1. Introduction
Bioluminescence and chemiluminescence refer to the process of visible light emission that occurs via exergonic chemical reactions, whereas fluorescence and phosphorescence involve remission of light from the singlet and triplet excited states, respectively [
1]. Bioluminescence has long been used to image biological process in vivo. It can be detected in tissue and living organisms, providing sensitive and invasive data regarding physiological conditions [
2].
Bacterial bioluminescence reaction involves the oxidation of long chain fatty aldehyde and reduced flavin mononucleotide (FMNH
2) with the emission of greenish-blue light [
3] (
Figure 1).
The genes coding for the enzymes and proteins responsible for the light- emitting reaction in the bioluminescent bacteria exist in a cluster forming an operon [
4]. The genes related to riboflavin (vitamin B
2) biosynthesis on the downstream of the
lux operon of
Photobacterium species were reported [
5,
6,
7] (
Figure 2). The biosynthesis of riboflavin is essential in the bioluminescent bacteria, as it is a precursor of FMNH
2 (riboflavin 5′-phosphate), which is the substrate of the bioluminescence reaction as shown in
Figure 1. The detection of the riboflavin genes just downstream of
luxG is particularly relevant to luminescence in
Photobacterium species as this genus produces the highest level of light intensities of any luminescent bacteria, with luciferase levels reaching up to 20% of the soluble proteins [
8].
The fact that the first gene (
ribE) next to
luxG codes for riboflavin synthase in
Photobacterium species is interested in the aspect of molecular genetics. This protein is thought to be created through gene duplication, as it shows about 30% amino acid identity with riboflavin synthase and is under the superfamily of riboflavin synthase [
6,
9]. The lumazine protein, found in most
Photobacterium species in 1970 [
10], was the first antenna protein that shortens the wavelength and amplifies the maximum bioluminescence intensity in
Photobacterium species [
11]. The lumazine protein found in
Photobacterium species are a paralog of riboflavin synthase which are devoid of enzymatic activity but bind the riboflavin synthase substrate, 6,7-dimethyl 8-ribityllumazine with high affinity, and serve as an optical transponder for bioluminescence emission [
11,
12].
They show differences in the quaternary structure of protein, the lumazine protein has a monomeric structure whereas the riboflavin synthase forms trimeric structure [
6,
13]. The peculiar characteristics of lumazine protein is the intramolecular sequence similarity between N-terminal and C-terminal domain half (
Figure 3a), and it was reported that the protein binds to one molecule of lumazine at N-terminal domain half [
14,
15] (
Figure 3b).
In this research, we inserted a gene coding for the lumazine protein into the pRFN4 plasmid, which contains the riboflavin biosynthesis genes of
Bacillus subtilis, in order to increase the fluorescence intensity since the lumazine chromophore binds to the lumazine protein with a high affinity. The pRFN4 recombinant plasmid containing whole riboflavin operon of
Bacillus subtilis were generated to produce 6,7-dimethyl 8-ribityllumazine by the missense mutation of F2A in
ribE gene for riboflavin synthase [
16] (
Figure 4).
Internal sequence similarity (
Figure 3a) as well as the comparison with the paralogous of riboflavin synthase (RS), whose three-dimensional structure of N-terminal domain half has been determined by NMR and X-ray crystallization [
17,
18] (
Figure 3b), suggest that lumazine protein folds into two domains with closely similar folding topology. Therefore, a truncated gene specifying the N-terminal domain half of the lumazine protein (N-LumP) was constructed and the cognate protein was expressed. Using the ligand of 6,7-dimethyl-8-ribityllumazine, the fluorescence characteristics of N-LumP was investigated and reveal that the N-terminal domain half itself can bind lumazine and emit fluorescence [
19]. In the paper, the gene coding for half of the N-terminal domain of lumazine protein (N-LumP), as the minimal version of fluorescent lumazine protein, as well as the whole gene coding for the whole lumazine protein (W-LumP) were amplified by polymerase chain reaction (PCR) and inserted to pRFN4 to generate recombinant plasmids for producing of fluorescent bacteria (
Figure 4).
In addition, we also inserted the DNA of the
lux promoter region from
P. leiognathi into the above recombinant plasmids to increase the expression of the lumazine protein gene. The
lux promoter DNA from bioluminescent bacteria of
Photobacterium species was amplified by PCR and ligated into pRFN4, Pl W-
lumP of pRFN4, and Pl N-
lumP of pRFN4 plasmids with the gene coding for whole lumazine protein (W-
lumP) or N-terminal domain half of lumazine protein (N-
lumP) from
P. leiognathi. Therefore, the recombinant plasmids were constructed into Pl
luxC promoter of pRFN4, Pl
luxC promoter of Pl W-
lumP of pRFN4, Pl
luxC promoter of Pl N-
lumP of pRFN4, respectively (
Figure 4). We transformed into the several
E. coli strains with the pRFN4 plasmid and the various recombined plasmids. Then, we conducted a spectroscopic study on the transformed
E. coli, evaluating the fluorescence intensity and examining the fluorescence of a single cell by confocal microscopy.
3. Results
In addition to the amino acids such as Ser 48, Thr 50, and Ala 66 at the binding sites in N-terminal half of lumazine protein, several studies have shown that Asn 101 and Ile 102 located beyond N-terminal region are involved in binding of the lumazine ligand [
14] (
Figure 3a,b). Therefore, the genes coding for N-terminal domain half of lumazine protein extending to the amino acid 112 (N-LumP) and for the whole lumazine protein (W-LumP) from
P. leiognathi were synthesized by PCR, ligated into pRFN4 vector to construct the recombinant plasmids of N-
lumP-pRFN4 and W-
lumP-pRFN4. We also conducted the PCR process inserting the
luxC promoter domain, whose template is the PlXba.pT7-3 plasmid containing the
lux genes
P. leiognathi [
20]. From these experiments, we generated that the recombinant plasmids, such as the Pl
luxC promoter of pRFN4, the Pl
luxC promoter of Pl W-
lumP of pRFN4, and the Pl
luxC promoter of Pl N-
lumP of pRFN4, had sizes of around 9.0 kbp, 9.5 kbp, and 9.3 kbp, respectively (
Figure 4).
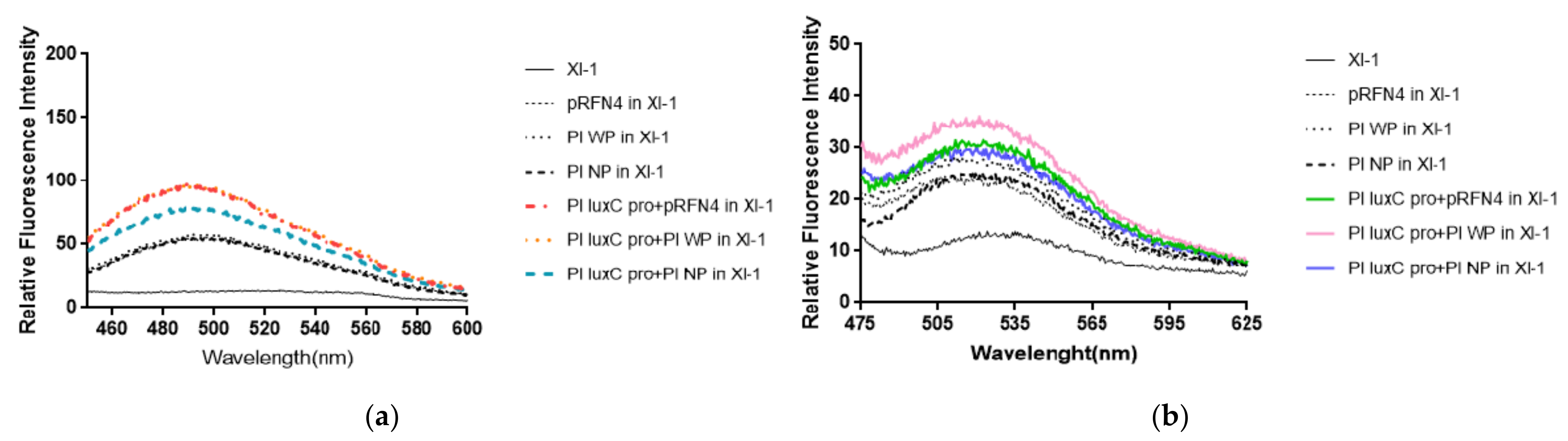
Fluorescent intensities from
E. coli cells harboring the recombinant plasmid containing the genes coding for the N-terminal domain half (N-LumP) and the whole lumazine protein (W-LumP) from
P. leiognathi were tested. The fluorescence intensities of supernatants of the cells in buffer A was measured with spectrofluorimeter. It was observed that the supernatant before sonication also show fluorescent, indicating that the fluorophore was present in the liquid media. Fluorescence was detected by checking for the presence of 6,7-dimethyl-8-ribityllumazine (excitation at 410 nm, molar absorptivity 10,300/M·cm) and of riboflavin (excitation at 450 nm, molar absorptivity 13,153/M·cm). We fixed the excitation wavelength of lumazine at 410 nm and measured the emission spectrum. The fluorescence intensity of
E. coli XL-1 Blue that was transformed with Pl W-
lumP of pRFN4 was about 1.2∼2 times stronger than that of
E. coli XL-1 Blue that was transformed with pRFN4 plasmid. The fluorescence intensity of
E. coli XL-1 Blue that was transformed with pRFN4 plasmid containing the DNA of the
luxC promoter region of
P. leiognathi ATCC25521 was about 1.5∼2 times stronger than that of
E. coli XL-1 Blue that was transformed with pRFN4 plasmid. Among all transformed
E. coli, the fluorescence intensity was the highest in the
E. coli XL-1 Blue that was transformed with the Pl W-
lumP of pRFN4 plasmid containing the DNA of the
luxC promoter region (
Figure 5a).
Compared to the E. coli XL-1 Blue that was transformed with pRFN4 plasmid, the fluorescence intensity was stronger in the E. coli XL-1 Blue that was transformed with Pl W-lumP of pRFN4. This result is due to the non-covalent bonding between the lumazine protein and its chromophore lumazine. In addition, the fluorescence intensity can be stimulated due to the increase of expression of the genes coding for the lumazine protein by the insertion of DNA containing the region of the lux promoter region from P. leiognathi.
Based on a previous study that reported that riboflavin competes with lumazine to bind with the lumazine protein [
13], we fixed the excitation wavelength of riboflavin at 450 nm and examined the emission spectrum. As you can see in
Figure 5a, we found that the emission spectrum of riboflavin excited at 450 nm was similar to that of lumazine shown in
Figure 5b. Compared to the
E. coli XL-1 Blue that was transformed with pRFN4 plasmid, the fluorescence intensities from riboflavin was stronger in the
E. coli XL-1 Blue that was transformed with Pl
luxC promoter of Pl W-
lumP of pRFN4 or Pl
luxC promoter of Pl N-
lumP of pRFN4.
The author’s previous studies [
20,
21] show that luminescence intensity was significantly enhanced over 1000 times in
E. coli 43R when transformed with the PlXba.pT7-3 plasmid among different
E. coli strains. According to the result, we expected a larger increase of fluorescence intensity in
E. coli 43R by the transformation of the pRFN4 recombinant plasmids compared to the transformation in
E. coli XL-1, whereby
E. coli transformed with pRFN4 plasmids show high fluorescence intensity (
Figure 6a,b).
We examined the emission spectrum in the transformed
E. coli 43R with the excitation wavelength fixed at 410 nm as we did in
E. coli XL-1. As a result, the
E. coli 43R transformed with Pl N-
lumP of the pRFN4 plasmid showed about 1.5∼2 times stronger fluorescence intensity than the
E. coli 43R transformed with the pRFN4 plasmid. Furthermore, the fluorescence intensity of the
E. coli 43R transformed with the Pl
luxC promoter of Pl N-
lumP of pRFN4 was about 10 times stronger than that of the
E. coli 43R transformed with the pRFN4 plasmid, and it was the strongest of all the transformed
E. coli 43R (
Figure 6a).
Accordingly, we suggest that this result is due to the increase of the binding of the lumazine ligand with the lumazine protein, as the
lux promoter from
P. leiognathi stimulates the expression of the gene coding the N-LumP. In addition, we fixed the excitation wavelength to that of riboflavin at 450 nm and examined the emission spectrum. In line with the emission spectrum of the excitation wavelength fixed to that of lumazine at 410 nm, the intensity of fluorescence was the strongest in the
E. coli 43R transformed with the Pl
luxC promoter of Pl N-
lumP of the pRFN4 plasmid when the excitation wavelength was fixed at 450 nm (
Figure 6b).
Taken together, we analyzed the average value of the emission spectrum of lumazine and riboflavin (
Figure 7a,b). The increase of the fluorescence intensity at 410 nm from the cell extract of
E. coli XL-1 Blue to
E. coli 43R was gradually increased by about 2∼6 times when transformed with the same plasmid (
Figure 7a). Moreover, in the analysis of the mean value of the emission spectrum of riboflavin excited at 450 nm, the increase of the fluorescence intensity from
E. coli XL-1 Blue to
E. coli 43R was sharp and varied by about 5∼6 times when transformed with the same plasmid (
Figure 7b). Therefore, the highest fluorescence intensity of the cell containing the recombinant plasmid of Pl
luxC promoter of Pl N-
lumP of pRFN4 has a value 250 times higher compared to the intensity from
E. coli XL or
E. coli 43R itself.
Finally, we checked the imaging of fluorescent
E. coli with confocal microscopy. The
E. coli 43R that was transformed using the recombination plasmid with the
lux gene of
Photobacterium species was incubated in a small amount of LB medium. We examined the single cell image of fluorescent
E. coli using DAPI (excitation 405 nm) and FITC (excitation 458 nm) filter sets in the super-resolution confocal laser scanning microscope LSM880. The
E. coli 43R that was not transformed did not show any fluorescence. On the other hand, some of the
E. coli 43R that was transformed with the recombinant plasmids of pRFN4 showed fluorescence (
Figure 8). We observed fluorescence in the single cell of
E. coli 43R transformed with the recombinant plasmids of pRFN4 that was inserted with the genes for W-LumP and N-LumP.
The fluorescence was stronger and observed in more cells compared to the
E. coli 43R transformed with only the pRFN4 plasmid. Furthermore, in the
E. coli 43R transformed with the recombinant plasmid that was additionally inserted with the DNA of the
P. leiognathi luxC promoter domain, the fluorescence was more intense and most of the cells expressed the fluorescence as shown in the image (
Figure 8). The fluorescence intensities and frequencies in a single cell of
E. coli transformed with the recombinant plasmids containing the gene coding for the lumazine protein from
P. leiognathi correlated with the result with the fluorescence intensity shown in
Figure 6.
4. Discussion
Bioluminescence imaging of a single cell is often complicated by the requirements of exogenous luciferin that can be poorly cell permeable or produce a high background signal [
3]. Gregor et al. have engineered an improve operon
ilux, which enable long term visualization of single bacterial cells while simultaneously providing information about cellular viability [
22]. Bacterial bioluminescent system and light emission is being applied sensitive and safe assay not only for prokaryote gene expression but single mammalian cells [
23,
24]. Gregor et al. reported high luminescence levels that support the autonomous bioluminescence microscopy of mammalian cells [
23].
We inserted the genes for the lumazine protein and the DNA of the lux promoter region of Photobacterium species into the pRFN4 plasmid that overproduces the chromophore lumazine, generating to various recombinant plasmids such as Pl W-lumP of pRFN4 and Pl N-lumP of pRFN4 plasmids that contain the lumazine protein gene of P. leiognathi. Then, these recombinant plasmids were transformed into the E. coli. We evaluated the fluorescence intensity using a fluorescence spectrometer and performed an imaging study of a single cell through confocal microscopy in each type of transformed E. coli. The E. coli cells that were transformed with the pRFN4 plasmids more clearly expressed the fluorescence by the overproduced lumazine ligand compared to the E. coli that was not transformed. The fluorescence was significantly stronger when E. coli was transformed with the recombinant plasmid that was also inserted with the lumazine protein gene of P. leiognathi with the luxC promoter. This gives spectroscopic evidence that the fluorescence intensity is increased when the lumazine protein is expressed along with the chromophore lumazine or riboflavin.
The recombinant plasmid DNA was transformed into
E. coli strain 43R and the fluorescence intensity of the bacteria was measured. As a result, the recombinant plasmid in which the gene coding for the lumazine protein and the DNA containing
lux promoter, the expression of this gene was so high to show fluorescence in single colony of
E. coli. The increasing variety of fluorescent proteins and FRET (fluorescence resonance energy transfer) has significantly impacted many molecular and cellular investigations [
25]. Therefore, the fluorescent system using bacterial bioluminescence genes can be utilized as a biosensor with high sensitivity and short analysis time.
In this study, we constructed series of pRFN4 recombinant plasmid derivatives, including Pl W-lumP of pRFN4, and Pl N-lumP of pRFN4 plasmids, with the gene coding for whole lumazine protein (W-LumP) or N-terminal domain half of lumazine protein (N-LumP) from P. leiognathi as well as Pl luxC promoter of pRFN4, Pl luxC promoter of Pl W-lumP of pRFN4, and Pl luxC promoter of Pl N-lumP of pRFN4 inserted the lux promoter DNA from bioluminescent bacteria of Photobacterium species, respectively. By inserting of the luxC promoter region into the plasmids, the fluorescent ligand largely bound the lumazine protein because of the high expression of the gene for lumazine protein by lux promoter in the transformed the E. coli. The fluorescent intensity of these transformed the Escherichia coli were measured by fluorescence spectrometer, and a single cell of the transformed E. coli was observed the appearance of fluorescence by using confocal microscopy. Notably, the expression of recombinant plasmids in E. coli 43R was so strong as to allow observation of the fluorescence in a single cell, raising the possibility for the generation of fluorescent bacteria which can provide the basis for a microbial biosensor.
There has been drastic interest in fluorescence for cellular and molecular imaging [
26,
27], and fluorescence is now a dominant methodology applied to biotechnology, flow cytometry, DNA sequencing, and genetic analysis. Fluorescence imaging can provide the information on the localization and measurement of intracellular molecules [
26]. Many genetically encoded proteins that possess fluorescence and FRET have been developed to better understand the spatiotemporal regulation of various cellular process [
26,
27]. Whole cell sensing systems based on bioluminescence have the benefit of detecting a variety of environmental pollutants [
2,
28]. Therefore, the results presented in this paper raise the possibility that the
Escherichia coli 43R harboring the recombinant plasmid inserted the genes coding for lumazine protein and riboflavin biosynthesis can be used as fluorescent bacteria for application in the field of bio-imaging.