Biosensors to Monitor Cell Activity in 3D Hydrogel-Based Tissue Models
Abstract
1. Introduction
1.1. Standard 2D Cultures and In Vivo Models
1.2. 3D Tissue Models
3D Hydrogel-Based Tissue Models
1.3. Importance of Integrating Biosensors with 3D Hydrogel-Based Tissue Models
1.4. Biosensors
- -
- biosensors to monitor the cellular metabolism, where glucose and oxygen consumptions as well as pH surrounding levels are monitored;
- -
- impedance biosensors, where the variations of cellular electrical properties are revealed;
- -
- biosensors to detect the secretion of specific molecules modulated by cell activity.
2. Biosensors to Monitor the Cellular Metabolism
2.1. PH Biosensors
2.1.1. Electrochemical Biosensors
2.1.2. ISFETs and LAPs
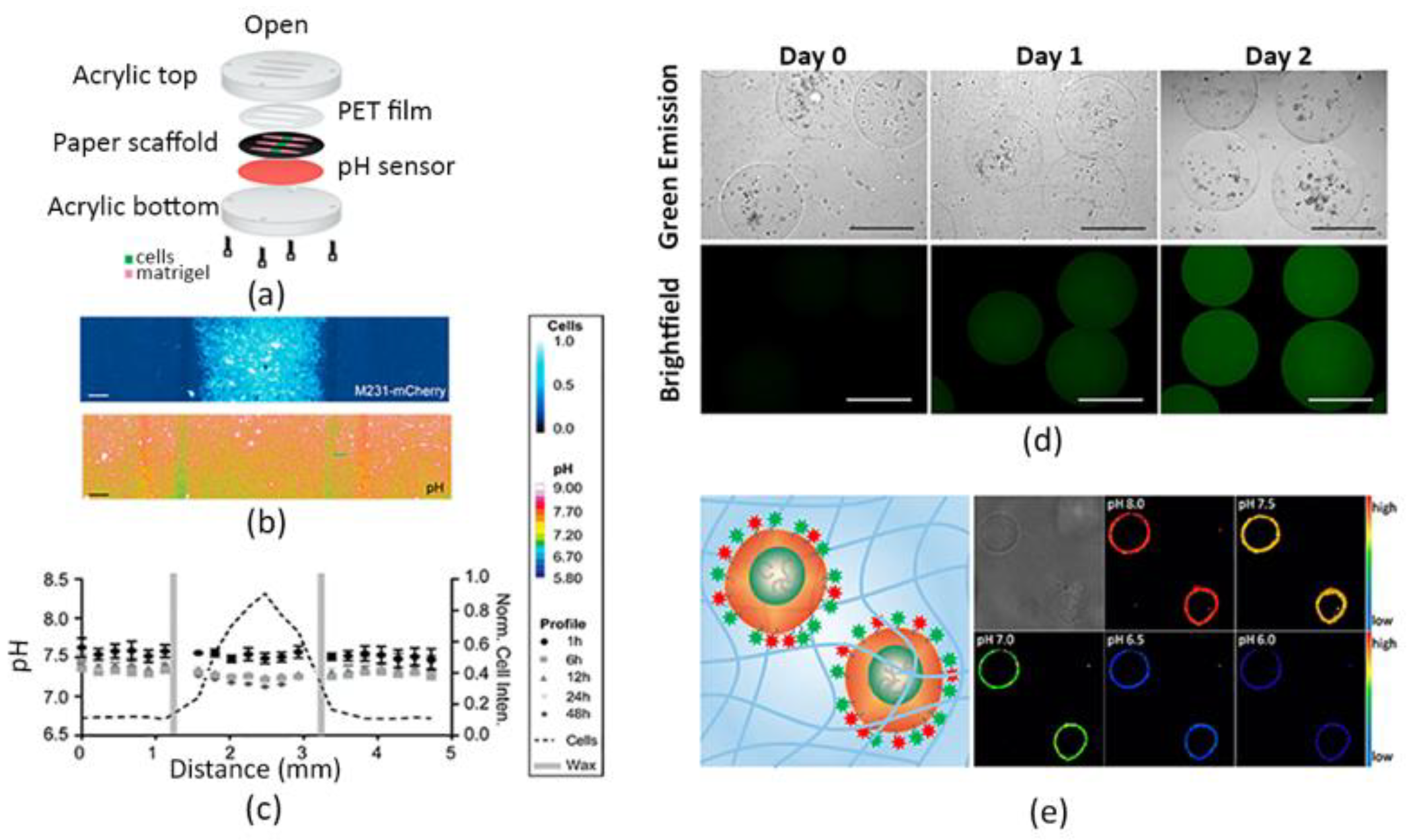
2.1.3. Optical Biosensors
2.2. Glucose Biosensors
2.2.1. Electrochemical Biosensors
2.2.2. Optical Biosensors
2.3. Oxygen Biosensors
2.3.1. Electrochemical Biosensors
2.3.2. Optical Biosensors
3. Biosensors to Monitor the Cellular Behavior: Impedance Biosensors
3.1. Working Principle
3.2. Impedance Biosensors
4. Biosensors to Detect Secreted Molecules
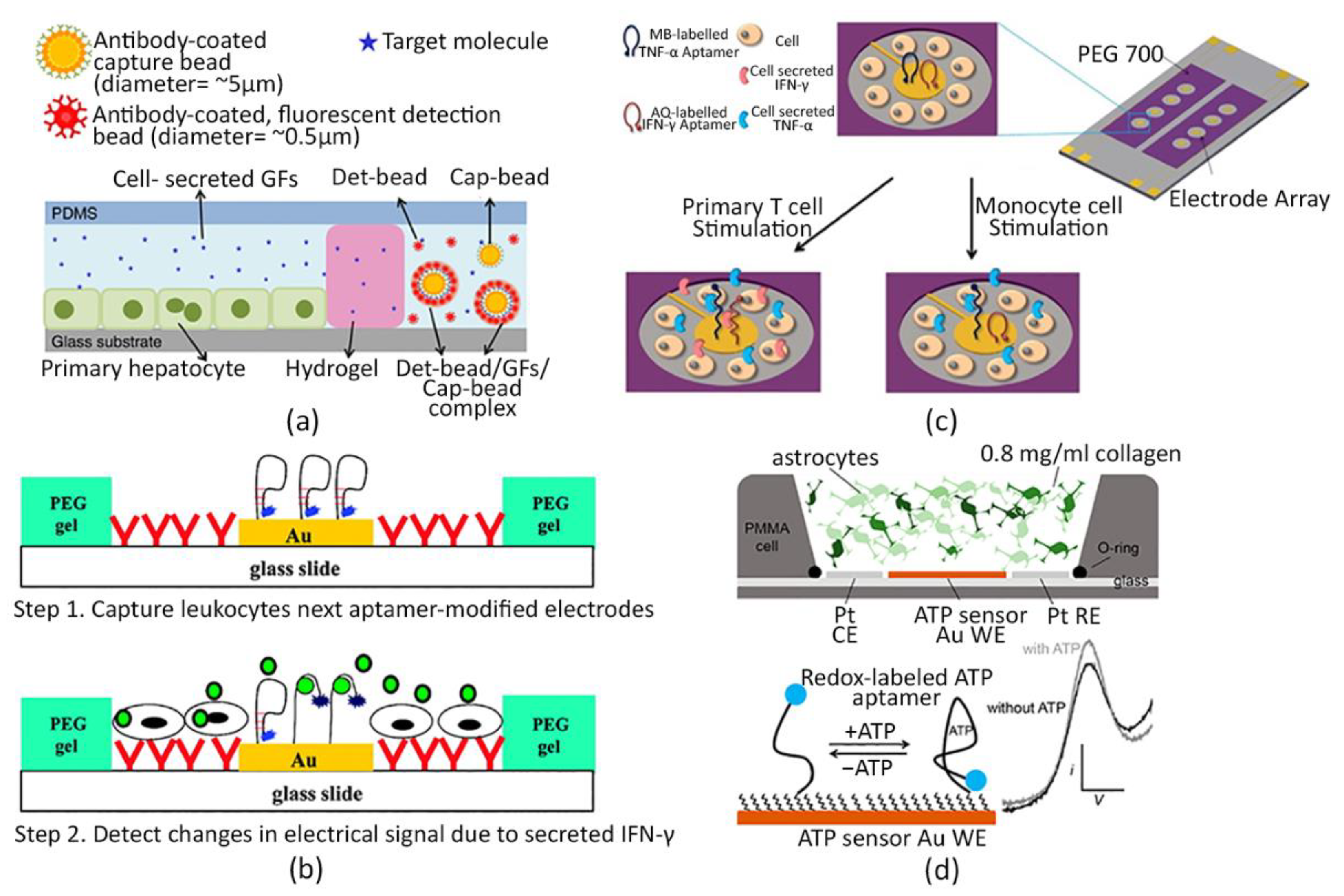
4.1. Antibody-Based Biosensors
4.2. Aptamer-Based Biosensors
5. Conclusions and Future Outlook
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hoarau-Véchot, J.; Rafii, A.; Touboul, C.; Pasquier, J. Halfway between 2D and animal models: Are 3D cultures the ideal tool to study cancer-microenvironment interactions? Int. J. Mol. Sci. 2018, 19, 181. [Google Scholar] [CrossRef]
- Das, V.; Bruzzese, F.; Konečný, P.; Iannelli, F.; Budillon, A.; Hajdúch, M. Pathophysiologically relevant in vitro tumor models for drug screening. Drug Discov. Today 2015, 20, 848–855. [Google Scholar] [CrossRef] [PubMed]
- Kapałczyńska, M.; Kolenda, T.; Przybyła, W.; Zajączkowska, M.; Teresiak, A.; Filas, V.; Ibbs, M.; Bliźniak, R.; Łuczewski, Ł.; Lamperska, K. 2D and 3D cell cultures–a comparison of different types of cancer cell cultures. Arch. Med. Sci. 2018, 14, 910–919. [Google Scholar] [CrossRef]
- Leong, D.T.; Ng, K.W. Probing the relevance of 3D cancer models in nanomedicine research. Adv. Drug Deliv. Rev. 2014, 79, 95–106. [Google Scholar] [CrossRef] [PubMed]
- Cukierman, E.; Pankov, R.; Stevens, D.R.; Yamada, K.M. Taking cell-matrix adhesions to the third dimension. Science 2001, 294, 1708–1712. [Google Scholar] [CrossRef]
- MJ, B.; Aylin, R.; Saira, I.M. Tissue architecture: The ultimate regulator of breast epithelial function. Curr. Opin. Cell Biol. 2003, 15, 753–762. [Google Scholar]
- Brien, L.E.O.; Zegers, M.M.P.; Mostov, K.E. Culture Models. Group 2002, 3, 1–7. [Google Scholar]
- Mulhall, H.J.; Hughes, M.P.; Kazmi, B.; Lewis, M.P.; Labeed, F.H. Epithelial cancer cells exhibit different electrical properties when cultured in 2D and 3D environments. Biochim. Biophys. Acta–Gen. Subj. 2013, 1830, 5136–5141. [Google Scholar] [CrossRef]
- Tay, C.Y.; Pal, M.; Yu, H.; Leong, W.S.; Tan, N.S.; Ng, K.W.; Venkatraman, S.; Boey, F.; Leong, D.T.; Tan, L.P. Bio-inspired micropatterned platform to steer stem cell differentiation. Small 2011, 7, 1416–1421. [Google Scholar] [CrossRef] [PubMed]
- Tay, C.Y.; Yu, H.; Pal, M.; Leong, W.S.; Tan, N.S.; Ng, K.W.; Leong, D.T.; Tan, L.P. Micropatterned matrix directs differentiation of human mesenchymal stem cells towards myocardial lineage. Exp. Cell Res. 2010, 316, 1159–1168. [Google Scholar] [CrossRef]
- Knight, E.; Przyborski, S. Advances in 3D cell culture technologies enabling tissue-like structures to be created in vitro. J. Anat. 2015, 227, 746–756. [Google Scholar] [CrossRef] [PubMed]
- Souza, A.G.; Ferreira, I.C. Advances in Cell Culture: More than a Century after Cultivating Cells. J. Biotechnol. Biomater. 2016, 6, 2–5. [Google Scholar] [CrossRef]
- Duval, K.; Grover, H.; Han, L.H.; Mou, Y.; Pegoraro, A.F.; Fredberg, J.; Chen, Z. Modeling physiological events in 2D vs. 3D cell culture. Physiology 2017, 32, 266–277. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Farach-Carson, M.C.; Jia, X. Three-dimensional in vitro tumor models for cancer research and drug evaluation. Biotechnol. Adv. 2014, 32, 1256–1268. [Google Scholar] [CrossRef]
- Fedi, A.; Vitale, C.; Ponschin, G.; Ayehunie, S.; Fato, M.; Scaglione, S. In vitro models replicating the human intestinal epithelium for absorption and metabolism studies: A systematic review. J. Control. Release 2021, 335, 247–268. [Google Scholar] [CrossRef]
- Edmondson, R.; Broglie, J.J.; Adcock, A.F.; Yang, L. Three-dimensional cell culture systems and their applications in drug discovery and cell-based biosensors. Assay Drug Dev. Technol. 2014, 12, 207–218. [Google Scholar] [CrossRef]
- Marrella, A.; Dondero, A.; Aiello, M.; Casu, B.; Olive, D.; Regis, S.; Bottino, C.; Caluori, G.; Castriconi, R.; Scaglione, S. Cell-laden hydrogel as a clinical-relevant 3D model for analyzing neuroblastoma growth, immunophenotype and susceptibility to therapies. Front. Immunol. 2019, 10, 1876. [Google Scholar] [CrossRef]
- Bhadriraju, K.; Chen, C.S. Engineering cellular microenvironments to improve cell-based drug testing. Drug Discov. Today 2002, 7, 612–620. [Google Scholar] [CrossRef]
- Meigs, L.; Smirnova, L.; Rovida, C.; Leist, M.; Hartung, T. Animal testing and its alternatives—The most important omics is economics. ALTEX 2018, 35, 275–305. [Google Scholar] [CrossRef]
- Romero, L.; Vela, J.M. Alternative Models in Drug Discovery and Development Part I: In Silico and In Vitro Models. Vivo Model. Drug Discov. 2014, 9783527333, 27–58. [Google Scholar] [CrossRef]
- Cheluvappa, R.; Scowen, P.; Eri, R. Ethics of animal research in human disease remediation, its institutional teaching; and alternatives to animal experimentation. Pharmacol. Res. Perspect. 2017, 5, e00332. [Google Scholar] [CrossRef] [PubMed]
- Liguori, G.R.; Jeronimus, B.F.; De Aquinas Liguori, T.T.; Moreira, L.F.P.; Harmsen, M.C. Ethical Issues in the Use of Animal Models for Tissue Engineering: Reflections on Legal Aspects, Moral Theory, Three Rs Strategies, and Harm’Benefit Analysis. Tissue Eng. Part C Methods 2017, 23, 850–862. [Google Scholar] [CrossRef] [PubMed]
- Leist, M.; Hasiwa, N.; Rovida, C.; Daneshian, M.; Basketter, D.; Kimber, I.; Clewell, H.; Gocht, T.; Goldberg, A.; Busquet, F.; et al. Consensus report on the future of animal-free systemic toxicity testing. ALTEX 2014, 31, 341–356. [Google Scholar] [CrossRef] [PubMed]
- Adler, S.; Basketter, D.; Creton, S.; Pelkonen, O.; Van Benthem, J.; Zuang, V.; Andersen, K.E.; Angers-Loustau, A.; Aptula, A.; Bal-Price, A.; et al. Alternative (non-animal) methods for cosmetics testing: Current status and future prospects-2010. Arch. Toxicol. 2011, 85, 367–485. [Google Scholar] [CrossRef] [PubMed]
- Hartung, T.; Blaauboer, B.J.; Bosgra, S.; Carney, E.; Coenen, J.; Conolly, R.B.; Corsini, E.; Green, S.; Faustman, E.M.; Gaspari, A.; et al. T4 report. An expert consortium review of the EC-commissioned report “Alternative (non-animal) methods for cosmetics testing: Current status and future prospects-2010”. ALTEX 2011, 28, 183–209. [Google Scholar] [CrossRef][Green Version]
- Dothel, G.; Vasina, V.; Barbara, G.; De Ponti, F. Animal models of chemically induced intestinal inflammation: Predictivity and ethical issues. Pharmacol. Ther. 2013, 139, 71–86. [Google Scholar] [CrossRef]
- Doke, S.K.; Dhawale, S.C. Alternatives to animal testing: A review. Saudi Pharm. J. 2015, 23, 223–229. [Google Scholar] [CrossRef]
- Baker, B.M.; Chen, C.S. Deconstructing the third dimension-how 3D culture microenvironments alter cellular cues. J. Cell Sci. 2012, 125, 3015–3024. [Google Scholar] [CrossRef]
- Horvath, P.; Aulner, N.; Bickle, M.; Davies, A.M.; Nery, E.D.; Ebner, D.; Montoya, M.C.; Östling, P.; Pietiäinen, V.; Price, L.S.; et al. Screening out irrelevant cell-based models of disease. Nat. Rev. Drug Discov. 2016, 15, 751–769. [Google Scholar] [CrossRef]
- Lancaster, M.A.; Renner, M.; Martin, C.A.; Wenzel, D.; Bicknell, L.S.; Hurles, M.E.; Homfray, T.; Penninger, J.M.; Jackson, A.P.; Knoblich, J.A. Cerebral organoids model human brain development and microcephaly. Nature 2013, 501, 373–379. [Google Scholar] [CrossRef]
- Abbott, A. Biology’s new dimension. Nature 2003, 424, 870–872. [Google Scholar] [CrossRef] [PubMed]
- Raghavan, S.; Ward, M.R.; Rowley, K.R.; Wold, R.M.; Takayama, S.; Buckanovich, R.J.; Mehta, G. Formation of stable small cell number three-dimensional ovarian cancer spheroids using hanging drop arrays for preclinical drug sensitivity assays. Gynecol. Oncol. 2015, 138, 181–189. [Google Scholar] [CrossRef] [PubMed]
- Herter, S.; Morra, L.; Schlenker, R.; Sulcova, J.; Fahrni, L.; Waldhauer, I.; Lehmann, S.; Reisländer, T.; Agarkova, I.; Kelm, J.M.; et al. A novel three-dimensional heterotypic spheroid model for the assessment of the activity of cancer immunotherapy agents. Cancer Immunol. Immunother. 2017, 66, 129–140. [Google Scholar] [CrossRef] [PubMed]
- Thoma, C.R.; Zimmermann, M.; Agarkova, I.; Kelm, J.M.; Krek, W. 3D cell culture systems modeling tumor growth determinants in cancer target discovery. Adv. Drug Deliv. Rev. 2014, 69–70, 29–41. [Google Scholar] [CrossRef] [PubMed]
- Longati, P.; Jia, X.; Eimer, J.; Wagman, A.; Witt, M.R.; Rehnmark, S.; Verbeke, C.; Toftgård, R.; Löhr, M.; Heuchel, R.L. 3D pancreatic carcinoma spheroids induce a matrix-rich, chemoresistant phenotype offering a better model for drug testing. BMC Cancer 2013, 13, 95. [Google Scholar] [CrossRef]
- Costa, E.C.; Moreira, A.F.; de Melo-Diogo, D.; Gaspar, V.M.; Carvalho, M.P.; Correia, I.J. 3D tumor spheroids: An overview on the tools and techniques used for their analysis. Biotechnol. Adv. 2016, 34, 1427–1441. [Google Scholar] [CrossRef]
- Markovitz-Bishitz, Y.; Tauber, Y.; Afrimzon, E.; Zurgil, N.; Sobolev, M.; Shafran, Y.; Deutsch, A.; Howitz, S.; Deutsch, M. A polymer microstructure array for the formation, culturing, and high throughput drug screening of breast cancer spheroids. Biomaterials 2010, 31, 8436–8444. [Google Scholar] [CrossRef]
- Asghar, W.; El Assal, R.; Shafiee, H.; Pitteri, S.; Paulmurugan, R.; Demirci, U. Engineering cancer microenvironments for in vitro 3-D tumor models. Mater. Today 2015, 18, 539–553. [Google Scholar] [CrossRef]
- Infanger, D.W.; Lynch, M.E.; Fischbach, C. Engineered Culture Models for Studies of Tumor- Microenvironment Interactions. Annu. Rev. Biomed. Eng. 2013, 15, 29–53. [Google Scholar] [CrossRef]
- Alemany-ribes, M.; Semino, C.E. Bioengineering 3D environments for cancer models. Adv. Drug Deliv. Rev. 2014, 79–80, 40–49. [Google Scholar] [CrossRef]
- Gebeyehu, A.; Surapaneni, S.K.; Huang, J.; Mondal, A.; Wang, V.Z.; Haruna, N.F.; Bagde, A.; Arthur, P.; Kutlehria, S.; Patel, N. Polysaccharide hydrogel based 3D printed tumor models for chemotherapeutic drug screening. Sci. Rep. 2021, 11, 372. [Google Scholar] [CrossRef] [PubMed]
- Shi, W.; Huang, J.; Fang, R.; Liu, M. Imparting functionality to the hydrogel by magnetic-field-induced nano-assembly and macro-response. ACS Appl. Mater. Interfaces 2020, 12, 5177–5194. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, F.-Y.; Han, H.-W.; Chen, X.-R.; Yang, C.-S.; Wei, Y.; Hsu, S. Non-viral delivery of an optogenetic tool into cells with self-healing hydrogel. Biomaterials 2018, 174, 31–40. [Google Scholar] [CrossRef]
- Young, A.T.; White, O.C.; Daniele, M.A. Rheological properties of coordinated physical gelation and chemical crosslinking in gelatin methacryloyl (GelMA) hydrogels. Macromol. Biosci. 2020, 20, 2000183. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Kankala, R.K.; Ou, C.; Chen, A.; Yang, Z. Advances in hydrogel-based vascularized tissues for tissue repair and drug screening. Bioact. Mater. 2022, 9, 198–220. [Google Scholar] [CrossRef]
- Alarçin, E.; Lee, T.Y.; Karuthedom, S.; Mohammadi, M.; Brennan, M.A.; Lee, D.H.; Marrella, A.; Zhang, J.; Syla, D.; Zhang, Y.S. Injectable shear-thinning hydrogels for delivering osteogenic and angiogenic cells and growth factors. Biomater. Sci. 2018, 6, 1604–1615. [Google Scholar] [CrossRef]
- Zhang, J.; Zheng, Y.; Lee, J.; Hua, J.; Li, S.; Panchamukhi, A.; Yue, J.; Gou, X.; Xia, Z.; Zhu, L. A pulsatile release platform based on photo-induced imine-crosslinking hydrogel promotes scarless wound healing. Nat. Commun. 2021, 12, 1670. [Google Scholar] [CrossRef]
- Zhu, D.; Li, Z.; Huang, K.; Caranasos, T.G.; Rossi, J.S.; Cheng, K. Minimally invasive delivery of therapeutic agents by hydrogel injection into the pericardial cavity for cardiac repair. Nat. Commun. 2021, 12, 1412. [Google Scholar] [CrossRef]
- Nicodemus, G.D.; Bryant, S.J. Cell encapsulation in biodegradable hydrogels for tissue engineering applications. Tissue Eng. Part B Rev. 2008, 14, 149–165. [Google Scholar] [CrossRef]
- Marrella, A.; Lagazzo, A.; Barberis, F.; Catelani, T.; Quarto, R.; Scaglione, S. Enhanced mechanical performances and bioactivity of cell laden-graphene oxide/alginate hydrogels open new scenario for articular tissue engineering applications. Carbon 2017, 115, 608–616. [Google Scholar] [CrossRef]
- Tibbitt, M.W.; Anseth, K.S. Hydrogels as extracellular matrix mimics for 3D cell culture. Biotechnol. Bioeng. 2009, 103, 655–663. [Google Scholar] [CrossRef] [PubMed]
- Unal, A.Z.; West, J.L. Synthetic ECM: Bioactive synthetic hydrogels for 3D tissue engineering. Bioconjug. Chem. 2020, 31, 2253–2271. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Shim, K.Y.; Kim, B.; Sung, J.H. Hydrogel-based three-dimensional cell culture for organ-on-a-chip applications. Biotechnol. Prog. 2017, 33, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Marrella, A.; Lagazzo, A.; Dellacasa, E.; Pasquini, C.; Finocchio, E.; Barberis, F.; Pastorino, L.; Giannoni, P.; Scaglione, S. 3D porous gelatin/PVA hydrogel as meniscus substitute using alginate micro-particles as porogens. Polymers 2018, 10, 380. [Google Scholar] [CrossRef] [PubMed]
- Song, R.; Murphy, M.; Li, C.; Ting, K.; Soo, C.; Zheng, Z. Current development of biodegradable polymeric materials for biomedical applications. Drug Des. Dev. Ther. 2018, 12, 3117. [Google Scholar] [CrossRef]
- Schnabel-Lubovsky, M.; Kossover, O.; Melino, S.; Nanni, F.; Talmon, Y.; Seliktar, D. Visualizing cell-laden fibrin-based hydrogels using cryogenic scanning electron microscopy and confocal microscopy. J. Tissue Eng. Regen. Med. 2019, 13, 587–598. [Google Scholar] [CrossRef]
- Sohn, H.W.; Tolar, P.; Brzostowski, J.; Pierce, S.K. Imaging and analysis of three-dimensional cell culture models. Methods Mol. Biol. 2010, 591, 159–183. [Google Scholar] [CrossRef]
- Marrella, A. 3D fluid-dynamic ovarian cancer model resembling systemic drug administration for efficacy assay. ALTEX 2020, 37, 82–94. [Google Scholar] [CrossRef]
- Hasan, A.; Nurunnabi, M.; Morshed, M.; Paul, A.; Polini, A.; Kuila, T.; Al Hariri, M.; Lee, Y.; Jaffa, A.A. Recent advances in application of biosensors in tissue engineering. Biomed Res. Int. 2014, 2014, 307519. [Google Scholar] [CrossRef]
- Li, Y.-C.E.; Lee, I. The current trends of biosensors in tissue engineering. Biosensors 2020, 10, 88. [Google Scholar] [CrossRef]
- Purohit, B.; Vernekar, P.R.; Shetti, N.P.; Chandra, P. Biosensor nanoengineering: Design, operation, and implementation for biomolecular analysis. Sensors Int. 2020, 1, 100040. [Google Scholar] [CrossRef]
- Kratz, S.R.A.; Höll, G.; Schuller, P.; Ertl, P.; Rothbauer, M. Latest trends in biosensing for microphysiological organs-on-a-chip and body-on-a-chip systems. Biosensors 2019, 9, 110. [Google Scholar] [CrossRef] [PubMed]
- Caballero, D.; Kaushik, S.; Correlo, V.M.; Oliveira, J.M.; Reis, R.L.; Kundu, S.C. Organ-on-chip models of cancer metastasis for future personalized medicine: From chip to the patient. Biomaterials 2017, 149, 98–115. [Google Scholar] [CrossRef] [PubMed]
- Kieninger, J.; Weltin, A.; Flamm, H.; Urban, G.A. Microsensor systems for cell metabolism-from 2D culture to organ-on-chip. Lab Chip 2018, 18, 1274–1291. [Google Scholar] [CrossRef] [PubMed]
- Bousse, L. Whole cell biosensors. Sens. Actuators B Chem. 1996, 34, 270–275. [Google Scholar] [CrossRef]
- Du, H.; Strohsahl, C.M.; Camera, J.; Miller, B.L.; Krauss, T.D. Sensitivity and specificity of metal surface-immobilized “molecular beacon” biosensors. J. Am. Chem. Soc. 2005, 127, 7932–7940. [Google Scholar] [CrossRef]
- Mehrotra, P. Biosensors and their applications A review. J. Oral Biol. Craniofacial Res. 2016, 6, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Alhadrami, H.A. Biosensors: Classifications, medical applications, and future prospective. Biotechnol. Appl. Biochem. 2018, 65, 497–508. [Google Scholar] [CrossRef] [PubMed]
- Hu, N.; Ha, D.; Wu, C.; Zhou, J.; Kirsanov, D.; Legin, A.; Wang, P. A LAPS array with low cross-talk for non-invasive measurement of cellular metabolism. Sens. Actuators A Phys. 2012, 187, 50–56. [Google Scholar] [CrossRef]
- Mandel, L.J. Chapter 8 Energy Metabolism of Cellular Activation, Growth, and Transformation. In The Role of Membranes in Cell Growth and Differentiation; Kleinzeller, A.B.T.-C.T., Ed.; Academic Press: Cambridge, MA, USA, 1986; Volume 27, pp. 261–291. [Google Scholar]
- Owicki, J.C.; Wallace Parce, J. Biosensors based on the energy metabolism of living cells: The physical chemistry and cell biology of extracellular acidification. Biosens. Bioelectron. 1992, 7, 255–272. [Google Scholar] [CrossRef]
- Lehmann, M.; Baumann, W.; Brischwein, M.; Ehret, R.; Kraus, M.; Schwinde, A.; Bitzenhofer, M.; Freund, I.; Wolf, B. Non-invasive measurement of cell membrane associated proton gradients by ion-sensitive field effect transistor arrays for microphysiological and bioelectronical applications. Biosens. Bioelectron. 2000, 15, 117–124. [Google Scholar] [CrossRef]
- Wolf, B.; Brischwein, M.; Baumann, W.; Ehret, R.; Kraus, M. Monitoring of cellular signalling and metabolism with modular sensor- technique: The physiocontrol-microsystem (PCM®). Biosens. Bioelectron. 1998, 13, 501–509. [Google Scholar] [CrossRef]
- Zhang, X.; Lin, Y.; Gillies, R.J. Tumor pH and its measurement. J. Nucl. Med. 2010, 51, 1167–1170. [Google Scholar] [CrossRef]
- Stock, C.; Schwab, A. Protons make tumor cells move like clockwork. Pflug. Arch. Eur. J. Physiol. 2009, 458, 981–992. [Google Scholar] [CrossRef] [PubMed]
- Singh, K.; Ohlan, A.; Saini, P.; Dhawan, S.K. pH sensor based on polyaniline and aniline–anthranilic acid copolymer films using quartz crystal microbalance and electronic absorption spectroscopy. Polym. Adv. Technol. 2008, 19, 229–236. [Google Scholar] [CrossRef]
- Martin, N.K.; Gaffney, E.A.; Gatenby, R.A.; Maini, P.K. Tumour-stromal interactions in acid-mediated invasion: A mathematical model. J. Theor. Biol. 2010, 267, 461–470. [Google Scholar] [CrossRef] [PubMed]
- Chu, B.; Wang, H.; Song, B.; Peng, F.; Su, Y.; He, Y. Fluorescent and Photostable Silicon Nanoparticles Sensors for Real-Time and Long-Term Intracellular pH Measurement in Live Cells. Anal. Chem. 2016, 88, 9235–9242. [Google Scholar] [CrossRef] [PubMed]
- Wike-Hooley, J.L.; van den Berg, A.P.; van der Zee, J.; Reinhold, H.S. Human tumour pH and its variation. Eur. J. Cancer Clin. Oncol. 1985, 21, 785–791. [Google Scholar] [CrossRef]
- Ges, I.A.; Ivanov, B.L.; Schaffer, D.K.; Lima, E.A.; Werdich, A.A.; Baudenbacher, F.J. Thin-film IrOx pH microelectrode for microfluidic-based microsystems. Biosens. Bioelectron. 2005, 21, 248–256. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.F.; Lee, G.B.; Wang, C.H.; Lee, H.H.; Liao, W.Y.; Chou, T.C. Microfluidic pH-sensing chips integrated with pneumatic fluid-control devices. Biosens. Bioelectron. 2006, 21, 1468–1475. [Google Scholar] [CrossRef] [PubMed]
- Milgrew, M.J.; Riehle, M.O.; Cumming, D.R.S. A large transistor-based sensor array chip for direct extracellular imaging. Sens. Actuators B Chem. 2005, 111–112, 347–353. [Google Scholar] [CrossRef]
- Yadavalli, V.K.; Pishko, M.V. Biosensing in microfluidic channels using fluorescence polarization. Anal. Chim. Acta 2004, 507, 123–128. [Google Scholar] [CrossRef]
- Liu, Z.; Liu, J.; Chen, T. Phenol red immobilized PVA membrane for an optical pH sensor with two determination ranges and long-term stability. Sens. Actuators B Chem. 2005, 107, 311–316. [Google Scholar] [CrossRef]
- Park, C.H.; Lee, S.; Pornnoppadol, G.; Nam, Y.S.; Kim, S.H.; Kim, B.J. Microcapsules Containing pH-Responsive, Fluorescent Polymer-Integrated MoS2: An Effective Platform for in Situ pH Sensing and Photothermal Heating. ACS Appl. Mater. Interfaces 2018, 10, 9023–9031. [Google Scholar] [CrossRef] [PubMed]
- Kattipparambil Rajan, D.; Patrikoski, M.; Verho, J.; Sivula, J.; Ihalainen, H.; Miettinen, S.; Lekkala, J. Optical non-contact pH measurement in cell culture with sterilizable, modular parts. Talanta 2016, 161, 755–761. [Google Scholar] [CrossRef] [PubMed]
- Moldero, I.L.; Chandra, A.; Cavo, M.; Mota, C.; Kapsokalyvas, D.; Gigli, G.; Moroni, L.; del Mercato, L.L. Probing the pH Microenvironment of Mesenchymal Stromal Cell Cultures on Additive-Manufactured Scaffolds. Small 2020, 16, 2002258. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Hou, T.; Pan, T. High-Performance Double-Gate. IEEE Trans. Electron Devices 2018, 65, 237–242. [Google Scholar] [CrossRef]
- Lai, C.S.; Yang, C.M.; Lu, T.F. PH sensitivity improvement on 8 nm thick hafnium oxide by post deposition annealing. Electrochem. Solid-State Lett. 2006, 9, 90–93. [Google Scholar] [CrossRef]
- Pan, T.M.; Liao, K.M. Structural properties and sensing characteristics of Y2O3 sensing membrane for pH-ISFET. Sens. Actuators B Chem. 2007, 127, 480–485. [Google Scholar] [CrossRef]
- Rani, R.A.; Sidek, O. ISFET pH sensor characterization: Towards biosensor microchip application. In Proceedings of the 2004 IEEE Region 10 Conference TENCON 2004, Chiang Mai, Thailand, 24 November 2004; Volume 500, pp. 660–663. [Google Scholar] [CrossRef]
- Lehmann, M.; Baumann, W.; Brischwein, M.; Gahle, H.J.; Freund, I.; Ehret, R.; Drechsler, S.; Palzer, H.; Kleintges, M.; Sieben, U.; et al. Simultaneous measurement of cellular respiration and acidification with a single CMOS ISFET. Biosens. Bioelectron. 2001, 16, 195–203. [Google Scholar] [CrossRef]
- Owicki, J.C.; Bousse, L.J.; Hafeman, D.G.; Kirk, G.L.; Olson, J.D.; Wada, H.G.; Parce, J.W. The Light-Addressable Potentiometric Sensor: Principles and Biological Applications. Annu. Rev. Biophys. Biomol. Struct. 1994, 23, 87–114. [Google Scholar] [CrossRef] [PubMed]
- Yoshinobu, T.; Schöning, M.J. Light-addressable potentiometric sensors (LAPS) for cell monitoring and biosensing. Curr. Opin. Electrochem. 2021, 28, 100727. [Google Scholar] [CrossRef]
- Hu, N.; Wu, C.; Ha, D.; Wang, T.; Liu, Q.; Wang, P. A novel microphysiometer based on high sensitivity LAPS and microfluidic system for cellular metabolism study and rapid drug screening. Biosens. Bioelectron. 2013, 40, 167–173. [Google Scholar] [CrossRef]
- Shaibani, P.M.; Etayash, H.; Naicker, S.; Kaur, K.; Thundat, T. Metabolic Study of Cancer Cells Using a pH Sensitive Hydrogel Nanofiber Light Addressable Potentiometric Sensor. ACS Sens. 2017, 2, 151–156. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.-M.; Yen, T.; Liu, H.-L.; Lin, Y.-J.; Lin, P.-Y.; Tsui, L.S.; Chen, C.-H.; Chen, Y.-P.; Hsu, Y.-C.; Lo, C.-H. A real-time mirror-LAPS mini system for dynamic chemical imaging and cell acidification monitoring. Sens. Actuators B Chem. 2021, 341, 130003. [Google Scholar] [CrossRef]
- Zhang, H.W.; Hu, X.B.; Qin, Y.; Jin, Z.H.; Zhang, X.W.; Liu, Y.L.; Huang, W.H. Conductive Polymer Coated Scaffold to Integrate 3D Cell Culture with Electrochemical Sensing. Anal. Chem. 2019, 91, 4838–4844. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.H.; Lin, J.L.; Wang, J.; Cui, Z.; Cui, Z. Development of high throughput optical sensor array for on-line pH monitoring in micro-scale cell culture environment. Biomed. Microdevices 2009, 11, 265–273. [Google Scholar] [CrossRef]
- Wikswo, J.P.; Block, F.E.; Cliffel, D.E.; Goodwin, C.R.; Marasco, C.C.; Markov, D.A.; McLean, D.L.; McLean, J.A.; McKenzie, J.R.; Reiserer, R.S.; et al. Engineering challenges for instrumenting and controlling integrated organ-on-chip systems. IEEE Trans. Biomed. Eng. 2013, 60, 682–690. [Google Scholar] [CrossRef]
- Shaegh, S.A.M.; De Ferrari, F.; Zhang, Y.S.; Nabavinia, M.; Mohammad, N.B.; Ryan, J.; Pourmand, A.; Laukaitis, E.; Sadeghian, R.B.; Nadhman, A.; et al. A microfluidic optical platform for real-time monitoring of pH and oxygen in microfluidic bioreactors and organ-on-chip devices. Biomicrofluidics 2016, 10, 044111. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Wu, S.; Yi, Z.; Zeng, F.; Wu, W.; Qiao, Y.; Zhao, X.; Cheng, X.; Tian, Y. Hydrogel-based fluorescent dual pH and oxygen sensors loaded in 96-well plates for high-throughput cell metabolism studies. Sensors 2018, 18, 564. [Google Scholar] [CrossRef] [PubMed]
- Hanson, M.A.; Ge, X.; Kostov, Y.; Brorson, K.A.; Moreira, A.R.; Rao, G. Comparisons of optical pH and dissolved oxygen sensors with traditional electrochemical probes during mammalian cell culture. Biotechnol. Bioeng. 2007, 97, 833–841. [Google Scholar] [CrossRef]
- Naciri, M.; Kuystermans, D.; Al-Rubeai, M. Monitoring pH and dissolved oxygen in mammalian cell culture using optical sensors. Cytotechnology 2008, 57, 245–250. [Google Scholar] [CrossRef]
- Arik, M.; Çelebi, N.; Onganer, Y. Fluorescence quenching of fluorescein with molecular oxygen in solution. J. Photochem. Photobiol. A Chem. 2005, 170, 105–111. [Google Scholar] [CrossRef]
- Buckler, K.J.; Vaughan-Jones, R.D. Application of a new pH-sensitive fluoroprobe (carboxy-SNARF-1) for intracellular pH measurement in small, isolated cells. Pflügers Arch. Eur. J. Physiol. 1990, 417, 234–239. [Google Scholar] [CrossRef]
- Kenney, R.M.; Boyce, M.W.; Whitman, N.A.; Kromhout, B.P.; Lockett, M.R. A pH-Sensing Optode for Mapping Spatiotemporal Gradients in 3D Paper-Based Cell Cultures. Anal. Chem. 2018, 90, 2376–2383. [Google Scholar] [CrossRef]
- Chandra, A.; Singh, N. Cell Microenvironment pH Sensing in 3D Microgels Using Fluorescent Carbon Dots. ACS Biomater. Sci. Eng. 2017, 3, 3620–3627. [Google Scholar] [CrossRef]
- Shangguan, J.; He, D.; He, X.; Wang, K.; Xu, F.; Liu, J.; Tang, J.; Yang, X.; Huang, J. Label-Free Carbon-Dots-Based Ratiometric Fluorescence pH Nanoprobes for Intracellular pH Sensing. Anal. Chem. 2016, 88, 7837–7843. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Xu, Z.; Zhang, C. Polyethyleneimine-Functionalized Fluorescent Carbon Dots: Water Stability, pH Sensing, and Cellular Imaging. ChemNanoMat 2015, 1, 122–127. [Google Scholar] [CrossRef]
- Du, F.; Ming, Y.; Zeng, F.; Yu, C.; Wu, S. A low cytotoxic and ratiometric fluorescent nanosensor based on carbon-dots for intracellular pH sensing and mapping. Nanotechnology 2013, 24, 365101. [Google Scholar] [CrossRef] [PubMed]
- Ke, G.; Zhu, Z.; Wang, W.; Zou, Y.; Guan, Z.; Jia, S.; Zhang, H.; Wu, X.; Yang, C.J. A cell-surface-anchored ratiometric fluorescent probe for extracellular pH sensing. ACS Appl. Mater. Interfaces 2014, 6, 15329–15334. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Jimenez Del Val, I.; Müller, C.; Wagtberg Sen, J.; Rasmussen, S.K.; Kontoravdi, C.; Weilguny, D.; Andersen, M.R. Amino acid and glucose metabolism in fed-batch CHO cell culture affects antibody production and glycosylation. Biotechnol. Bioeng. 2015, 112, 521–535. [Google Scholar] [CrossRef] [PubMed]
- Quek, L.-E.; Dietmair, S.; Krömer, J.O.; Nielsen, L.K. Metabolic flux analysis in mammalian cell culture. Metab. Eng. 2010, 12, 161–171. [Google Scholar] [CrossRef] [PubMed]
- Boero, C.; Carrara, S.; Del Vecchio, G.; Calzà, L.; De Micheli, G. Highly sensitive carbon nanotube-based sensing for lactate and glucose monitoring in cell culture. IEEE Trans. Nanobiosci. 2011, 10, 59–67. [Google Scholar] [CrossRef] [PubMed]
- Borgmann, S.; Schulte, A.; Neugebauer, S.; Schuhmann, W. Amperometric Biosensors; Wiley: Hoboken, NJ, USA, 2012; Volume 13, ISBN 9783527644117. [Google Scholar]
- Heller, A.; Feldman, B. Electrochemical Glucose Sensors and Their Applications in Diabetes Management. Chem. Rev. 2008, 108, 2482–2505. [Google Scholar] [CrossRef]
- Obregón, R.; Ahadian, S.; Ramón-Azcón, J.; Chen, L.; Fujita, T.; Shiku, H.; Chen, M.; Matsue, T. Non-invasive measurement of glucose uptake of skeletal muscle tissue models using a glucose nanobiosensor. Biosens. Bioelectron. 2013, 50, 194–201. [Google Scholar] [CrossRef]
- Chang, D.K.; Goel, A.; Ricciardiello, L.; Lee, D.H.; Chang, C.L.; Carethers, J.M.; Boland, C.R. Effect of H2O2 on cell cycle and survival in DNA mismatch repair-deficient and -proficient cell lines. Cancer Lett. 2003, 195, 243–251. [Google Scholar] [CrossRef]
- Pereira Rodrigues, N.; Sakai, Y.; Fujii, T. Cell-based microfluidic biochip for the electrochemical real-time monitoring of glucose and oxygen. Sens. Actuators B Chem. 2008, 132, 608–613. [Google Scholar] [CrossRef]
- Weltin, A.; Slotwinski, K.; Kieninger, J.; Moser, I.; Jobst, G.; Wego, M.; Ehret, R.; Urban, G.A. Cell culture monitoring for drug screening and cancer research: A transparent, microfluidic, multi-sensor microsystem. Lab Chip 2014, 14, 138–146. [Google Scholar] [CrossRef]
- Kemas, A.M.; Youhanna, S.; Zandi Shafagh, R.; Lauschke, V.M. Insulin-dependent glucose consumption dynamics in 3D primary human liver cultures measured by a sensitive and specific glucose sensor with nanoliter input volume. FASEB J. 2021, 35, e21305. [Google Scholar] [CrossRef]
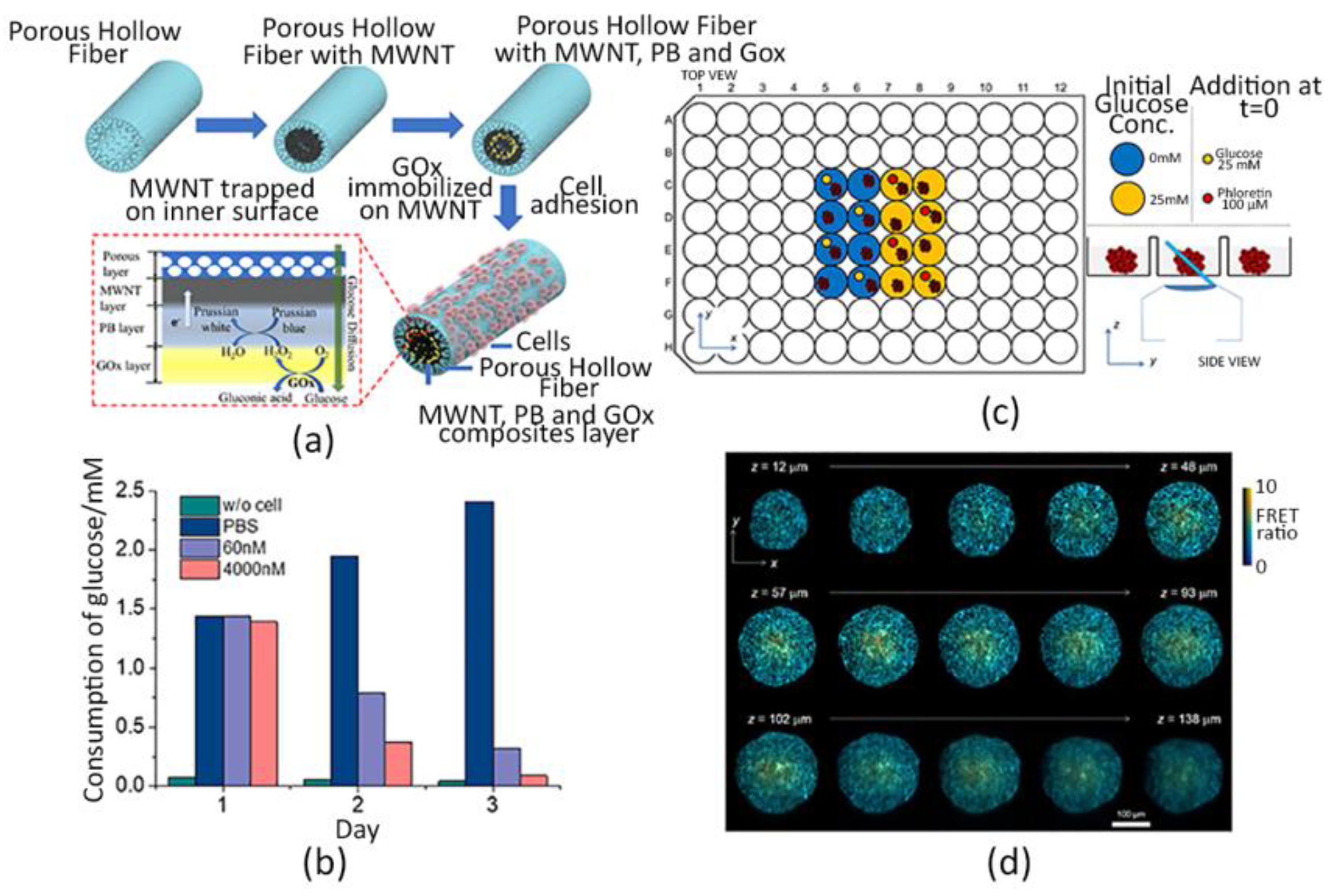
- Ma, Z.; Luo, Y.; Zhu, Q.; Jiang, M.; Pan, M.; Xie, T.; Huang, X.; Chen, D. In-situ monitoring of glucose metabolism in cancer cell microenvironments based on hollow fiber structure. Biosens. Bioelectron. 2020, 162, 112261. [Google Scholar] [CrossRef]
- Bavli, D.; Prill, S.; Ezra, E.; Levy, G.; Cohen, M.; Vinken, M.; Vanfleteren, J.; Jaeger, M.; Nahmias, Y. Real-time monitoring of metabolic function in liver-onchip microdevices tracks the dynamics of Mitochondrial dysfunction. Proc. Natl. Acad. Sci. USA. 2016, 113, E2231–E2240. [Google Scholar] [CrossRef] [PubMed]
- Qian, L.H.; Chen, M.W. Ultrafine nanoporous gold by low-temperature dealloying and kinetics of nanopore formation. Appl. Phys. Lett. 2007, 91, 083105. [Google Scholar] [CrossRef]
- Figueiredo, L.; Pace, R.; D’Arros, C.; Réthoré, G.; Guicheux, J.; Le Visage, C.; Weiss, P. Assessing glucose and oxygen diffusion in hydrogels for the rational design of 3D stem cell scaffolds in regenerative medicine. J. Tissue Eng. Regen. Med. 2018, 12, 1238–1246. [Google Scholar] [CrossRef] [PubMed]
- Farrell, M.J.; Shin, J.I.; Smith, L.J.; Mauck, R.L. Functional consequences of glucose and oxygen deprivation onengineered mesenchymal stem cell-based cartilage constructs. Osteoarthr. Cartil. 2015, 23, 134–142. [Google Scholar] [CrossRef] [PubMed]
- Li, I.T.; Pham, E.; Truong, K. Protein biosensors based on the principle of fluorescence resonance energy transfer for monitoring cellular dynamics. Biotechnol. Lett. 2006, 28, 1971–1982. [Google Scholar] [CrossRef] [PubMed]
- Lorenzo, C.; Frongia, C.; Jorand, R.; Fehrenbach, J.; Weiss, P.; Maandhui, A.; Gay, G.; Ducommun, B.; Lobjois, V. Live cell division dynamics monitoring in 3D large spheroid tumor models using light sheet microscopy. Cell Div. 2011, 6, 22. [Google Scholar] [CrossRef] [PubMed]
- Maioli, V.; Chennell, G.; Sparks, H.; Lana, T.; Kumar, S.; Carling, D.; Sardini, A.; Dunsby, C. Time-lapse 3-D measurements of a glucose biosensor in multicellular spheroids by light sheet fluorescence microscopy in commercial 96-well plates. Sci. Rep. 2016, 6, 37777. [Google Scholar] [CrossRef] [PubMed]
- Lewis, D.M.; Blatchley, M.R.; Park, K.M.; Gerecht, S. O2-controllable hydrogels for studying cellular responses to hypoxic gradients in three dimensions in vitro and in vivo. Nat. Protoc. 2017, 12, 1620–1638. [Google Scholar] [CrossRef]
- Schmitz, C.; Pepelanova, I.; Seliktar, D.; Potekhina, E.; Belousov, V.V.; Scheper, T.; Lavrentieva, A. Live reporting for hypoxia: Hypoxia sensor–modified mesenchymal stem cells as in vitro reporters. Biotechnol. Bioeng. 2020, 117, 3265–3276. [Google Scholar] [CrossRef] [PubMed]
- Rouwkema, J.; Koopman, B.F.J.M.; Blitterswijk, C.A.V.; Dhert, W.J.A.; Malda, J. Supply of nutrients to cells in engineered tissues. Biotechnol. Genet. Eng. Rev. 2009, 26, 163–178. [Google Scholar] [CrossRef] [PubMed]
- Christopher W Pugh; Peter J Ratcliffe Regulation of angiogenesis by hypoxia: Role of the HIF system. Nat. Med. 2003, 9, 677–684. [CrossRef] [PubMed]
- Magliaro, C.; Mattei, G.; Iacoangeli, F.; Corti, A.; Piemonte, V.; Ahluwalia, A.; Lewis, D.M.; Park, K.M.; Tang, V.; Xu, Y.; et al. Hypoxia in Static and Dynamic 3D Culture Systems for Tissue Engineering of Bone. Free Radic. Biol. Med. 2016, 14, 1331–1340. [Google Scholar] [CrossRef]
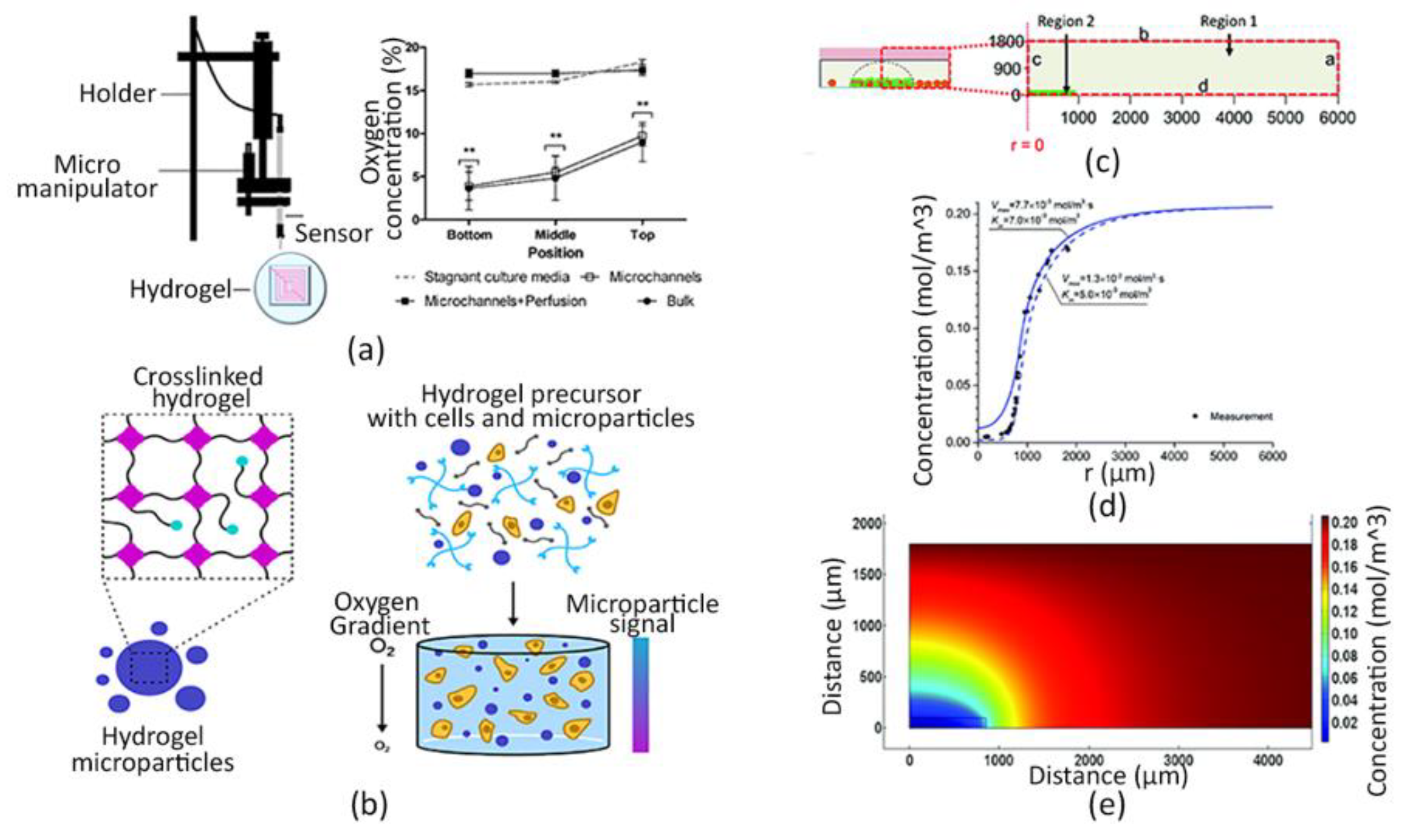
- Acosta, M.A.; Ymele-Leki, P.; Kostov, Y.V.; Leach, J.B. Fluorescent microparticles for sensing cell microenvironment oxygen levels within 3D scaffolds. Biomaterials 2009, 30, 3068–3074. [Google Scholar] [CrossRef] [PubMed]
- Westfall, S.D.; Sachdev, S.; Das, P.; Hearne, L.B.; Hannink, M.; Roberts, R.M.; Ezashi, T. Identification of oxygen-sensitive transcriptional programs in human embryonic stem cells. Stem Cells Dev. 2008, 17, 869–881. [Google Scholar] [CrossRef] [PubMed]
- O’Driscoll, S.W.; Fitzsimmons, J.S.; Commisso, C.N. Role of oxygen tension during cartilage formation by periosteum. J. Orthop. Res. 1997, 15, 682–687. [Google Scholar] [CrossRef] [PubMed]
- Mohyeldin, A.; Garzón-Muvdi, T.; Quiñones-Hinojosa, A. Oxygen in stem cell biology: A critical component of the stem cell niche. Cell Stem Cell 2010, 7, 150–161. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Zhu, L.; Fan, M. Oxygen, a Key Factor Regulating Cell Behavior during Neurogenesis and Cerebral Diseases. Front. Mol. Neurosci. 2011, 4, 5. [Google Scholar] [CrossRef]
- Carreau, A.; Hafny-Rahbi, B.E.; Matejuk, A.; Grillon, C.; Kieda, C. Why is the partial oxygen pressure of human tissues a crucial parameter? Small molecules and hypoxia. J. Cell. Mol. Med. 2011, 15, 1239–1253. [Google Scholar] [CrossRef] [PubMed]
- Pouysségur, J.; Dayan, F.; Mazure, N.M. Hypoxia signalling in cancer and approaches to enforce tumour regression. Nature 2006, 441, 437–443. [Google Scholar] [CrossRef]
- Harrison, B.S.; Eberli, D.; Lee, S.J.; Atala, A.; Yoo, J.J. Oxygen producing biomaterials for tissue regeneration. Biomaterials 2007, 28, 4628–4634. [Google Scholar] [CrossRef]
- Hossmann, K.-A. Pathophysiology and Therapy of Experimental Stroke. Cell. Mol. Neurobiol. 2006, 26, 1055–1081. [Google Scholar] [CrossRef] [PubMed]
- Heathman, T.R.J.; Nienow, A.W.; McCall, M.J.; Coopman, K.; Kara, B.; Hewitt, C.J. The translation of cell-based therapies: Clinical landscape and manufacturing challenges. Regen. Med. 2015, 10, 49–64. [Google Scholar] [CrossRef] [PubMed]
- Beckers, S.; Noor, F.; Müller-Vieira, U.; Mayer, M.; Strigun, A.; Heinzle, E. High throughput, non-invasive and dynamic toxicity screening on adherent cells using respiratory measurements. Toxicol. In Vitro 2010, 24, 686–694. [Google Scholar] [CrossRef]
- Wouters, A.; Pauwels, B.; Lambrechts, H.A.J.; Pattyn, G.G.O.; Ides, J.; Baay, M.; Meijnders, P.; Dewilde, S.; Vermorken, J.B.; Lardon, F. Chemoradiation interactions under reduced oxygen conditions: Cellular characteristics of an in vitro model. Cancer Lett. 2009, 286, 180–188. [Google Scholar] [CrossRef]
- Browne, S.M.; Daud, H.; Murphy, W.G.; Al-Rubeai, M. Measuring dissolved oxygen to track erythroid differentiation of hematopoietic progenitor cells in culture. J. Biotechnol. 2014, 187, 135–138. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.C.; Luk, H.N.; Lin, Y.T.T.; Yuan, C.Y. A Clark-type oxygen chip for in situ estimation of the respiratory activity of adhering cells. Talanta 2010, 81, 228–234. [Google Scholar] [CrossRef] [PubMed]
- Kieninger, J.; Aravindalochanan, K.; Sandvik, J.A.; Pettersen, E.O.; Urban, G.A. Pericellular oxygen monitoring with integrated sensor chips for reproducible cell culture experiments. Cell Prolif. 2014, 47, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Liebisch, F.; Weltin, A.; Marzioch, J.; Urban, G.A.; Kieninger, J. Zero-consumption Clark-type microsensor for oxygen monitoring in cell culture and organ-on-chip systems. Sens. Actuators B Chem. 2020, 322, 128652. [Google Scholar] [CrossRef]
- Kieninger, J.; Tamari, Y.; Enderle, B.; Jobst, G.; Sandvik, J.A.; Pettersen, E.O.; Urban, G.A. Sensor access to the cellular microenvironment using the sensing cell culture flask. Biosensors 2018, 8, 44. [Google Scholar] [CrossRef] [PubMed]
- Wilson, R.L.; Connell, J.P.; Grande-Allen, K.J. Monitoring Oxygen Levels within Large, Tissue-Engineered Constructs Using Porphyin-Hydrogel Microparticles. ACS Biomater. Sci. Eng. 2019, 5, 4522–4530. [Google Scholar] [CrossRef]
- Magliaro, C.; Mattei, G.; Iacoangeli, F.; Corti, A.; Piemonte, V.; Ahluwalia, A. Oxygen consumption characteristics in 3D constructs depend on cell density. Front. Bioeng. Biotechnol. 2019, 7, 251. [Google Scholar] [CrossRef] [PubMed]
- Weltin, A.; Hammer, S.; Noor, F.; Kaminski, Y.; Kieninger, J.; Urban, G.A. Accessing 3D microtissue metabolism: Lactate and oxygen monitoring in hepatocyte spheroids. Biosens. Bioelectron. 2017, 87, 941–948. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, L.; Le Visage, C.; Weiss, P.; Yang, J. Quantifying oxygen levels in 3D bioprinted cell-laden thick constructs with perfusable microchannel networks. Polymers 2020, 12, 1260. [Google Scholar] [CrossRef] [PubMed]
- Marland, J.R.K.; Gray, M.E.; Dunare, C.; Blair, E.O.; Tsiamis, A.; Sullivan, P.; González-Fernández, E.; Greenhalgh, S.N.; Gregson, R.; Clutton, R.E. Real-time measurement of tumour hypoxia using an implantable microfabricated oxygen sensor. Sens. Bio-Sens. Res. 2020, 30, 100375. [Google Scholar] [CrossRef]
- Wolfbeis, O.S. Luminescent sensing and imaging of oxygen: Fierce competition to the Clark electrode. BioEssays 2015, 37, 921–928. [Google Scholar] [CrossRef] [PubMed]
- Otero, J.; Ulldemolins, A.; Farré, R.; Almendros, I. Oxygen Biosensors and Control in 3D Physiomimetic Experimental Models. Antioxidants 2021, 10, 1165. [Google Scholar] [CrossRef] [PubMed]
- Rivera, K.R.; Pozdin, V.A.; Young, A.T.; Erb, P.D.; Wisniewski, N.A.; Magness, S.T.; Daniele, M. Integrated phosphorescence-based photonic biosensor (iPOB) for monitoring oxygen levels in 3D cell culture systems. Biosens. Bioelectron. 2019, 123, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Boyce, M.W.; Kenney, R.M.; Truong, A.S.; Lockett, M.R. Quantifying oxygen in paper-based cell cultures with luminescent thin film sensors Young Investigators in Analytical and Bioanalytical Science. Anal. Bioanal. Chem. 2016, 408, 2985–2992. [Google Scholar] [CrossRef]
- Boyce, M.W.; Simke, W.C.; Kenney, R.M.; Lockett, M.R. Generating linear oxygen gradients across 3D cell cultures with block-layered oxygen controlled chips (BLOCCs). Anal. Methods 2019, 12, 18–24. [Google Scholar] [CrossRef] [PubMed]
- Wolff, P.; Heimann, L.; Liebsch, G.; Meier, R.J.; Gutbrod, M.; van Griensven, M.; Balmayor, E.R. Oxygen-distribution within 3-D collagen I hydrogels for bone tissue engineering. Mater. Sci. Eng. C 2019, 95, 422–427. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Liu, B.; Biosciences, M.; Biosciences, M.; Papkovsky, D.B.; Zhdanov, A.V. Polymer-encapsulated organic nanoparticles for fluorescence and photoacoustic imaging. Free Radic. Biol. Med. 2016, 101, 6570–6597. [Google Scholar] [CrossRef] [PubMed]
- Koduri, M.P.; Goudar, V.S.; Shao, Y.W.; Hunt, J.A.; Henstock, J.R.; Curran, J.; Tseng, F.G. Fluorescence-Based Nano-Oxygen Particles for Spatiometric Monitoring of Cell Physiological Conditions. ACS Appl. Mater. Interfaces 2018, 10, 30163–30171. [Google Scholar] [CrossRef]
- Vitale, C.; Fedi, A.; Marrella, A.; Varani, G.; Fato, M.; Scaglione, S. 3D perfusable hydrogel recapitulating the cancer dynamic environment to in vitro investigate metastatic colonization. Polymers 2020, 12, 2467. [Google Scholar] [CrossRef] [PubMed]
- Id, A.M.; Fedi, A.; Varani, G.; Vaccari, I.; Id, M.F.; Firpo, G.; Guida, P.; Aceto, N.; Id, S.S. High blood flow shear stress values are associated with circulating tumor cells cluster disaggregation in a multi-channel microfluidic device. PLoS ONE 2021, 16, e0245536. [Google Scholar] [CrossRef]
- Pulsoni, I.; Lubda, M.; Aiello, M.; Fedi, A.; Marzagalli, M.; von Hagen, J.; Scaglione, S. Comparison Between Franz Diffusion Cell and a novel Micro-physiological System for In Vitro Penetration Assay Using Different Skin Models. SLAS Technol. 2022, in press. [Google Scholar] [CrossRef] [PubMed]
- Lesher-Pérez, S.C.; Kim, G.A.; Kuo, C.H.; Leung, B.M.; Mong, S.; Kojima, T.; Moraes, C.; Thouless, M.D.; Luker, G.D.; Takayama, S. Dispersible oxygen microsensors map oxygen gradients in three-dimensional cell cultures. Biomater. Sci. 2017, 5, 2106–2113. [Google Scholar] [CrossRef] [PubMed]
- Dmitriev, R.I.; Papkovsky, D.B. Optical probes and techniques for O2 measurement in live cells and tissue. Cell. Mol. Life Sci. 2012, 69, 2025–2039. [Google Scholar] [CrossRef]
- Leedale, J.; Herrmann, A.; Bagnall, J.; Fercher, A.; Papkovsky, D.; Sée, V.; Bearon, R.N. Modeling the dynamics of hypoxia inducible factor-1α (HIF-1α) within single cells and 3D cell culture systems. Math. Biosci. 2014, 258, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Papkovsky, D.B.; Zhdanov, A.V. Phosphorescence based O2 sensors Essential tools for monitoring cell and tissue oxygenation and its impact on metabolism. Free Radic. Biol. Med. 2016, 101, 202–210. [Google Scholar] [CrossRef] [PubMed]
- Asphahani, F.; Zhang, M. Cellular impedance biosensors for drug screening and toxin detection. Analyst 2007, 132, 835–841. [Google Scholar] [CrossRef] [PubMed]
- Hu, N.; Wang, T.; Wang, Q.; Zhou, J.; Zou, L.; Su, K.; Wu, J.; Wang, P. High-performance beating pattern function of human induced pluripotent stem cell-derived cardiomyocyte-based biosensors for hERG inhibition recognition. Biosens. Bioelectron. 2015, 67, 146–153. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Wu, C.; Cai, H.; Hu, N.; Zhou, J.; Wang, P. Cell-Based Biosensors and Their Application in Biomedicine. Chem. Rev. 2014, 114, 6423–6461. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.A.; Yin, T.-I.; Reyes, D.; Urban, G.A. Microfluidic Chip with Integrated Electrical Cell-Impedance Sensing for Monitoring Single Cancer Cell Migration in Three-Dimensional Matrixes. Anal. Chem. 2013, 85, 11068–11076. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Wu, C.; Tu, J.; Ling, Y.; Hu, N.; Zhang, Y.; Su, K.; Wang, P. Assessment of cadmium-induced hepatotoxicity and protective effects of zinc against it using an improved cell-based biosensor. Sens. Actuators A Phys. 2013, 199, 156–164. [Google Scholar] [CrossRef]
- Zou, L.; Wu, C.; Wang, Q.; Zhou, J.; Su, K.; Li, H.; Hu, N.; Wang, P. An improved sensitive assay for the detection of PSP toxins with neuroblastoma cell-based impedance biosensor. Biosens. Bioelectron. 2015, 67, 458–464. [Google Scholar] [CrossRef]
- Heileman, K.; Daoud, J.; Tabrizian, M. Dielectric spectroscopy as a viable biosensing tool for cell and tissue characterization and analysis. Biosens. Bioelectron. 2013, 49, 348–359. [Google Scholar] [CrossRef]
- Montaño-Figueroa, A.G.; Wheelis, S.E.; Hedden, B.M.; Alshareef, N.H.; Dammanna, D.; Shaik, H.; Rodrigues, D.C.; Quevedo-Lopez, M. Detection of apoptotic and live pre-osteoblast cell line using impedance-based biosensors with variable electrode design. Biosens. Bioelectron. 2019, 128, 37–44. [Google Scholar] [CrossRef]
- Xia, N.; Huang, Y.; Cui, Z.; Liu, S.; Deng, D.; Liu, L.; Wang, J. Impedimetric biosensor for assay of caspase-3 activity and evaluation of cell apoptosis using self-assembled biotin-phenylalanine network as signal enhancer. Sens. Actuators B Chem. 2020, 320, 128436. [Google Scholar] [CrossRef]
- Anh-Nguyen, T.; Tiberius, B.; Pliquett, U.; Urban, G.A. An impedance biosensor for monitoring cancer cell attachment, spreading and drug-induced apoptosis. Sens. Actuators A Phys. 2016, 241, 231–237. [Google Scholar] [CrossRef]
- Benson, K.; Cramer, S.; Galla, H.-J. Impedance-based cell monitoring: Barrier properties and beyond. Fluids Barriers CNS 2013, 10, 5. [Google Scholar] [CrossRef]
- Elbrecht, D.H.; Long, C.J.; Hickman, J.J. Transepithelial/endothelial Electrical Resistance (TEER) theory and ap-plications for microfluidic body-on-a-chip devices. J. Rare Dis. Res. Treat. 2016, 1, 1. [Google Scholar]
- Srinivasan, B.; Kolli, A.R.; Esch, M.B.; Abaci, H.E.; Shuler, M.L.; Hickman, J.J. TEER Measurement Techniques for In Vitro Barrier Model Systems. J. Lab. Autom. 2015, 20, 107–126. [Google Scholar] [CrossRef]
- Thakur, M.; Mergel, K.; Weng, A.; Frech, S.; Gilabert-Oriol, R.; Bachran, D.; Melzig, M.F.; Fuchs, H. Real time monitoring of the cell viability during treatment with tumor-targeted toxins and saponins using impedance measurement. Biosens. Bioelectron. 2012, 35, 503–506. [Google Scholar] [CrossRef] [PubMed]
- Canali, C.; Heiskanen, A.; Martinsen, Ø.G.; Mohanty, S.; Dufva, M.; Wolff, A.; Emnéus, J. Impedance-based monitoring for tissue engineering applications. In Proceedings of the II Latin American Conference on Bioimpedance, Montevideo, Uruguay, 30 September–2 October 2015; Springer: Berlin/Heidelberg, Germany, 2016; pp. 36–39. [Google Scholar]
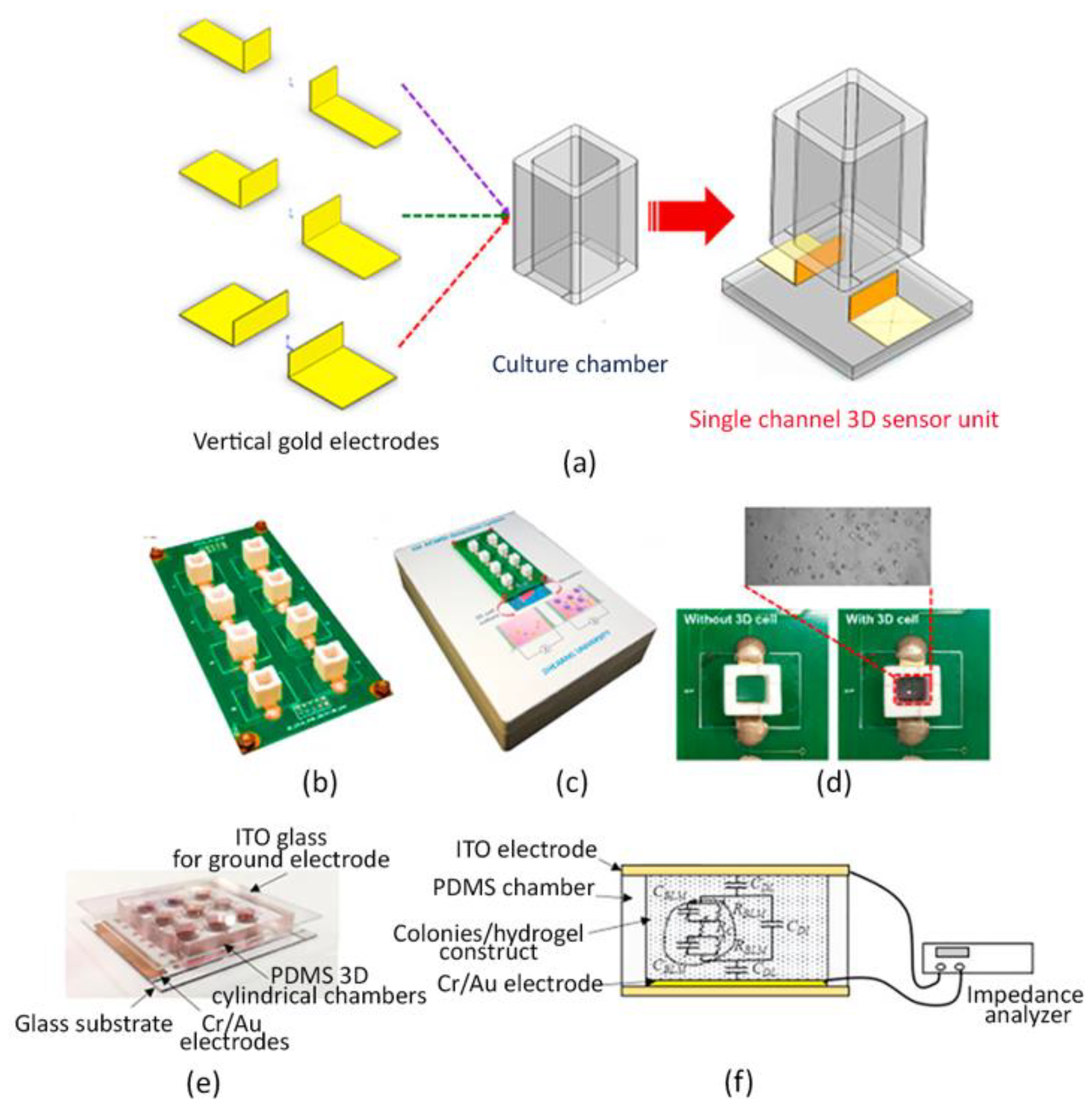
- Pan, Y.; Hu, N.; Wei, X.; Gong, L.; Zhang, B.; Wan, H.; Wang, P. 3D cell-based biosensor for cell viability and drug assessment by 3D electric cell/matrigel-substrate impedance sensing. Biosens. Bioelectron. 2019, 130, 344–351. [Google Scholar] [CrossRef] [PubMed]
- Lei, K.F.; Liu, T.K.; Tsang, N.M. Towards a high throughput impedimetric screening of chemosensitivity of cancer cells suspended in hydrogel and cultured in a paper substrate. Biosens. Bioelectron. 2018, 100, 355–360. [Google Scholar] [CrossRef] [PubMed]
- Lei, K.F.; Wu, Z.M.; Huang, C.H. Impedimetric quantification of the formation process and the chemosensitivity of cancer cell colonies suspended in 3D environment. Biosens. Bioelectron. 2015, 74, 878–885. [Google Scholar] [CrossRef]
- Jeddi, I.; Revzin, A. Sensing cell-secreted molecules. Bioanal. Rev. 2012, 4, 87–95. [Google Scholar] [CrossRef]
- Maruvada, P.; Wang, W.; Wagner, P.D.; Srivastava, S. Biomarkers in molecular medicine: Cancer detection and diagnosis. Biotechniques 2005, 38, 9–15. [Google Scholar] [CrossRef]
- Gnecchi, M.; Zhang, Z.; Ni, A.; Dzau, V.J. Paracrine mechanisms in adult stem cell signaling and therapy. Circ. Res. 2008, 103, 1204–1219. [Google Scholar] [CrossRef]
- Lander, A.D. How cells know where they are. Science 2013, 339, 923–927. [Google Scholar] [CrossRef]
- Stastna, M.; Van Eyk, J.E. Secreted proteins as a fundamental source for biomarker discovery. Proteomics 2012, 12, 722–735. [Google Scholar] [CrossRef] [PubMed]
- Borish, L.C.; Steinke, J.W. 2. Cytokines and chemokines. J. Allergy Clin. Immunol. 2003, 111, 460–475. [Google Scholar] [CrossRef] [PubMed]
- Turner, M.D.; Nedjai, B.; Hurst, T.; Pennington, D.J. Cytokines and chemokines: At the crossroads of cell signalling and inflammatory disease. Biochim. Biophys. Acta Mol. Cell Res. 2014, 1843, 2563–2582. [Google Scholar] [CrossRef] [PubMed]
- Rothenberg, E.V. Cell lineage regulators in B and T cell development. Nat. Immunol. 2007, 8, 441–444. [Google Scholar] [CrossRef]
- Brando, B.; Barnett, D.; Janossy, G.; Mandy, F.; Autran, B.; Rothe, G.; Scarpati, B.; D’Avanzo, G.; D’Hautcourt, J.L.; Lenkei, R.; et al. Cytofluorometric methods for assessing absolute numbers of cell subsets in blood. Commun. Clin. Cytom. 2000, 42, 327–346. [Google Scholar] [CrossRef]
- Cox, J.H.; Ferrari, G.; Janetzki, S. Measurement of cytokine release at the single cell level using the ELISPOT assay. Methods 2006, 38, 274–282. [Google Scholar] [CrossRef]
- Karlsson, A.C.; Martin, J.N.; Younger, S.R.; Bredt, B.M.; Epling, L.; Ronquillo, R.; Varma, A.; Deeks, S.G.; McCune, J.M.; Nixon, D.F.; et al. Comparison of the ELISPOT and cytokine flow cytometry assays for the enumeration of antigen-specific T cells. J. Immunol. Methods 2003, 283, 141–153. [Google Scholar] [CrossRef]
- Mäkitalo, B.; Andersson, M.; Areström, I.; Karlén, K.; Villinger, F.; Ansari, A.; Paulie, S.; Thorstensson, R.; Ahlborg, N. ELISpot and ELISA analysis of spontaneous, mitogen-induced and antigen-specific cytokine production in cynomolgus and rhesus macaques. J. Immunol. Methods 2002, 270, 85–97. [Google Scholar] [CrossRef]
- Steffen, M.J.; Ebersole, J.L. Sequential ELISA for cytokine levels in limited volumes of biological fluids. Biotechniques 1996, 21, 504–509. [Google Scholar] [CrossRef] [PubMed]
- Son, K.J.; Rahimian, A.; Shin, D.S.; Siltanen, C.; Patel, T.; Revzin, A. Microfluidic compartments with sensing microbeads for dynamic monitoring of cytokine and exosome release from single cells. Analyst 2016, 141, 679–688. [Google Scholar] [CrossRef] [PubMed]
- Shin, S.R.; Kilic, T.; Zhang, Y.S.; Avci, H.; Hu, N.; Kim, D.; Branco, C.; Aleman, J.; Massa, S.; Silvestri, A.; et al. Label-Free and Regenerative Electrochemical Microfluidic Biosensors for Continual Monitoring of Cell Secretomes. Adv. Sci. 2017, 4, 1600522. [Google Scholar] [CrossRef] [PubMed]
- Cappi, G.; Spiga, F.M.; Moncada, Y.; Ferretti, A.; Beyeler, M.; Bianchessi, M.; Decosterd, L.; Buclin, T.; Guiducci, C. Label-Free detection of tobramycin in serum by transmission-localized surface plasmon resonance. Anal. Chem. 2015, 87, 5278–5285. [Google Scholar] [CrossRef] [PubMed]
- Fortier, M.H.; Bonneil, E.; Goodley, P.; Thibault, P. Integrated microfluidic device for mass spectrometry-based proteomics and its application to biomarker discovery programs. Anal. Chem. 2005, 77, 1631–1640. [Google Scholar] [CrossRef] [PubMed]
- Sivanesan, A.; Izake, E.L.; Agoston, R.; Ayoko, G.A.; Sillence, M. Reproducible and label-free biosensor for the selective extraction and rapid detection of proteins in biological fluids. J. Nanobiotechnol. 2015, 13, 43. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wu, Y.; Tu, Y.; Liu, S. A reusable electrochemical immunosensor for carcinoembryonic antigen via molecular recognition of glycoprotein antibody by phenylboronic acid self-assembly layer on gold. Analyst 2008, 133, 485–492. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.J.; Bi, L.J.; Zhang, X.E.; Zhou, Y.F.; Zhang, J.B.; Chen, Y.Y.; Li, W.; Zhang, Z.P. Reversible immobilization of proteins with streptavidin affinity tags on a surface plasmon resonance biosensor chip. Anal. Bioanal. Chem. 2006, 386, 1321–1326. [Google Scholar] [CrossRef] [PubMed]
- Toner, M.; Irimia, D. Blood-on-a-chip. Annu. Rev. Biomed. Eng. 2005, 7, 77–103. [Google Scholar] [CrossRef] [PubMed]
- Blawas, A.S.; Reichert, W.M. Protein patterning. Biomaterials 1998, 19, 595–609. [Google Scholar] [CrossRef]
- Kane, R.S.; Takayama, S.; Ostuni, E.; Ingber, D.E.; Whitesides, G.M. Patterning proteins and cells using soft lithography. Biomater. Silver Jubil. Compend. 1999, 20, 161–174. [Google Scholar] [CrossRef]
- Whitesides, G.M.; Ostuni, E.; Jiang, X.; Ingber, D.E. S Oft L Ithography in B Iology. Annu. Rev. Biomed. Eng. 2001, 3, 335–373. [Google Scholar] [CrossRef] [PubMed]
- Shirasaki, Y.; Yamagishi, M.; Suzuki, N.; Izawa, K.; Nakahara, A.; Mizuno, J.; Shoji, S.; Heike, T.; Harada, Y.; Nishikomori, R.; et al. Real-time single-cell imaging of protein secretion. Sci. Rep. 2014, 4, 4736. [Google Scholar] [CrossRef]
- Love, J.C.; Ronan, J.L.; Grotenbreg, G.M.; Van Der Veen, A.G.; Ploegh, H.L. A microengraving method for rapid selection of single cells producing antigen-specific antibodies. Nat. Biotechnol. 2006, 24, 703–707. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.H.; Chen, L.J.; Verkhoturov, S.V.; Schweikert, E.A.; Revzin, A. The use of glass substrates with bi-functional silanes for designing micropatterned cell-secreted cytokine immunoassays. Biomaterials 2011, 32, 5478–5488. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yan, J.; Revzin, A. Chapter 22 Micropatterned Biosensing Surfaces for Detection of Cell-Secreted Inflammatory Signals. In Biosensors and Molecolar Technology for Cancer Diagnostics; Herold, K.E., Rasooly, A., Eds.; CRC Press, Taylor & Francis Group: Boca Raton, FL, USA, 2012; pp. 414–440. [Google Scholar] [CrossRef]
- Zhu, H.; Stybayeva, G.; Silangcruz, J.; Yan, J.; Ramanculov, E.; Dandekar, S.; George, M.D.; Revzin, A. Detecting cytokine release from single T-cells. Anal. Chem. 2009, 81, 8150–8156. [Google Scholar] [CrossRef]
- Folch, A.; Toner, M. Microengineering of cellular interactions. Annu. Rev. Biomed. Eng. 2000, 2, 227–256. [Google Scholar] [CrossRef] [PubMed]
- Bange, A.; Halsall, H.B.; Heineman, W.R. Microfluidic immunosensor systems. Biosens. Bioelectron. 2005, 20, 2488–2503. [Google Scholar] [CrossRef]
- Berthuy, O.I.; Blum, L.J.; Marquette, C.A. Cancer-cells on chip for label-free detection of secreted molecules. Biosensors 2016, 6, 2. [Google Scholar] [CrossRef]
- Sadik, O.A.; Aluoch, A.O.; Zhou, A. Status of biomolecular recognition using electrochemical techniques. Biosens. Bioelectron. 2009, 24, 2749–2765. [Google Scholar] [CrossRef]
- Xu, X.; Zhang, S.; Chen, H.; Kong, J. Integration of electrochemistry in micro-total analysis systems for biochemical assays: Recent developments. Talanta 2009, 80, 8–18. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.S.; Aleman, J.; Shin, S.R.; Kilic, T.; Kim, D.; Shaegh, S.A.M.; Massa, S.; Riahi, R.; Chae, S.; Hu, N.; et al. Multisensor-integrated organs-on-chips platform for automated and continual in situ monitoring of organoid behaviors. Proc. Natl. Acad. Sci. USA 2017, 114, E2293–E2302. [Google Scholar] [CrossRef] [PubMed]
- Skardal, A.; Murphy, S.V.; Devarasetty, M.; Mead, I.; Kang, H.-W.; Seol, Y.-J.; Zhang, Y.S.; Shin, S.-R.; Zhao, L.; Aleman, J. Multi-tissue interactions in an integrated three-tissue organ-on-a-chip platform. Sci. Rep. 2017, 7, 8837. [Google Scholar] [CrossRef] [PubMed]
- Ortega, M.A.; Fernández-Garibay, X.; Castaño, A.G.; De Chiara, F.; Hernández-Albors, A.; Balaguer-Trias, J.; Ramón-Azcón, J. Muscle-on-a-chip with an on-site multiplexed biosensing system for in situ monitoring of secreted IL-6 and TNF-α. Lab Chip 2019, 19, 2568–2580. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Stybayeva, G.; MacAl, M.; Ramanculov, E.; George, M.D.; Dandekar, S.; Revzin, A. A microdevice for multiplexed detection of T-cell-secreted cytokines. Lab Chip 2008, 8, 2197–2205. [Google Scholar] [CrossRef]
- Ma, C.; Fan, R.; Ahmad, H.; Shi, Q.; Comin-Anduix, B.; Chodon, T.; Koya, R.C.; Liu, C.C.; Kwong, G.A.; Radu, C.G.; et al. A clinical microchip for evaluation of single immune cells reveals high functional heterogeneity in phenotypically similar T cells. Nat. Med. 2011, 17, 738–743. [Google Scholar] [CrossRef]
- Bradshaw, E.M.; Kent, S.C.; Tripuraneni, V.; Orban, T.; Ploegh, H.L.; Hafler, D.A.; Love, J.C. Concurrent detection of secreted products from human lymphocytes by microengraving: Cytokines and antigen-reactive antibodies. Clin. Immunol. 2008, 129, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhyay, P.K.; Gierahn, T.M.; Roederer, M.; Love, J.C. Single-cell technologies for monitoring immune systems. Nat. Immunol. 2014, 15, 128–135. [Google Scholar] [CrossRef]
- Riahi, R.; Shaegh, S.A.M.; Ghaderi, M.; Zhang, Y.S.; Shin, S.R.; Aleman, J.; Massa, S.; Kim, D.; Dokmeci, M.R.; Khademhosseini, A. Automated microfluidic platform of bead-based electrochemical immunosensor integrated with bioreactor for continual monitoring of cell secreted biomarkers. Sci. Rep. 2016, 6, 24598. [Google Scholar] [CrossRef]
- Zhou, Q.; Kwa, T.; Gao, Y.; Liu, Y.; Rahimian, A.; Revzin, A. On-chip regeneration of aptasensors for monitoring cell secretion. Lab Chip 2014, 14, 276–279. [Google Scholar] [CrossRef]
- Dishinger, J.F.; Kennedy, R.T. Serial immunoassays in parallel on a microfluidic chip for monitoring hormone secretion from living cells. Anal. Chem. 2007, 79, 947–954. [Google Scholar] [CrossRef]
- Cohen, N.; Sabhachandani, P.; Golberg, A.; Konry, T. Approaching near real-time biosensing: Microfluidic microsphere based biosensor for real-time analyte detection. Biosens. Bioelectron. 2015, 66, 454–460. [Google Scholar] [CrossRef]
- Xu, Y.; Wang, E. Electrochemical biosensors based on magnetic micro/nano particles. Electrochim. Acta 2012, 84, 62–73. [Google Scholar] [CrossRef]
- Son, K.J.; Gheibi, P.; Stybayeva, G.; Rahimian, A.; Revzin, A. Detecting cell-secreted growth factors in microfluidic devices using bead-based biosensors. Microsystems Nanoeng. 2017, 3, 17025. [Google Scholar] [CrossRef]
- Bini, A.; Minunni, M.; Tombelli, S.; Centi, S.; Mascini, M. Analytical performances of aptamer-based sensing for thrombin detection. Anal. Chem. 2007, 79, 3016–3019. [Google Scholar] [CrossRef]
- Ferapontova, E.E.; Olsen, E.M.; Gothelf, K.V. An RNA aptamer-based electrochemical biosensor for detection of theophylline in serum. J. Am. Chem. Soc. 2008, 130, 4256–4258. [Google Scholar] [CrossRef]
- McCauley, T.G.; Hamaguchi, N.; Stanton, M. Aptamer-based biosensor arrays for detection and quantification of biological macromolecules. Anal. Biochem. 2003, 319, 244–250. [Google Scholar] [CrossRef]
- Tombelli, S.; Minunni, M.; Luzi, E.; Mascini, M. Aptamer-based biosensors for the detection of HIV-1 Tat protein. Bioelectrochemistry 2005, 67, 135–141. [Google Scholar] [CrossRef]
- Wang, J.; Jiang, Y.; Zhou, C.; Fang, X. Aptamer-based ATP assay using a luminescent light switching complex. Anal. Chem. 2005, 77, 3542–3546. [Google Scholar] [CrossRef]
- Ruigrok, V.J.B.; Levisson, M.; Eppink, M.H.M.; Smidt, H.; van der Oost, J. Alternative affinity tools: More attractive than antibodies? Biochem. J. 2011, 436, 1–13. [Google Scholar] [CrossRef]
- Maehashi, K.; Katsura, T.; Kerman, K.; Takamura, Y.; Matsumoto, K.; Tamiya, E. Label-free protein biosensor based on aptamer-modified carbon nanotube field-effect transistors. Anal. Chem. 2007, 79, 782–787. [Google Scholar] [CrossRef]
- Liss, M.; Petersen, B.; Wolf, H.; Prohaska, E. An aptamer-based quartz crystal protein biosensor. Anal. Chem. 2002, 74, 4488–4495. [Google Scholar] [CrossRef]
- Hansen, J.A.; Wang, J.; Kawde, A.N.; Xiang, Y.; Gothelf, K.V.; Collins, G. Quantum-dot/aptamer-based ultrasensitive multi-analyte electrochemical biosensor. J. Am. Chem. Soc. 2006, 128, 2228–2229. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Lubin, A.A.; Heeger, A.J.; Plaxco, K.W. Label-Free Electronic Detection of Thrombin in Blood Serum by Using an Aptamer-Based Sensor. Angew. Chem. 2005, 117, 5592–5595. [Google Scholar] [CrossRef]
- Du, L.; Wu, C.; Peng, H.; Zou, L.; Zhao, L.; Huang, L.; Wang, P. Piezoelectric olfactory receptor biosensor prepared by aptamer-assisted immobilization. Sens. Actuators B Chem. 2013, 187, 481–487. [Google Scholar] [CrossRef]
- Liu, Y.; Tuleouva, N.; Ramanculov, E.; Revzin, A. Aptamer-based electrochemical biosensor for interferon gamma detection. Anal. Chem. 2010, 82, 8131–8136. [Google Scholar] [CrossRef]
- Kara, P.; de la Escosura-Muñiz, A.; Maltez-da Costa, M.; Guix, M.; Ozsoz, M.; Merkoçi, A. Aptamers based electrochemical biosensor for protein detection using carbon nanotubes platforms. Biosens. Bioelectron. 2010, 26, 1715–1718. [Google Scholar] [CrossRef]
- Jayasena, S.D. Aptamers: An emerging class of molecules that rival antibodies in diagnostics. Clin. Chem. 1999, 45, 1628–1650. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Zhu, X.; Lu, P.Y.; Rosato, R.R.; Tan, W.; Zu, Y. Oligonucleotide aptamers: New tools for targeted cancer therapy. Mol. Ther. Nucleic Acids 2014, 3, e182. [Google Scholar] [CrossRef]
- Shaban, S.M.; Kim, D.-H. Recent Advances in Aptamer Sensors. Sensors 2021, 21, 979. [Google Scholar] [CrossRef] [PubMed]
- Li, J.J.; Fang, X.; Tan, W. Molecular aptamer beacons for real-time protein recognition. Biochem. Biophys. Res. Commun. 2002, 292, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Piorek, B.D.; Plaxco, K.W.; Heegert, A.J. A reagentless signal-on architecture for electronic, aptamer-based sensors via target-induced strand displacement. J. Am. Chem. Soc. 2005, 127, 17990–17991. [Google Scholar] [CrossRef]
- Radi, A.E.; Acero Sánchez, J.L.; Baldrich, E.; O’Sullivan, C.K. Reagentless, reusable, ultrasensitive electrochemical molecular beacon aptasensor. J. Am. Chem. Soc. 2006, 128, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.H.; Hilton, J.P.; Lin, Q. Emerging applications of aptamers to micro- and nanoscale biosensing. Microfluid. Nanofluidics 2009, 6, 347–362. [Google Scholar] [CrossRef]
- Liu, Y.; Yan, J.; Howland, M.C.; Kwa, T.; Revzin, A. Micropatterned aptasensors for continuous monitoring of cytokine release from human leukocytes. Anal. Chem. 2011, 83, 8286–8292. [Google Scholar] [CrossRef]
- Liu, Y.; Kwa, T.; Revzin, A. Simultaneous detection of cell-secreted TNF-α and IFN-γ using micropatterned aptamer-modified electrodes. Biomaterials 2012, 33, 7347–7355. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, Y.; Matharu, Z.; Rahimian, A.; Revzin, A. Detecting multiple cell-secreted cytokines from the same aptamer-functionalized electrode. Biosens. Bioelectron. 2015, 64, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Kwa, T.; Zhou, Q.; Gao, Y.; Rahimian, A.; Kwon, L.; Liu, Y.; Revzin, A. Reconfigurable microfluidics with integrated aptasensors for monitoring intercellular communication. Lab Chip 2014, 14, 1695–1704. [Google Scholar] [CrossRef]
- Shin, S.R.; Zhang, Y.S.; Kim, D.J.; Manbohi, A.; Avci, H.; Silvestri, A.; Aleman, J.; Hu, N.; Kilic, T.; Keung, W.; et al. Aptamer-Based Microfluidic Electrochemical Biosensor for Monitoring Cell-Secreted Trace Cardiac Biomarkers. Anal. Chem. 2016, 88, 10019–10027. [Google Scholar] [CrossRef]
- Lee, J.; Mehrotra, S.; Zare-Eelanjegh, E.; Rodrigues, R.O.; Akbarinejad, A.; Ge, D.; Amato, L.; Kiaee, K.; Fang, Y.C.; Rosenkranz, A.; et al. A Heart-Breast Cancer-on-a-Chip Platform for Disease Modeling and Monitoring of Cardiotoxicity Induced by Cancer Chemotherapy. Small 2020, 2004258, 2004258. [Google Scholar] [CrossRef]
- Shin, Y.; Han, S.; Jeon, J.S.; Yamamoto, K.; Zervantonakis, I.K.; Sudo, R.; Kamm, R.D.; Chung, S. Microfluidic assay for simultaneous culture of multiple cell types on surfaces or within hydrogels. Nat. Protoc. 2012, 7, 1247–1259. [Google Scholar] [CrossRef] [PubMed]
- Marrella, A.; Buratti, P.; Markus, J.; Firpo, G.; Pesenti, M.; Landry, T.; Ayehunie, S.; Scaglione, S.; Kandarova, H.; Aiello, M. In vitro demonstration of intestinal absorption mechanisms of different sugars using 3d organotypic tissues in a fluidic device. ALTEX 2020, 37, 255–264. [Google Scholar] [CrossRef]
- Schoukroun-Barnes, L.R.; Wagan, S.; Liu, J.; Leach, J.B.; White, R.J. Biocompatible hydrogel membranes for the protection of RNA aptamer-based electrochemical sensors. Smart Biomed. Physiol. Sens. Technol. X 2013, 8719, 87190I. [Google Scholar] [CrossRef]
- Santos-Cancel, M.; White, R.J. Collagen Membranes with Ribonuclease Inhibitors for Long-Term Stability of Electrochemical Aptamer-Based Sensors Employing RNA. Anal. Chem. 2017, 89, 5598–5604. [Google Scholar] [CrossRef] [PubMed]
- Rowe, A.A.; Miller, E.A.; Plaxco, K.W. Reagentless measurement of aminoglycoside antibiotics in blood serum via an electrochemical, ribonucleic acid aptamer-based biosensor. Anal. Chem. 2010, 82, 7090–7095. [Google Scholar] [CrossRef] [PubMed]
- Ferapontova, E.E.; Gothelf, K.V. Optimization of the electrochemical RNA-aptamer based biosensor for theophylline by using a methylene blue redox label. Electroanalysis 2009, 21, 1261–1266. [Google Scholar] [CrossRef]
- Verderio, C.; Matteoli, M. ATP in neuron-glia bidirectional signalling. Brain Res. Rev. 2011, 66, 106–114. [Google Scholar] [CrossRef]
- Savtchouk, I.; Volterra, A. Gliotransmission: Beyond black-and-white. J. Neurosci. 2018, 38, 14–25. [Google Scholar] [CrossRef]
- Santos-Cancel, M.; Simpson, L.W.; Leach, J.B.; White, R.J. Direct, Real-Time Detection of Adenosine Triphosphate Release from Astrocytes in Three-Dimensional Culture Using an Integrated Electrochemical Aptamer-Based Sensor. ACS Chem. Neurosci. 2019, 10, 2070–2079. [Google Scholar] [CrossRef]
| Pre-Clinical Model | Main Advantages | Main Disadvantages |
|---|---|---|
| 2D cell cultures | Simple to use | Limited or altered cell–cell and cell–extracellular matrix (ECM) interactions |
| Cheap | Altered cell morphology, proliferation, and differentiation | |
| Standard | Overestimated drugs response | |
| Lack of metabolic gradients | ||
| Oversimplified | ||
| Animal models | High complexity | Time-consuming, laborious, expensive |
| Species-specific responses | ||
| Ethical issues | ||
| 3D cellular spheroids | Cheap | Lack of surrounding ECM |
| Metabolic gradients | Susceptibility to physical deterioration | |
| Proper cell–cell interactions | ||
| In vivo-like cell morphology and proliferation | ||
| 3D hydrogel-based tissue models | Surrounding ECM with tunable properties | Batch-to-batch variability |
| Reproduction of key mechanical and biochemical features of human tissues | Difficult to monitor cell activity with traditional tools | |
| Proper cell–cell and cell–ECM interactions |
| Biosensors to Monitor Cell Activity in 3D Hydrogel-Based Tissue Models | ||||
|---|---|---|---|---|
| Typology | Method or Technology of Detection | Advantages | Disadvantages | Ref |
| pH | Electrochemical | Fast and accurate | Require large sample volume and physical contact; provide only an average value | [85,86,87] |
| ISFETs and LAPs | Sensitive and repetitive measurements; small sample volume | Require physical contact; limited applicability with 3D scaffolds | [64,88,89,90,91,92,93,94,95,96,97,98] | |
| Optical | Low costs; absence of immune and electrical interference; non-invasive sensing method; consistent, reliable, real-time, 3D measurements | Intrinsic cytotoxicity and photobleaching (fluorescent molecules); dye leaching; laborious (fluorophores with micro- or nano-particles) | [86,99,100,101,102,103,104,105,106,107,108,109,110,111,112] | |
| Glucose | Electrochemical | High selectivity and sensitivity; low costs | Chemical and thermal instability of the enzymes, low versatility (enzyme-based); measure only culture medium concentration; limited lifetime, invasive, single-point measurement (microneedles) | [64,98,115,118,119,120,121,122,123,124,125,126,127] |
| Optical | Real-time; 3D measurements | Limited applicability | [128,129,130] | |
| Oxygen | Electrochemical | Real-time measurements; low costs | Susceptibility to contaminations (non-Clark-type); measure only culture medium concentration | [149,150,151,152,155] |
| Optical | Effective; reliable; real-time; 3D measurements | Invasive and single-point measurement (optical fiber); provide only planar measurements (sensing film); laborious (fluorophores with micro- or nano-particles) | [136,156,157,158,159,160,161,162,163,164,165,169,170,171,172] | |
| Impedance | - | Automation-compatible, label-free, real-time technology; versatility; non- invasive | May require multiparametric approach to for the interpretation of the data obtained (cause of impedance variations, such as hydrogel matrix degradation or cell migration, not detected) | [173,174,175,176,177,178,179,180,181,182,183,184,185,186,187,188,189,190] |
| Secreted molecules | Antibody-based | High specificity; quantification of molecules; small sample volume | Fast saturation limiting continuous analyses (need of reconfigurable systems or washing processes) | [205,222,225,226,227,232,237] |
| Aptamer-based | Real-time; high specificity; quantification of molecules; small sample volume; high thermal and chemical stability; high sensibility; easily editable; simple conjugation with different labels | Sensibility to serum proteins of culture media (need of samples or aptamers pretreatments) | [258,260,261,262,263,272] | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fedi, A.; Vitale, C.; Giannoni, P.; Caluori, G.; Marrella, A. Biosensors to Monitor Cell Activity in 3D Hydrogel-Based Tissue Models. Sensors 2022, 22, 1517. https://doi.org/10.3390/s22041517
Fedi A, Vitale C, Giannoni P, Caluori G, Marrella A. Biosensors to Monitor Cell Activity in 3D Hydrogel-Based Tissue Models. Sensors. 2022; 22(4):1517. https://doi.org/10.3390/s22041517
Chicago/Turabian StyleFedi, Arianna, Chiara Vitale, Paolo Giannoni, Guido Caluori, and Alessandra Marrella. 2022. "Biosensors to Monitor Cell Activity in 3D Hydrogel-Based Tissue Models" Sensors 22, no. 4: 1517. https://doi.org/10.3390/s22041517
APA StyleFedi, A., Vitale, C., Giannoni, P., Caluori, G., & Marrella, A. (2022). Biosensors to Monitor Cell Activity in 3D Hydrogel-Based Tissue Models. Sensors, 22(4), 1517. https://doi.org/10.3390/s22041517









