A Conditional GAN for Generating Time Series Data for Stress Detection in Wearable Physiological Sensor Data
Abstract
:1. Introduction
2. Related Work
2.1. Detecting Stress-Related Events from Physiological Time Series Measurement Data
2.2. Conditional GANs for Time Series Data
2.3. Data Augmentation for Physiological Time Series Measurement Data
3. Methodology
3.1. Data Description
3.2. Data Acquisition Campaign in a Controlled Laboratory Environment
Setup
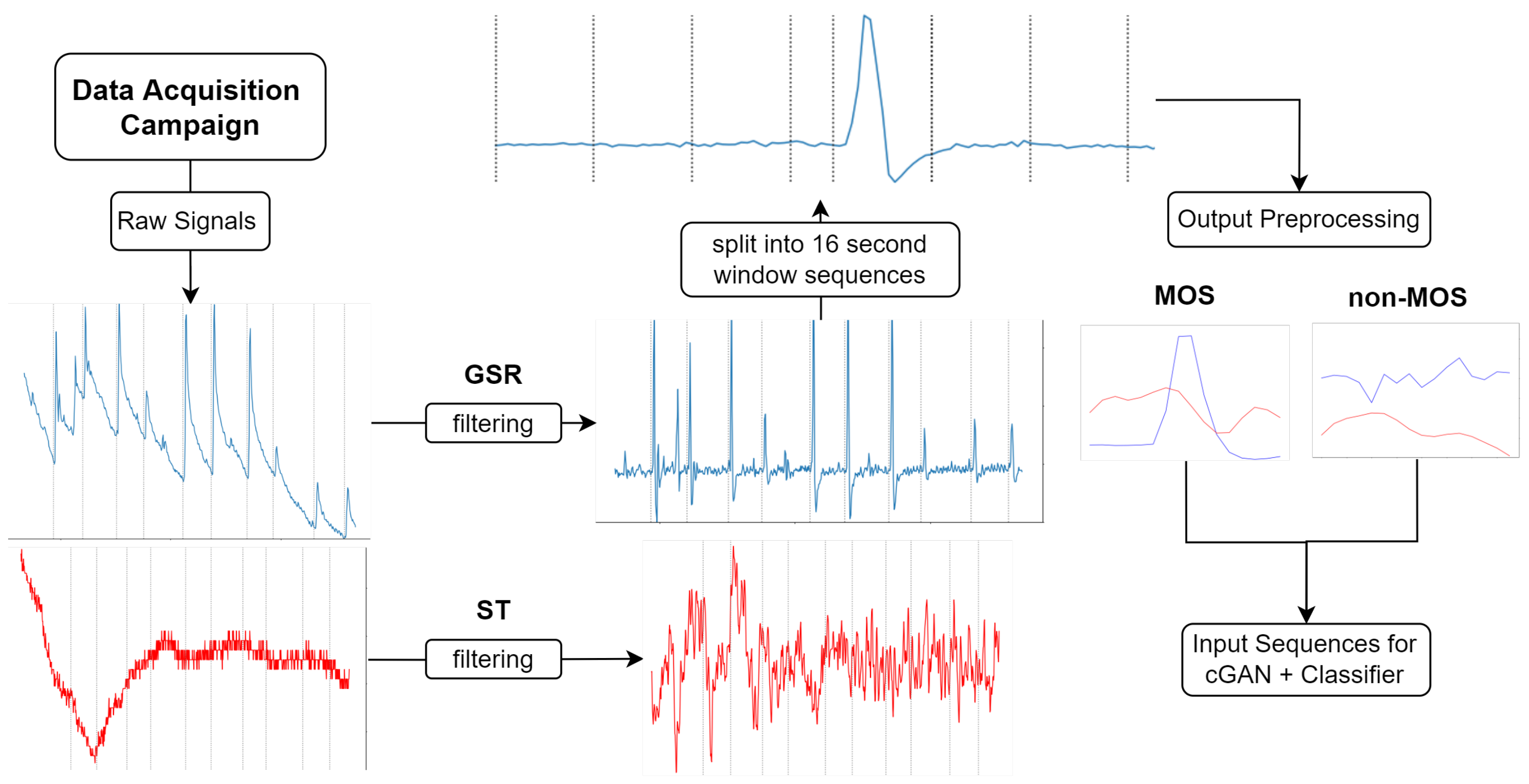
3.3. Data Processing
3.3.1. Train-Test Split
3.4. GAN Architecture and Model Training
3.4.1. Temporal Fully Convolutional Networks
3.4.2. LSTM Network
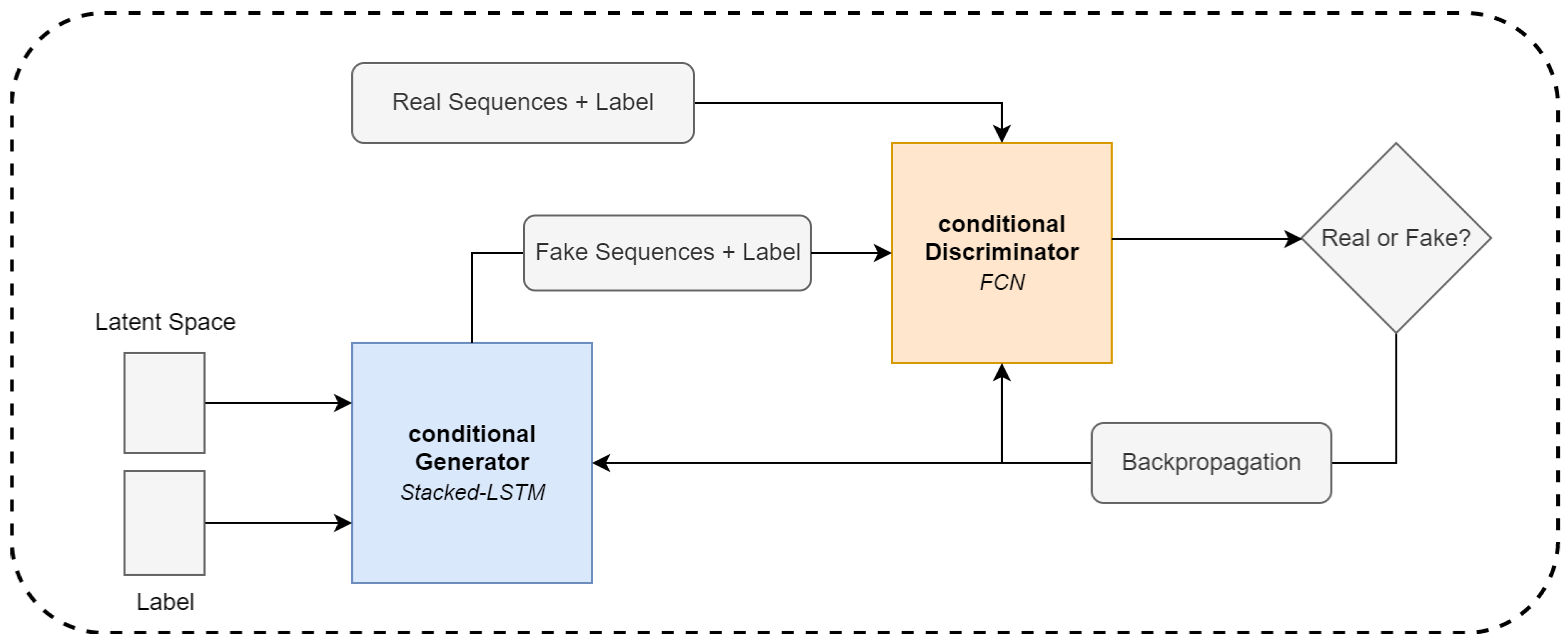
3.4.3. Conditional GAN
3.4.4. Model Training
3.5. Evaluation
- Discriminability of synthetic and real sequences, which means that we want to show that our generated data are no longer distinguishable from real data samples;
- Variety of synthetic sequences, where we want to show that our generated data cover as many different modes of our real dataset as possible;
- Quality of the generated sequences, where we want to show that the generator captured the dynamic features of our real dataset.
3.5.1. Visual Evaluation
3.5.2. Statistical Evaluation
3.5.3. Classifier Architecture
4. Experiments and Results
4.1. Generated Moments of Stress
4.2. t-sne Results
4.3. Expert Assessment Experiment
4.4. Classifying Moments of Stress
- Recurrent Conditional GAN (RCGAN) [18], where two recurrent networks as generator and discriminator are used. There is also the possibility to add label information in the generation process.
- TimeGAN [43] is a GAN framework for generated time series data. Different supervised and unsupervised loss functions are combined to generate the data.
4.4.1. Train on Generated, Test on Real
4.4.2. Data Augmentation Results
4.5. Classifier Two-Sample Test
5. Discussion and Future Research
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Giannakakis, G.; Grigoriadis, D.; Giannakaki, K.; Simantiraki, O.; Roniotis, A.; Tsiknakis, M. Review on psychological stress detection using biosignals. IEEE Trans. Affect. Comput. 2019, 13, 440–460. [Google Scholar] [CrossRef]
- Chrousos, G.P. Stress and disorders of the stress system. Nat. Rev. Endocrinol. 2009, 5, 374–381. [Google Scholar] [CrossRef] [PubMed]
- Cho, D.; Ham, J.; Oh, J.; Park, J.; Kim, S.; Lee, N.K.; Lee, B. Detection of Stress Levels from Biosignals Measured in Virtual Reality Environments Using a Kernel-Based Extreme Learning Machine. Sensors 2017, 17, 2435. [Google Scholar] [CrossRef]
- Shorten, C.; Khoshgoftaar, T.M. A survey on image data augmentation for deep learning. J. Big Data 2019, 6, 60. [Google Scholar] [CrossRef]
- Zhu, J.Y.; Park, T.; Isola, P.; Efros, A.A. Unpaired Image-to-Image Translation using Cycle-Consistent Adversarial Networks. arXiv 2017, arXiv:1703.10593. [Google Scholar]
- Mirza, M.; Osindero, S. Conditional generative adversarial nets. arXiv 2014, arXiv:1411.1784. [Google Scholar]
- Karras, T.; Laine, S.; Aila, T. A Style-Based Generator Architecture for Generative Adversarial Networks. arXiv 2018, arXiv:1812.04948. [Google Scholar]
- Frid-Adar, M.; Klang, E.; Amitai, M.; Goldberger, J.; Greenspan, H. Synthetic data augmentation using GAN for improved liver lesion classification. In Proceedings of the 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), Washington, DC, USA, 4–7 April 2018; IEEE: Piscataway, NJ, USA, 2018; pp. 289–293. [Google Scholar]
- Ismail Fawaz, H.; Forestier, G.; Weber, J.; Idoumghar, L.; Muller, P.A. Deep learning for time series classification: A review. Data Min. Knowl. Discov. 2019, 33, 917–963. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Yan, W.; Oates, T. Time Series Classification from Scratch with Deep Neural Networks: A Strong Baseline. arXiv 2016, arXiv:cs.LG/1611.06455. [Google Scholar]
- Can, Y.S.; Arnrich, B.; Ersoy, C. Stress detection in daily life scenarios using smart phones and wearable sensors: A survey. J. Biomed. Inform. 2019, 92, 103139. [Google Scholar] [CrossRef]
- Kyriakou, K.; Resch, B.; Sagl, G.; Petutschnig, A.; Werner, C.; Niederseer, D.; Liedlgruber, M.; Wilhelm, F.H.; Osborne, T.; Pykett, J. Detecting Moments of Stress from Measurements of Wearable Physiological Sensors. Sensors 2019, 19, 3805. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Castaldo, R.; Montesinos, L.; Melillo, P.; Massaro, S.; Pecchia, L. To What Extent Can We Shorten HRV Analysis in Wearable Sensing? A Case Study on Mental Stress Detection. In EMBEC & NBC 2017; Springer: Singapore, 2017; pp. 643–646. [Google Scholar]
- de Santos Sierra, A.; Ávila, C.S.; Casanova, J.G.; del Pozo, G.B.; Vera, V.J. Two stress detection schemes based on physiological signals for real-time applications. In Proceedings of the 2010 Sixth International Conference on Intelligent Information Hiding and Multimedia Signal Processing, Darmstadt, Germany, 15–17 October 2010; IEEE: Piscataway, NJ, USA, 2010; pp. 364–367. [Google Scholar]
- Bobade, P.; Vani, M. Stress detection with machine learning and deep learning using multimodal physiological data. In Proceedings of the 2020 Second International Conference on Inventive Research in Computing Applications (ICIRCA), Coimbatore, India, 15–17 July 2020; IEEE: Piscataway, NJ, USA, 2020; pp. 51–57. [Google Scholar]
- Albertetti, F.; Simalastar, A.; Rizzotti-Kaddouri, A. Stress detection with deep learning approaches using physiological signals. In Proceedings of the International Conference on IoT Technologies for HealthCare, Viana do Castello, Portugal, 3 December 2020; Springer International Publishing: Cham, Switzerland, 2020; pp. 95–111. [Google Scholar]
- Ramponi, G.; Protopapas, P.; Brambilla, M.; Janssen, R. T-CGAN: Conditional Generative Adversarial Network for Data Augmentation in Noisy Time Series with Irregular Sampling. arXiv 2018, arXiv:1811.08295. [Google Scholar]
- Esteban, C.; Hyland, S.L.; Rätsch, G. Real-valued (medical) time series generation with recurrent conditional gans. arXiv 2017, arXiv:1706.02633. [Google Scholar]
- Iwana, B.K.; Uchida, S. An empirical survey of data augmentation for time series classification with neural networks. PLoS ONE 2021, 16, e0254841. [Google Scholar] [CrossRef]
- Wen, Q.; Sun, L.; Yang, F.; Song, X.; Gao, J.; Wang, X.; Xu, H. Time Series Data Augmentation for Deep Learning: A Survey. In Proceedings of the Thirtieth International Joint Conference on Artificial Intelligence, Virtual Event/Montreal, Canada, 19–27 August 2021. [Google Scholar]
- Um, T.T.; Pfister, F.M.J.; Pichler, D.; Endo, S.; Lang, M.; Hirche, S.; Fietzek, U.; Kulić, D. Data augmentation of wearable sensor data for parkinson’s disease monitoring using convolutional neural networks. In Proceedings of the 19th ACM International Conference on Multimodal Interaction, Glasgow, UK, 13–17 November 2017; ACM: New York, NY, USA, 2017; pp. 216–220. [Google Scholar]
- Haradal, S.; Hayashi, H.; Uchida, S. Biosignal Data Augmentation Based on Generative Adversarial Networks. In Proceedings of the 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 18–21 July 2018; pp. 368–371. [Google Scholar] [CrossRef]
- Kiyasseh, D.; Tadesse, G.A.; Thwaites, L.; Zhu, T.; Clifton, D. PlethAugment: GAN-based PPG augmentation for medical diagnosis in low-resource settings. IEEE J. Biomed. Health Inform. 2020, 24, 3226–3235. [Google Scholar] [CrossRef] [PubMed]
- Furdui, A.; Zhang, T.; Worring, M.; Cesar, P.; El Ali, A. AC-WGAN-GP: Augmenting ECG and GSR Signals Using Conditional Generative Models for Arousal Classification. In Proceedings of the Adjunct Proceedings of the 2021 ACM International Joint Conference on Pervasive and Ubiquitous Computing and Proceedings of the 2021 ACM International Symposium on Wearable Computers, Virtual, 21–26 September 2021; Association for Computing Machinery: New York, NY, USA, 2021; pp. 21–22. [Google Scholar] [CrossRef]
- Yang, D.; Hong, S.; Jang, Y.; Zhao, T.; Lee, H. Diversity-sensitive conditional generative adversarial networks. arXiv 2019, arXiv:1901.09024. [Google Scholar]
- Empatica Inc. E4 Wristband. Available online: https://www.empatica.com/research/e4/ (accessed on 1 July 2022).
- Christopoulos, G.I.; Uy, M.A.; Yap, W.J. The Body and the Brain: Measuring Skin Conductance Responses to Understand the Emotional Experience. Organ. Res. Methods 2019, 22, 394–420. [Google Scholar] [CrossRef]
- Schumm, J.; Bächlin, M.; Setz, C.; Arnrich, B.; Roggen, D.; Tröster, G. Effect of movements on the electrodermal response after a startle event. Methods Inf. Med. 2008, 47, 186–191. [Google Scholar]
- Chênes, C.; Chanel, G.; Soleymani, M.; Pun, T. Highlight detection in movie scenes through inter-users, physiological linkage. In Social Media Retrieval; Springer: London, UK, 2013; pp. 217–237. [Google Scholar]
- Rácz, A.; Bajusz, D.; Héberger, K. Effect of dataset size and train/test split ratios in QSAR/QSPR multiclass classification. Molecules 2021, 26, 1111. [Google Scholar] [CrossRef]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets. In Advances in Neural Information Processing Systems; Curran Associates, Inc.: Red Hook, NY, USA, 2014; Volume 27, pp. 2672–2680. [Google Scholar]
- Karim, F.; Majumdar, S.; Darabi, H.; Chen, S. LSTM fully convolutional networks for time series classification. IEEE Access 2017, 6, 1662–1669. [Google Scholar] [CrossRef]
- Hochreiter, S.; Schmidhuber, J. Long short-term memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef] [PubMed]
- Pascanu, R.; Mikolov, T.; Bengio, Y. On the difficulty of training Recurrent Neural Networks. arXiv 2013, arXiv:cs.LG/1211.5063. [Google Scholar]
- Wiatrak, M.; Albrecht, S.V.; Nystrom, A. Stabilizing Generative Adversarial Networks: A Survey. arXiv 2020, arXiv:cs.LG/1910.00927. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. arXiv 2017, arXiv:cs.LG/1412.6980. [Google Scholar]
- Radford, A.; Metz, L.; Chintala, S. Unsupervised Representation Learning with Deep Convolutional Generative Adversarial Networks. arXiv 2016, arXiv:cs.LG/1511.06434. [Google Scholar]
- Borji, A. Pros and Cons of GAN Evaluation Measures. arXiv 2018, arXiv:cs.CV/1802.03446. [Google Scholar] [CrossRef] [Green Version]
- Van der Maaten, L.; Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Lopez-Paz, D.; Oquab, M. Revisiting classifier two-sample tests. arXiv 2018, arXiv:1610.06545. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Systems. 2015. Available online: tensorflow.org (accessed on 6 July 2022).
- Yoon, J.; Jarrett, D.; Van der Schaar, M. Time-series generative adversarial networks. In Advances in Neural Information Processing Systems; Curran Associates, Inc.: Red Hook, NY, USA, 2019; Volume 32. [Google Scholar]
- Arjovsky, M.; Chintala, S.; Bottou, L. Wasserstein generative adversarial networks. In Proceedings of the International Conference on Machine Learning, Sydney, Australia, 6–11 August 2017; pp. 214–223. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Settles, B. Active Learning Literature Survey 2009; Technical Report; University of Wisconsin-Madison Department of Computer Sciences: Madison, WI, USA, 2009. [Google Scholar]
- Lazarus, R.S. Emotion and Adaptation; Oxford University Press: Oxford, UK, 1991. [Google Scholar]





| FCN | Recall | Precision | F1 | Accuracy |
| baseline | 0.4881 | 0.8542 | 0.6212 | 0.84 |
| RCGAN TGTR | 0.5357 | 0.7377 | 0.6207 | 0.8382 |
| RCGAN DAug | 0.5833 | 0.7903 | 0.6712 | 0.8588 |
| TimeGAN TGTR | 0.5833 | 0.6203 | 0.6012 | 0.8088 |
| TimeGAN DAug | 0.6429 | 0.71 | 0.6750 | 0.8471 |
| Ours TGTR | 0.5238 | 0.7719 | 0.6241 | 0.84 |
| Ours DAug | 0.7262 | 0.7439 | 0.7349 | 0.8676 |
| LSTM | Recall | Precision | F1 | Accuracy |
| baseline | 0.5357 | 0.8654 | 0.6618 | 0.8647 |
| RCGAN TGTR | 0.4762 | 0.6250 | 0.5405 | 0.8000 |
| RCGAN DAug | 0.6190 | 0.7324 | 0.6709 | 0.8500 |
| TimeGAN TGTR | 0.5833 | 0.6533 | 0.6163 | 0.8206 |
| TimeGAN DAug | 0.5952 | 0.8065 | 0.6849 | 0.8647 |
| Ours TGTR | 0.6786 | 0.7600 | 0.7170 | 0.8618 |
| Ours DAug | 0.7262 | 0.8243 | 0.7721 | 0.88 |
| Accuracy | |
|---|---|
| Real/Generated | 0.4575 |
| Recall | Precision | F1 | Accuracy | |
|---|---|---|---|---|
| All Sequences | 0.7567 | 0.7814 | 0.7487 | 0.8175 |
| Real | 0.74 | 0.7019 | 0.6973 | 0.765 |
| Generated | 0.7733 | 0.8816 | 0.8065 | 0.870 |
| Neural Net | LSTM | |
|---|---|---|
| CTST LSTM-FCN | 0.6221 | 0.5903 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ehrhart, M.; Resch, B.; Havas, C.; Niederseer, D. A Conditional GAN for Generating Time Series Data for Stress Detection in Wearable Physiological Sensor Data. Sensors 2022, 22, 5969. https://doi.org/10.3390/s22165969
Ehrhart M, Resch B, Havas C, Niederseer D. A Conditional GAN for Generating Time Series Data for Stress Detection in Wearable Physiological Sensor Data. Sensors. 2022; 22(16):5969. https://doi.org/10.3390/s22165969
Chicago/Turabian StyleEhrhart, Maximilian, Bernd Resch, Clemens Havas, and David Niederseer. 2022. "A Conditional GAN for Generating Time Series Data for Stress Detection in Wearable Physiological Sensor Data" Sensors 22, no. 16: 5969. https://doi.org/10.3390/s22165969
APA StyleEhrhart, M., Resch, B., Havas, C., & Niederseer, D. (2022). A Conditional GAN for Generating Time Series Data for Stress Detection in Wearable Physiological Sensor Data. Sensors, 22(16), 5969. https://doi.org/10.3390/s22165969








