A Review on Computer Aided Diagnosis of Acute Brain Stroke
Abstract
:1. Introduction
1.1. Review Objective
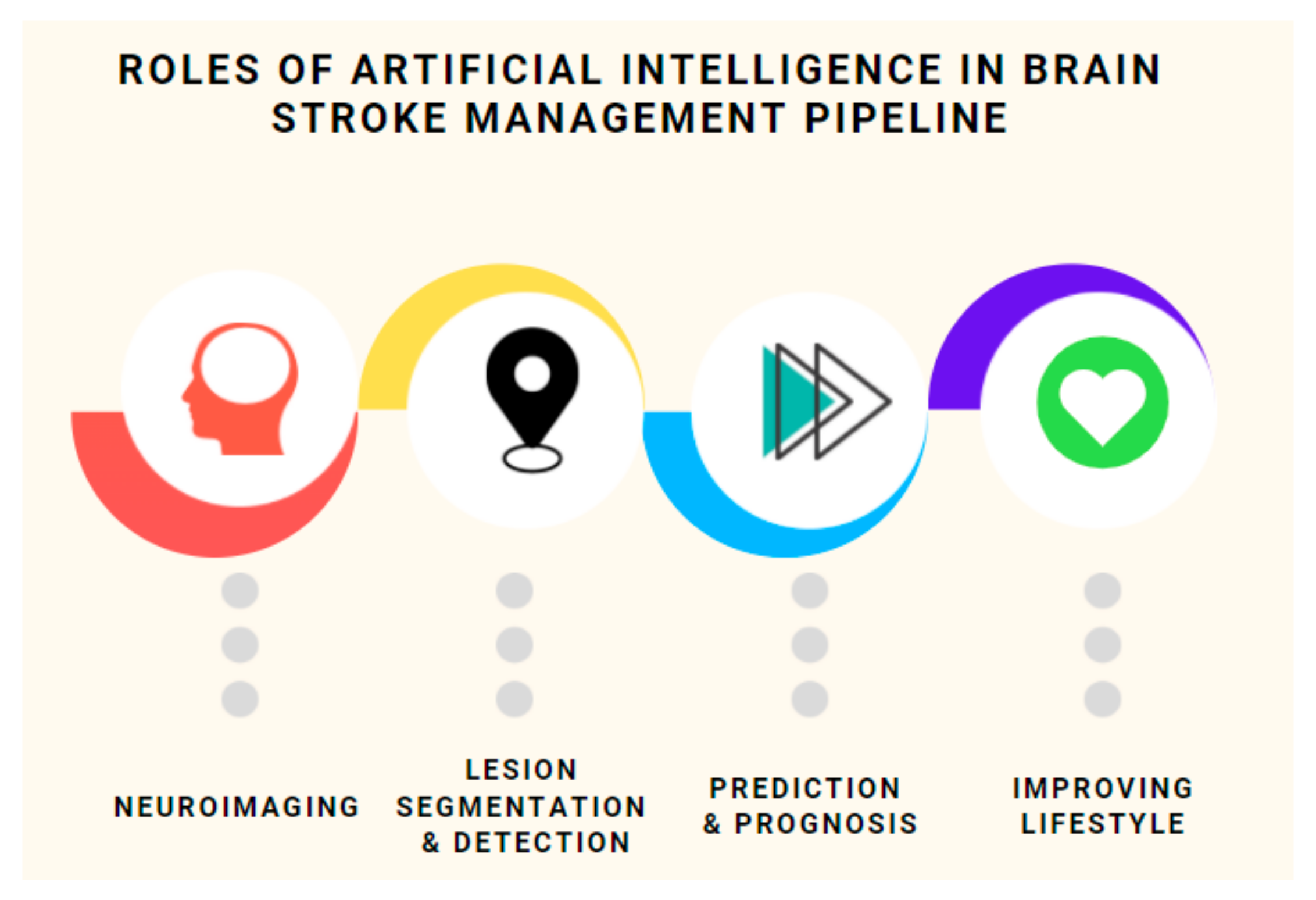
- A comprehensive overview of various modalities involved in neuroimaging, their characteristics, and requirement. We compare the most prominent ones and make remarks on their suitability, accessibility and viability. This will be useful in prioritizing future research avenues;
- An all–inclusive overview of a host of recent techniques (with special focus on prognosis) for stroke classification, detection and lesion segmentation categorized on the basis of modality used, techniques employed, datasets used (with benchmarks) (see Table 1) and the challenges faced;
- The areas of plausible future research.
1.2. Article Search
1.3. Selection of Articles
1.4. Analysis of Articles
1.5. Paper Structure
2. Brief Perspective on Brain Stroke Imaging
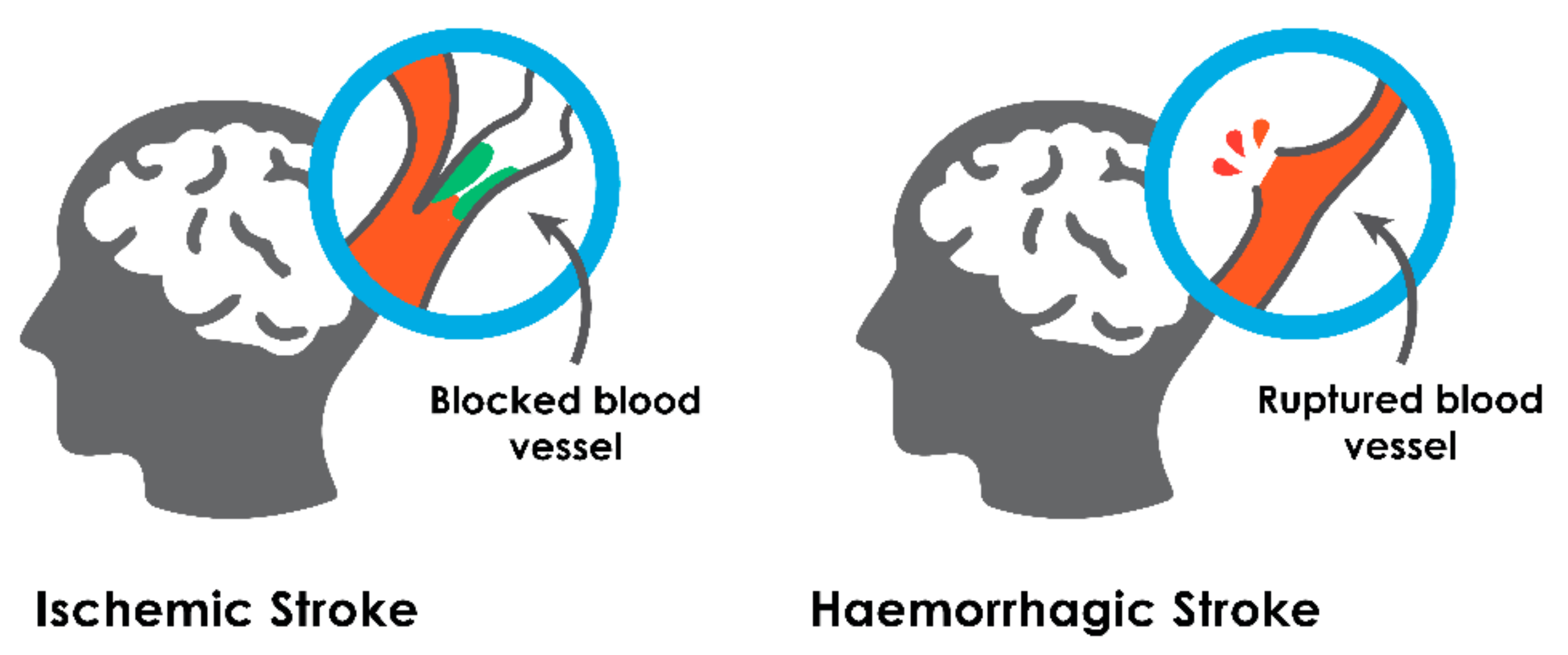
2.1. Ischemic Stroke
2.2. Hemorrhagic Stroke
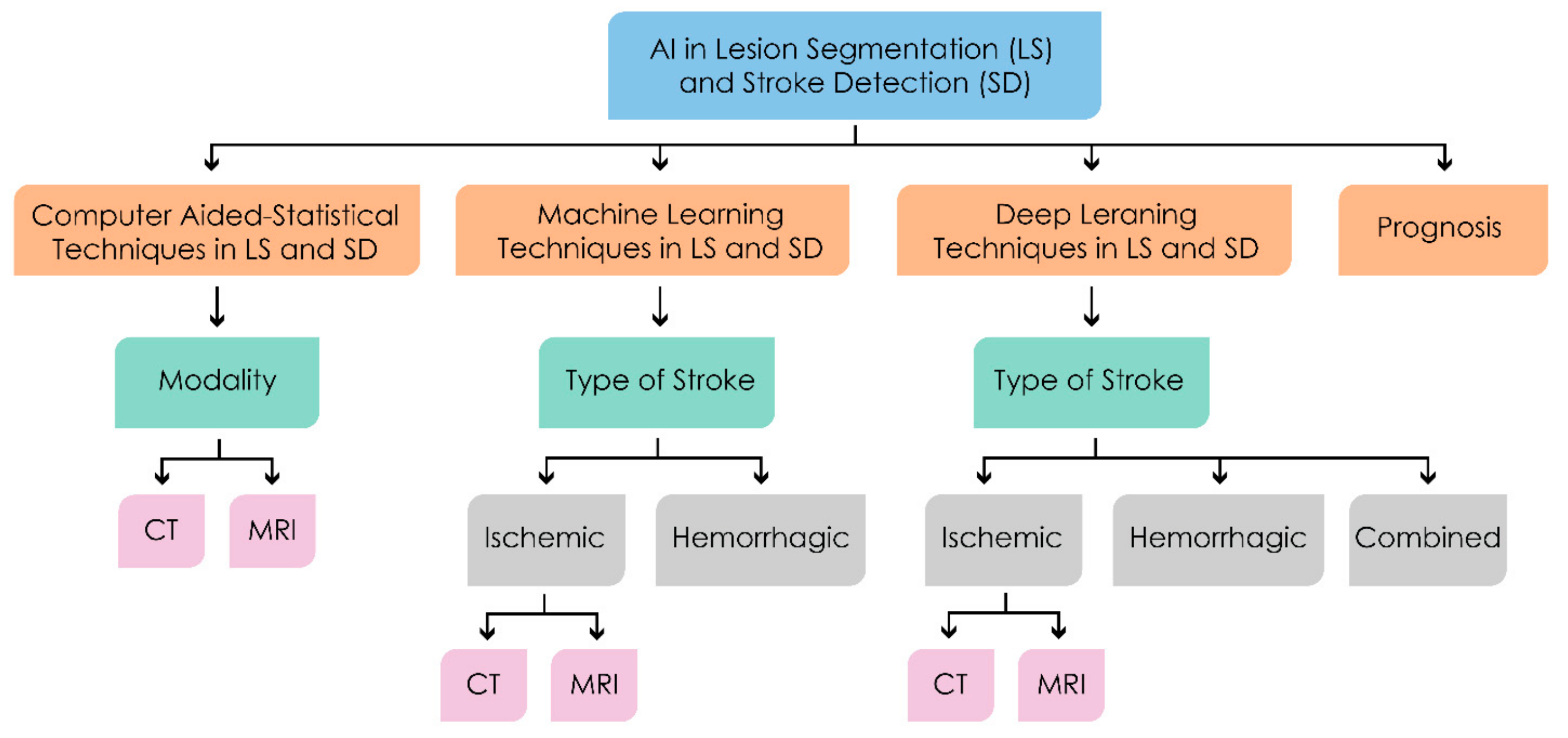
3. Machine Intelligence in Lesion Segmentation and Stroke Detection
3.1. Computer Aided—Statistical Techniques
3.1.1. CT Based Methods
3.1.2. MRI Based Methods
3.2. Machine Learning Methods
3.2.1. Ischemic Stroke
- CT based methods:
- MRI based methods:
3.2.2. Hemorrhagic Strokes
3.3. Deep Learning Methods
3.3.1. Ischemic Strokes
- CT based methods:
- MRI based methods:
3.3.2. Hemorrhagic Stroke
3.3.3. Combined Stroke
4. Discussion
4.1. Non–ML/DL Based Techniques
4.2. ML Based Techniques
4.3. DL Based Techniques
4.4. Preferred Choice of Diagnostic Imaging
4.5. Time Complexity
4.6. Prognosis
5. Challenges and Future Directions
- IoT based personalized AI: AI being the main protagonist of Industry 4.0, having far–reaching implications, especially in healthcare. Hyper personalization of healthcare could provide tailor–made diagnostics and would vastly improve early detection of disease.
- Creation of a large hetero public database: The dataset that we have addressed consists of few images for train and test, with regard to particular domain or region. A larger public dataset would assist to better cover major areas.
- Remote patient monitoring through federated learning: Figure 10 shows a prototype for remote patient monitoring with cloud–based AI Models. Wearable modes with continuous monitoring of biomarkers with easy transfer of meta–data to cloud through phones for collective learning and personalized prediction would be helpful. These could act as a digital expert to assist in patient diagnosis and prognosis.
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Virani, S.S.; Alonso, A.; Aparicio, H.J.; Benjamin, E.J.; Bittencourt, M.S.; Callaway, C.W.; Carson, A.P.; Chamberlain, A.M.; Cheng, S.; Delling, F.N.; et al. Heart Disease and Stroke Statistics—2021 Update: A Report from the American Heart Association. Available online: https://www.heart.org/-/media/PHD-Files-2/Science-News/2/2021-Heart-and-Stroke-Stat-Update/2021_Stat_Update_factsheet_Global_Burden_of_Disease.pdf (accessed on 27 January 2021). [CrossRef]
- GBD 2016 Stroke Collaborators. Global, regional, and national burden of stroke, 1990–2016: A systematic analysis for the Global Burden of Disease Study 2016. Lancet 2019, 18, 439–458. [Google Scholar] [CrossRef] [Green Version]
- Feigin, V.L.; Norrving, B.; Mensah, G.A. Global Burden of Stroke. Circ. Res. 2017, 120, 439–448. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, M.J.; Xavier, D.; Liu, L.; Zhang, H.; Chin, S.L.; Rao-Melacini, P.; Rangarajan, S.; Islam, S.; Pais, P.; McQueen, M.J.; et al. Risk factors for ischaemic and intracerebral haemorrhagic stroke in 22 countries (the INTERSTROKE study): A case-control study. Lancet 2010, 376, 112–123. [Google Scholar] [CrossRef]
- Campbell, B.C.V.; De Silva, D.A.; MacLeod, M.R.; Coutts, S.B.; Schwamm, L.H.; Davis, S.M.; Donnan, G.A. Ischaemic stroke. Nat. Rev. Dis. Primers 2019, 5, 70. [Google Scholar] [CrossRef]
- Switzer, A.J.; Hess, D.C.; Nichols, F.T.; Adams, R.J. Pathophysiology and treatment of stroke in sickle-cell disease: Present and future. Lancet Neurol. 2006, 5, 501–512. [Google Scholar] [CrossRef]
- Krishnamurthi, R.V.; Feigin, V.L.; Forouzanfar, M.H.; Mensah, A.G.; Connor, M.; Bennett, A.D.; Moran, E.A.; Sacco, R.L.; Anderson, L.M.; Truelsen, T.; et al. Global and regional burden of first–ever ischaemic and haemorrhagic stroke during 1990–2010: Findings from the Global Burden of Disease Study. Lancet Glob. Health 2013, 1, e259–e281. [Google Scholar] [CrossRef] [Green Version]
- Sagar, R.; Dandona, R.; Gururaj, G.; Dhaliwal, R.S.; Singh, A.; Ferrari, A.; Dua, T.; Ganguli, A.; Varghese, M.; Chakma, J.K.; et al. The burden of neurological disorders across the states of India: The Global Burden of Disease Study 1990–2019. Lancet Psychiatry 2020, 7, 148–161. [Google Scholar] [CrossRef] [Green Version]
- Kamalakannan, S.; Gudlavalleti, A.; Gudlavalleti, V.S.M.; Goenka, S.; Kuper, H. Incidence & prevalence of stroke in India: A systematic review. Indian J. Med. Res. 2017, 146, 175–185. [Google Scholar] [CrossRef]
- Donkor, E.S. Stroke in the21st Century: A Snapshot of the Burden, Epidemiology, and Quality of Life. Stroke Res. Treat. 2018, 2018, 3238165. [Google Scholar] [CrossRef] [Green Version]
- Saver, J.L. Penumbral salvage and thrombolysis outcome: A drop of brain, a week of life. Brain 2017, 140, 519–522. [Google Scholar] [CrossRef] [Green Version]
- Meretoja, A.; Keshtkaran, M.; Tatlisumak, T.; Donnan, G.A.; Churilov, L. Endovascular therapy for ischemic stroke: Save a minute-save a week. Neurology 2017, 88, 2123–2127. [Google Scholar] [CrossRef]
- Powers, W.J.; Rabinstein, A.A.; Ackerson, T.; Adeoye, O.M.; Bambakidis, N.C.; Becker, K.; Biller, J.; Brown, M.; Demaerschalk, B.M.; Hoh, B.; et al. 2018 Guidelines for the Early Management of Patients with Acute Ischemic Stroke: A Guideline for Healthcare Professionals from the American Heart Association/American Stroke Association. Stroke 2018, 49, e46–e99. [Google Scholar] [CrossRef]
- Lindsay, P.; Furie, K.L.; Davis, S.M.; Donnan, G.; Norrving, B. World Stroke Organization Global Stroke Services Guidelines and Action Plan. Int. J. Stroke 2014, 9, 4–13. [Google Scholar] [CrossRef] [Green Version]
- Brazzelli, M.; Sandercock, P.A.; Chappell, F.M.; Celani, M.G.; Righetti, E.; Arestis, N.; Wardlaw, J.M.; Deeks, J.J. Magnetic resonance imaging versus computed tomography for detection of acute vascular lesions in patients presenting with stroke symptoms. Cochrane Database Syst. Rev. 2009, 4, CD007424. [Google Scholar] [CrossRef]
- Clinical Guidelines for Stroke Management. Available online: https://informme.org.au/en/Guidelines/Clinical-Guidelines-for-Stroke-Management (accessed on 3 October 2021).
- Bivard, A.; Churilov, L.; Parsons, M. Artificial intelligence for decision support in acute stroke—Current roles and potential. Nat. Rev. Neurol. 2020, 16, 575–585. [Google Scholar] [CrossRef]
- Rowley, H.; Vagal, A. Stroke and Stroke Mimics: Diagnosis and Treatment. In Diseases of the Brain, Head and Neck, Spine 2020–2023: Diagnostic Imaging; IDKD Springer Series; Springer: Berlin/Heidelberg, Germany, 2020; pp. 25–36. [Google Scholar]
- Allen, M.L.; Anton, N.; Hasso, J.; Handwerker, J.; Hamed, F. Sequence-specific MR Imaging Findings that Are Useful in Dating Ischemic Stroke. RadioGraphics 2012, 32, 1285–1297. [Google Scholar] [CrossRef] [Green Version]
- Moreau, F.; Asdaghi, N.; Modi, J.; Goyal, M.; Coutts, S.B. Magnetic Resonance Imaging versus Computed Tomography in Transient Ischemic Attack and Minor Stroke: The More You See the More You Know. Cerebrovasc. Dis. Extra 2013, 3, 130–136. [Google Scholar] [CrossRef]
- National Institutes of Health. MRI More Sensitive Than CT in Diagnosing Most Common Form of Acute Stroke, Finds NIH Study. Available online: https://www.nih.gov/news-events/news-releases/mri-more-sensitive-ct-diagnosing-most-common-form-acute-stroke-finds-nih-study (accessed on 4 October 2021).
- Hopyan, J.; Ciarallo, A.; Dowlatshahi, D.; Howard, P.; John, V.; Yeung, R.; Zhang, L.; Kim, J.; Macfarlane, G.; Lee, T.-Y.; et al. Certainty of Stroke Diagnosis: Incremental Benefit with CT Perfusion over Noncontrast CT and CT Angiography. Radiology 2010, 255, 142–153. [Google Scholar] [CrossRef]
- Karthik, R.; Menaka, R.; Johnson, A.; Anand, S. Neuroimaging and deep learning for brain stroke detection–A review of recent advancements and future prospects. Comput. Methods Programs Biomed. 2020, 197, 105728. [Google Scholar] [CrossRef]
- Karthik, R.; Menaka, R. Computer-aided detection and characterization of stroke lesion—A short review on the current state-of-the art methods. Imaging Sci. J. 2017, 66, 836–843. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, S.; Li, C.; Wang, J. Review: Application of Deep Learning Method on Ischemic Stroke Lesion Segmentation. J. Shanghai Jiaotong Univ. 2021, 1–13. [Google Scholar] [CrossRef]
- Sirsat, M.S.; Fermé, E.; Câmara, J. Machine Learning for Brain Stroke: A Review. J. Stroke Cerebrovasc. Dis. 2020, 29, 105162. [Google Scholar] [CrossRef]
- Wadhwa, A.; Bhardwaj, A.; Verma, V.S. A review on brain tumor segmentation of MRI images. Magn. Reson. Imaging 2019, 61, 247–259. [Google Scholar] [CrossRef]
- Magnetic Resonance Imaging (MRI) of the Brain and Spine: Basics. Available online: https://case.edu/med/neurology/NR/MRI%20Basics.htm (accessed on 4 October 2021).
- Niknejad, M.; Bell, D. Apparent Diffusion Coefficient. Available online: https://radiopaedia.org/articles/apparent-diffusion-coefficient-1 (accessed on 4 October 2021).
- Chalela, J.A.; Kidwell, C.S.; Nentwich, L.M.; Luby, M.; Butman, J.A.; Demchuk, A.M.; Hill, M.D.; Patronas, N.; Latour, L.; Warach, S. Magnetic resonance imaging and computer tomography in emergency assessment of patients with suspected acute stroke: A prospective comparison. Lancet 2007, 369, 293–298. [Google Scholar] [CrossRef] [Green Version]
- Mainali, S.; Wahba, M.; Elijovich, L. Detection of Early Ischemic Changes in Noncontrast CT Head Improved with “Stroke Windows”. ISRN Neurosci. 2014, 2014, 654980. [Google Scholar] [CrossRef]
- Tee, Y.J.; Gaillard, F. Ischemic Stroke. Available online: https://radiopaedia.org/articles/ischaemic-stroke (accessed on 4 October 2021).
- Macellari, F.; Paciaroni, M.; Agnelli, G.; Caso, V. Neuroimaging in Intracerebral Hemorrhage. Stroke 2014, 45, 903–908. [Google Scholar] [CrossRef] [Green Version]
- Smith, E.E.; Rosand, J.; Greenberg, S.M. Imaging of Hemorrhagic Stroke. Magn. Reson. Imaging Clin. N. Am. 2006, 14, 127–140. [Google Scholar] [CrossRef] [PubMed]
- Kang, B.K.; Na, D.G.; Ryoo, J.W.; Byun, H.S.; Roh, H.G.; Pyeun, Y.S. Diffusion-Weighted MR Imaging of Intracerebral Hemorrhage. Korean J. Radiol. 2001, 2, 183–191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- French, B.R.; Boddepalli, R.S.; Govindarajan, R. Acute Ischemic Stroke: Current Status and Future Directions. Mo. Med. 2016, 113, 480–486. [Google Scholar] [PubMed]
- Lebedev, G.; Meshcheryakova, A.; Kurenkov, V.; Kluganov, V.; Sologubov, A.; Logacheva, N.; Radzievskiy, G.; Klimenko, H. Application of artificial intelligence methods to recognize pathologies on photographs of fundus. Procedia Comput. Sci. 2020, 176, 1823–1828. [Google Scholar] [CrossRef]
- Tang, F.-H.; Ng, K.S.; Chow, H.K.D. An image feature approach for computer-aided detection of ischemic stroke. Comput. Biol. Med. 2011, 41, 529–536. [Google Scholar] [CrossRef]
- Sajjadi, M.; Amirfattahi, R.; Ahmadzadeh, M.R.; Saghafi, M.A. A new filter bank algorithm for enhancement of early signs of ischemic stroke in brain CT images. In Proceedings of the 2011 IEEE International Conference on Signal and Image Processing Applications (ICSIPA), Kuala Lumpur, Malaysia, 16–18 November 2011; pp. 384–389. [Google Scholar] [CrossRef]
- Nowinski, W.L.; Gupta, V.; Qian, G.; He, J.; Poh, L.E.; Ambrosius, W.; Chrzan, R.M.; Polonara, G.; Mazzoni, C.; Mol, M.; et al. Automatic Detection, Localization, and Volume Estimation of Ischemic Infarcts in Noncontrast Computed Tomographic Scans. Investig. Radiol. 2013, 48, 661–670. [Google Scholar] [CrossRef]
- Filho, P.P.R.; Sarmento, R.M.; Holanda, G.B.; Lima, D.D.A. New approach to detect and classify stroke in skull CT images via analysis of brain tissue densities. Comput. Methods Programs Biomed. 2017, 148, 27–43. [Google Scholar] [CrossRef]
- Flottmann, F.; Broocks, G.; Faizy, T.D.; Ernst, M.; Forkert, N.D.; Grosser, M.; Thomalla, G.; Siemonsen, S.; Fiehler, J.; Kemmling, A. CT-perfusion stroke imaging: A threshold free probabilistic approach to predict infarct volume compared to traditional ischemic thresholds. Sci. Rep. 2017, 7, 6679. [Google Scholar] [CrossRef]
- Sakai, Y.; Delman, B.; Fifi, J.T.; Tuhrim, S.; Wheelwright, D.; Doshi, A.H.; Mocco, J.; Nael, K. Estimation of Ischemic Core Volume Using Computed Tomographic Perfusion. Stroke 2018, 49, 2345–2352. [Google Scholar] [CrossRef]
- Lo, C.-M.; Hung, P.-H.; Hsieh, K.L.-C. Computer-Aided Detection of Hyperacute Stroke Based on Relative Radiomic Patterns in Computed Tomography. Appl. Sci. 2019, 9, 1668. [Google Scholar] [CrossRef] [Green Version]
- Kamalian, S.; Kamalian, S.; Konstas, A.A.; Maas, M.B.; Payabvash, S.; Pomerantz, S.R.; Schaefer, P.W.; Furie, K.L.; González, R.G.; Lev, M.H. CT Perfusion Mean Transit Time Maps Optimally Distinguish Benign Oligemia from True “At-Risk” Ischemic Penumbra, but Thresholds Vary by Postprocessing Technique. Am. J. Neuroradiol. 2011, 33, 545–549. [Google Scholar] [CrossRef]
- Kheradmand, A.; Fisher, M.; Paydarfar, D. Ischemic Stroke in Evolution: Predictive Value of Perfusion Computed Tomography. J. Stroke Cerebrovasc. Dis. 2014, 23, 836–843. [Google Scholar] [CrossRef]
- Kawiorski, M.M.; Vicente, A.; Lourido, D.; Muriel, A.; Fandiño, E.; Méndez, J.C.; Sánchez-González, V.; Aguado, A.; Álvarez-Velasco, R.; De Leciñana, M.A. Good Clinical and Radiological Correlation from Standard Perfusion Computed Tomography Accurately Identifies Salvageable Tissue in Ischemic Stroke. J. Stroke Cerebrovasc. Dis. 2016, 25, 1062–1069. [Google Scholar] [CrossRef]
- Bhadauria, H.S.; Dewal, M.L. Intracranial hemorrhage detection using spatial fuzzy c-mean and region-based active contour on brain CT imaging. Signal Image Video Process. 2012, 8, 357–364. [Google Scholar] [CrossRef]
- De Haan, B.; Clas, P.; Juenger, H.; Wilke, M.; Karnath, H.-O. Fast semi-automated lesion demarcation in stroke. NeuroImage Clin. 2015, 9, 69–74. [Google Scholar] [CrossRef]
- Yahiaoui, A.F.Z.; Bessaid, A. Segmentation of ischemic stroke area from CT brain images. In Proceedings of the 2016 International Symposium on Signal, Image, Video and Communications (ISIVC), Tunis, Tunisia, 21–23 November 2016; pp. 13–17. [Google Scholar]
- Reboucas, E.D.S.; Braga, A.M.; Sarmento, R.M.; Marques, R.; Filho, P.P.R. Level Set Based on Brain Radiological Densities for Stroke Segmentation in CT Images. In Proceedings of the 2017 IEEE 30th International Symposium on Computer-Based Medical Systems (CBMS), Thessaloniki, Greece, 22–24 June 2017; pp. 391–396. [Google Scholar]
- Kumar, I.; Bhatt, C.; Singh, K.U. Entropy based automatic unsupervised brain intracranial hemorrhage segmentation using CT images. J. King Saud Univ. Comput. Inf. Sci. 2020, in press. [Google Scholar] [CrossRef]
- Vasconcelos, F.F.; Sarmento, R.M.; Filho, P.P.R.; de Albuquerque, V.H.C. Artificial intelligence techniques empowered edge-cloud architecture for brain CT image analysis. Eng. Appl. Artif. Intell. 2020, 91, 103585. [Google Scholar] [CrossRef]
- Nabizadeh, N.; John, N.; Wright, C. Histogram-based gravitational optimization algorithm on single MR modality for automatic brain lesion detection and segmentation. Expert Syst. Appl. 2014, 41, 7820–7836. [Google Scholar] [CrossRef]
- Ghosh, N.; Sun, Y.; Bhanu, B.; Ashwal, S.; Obenaus, A. Automated detection of brain abnormalities in neonatal hypoxia ischemic injury from MR images. Med. Image Anal. 2014, 18, 1059–1069. [Google Scholar] [CrossRef] [Green Version]
- Cauley, K.A.; Mongelluzzo, G.J.; Fielden, S.W. Automated Estimation of Acute Infarct Volume from Noncontrast Head CT Using Image Intensity Inhomogeneity Correction. Int. J. Biomed. Imaging 2019, 2019, 1720270. [Google Scholar] [CrossRef] [Green Version]
- Ledig, C.; Heckemann, R.; Hammers, A.; Lopez, J.C.; Newcombe, V.; Makropoulos, A.; Lötjönen, J.; Menon, D.K.; Rueckert, D. Robust whole-brain segmentation: Application to traumatic brain injury. Med. Image Anal. 2015, 21, 40–58. [Google Scholar] [CrossRef] [Green Version]
- Nazari-Farsani, S.; Nyman, M.; Karjalainen, T.; Bucci, M.; Isojärvi, J.; Nummenmaa, L. Automated segmentation of acute stroke lesions using a data-driven anomaly detection on diffusion weighted MRI. J. Neurosci. Methods 2020, 333, 108575. [Google Scholar] [CrossRef]
- Moeskops, P.; Viergever, M.A.; Mendrik, A.M.; De Vries, L.S.; Benders, M.J.N.L.; Isgum, I. Automatic Segmentation of MR Brain Images with a Convolutional Neural Network. IEEE Trans. Med. Imaging 2016, 35, 1252–1261. [Google Scholar] [CrossRef] [Green Version]
- Puonti, O.; van Leemput, K. Simultaneous whole-brain segmentation and white matter lesion detection using contrast-adaptive probabilistic models. In Proceedings of the Brainles 2015 MICCAI Workshop, Munich, Germany, 5 October 2015; pp. 12–19. [Google Scholar]
- Si, T.; De, A.; Bhattacharjee, A.K. Segmentation of Brain MRI Using Wavelet Transform and Grammatical Bee Colony. J. Circuits Syst. Comput. 2018, 27, 1850108. [Google Scholar] [CrossRef]
- Haeck, T.; Maes, F.; Suetens, P. ISLES Challenge 2015: Automated Model-Based Segmentation of Ischemic Stroke in MR Images. In BrainLes 2015, Proceedings of the First International Workshop, Held in Conjunction with MICCAI 2015, Munich, Germany, 5 October 2015; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2016; pp. 246–253. [Google Scholar]
- Ji, Z.; Xia, Y.; Zheng, Y. Robust generative asymmetric GMM for brain MR image segmentation. Comput. Methods Programs Biomed. 2017, 151, 123–138. [Google Scholar] [CrossRef] [PubMed]
- Kamnitsas, K.; Ledig, C.; Newcombe, V.; Simpson, J.P.; Kane, A.D.; Menon, D.K.; Rueckert, D.; Glocker, B. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 2017, 36, 61–78. [Google Scholar] [CrossRef] [PubMed]
- Rajini, N.H.; Bhavani, R. Computer aided detection of ischemic stroke using segmentation and texture features. Measurement 2013, 46, 1865–1874. [Google Scholar] [CrossRef]
- Chen, Y.; Dhar, R.; Heitsch, L.; Ford, A.; Fernandez–Cade, I.; Carrera, C.; Montaner, J.; Lin, W.; Shen, D.; An, H.; et al. Automated quantification of cerebral edema following hemispheric infarction: Application of a machine-learning algorithm to evaluate CSF shifts on serial head CTs. NeuroImage Clin. 2016, 12, 673–680. [Google Scholar] [CrossRef] [Green Version]
- Dhar, R.; Chen, Y.; An, H.; Lee, J.-M. Application of Machine Learning to Automated Analysis of Cerebral Edema in Large Cohorts of Ischemic Stroke Patients. Front. Neurol. 2018, 9, 687. [Google Scholar] [CrossRef]
- Guberina, N.; Dietrich, U.; Radbruch, A.; Goebel, J.; Deuschl, C.; Ringelstein, A.; Köhrmann, M.; Kleinschnitz, C.; Forsting, M.; Mönninghoff, C. Detection of early infarction signs with machine learning-based diagnosis by means of the Alberta Stroke Program Early CT score (ASPECTS) in the clinical routine. Neuroradiology 2018, 60, 889–901. [Google Scholar] [CrossRef]
- Fuchigami, T.; Akahori, S.; Okatani, T.; Li, Y. A hyperacute stroke segmentation method using 3D UNet integrated with physicians’ knowledge for NCCT. Proc. SPIE 2020, 11314, 113140G. [Google Scholar]
- Carey, L.M.; Seitz, R.J.; Parsons, M.; Levi, C.; Farquharson, S.; Tournier, J.-D.; Palmer, S.; Connelly, A. Beyond the lesion: Neuroimaging foundations for post-stroke recovery. Future Neurol. 2013, 8, 507–527. [Google Scholar] [CrossRef] [Green Version]
- Hirano, T. Searching for Salvageable Brain: The Detection of Ischemic Penumbra Using Various Imaging Modalities? J. Stroke Cerebrovasc. Dis. 2014, 23, 795–798. [Google Scholar] [CrossRef]
- Maier, O.; Schröder, C.; Forkert, N.D.; Martinetz, T.; Handels, H. Correction: Classifiers for Ischemic Stroke Lesion Segmentation: A Comparison Study. PLoS ONE 2016, 11, e0149828. [Google Scholar] [CrossRef]
- Mitra, J.; Bourgeat, P.; Fripp, J.; Ghose, S.; Rose, S.; Salvado, O.; Connelly, A.; Campbell, B.; Palmer, S.; Sharma, G.; et al. Lesion segmentation from multimodal MRI using random forest following ischemic stroke. NeuroImage 2014, 98, 324–335. [Google Scholar] [CrossRef]
- Bharathi, P.G.; Agrawal, A.; Sundaram, P.; Sardesai, S.; Praveen, G. Combination of hand-crafted and unsupervised learned features for ischemic stroke lesion detection from Magnetic Resonance Images. Biocybern. Biomed. Eng. 2019, 39, 410–425. [Google Scholar] [CrossRef]
- Yoo, A.J.; Barak, E.R.; Copen, W.A.; Kamalian, S.; Gharai, L.R.; Pervez, M.A.; Schwamm, L.H.; González, R.G.; Schaefer, P.W. Combining Acute Diffusion–Weighted Imaging and Mean Transmit Time Lesion Volumes with National Institutes of Health Stroke Scale Score Improves the Prediction of Acute Stroke. Stroke 2010, 41, 1728–1735. [Google Scholar] [CrossRef] [Green Version]
- Maier, O.; Wilms, M.; von der Gablentz, J.; Krämer, U.M.; Münte, T.F.; Handels, H. Extra Tree forests for sub–acute ischemic stroke lesion segmentation in MR sequences. J. Neurosci. Methods 2015, 240, 89–100. [Google Scholar] [CrossRef]
- Bouts, M.; Tiebosch, I.A.C.W.; Van Der Toorn, A.; Viergever, A.M.; Wu, O.; Dijkhuizen, R.M. Early Identification of Potentially Salvageable Tissue with MRI–Based Predictive Algorithms after Experimental Ischemic Stroke. Br. J. Pharmacol. 2013, 33, 1075–1082. [Google Scholar] [CrossRef]
- Muschelli, J. Prediction of ischemic lesions using local image properties and random forests. In Proceedings of the Ischemic Stroke Lesion Segmentation (ISLES) challenge–MICCAI, Munich, Germany, 5 October 2015; pp. 63–68. [Google Scholar]
- Mahmood, Q.; Basit, A. Automatic Ischemic Stroke Lesion Segmentation in Multi-spectral MRI Images Using Random Forests Classifier. In BrainLes 2015, Proceedings of the First International Workshop, Held in Conjunction with MICCAI 2015, Munich, Germany, 5 October 2015; Lecture Notes in Computer Science; Springer: Cham, Switzerland, 2015; pp. 266–274. [Google Scholar]
- Jesson, A.; Arbel, T. Hierarchical segmentation of normal and lesional structures combining an ensemble of probabilistic local classifiers and regional random forest classification. In Proceedings of the Ischemic Stroke Lesion Segmentation (ISLES) challenge–MICCAI, Munich, Germany, 5 October 2015; pp. 57–62. [Google Scholar]
- Halme, H.-L.; Korvenoja, A.; Salli, E. ISLES (SISS) Challenge 2015: Segmentation of Stroke Lesions Using Spatial Normalization, Random Forest Classification and Contextual Clustering. In BrainLes 2015, Proceedings of the First International Workshop, Held in Conjunction with MICCAI 2015, Munich, Germany, 5 October 2015; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2016; pp. 211–221. [Google Scholar]
- Jerman, T.; Galimzianova, A.; Pernuš, F.; Likar, B.; Špiclin, Ž. Combining Unsupervised and Supervised Methods for Lesion Segmentation. In BrainLes 2015, Proceedings of the First International Workshop, Held in Conjunction with MICCAI 2015, Munich, Germany, 5 October 2015; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2016; pp. 45–56. [Google Scholar]
- McKinley, R.; Häni, L.; Wiest, R.; Reyes, M. Segmenting the Ischemic Penumbra: A Decision Forest Approach with Automatic Threshold Finding. In BrainLes 2015, Proceedings of the First International Workshop, Held in Conjunction with MICCAI 2015, Munich, Germany, 5 October 2015; Springer: Cham, Switzerland, 2015; pp. 275–283. [Google Scholar] [CrossRef]
- Robben, D.; Christiaens, D.; Rangarajan, J.R.; Gelderblom, J.; Joris, P.; Maes, F.; Suetens, P. A Voxel–Wise, Cascaded Classification Approach to Ischemic Stroke Lesion Segmentation. In BrainLes 2015, Proceedings of the First International Workshop, Held in Conjunction with MICCAI 2015, Munich, Germany, 5 October 2015; Lecture Notes in Computer Science; Springer: Cham, Switzerland, 2015; Volume 9556, pp. 254–265. [Google Scholar]
- Chen, G.; Li, Q.; Shi, F.; Rekik, I.; Pan, Z. RFDCR: Automated brain lesion segmentation using cascaded random forests with dense conditional random fields. NeuroImage 2020, 211, 116620. [Google Scholar] [CrossRef]
- Goetz, M.; Weber, C.; Kolb, C.; Maier-Hein, K. Input Data Adaptive Learning (IDAL) for Sub-acute Ischemic Stroke Lesion Segmentation. In BrainLes 2015, Proceedings of the First International Workshop, Held in Conjunction with MICCAI 2015, Munich, Germany, 5 October 2015; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2016; Volume 9556, pp. 284–295. [Google Scholar]
- Griffanti, L.; Zamboni, G.; Khan, A.; Li, L.; Bonifacio, G.; Sundaresan, V.; Schulz, U.G.; Kuker, W.; Battaglini, M.; Rothwell, P.M.; et al. BIANCA (Brain Intensity AbNormality Classification Algorithm): A new tool for automated segmentation of white matter hyperintensities. NeuroImage 2016, 141, 191–205. [Google Scholar] [CrossRef] [Green Version]
- Griffis, J.C.; Allendorfer, J.B.; Szaflarski, J.P. Voxel-based Gaussian naïve Bayes classification of ischemic stroke lesions in individual T1-weighted MRI scans. J. Neurosci. Methods 2016, 257, 97–108. [Google Scholar] [CrossRef] [Green Version]
- Karthik, R.; Menaka, R. A multi-scale approach for detection of ischemic stroke from brain MR images using discrete curvelet transformation. Measurement 2017, 100, 223–232. [Google Scholar] [CrossRef]
- Pereira, S.; Meier, R.; McKinley, R.; Wiest, R.; Alves, V.; Silva, C.A.; Reyes, M. Enhancing interpretability of automatically extracted machine learning features: Application to a RBM-Random Forest system on brain lesion segmentation. Med. Image Anal. 2018, 44, 228–244. [Google Scholar] [CrossRef]
- Lin, C.-H.; Hsu, K.-C.; Johnson, K.R.; Luby, M.; Fann, Y.C. Applying density-based outlier identifications using multiple datasets for validation of stroke clinical outcomes. Int. J. Med. Inform. 2019, 132, 103988. [Google Scholar] [CrossRef]
- Subudhi, A.K.; Acharya, U.R.; Dash, M.; Jena, S.; Sabut, S. Automated approach for detection of ischemic stroke using Delaunay Triangulation in brain MRI images. Comput. Biol. Med. 2018, 103, 116–129. [Google Scholar] [CrossRef]
- Peixoto, S.A.; Filho, P.P.R. Neurologist-level classification of stroke using a Structural Co-Occurrence Matrix based on the frequency domain. Comput. Electr. Eng. 2018, 71, 398–407. [Google Scholar] [CrossRef]
- Garg, R.; Oh, E.; Naidech, A.; Kording, K.; Prabhakaran, S. Automating Ischemic Stroke Subtype Classification Using Machine Learning and Natural Language Processing. J. Stroke Cerebrovasc. Dis. 2019, 28, 2045–2051. [Google Scholar] [CrossRef]
- Chen, H.; Khan, S.; Kou, B.; Nazir, S.; Liu, W.; Hussain, A. A Smart Machine Learning Model for the Detection of Brain Hemorrhage Diagnosis Based Internet of Things in Smart Cities. Complexity 2020, 2020, 3047869. [Google Scholar] [CrossRef]
- Gillebert, C.; Humphreys, G.W.; Mantini, D. Automated delineation of stroke lesions using brain CT images. NeuroImage Clin. 2014, 4, 540–548. [Google Scholar] [CrossRef] [Green Version]
- Bentley, P.; Ganesalingam, J.; Jones, A.L.C.; Mahady, K.; Epton, S.; Rinne, P.; Sharma, P.; Halse, O.; Mehta, A.; Rueckert, D. Prediction of stroke thrombolysis outcome using CT brain machine learning. NeuroImage Clin. 2014, 4, 635–640. [Google Scholar] [CrossRef] [Green Version]
- Chin, C.-L.; Lin, B.-J.; Wu, G.-R.; Weng, T.-C.; Yang, C.-S.; Su, R.-C.; Pan, Y.-J. An automated early ischemic stroke detection system using CNN deep learning algorithm. In Proceedings of the 2017 IEEE 8th International Conference on Awareness Science and Technology (iCAST), Taichung, Taiwan, 8 November 2017; pp. 368–372. [Google Scholar]
- Lisowska, A.; O’Neil, A.; Dilys, V.; Daykin, M.; Beveridge, E.; Muir, K.; Mclaughlin, S.; Poole, I. Context-Aware Convolutional Neural Networks for Stroke Sign Detection in Non-contrast CT Scans. In Proceedings of the Annual Conference on Medical Image Understanding and Analysis, 21st Annual Conference, MIUA 2017, Communications in Computer and Information Science. Edinburgh, UK, 11–13 July 2017; Springer: Berlin/Heidelberg, Germany, 2017; Volume 723, pp. 494–505. [Google Scholar]
- Abulnaga, S.M.; Rubin, J. Ischemic Stroke Lesion Segmentation in CT Perfusion Scans using Pyramid Pooling and Focal Loss. arXiv 2018, arXiv:1811.01085. [Google Scholar]
- Lucas, C.; Kemmling, A.; Bouteldja, N.; Aulmann, L.F.; Mamlouk, A.M.; Heinrich, M.P. Learning to Predict Ischemic Stroke Growth on Acute CT Perfusion Data by Interpolating Low-Dimensional Shape Representations. Front. Neurol. 2018, 9, 989. [Google Scholar] [CrossRef] [Green Version]
- Vargas, J.; Spiotta, A.; Chatterjee, A.R. Initial Experiences with Artificial Neural Networks in the Detection of Computed Tomography Perfusion Deficits. World Neurosurg. 2018, 124, e10–e16. [Google Scholar] [CrossRef]
- Barman, A.; Inam, M.E.; Lee, S.; Savitz, S.; Sheth, S.; Giancardo, L. Determining Ischemic Stroke from CT-Angiography Imaging Using Symmetry-Sensitive Convolutional Networks. In Proceedings of the 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), Taichung, Taiwan, 8–10 November 2019; pp. 1873–1877. [Google Scholar]
- Clèrigues, A.; Valverde, S.; Bernal, J.; Freixenet, J.; Oliver, A.; Llado, X. Acute ischemic stroke lesion core segmentation in CT perfusion images using fully convolutional neural networks. Comput. Biol. Med. 2019, 115, 103487. [Google Scholar] [CrossRef]
- Shinohara, Y.; Takahashi, N.; Lee, Y.; Ohmura, T.; Kinoshita, T. Development of a deep learning model to identify hyperdense MCA sign in patients with acute ischemic stroke. Jpn. J. Radiol. 2020, 38, 112–117. [Google Scholar] [CrossRef] [PubMed]
- Barros, S.R.; Tolhuisen, M.L.; Boers, A.M.; Jansen, I.; Ponomareva, E.; Dippel, D.W.J.; Lught, A.V.D.; Oostenbrugge, R.J.V.; Zwam, W.H.V.; Berkhemer, O.A.; et al. Automatic segmentation of cerebral infarcts in follow-up computed tomography images with convolutional neural networks. J. NeuroInterv. Surg. 2020, 12, 848–852. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Öman, O.; Mäkelä, T.; Salli, E.; Savolainen, S.; Kangasniemi, M. 3D convolutional neural networks applied to CT angiography in the detection of acute ischemic stroke. Eur. Radiol. Exp. 2019, 3, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, X.; Luo, W.; Hu, J.; Guo, S.; Huang, W.; Scott, M.R.; Wiest, R.; Dahlweid, M.; Reyes, M. Brain SegNet: 3D local refinement network for brain lesion segmentation. BMC Med. Imaging 2020, 20, 17. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.; Vaidyanathan, N.R.; Jose, V.J.M.; Ren, H. Ischemic Stroke Lesion Segmentation Using Adversarial Learning. Fundam. Softw. Eng. 2019, 11383, 292–300. [Google Scholar] [CrossRef]
- Bertels, J.; Robben, D.; Vandermeulen, D.; Suetens, P. Contra-Lateral Information CNN for Core Lesion Segmentation Based on Native CTP in Acute Stroke. In Proceedings of the International MICCAI Brainlesion Workshop, 4th International Workshop, BrainLes 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, 16 September 2018; Lecture Notes in Computer Science. Springer: Cham, Switzerland, 2019; pp. 263–270. [Google Scholar] [CrossRef]
- Kuang, H.; Menon, B.K.; Sohn, S.I.; Qiu, W. EIS-Net: Segmenting early infarct and scoring ASPECTS simultaneously on non-contrast CT of patients with acute ischemic stroke. Med. Image Anal. 2021, 70, 101984. [Google Scholar] [CrossRef]
- Avetisian, M.; Kokh, V.; Tuzhilin, A.; Umerenkov, D. Radiologist-level stroke classification on non-contrast CT scans with Deep U-Net. arXiv 2020, arXiv:2003.14287. [Google Scholar] [CrossRef] [Green Version]
- Robben, D.; Boers, A.M.; Marquering, H.; Langezaal, L.L.; Roos, Y.B.; van Oostenbrugge, R.J.; van Zwam, W.H.; Dippel, D.W.; Majoie, C.B.; van der Lugt, A.; et al. Prediction of final infarct volume from native CT perfusion and treatment parameters using deep learning. Med. Image Anal. 2020, 59, 101589. [Google Scholar] [CrossRef]
- Wang, C.-W.; Lee, J.-H. Stroke Lesion Segmentation of 3D Brain MRI Using Multiple Random Forests and 3D Registration. Adv. Data Min. Appl. 2016, 9556, 222–232. [Google Scholar]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar] [CrossRef] [Green Version]
- Havaei, M.; Dutil, F.; Pal, C.; Larochelle, H.; Jodoin, P.-M. A Convolutional Neural Network Approach to Brain Tumor Segmentation. In Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries; Springer: Berlin/Heidelberg, Germany, 2016; Volume 9556, pp. 195–208. [Google Scholar]
- Stier, N.; Vincent, N.; Liebeskind, D.; Scalzo, F. Deep learning of tissue fate features in acute ischemic stroke. In Proceedings of the 2015 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Washington, DC, USA, 9–12 November 2015; Volume 2015, pp. 1316–1321. [Google Scholar]
- Dou, Q.; Chen, H.; Yu, L.; Zhao, L.; Qin, J.; Wang, D.; Mok, V.; Shi, L.; Heng, P.A. Automatic Detection of Cerebral Microbleeds From MR Images via 3D Convolutional Neural Networks. IEEE Trans. Med. Imaging 2016, 35, 1182–1195. [Google Scholar] [CrossRef]
- Choi, Y.; Kwon, Y.; Lee, H.; Kim, B.J.; Paik, M.C.; Won, J.-H. Ensemble of Deep Convolutional Neural Networks for Prognosis of Ischemic Stroke. In Computer Vision; Springer: Berlin/Heidelberg, Germany, 2016; Volume 10154, pp. 231–243. [Google Scholar]
- Wang, G.; Song, T.; Dong, Q.; Cui, M.; Huang, N.; Zhang, S. Automatic ischemic stroke lesion segmentation from computed tomography perfusion images by image synthesis and attention–based deep neural networks. Med. Image Anal. 2020, 65, 101787. [Google Scholar] [CrossRef]
- Chen, L.; Bentley, P.; Rueckert, D. Fully automatic acute ischemic lesion segmentation in DWI using convolutional neural networks. NeuroImage Clin. 2017, 15, 633–643. [Google Scholar] [CrossRef]
- Lucas, C.; Maier, O.; Heinrich, M.P. Shallow Fully-Connected Neural Networks for Ischemic Stroke-Lesion Segmentation in MRI. In Informatik Aktuell; Springer: Berlin/Heidelberg, Germany, 2017; pp. 261–266. [Google Scholar]
- Alex, V.; Vaidhya, K.; Thirunavukkarasu, S.; Kesavadas, C.; Krishnamurthi, G. Semi-supervised Learning using Denoising Autoencoders for Brain Lesion Detection and Segmentation. J. Med. Imaging 2017, 4, 041311. [Google Scholar] [CrossRef]
- Giacalone, M.; Rasti, P.; Debs, N.; Frindel, C.; Cho, T.-H.; Grenier, E.; Rousseau, D. Local spatio-temporal encoding of raw perfusion MRI for the prediction of final lesion in stroke. Med. Image Anal. 2018, 50, 117–126. [Google Scholar] [CrossRef]
- Lucas, C.; Kemmling, A.; Mamlouk, A.M.; Heinrich, M.P. Multi-scale neural network for automatic segmentation of ischemic strokes on acute perfusion images. In Proceedings of the 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), Washington, DC, USA, 4–7 April 2018; pp. 1118–1121. [Google Scholar]
- Bento, M.; Souza, R.; Salluzzi, M.; Rittner, L.; Zhang, Y.; Frayne, R. Automatic identification of atherosclerosis subjects in a heterogeneous MR brain imaging data set. Magn. Reson. Imaging 2019, 62, 18–27. [Google Scholar] [CrossRef]
- Song, T. Generative Model-Based Ischemic Stroke Lesion Segmentation. arXiv 2019, arXiv:1906.02392. [Google Scholar]
- Liu, Z.; Cao, C.; Ding, S.; Han, T.; Liu, S. Towards Clinical Diagnosis: Automated Stroke Lesion Segmentation on Multi-Spectral MR Image Using Convolutional Neural Network. IEEE Access 2018, 6, 57006–57016. [Google Scholar] [CrossRef]
- Zhang, R.; Zhao, L.; Lou, W.; Abrigo, J.; Mok, V.C.T.; Chu, W.C.W.; Wang, D.; Shi, L. Automatic Segmentation of Acute Ischemic Stroke From DWI Using 3-D Fully Convolutional DenseNets. IEEE Trans. Med. Imaging 2018, 37, 2149–2160. [Google Scholar] [CrossRef]
- Chen, H.; Dou, Q.; Yu, L.; Qin, J.; Heng, P.A. VoxResNet: Deep voxelwise residual networks for brain segmentation from 3D MR images. NeuroImage 2018, 170, 446–455. [Google Scholar] [CrossRef]
- Li, H.; Jiang, G.; Zhang, J.; Wang, R.; Wang, Z.; Zheng, W.-S.; Menze, B. Fully convolutional network ensembles for white matter hyperintensities segmentation in MR images. NeuroImage 2018, 183, 650–665. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Praveen, G.; Agrawal, A.; Sundaram, P.; Sardesai, S. Ischemic stroke lesion segmentation using stacked sparse autoencoder. Comput. Biol. Med. 2018, 99, 38–52. [Google Scholar] [CrossRef] [PubMed]
- Alex, V.; Safwan, K.P.M.; Chennamsetty, S.S.; Krishnamurthi, G. Generative adversarial networks for brain lesion detection. In Proceedings of the SPIE 10133, Medical Imaging 2017, Indian Institute of Technology, Madras, India, 24 February 2017. [Google Scholar] [CrossRef]
- Li, H.; Zhygallo, A.; Menze, B. Automatic Brain Structures Segmentation Using Deep Residual Dilated U-Net. arXiv 2018, arXiv:1811.04312. [Google Scholar]
- Luna, M.; Park, S.H. 3D Patchwise U-Net with Transition Layers for MR Brain Segmentation. In Proceedings of the International MICCAI Brainlesion Workshop, 4th International Workshop, BrainLes 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, 16 September 2018; Springer: Cham, Switzerland, 2019; pp. 394–403. [Google Scholar] [CrossRef]
- Winzeck, S.; Mocking, S.; Bezerra, R.; Bouts, M.; McIntosh, E.; Diwan, I.; Garg, P.; Chutinet, A.; Kimberly, W.; Copen, W.; et al. Ensemble of Convolutional Neural Networks Improves Automated Segmentation of Acute Ischemic Lesions Using Multiparametric Diffusion-Weighted MRI. Am. J. Neuroradiol. 2019, 40, 938–945. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Chen, S.; Zhang, F.; Wu, F.-X.; Pan, Y.; Wang, J. Deep convolutional neural network for automatically segmenting acute ischemic stroke lesion in multi-modality MRI. Neural Comput. Appl. 2020, 32, 6545–6558. [Google Scholar] [CrossRef]
- Karthik, R.; Gupta, U.; Jha, A.; Rajalakshmi, R.; Menaka, R. A deep supervised approach for ischemic lesion segmentation from multimodal MRI using Fully Convolutional Network. Appl. Soft Comput. 2019, 84, 105685. [Google Scholar] [CrossRef]
- Li, H.; Li, A.; Wang, M. A novel end-to-end brain tumor segmentation method using improved fully convolutional networks. Comput. Biol. Med. 2019, 108, 150–160. [Google Scholar] [CrossRef]
- Malla, C.U.P.; Hernández, M.D.C.V.; Rachmadi, M.F.; Komura, T. Evaluation of Enhanced Learning Techniques for Segmenting Ischaemic Stroke Lesions in Brain Magnetic Resonance Perfusion Images Using a Convolutional Neural Network Scheme. Front. Aging Neurosci. 2019, 13, 33. [Google Scholar] [CrossRef] [Green Version]
- Yang, H.; Huang, W.; Qi, K.; Li, C.; Liu, X.; Wang, M.; Zheng, H.; Wang, S. CLCI-Net: Crosslevel fusion and context inference networks for lesion segmentation of chronic stroke. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Shenzhen, China, 13–17 October 2019; pp. 266–274. [Google Scholar]
- Qi, K.; Yang, H.; Li, C.; Liu, Z.; Wang, M.; Liu, Q.; Wang, S. X-net: Brain stroke lesion segmentation based on depthwise separable convolution and long-range dependencies. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Shenzhen, China, 13–17 October 2019; pp. 247–255. [Google Scholar] [CrossRef] [Green Version]
- Liu, L.; Wu, F.-X.; Wang, J. Efficient multi-kernel DCNN with pixel dropout for stroke MRI segmentation. Neurocomputing 2019, 350, 117–127. [Google Scholar] [CrossRef]
- Chin, D.; Roderick, W.; Wang, K.M. Using Cascaded Networks for Post-Stroke Lesion Detection in the ATLAS Dataset. Available online: http://cs230.stanford.edu/projects_spring_2018/reports/8288136.pdf (accessed on 4 October 2021).
- Wang, Y.-R.; Wang, H.; Chen, S.; Katsaggelos, A.K.; Martersteck, A.; Higgins, J.; Hill, V.B.; Parrish, T. A 3D Cross-Hemisphere Neighborhood Difference Convnet for Chronic Stroke Lesion Segmentation. In Proceedings of the 2019 IEEE International Conference on Image Processing (ICIP), Taipei, Taiwan, 22–25 September 2019; pp. 1545–1549. [Google Scholar]
- Wang, Y.; Katsaggelos, A.K.; Wang, X.; Parrish, T.B. A deep symmetry convnet for stroke lesion segmentation. In Proceedings of the 2016 IEEE International Conference on Image Processing (ICIP), Phoenix, AZ, USA, 3 August 2016; pp. 111–115. [Google Scholar]
- Rajan, R.; Sathish, R.; Sheet, D. Significance of Residual Learning and Boundary Weighted Loss in Ischaemic Stroke Lesion Segmentation. arXiv 2019, arXiv:1908.04840. [Google Scholar]
- Liu, L.; Kurgan, L.; Wu, F.-X.; Wang, J. Attention convolutional neural network for accurate segmentation and quantification of lesions in ischemic stroke disease. Med. Image Anal. 2020, 65, 101791. [Google Scholar] [CrossRef]
- Zhang, L.; Song, R.; Wang, Y.; Zhu, C.; Liu, J.; Yang, J.; Liu, L. Ischemic Stroke Lesion Segmentation Using Multi-Plane Information Fusion. IEEE Access 2020, 8, 45715–45725. [Google Scholar] [CrossRef]
- Amin, J.; Sharif, M.; Gul, N.; Raza, M.; Anjum, M.A.; Nisar, M.W.; Bukhari, S.A.C. Brain Tumor Detection by Using Stacked Autoencoders in Deep Learning. J. Med. Syst. 2019, 44, 32. [Google Scholar] [CrossRef]
- Bui, T.D.; Ahn, S.; Lee, Y.; Shin, J. A Skip-Connected 3D DenseNet Networks with Adversarial Training for Volumetric Segmentation. In International MICCAI Brainlesion Workshop, Proceedings of the 4th International Workshop, BrainLes 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, 16 September 2018; Springer: Cham, Switzerland, 2018; pp. 378–384. [Google Scholar] [CrossRef]
- Joshi, S.; Gore, S. Ishemic Stroke Lesion Segmentation by Analyzing MRI Images Using Dilated and Transposed Convolutions in Convolutional Neural Networks. In Proceedings of the 2018 Fourth International Conference on Computing Communication Control and Automation (ICCUBEA), Pune, India, 16–18 August 2018; pp. 1–5. [Google Scholar]
- Gupta, A.; Vupputuri, A.; Ghosh, N. Delineation of Ischemic Core and Penumbra Volumes from MRI using MSNet Architecture. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; Volume 2019, pp. 6730–6733. [Google Scholar]
- Kumar, A.; Upadhyay, N.; Ghosal, P.; Chowdhury, T.; Das, D.; Mukherjee, A.; Nandi, D. CSNet: A new DeepNet framework for ischemic stroke lesion segmentation. Comput. Methods Programs Biomed. 2020, 193, 105524. [Google Scholar] [CrossRef]
- Sathish, R.; Rajan, R.; Vupputuri, A.; Ghosh, N.; Sheet, D. Adversarially Trained Convolutional Neural Networks for Semantic Segmentation of Ischaemic Stroke Lesion using Multisequence Magnetic Resonance Imaging. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; Volume 2019, pp. 1010–1013. [Google Scholar]
- Phong, T.D.; Duong, H.N.; Nguyen, H.T.; Trong, N.T.; Nguyen, V.H.; Van Hoa, T.; Snasel, V. Brain Hemorrhage Diagnosis by Using Deep Learning. In Proceedings of the 2017 International Conference on Machine Learning and Soft Computing, Ho Chi Minh City, Vietnam, 13–16 January 2017; ACM Press: New York, NY, USA, 2017; pp. 34–39. [Google Scholar]
- Majumdar, A.; Brattain, L.; Telfer, B.; Farris, C.; Scalera, J. Detecting Intracranial Hemorrhage with Deep Learning. In Proceedings of the 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 17–21 July 2018; Volume 2018, pp. 583–587. [Google Scholar]
- Arbabshirani, M.R.; Fornwalt, B.K.; Mongelluzzo, G.J.; Suever, J.D.; Geise, B.D.; Patel, A.A.; Moore, G.J. Advanced machine learning in action: Identification of intracranial hemorrhage on computed tomography scans of the head with clinical workflow integration. NPJ Digit. Med. 2018, 1, 9. [Google Scholar] [CrossRef]
- Kuo, W.; Hӓne, C.; Mukherjee, P.; Malik, J.; Yuh, E.L. Expert-level detection of acute intracranial hemorrhage on head computed tomography using deep learning. Proc. Natl. Acad. Sci. USA 2019, 116, 22737–22745. [Google Scholar] [CrossRef] [Green Version]
- Patel, A.; Schreuder, F.H.B.M.; Klijn, C.J.M.; Prokop, M.; Van Ginneken, B.; Marquering, H.; Roos, Y.B.W.E.M.; Baharoglu, M.I.; Meijer, F.J.A.; Manniesing, R. Intracerebral Haemorrhage Segmentation in Non-Contrast CT. Sci. Rep. 2019, 9, 17858. [Google Scholar] [CrossRef]
- Cho, J.; Park, K.-S.; Karki, M.; Lee, E.; Ko, S.; Kim, J.K.; Lee, D.E.; Choe, J.; Son, J.; Kim, M.; et al. Improving Sensitivity on Identification and Delineation of Intracranial Hemorrhage Lesion Using Cascaded Deep Learning Models. J. Digit. Imaging 2019, 32, 450–461. [Google Scholar] [CrossRef]
- Patel, A.; van de Leemput, S.C.; Prokop, M.; Van Ginneken, B.; Manniesing, R. Image Level Training and Prediction: Intracranial Hemorrhage Identification in 3D Non-Contrast CT. IEEE Access 2019, 7, 92355–92364. [Google Scholar] [CrossRef]
- Barros, R.S.; Van Der Steen, W.E.; Boers, A.M.; Zijlstra, I.; Berg, R.V.D.; El Youssoufi, W.; Urwald, A.; Verbaan, D.; Vandertop, P.; Majoie, C.; et al. Automated segmentation of subarachnoid hemorrhages with convolutional neural networks. Inform. Med. Unlocked 2020, 19, 100321. [Google Scholar] [CrossRef]
- Lee, J.Y.; Kim, J.S.; Kim, T.Y.; Kim, Y.S. Detection and classification of intracranial haemorrhage on CT images using a novel deep-learning algorithm. Sci. Rep. 2020, 10, 20546. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Holanda, G.; Souza, L.F.d.F.; Silva, H.; Gomes, A.; Silva, I.; Ferreira, M.; Jia, C.; Han, T.; de Albuquerque, V.H.C.; et al. Deep Learning-Enhanced Internet of Medical Things to Analyze Brain CT Scans of Hemorrhagic Stroke Patients: A New Approach. IEEE Sens. J. 2020, 21, 24941–24951. [Google Scholar] [CrossRef]
- Li, L.; Wei, M.; Liu, B.; Atchaneeyasakul, K.; Zhou, F.; Pan, Z.; Kumar, S.A.; Zhang, J.Y.; Pu, Y.; Liebeskind, D.S.; et al. Deep Learning for Hemorrhagic Lesion Detection and Segmentation on Brain CT Images. IEEE J. Biomed. Health Inform. 2021, 25, 1646–1659. [Google Scholar] [CrossRef] [PubMed]
- Arab, A.; Chinda, B.; Medvedev, G.; Siu, W.; Guo, H.; Gu, T.; Moreno, S.; Hamarneh, G.; Ester, M.; Song, X. A fast and fully-automated deep-learning approach for accurate hemorrhage segmentation and volume quantification in non-contrast whole-head CT. Sci. Rep. 2020, 10, 19389. [Google Scholar] [CrossRef]
- Grewal, M.; Srivastava, M.M.; Kumar, P.; Varadarajan, S. RADnet: Radiologist level accuracy using deep learning for hemorrhage detection in CT scans. In Proceedings of the 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), Washington, DC, USA, 4–7 April 2018; pp. 281–284. [Google Scholar]
- Burduja, M.; Ionescu, R.; Verga, N. Accurate and Efficient Intracranial Hemorrhage Detection and Subtype Classification in 3D CT Scans with Convolutional and Long Short-Term Memory Neural Networks. Sensors 2020, 20, 5611. [Google Scholar] [CrossRef]
- Pereira, D.R.; Filho, P.P.R.; de Rosa, G.H.; Papa, J.P.; de Albuquerque, V.H.C. Stroke Lesion Detection Using Convolutional Neural Networks. In Proceedings of the 2018 International Joint Conference on Neural Networks (IJCNN), Rio de Janeiro, Brazil, 8–13 July2018; pp. 1–6. [Google Scholar]
- Marbun, J.T.; Seniman; Andayani, U. Classification of stroke disease using convolutional neural network. J. Phys. Conf. Ser. 2018, 978, 012092. [Google Scholar] [CrossRef]
- Dourado, C.M.J.M., Jr.; da Silva, S.P.P.; da Nóbrega, R.V.M.; Barros, A.C.D.S.; Filho, P.P.R.; de Albuquerque, V.H.C. Deep learning IoT system for online stroke detection in skull computed tomography images. Comput. Netw. 2019, 152, 25–39. [Google Scholar] [CrossRef]
- Kuang, H.; Menon, B.K.; Qiu, W. Segmenting Hemorrhagic and Ischemic Infarct Simultaneously from Follow-Up Non-Contrast CT Images in Patients with Acute Ischemic Stroke. IEEE Access 2019, 7, 39842–39851. [Google Scholar] [CrossRef]
- Xue, Y.; Farhat, F.G.; Boukrina, O.; Barrett, A.; Binder, J.R.; Roshan, U.W.; Graves, W.W. A multi-path 2.5 dimensional convolutional neural network system for segmenting stroke lesions in brain MRI images. NeuroImage Clin. 2020, 25, 102118. [Google Scholar] [CrossRef]
- Bria, A.; Marrocco, C.; Tortorella, F. Addressing class imbalance in deep learning for small lesion detection on medical images. Comput. Biol. Med. 2020, 120, 10373. [Google Scholar] [CrossRef]
- Sipser, M. Introduction to the Theory of Computation. ACM Sigact News 1996, 27, 27–29. [Google Scholar] [CrossRef]
- Huang, K.; Rhee, D.J.; Ger, R.; Layman, R.; Yang, J.; Cardenas, C.E.; Court, L.E. Impact of slice thickness, pixel size, and CT dose on the performance of automatic contouring algorithms. J. Appl. Clin. Med. Phys. 2021, 22, 168–174. [Google Scholar] [CrossRef]







| Modality | Database | Data Size | Area | Classes | Ground Truth | Data Info |
|---|---|---|---|---|---|---|
| MRI | ISLES 2015 | SISS: 28(train) 36(test) SPES: 30(train) 20(test) | SISS: sub–acute ischemic stroke lesion segmentation SPES: acute stroke outcome/penumbra estimation | SISS: Lesions were classified as sub–acute infarct and Infarct lesions. SPES: target mismatch = perfusion–restriction label minus diffusion–restriction label | SISS: Segmentation by an Expert | http://www.isles-challenge.org/ISLES2015 (accessed on 4 October 2021). |
| MRI | ISLES 2016 | 35(train) 19(test) | Dataset provides a regression and segmentation and a task: Task 1: prediction of Lesion outcome Task 2: prediction of Clinical outcome | MODIFIED RANKIN SCALE (MRS) The 90 days mRS is a scale to assess the degree of disability 90 days after a stroke incidence (Task II assessment) (Grade: G)
| Final lesion volume (Task 1) as manually and the clinical mRM score (Task 2) denoting the extent of disability | http://www.isles-challenge.org/ISLES2016/ Highest IDC = 3.37 |
| MRI– (DWI, ADC) | ISLES 2017 | 43(train) 32(test) | Acute ischemic stroke (Challenge for stroke lesions segmentation, core and penumbra separation) | Ground–truth segmentation maps manually drawn on scans | Lesion outcome (prediction) based on acute MRI data. http://www.isles-challenge.org/ISLES2017/ Highest IDC = 4.53 | |
| CBF, MTT, CBV, TMAX, CTP | ISLES 2018 | 63(train) 40(test) | Penumbra–core separation using CT | Expert segmentations of the infarct lesions. | Acute ischemic stroke patients with 8 hrs. of stroke onset and MRI DWI within 3 h. after CTP. http://www.isles-challenge.org/ |
| Inclusion | Exclusion |
|---|---|
| Studies pertaining to | Studies pertaining to |
| 1. CT and MRI (including variants) | 1. Treatment of strokes (Exclusively) |
| 2. Ischemic and hemorrhagic strokes | 2. Pure Statistical and Biological methods of treatment. |
| 3. Measurement of the degree of the infarct and damage. | 3. Technical working and advancement of algorithms |
| 4. Prognosis of strokes and the likelihood of damage | 4. Lesions extraneous to strokes |
| 5. Lesion detection and segmentation (core and penumbra region) | |
| 6. ML and DL techniques for segmentation of lesion regions | |
| 7. Latest architectures in DL techniques and factorization techniques for feature–specific algorithms. |
| Modality | Description |
|---|---|
| NCCT (CT) | CT uses a beam of X–rays followed by a process of high–powered computers to generate images of soft tissues and bones. Overall sensitivity of 57–71% and 12% in the first 24 h, 3 h respectively [31,32] |
| Perfusion CT | These scans help identify areas adequately supplied with blood (perfused) and provide detailed information about blood flow to the brain. Regions which demonstrate matched defects in MTT and CBV represent the unsalvageable infarct core, whereas regions with prolonged MTT, but preserved CBV are considered to be the ischemic penumbra, and are potentially salvageable [32] |
| Angiography CT | CT angiography is a type of medical test that combines a CT scan with an injection of a special dye to produce pictures of blood vessels and tissues. Within an intracranial vessel it may also identify thrombus, and may guide for intra–arterial thrombolysis or clot retrieval [32] |
| MRI | MRI is based on the magnetization properties of atomic nuclei. Protons in the water nuclei of tissues are excited and relaxed, and subsequently capturing the released energy. Based on the relaxation time, T1 and T2 tissues are characterized [32]. |
| T1 weighted (MRI) | Characterized by shorter relaxation time. Following noticeable changes in scans [32]
|
| T2 weighted (MRI) | Characterized by longer relaxation time. Following noticeable changes in scans [32]
|
| Flair (MRI) | Characterized by longer relaxation time than T2 weighted images. Following noticeable changes in scans [32]
|
| DWI (MRI) | Detect the random movements of water protons. Spontaneous movements, rapidly become restricted in ischemic brain tissue which appear bright in scans. It is an extremely sensitive method for detecting acute stroke. [32] Apparent diffusion coefficient (ADC) is a measure of the magnitude of diffusion (of water molecules) within tissue. Rough values (10−6 mm2/s):
|
| Stroke | Ischemic | Hemorrhagic | ||||
|---|---|---|---|---|---|---|
| Modality | Acute (0–7 days) | Subacute (1–3 Weeks) | Chronic (>3 Weeks) | Acute (0–7 Days) | Subacute (1–3 Weeks) | Chronic (>3 Weeks) |
| NCCT | Loss of grey–white matter differentiation, and hypo attenuation (low density, obstruction) of deep nuclei [31,32] | Attenuation of the cortex [31,32] | Hypo density region [31,32] | Hyper dense with fluid levels [33] | Less intense with ring–like profile [33] | Iso dense or modest confined hypo density [33] |
| T1 | Low T1 signal [32] | Low T1 signal [32] | Low T1 signal [32] | Iso intensity or slight hypo intensity with thin hyper intense rim in the periphery [32,33] | Hyper intensity [32,33] | Hypo intensity [32,33] |
| T2 | infarct remains Hyper intense [32] | Hyper intensity [32] | High T2 signal [32] | Hypo intense with hyper intense perilesional rim [32,33] | Hyper intensity [32,33] | Hypo intensity [32] |
| DWI | Decreased ADC values with maximal signal reduction within 1 to 4 days marked with hyper intensity [32] | First ADC values rise and return close to baseline, despite normal ADC values irreversible tissue necrosis is present (DWI remains hyper intense) [32] | ADC signal high [32] | ADC: 0.70 [35] | ADC: 0.72 [35] | ADC: 2.56 [35] |
| Articles | Modality | Technique | Outcome | Year |
|---|---|---|---|---|
| Tang et al. [38] | CT | Image texture analysis through Circular Adaptive Region of Interest method | SROC: [0.99–0.94] | 2011 |
| Sajjadi et al. [39] | CT | Translation–invariant wavelet for image enhancement | Higher information image extracted | 2011 |
| Nowinski et al. [40] | NCCT | Analyzing hemisphere attenuation values using percentile difference ratios | SAcc is 83.2%. The early detection accuracy (<3 h) is 78.4%. | 2013 |
| Filho et al. [41] | CT | Analysis of brain tissue density | 2017 | |
| Flottman et al. [42] | CT | Novel threshold free method | 2017 | |
| Lo et al. [44] | NCCT | Local contract enhancement using Ranklet Transformation and probability based detection | GLCM Ranklet SACC 71% 81% | 2019 |
| Bhaduria et al. [48] | CT | Segmenting through the features of both fuzzy clustering and region–based active contour model | SDC: 0.92 | 2014 |
| Haan et al. [49] | CT, DWI, T2FLAIR | Clustering algorithm for lesion demarcation in AIS | Reduced processing time to on average 17.8 min/patient | 2015 |
| YAHIAOUI et al. [50] | CT | Differentiation of brain pathology area (hypodense) from its adjacent normal parenchym (i.e., contrast enhancement) using Laplacian Pyramid | Laplacian Pyramid algorithm gives Better and faster (10.46 s) result than DWT, especially in small sized lesions. | 2016 |
| Reboucas et al. [51] | CT | Level set based approach on brain densities (radiological) method to generate stroke segmentation | Segmentation time and SACC LSBRD (proposed) 1.76, 99% Watershed 3.10, 92% Region Growing 4.81, 93% | 2017 |
| Kumar et al. [52] | CT | Entropy based segmentation | SACC: 99.87 (avg) | 2020 |
| Vasconcelos et al. [53] | CT | Adaptive Brain Tissue Density Analysis | CACC: 98.13% | 2020 |
| Nabizadeh et al. [54] | MRI | Histogram–based gravitational optimization algorithm | SACC: 91.5%(strokes) | 2014 |
| Ghosh et al. [55] | Hierarchical Region Splitting, Symmetry Integrated Region Growing and Modified Watershed Segmentation | 2014 | ||
| Ledig et al. [57] | MRI | Refinement using Multi–Atlas Label based context with Expectation–Maximization. | 64.7% SACC using acute–phase | 2015 |
| Farsani et al. [58] | MRI | Diffusion restricted characterisitics | CACC: 73% | 2016 |
| Moeskops et al. [59] | MRI | CNN | SDC: 0.84–0.91 | 2017 |
| Ji et al. [63] | MRI | Gaussian Mixture Model | SACC: 5% more than baseline model | 2017 |
| Kamnitsas et al. [64] | MRI | A 11 layered dual pathway architecture for joint processing of adjacent image patches (DeepMedic) | SDC on training data of BRATS 2015 DeepMedic + CRF 89 .8 DeepMedic 89 .7 | 2017 |
| Articles | Modality | Technique | Outcome | Year |
|---|---|---|---|---|
| Filho et al. [41] | CT | Feature extraction based on density patterns (radiological) and classification of strokes through Bayesian, SVM, kNN, MLP, and OPF classifiers | Fastest extraction time IACC: 99.30% | 2017 |
| Rajini et al. [65] | CT | Symmetry (mid line shift) based segmentation; image texture analysis using GLCC and classification using SVM, k–NN, ANN, decision tree | SVM IACC: 98% | 2013 |
| Maier et al. [72] | MRI | Generalized Linear Models, RFs and CNN are evaluated and compared with each other for sub–acute ischemic stroke patients | AdaBoost IDC: 0.69 | 2015 |
| Mitra et al. [73] | FLAIR MRI | Bayesian–Markov Random Field and RF | IDC: 0.60 ± 0.12 | 2014 |
| Bharathi et al. [74] | MRI T1, T2, DWI and FLAIR | Feature Extraction using GLCM and unsupervised extraction Kmeans clustering; and training RF classifier for detection of ischemic stroke lesion | IDC: 0.88 IACC 0.82 | 2019 |
| Maier et al. [76] | T1w, T2w, FLAIR and DWI | Extra Tree Forest framework for voxel–wise classification | IDC: 0.65 ± 0.18 | 2015 |
| McKinley et al. [83] | MRI T1, T2 | Spatial Random Forest | ISLES (leave one out) IDC: 0.85 (±0.06) | 2015 |
| Robben et al. [84] | T1w– and T2w, Flair and DWI | cascaded extremely randomized trees | IDC SISS 0.57 ± 0.28 SPES 0.82 ± 0.07 | 2016 |
| Chen et al. [85] | MRI | random forests (cascaded) with dense conditional randomfields | ISLES 2015/BRATS 2018 IDC of 0.51 ± 0.29/0.86 | 2020 |
| Griffanti et al. [87] | T2 and FLAIR | k–nearest neighbor | IICC: 0.99 | 2016 |
| Griffis et al. [88] | T1 | Gaussian naïve Bayes | IDC 0.66 | 2016 |
| Karthik et al. [89] | MRI | Multidirectional features based on Discrete curvelet transform and watershed algorithm for fetching the ROI and then applying support vector machines to develop the classification system. | IACC 99.1% | 2017 |
| Pereira et al. [90] | MRI | Unsupervised feature learning through RBM with RF classifier | IDC 0.81 ± 0.84 | 2018 |
| Lin et al. [91] | CT | DBSCAN, hierarchical DBSCAN (HDBSCAN) and local outlier factor (LOF) for identification of erroneous stroke detection | DBSCAN (Avg) IACC 96.9 | 2019 |
| Subudhia et al. [92] | MRI | Delaunay triangulation based segmentation optimized by Darwinian particle swarm optimization | IACC of 0.95 | 2018 |
| Peixoto et al. [93] | CT | SCM, SVM, MLP | ISPEC = 99.1%[highest] | 2018 |
| Garg et al. [94] | Electronic Data (NLP) | Classification of Ischemic Stroke Subtype (TOAST) using ML (RF, GBM, KNN, XGBOOST, SVM, Extra Trees) and NLP | Kappa stacking: combined data = 0.57 | 2019 |
| Articles | Modality | Technique | Loss Function | Outcome | Year |
|---|---|---|---|---|---|
| Lisowska et al. [99] | NCCT | Bilateral CNN + Atlas | squared hinge loss | IAUC: 0.964 | 2020 |
| Abulnaga et al. [100] | CTP | Pyramid Scene Parsing Network | Focal Loss | IDC:0.54 ± 0.009 | 2017 |
| Vargas et al. [102] | CTP | CNN LSTM [Train 356, Validation 40] | IACC: 87.5% | 2018 | |
| Barman et al. [103] | CT A | DeepSymNet Two identical CNNs with 3 Inception module for learning the low and high level volume 3D representation common to the two brain hemispheres. | L–1 difference | IAUC: 91.4% | 2019 |
| Clèrigues et al. [104] | CT, CT–PWI CBF, CBV& MTT | DL based segmentation approach using 2D patch based for of the acute stroke lesion core. | To minimize the effects of class imbalance Generalized Dice Loss (GDL) with the cross entropy loss. | IDSC improvement of 4.5% over the baseline [ISLES 2018] | 2019 |
| Shinohara et al. [105] | NCCT | Xception architecture pre–trained on the ImageNet database | classification loss | ISPEC: 89.7% IACC: 86.5% | 2020 |
| Barros et al. [106] | NCCT | CNN with two convolutional layers (256 nodes, 64/128 feature resp.) followed by 2 FCN. Each dense layer has. Max–polling layer with a 2 × 2 kernel and a 2 × 2 stride. | Severe IACC: 0.98 Intermediate IACC: 0.93 Subtle IACC: 0.66 | 2019 | |
| Oman et al. [107] | CTA, NCCT | 3D CNN | IDC: 0.61 | 2019 | |
| Hu et al. [108] | 3D MRI | 3D residual framework | Focal Loss | BRATS 2015 IDC: 0.86 (whole) | 2020 |
| Bertels et al. [110] | CTP | Contra Lateral Information CNN | Binary cross–entropy | IDC: 0.45 [ISLES 2018] | 2018 |
| Kuang et al. [111] | NCCT | EIS–Net Triple–CNN with three triple encoders and one de–coder with multi–level attention gate modules. | Combination of weighted binary cross entropy and Generalized Dice–Coefficient. | IACC: EIS–Net 85.7% | 2021 |
| Avetisian et al. [112] | NCCT | Dual Path Network which fusing the features of Res–Nets and densely–connected networks | Focal Loss | IDC: 0.703 | 2020 |
| Wang et al. [114] | MRI | 3D RF trained on ISLES dataset | Hybrid loss function | IDC: 0.16 ± 0.31 [test] | 2016 |
| Havaei M [116] | T1, T2, T1C and Flair | CNN (two pathways cascaded architecture) | cross–entropy loss | SISS IDC: 0.69 SPES IDC: 0.85 | 2020 |
| Chen et al. [121] | DWI | CNN | Cross Entropy | IDC: 0.67 [avg] | 2016 |
| Lucas et al. [121] | FLAIR, DWI, T1, and T2 | FCNN–MatConvNet | cross–entropy loss | IDC: 0.59 | 2017 |
| Alex et al. [123] | T1, T2,T1C FLAIR | Stacked denoising autoencoders | High and Low Grade Glioma | 2017 | |
| Lucas et al. [125] | MRI | Res–UNets | Weighted sum of a classification and soft QDice metric | 33% lower surface distance than U–Net | 2017 |
| Liu et al. [128] | MRI | FCN (Res–FCN) | Customized Loss Function | IDC: 0.645 | 2018 |
| Zhang et al. [129] | DWI | 3D FC–DenseNet | Customized Loss function + Dice Loss function | IDC: 0.79 [Best] | 2018 |
| Chen et al. [130] | 3D MRI | VoxResNet: Stacked residual modules with convolutional/de–convolutional (total 25 volumetric) | spatial information loss | IDC GM 86.15 WM 89.46 CSF 84.25 | 2018 |
| Li et al. [131] | MRI FLAIR | Two convolutional layers are repeatedly employed, each with ReLU and a 2 × 2 (max pooling), down–sampling with stride 2 | Dice Loss | MICCAI 2017 IDSC: 0.80 | 2018 |
| Praveen et al. [132] | FLAIR, DWI, T1, and T2 | Stacked Sparse autoencoder layers and support vector machine classifier as the output layer. | Mean Squared Loss | ISLES 2015 IDC: 0.943 ± 0.057 | 2018 |
| Li et al. [134] | CT, DPWI, CBF | Deep Residual Dilated U-Net | Cross–entropy loss | MICCAI IDC: 0.81 | 2018 |
| Luna et al. [135] | MRI | 3D CNN | normalized categorical cross entropy loss | MRBrainS18 Weighted DC 4.44 | 2019 |
| Winzeck et al. [136] | MRI | Ensemble Res–CNN: | Costumed Loss Function | IDC: 82.2% | 2019 |
| Li et al. [139] | T1, T2, T1c and FLAIR | U–Net structure with a new cross–layer architecture (up skip connection) and incorporating inception modules | DSC | [train] IDC: BRATS 15 0.89 BRATS 17 0.876 | 2018 |
| Malla et al. [140] | MRI | CNN [Deepmedic] | Dice Similarity Coefficient | 17% improved IDC: over BS | 2019 |
| Yang et al. [141] | T1 MRI | Cross–level fusion with context (inference) network for stroke lesion segmentation (chronic) | DLF | ATLAS IDC: 0.58 | 2019 |
| Qi et al. [142] | MRI | X–Net (a nonlocal operation to capture long–ranged dependencies) or the chronic stroke lesion segmentation | DLF | ATLAS IDC: 0.48 | 2019 |
| Liu et al. [143] | MRI | multi-kernel DCNN with pixel dropout | DLF | SPES IDC: 0.79 | 2019 |
| Chin et al. [144] | MRI | Cascaded Networks (U-Net) | Train (Private Dataset) IDC: 0.44 | 2020 | |
| Liu et al. [148] | MRI | Attention–based DRANet. | DLF | (748 Images Sub-acute) IDC: 0.76 (Best) | 2016 |
| ZHANG et al. [149] | DWI | A triple–branch DSN architecture with a multi–plane fusion network | Customized Loss Function | ISLES 2015 SSIS IDC: 0.62 | 2020 |
| Amin et al. [150] | MRI | Auto encoders [segmentation] | IDC: 0.96 (BRATS) | 2020 | |
| Bui et al. [151] | MRI | 3D Dense Net | modified DLF | MRBrainS18 IDC 0.87 | 2019 |
| Joshi et al. [152] | DWI-MRI | Dilated and Transposed CNN | Binary cross entropy plus the dice loss | ISLES 2015–2017 (train 25000, validation 4000) IDC: 0.85 (validation) and IJACD: 0.78 | 2018 |
| Gupta et al. [153] | MRI | Multi–Sequence Network architecture: Conv. Layers, Pooling Layers (2 × 2), Up sampling layers (2 × 2), Dropout Layers, | Binary Cross–entropy | ISLES 2015 Core Esti IDC: 0.68 Penumbra Esti. IDC: 0.82 | 2019 |
| Kumar et al. [154] | MRI | Classifier–Segmenter network (modified UNet for segmentation) | multi–scale loss function (customized) | ISLES 2017–SPES dataset IDC: 0.83 | 2020 |
| Satish et al. [155] | DWI, PWI | Adversarial Architecture: Encoder–decoder as segmentor. Discriminators: CNN | cross–entropy | ISLES 2015 IDC: 0.82 | 2020 |
| Modality | Articles | Technique | Loss Function | Outcome | Year |
|---|---|---|---|---|---|
| CT | Phong et al. [156] | LeNet, GoogLeNet, and Inception–ResNet Private Dataset of 1700 records | F1 Score 0.997 (LeNet) | 2017 | |
| CT | Majumdar et al. [157] | 9 (3 × 3) convolutional blocks, (2 × 2) max–pooling, BN and ReLU | 81% HSENS per lesion 98% HSPEC per case | 2018 | |
| NCCT | Patel et al. [160] | CNN with two distinct pathways integrating contextual information | categorical cross entropy | HDC: 0.91 | 2019 |
| CT | Cho et al. [161] | FCN–8s | HACC: 98.28% | 2019 | |
| CT | Patel et al. [162] | CNN and RNN | Binary Cross Entropy | HACC: 0.87 | 2019 |
| NCCT | Barros et al. [163] | CNN | HDC: 0.63 ± 0.16 | 2020 | |
| NCCT | Lee et al. [164] | CNN | HAUC: 0.903 | 2020 | |
| CT | Xu et al. [165] | Masked RNN and ML | Model[Resnet50+ MLP+MobileNET] IACC: EM 99.89 [Best] | 2020 | |
| CT | Li et al. [166] | Pre–trained Dilated UNet | HDC: 0.8033 | 2021 | |
| NCCT | Arab et al. [167] | U–Net with deep supervision. Encoder: Residual block Decoder: Convl layers | Dice similarity coefficients | HDC: 0.84 ± 0.06 | 2020 |
| CT | Grewal et al. [168] | Recurrent Attention DenseNet, bidirectional LSTM layer | HACC: 0.8182 | 2018 | |
| NCCT | Burduja et al. [169] | CNN & LSTM | Binary cross–entropy | HLL: 0.04989 | 2020 |
| Articles | Modality | Technique | Prediction | Year |
|---|---|---|---|---|
| Rebouças et al. [51] | CT | Feature extraction based on density patterns (radiological) and classification of strokes through kNN, SVM, MLP, OPF and Bayesian classifiers | Identify & classify the occurrence of strokes (extent and severity). | 2017 |
| Robben et al. [84] | CTP | Modifed DeepMedic | Final infarct volume | 2019 |
| Bentley et al. [97] | CT | SVM with an HAUC: 0.744 | Predict symptomatic intracranial hemorrhage | 2014 |
| Stier et al. [117] | Tmax MRI | CNN with 2 Conv layers, 2 6x6 pooling layers, trained with 100 epochs for Binary prediction | Tissue Fate Features in AIS | 2016 |
| Choi et al. [119] | MRI | Lesion outcome prediction—3D Res U–Net—CNN Clinical outcome prediction–CNN–Log Regression | Automated prognosis for post–treatment ischemic stroke | 2016 |
| Chen et al. [121] | CT | CNN | Early stroke detection (ischemic) system with CNN | 2017 |
| Lucas et al. [122] | CT | 3D U–net appended with Convolutional auto–encoder | 2018 | |
| Lucas et al. [125] | CT | 3D UNets | Predict Ischemic Stroke Growth | 2018 |
| Bento et al. [126] | SVM IACC: 97.5% | Early identification of Carotidartery Atherosclerosis | 2019 | |
| Song et al. [127] | GAN IDC: 0.624 | Prediction of perfusion parameters | 2019 | |
| Giacalone et al. [124] | SVM IPRES: 95% | Final lesion prediction | 2018 | |
| Arbabshirani et al. [158] | CT | DCNN HAUC: 0.846 | Detecting of ICH based on clinical database of brain CT images |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Inamdar, M.A.; Raghavendra, U.; Gudigar, A.; Chakole, Y.; Hegde, A.; Menon, G.R.; Barua, P.; Palmer, E.E.; Cheong, K.H.; Chan, W.Y.; et al. A Review on Computer Aided Diagnosis of Acute Brain Stroke. Sensors 2021, 21, 8507. https://doi.org/10.3390/s21248507
Inamdar MA, Raghavendra U, Gudigar A, Chakole Y, Hegde A, Menon GR, Barua P, Palmer EE, Cheong KH, Chan WY, et al. A Review on Computer Aided Diagnosis of Acute Brain Stroke. Sensors. 2021; 21(24):8507. https://doi.org/10.3390/s21248507
Chicago/Turabian StyleInamdar, Mahesh Anil, Udupi Raghavendra, Anjan Gudigar, Yashas Chakole, Ajay Hegde, Girish R. Menon, Prabal Barua, Elizabeth Emma Palmer, Kang Hao Cheong, Wai Yee Chan, and et al. 2021. "A Review on Computer Aided Diagnosis of Acute Brain Stroke" Sensors 21, no. 24: 8507. https://doi.org/10.3390/s21248507
APA StyleInamdar, M. A., Raghavendra, U., Gudigar, A., Chakole, Y., Hegde, A., Menon, G. R., Barua, P., Palmer, E. E., Cheong, K. H., Chan, W. Y., Ciaccio, E. J., & Acharya, U. R. (2021). A Review on Computer Aided Diagnosis of Acute Brain Stroke. Sensors, 21(24), 8507. https://doi.org/10.3390/s21248507









