The statistical methodology comprises the following steps: (1) Identify structural breaks in all time series. (2) Extract features directly from the time series. (3) Compute classification variables using the additive seasonal Holt–Winters approach. (4) Compute classification variables by means of seasonal ARIMA. (5) Compute classification variables according to the Wold decomposition. (6) Determine the optimum number of classes and establish the class for each sensor, using PCA and k-means algorithm. (7) Check the classification of sensors using sPLS-DA and identify the optimal subset of variables that best discriminate between classes, per method. (8) Check the classification of sensors by means of the RF algorithm and identify the optimal set of variables calculated from each method, that best discriminate between the classes. (9) Propose a methodology for selecting a subset of representative sensors for future long-term monitoring experiments in the museum.
3.3.2. Calculation of Classification Variables—Method M1
This method consists of computing features such as mean, median and maximum, among others, from estimates of the Auto Correlation Function (ACF), Partial Auto Correlation Function (PACF), spectral density and Moving Range (MR) [
45,
46]. The ACF and PACF correlograms of the observed time series are commonly used to fit Auto Regressive Moving Average (ARMA) models. Firstly, a set of variables denoted as Type 1 comprised the mean, MR and PACF, which were estimated for the 8 stages of
values. The average of
was calculated to capture the level or position in each stage of the time series. The mean of MR with order 2 (i.e., average range over 2 past values) was computed to identify sudden shifts or increases in the level of
. For each stage of
, the sample PACF parameters (
at lag
l) were estimated for the first four lags (
l = 1, 2, 3, 4), which are usually the most important ones for capturing the relevant information in time series.
Secondly, another set of variables called Type 2 comprised spectral density and ACF, which were estimated for values after applying the logarithm transformation and regular differencing. The objective of using this transformation and differencing was to stabilize the variances and remove the trend of , so that the computed variables provide information about the seasonal component of the time series. Spectral density was estimated using the periodogram of observed time series ( of signals w). The maximum peak of the periodogram and its frequency were identified. Values of ACF ( at lag l ) were estimated to analyze the correlation between values with the lagged values of the same observed time series at each lag, for the first 72 lags. This criterion was used because the values of the ACF correlogram for further lags were comprised within the limits of a 95% confidence interval in the correlogram.
The different steps involved in M1 are depicted in
Figure 4. Firstly, the 27 time series were split according to the climatic stages observed: Wr1, Cd, Tr, Ht and Wr2 (Data 2). Secondly, some of the main stages (Data 2) were subdivided according to the structural breaks (SB) identified in Wr1, Cd and Ht (Data 3). In the third step (Data 4), the logarithm transformation was applied and, next, one regular differentiation (Data 5). The fifth step consists of applying the formulas of Type 2 variables to
: maximum of periodogram (
) and its frequency (
w), as well as mean, median, range and variance of the sample ACF for the first 72 lags (
,
,
and
, with
). Finally, the formulas of Type 1 variables were applied to
values (Data 3): mean of
(
), mean of MR of order 2 (
) and PACF for the first four lags (
,
,
and
).
The estimates of MR values were computed with the R software according to the function
rollrange from the
QuantTools package [
40]. The sample ACF and sample PACF values were calculated with the function
acf (
stats) [
47] and
pacf (
tseries) [
36], respectively. The values of the periodogram and their frequencies were obtained with the function
spectrum (
stats).
3.3.3. Calculation of Classification Variables—Method M2
The Holt–Winters method (H-W) [
48] is a type of exponential smoothing that is used for forecasting in time series analysis. The seasonal H-W (SH-W) method uses a smoothing equation for each component of a given time series: the level, slope and seasonality, which are denoted at time
as
,
and
, respectively. The additive SH-W prediction function of a time series of T is given by Equation (
1), where
p denotes the number of observations per period and
k is the integer part of
[
49,
50]. This equation was implemented with the conditions:
,
,
and
. When the algorithm converges,
a corresponds to the last level of
,
b is the last slope of
and
–
are the last seasonal contributions of each
. A forecast of
based on all the data up to time
is denoted by
; a simpler notation for
is
.
The method M2 consists of fitting additive SH-W equations in two steps. Firstly, for each stage of the 27 time series of T, the classification variables are the last level of smoothing components of the additive SH-W method per sensor: level (a), slope (b) and seasonal components (). The method was fitted by considering as the number of observations per day. These smoothing components were called as Type 3 variables. Secondly, by considering the complete time series, the first 24 predictions of for each unique additive SH-W model per sensor were regarded as additional classification variables, which were denoted as Type 5 variables.
Although a residual analysis is not necessary when using SH-W method, estimates of the features from residuals were also computed per stage of the time series (Type 4 variables): sum of squared estimate of errors (SSE), maximum of periodogram (
) and its frequency (
w) and several parameters (mean, median, range and variance) of sample ACF for 72 lags (
,
,
and
, with
). Moreover, the Kolmogorov–Smirnov (KS) normality test [
51] and Shapiro–Wilk test (SW) [
52,
53,
54] were applied in order to extract further information from the different stages of the observed time series. The statistic of the KS test (denoted as Dn) was used to compare the empirical distribution function of the residuals with the cumulative distribution function of the normal model. Likewise, the statistic of the SW test (Wn) was employed to detect deviations from normality, due to skewness and/or kurtosis. The statistics of both tests were also used as classification variables because they provide information about deviation from normality for the residuals derived from the SH-W method.
The steps involved in M2 are depicted in
Figure 5. Firstly, the different time series were split according to the climatic stages observed (Data 2) and the structural breaks (SB) identified (Data 3). Secondly, the seasonal H-W method was applied to Data 3 in order to obtain the last level of smoothing components (Type 3 variables) and then the model residuals. The third step consisted of applying the formulas of Type 4 variables to the residuals. Finally, the seasonal H-W method was applied to Data 1 in order to obtain the first 24 predictions of T (Type 5 variables).
The
HoltWinters function (
stats) was used to fit the Additive SH-W method. The
shapiro.test (
stats) and
ks.test (
dgof) [
55] were used to apply the normality tests. Values of the sample ACF and sample PACF were computed with the functions
acf (
stats) and
pacf (
tseries), respectively. Values of the periodogram and their frequencies were calculated with the function
spectrum (
stats).
3.3.4. Calculation of Classification Variables—Method M3
The ARMA model is also known as the Box–Jenkins approach, which focuses on the conditional mean of the time series and assumes that it is stationary [
44]. By contrast, the ARIMA model can be used when a time series is not stationary. It employs regular differencing of the time series prior to fitting an ARMA model. ARIMA
models employ
p Auto Regressive (AR) terms,
d regular differences and
q Moving Average (MA) terms. Parameters of the AR component are denoted as
(
) and parameters of the MA component as
(
). The error terms
are assumed to be a sequence of data not autocorrelated with a null mean, which is called White Noise (WN) [
44]. In addition, if a given time series is assumed to follow an ARIMA process, the conditional variance of residuals is supposed to be constant. If this is not the case, then it is assumed that an ARCH effect exists in the time series. Two of the most important models for capturing such changing conditional variance are the ARCH and Generalized ARCH (GARCH) models [
41].
Seasonal ARIMA
models are more appropriate in this case given the marked daily cycles.
P refers to the number of seasonal AR (SAR) terms,
D to the number of difference necessary to obtain a stationary time series,
Q to the number of seasonal MA (SMA) terms and
S to the number of observations per period (
in this case). Parameters of the SAR component are denoted as
(
) and the SMA component as
(
). The error terms
are assumed to be a WN sequence [
44].
A seasonal ARIMA
model is given by Equation (
2), where the polynomial
is the regular AR operator of order
p,
is the regular MA operator of order
q,
is the seasonal AR operator (SAR) of order
P,
is the seasonal MA operator (SMA) of order
Q and
B is the backshift operator (i.e.,
, for
and
, for
). Furthermore,
represents the seasonal differences while
accounts for the regular differences, so that
is defined as
and
as
[
41]. In this study,
, which was obtained by differentiating the series once regularly (
) and once seasonally (
). Thus,
.
For each stage of the time series, a common seasonal ARIMA model was fitted for the 27 observed time series (see
Table 1). Firstly, each observed stage of the time series
was checked to determine if it could be regarded as stationary, which implies that the mean and variance are constant over time
t and the covariance between one observation and another (in lagged
l steps) from the same time series does not depend on
t [
44]. The ACF and PACF correlograms were used to examine this condition. Furthermore, the Augmented Dickey–Fuller (ADF) test [
56] was applied for checking the null hypothesis of non stationarity, as well as the Lagrange multiplier (LM) test [
57] for examining the null hypothesis about the absence of ARCH effect. In addition, the autocorrelation Ljung–Box Q (LBQ) test [
58] was applied for inspecting the null hypothesis of independence in a given time series. This LBQ test was carried out on the different lags from
to
, where
is the sum of the number of AR, MA, SAR and SMA terms of the seasonal ARIMA models. It was applied to the time series of the model residuals and the squared residuals.
The condition of stationarity is necessary when fitting an ARMA model. For this purpose, logarithmic transformation, one regular differentiation (
) and one seasonal differentiation (
) were applied to all time series of
, in order to stabilize the variance and remove both the trend in mean and seasonal trend [
59]. Seasonal differentiation was applied to the observed time series
, and the results were denoted as
(
), being
.
In order to determine the appropriate values of
and
, the corrected Akaike’s Information Criterion (
) [
49] was used, which checks how well a model fits the time series using the restriction
and
. The most successful model for each stage of the different observed time series
was chosen according to the lowest
value. Next, the maximum likelihood estimation method was used to estimate the parameters of the seasonal ARIMA models [
44]. Different tests were used to determine whether model assumptions were fulfilled. After computing the model residuals, ADF and LBQ [
58] tests were applied to the residuals and their squared values, for 48 lags, in order to evaluate the condition of WN process. The ACF and PACF correlograms were also used. The next step was to evaluate the absence of Arch effects in the residuals. For this purpose, the LM test was applied to the residuals and their squared values [
60,
61]. Although the normality of errors is not an assumption for fitting ARIMA models, the distribution of residuals derived from the fitted models were compared with the normal distribution by means of the Q-Q normal scores plots, as well as the SW and KS normality tests.
Given that the errors of all models cannot be regarded as WN in this case, it is possible that the model residuals contain useful information about the performance of the different time series. In order to extract further information from the residuals, some features were calculated using ACF, PACF and statistics of normality tests, among others. They were used as additional classification variables.
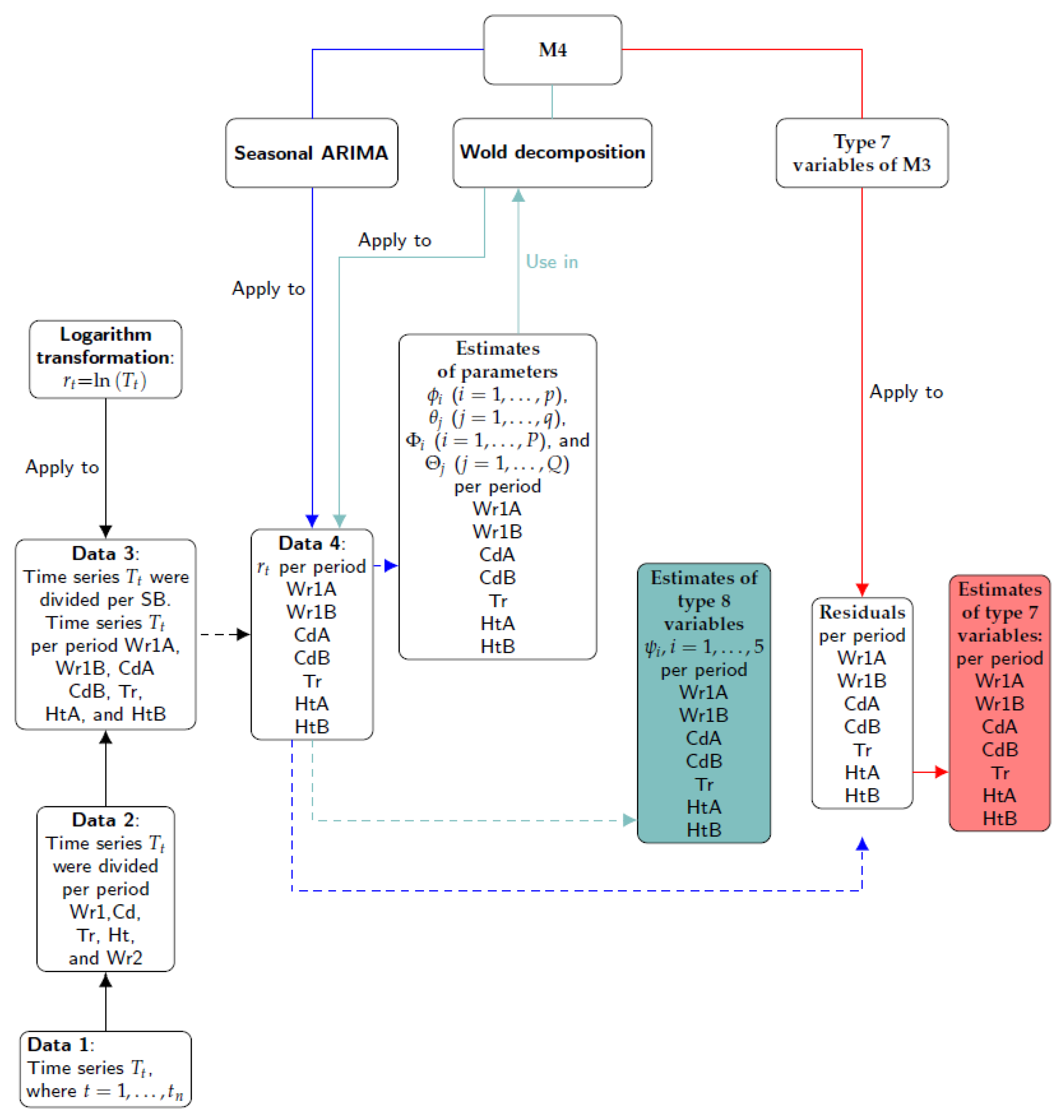
The steps involved in Method 3 are illustrated in
Figure 6. Firstly, the different time series were split according to the climatic stages observed (Data 2) and the structural breaks (SB) identified (Data 3). Secondly, the logarithm transformation was applied to Data 3 (the result is denoted as Data 4). The third step consisted of applying the seasonal ARIMA model to Data 4 in order to obtain the estimates of model coefficients:
,
,
and
. These parameters were denoted as Type 6 variables. Next, different features (Type 7 variables) were computed from the residuals: variance (
), maximum of periodogram (
) and its frequency
w. For the set of 72 lags, additional features were computed from sample ACF values: mean (
), median (
), variance (
) and range (
), with
. Finally, the first four values of sample PACF (
,
,
and
) were computed, as well as the statistics of the KS normality (Dn) and SW (Wn) tests.
In order to choose a seasonal ARIMA model and the estimations of the model parameters for each sensor, the
arima (
stats) and
auto.arima (
forecast) functions [
33,
34] were used. The ADF test was computed using the
adf.test (
aTSA) [
32]. The LBQ test was applied by means of the
Box.test function (
stats). The LM test was carried out using the
arch.test function (
aTSA). The SW and KS normality tests were applied using the
shapiro.test (
stats) and
ks.test functions (
dgof), respectively.
3.3.5. Calculation of Classification Variables—Method M4
The Wold decomposition establishes that any covariance stationary process can be written as the sum of a non-deterministic and deterministic process. This decomposition, which is unique, is a linear combination of lags of a WN process and a second process whose future values can be predicted exactly by some linear functions of past observations. If the time series
is purely non-deterministic, then it can be written as a linear combination of lagged values of a WN process (MA(
∞) representation), that is,
, where
,
and
is a WN [
41]. Although the Wold decomposition depends on an infinite number of parameters, the values of coefficients of the decomposition decay rapidly to zero.
For Method 3, a unique model was fitted for the 27 sensors in the same stage of the time series because it is necessary to have the same number of classification variables per sensor, in order to apply later the sPLS-DA method. It is not possible to work with ’the best model’ per sensor in each stage. In order to obtain the same number of variables using the ’best model’ per sensor, the Wold decomposition was applied to each sensor. Hence, Method 4 consists of obtaining the Wold decomposition for the ARMA models. Firstly, different seasonal ARIMA models were fitted iteratively to time series per sensor and stage of the time series, and the most successful model was determined.
As an illustration, consider that a time series follows a seasonal ARIMA process. Now, consider that and , with . Then, the time series follows an ARMA process, which can be decomposed according to the Wold approach by obtaining the polynomials and that determine the best ARMA model. In summary, for each seasonal ARIMA , it was possible to find the best ARMA model and its Wold decomposition.
The analysis of residuals of the different models fitted suggests that the condition of not autocorrelation is not fulfilled in all cases. Nonetheless, the Wold decomposition of each model was fitted independently, in order to have the ’best seasonal ARIMA model’ per sensor and the same number of parameters per sensor. For each model, the first five coefficients of the Wold decomposition were calculated and used as classification variables. In all cases, the most successful seasonal ARIMA model per sensor used and .
The steps involved in Method 4 are illustrated in
Figure 7. Firstly, the different time series were split according to the climatic stages observed (Data 2) and the structural breaks (SB) identified (Data 3). Secondly, the logarithm transformation was applied to Data 3 (the result is denoted as Data 4). The third step consisted of applying the seasonal ARIMA model to Data 4 in order to obtain the estimates of parameters and their residuals. Next, the same formulas of Type 7 variables used in M3 were applied to the residuals. Finally, the Wold decomposition was determined using the estimates of parameters of seasonal ARIMA models. The first five coefficients of the MA weights (i.e.,
) were denoted as Type 8 variables.
Apart from the same functions used for M3, this method employed ARMAtoMA (stats). One function was created for reducing a polynomial with AR and SAR components to a polynomial with just AR component. Likewise, another function converted a polynomial with MA and SMA components to one with MA component.
3.3.6. Determination of Number of Classes and Class per Sensor Using PCA and K-Means Algorithm
All classification variables calculated as described above for each sensor were arranged as a matrix called ’total classification dataset’ (TCD) with 27 rows (sensors) by 671 columns corresponding to the classification variables from the four methods. The total number of variables was 88, 296, 143 and 139 from M1, M2, M3 and M4, respectively. The multivariate analysis of the TCD matrix would allow the identification of different microclimates in the archaeological museum. It was checked that the statistical distribution of some classification variables was strongly skewed, which recommends to apply a data pretreatment prior to the multivariate analysis. For those variables with a strongly skewed distribution, different standard (simple) Box–Cox transformations [
62] were applied with the goal of finding a simple transformation leading to a normal distribution. In particular, the Box–Cox transformations were used on those classification variables with a Fisher coefficient of kurtosis [
63] or with a Fisher–Pearson standardized moment coefficient of skewness [
63] outside the interval from −2.0 to 2.0. Before applying the Box–Cox transformations, an absolute value function was used for variables with a negative skewness that fulfilled one of the aforementioned conditions. The skewness statistic was computed for each variable in order to check the asymmetry of the probability distribution. The kurtosis parameter indicates which variables were heavy-tailed or light-tailed, relative to a normal distribution. Moreover, the estimates of kurtosis were useful measures for identifying outliers in the classification variables. The functions
kurtosis and
skewness (
PerformanceAnalytics) [
38] were used to compute the coefficients of kurtosis and skewness, while
boxcoxfit (
geoR) [
64] was employed to apply different Box–Cox transformations. The function
prcomp (
stats) was used to carry out PCA.
Those values of a given classification variable that clearly departed from a straight line on the normal probability plot were removed and regarded as missing data. These were estimated using the NIPALS algorithm [
65] implemented in the
mixOmics package [
31], which is able to cope with this drawback and returns accurate results [
66]. After the data normalization, all variables were mean-centered and scaled to unit variance, which is the common pretreatment in PCA. Next, PCA was carried out to reduce the dimensionality of the TCD matrix. Each observation (sensor) was projected onto the first few principal components to obtain lower-dimensional data, while preserving as much of the data variation as possible.
Given that the two first components maximize the variance of the projected observations (TCD), only two components were employed to run the k-means clustering. This method is a centroid-based algorithm that computes the distance between each sensor and the different centroids, one per cluster or class. The algorithm determines
K clusters so that the intra-cluster variation is as small as possible. However, prior to applying this method, the number of clusters
K has to be determined, which depends on the type of clustering and previous knowledge about the time series of T. For this purpose, different criteria can be used [
39,
67]. Such methods do not always agree exactly in their estimation of the optimal number of clusters, but they tend to narrow the range of possible values. The
NbClust function of the
NbClust package [
39] incorporates 30 different indices for determining the number of clusters [
67]. This function claims to use the best clustering scheme from the different results obtained, by varying all combinations of the number of clusters, distance measures and clustering methods. It allows the user to identify the value
K in which more indices coincide, providing assurance that a good choice is being made.
In this clustering method, the measure used to determine the internal variance of each cluster was the sum of the squared Euclidean distances between each sensor and each centroid. The distances were used to assign each sensor to a cluster. For this purpose, the k-means algorithm of Hartigan and Wong [
68] was applied by means of the function
kmeans (
stats). It performs better than the algorithms proposed by MacQueen [
69], Lloyd [
70] and Forgy [
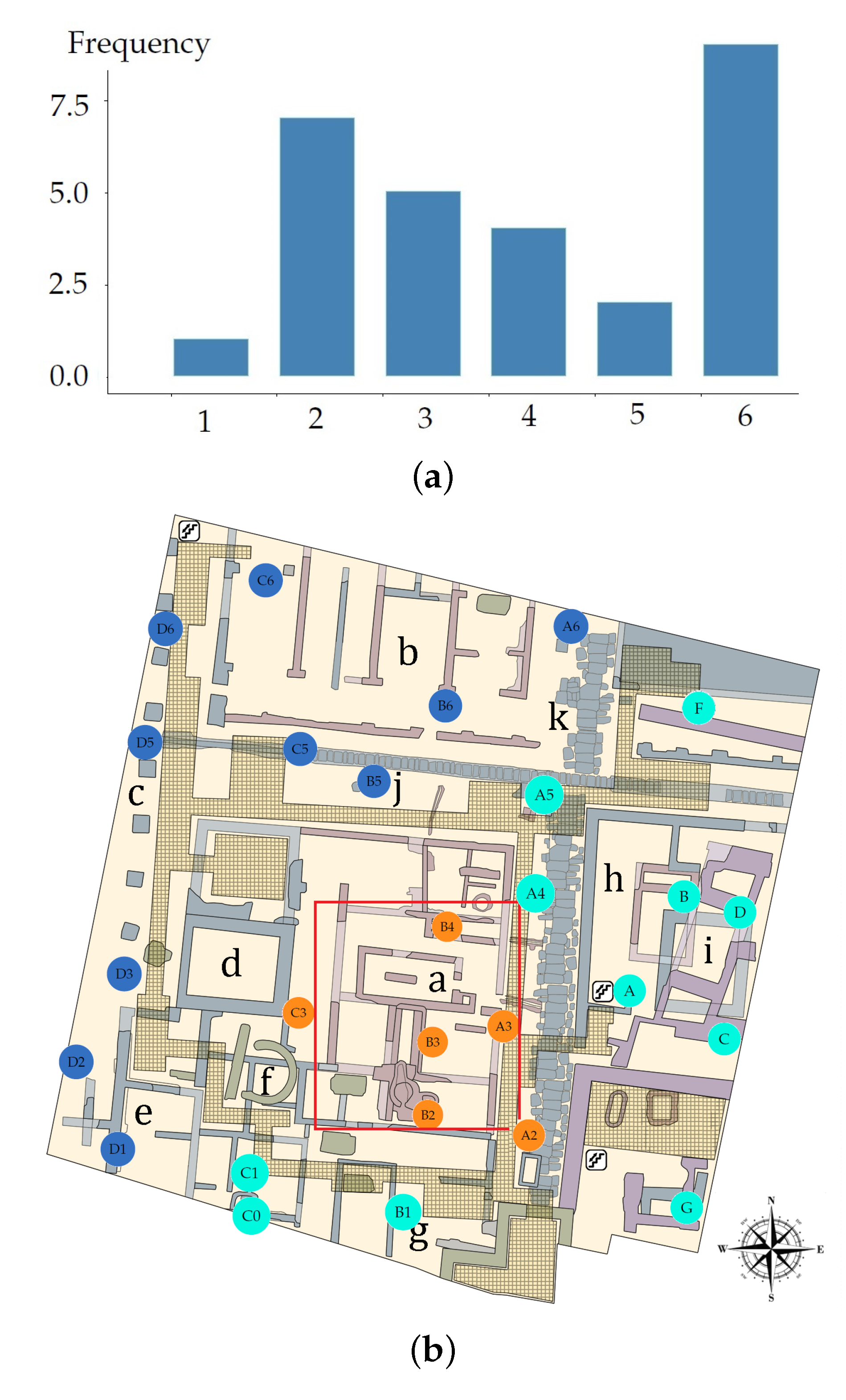
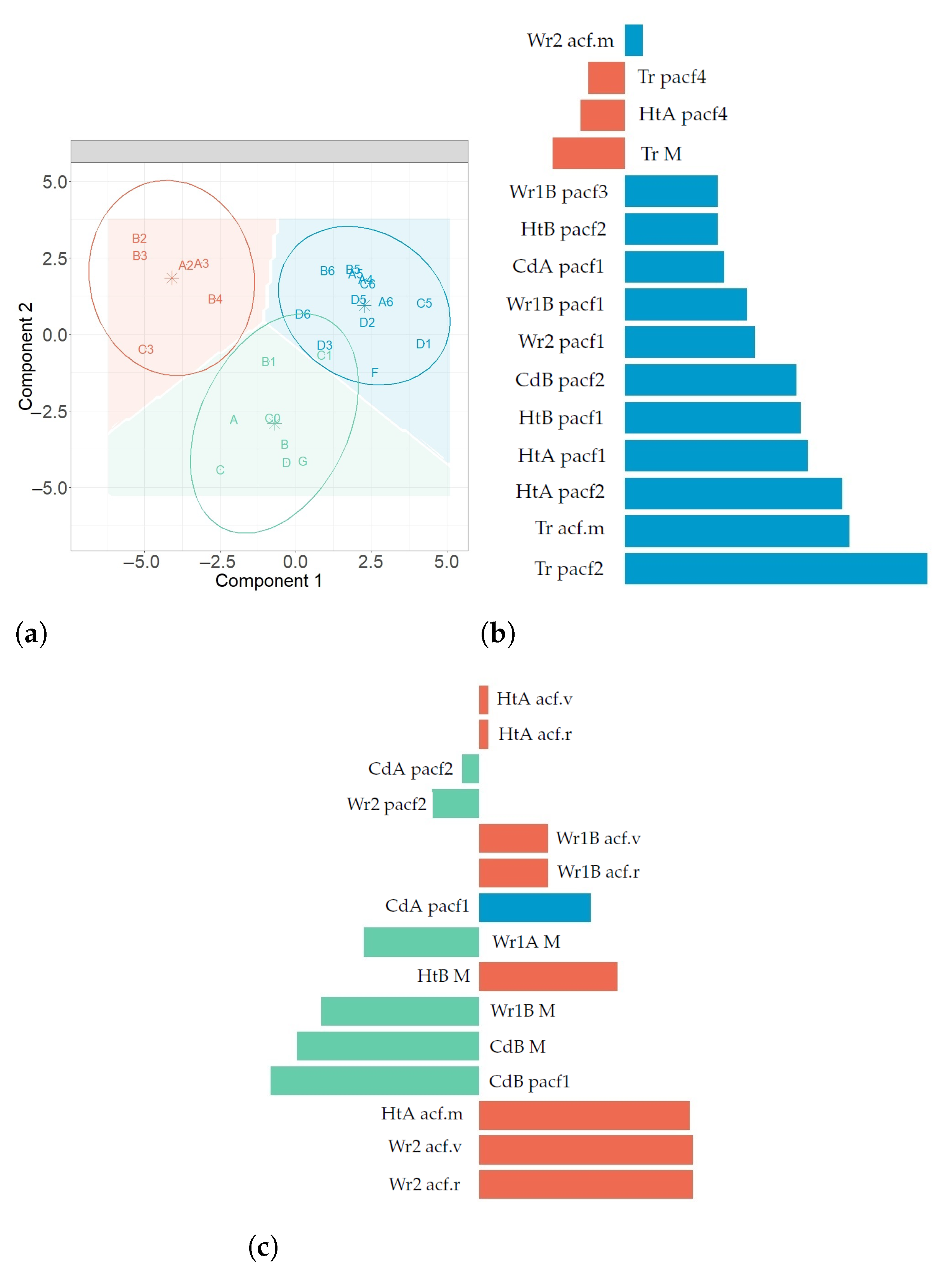
71]. However, when the algorithm of Hartigan and Wong is carried out, it is often recommended to try several random starts. In the present study, 100 random starts were employed. This algorithm guarantees that, at each step, the total intra-variance of the clusters is reduced until reaching a local optimum. Results from the k-means algorithm depend on the initial random assignment. For this reason, the algorithm was run 100 times, each with a different initial assignment. The final result was the one leading to a classification with the lowest total variance value. By comparing the classification obtained with the position of sensors in the museum (
Figure 8a), the three zones were denoted as North West (NW), South East (SE) and Skylight (Sk).
3.3.7. Sensor Classification Using sPLS-DA
Partial Least Squares (PLS) regression [
72] is a multivariate regression method which relates two data matrices (predictors and answer). PLS maximizes the covariance between latent components from these two datasets. A latent component is a linear combination of variables. The weight vectors used to calculate the linear combinations are called loading vectors.
Penalties such as Lasso and Ridge [
73] have been applied to the weight vectors in PLS for variable selection in order to improve the interpretability when dealing with a large number of variables [
74,
75,
76]. Chung and Keles [
77] extended the sparse PLS [
76] to classification problems (SPLSDA and SGPLS) and demonstrated that both SPLSDA and SGPLS improved classification accuracy compared to classical PLS [
78,
79,
80]. Le Cao et al. [
81] introduced a sparse version of the PLS algorithm for discrimination purposes (sPLS-Discriminant Analysis, sPLS-DA) which is an extension of the sPLS proposed by Lê Cao et al. [
74,
75]. They showed that sPLS-DA has very satisfying predictive performances and is able to select the most informative variables. Contrary to the two-stages approach (SPLSDA) proposed by Chung and Keles [
77], sPLS-DA performs variable selection and classification in a single-step procedure. In order to classify the sensors and improve the interpretability of results, sPLS-DA was applied to the different classification datasets.
Since the original PLS algorithm proposed by Wold [
72], many variants have arisen (e.g., PLS1, PLS2, PLS-A, PLS-SVD [
82] and SIMPLS [
83]), depending on how the regressor matrix (
) and response matrix (
) are deflated. Alternatives exist whether
and
are deflated separately or directly, using the cross product
and the Singular Value Decomposition (SVD). A hybrid PLS with SVD is used in the version sPLS-DA [
74]. For sPLS-DA, a regressor matrix is denoted as
(
,
,
or
in this case), with dimension
. The number of rows (sensors) is
and the columns correspond to the classification variables
p (
,
,
or
). A response qualitative vector denoted as
has length
n and it indicates the class of each sensor, with values coded as 1 (for NW), 2 (SE) and 3 (Sk).
sPLS-DA was carried out using Lasso penalization of the loading vectors associated to
[
84] using a hybrid PLS with SVD decomposition [
85]. The penalty function is included in the objective function of PLS-DA, which corresponds to PLS carried out using a response matrix
with values of either 0 or 1, created with the values of response vector
. Thus, this vector was converted into a dummy matrix
with dimension
, being
the number of sensors and
the number of sensor classes.
Regarding the optimization problem of sPLS-DA, in this case:
is a matrix with
p variables and
n sensors,
is a response vector (classes
) and
is an indicator matrix, where
, with
. The sPLS-DA method modeled
and
as
and
, where
and
are matrices that contain the regression coefficients of
and
on the
H latent components associated to
, while
and
are random errors. Furthermore, each component of
is a combination of selected variables, where
, and each vector
was computed sequentially as
, where
is the orthogonal projection of
on subspace
and
is the solution the optimization problem according to Equation (
3), subject to
.
The optimization problem minimizes the Frobenius norm between the current cross product matrix (
) and the loading vectors (
and
), where
and
is the orthogonal projection of
on subspace
. Furthermore,
, and
, defined as
, is the Lasso penalty function [
75,
81]. This optimization problem is solved iteratively based on the PLS algorithm [
86]. The SVD decomposition of matrix
is subsequently deflated for each iteration
h. This matrix is computed as
, where
and
are orthonormal matrices and
is a diagonal matrix whose diagonal elements are called the singular values. During the deflation step of PLS,
, because
and
are computed separately, and the new matrix is called
. At each step, a new matrix
is computed and decomposed by SVD [
74]. Furthermore, the soft-thresholding function
, with
, was used in penalizing loading vectors
to perform variable selection in regressor matrix, thus
[
74].
Rohart et al. [
30] implemented an algorithm to solve the optimization problem in Equation (
3), where the parameter
needs to be tuned and the algorithm chooses
among a finite set of values. It is possible to find a value
for any null elements in a loading vector. These authors implemented the algorithm using the number of non-zero elements of the loading vector as input, which corresponds to the number of selected variables for each component. They implemented sPLS-DA in the
R package
mixOmics, which provides functions such as
perf,
tune.splsda and
splsda, in order to determine the number of components and elements different to zero in the loading vector before running the final model.
In order to compare the performance of models constructed with a different number of components, 1000 training and test datasets were simulated and the sPLS-DA method (for a maximum number of 10 components) was tuned by three-fold cross-validation (CV) for
. The
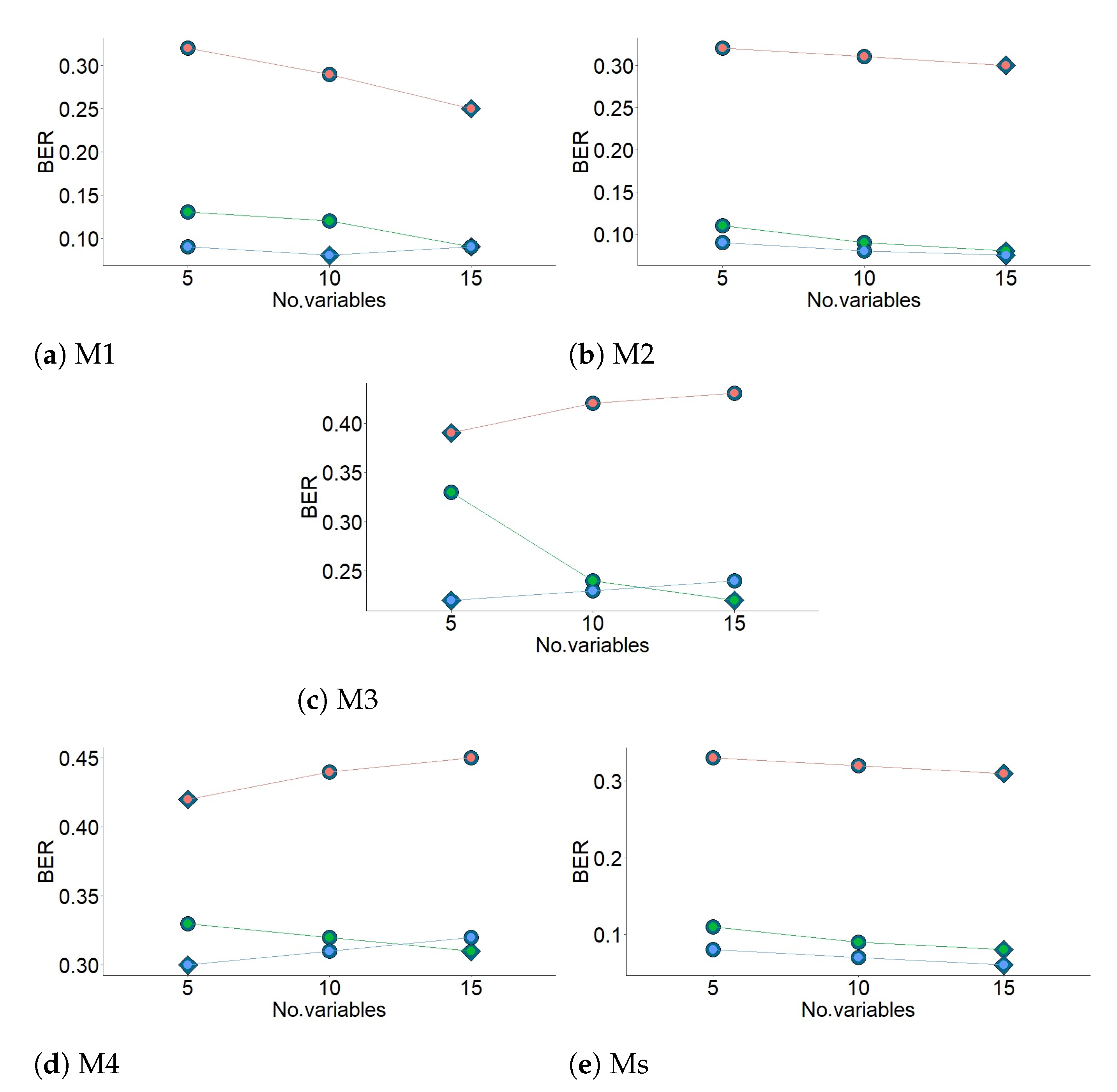
perf function outputs the optimal number of components that achieve the best performance based on both types of classification error rate (CER): Balanced Error Rate (BER) and the Overall classification error rate (Overall). BER is the average proportion of wrongly classified sensors in each class, weighted by the number of sensors. In most cases, the results from sPLS-DA were better (or very similar) using Overall than when using BER. However, BER was preferred to Overall because it is less biased towards majority classes during the performance assessment of sPLS-DA. In this step, three different prediction distances were used, maximum, centroid and Mahalanobis [
30], in order to determine the predicted class per sensor for each of the test datasets. For each prediction distance, both Overall and BER were computed.
The maximum number of components found in the present work was three (
) when BER was used instead of Overall. Furthermore, when sPLS-DA used classification variables from M2 and M3, two components led to the lowest BER values. Among the three prediction distances calculated (i.e., maximum, centroid and Mahalanobis), it was found that centroid performed better for the classification. Thus, this distance was used to determine the number of selected variables and to run the final model. Details of distances can be found in the Supplementary Materials of [
30].
In order to compare the performance of diverse models with different penalties, 1000 training and test datasets were simulated and the sPLS-DA method was carried out by three-fold CV for . The performance was measured via BER, and it was assessed for each value of a grid (keepX) from Component 1 to H, one component at a time. The different grids of values of the number of variables were carefully chosen to achieve a trade-off between resolution and computational time. Firstly, a two coarse tuning grids were assessed before setting a finer grid. The algorithm used the same grids of keepX argument in tune.splsda function to tune each component.
Once the optimal parameters were chosen (i.e., number of components and number of variables to select), the final sPLS-DA model was run on the whole dataset . The performance of this model in terms of BER was estimated using repeated CV.
In summary, three-fold CV was carried out for a maximum of ten components, using the three distances and both types of CER. The optimal number of components was obtained by using the BER and centroid distance. Next, the optimal number of variables was identified by carrying out the second three-fold CV. It was run using BER, centroid distance and values of three grids with different number of variables. Next, when both optimal numbers were obtained, the final model was computed.
Regarding the most relevant variables for explaining the classification of sensors, there are many different criteria [
86,
87,
88]. The first measure selected is the relative importance of each variable for each component and another is the accumulated importance of each variable from components. Both measures were employed in this research.
Lê Cao et al. [
81] applied sPLS-DA and selected only those variables with a non-zero weight. The sparse loading vectors are orthogonal to each other, which leads to uniquely selected variables across all dimensions. Hence, one variable might be influential in one component, but not in the other. Considering the previous argument and that the maximum number of components was three (
h = 1, 2, 3), Variable Importance in Projection (
) [
86] was used to select the most important ones. It is defined using loading vectors and the correlations of all response variables, for each component.
denotes the relative importance of variable
for component
h in the prediction model. Variables with
are the most relevant for explaining the classification of sensors.
was calculated using the
vip function (
mixOmics). Although the assumption of the sparse loading vectors being orthogonal was considered, in practice, some selected variables were common in two components. Then, a second measure of
[
88] was employed:
denotes the overall importance of variable
on all responses (one per class) cumulatively over all components. It is defined using the loading vectors and the sum of squares per component. Variables with
are the most relevant for explaining the classification of sensors. The selected variables were ranked according to both types of VIPs, which are discussed below for each stage of the time series.
3.3.8. Sensor Classification Using Random Forest Algorithm
The RF algorithm [
89] handles big datasets with high dimensionality. It consists of a large number of individual decision trees that were trained with a different sample of classification variables (
) generated by bootstrapping. The overall prediction from the algorithm was determined according to the predictions from individual decision trees. The class which receives most of the votes was selected as the prediction from each sensor. In addition, it can be used for identifying the most important variables.
An advantage of using the bootstrap resampling is that random forests have an Out-Of-Bag (OOB) sample that provides a reasonable approximation of the test error, which allows a built-in validation set that does not require an external data subset for validation. The following steps were carried out to obtain the prediction (output) from the algorithm, using the different classification data. Firstly, from a classification dataset, B random samples with replacements (bootstrap samples) were chosen, as well as one sample of 17 sensors. Next, from each bootstrap sample, a decision tree was grown. At each node, m variables out of total p were randomly selected without replacement. Each tree used an optimal number of variables that was determined by comparing the OOB classification error of a model, based on the number of predictors evaluated. Each node was divided using the variable that provided the best split according to the value of a variable importance measure (Gini index). Each tree grew to its optimal number of nodes. The optimal value of this hyper-parameter was obtained by comparing the OOB classification error of a model, based on the minimum size of the terminal nodes. From each bootstrap sample, a decision tree came up with a set of rules for classifying the sensors. Finally, the predicted class for each sensor was determined using those trees which excluded the sensor from its bootstrap sample. Each sensor was assigned to the class that received the majority of votes.
If the number of trees is high enough, the OOB classification error is roughly equivalent to the leave-one-out cross-validation error. Furthermore, RF does not produce overfitting problems when increasing the number of trees created in the process. According to previous arguments, 1500 trees were created for all cases to run the algorithm and the OOB classification error was used as an estimate of the test error. Prior to running the final RF algorithm, the optimal values of the number of predictors and the terminal nodes were determined (i.e., those corresponding to a stable OOB error).
In order to select the most important variables, the methodology proposed by Han et al. [
90] was used, which is based on two indices: Mean Decrease Accuracy (MDA) and Mean Decrease in Gini (MDG). The combined information is denoted as MDAMDG. Some reasons for using the methodology were: (1) The OOB error usually gives fair estimations compared to the usual alternative test set error, even if it is considered to be a little bit optimistic. (2) The use of both indices is more robust than considering any individual one [
90]. MDA corresponds to the average of the differences between OOB error before permuting the values of the variable and OOB error after permuting the values of the variable for all trees. Because a random forest is an ensemble of individual decision trees, the expected error rate called Gini impurity [
91] is used to calculate MDG. For classification, the node impurity is measured by the Gini index, and the MDG index is based on this. The former is the sum of a variable’s total decrease in node impurity, weighted by the proportion of samples reaching that node in each individual decision tree in the random forest [
92]. The higher is the MDG, the greater is the contribution of a given variable in the classification of sensors. The functions
randomForest and
importance [
93] were used to carried out the RF algorithm.