the general population of Finland. J. Psychosom. Res. 1999, 46, 75–82. [Google Scholar] [CrossRef]
- Greenaway, K.H.; Kalokerinos, E.K.; Williams, L.A. Context is everything (in emotion research). Soc. Personal. Psychol. Compass 2018, 12, e12393. [Google Scholar] [CrossRef]
- Saganowski, S.; Dutkowiak, A.; Dziadek, A.; Dzieżyc, M.; Komoszyńska, J.; Michalska, W.; Polak, A.; Ujma, M.; Kazienko, P. Emotion recognition using wearables: A systematic literature review-work-in-progress. In Proceedings of the 2020 IEEE International Conference on Pervasive Computing and Communications Workshops (PerCom Workshops), Austin, TX, USA, 23–27 March 2020; pp. 1–6. [Google Scholar]
- Peake, J.M.; Kerr, G.; Sullivan, J.P. A critical review of consumer wearables, mobile applications, and equipment for providing biofeedback, monitoring stress, and sleep in physically active populations. Front. Physiol. 2018, 9, 743. [Google Scholar] [CrossRef]
- Shu, L.; Xie, J.; Yang, M.; Li, Z.; Li, Z.; Liao, D.; Xu, X.; Yang, X. A review of emotion recognition using physiological signals. Sensors 2018, 18, 2074. [Google Scholar] [CrossRef] [Green Version]
- Kutt, K.; Nalepa, G.J.; Giżycka, B.; Jemiolo, P.; Adamczyk, M. Bandreader-a mobile application for data acquisition from wearable devices in affective computing experiments. In Proceedings of the 2018 11th International Conference on Human System Interaction (HSI), Gdansk, Poland, 4–6 July 2018; pp. 42–48. [Google Scholar] [CrossRef]
- Vallejo-Correa, P.; Monsalve-Pulido, J.; Tabares-Betancur, M. A systematic mapping review of context-aware analysis and its approach to mobile learning and ubiquitous learning processes. Comput. Sci. Rev. 2021, 39, 100335. [Google Scholar] [CrossRef]
- Bardram, J.E.; Matic, A. A decade of ubiquitous computing research in mental health. IEEE Pervasive Comput. 2020, 19, 62–72. [Google Scholar] [CrossRef]
- Cárdenas-Robledo, L.A.; Peña-Ayala, A. Ubiquitous learning: A systematic review. Telemat. Inform. 2018, 35, 1097–1132. [Google Scholar] [CrossRef]
- Paré, G.; Kitsiou, S. Methods for literature reviews. In Handbook of eHealth Evaluation: An Evidence-Based Approach [Internet]; University of Victoria: Victoria, BC, Canada, 2017. [Google Scholar]
- Okoli, C. A guide to conducting a standalone systematic literature review. Commun. Assoc. Inf. Syst. 2015, 37, 43. [Google Scholar] [CrossRef] [Green Version]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G. PRISMA 2009 flow diagram. PRISMA Statement 2009, 6, 97. [Google Scholar]
- Liberati, A.; Altman, D.; Tetzlaff, J.; Mulrow, C.; Gøtzsche, P.; Ioannidis, J.; Clarke, M.; Devereaux, P. The PRISMA statement for reporting systematic and meta-analyses of studies that evaluate interventions. PLoS Med. 2009, 6, 1–28. [Google Scholar]
- Jemioło, P.; Storman, D.; Mamica, M.; Szymkowski, M.; Orzechowski, P.; Dranka, W. Emotion Recognition from Cardiovascular Signals Using Artificial Intelligence—A Systematic Review. Protocol Registration. Available online: Https://osf.io/nj7ut (accessed on 12 February 2022).
- Jemioło, P.; Storman, D.; Mamica, M.; Szymkowski, M.; Orzechowski, P. Automated Affect and Emotion Recognition from Cardiovascular Signals—A Systematic Overview of the Field. In Proceedings of the Hawaii International Conference on System Sciences, Maui, HI, USA, 4–7 January 2022; ScholarSpace: Maui, HI, USA, 2022; pp. 4047–4056. [Google Scholar] [CrossRef]
- Jemioło, P.; Storman, D.; Mamica, M.; Szymkowski, M.; Orzechowski, P.; Dranka, W. Emotion Recognition from Cardiovascular Signals Using Artificial Intelligence—A Systematic Review. Supplementary Information. Available online: Https://osf.io/kzj8y/ (accessed on 12 February 2022).
- Konar, A.; Chakraborty, A. Emotion Recognition: A Pattern Analysis Approach; John Wiley & Sons: New York, NY, USA, 2015; pp. 1–45. [Google Scholar]
- Kim, K.H.; Bang, S.W.; Kim, S.R. Emotion recognition system using short-term monitoring of physiological signals. Med. Biol. Eng. Comput. 2004, 42, 419–427. [Google Scholar] [CrossRef]
- Copeland, B. Artificial Intelligence: Definition, Examples, and Applications. Available online: Https://www.britannica.com/technology/artificial-intelligence (accessed on 10 May 2021).
- Craik, A.; He, Y.; Contreras-Vidal, J.L. Deep learning for electroencephalogram (EEG) classification tasks: A review. J. Neural Eng. 2019, 16, 031001. [Google Scholar]
- Botchkarev, A. Performance metrics (error measures) in machine learning regression, forecasting and prognostics: Properties and typology. arXiv 2018, arXiv:1809.03006. [Google Scholar]
- McNames, J.; Aboy, M. Statistical modeling of cardiovascular signals and parameter estimation based on the extended Kalman filter. IEEE Trans. Biomed. Eng. 2007, 55, 119–129. [Google Scholar] [CrossRef] [Green Version]
- Ouzzani, M.; Hammady, H.; Fedorowicz, Z.; Elmagarmid, A. Rayyan—A web and mobile app for systematic reviews. Syst. Rev. 2016, 5, 210. [Google Scholar] [CrossRef] [Green Version]
- Whiting, P.F.; Rutjes, A.W.; Westwood, M.E.; Mallett, S.; Deeks, J.J.; Reitsma, J.B.; Leeflang, M.M.; Sterne, J.A.; Bossuyt, P.M. QUADAS-2: A revised tool for the quality assessment of diagnostic accuracy studies. Ann. Intern. Med. 2011, 155, 529–536. [Google Scholar]
- Wolff, R.F.; Moons, K.G.; Riley, R.D.; Whiting, P.F.; Westwood, M.; Collins, G.S.; Reitsma, J.B.; Kleijnen, J.; Mallett, S. PROBAST: A tool to assess the risk of bias and applicability of prediction model studies. Ann. Intern. Med. 2019, 170, 51–58. [Google Scholar] [CrossRef] [Green Version]
- Benton, M.J.; Hutchins, A.M.; Dawes, J.J. Effect of menstrual cycle on resting metabolism: A systematic review and meta-analysis. PLoS ONE 2020, 15, e0236025. [Google Scholar] [CrossRef] [PubMed]
- Koelstra, S.; Muhl, C.; Soleymani, M.; Lee, J.S.; Yazdani, A.; Ebrahimi, T.; Pun, T.; Nijholt, A.; Patras, I. Deap: A database for emotion analysis; using physiological signals. IEEE Trans. Affect. Comput. 2011, 3, 18–31. [Google Scholar] [CrossRef] [Green Version]
- Soleymani, M.; Lichtenauer, J.; Pun, T.; Pantic, M. A multimodal database for affect recognition and implicit tagging. IEEE Trans. Affect. Comput. 2011, 3, 42–55. [Google Scholar] [CrossRef] [Green Version]
- Abadi, M.K.; Subramanian, R.; Kia, S.M.; Avesani, P.; Patras, I.; Sebe, N. DECAF: MEG-based multimodal database for decoding affective physiological responses. IEEE Trans. Affect. Comput. 2015, 6, 209–222. [Google Scholar] [CrossRef]
- Correa, J.A.M.; Abadi, M.K.; Sebe, N.; Patras, I. Amigos: A dataset for affect, personality and mood research on individuals and groups. IEEE Trans. Affect. Comput. 2021, 12, 479–493. [Google Scholar] [CrossRef] [Green Version]
- Subramanian, R.; Wache, J.; Abadi, M.K.; Vieriu, R.L.; Winkler, S.; Sebe, N. ASCERTAIN: Emotion and personality recognition using commercial sensors. IEEE Trans. Affect. Comput. 2016, 9, 147–160. [Google Scholar] [CrossRef]
- Kim, J.; André, E. Emotion recognition based on physiological changes in music listening. IEEE Trans. Pattern Anal. Mach. Intell. 2008, 30, 2067–2083. [Google Scholar] [CrossRef]
- Quiroz, J.C.; Geangu, E.; Yong, M.H. Emotion recognition using smart watch sensor data: Mixed-design study. JMIR Ment. Health 2018, 5, e10153. [Google Scholar] [CrossRef] [Green Version]
- Pinto, J. Exploring Physiological Multimodality for Emotional Assessment; Instituto Superior Técnico (IST): Lisboa, Portugal, 2019. [Google Scholar]
- Yang, W.; Rifqi, M.; Marsala, C.; Pinna, A. Physiological-based emotion detection and recognition in a video game context. In Proceedings of the 2018 International Joint Conference on Neural Networks (IJCNN), Rio de Janeiro, Brazil, 8–13 July 2018; pp. 1–8. [Google Scholar]
- Gupta, R.; Khomami Abadi, M.; Cárdenes Cabré, J.A.; Morreale, F.; Falk, T.H.; Sebe, N. A quality adaptive multimodal affect recognition system for user-centric multimedia indexing. In Proceedings of the 2016 ACM on International Conference on Multimedia Retrieval, New York, NY, USA, 6–9 June 2016; pp. 317–320. [Google Scholar]
- Marín-Morales, J.; Higuera-Trujillo, J.L.; Greco, A.; Guixeres, J.; Llinares, C.; Scilingo, E.P.; Alcañiz, M.; Valenza, G. Affective computing ual reality: Emotion recognition from brain and heartbeat dynamics using wearable sensors. Sci. Rep. 2018, 8, 13657. [Google Scholar]
- Hsu, Y.L.; Wang, J.S.; Chiang, W.C.; Hung, C.H. Automatic ECG-based emotion recognition in music listening. IEEE Trans. Affect. Comput. 2017, 11, 85–99. [Google Scholar] [CrossRef]
- Schmidt, P.; Reiss, A.; Duerichen, R.; Marberger, C.; Van Laerhoven, K. Introducing wesad, a multimodal dataset for wearable stress and affect detection. In Proceedings of the 20th ACM International Conference on Multimodal Interaction, Boulder, CO, USA, 16–20 October 2018; pp. 400–408. [Google Scholar]
- Song, T.; Zheng, W.; Lu, C.; Zong, Y.; Zhang, X.; Cui, Z. MPED: A multi-modal physiological emotion database for discrete emotion recognition. IEEE Access 2019, 7, 12177–12191. [Google Scholar] [CrossRef]
- Ranganathan, H.; Chakraborty, S.; Panchanathan, S. Multimodal emotion recognition using deep learning architectures. In Proceedings of the 2016 IEEE Winter Conference on Applications of Computer Vision (WACV), Lake Placid, NY, USA, 7–10 March 2016; pp. 1–9. [Google Scholar]
- Katsigiannis, S.; Ramzan, N. DREAMER: A database for emotion recognition through EEG and ECG signals from wireless low-cost off-the-shelf devices. IEEE J. Biomed. Health Inform. 2017, 22, 98–107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, W.; Liu, G.; Wen, W. MAPD: A Multi-subject Affective Physiological Database. In Proceedings of the 2014 Seventh International Symposium on Computational Intelligence and Design, Hangzhou, China, 13–14 December 2014; Volume 2, pp. 585–589. [Google Scholar]
- Yannakakis, G.N.; Martínez, H.P.; Jhala, A. Towards affective camera control in games. User Model. User-Adapt. Interact. 2010, 20, 313–340. [Google Scholar] [CrossRef] [Green Version]
- McInnes, M.D.; Moher, D.; Thombs, B.D.; McGrath, T.A.; Bossuyt, P.M.; Clifford, T.; Cohen, J.F.; Deeks, J.J.; Gatsonis, C.; Hooft, L.; et al. Preferred reporting items for a systematic review and meta-analysis of diagnostic test accuracy studies: The PRISMA-DTA statement. JAMA 2018, 319, 388–396. [Google Scholar] [CrossRef] [PubMed]
- Wierzbicka, A. Defining emotion concepts. Cogn. Sci. 1992, 16, 539–581. [Google Scholar] [CrossRef]
- Wierzbicka, A. Emotion, language, and cultural scripts. In Emotion and Culture: Empirical Studies of Mutual Influence; American Psychological Association: Washington, DC, USA, 1994; pp. 133–196. [Google Scholar]
- Cook, D.A.; Levinson, A.J.; Garside, S. Method and reporting quality in health professions education research: A systematic review. Med. Educ. 2011, 45, 227–238. [Google Scholar] [CrossRef]
- Wijasena, H.Z.; Ferdiana, R.; Wibirama, S. A Survey of Emotion Recognition using Physiological Signal in Wearable Devices. In Proceedings of the 2021 International Conference on Artificial Intelligence and Mechatronics Systems (AIMS), Bandung, Indonesia, 28–30 April 2021; pp. 1–6. [Google Scholar]
- Saganowski, S.; Kazienko, P.; Dziezyc, M.; Jakimow, P.; Komoszynska, J.; Michalska, W.; Dutkowiak, A.; Polak, A.; Dziadek, A.; Ujma, M. Consumer Wearables and Affective Computing for Wellbeing Support. In Proceedings of the MobiQuitous 2020—17th EAI International Conference on Mobile and Ubiquitous Systems: Computing, Networking and Services, Darmstadt, Germany, 7–9 December 2020. [Google Scholar]
- Schmidt, P.; Reiss, A.; Dürichen, R.; Laerhoven, K.V. Wearable-Based Affect Recognition—A Review. Sensors 2019, 19, 4079. [Google Scholar] [CrossRef] [Green Version]
- Merone, M.; Soda, P.; Sansone, M.; Sansone, C. ECG databases for biometric systems: A systematic review. Expert Syst. Appl. 2017, 67, 189–202. [Google Scholar] [CrossRef]
- Da Silva, H.P.; Lourenço, A.; Fred, A.; Raposo, N.; Aires-de Sousa, M. Check Your Biosignals Here: A new dataset for off-the-person ECG biometrics. Comput. Methods Programs Biomed. 2014, 113, 503–514. [Google Scholar] [CrossRef]
- Hong, S.; Zhou, Y.; Shang, J.; Xiao, C.; Sun, J. Opportunities and challenges of deep learning methods for electrocardiogram data: A systematic review. Comput. Biol. Med. 2020, 122, 103801. [Google Scholar] [CrossRef]
- Santamaria-Granados, L.; Munoz-Organero, M.; Ramirez-Gonzalez, G.; Abdulhay, E.; Arunkumar, N. Using deep convolutional neural network for emotion detection on a physiological signals dataset (AMIGOS). IEEE Access 2018, 7, 57–67. [Google Scholar] [CrossRef]
- Soroush, M.Z.; Maghooli, K.; Setarehdan, S.K.; Nasrabadi, A.M. A review on EEG signals based emotion recognition. Int. Clin. Neurosci. J. 2017, 4, 118. [Google Scholar]
- Suhaimi, N.S.; Mountstephens, J.; Teo, J. EEG-based emotion recognition: A state-of-the-art review of current trends and opportunities. Comput. Intell. Neurosci. 2020, 2020, 8875426. [Google Scholar] [CrossRef] [PubMed]
- Wagh, K.P.; Vasanth, K. Electroencephalograph (EEG) based emotion recognition system: A review. In Innovations in Electronics and Communication Engineering; Springer: Berlin/Heidelberg, Germany, 2019; pp. 37–59. [Google Scholar]
- Egger, M.; Ley, M.; Hanke, S. Emotion recognition from physiological signal analysis: A review. Electron. Notes Theor. Comput. Sci. 2019, 343, 35–55. [Google Scholar] [CrossRef]
- Jerritta, S.; Murugappan, M.; Nagarajan, R.; Wan, K. Physiological signals based human emotion recognition: A review. In Proceedings of the 2011 IEEE 7th International Colloquium on Signal Processing and its Applications, Penang, Malaysia, 4–6 March 2011; pp. 410–415. [Google Scholar]
- Ali, M.; Mosa, A.H.; Al Machot, F.; Kyamakya, K. Emotion recognition involving physiological and speech signals: A comprehensive review. In Recent Advances in Nonlinear Dynamics and Synchronization; Springer: Berlin/Heidelberg, Germany, 2018; pp. 287–302. [Google Scholar]
- Szwoch, W. Using physiological signals for emotion recognition. In Proceedings of the 2013 6th International Conference on Human System Interactions (HSI), Sopot, Poland, 6–8 June 2013; pp. 556–561. [Google Scholar]
- Callejas-Cuervo, M.; Martínez-Tejada, L.A.; Alarcón-Aldana, A.C. Emotion recognition techniques using physiological signals and video games-Systematic review. Rev. Fac. Ing. 2017, 26, 19–28. [Google Scholar] [CrossRef] [Green Version]
- Marechal, C.; Mikolajewski, D.; Tyburek, K.; Prokopowicz, P.; Bougueroua, L.; Ancourt, C.; Wegrzyn-Wolska, K. Survey on AI-Based Multimodal Methods for Emotion Detection. In High-Performance Modelling and Simulation for Big Data Application; Springer: Berlin/Heidelberg, Germany, 2018; pp. 307–324. [Google Scholar]
- Picard, R.W. Affective Computing; MIT Press: Cambridge, MA, USA, 1997. [Google Scholar]
- Amira, T.; Dan, I.; Az-eddine, B.; Ngo, H.H.; Said, G.; Katarzyna, W.W. Monitoring chronic disease at home using connected devices. In Proceedings of the 2018 13th Annual Conference on System of Systems Engineering (SoSE), Paris, France, 19–22 June 2018; pp. 400–407. [Google Scholar]
- Khan, K.S.; Kunz, R.; Kleijnen, J.; Antes, G. Five steps to conducting a systematic review. J. R. Soc. Med. 2003, 96, 118–121. [Google Scholar] [CrossRef]
- Enhancing the Quality and Transparency of Health Research. Available online: Https://www.equator-network.org/ (accessed on 12 February 2022).
- Giżycka, B.; Jemioło, P.; Domarecki, S.; Świder, K.; Wiśniewski, M.; Mielczarek, Ł. A Thin Light Blue Line—Towards Balancing Educational and Recreational Values of Serious Games. In Proceedings of the 3rd Workshop on Affective Computing and Context Awareness in Ambient Intelligence, Cartagena, Spain, 11–12 November 2019; Technical University of Aachen: Aachen, Germany, 2019.
- Jemioło, P.; Giżycka, B.; Nalepa, G.J. Prototypes of arcade games enabling affective interaction. In Proceedings of the International Conference on Artificial Intelligence and Soft Computing, Zakopane, Poland, 16–20 June 2019; Springer: Berlin/Heidelberg, Germany, 2019; pp. 553–563. [Google Scholar] [CrossRef]
- Nalepa, G.J.; Kutt, K.; Giżycka, B.; Jemioło, P.; Bobek, S. Analysis and use of the emotional context with wearable devices for games and intelligent assistants. Sensors 2019, 19, 2509. [Google Scholar] [CrossRef] [Green Version]
- Benovoy, M.; Cooperstock, J.R.; Deitcher, J. Biosignals analysis and its application in a performance setting. In Proceedings of the International Conference on Bio-Inspired Systems and Signal Processing, Madeira, Portugal, 3–4 January 2008; pp. 253–258. [Google Scholar]
- Mera, K.; Ichimura, T. Emotion analyzing method using physiological state. In Proceedings of the International Conference on Knowledge-Based and Intelligent Information and Engineering Systems, Wellington, New Zealand, 20–25 September 2004; Springer: Berlin/Heidelberg, Germany, 2004; pp. 195–201. [Google Scholar]
- Wang, Y.; Mo, J. Emotion feature selection from physiological signals using tabu search. In Proceedings of the 2013 25th Chinese Control and Decision Conference (CCDC), Guiyang, China, 25–27 May 2013; pp. 3148–3150. [Google Scholar]
- Guendil, Z.; Lachiri, Z.; Maaoui, C.; Pruski, A. Emotion recognition from physiological signals using fusion of wavelet based features. In Proceedings of the 2015 7th International Conference on Modelling, Identification and Control (ICMIC), Sousse, Tunisia, 18–20 December 2015; pp. 1–6. [Google Scholar]
- Zong, C.; Chetouani, M. Hilbert-Huang transform based physiological signals analysis for emotion recognition. In Proceedings of the 2009 IEEE International Symposium on Signal Processing and Information Technology (ISSPIT), Ajman, United Arab Emirates, 14–17 December 2009; pp. 334–339. [Google Scholar]
- Joesph, C.; Rajeswari, A.; Premalatha, B.; Balapriya, C. Implementation of physiological signal based emotion recognition algorithm. In Proceedings of the 2020 IEEE 36th International Conference on Data Engineering (ICDE), Dallas, TX, USA, 20–24 April 2020; pp. 2075–2079. [Google Scholar]
- Leon, E.; Clarke, G.; Sepulveda, F.; Callaghan, V. Neural network-based improvement in class separation of physiological signals for emotion classification. In Proceedings of the IEEE Conference on Cybernetics and Intelligent Systems, Singapore, 1–3 December 2004; Volume 2, pp. 724–728. [Google Scholar]
- Siow, S.C.; Loo, C.K.; Tan, A.W.; Liew, W.S. Adaptive Resonance Associative Memory for multi-channel emotion recognition. In Proceedings of the 2010 IEEE EMBS Conference on Biomedical Engineering and Sciences (IECBES), Kuala Lumpur, Malaysia, 30 November—2 December 2010; pp. 359–363. [Google Scholar]
- Perez-Rosero, M.S.; Rezaei, B.; Akcakaya, M.; Ostadabbas, S. Decoding emotional experiences through physiological signal processing. In Proceedings of the 2017 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), New Orleans, LA, USA, 5–9 March 2017; pp. 881–885. [Google Scholar]
- Sokolova, M.V.; Fernández-Caballero, A.; López, M.T.; Martínez-Rodrigo, A.; Zangróniz, R.; Pastor, J.M. A distributed architecture for multimodal emotion identification. In Proceedings of the 13th International Conference on Practical Applications of Agents and Multi-Agent Systems, Salamanca, Spain, 3–5 June 2015; Springer: Berlin/Heidelberg, Germany, 2015; pp. 125–132. [Google Scholar]
- Shirahama, K.; Grzegorzek, M. Emotion recognition based on physiological sensor data using codebook approach. In Proceedings of the Conference of Information Technologies in Biomedicine, Kamien Slaski, Poland, 20–22 June 2016; Springer: Berlin/Heidelberg, Germany, 2016; pp. 27–39. [Google Scholar]
- Gong, P.; Ma, H.T.; Wang, Y. Emotion recognition based on the multiple physiological signals. In Proceedings of the 2016 IEEE International Conference on Real-time Computing and Robotics (RCAR), Angkor Wat, Cambodia, 6–10 June 2016; pp. 140–143. [Google Scholar]
- Jain, M.; Saini, S.; Kant, V. A hybrid approach to emotion recognition system using multi-discriminant analysis & k-nearest neighbour. In Proceedings of the 2017 International Conference on Advances in Computing, Communications and Informatics (ICACCI), Udupi, India, 13–16 September 2017; pp. 2251–2256. [Google Scholar]
- Guendil, Z.; Lachiri, Z.; Maaoui, C.; Pruski, A. Multiresolution framework for emotion sensing in physiological signals. In Proceedings of the 2016 2nd International Conference on Advanced Technologies for Signal and Image Processing (ATSIP), Monastir, Tunisia, 21–23 March 2016; pp. 793–797. [Google Scholar]
- Wong, W.M.; Tan, A.W.; Loo, C.K.; Liew, W.S. PSO optimization of synergetic neural classifier for multichannel emotion recognition. In Proceedings of the 2010 Second World Congress on Nature and Biologically Inspired Computing (NaBIC), Kitakyushu, Japan, 15–17 December 2010; pp. 316–321. [Google Scholar]
- Guo, X. Study of emotion recognition based on electrocardiogram and RBF neural network. Procedia Eng. 2011, 15, 2408–2412. [Google Scholar]
- Zhu, H.; Han, G.; Shu, L.; Zhao, H. ArvaNet: Deep Recurrent Architecture for PPG-Based Negative Mental-State Monitoring. IEEE Trans. Comput. Soc. Syst. 2020, 8, 179–190. [Google Scholar] [CrossRef]
- Wu, C.H.; Kuo, B.C.; Tzeng, G.H. Factor analysis as the feature selection method in an Emotion Norm Database. In Proceedings of the Asian Conference on Intelligent Information and Database Systems, Bangkok, Thailand, 7–9 April 2014; Springer: Berlin/Heidelberg, Germany, 2014; pp. 332–341. [Google Scholar]
- Akbulut, F.P.; Perros, H.G.; Shahzad, M. Bimodal affect recognition based on autoregressive hidden Markov models from physiological signals. Comput. Methods Programs Biomed. 2020, 195, 105571. [Google Scholar]
- Takahashi, M.; Kubo, O.; Kitamura, M.; Yoshikawa, H. Neural network for human cognitive state estimation. In Proceedings of the IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS’94), Munich, Germany, 12–16 September 1994; Volume 3, pp. 2176–2183. [Google Scholar]
- Gao, Y.; Barreto, A.; Adjouadi, M. An affective sensing approach through pupil diameter processing and SVM classification. Biomed. Sci. Instrum. 2010, 46, 326–330. [Google Scholar] [PubMed]
- Mohino-Herranz, I.; Gil-Pita, R.; Rosa-Zurera, M.; Seoane, F. Activity recognition using wearable physiological measurements: Selection of features from a comprehensive literature study. Sensors 2019, 19, 5524. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bonarini, A.; Costa, F.; Garbarino, M.; Matteucci, M.; Romero, M.; Tognetti, S. Affective videogames: The problem of wearability and comfort. In Proceedings of the International Conference on Human-Computer Interaction, Orlando, FL, USA, 9–14 July 2011; Springer: Berlin/Heidelberg, Germany, 2011; pp. 649–658. [Google Scholar]
- Alqahtani, F.; Katsigiannis, S.; Ramzan, N. ECG-based affective computing for difficulty level prediction in intelligent tutoring systems. In Proceedings of the 2019 UK/China Emerging Technologies (UCET), Glasgow, UK, 21–22 August 2019; pp. 1–4. [Google Scholar]
- Xu, J.; Hu, Z.; Zou, J.; Bi, A. Intelligent emotion detection method based on deep learning in medical and health data. IEEE Access 2019, 8, 3802–3811. [Google Scholar] [CrossRef]
- Wendt, C.; Popp, M.; Karg, M.; Kuhnlenz, K. Physiology and HRI: Recognition of over-and underchallenge. In Proceedings of the RO-MAN 2008—The 17th IEEE International Symposium on Robot and Human Interactive Communication, Munich, Germany, 1–3 August 2008; pp. 448–452. [Google Scholar]
- Omata, M.; Moriwaki, K.; Mao, X.; Kanuka, D.; Imamiya, A. Affective rendering: Visual effect animations for affecting user arousal. In Proceedings of the 2012 International Conference on Multimedia Computing and Systems, Tangiers, Morocco, 10–12 May 2012; pp. 737–742. [Google Scholar]
- van den Broek, E.L.; Schut, M.H.; Westerink, J.H.; Tuinenbreijer, K. Unobtrusive sensing of emotions (USE). J. Ambient Intell. Smart Environ. 2009, 1, 287–299. [Google Scholar] [CrossRef]
- Quiroz, J.C.; Yong, M.H.; Geangu, E. Emotion-recognition using smart watch accelerometer data: Preliminary findings. In Proceedings of the 2017 ACM International Joint Conference on Pervasive and Ubiquitous Computing, Maui, HI, USA, 11–15 September 2017; pp. 805–812. [Google Scholar]
- Althobaiti, T.; Katsigiannis, S.; West, D.; Bronte-Stewart, M.; Ramzan, N. Affect detection for human-horse interaction. In Proceedings of the 2018 21st Saudi Computer Society National Computer Conference (NCC), Riyadh, Saudi Arabia, 25–26 April 2018; pp. 1–6. [Google Scholar]
- Dobbins, C.; Fairclough, S. Detecting negative emotions during real-life driving via dynamically labelled physiological data. In Proceedings of the 2018 IEEE International Conference on Pervasive Computing and Communications Workshops (PerCom Workshops), Athens, Greece, 19–23 March 2018; pp. 830–835. [Google Scholar]
- Jo, Y.; Lee, H.; Cho, A.; Whang, M. Emotion Recognition Through Cardiovascular Response in Daily Life Using KNN Classifier. In Advances in Computer Science and Ubiquitous Computing; Springer: Berlin/Heidelberg, Germany, 2017; pp. 1451–1456. [Google Scholar]
- Hamdi, H.; Richard, P.; Suteau, A.; Allain, P. Emotion assessment for affective computing based on physiological responses. In Proceedings of the 2012 IEEE International Conference on Fuzzy Systems, Brisbane, QLD, Australia, 10–15 June 2012; pp. 1–8. [Google Scholar]
- Moghimi, S.; Chau, T.; Guerguerian, A.M. Using prefrontal cortex near-infrared spectroscopy and autonomic nervous system activity for identifying music-induced emotions. In Proceedings of the 2013 6th International IEEE/EMBS Conference on Neural Engineering (NER), San Diego, CA, USA, 6–8 November 2013; pp. 1283–1286. [Google Scholar]
- Zhang, Z.; Tanaka, E. Affective computing using clustering method for mapping human’s emotion. In Proceedings of the 2017 IEEE International Conference on Advanced Intelligent Mechatronics (AIM), Munich, Germany, 3–7 July 2017; pp. 235–240. [Google Scholar]
- Reinerman-Jones, L.; Taylor, G.; Cosenzo, K.; Lackey, S. Analysis of multiple physiological sensor data. In Proceedings of the International Conference on Foundations of Augmented Cognition, Orlando, FL, USA, 9–14 July 2011; Springer: Berlin/Heidelberg, Germany, 2011; pp. 112–119. [Google Scholar]
- Roza, V.; Postolache, O.; Groza, V.; Pereira, J.D. Emotions Assessment on Simulated Flights. In Proceedings of the 2019 IEEE International Symposium on Medical Measurements and Applications (MeMeA), Istanbul, Turkey, 26–28 June 2019; pp. 1–6. [Google Scholar]
- Savran, A.; Ciftci, K.; Chanel, G.; Mota, J.; Viet, L.; Sankur, B.; Akarun, L.; Caplier, A.; Rombaut, M. Emotion detection in the loop from brain signals and facial images. In Proceedings of the eNTERFACE 2006 Workshop, Dubrovnik, Croatia, 17 July—11 August 2006. [Google Scholar]
| Variable (No. of Experiments Available for Calculations) | No (%) Mean (Range) | |
|---|---|---|
| Type of stimuli (21) | ||
| Video (music, movie, ads) | 11 (52.38) | |
| Audio (music excerpts) | 4 (19.05) | |
| Game (FIFA 2016, Maze-Ball) | 2 (9.52) | |
| Virtual Reality (videos, scenes) | 2 (9.52) | |
| Self elicitation (actors) | 1 (4.76) | |
| Mixed (TSST 1, video and meditation in one experiment) | 1 (4.76) | |
| Length of stimuli [seconds] (17) | 304.60 (32–1200) | |
| No. of stimuli in dataset (20) | 27.70 (4–144) | |
| No. of elicited emotions [classes] (18) | 6.06 (3–23) | |
| Variable (No. of Datasets Available for Calculations) 1 | No (%) Mean (Range) | |
|---|---|---|
| Used devices (16) | ||
| Shimmer 2R | 3 (18.75) | |
| BIOPAC MP150 | 3 (18.75) | |
| Biosemi ActiveTwo | 2 (12.50) | |
| NeXus-10 | 1 (6.25) | |
| ProComp Infiniti | 1 (6.25) | |
| BIOPAC BioNomadix | 1 (6.25) | |
| BItalino | 1 (6.25) | |
| RespiBAN Professional | 1 (6.25) | |
| B-Alert x10 | 1 (6.25) | |
| Empatica E4 | 1 (6.25) | |
| Polar H7 | 1 (6.25) | |
| Zephyr BioHarness | 1 (6.25) | |
| IOM Biofeedback | 1 (6.25) | |
| CV 2 signals recorded (18) | ||
| ECG 3 | 15 (83.33) | |
| HR 4 | 3 (16.67) | |
| BVP 5 | 3 (16.67) | |
| PPG 6 | 1 (5.56) | |
| Sampling frequency [Hz] (12) | 543.31 (32–2048) | |
| Length of baseline recording [seconds] (7) | 292.14 (5–1200) | |
| Variable (No. of Experiments Available for Calculations) | No. (%) Mean (Range) |
|---|---|
| Participating people (21) | 916 43.62 (3–250) |
| Eligible people (20) | 812 40.60 (3–250) |
| Age (18) | 23.8 (17–47) |
| Percentage of females (16) | 45.13 (0–86) |
| Ethnicity (4) | |
| Chinese | 2 (9.52) |
| European | 2 (9.52) |
| Study ID | RoB Item 1 | Overall Quality 2 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | Scenario 1 | Scenario 2 | |
| [75] | NR | PY | PY | Y | Y | Y | Y | PN | Low | Low |
| [76] | NR | PY | PY | Y | PN | Y | Y | NR | Low | Low |
| [77] | NR | NR | NR | Y | PN | Y | Y | PN | Low | Low |
| [78] | NR | Y | PY | Y | Y | Y | Y | PY | Unclear | High |
| [79] | NR | PY | PN | PY | PN | Y | Y | N | Low | Low |
| [80] | NR | PN | N | PN | PN | N | N | PY | Low | Low |
| [81] | NR | Y | PN | PY | Y | Y | Y | PN | Low | Low |
| [82] | NR | Y | Y | PY | Y | Y | Y | PN | Low | Low |
| [83] | NR | NR | NR | PN | PN | Y | Y | NR | Low | Low |
| [84] | NR | NR | NR | NR | PN | Y | Y | N | Low | Low |
| [85] | NR | Y | Y | Y | Y | Y | Y | Y | Unclear | High |
| [86] | NR | PY | PY | NR | Y | Y | Y | NR | Unclear | Unclear |
| [87] | NR | Y | Y | NR | Y | Y | N | NR | Low | Low |
| [88] | NR | Y | Y | Y | Y | Y | PN | Y | Low | Low |
| [89] | NR | Y | PN | NR | PN | Y | Y | NR | Low | Low |
| [90] | NR | PY | PY | PY | Y | Y | Y | NR | Unclear | Unclear |
| [91] | NR | PN | PY | NR | Y | Y | Y | NR | Low | Low |
| [92] | NR | Y | NR | Y | Y | NR | N | NR | Low | Low |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jemioło, P.; Storman, D.; Mamica, M.; Szymkowski, M.; Żabicka, W.; Wojtaszek-Główka, M.; Ligęza, A. Datasets for Automated Affect and Emotion Recognition from Cardiovascular Signals Using Artificial Intelligence— A Systematic Review. Sensors 2022, 22, 2538. https://doi.org/10.3390/s22072538
Jemioło P, Storman D, Mamica M, Szymkowski M, Żabicka W, Wojtaszek-Główka M, Ligęza A. Datasets for Automated Affect and Emotion Recognition from Cardiovascular Signals Using Artificial Intelligence— A Systematic Review. Sensors. 2022; 22(7):2538. https://doi.org/10.3390/s22072538
Chicago/Turabian StyleJemioło, Paweł, Dawid Storman, Maria Mamica, Mateusz Szymkowski, Wioletta Żabicka, Magdalena Wojtaszek-Główka, and Antoni Ligęza. 2022. "Datasets for Automated Affect and Emotion Recognition from Cardiovascular Signals Using Artificial Intelligence— A Systematic Review" Sensors 22, no. 7: 2538. https://doi.org/10.3390/s22072538
APA StyleJemioło, P., Storman, D., Mamica, M., Szymkowski, M., Żabicka, W., Wojtaszek-Główka, M., & Ligęza, A. (2022). Datasets for Automated Affect and Emotion Recognition from Cardiovascular Signals Using Artificial Intelligence— A Systematic Review. Sensors, 22(7), 2538. https://doi.org/10.3390/s22072538






