Electromyography Parameter Variations with Electrocardiography Noise
Abstract
:1. Introduction
2. Materials and Methods
2.1. Simulated EMG and ECG Dataset
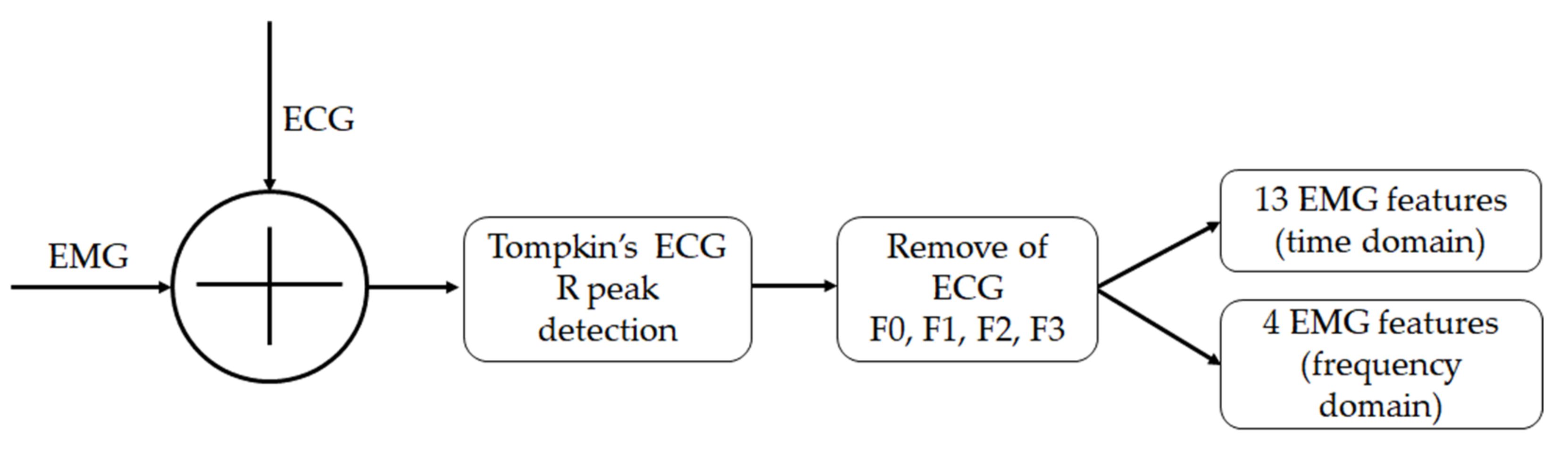
2.2. ECG R-Peak Detection
| EMG | m | ECG | Dm.1 | Dm.2 | Dm.10 | Dm.20 |
|---|---|---|---|---|---|---|
| EMG + ECG × 1 | EMG + ECG × 0.5 | EMG + ECG × 0.1 | EMG + ECG × 0.05 | |||
| EMG1 | 1 | A02C | D1.1 | D1.2 | D1.10 | D1.20 |
| 2 | A03C | D2.1 | D2.2 | D2.10 | D2.20 | |
| 3 | B02C | D3.1 | D3.2 | D3.10 | D3.20 | |
| 4 | B03C | D4.1 | D4.2 | D4.10 | D4.20 | |
| EMG2 | 5 | A02C | D5.1 | D5.2 | D5.10 | D5.20 |
| 6 | A03C | D6.1 | D6.2 | D6.10 | D6.20 | |
| 7 | B02C | D7.1 | D7.2 | D7.10 | D7.20 | |
| 8 | B03C | D8.1 | D8.2 | D8.10 | D8.20 |
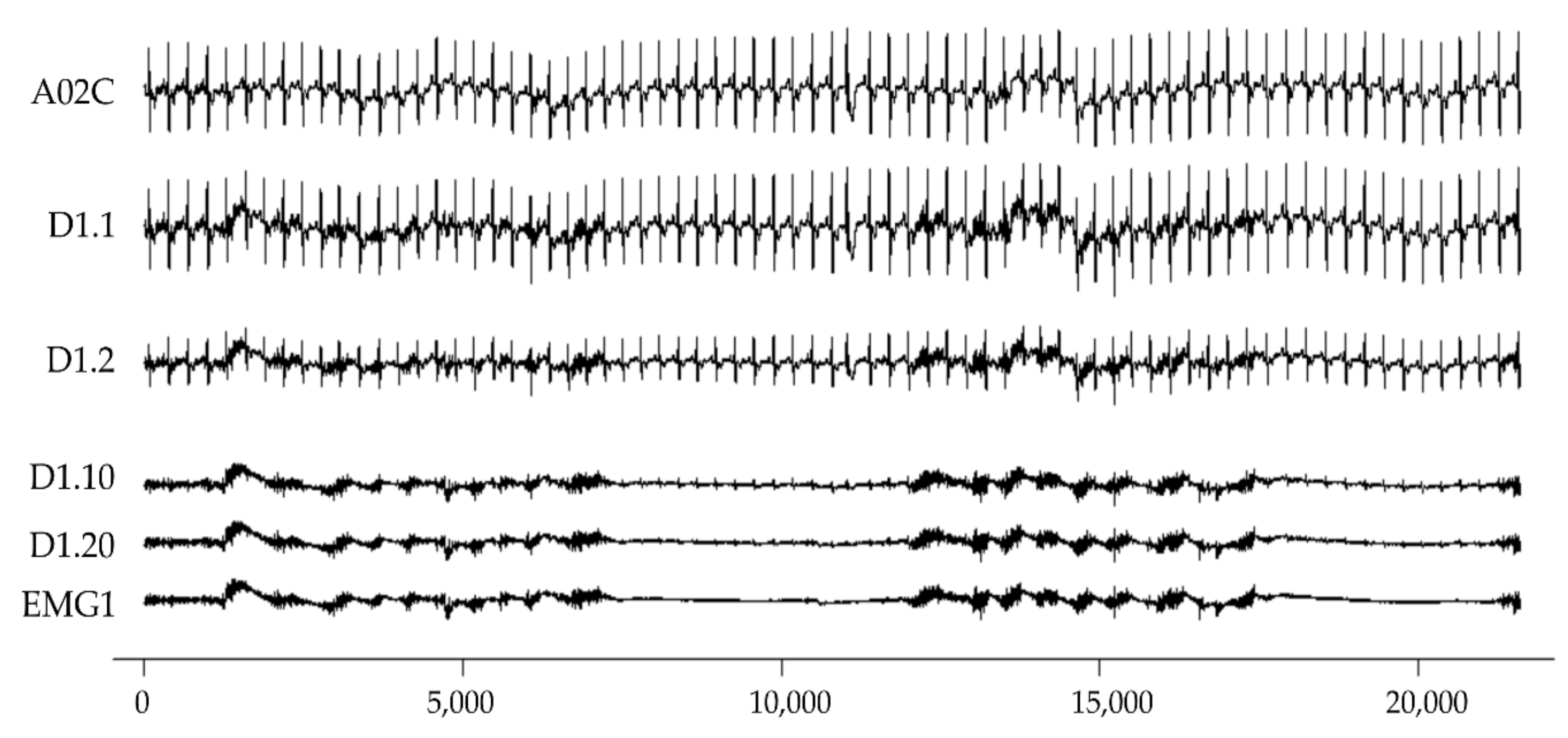
2.3. ECG Deletion

2.4. EMG Parameters
2.5. Statistics
| Number | Feature Name (Abbreviation) | Formula |
| Time-Domain | ||
| 7 | Average amplitude abs value (AAV) | |
| 8 | Standard deviation (STD) | |
| 9 | Integrated EMG (IEMG) | |
| 10 | MAV1 type-1 | |
| 11 | Simple square integral (SSI) | |
| 12 | Root mean square (RMS) | |
| 13 | LOG (log detector) | |
| 14 | Waveform length (WL) | |
| 15 | Average amplitude change (AAC) | |
| 16 | Median differential value (MDV) | |
| 17 | Difference absolute standard deviation value (DASDV) | |
| 18 | Amplitude of the first burst (AFB) | |
| 19 | Zero crossing (ZC) | |
| Number | Feature Name (Abbreviation) | Formula |
| Frequency-Domain | ||
| 20 | Total power (TTP) | |
| 21 | median frequency (MDF) | |
| 22 | Max peak frequency (PKF) | ), t = 1, …, M |
| 23 | Amplitude of peak frequency (PKF.amp) | ), t = 1, …, M |
3. Results
3.1. R-Peaks Detection Performance
3.2. Similarity for All Features in Different EMG/ECG SNR
3.3. Similarity for Time-Domain Features in Different EMG/ECG SNR
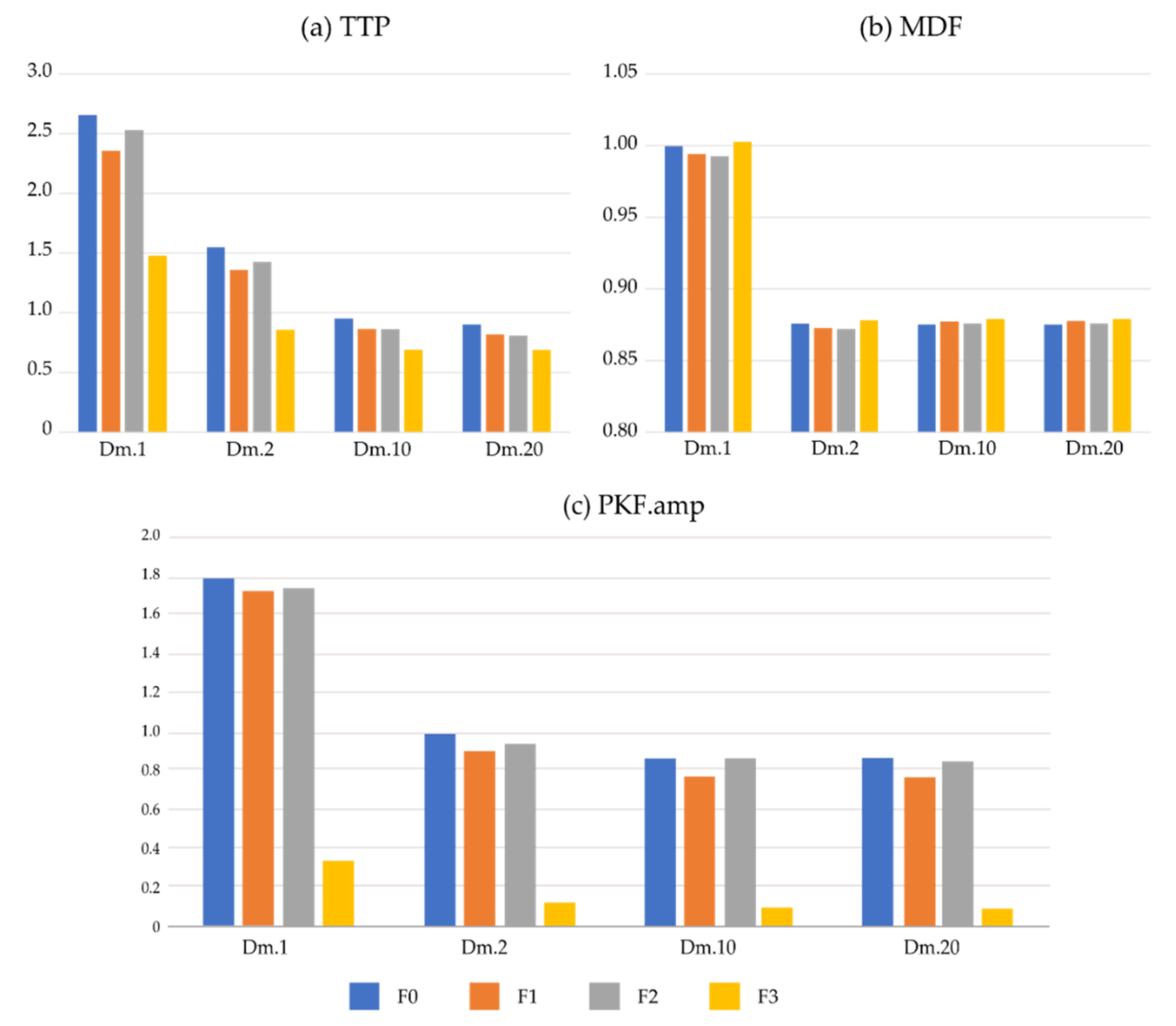
3.4. Similarity for Frequency-Domain Features in Different EMG/ECG
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
References
- Khushaba, R.N.; Al-Timemy, A.; Kodagoda, S. Influence of Multiple Dynamic Factors on the Performance of Myoelectric Pattern Recognition. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2015, 2015, 1679–1682. [Google Scholar] [CrossRef] [PubMed]
- Mengarelli, A.; Tigrini, A.; Fioretti, S.; Cardarelli, S.; Verdini, F. On the Use of Fuzzy and Permutation Entropy in Hand Gesture Characterization from EMG Signals: Parameters Selection and Comparison. Appl. Sci. 2020, 10, 7144. [Google Scholar] [CrossRef]
- Tigrini, A.; Pettinari, L.A.; Verdini, F.; Fioretti, S.; Mengarelli, A. Shoulder Motion Intention Detection through Myoelectric Pattern Recognition. IEEE Sens. Lett. 2021, 5, 6001904. [Google Scholar] [CrossRef]
- Parajuli, N.; Sreenivasan, N.; Bifulco, P.; Cesarelli, M.; Savino, S.; Niola, V.; Esposito, D.; Hamilton, T.J.; Naik, G.R.; Gunawardana, U.; et al. Real-Time EMG Based Pattern Recognition Control for Hand Prostheses: A Review on Existing Methods, Challenges and Future Implementation. Sensors 2019, 19, 4596. [Google Scholar] [CrossRef] [Green Version]
- Toledo-Pérez, D.C.; Rodríguez-Reséndiz, J.; Gómez-Loenzo, R.A.; Jauregui-Correa, J.C. Support Vector Machine-Based EMG Signal Classification Techniques: A Review. Appl. Sci. 2019, 9, 4402. [Google Scholar] [CrossRef] [Green Version]
- Phinyomark, A.; Phukpattaranont, P.; Limsakul, C. Feature Reduction and Selection for EMG Signal Classification. Expert Syst. Appl. 2012, 39, 7420–7431. [Google Scholar] [CrossRef]
- Qin, P.; Shi, X. Evaluation of Feature Extraction and Classification for Lower Limb Motion Based on SEMG Signal. Entropy 2020, 22, 852. [Google Scholar] [CrossRef]
- Tuncer, T.; Dogan, S.; Subasi, A. Surface EMG Signal Classification Using Ternary Pattern and Discrete Wavelet Transform Based Feature Extraction for Hand Movement Recognition. Biomed. Signal Process. Control 2020, 58, 101872. [Google Scholar] [CrossRef]
- Sahu, G.; Chaurasia, N.; Suwalka, P.P.; Bajaj, V.; Kumar, A. HHT Based Features for Discrimination of EMG Signals. In Proceedings of the Information Systems Design and Intelligent Applications; Mandal, J.K., Satapathy, S.C., Kumar Sanyal, M., Sarkar, P.P., Mukhopadhyay, A., Eds.; Springer: New Delhi, India, 2015; pp. 95–103. [Google Scholar]
- Chen, X.; Chen, J.; Liang, J.; Li, Y.; Courtney, C.A.; Yang, Y. Entropy-Based Surface Electromyogram Feature Extraction for Knee Osteoarthritis Classification. IEEE Access 2019, 7, 164144–164151. [Google Scholar] [CrossRef]
- Willigenburg, N.W.; Daffertshofer, A.; Kingma, I.; van Dieën, J.H. Removing ECG Contamination from EMG Recordings: A Comparison of ICA-Based and Other Filtering Procedures. J. Electromyogr. Kinesiol. 2012, 22, 485–493. [Google Scholar] [CrossRef] [Green Version]
- Abbaspour, S.; Fallah, A. Removing ECG Artifact from the Surface EMG Signal Using Adaptive Subtraction Technique. J. Biomed. Phys. Eng. 2014, 4, 33–38. [Google Scholar]
- Christov, I.; Raikova, R.; Angelova, S. Separation of Electrocardiographic from Electromyographic Signals Using Dynamic Filtration. Med. Eng. Phys. 2018, 57, 1–10. [Google Scholar] [CrossRef]
- Christov, I.; Gotchev, A.; Bortolan, G.; Neycheva, T.; Raikova, R.; Schmid, R. Separation of the Electromyographic from the Electrocardiographic Signals and Vice Versa. A Topical Review of the Dynamic Procedure. Int. J. Bioautomation 2020, 24, 289–317. [Google Scholar] [CrossRef]
- Costa Junior, J.D.; de Seixas, J.M.; Miranda de Sá, A.M.F.L. A Template Subtraction Method for Reducing Electrocardiographic Artifacts in EMG Signals of Low Intensity. Biomed. Signal Process. Control 2019, 47, 380–386. [Google Scholar] [CrossRef]
- Xu, L.; Peri, E.; Vullings, R.; Rabotti, C.; Van Dijk, J.P.; Mischi, M. Comparative Review of the Algorithms for Removal of Electrocardiographic Interference from Trunk Electromyography. Sensors 2020, 20, 4890. [Google Scholar] [CrossRef]
- Petersen, E.; Sauer, J.; Graßhoff, J.; Rostalski, P. Removing Cardiac Artifacts from Single-Channel Respiratory Electromyograms. IEEE Access 2020, 8, 30905–30917. [Google Scholar] [CrossRef]
- Modak, S.; Taha, L.Y.; Abdel-Raheem, E. A Novel Method of QRS Detection Using Time and Amplitude Thresholds with Statistical False Peak Elimination. IEEE Access 2021, 9, 46079–46092. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, Z.; Li, Z. An Improved Real-Time R-Wave Detection Efficient Algorithm in Exercise ECG Signal Analysis. J. Healthc. Eng. 2020, 2020, e8868685. [Google Scholar] [CrossRef]
- Moody, G.B.; Muldrow, W.; Mark, R.G. The MIT-BIH Noise Stress Test Database. Comput. Cardiol. 1984, 11, 381–384. [Google Scholar]
- Elgendi, M. TERMA Framework for Biomedical Signal Analysis: An Economic-Inspired Approach. Biosensors 2016, 6, 55. [Google Scholar] [CrossRef] [Green Version]
- Romagnoli, S.; Marcantoni, I.; Campanella, K.; Sbrollini, A.; Morettini, M.; Burattini, L. Ensemble Empirical Mode Decomposition for Efficient R-Peak Detection in Electrocardiograms Acquired by Portable Sensors during Sport Activity. In Proceedings of the 2021 IEEE International Symposium on Medical Measurements and Applications (MeMeA), Lausanne, Switzerland, 23–25 June 2021; pp. 1–6. [Google Scholar] [CrossRef]
- Benitez, D.S.; Gaydecki, P.A.; Zaidi, A.; Fitzpatrick, A.P. A New QRS Detection Algorithm Based on the Hilbert Transform. In Proceedings of the Computers in Cardiology, Cambridge, MA, USA,, 24–27 September 2000; 27. [Google Scholar]
- Li, H.; Tan, J. Body Sensor Network Based Context Aware QRS Detection. In Proceedings of the 2006 International Conference of the IEEE Engineering in Medicine and Biology Society, New York, NY, USA, 30 August–3 September 2006; pp. 3266–3269. [Google Scholar]
- Plesnik, E.; Malgina, O.; Tasič, J.F.; Zajc, M. Detection of the Electrocardiogram Fiducial Points in the Phase Space Using the Euclidian Distance Measure. Med. Eng. Phys. 2012, 34, 524–529. [Google Scholar] [CrossRef]
- Elgendi, M. Fast QRS Detection with an Optimized Knowledge-Based Method: Evaluation on 11 Standard ECG Databases. PLoS ONE 2013, 8, e73557. [Google Scholar] [CrossRef] [Green Version]
- Dohare, A.K.; Kumar, V.; Kumar, R. An Efficient New Method for the Detection of QRS in Electrocardiogram. Comput. Electr. Eng. 2014, 40, 1717–1730. [Google Scholar] [CrossRef]
- Yakut, Ö.; Bolat, E.D. An Improved QRS Complex Detection Method Having Low Computational Load. Biomed. Signal Process. Control 2018, 42, 230–241. [Google Scholar] [CrossRef]
- Rahul, J.; Sora, M.; Sharma, L.D. A Novel and Lightweight P, QRS, and T Peaks Detector Using Adaptive Thresholding and Template Waveform. Comput. Biol. Med. 2021, 132, 104307. [Google Scholar] [CrossRef]
- Goldberger, A.L.; Amaral, L.A.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: Components of a New Research Resource for Complex Physiologic Signals. Circulation 2000, 101, E215–E220. [Google Scholar] [CrossRef] [Green Version]
- Pan, J.; Tompkins, W.J. A Real-Time QRS Detection Algorithm. IEEE Trans. Biomed. Eng. 1985, 32, 230–236. [Google Scholar] [CrossRef]
- Detect_Rpeaks Function-RDocumentation. Available online: https://www.rdocumentation.org/packages/rsleep/versions/1.0.4/topics/detect_rpeaks (accessed on 26 July 2022).
- Zivanovic, M.; González-Izal, M. Nonstationary Harmonic Modeling for ECG Removal in Surface EMG Signals. IEEE Trans. Biomed. Eng. 2012, 59, 1633–1640. [Google Scholar] [CrossRef]
- Sanei, S.; Lee, T.K.M.; Abolghasemi, V. A New Adaptive Line Enhancer Based on Singular Spectrum Analysis. IEEE Trans. Biomed. Eng. 2012, 59, 428–434. [Google Scholar] [CrossRef]
- Sbrollini, A.; Strazza, A.; Candelaresi, S.; Marcantoni, I.; Morettini, M.; Fioretti, S.; Di Nardo, F.; Burattini, L. Surface Electromyography Low-Frequency Content: Assessment in Isometric Conditions after Electrocardiogram Cancellation by the Segmented-Beat Modulation Method. Inform. Med. Unlocked 2018, 13, 71–80. [Google Scholar] [CrossRef]
- Barrios-Muriel, J.; Romero, F.; Alonso, F.J.; Gianikellis, K. A Simple SSA-Based de-Noising Technique to Remove ECG Interference in EMG Signals. Biomed. Signal Process. Control 2016, 30, 117–126. [Google Scholar] [CrossRef]
- Oo, T.; Phukpattaranont, P. Accounting for SNR in an Algorithm Using Wavelet Transform to Remove ECG Interference from EMG Signals. Fluct. Noise Lett. 2020, 19, 2050001. [Google Scholar] [CrossRef]
- Abbaspour, S.; Fallah, A. A Combination Method for Electrocardiogram Rejection from Surface Electromyogram. Open Biomed. Eng. J. 2014, 8, 13–19. [Google Scholar] [CrossRef] [Green Version]
- Fariha, M.A.Z.; Ikeura, R.; Hayakawa, S.; Tsutsumi, S. Analysis of Pan-Tompkins Algorithm Performance with Noisy ECG Signals. J. Phys. Conf. Ser. 2020, 1532, 012022. [Google Scholar] [CrossRef]
| Author | Year | Se% | +P% |
|---|---|---|---|
| Pan-Tompkins [12] | 1985 | 74.46 | 93.67 |
| Benitez DS, et al. [23] | 2000 | 93.48 | 90.60 |
| Li, H. and Tan, [24] | 2006 | 90.66 | 87.19 |
| Plesnik et al. [25] | 2012 | 72.11 | 82.48 |
| Elgendi, M [26] | 2013 | 95.39 | 90.25 |
| Dohare, et al. [27] | 2014 | 88.20 | 89.19 |
| Yakut, Ö. and Bolat, E. D. [28] | 2018 | 93.62 | 94.52 |
| Rahul, J., et al. [29] | 2021 | 97.58 | 96.04 |
| A | |||||||
| ECG | TP | FP | FN | Se% | +P% | Median (Sec) | STD (Sec) |
| A02C | 73 | n.a. | n.a. | 100 | 100 | 0.836 | 0.042 |
| D1.1.R | 73 | 0 | 0 | 100 | 100 | 0.830 | 0.045 |
| D1.2.R | 73 | 1 | 0 | 100 | 98.6 | 0.830 | 0.087 |
| D1.10.R | 55 | 35 | 18 | 75.3 | 61.1 | 0.433 | 0.791 |
| D1.20.R | 32 | 30 | 41 | 43.0 | 51.6 | 0.294 | 2.183 |
| D5.1.R | 73 | 0 | 0 | 100 | 100 | 0.833 | 0.048 |
| D5.2.R | 73 | 3 | 0 | 100 | 96.1 | 0.833 | 0.133 |
| D5.10.R | 30 | 22 | 43 | 41.1 | 57.7 | 0.286 | 2.021 |
| D5.20.R | 24 | 33 | 49 | 32.9 | 42.1 | 0.254 | 2.028 |
| B | |||||||
| ECG | TP | FP | FN | Se% | +P% | Median (Sec) | STD (Sec) |
| A03C | 73 | n.a. | n.a. | 100 | 100 | 0.836 | 0.032 |
| D2.1.R | 73 | 0 | 0 | 100 | 100 | 0.833 | 0.031 |
| D2.2.R | 72 | 4 | 1 | 98.6 | 94.7 | 0.825 | 0.132 |
| D2.10.R | 33 | 34 | 40 | 45.2 | 49.3 | 0.313 | 2.091 |
| D2.20.R | 30 | 25 | 43 | 41.1 | 54.5 | 0.295 | 2.128 |
| D6.1.R | 73 | 0 | 0 | 100 | 100 | 0.836 | 0.034 |
| D6.2.R | 72 | 16 | 1 | 98.6 | 81.8 | 0.811 | 0.233 |
| D6.10.R | 40 | 19 | 33 | 54.8 | 67.8 | 0.255 | 1.989 |
| D6.20.R | 25 | 36 | 48 | 34.2 | 41 | 0.243 | 1.929 |
| C | |||||||
| ECG | TP | FP | FN | Se% | +P% | Median (Sec) | STD (Sec) |
| B02C | 65 | n.a. | n.a. | 100 | 100 | 0.897 | 0.296 |
| D3.1.R | 65 | 1 | 0 | 100 | 98.5 | 0.9 | 0.298 |
| D3.2.R | 65 | 1 | 0 | 100 | 98.5 | 0.897 | 0.298 |
| D3.10.R | 39 | 35 | 26 | 60 | 52.7 | 0.477 | 0.733 |
| D3.20.R | 25 | 39 | 40 | 38.5 | 39.1 | 0.291 | 2.143 |
| D7.1.R | 65 | 0 | 0 | 100 | 100 | 0.897 | 0.296 |
| D7.2.R | 65 | 4 | 0 | 100 | 94.2 | 0.881 | 0.311 |
| D7.10.R | 29 | 37 | 36 | 44.6 | 43.9 | 0.377 | 0.926 |
| D7.20.R | 19 | 39 | 46 | 29.2 | 32.8 | 0.261 | 1.971 |
| D | |||||||
| ECG | TP | FP | FN | Se% | +P% | Median (Sec) | STD (Sec) |
| B03C | 66 | n.a. | n.a. | 100 | 100 | 0.897 | 0.303 |
| D4.1.R | 65 | 1 | 1 | 98.5 | 98.5 | 0.894 | 0.305 |
| D4.2.R | 61 | 3 | 5 | 92.4 | 95.3 | 0.883 | 0.414 |
| D4.10.R | 27 | 43 | 39 | 40.9 | 38.6 | 0.305 | 1.337 |
| D4.20.R | 23 | 42 | 43 | 34.8 | 35.4 | 0.275 | 2.141 |
| D8.1.R | 63 | 3 | 3 | 95.5 | 95.5 | 0.886 | 0.372 |
| D8.2.R | 50 | 17 | 16 | 75.8 | 74.6 | 0.863 | 0.619 |
| D8.10.R | 20 | 39 | 46 | 30.3 | 33.9 | 0.258 | 1.961 |
| D8.20.R | 19 | 43 | 47 | 28.8 | 30.6 | 0.254 | 1.927 |
| Features | D1.1 | D1.2 | D1.10 | D1.20 | EMG1 |
|---|---|---|---|---|---|
| AAV | 0.2668 | 0.1555 | 0.0859 | 0.0813 | 0.0798 |
| STD | 0.3152 | 0.1665 | 0.0817 | 0.0792 | 0.0787 |
| IEMG | 5789.02 | 3362.19 | 1862.9 | 1766.47 | 1730.78 |
| MAV | 0.268 | 0.1556 | 0.0862 | 0.0817 | 0.0801 |
| MAV1 | 0.2704 | 0.1545 | 0.0821 | 0.0773 | 0.0755 |
| SSI | 3685 | 1121.82 | 303.84 | 278.63 | 270.53 |
| VAR | 0.1666 | 0.0124 | 0.0008 | 0.0008 | 0.0008 |
| ZC | 52 | 54 | 40 | 32 | 30 |
| Signals | Dm.1 | Dm.2 | Dm.10 | Dm.20 |
|---|---|---|---|---|
| ALL | 1.760 (4.569) | 1.300 (1.145) | 0.978 (0.247) | 0.944 (0.236) |
| F0 | 2.877 (6.122) | 1.763 (1.371) | 1.069 (0.043) | 1.015 (0.017) |
| F1 | 1.986 (4.927) | 1.299 (1.164) | 0.874 (0.107) | 0.852 (0.096) |
| F2 | 2.156 (4.878) | 1.438 (1.124) | 0.997 (0.035) | 0.953 (0.035) |
| F3 | 1.287 (0.652) | 0.669 (0.373) | 0.529 (0.402) | 0.528 (0.401) |
| Method Pair | Dm.1 | Dm.2 | Dm.10 | Dm.20 |
|---|---|---|---|---|
| F0–F1 | 0.015 | 0.001 | <0.001 | <0.001 |
| F0–F2 | 0.025 | 0.006 | <0.001 | <0.001 |
| F0–F3 | 0.026 | 0.006 | 0.004 | 0.006 |
| F1–F2 | <0.001 | <0.001 | 0.001 | 0.001 |
| F1–F3 | 0.059 | 0.025 | 0.073 | 0.111 |
| F2–F3 | 0.044 | 0.012 | 0.012 | 0.021 |
| (A) | ||||
| F0 | Dm.1 | Dm.2 | Dm.10 | Dm.20 |
| AAV | 2.881 | 1.767 | 1.07 | 1.022 |
| STD | 3.939 | 2.103 | 1.044 | 1.008 |
| IEMG | 2.877 | 1.763 | 1.069 | 1.021 |
| MAV1 | 2.942 | 1.792 | 1.073 | 1.023 |
| SSI | 12.872 | 3.971 | 1.12 | 1.03 |
| RMS | 3.411 | 1.93 | 1.058 | 1.015 |
| LOG | 2.48 | 1.629 | 1.082 | 1.031 |
| WL | 1.529 | 1.218 | 1.023 | 1.008 |
| AAC | 1.529 | 1.218 | 1.023 | 1.008 |
| MDV | 1.44 | 1.197 | 0.997 | 0.994 |
| DASDV | 1.466 | 1.138 | 1.006 | 1.001 |
| AFB | 22.446 | 5.943 | 1.072 | 1.005 |
| ZC | 1.542 | 1.452 | 1.148 | 1.06 |
| (B) | ||||
| F1 | Dm.1 | Dm.2 | Dm.10 | Dm.20 |
| AAV | 1.986 | 1.299 | 0.888 | 0.861 |
| STD | 2.628 | 1.558 | 0.971 | 0.94 |
| IEMG | 2.001 | 1.295 | 0.884 | 0.857 |
| MAV1 | 2.036 | 1.302 | 0.867 | 0.836 |
| SSI | 5.908 | 2.138 | 0.859 | 0.806 |
| RMS | 2.305 | 1.423 | 0.926 | 0.898 |
| LOG | 1.725 | 1.111 | 0.685 | 0.692 |
| WL | 1.179 | 0.985 | 0.796 | 0.778 |
| AAC | 1.179 | 0.985 | 0.796 | 0.778 |
| MDV | 1.033 | 0.871 | 0.675 | 0.636 |
| DASDV | 1.753 | 1.204 | 0.874 | 0.852 |
| AFB | 19.306 | 5.332 | 0.952 | 0.883 |
| ZC | 1.56 | 1.389 | 1.055 | 0.992 |
| (C) | ||||
| F2 | Dm.1 | Dm.2 | Dm.10 | Dm.20 |
| AAV | 2.156 | 1.438 | 1.017 | 0.98 |
| STD | 2.682 | 1.572 | 0.977 | 0.948 |
| IEMG | 2.228 | 1.456 | 1.016 | 0.98 |
| MAV1 | 2.276 | 1.475 | 1.014 | 0.975 |
| SSI | 6.512 | 2.381 | 0.997 | 0.932 |
| RMS | 2.418 | 1.502 | 0.998 | 0.965 |
| LOG | 1.956 | 1.387 | 1.04 | 0.998 |
| WL | 1.362 | 1.12 | 0.947 | 0.925 |
| AAC | 1.362 | 1.12 | 0.947 | 0.925 |
| MDV | 1.282 | 1.06 | 0.989 | 0.989 |
| DASDV | 2.033 | 1.326 | 0.936 | 0.911 |
| AFB | 19.239 | 5.315 | 0.948 | 0.88 |
| ZC | 1.593 | 1.397 | 1.033 | 0.953 |
| (D) | ||||
| F3 | Dm.1 | Dm.2 | Dm.10 | Dm.20 |
| AAV | 0.944 | 0.565 | 0.428 | 0.426 |
| STD | 1.566 | 0.776 | 0.631 | 0.63 |
| IEMG | 0.939 | 0.563 | 0.427 | 0.425 |
| MAV1 | 0.977 | 0.592 | 0.45 | 0.448 |
| SSI | 1.717 | 0.451 | 0.281 | 0.28 |
| RMS | 1.269 | 0.669 | 0.529 | 0.528 |
| LOG | 0.519 | 0.39 | 0.301 | 0.299 |
| WL | 1.316 | 1.086 | 0.995 | 0.993 |
| AAC | 1.316 | 1.086 | 0.995 | 0.993 |
| MDV | 1.287 | 1.082 | 0.925 | 0.923 |
| DASDV | 1.219 | 1.024 | 0.98 | 0.98 |
| AFB | 3.242 | 0.59 | 0.415 | 0.399 |
| ZC | 1.769 | 1.733 | 1.687 | 1.674 |
| PKF | Dm.1 | Dm.2 | Dm.10 | Dm.20 |
|---|---|---|---|---|
| F0 | 999.8 | 541.7 | 540.7 | 0.9 |
| F1 | 5.5 | 2.4 | 0.9 | 0.9 |
| F2 | 541.8 | 2.5 | 540.7 | 0.9 |
| F3 | 604.7 | 693.6 | 1117.7 | 667.7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chang, K.-M.; Liu, P.-T.; Wei, T.-S. Electromyography Parameter Variations with Electrocardiography Noise. Sensors 2022, 22, 5948. https://doi.org/10.3390/s22165948
Chang K-M, Liu P-T, Wei T-S. Electromyography Parameter Variations with Electrocardiography Noise. Sensors. 2022; 22(16):5948. https://doi.org/10.3390/s22165948
Chicago/Turabian StyleChang, Kang-Ming, Peng-Ta Liu, and Ta-Sen Wei. 2022. "Electromyography Parameter Variations with Electrocardiography Noise" Sensors 22, no. 16: 5948. https://doi.org/10.3390/s22165948
APA StyleChang, K.-M., Liu, P.-T., & Wei, T.-S. (2022). Electromyography Parameter Variations with Electrocardiography Noise. Sensors, 22(16), 5948. https://doi.org/10.3390/s22165948






