Deep-Learning-Based Estimation of the Spatial QRS-T Angle from Reduced-Lead ECGs
Abstract
:1. Introduction
2. Conventional Approach for QRS-T Angle Estimation
3. Deep-Learning-Based Approach for QRS-T Angle Estimation
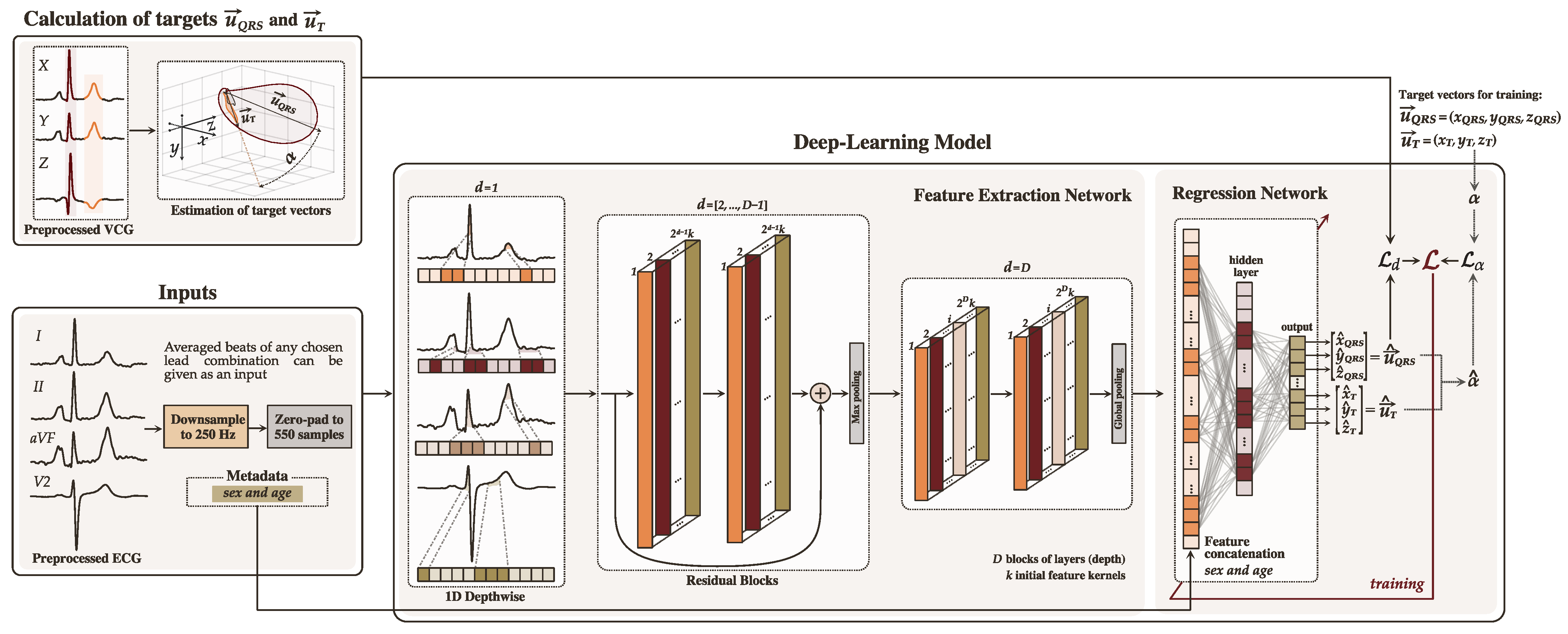
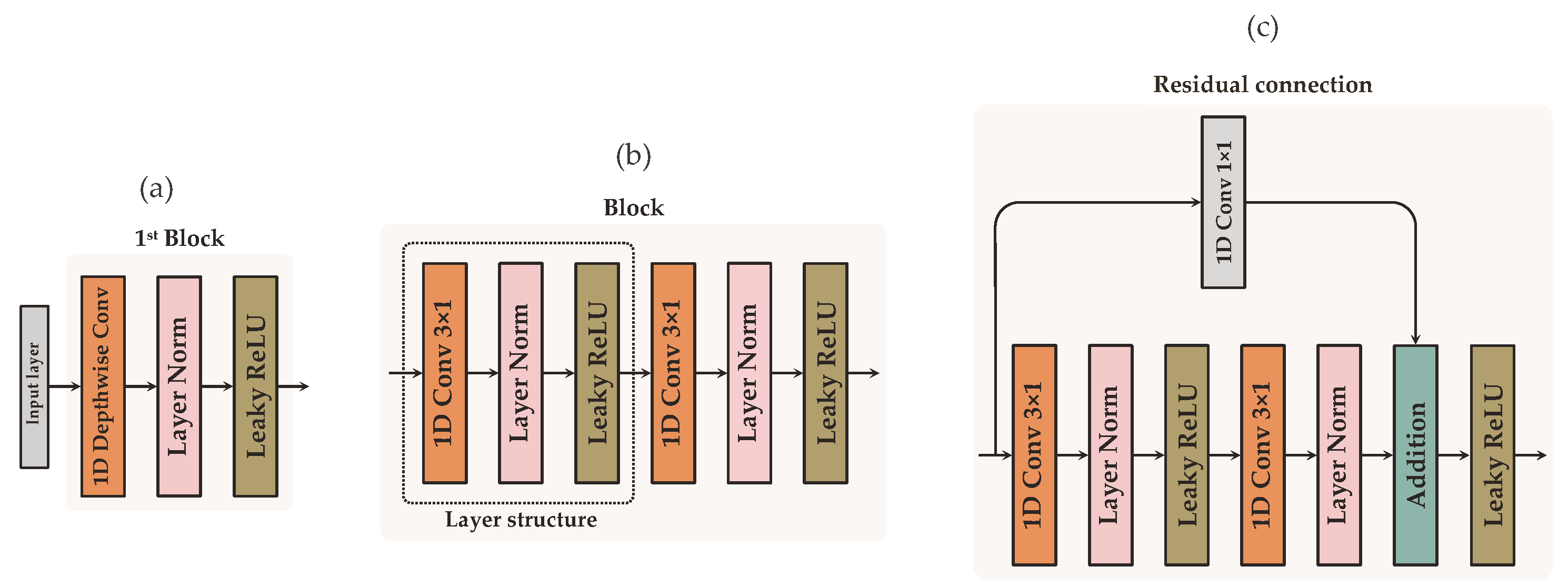
3.1. Deep Learning Model Architecture
3.1.1. Feature Extraction Network
3.1.2. Regression Network
3.2. Loss Function
3.3. Tuning of Hyperparameters
4. Data
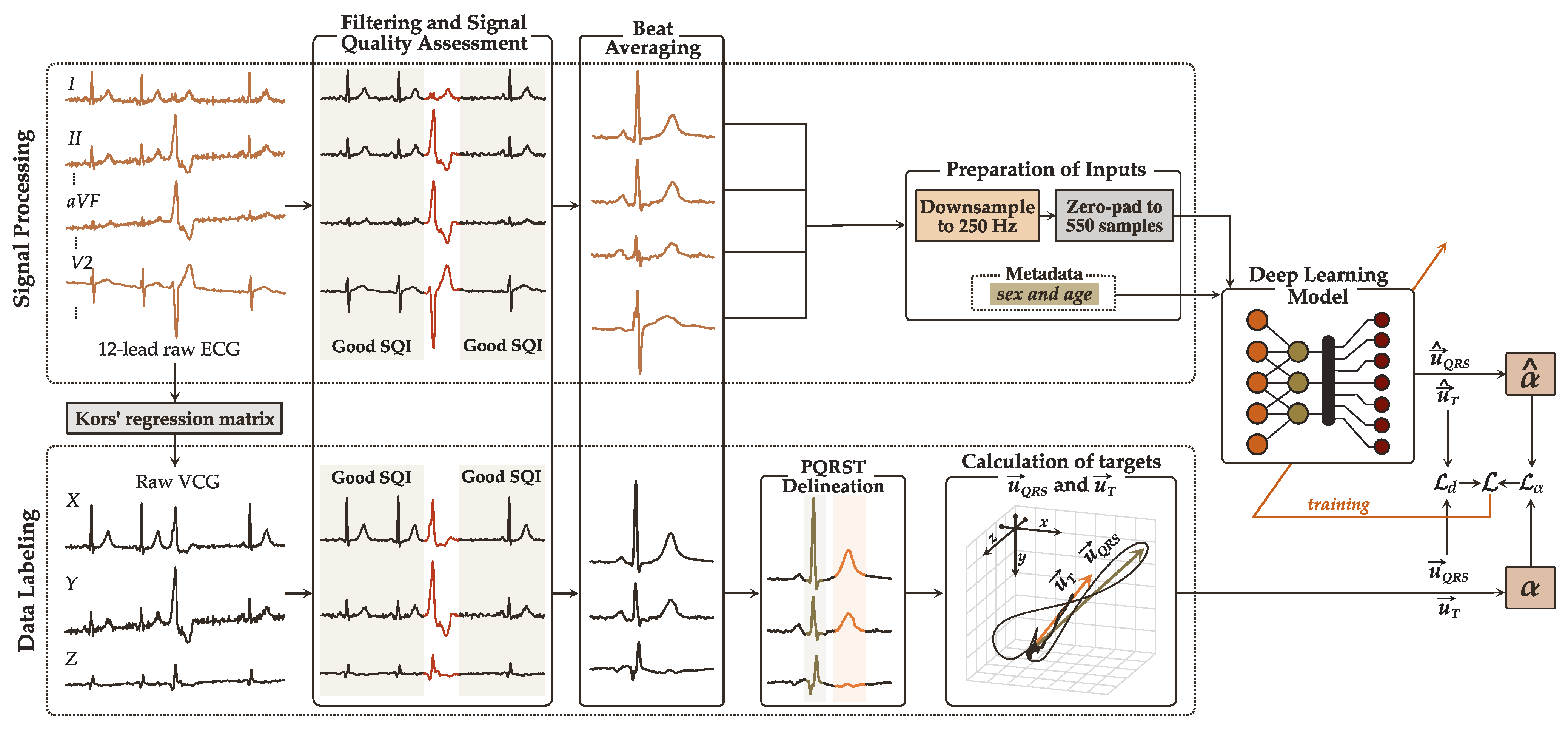
4.1. Data Preparation and Labeling
4.1.1. Signal Preprocessing
4.1.2. Data Labeling
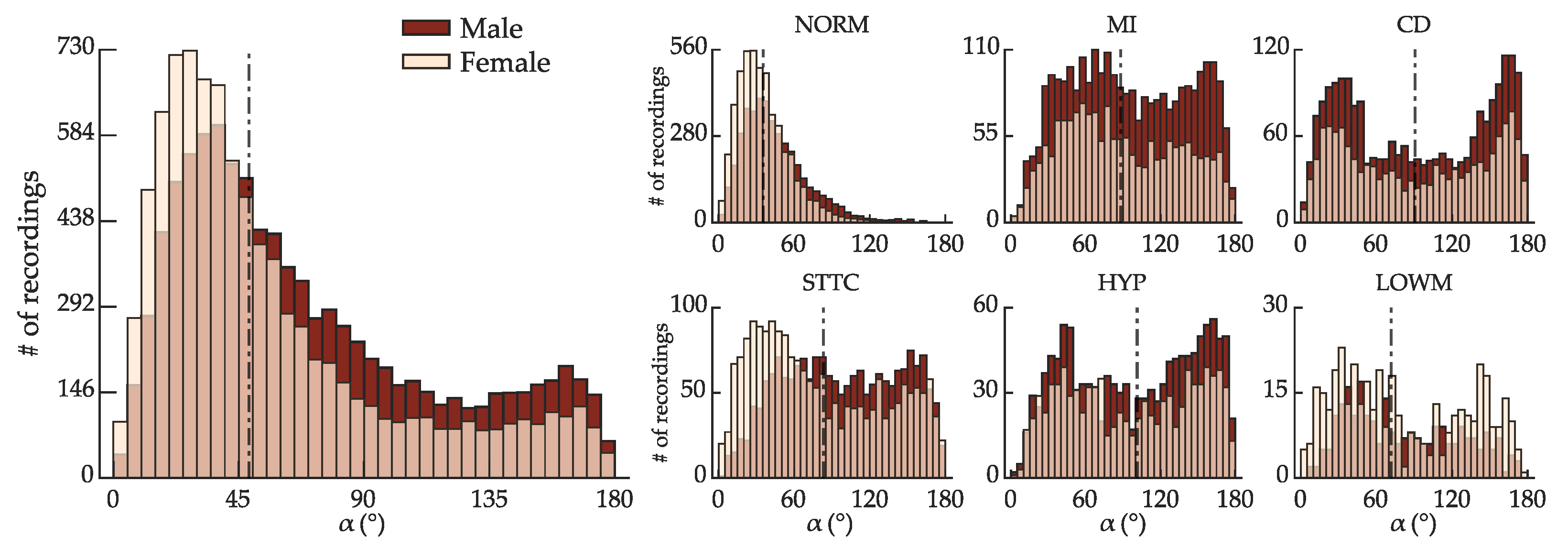
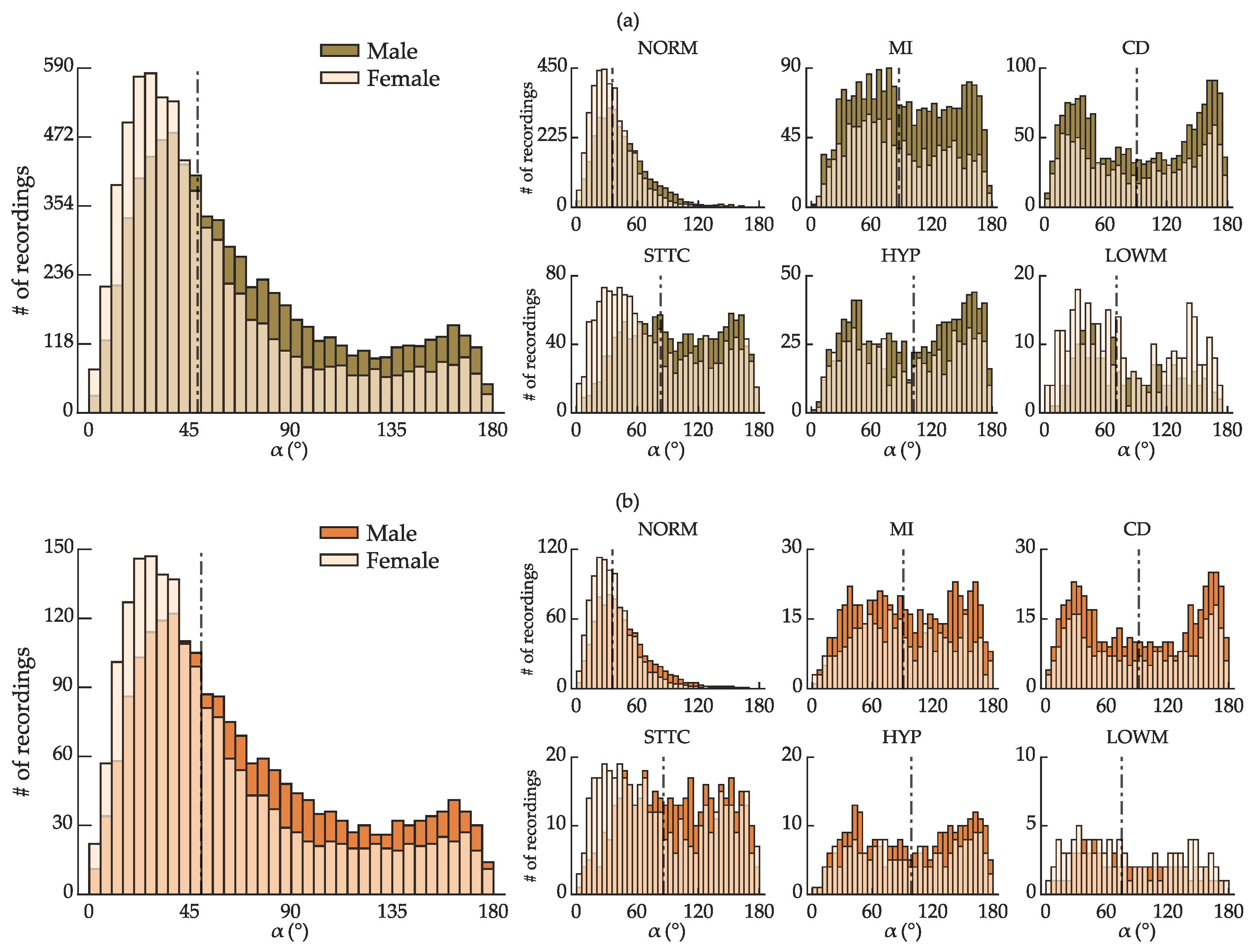
4.2. Exploratory Data Analysis
- Sex-related morphological differences in the ECG may influence the decision of the regression network (see Section 3.1.2); thus, the training set must be proportioned in terms of sex.
- Each of the morphological classes is characterized by distinctive morphological traits. Since contrastive ECG morphologies can still exhibit QRS-T angles of comparable range, the training set must include a diversity of morphologies to prevent the model from associating a specific range of QRS-T angles with just one subset of particular morphological traits.
- Randomly splitting the data without considering the uneven distribution of within specific ranges could result in a disproportionate depiction of specific ranges in the training set, leading to higher errors in other ranges.
4.3. Training and Validation Sets
5. Experiments and Performance Evaluation
5.1. Selection of Subsets of ECG Leads
5.2. Performance Metrics
6. Results
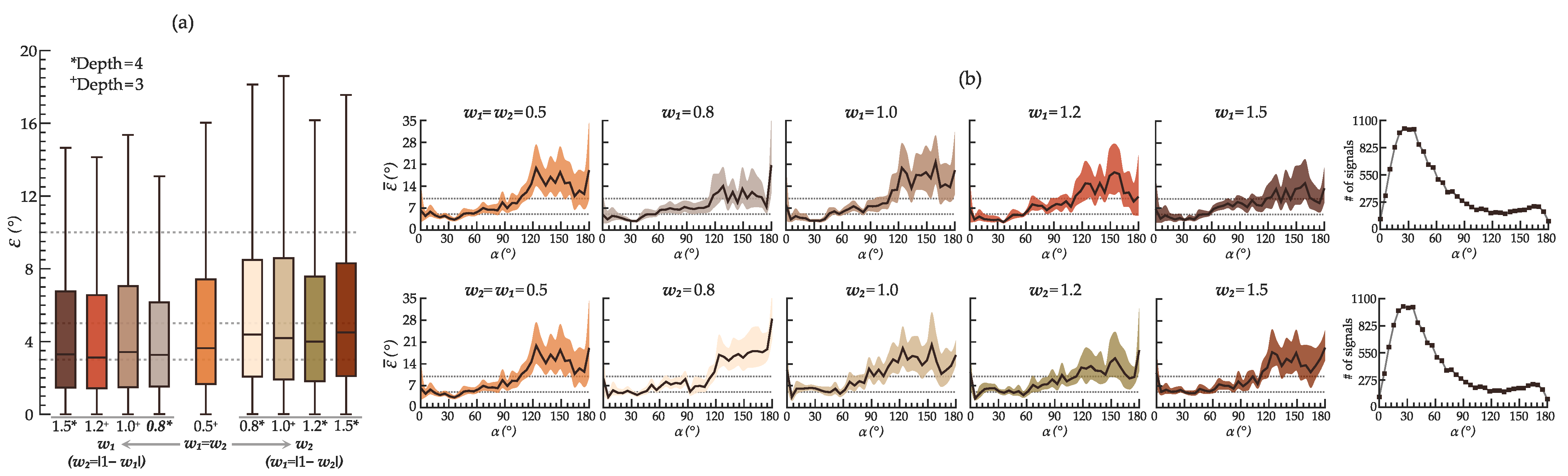
6.1. Influence of Hyperparameter Tuning on the Model Performance
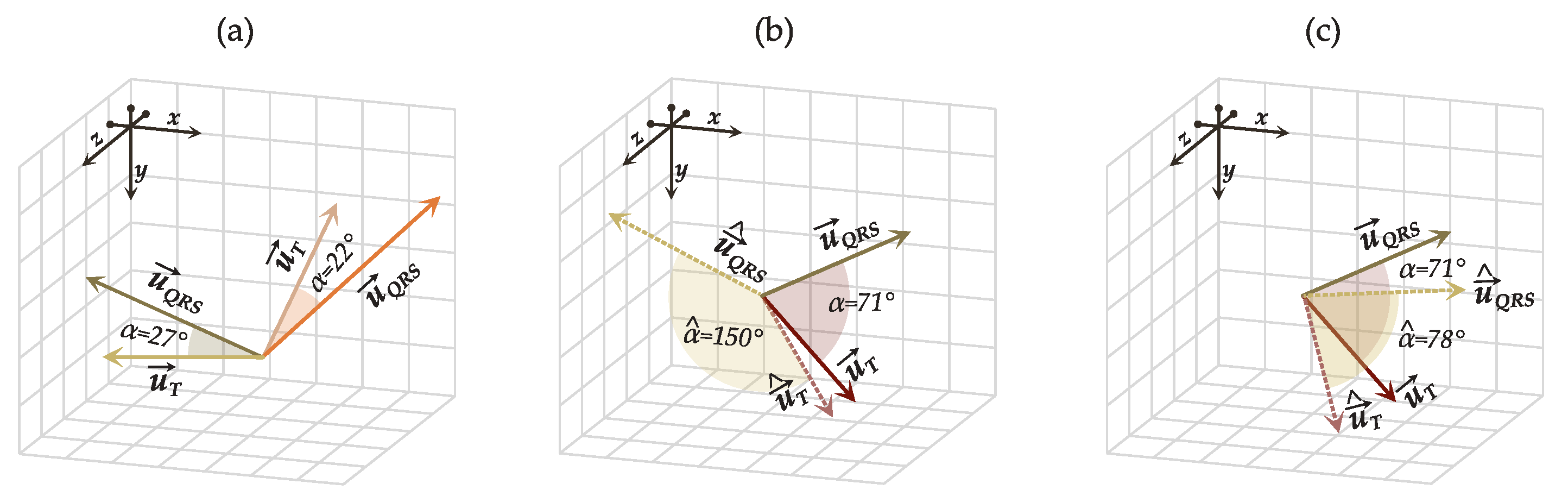
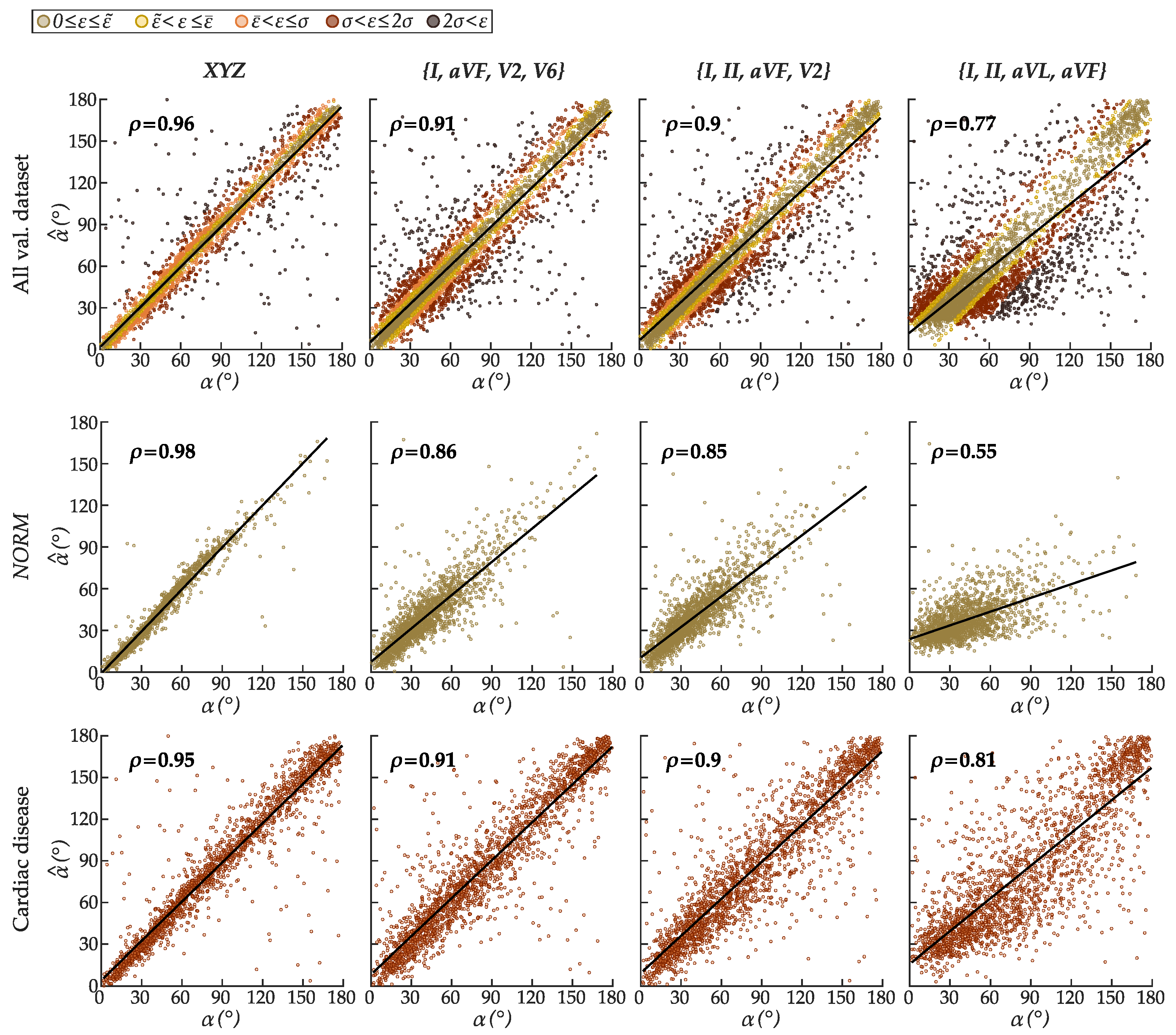
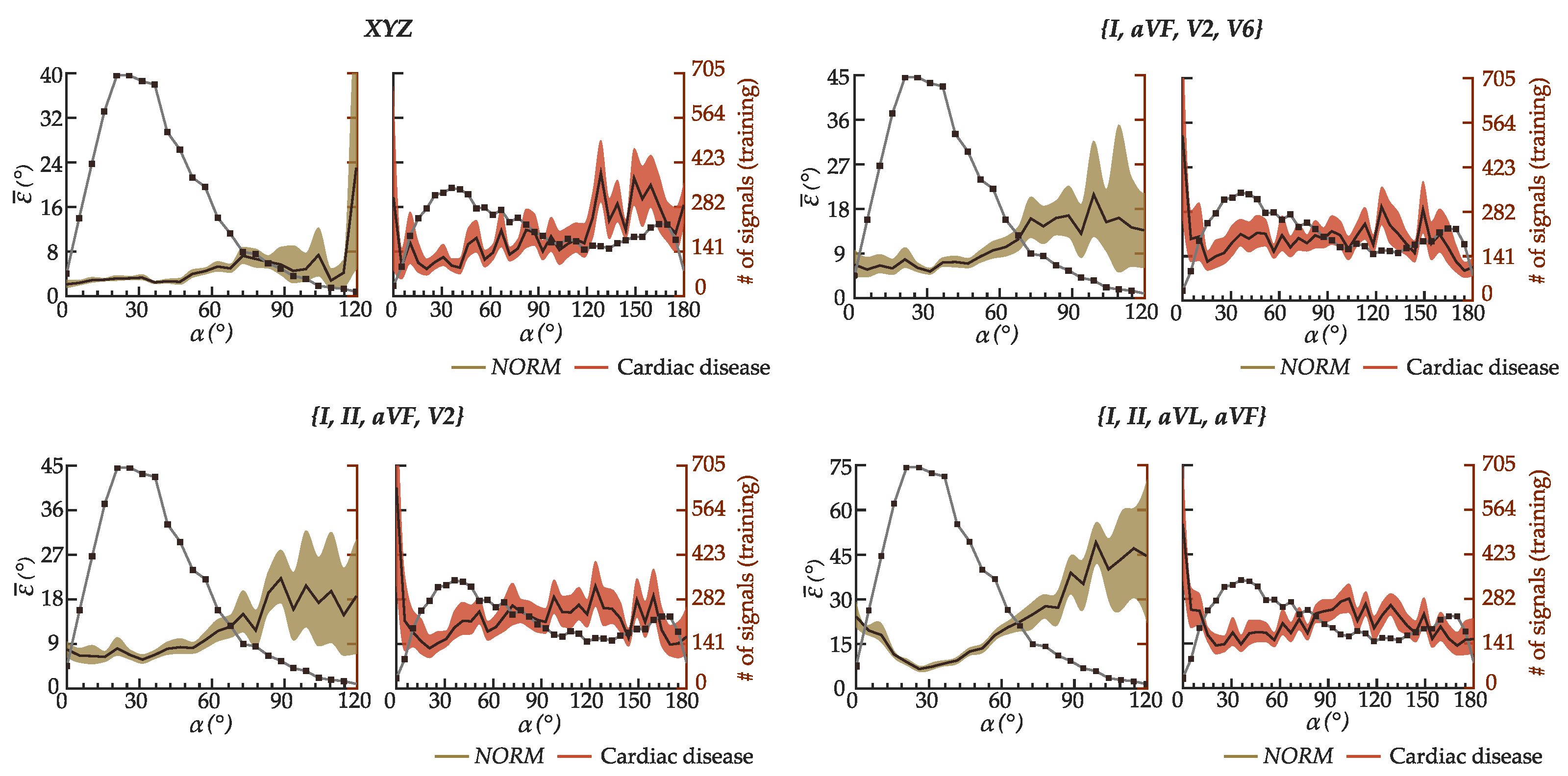
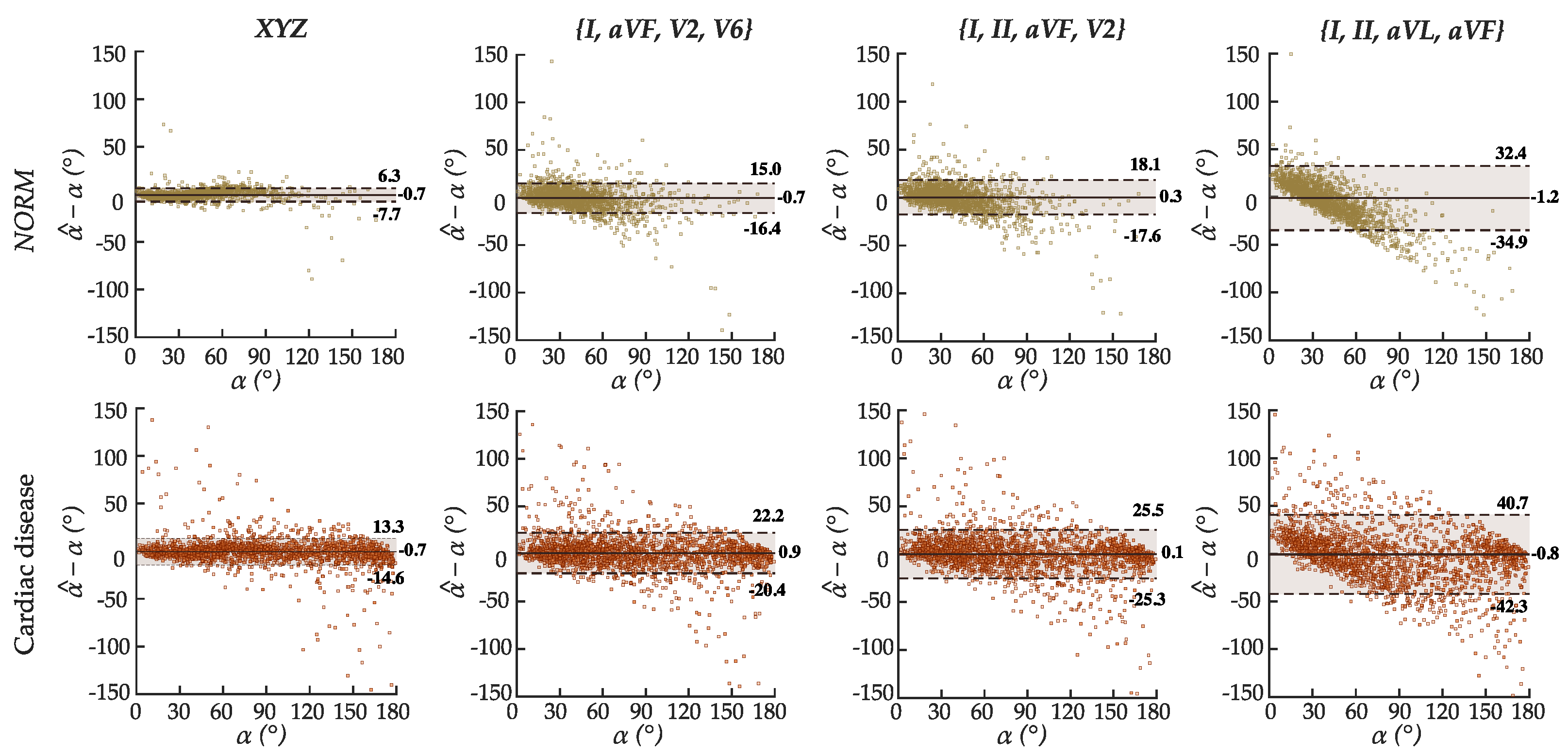
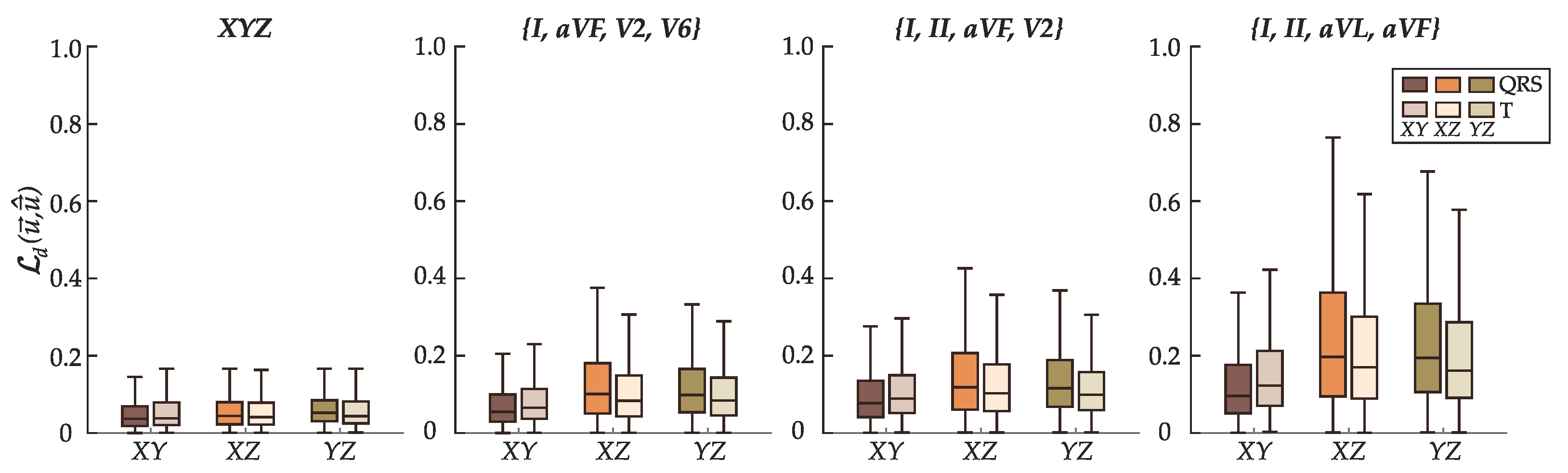
6.2. Performance of the Best Configurations on Estimating the Spatial QRS-T Angle
7. Discussion
7.1. Summary and Significance
7.2. Considerations on the Model Architecture
7.3. Considerations on the Attained Results
7.4. Suitability for ECG Consumer Healthcare Devices
7.5. Limitations and Future Directions
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| CD | Conduction Disturbance |
| CNN | Convolutional neural network |
| CNN1D | 1D convolutional neural network |
| ECG | Electrocardiogram |
| HYP | Hypertrophy |
| LOWM | Low magnitude (i.e., flat) T waves |
| MI | Myocardial Infarction |
| NORM | Normal |
| SCD | Sudden Cardiac Death |
| STTC | Change in ST-T segment |
| TCRT | Total cosine R to T |
| VCG | Vectocardiogram |
References
- Waks, J.W.; Sitlani, C.M.; Soliman, E.Z.; Kabir, M.; Ghafoori, E.; Biggs, M.L.; Henrikson, C.A.; Sotoodehnia, N.; Biering-Sørensen, T.; Agarwal, S.K.; et al. Global Electric Heterogeneity Risk Score for Prediction of Sudden Cardiac Death in the General Population. Circulation 2016, 133, 2222–2234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hayashi, M.; Shimizu, W.; Albert, C.M. The Spectrum of Epidemiology Underlying Sudden Cardiac Death. Circ. Res. 2015, 116, 1887–1906. [Google Scholar] [CrossRef]
- Srinivasan, N.T.; Schilling, R.J. Sudden Cardiac Death and Arrhythmias. Arrhythmia Electrophysiol. Rev. 2018, 7, 111. [Google Scholar] [CrossRef]
- Osadchii, O.E. Role of abnormal repolarization in the mechanism of cardiac arrhythmia. Acta Physiol. 2017, 220, 1–71. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haïssaguerre, M.; Derval, N.; Sacher, F.; Jesel, L.; Deisenhofer, I.; de Roy, L.; Pasquié, J.L.; Nogami, A.; Babuty, D.; Yli-Mayry, S.; et al. Sudden Cardiac Arrest Associated with Early Repolarization. N. Engl. J. Med. 2008, 358, 2016–2023. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kannel, W.B.; Anderson, K.; McGee, D.L.; Degatano, L.S.; Stampfer, M.J. Nonspecific electrocardiographic abnormality as a predictor of coronary heart disease: The Framingham Study. Am. Heart J. 1987, 113, 370–376. [Google Scholar] [CrossRef]
- Alders, M.; Bikker, H.; Christiaans, I. Long QT Syndrome; GeneReviews; University of Washington: Seattle, WA, USA, 20 February 2003; Updated 8 February 2018. Available online: https://www.ncbi.nlm.nih.gov/books/NBK1129/ (accessed on 29 June 2022).
- Oehler, A.; Feldman, T.; Henrikson, C.A.; Tereshchenko, L.G. QRS-T Angle: A Review. Ann. Noninvasive Electrocardiol. 2014, 19, 534–542. [Google Scholar] [CrossRef]
- Zhang, X.; Zhu, Q.; Zhu, L.; Jiang, H.; Xie, J.; Huang, W.; Xu, B. Spatial/Frontal QRS-T Angle Predicts All-Cause Mortality and Cardiac Mortality: A Meta-Analysis. PloS ONE 2015, 10, e0136174. [Google Scholar] [CrossRef]
- Chua, K.C.; Teodorescu, C.; Reinier, K.; Uy-Evanado, A.; Aro, A.L.; Nair, S.G.; Chugh, H.; Jui, J.; Chugh, S.S. Wide QRS-T Angle on the 12-Lead ECG as a Predictor of Sudden Death Beyond the LV Ejection Fraction. J. Cardiovasc. Electrophysiol. 2016, 27, 833–839. [Google Scholar] [CrossRef] [Green Version]
- Acar, B.; Yi, G.; Hnatkova, K.; Malik, M. Spatial, temporal and wavefront direction characteristics of 12-lead T-wave morphology. Med. Biol. Eng. Comput. 1999, 37, 574–584. [Google Scholar] [CrossRef]
- KORS, J.A.; HERPEN, G.V.; SITTIG, A.C.; BEMMEL, J.H.V. Reconstruction of the Frank vectorcardiogram from standard electrocardiographic leads: Diagnostic comparison of different methods. Eur. Heart J. 1990, 11, 1083–1092. [Google Scholar] [CrossRef] [PubMed]
- Jaros, R.; Martinek, R.; Danys, L. Comparison of Different Electrocardiography with Vectorcardiography Transformations. Sensors 2019, 19, 3072. [Google Scholar] [CrossRef] [Green Version]
- Maheshwari, S.; Acharyya, A.; Schiariti, M.; Puddu, P.E. Frank vectorcardiographic system from standard 12 lead ECG: An effort to enhance cardiovascular diagnosis. J. Electrocardiol. 2016, 49, 231–242. [Google Scholar] [CrossRef]
- Chen, Y.W.; Jain, L.C. (Eds.) Deep Learning in Healthcare; Springer International Publishing: Berlin/Heidelberg, Germany, 2020. [Google Scholar] [CrossRef]
- Wu, M.; Lu, Y.; Yang, W.; Wong, S.Y. A Study on Arrhythmia via ECG Signal Classification Using the Convolutional Neural Network. Front. Comput. Neurosci. 2021, 14. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, C.H.; Li, Y.S.; Hwang, B.J.; Hsiao, C.H. Detection of Atrial Fibrillation Using 1D Convolutional Neural Network. Sensors 2020, 20, 2136. [Google Scholar] [CrossRef] [Green Version]
- Hannun, A.Y.; Rajpurkar, P.; Haghpanahi, M.; Tison, G.H.; Bourn, C.; Turakhia, M.P.; Ng, A.Y. Cardiologist-level arrhythmia detection and classification in ambulatory electrocardiograms using a deep neural network. Nat. Med. 2019, 25, 65–69. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Zhang, L.; Wan, D.; Xie, Y.; Deng, H. Detection of sleep apnea syndrome by CNN based on ECG. J. Phys. Conf. Ser. 2021, 1757, 012043. [Google Scholar] [CrossRef]
- Chang, H.Y.; Yeh, C.Y.; Lee, C.T.; Lin, C.C. A Sleep Apnea Detection System Based on a One-Dimensional Deep Convolution Neural Network Model Using Single-Lead Electrocardiogram. Sensors 2020, 20, 4157. [Google Scholar] [CrossRef]
- Grande-Fidalgo, A.; Calpe, J.; Redón, M.; Millán-Navarro, C.; Soria-Olivas, E. Lead Reconstruction Using Artificial Neural Networks for Ambulatory ECG Acquisition. Sensors 2021, 21, 5542. [Google Scholar] [CrossRef]
- Sohn, J.; Yang, S.; Lee, J.; Ku, Y.; Kim, H.C. Reconstruction of 12-Lead Electrocardiogram from a Three-Lead Patch-Type Device Using a LSTM Network. Sensors 2020, 20, 3278. [Google Scholar] [CrossRef]
- Wagner, P.; Strodthoff, N.; Bousseljot, R.D.; Kreiseler, D.; Lunze, F.I.; Samek, W.; Schaeffter, T. PTB-XL, a large publicly available electrocardiography dataset. Sci. Data 2020, 7, 154. [Google Scholar] [CrossRef] [PubMed]
- Acar, B.; Koymen, H. SVD-based on-line exercise ECG signal orthogonalization. IEEE Trans. Biomed. Eng. 1999, 46, 311–321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Man, S.; Maan, A.C.; Schalij, M.J.; Swenne, C.A. Vectorcardiographic diagnostic & prognostic information derived from the 12-lead electrocardiogram: Historical review and clinical perspective. J. Electrocardiol. 2015, 48, 463–475. [Google Scholar] [CrossRef] [PubMed]
- Young, W.J.; van Duijvenboden, S.; Ramírez, J.; Jones, A.; Tinker, A.; Munroe, P.B.; Lambiase, P.D.; Orini, M. A Method to Minimise the Impact of ECG Marker Inaccuracies on the Spatial QRS-T angle: Evaluation on 1, 512 Manually Annotated ECGs. Biomed. Signal Process. Control. 2021, 64, 102305. [Google Scholar] [CrossRef]
- Hnatkova, K.; Seegers, J.; Barthel, P.; Novotny, T.; Smetana, P.; Zabel, M.; Schmidt, G.; Malik, M. Clinical value of different QRS-T angle expressions. Europace 2018, 20, 1352–1361. [Google Scholar] [CrossRef] [Green Version]
- Laguna, P.; Cortés, J.P.M.; Pueyo, E. Techniques for ventricular repolarization instability assessment from the ECG. Proc. IEEE 2016, 104, 392–415. [Google Scholar] [CrossRef] [Green Version]
- Mieszczanska, H.; Pietrasik, G.; Piotrowicz, K.; McNitt, S.; Moss, A.J.; Zareba, W. Gender-Related Differences in Electrocardiographic Parameters and Their Association With Cardiac Events in Patients After Myocardial Infarction. Am. J. Cardiol. 2008, 101, 20–24. [Google Scholar] [CrossRef] [Green Version]
- Sotobata, I.; Richman, H.; Simonson, E.; Fukomoto, A. Sex Differences in the Vectorcardiogram. Circulation 1968, 37, 438–448. [Google Scholar] [CrossRef] [Green Version]
- Chaudhry, S.; Muthurajah, J.; Lau, K.; Xiao, H.B. The effect of ageing on the frontal QRS-T angle on the 12-lead ECG. Br. J. Cardiol. 2019, 26, 157–158. [Google Scholar] [CrossRef]
- Augustauskas, R.; Lipnickas, A.; Surgailis, T. Segmentation of Drilled Holes in Texture Wooden Furniture Panels Using Deep Neural Network. Sensors 2021, 21, 3633. [Google Scholar] [CrossRef]
- Augustauskas, R.; Lipnickas, A. Pixel-level Road Pavement Defects Segmentation Based on Various Loss Functions. In Proceedings of the 2021 11th IEEE International Conference on Intelligent Data Acquisition and Advanced Computing Systems: Technology and Applications (IDAACS), Cracow, Poland, 22–25 September 2021; IEEE: Piscataway, NJ, USA, 2021. [Google Scholar] [CrossRef]
- Goldberger, A.L.; Amaral, L.A.N.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodrigues, A.; Augustauskas, R. Github Repository: DeepLearningSpatialQRS-Tangle. 2022. Available online: https://github.com/anarasr/DeepLearningSpatialQRS-Tangle (accessed on 15 July 2022).
- Orphanidou, C.; Bonnici, T.; Charlton, P.; Clifton, D.; Vallance, D.; Tarassenko, L. Signal Quality Indices for the Electrocardiogram and Photoplethysmogram: Derivation and Applications to Wireless Monitoring. IEEE J. Biomed. Health Inform. 2015, 19, 832–838. [Google Scholar] [CrossRef] [PubMed]
- Pilia, N.; Nagel, C.; Lenis, G.; Becker, S.; Dössel, O.; Loewe, A. ECGdeli-An open source ECG delineation toolbox for MATLAB. SoftwareX 2021, 13, 100639. [Google Scholar] [CrossRef]
- Kligfield, P.; Gettes, L.S.; Bailey, J.J.; Childers, R.; Deal, B.J.; Hancock, E.W.; van Herpen, G.; Kors, J.A.; Macfarlane, P.; Mirvis, D.M.; et al. Recommendations for the Standardization and Interpretation of the Electrocardiogram. J. Am. Coll. Cardiol. 2007, 49, 1109–1127. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dilaveris, P.; Gialafos, E.; Pantazis, A.; Synetos, A.; Triposkiadis, F.; Gialafos, J. The spatial QRS-T angle as a marker of ventricular repolarisation in hypertension. J. Hum. Hypertens. 2000, 15, 63–70. [Google Scholar] [CrossRef]
- Hall, P.; Horowitz, J. A simple bootstrap method for constructing nonparametric confidence bands for functions. Ann. Stat. 2013, 41, 1892–1921. [Google Scholar] [CrossRef] [Green Version]
- Efron, B. The Jackknife, the Bootstrap and Other Resampling Plans; Society for Industrial and Applied Mathematics: Philadelphia, PA, USA, 1982. [Google Scholar] [CrossRef]
- Ribeiro, A.H.; Ribeiro, M.H.; Paixão, G.M.M.; Oliveira, D.M.; Gomes, P.R.; Canazart, J.A.; Ferreira, M.P.S.; Andersson, C.R.; Macfarlane, P.W.; Meira, W.; et al. Automatic diagnosis of the 12-lead ECG using a deep neural network. Nat. Commun. 2020, 11, 1760. [Google Scholar] [CrossRef] [Green Version]
- Vondrak, J.; Penhaker, M. Review of Processing Pathological Vectorcardiographic Records for the Detection of Heart Disease. Front. Physiol. 2022, 13. [Google Scholar] [CrossRef]
- Voulgari, C.; Pagoni, S.; Tesfaye, S.; Tentolouris, N. The Spatial QRS-T Angle: Implications in Clinical Practice. Curr. Cardiol. Rev. 2013, 9, 197–210. [Google Scholar] [CrossRef]
- AliveCor, I. KardiaMobile; AliveCor Inc., Medical Device Company. Headquarters: Mountain View, CA, USA, 2022; Available online: https://store.kardia.com/products/kardiamobile (accessed on 16 July 2022).
- Rodrigues, A.; Paliakaitė, B.; Daukantas, S.; Sološenko, A.; Petrėnas, A.; Marozas, V. Personalized Evaluation of Life-threatening Conditions in Chronic Kidney Disease Patients: The Concept of Wearable Technology and Case Analysis. In Proceedings of the 15th International Joint Conference on Biomedical Engineering Systems and Technologies, Vienna, Austria, 9–11 February 2022; SCITEPRESS—Science and Technology Publications: Setúbal, Portugal, 2022. [Google Scholar] [CrossRef]
- Bacevicius, J.; Abramikas, Z.; Dvinelis, E.; Audzijoniene, D.; Petrylaite, M.; Marinskiene, J.; Staigyte, J.; Karuzas, A.; Juknevicius, V.; Jakaite, R.; et al. High Specificity Wearable Device With Photoplethysmography and Six-Lead Electrocardiography for Atrial Fibrillation Detection Challenged by Frequent Premature Contractions: DoubleCheck-AF. Front. Cardiovasc. Med. 2022, 9. [Google Scholar] [CrossRef]
- Nigusse, A.B.; Mengistie, D.A.; Malengier, B.; Tseghai, G.B.; Langenhove, L.V. Wearable Smart Textiles for Long-Term Electrocardiography Monitoring—A Review. Sensors 2021, 21, 4174. [Google Scholar] [CrossRef] [PubMed]
- Corporation, B. Bittium OmegaSnap™ ECG Electrodes; Bittium Corporation. Headquarters: Oulu, Finland, 2022; Available online: https://www.bittium.com/medical/bittium-ecg-electrodes/ (accessed on 23 June 2022).
- Partners, D. Viscero—ECG Vest; Design Parterns Ltd. Headquarters: Dublin, Ireland, 2022; Available online: https://www.designpartners.com/projects/viscero-ecg-vest/ (accessed on 23 June 2022).
- Jain, U.; Butchy, A.; Leasure, M.; Covalesky, V.; Mccormick, D.; Mintz, G. 12-Lead ECG Reconstruction via Combinatoric Inclusion of Fewer Standard ECG Leads with Implications for Lead Information and Significance. In Proceedings of the 15th International Joint Conference on Biomedical Engineering Systems and Technologies, Vienna, Austria, 9–11 February 2022; SCITEPRESS—Science and Technology Publications: Setúbal, Portugal, 2022. [Google Scholar] [CrossRef]
- Rijnbeek, P.R.; Kors, J.A.; Witsenburg, M. Minimum Bandwidth Requirements for Recording of Pediatric Electrocardiograms. Circulation 2001, 104, 3087–3090. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abdou, A.; Krishnan, S. Horizons in Single-Lead ECG Analysis From Devices to Data. Front. Signal Process. 2022, 2. [Google Scholar] [CrossRef]
- Lipton, J.A.; Nelwan, S.P.; van Domburg, R.T.; Kors, J.A.; Elhendy, A.; Schinkel, A.F.; Poldermans, D. Abnormal spatial QRS-T angle predicts mortality in patients undergoing dobutamine stress echocardiography for suspected coronary artery disease. Coron. Artery Dis. 2010, 21, 26–32. [Google Scholar] [CrossRef]
- Sweda, R.; Sabti, Z.; Strebel, I.; Kozhuharov, N.; Wussler, D.; Shrestha, S.; Flores, D.; Badertscher, P.; Lopez-Ayala, P.; Zimmermann, T.; et al. Diagnostic and prognostic values of the QRS-T angle in patients with suspected acute decompensated heart failure. ESC Heart Fail. 2020, 7, 1817–1829. [Google Scholar] [CrossRef]
- Borleffs, C.J.W.; Scherptong, R.W.; Man, S.C.; van Welsenes, G.H.; Bax, J.J.; van Erven, L.; Swenne, C.A.; Schalij, M.J. Predicting Ventricular Arrhythmias in Patients With Ischemic Heart Disease. Circ. Arrhythmia Electrophysiol. 2009, 2, 548–554. [Google Scholar] [CrossRef] [Green Version]
| Subset of Leads | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| XYZ | {I, aVF, V2, V6} | {I, II, aVF, V2} | {I, II, aVL, aVR} | ||||||||||
| Recordings | Ranges of | RMSE | RMSE | RMSE | RMSE | ||||||||
| All val. dataset | 12.2° | 5.8° | 3.3° | 17.2° | 10.3° | 6.4° | 18.4° | 11.4° | 7.3° | 25.4° | 17.9° | 12.7° | |
| 9.2° | 4.7° | 2.9° | 15.4° | 9.8° | 6.3° | 16.0° | 10.5° | 7.1° | 22.8° | 16.6° | 12.2° | ||
| NORM | 6.1° | 3.4° | 2.5° | 13.5° | 8.3° | 5.5° | 14.1° | 9.0° | 6.1° | 21.0° | 14.9° | 11.1° | |
| 4.6° | 3.0° | 2.4° | 11.0° | 7.2° | 5.1° | 11.1° | 7.6° | 5.7° | 15.2° | 11.7° | 9.8° | ||
| Cardiac disease | 16.8° | 8.7° | 4.9° | 20.5° | 12.2° | 7.3° | 21.8° | 13.7° | 8.7° | 29.1° | 20.7° | 14.3° | |
| 12.8° | 7.2° | 4.2° | 18.4° | 12.1° | 8.1° | 19.6° | 13.3° | 9.5° | 27.7° | 20.6° | 15.0° | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Santos Rodrigues, A.; Augustauskas, R.; Lukoševičius, M.; Laguna, P.; Marozas, V. Deep-Learning-Based Estimation of the Spatial QRS-T Angle from Reduced-Lead ECGs. Sensors 2022, 22, 5414. https://doi.org/10.3390/s22145414
Santos Rodrigues A, Augustauskas R, Lukoševičius M, Laguna P, Marozas V. Deep-Learning-Based Estimation of the Spatial QRS-T Angle from Reduced-Lead ECGs. Sensors. 2022; 22(14):5414. https://doi.org/10.3390/s22145414
Chicago/Turabian StyleSantos Rodrigues, Ana, Rytis Augustauskas, Mantas Lukoševičius, Pablo Laguna, and Vaidotas Marozas. 2022. "Deep-Learning-Based Estimation of the Spatial QRS-T Angle from Reduced-Lead ECGs" Sensors 22, no. 14: 5414. https://doi.org/10.3390/s22145414
APA StyleSantos Rodrigues, A., Augustauskas, R., Lukoševičius, M., Laguna, P., & Marozas, V. (2022). Deep-Learning-Based Estimation of the Spatial QRS-T Angle from Reduced-Lead ECGs. Sensors, 22(14), 5414. https://doi.org/10.3390/s22145414








