A Systematic Review of Machine Learning Techniques in Hematopoietic Stem Cell Transplantation (HSCT)
Abstract
1. Introduction
1.1. Background: Machine Learning (ML)
1.2. Applying ML Techniques in the Context of Hematopoietic Stem Cell Transplantation (HSCT)
2. Methods
2.1. Search Strategy
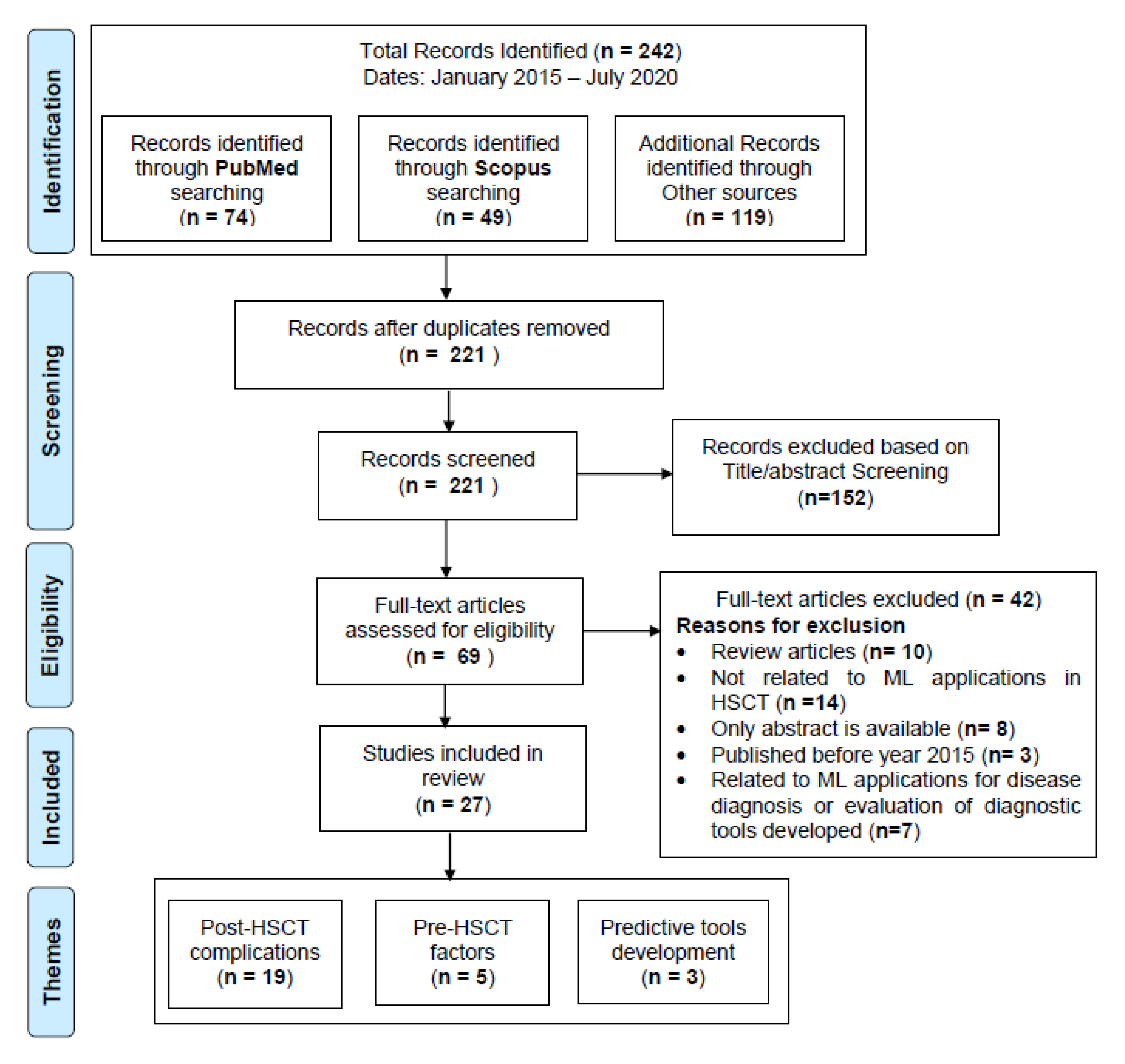
2.2. Study Selection
2.3. Data Extraction and Evaluation
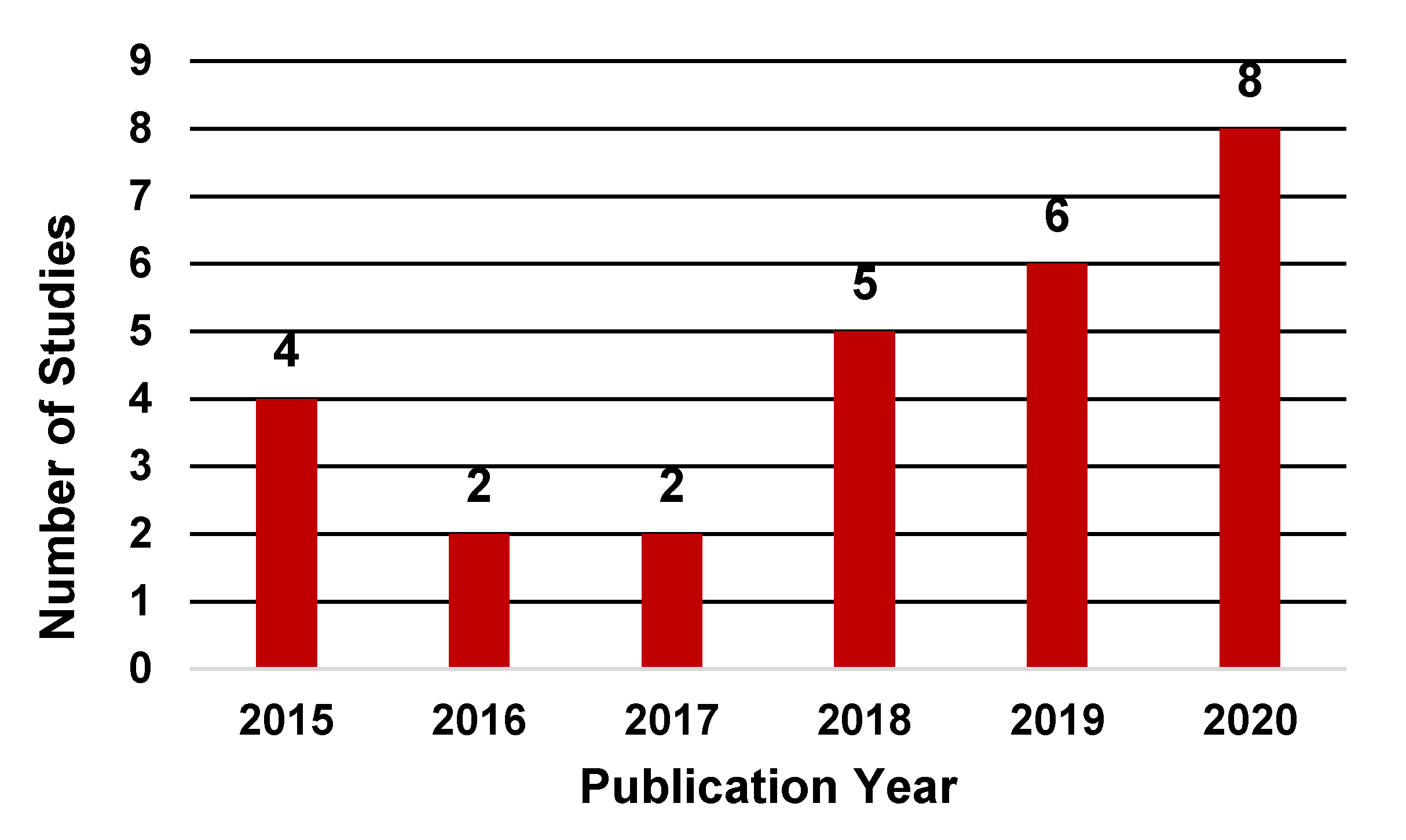
3. Results
3.1. Major Themes Identified
3.1.1. Post-HSCT Complications
3.1.2. Pre-transplant Factors
3.1.3. Predictive Tools Development
4. Discussion
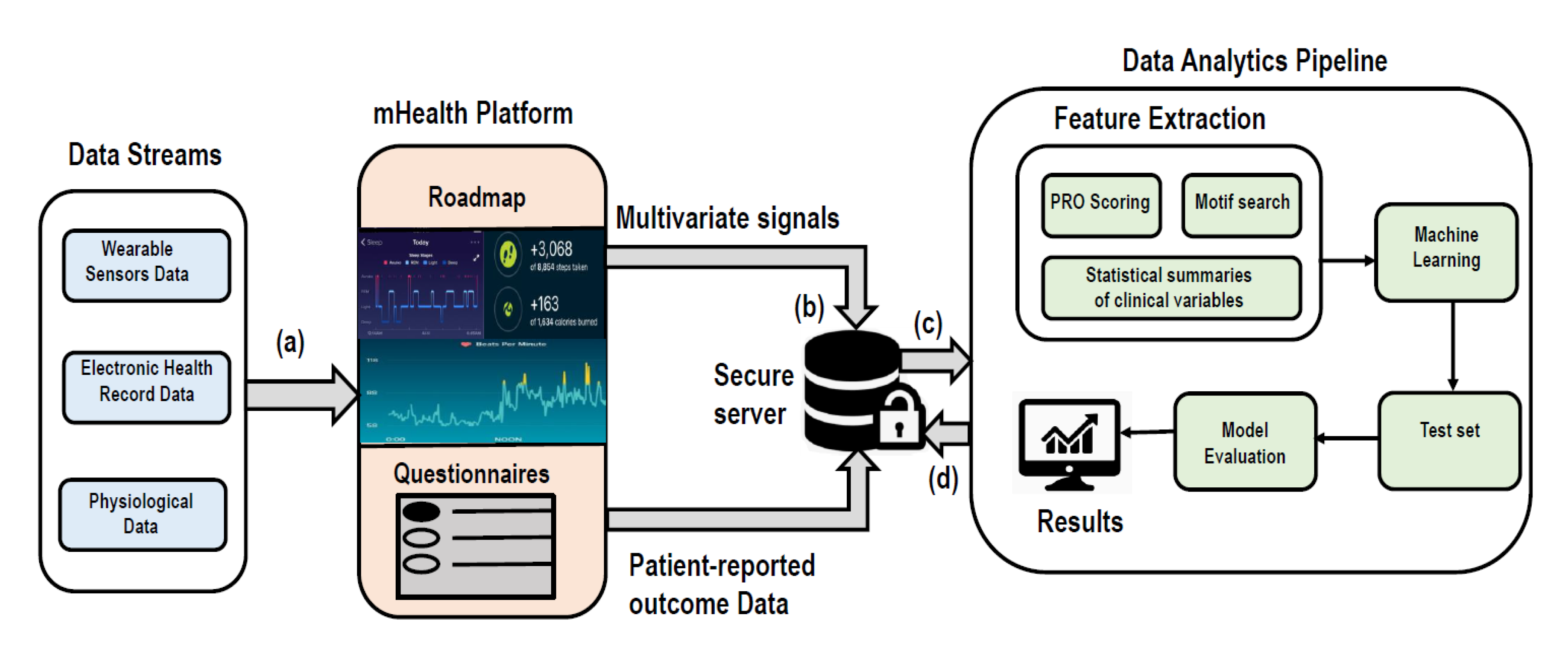
5. Case Study: Roadmap 2.0
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ML | Machine Learning |
| AI | Artificial intelligence |
| GVHD | Graft-versus-host-disease |
| ATS | Adaptive treatment strategies |
| HSCT | Hematopoietic stem cell transplantation |
| HLA | human leukocyte antigen |
| SVM | Support vector Machines |
| EHR | Electronic health record |
| mHealth | Mobile health |
| PRISMA | Preferred Reporting Items for Systematic Reviews and Meta-Analyses. |
| LR | Logistic Regression |
| RF | Random Forest |
| PCA | Principal Component Analysis |
| BLSTM | Bidirectional Long-short-term-memory |
| ADT | Alternating Decision Tree |
| NB | Naïve Bayes |
| MLP | Multiplayer perceptron |
| RL | Reinforcement Learning |
| CART | Classification and Regression Trees |
| BRT | Boosted Regression Trees |
| SR | Spline regression |
| BART | Bayesian additive regression trees |
| NN | Neural networks |
| k-NN | k-nearest neighbor |
| LDA | Linear Discriminant analysis |
| SDA | Shrinkage Discriminant analysis |
| RSF | Random survival Forest |
| BN | Bayesian Network |
| GBM | Gradient Boosting Machines |
| DT | Decision Tree |
| BL | Bayesian Learners |
| EL | Ensemble learners |
| AML | Acute Myeloid Leukemia |
| CBC | Complete blood count |
| WBC | White blood clount |
| rDRI | refined disease risk index |
| MM | multiple myeloma |
| NRM | Non-relapse mortality |
| SOM | Self-organizing map |
| GVHD-DE | Graft-versus-host-disease- dry eye |
| VT | Verification Typing |
| BDT | Boosted decision Trees |
| AUC | Area under curve |
| GPS | Generalized path seeker |
| API | Application programming Interface |
| PRO | Patient reported outcome |
| HRQOL | Health related quality of life |
| HIPPA | Health Insurance Portability and Accountability Act |
References
- Mitchell, T. Machine Learning, 1st ed.; McGraw Hill: New York, NY, USA, 1997. [Google Scholar]
- Mitchell, T. Machine learning and data mining. Commun. ACM 1999, 42, 30–36. [Google Scholar] [CrossRef]
- Beam, A.L.; Kohane, I.S. Big Data and Machine Learning in Health Care. JAMA 2018, 319, 1317–1318. [Google Scholar] [CrossRef] [PubMed]
- Rajkomar, A.; Dean, J.; Kohane, I. Machine Learning in Medicine. N. Engl. J. Med. 2019, 380, 1347–1358. [Google Scholar] [CrossRef]
- Ciganović, I.; Pluškoski, A.; Jovanović, M.D. Autonomous car driving-one possible implementation using machine learning algorithm. In Proceedings of the 5th International Conference on Electrical, Electronic and Computing Engineering, Palić, Serbia, 11–14 June 2018; Volume 1, pp. 1016–1021. [Google Scholar]
- Gulshan, V.; Peng, L.; Coram, M.; Stumpe, M.C.; Wu, D.; Narayanaswamy, A.; Venugopalan, S.; Widner, K.; Madams, T.; Cuadros, J.; et al. Development and Validation of a Deep Learning Algorithm for Detection of Diabetic Retinopathy in Retinal Fundus Photographs. JAMA 2016, 316, 2402–2410. [Google Scholar] [CrossRef]
- Esteva, A.; Kuprel, B.; Novoa, R.A.; Ko, J.; Swetter, S.M.; Blau, H.M.; Thrun, S. Dermatologist-level classification of skin cancer with deep neural networks. Nat. Cell Biol. 2017, 542, 115–118. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Schuster, M.; Chen, Z.; Le, Q.V.; Norouzi, M.; Macherey, W.; Klingner, J. Google’s neural machine translation system: Bridging the gap between human and machine translation. arXiv 2016, arXiv:1609.08144. [Google Scholar]
- Taigman, Y.; Yang, M.; Ranzato, M.; Wolf, L. DeepFace: Closing the Gap to Human-Level Performance in Face Verification. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Columbus, OH, USA, 23–28 June 2014; pp. 1701–1708. [Google Scholar]
- Cirillo, D.; Valencia, A. Big data analytics for personalized medicine. Curr. Opin. Biotechnol. 2019, 58, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.-Y.; Cheng, C.-W.; Kaddi, C.D.; Venugopalan, J.; Hoffman, R.; Wang, M.D. Omic and Electronic Health Record Big Data Analytics for Precision Medicine. IEEE Trans. Biomed. Eng. 2016, 64, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Appelbaum, F.R. Haematopoietic cell transplantation as immunotherapy. Nat. Cell Biol. 2001, 411, 385–389. [Google Scholar] [CrossRef]
- Copelan, E.A. Hematopoietic stem-cell transplantation. N. Engl. J. Med. 2006, 354, 1813–1826. [Google Scholar] [CrossRef] [PubMed]
- Muhsen, I.N.; Jagasia, M.; Toor, A.A.; Hashmi, S. Registries and artificial intelligence: Investing in the future of hematopoietic cell transplantation. Bone Marrow Transplant. 2018, 54, 477–480. [Google Scholar] [CrossRef] [PubMed]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; Group, T.P. Preferred Reporting Items for Systematic Reviews and Meta-analysis (PRISMA). Available online: http://prisma-statement.org/prismastatement/flowdiagram.aspx (accessed on 10 September 2020).
- Lu, C.C.; Li, J.L.; Wang, Y.F.; Ko, B.S.; Tang, J.L.; Lee, C.C. A BLSTM with Attention Network for Predicting Acute Myeloid Leukemia Patient’s Prognosis using Comprehensive Clinical Parameters. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; pp. 2455–2458. [Google Scholar]
- Fuse, K.; Uemura, S.; Tamura, S.; Suwabe, T.; Katagiri, T.; Tanaka, T.; Kuroha, T. Patient-based prediction algorithm of relapse after allo-HSCT for acute Leukemia and its usefulness in the decision-making process using a machine learning approach. Cancer Med. 2019, 8, 5058–5067. [Google Scholar] [CrossRef] [PubMed]
- Goswami, C.; Poonia, S.; Kumar, L.; Sengupta, D. Staging System to Predict the Risk of Relapse in Multiple Myeloma Patients Undergoing Autologous Stem Cell Transplantation. Front. Oncol. 2019, 9, 633. [Google Scholar] [CrossRef] [PubMed]
- Ritari, J.; Hyvärinen, K.; Koskela, S.; Itälä-Remes, M.; Niittyvuopio, R.; Nihtinen, A.; Salmenniemi, U.; Putkonen, M.; Volin, L.; Kwan, T.; et al. Genomic prediction of relapse in recipients of allogeneic haematopoietic stem cell transplantation. Leukemia 2018, 33, 240–248. [Google Scholar] [CrossRef]
- Marino, S.R.; Lee, S.M.; Binkowski, T.A.; Wang, T.; Haagenson, M.; Wang, H.-L.; Maiers, M.; Spellman, S.; Van Besien, K.; Lee, S.J.; et al. Identification of high-risk amino-acid substitutions in hematopoietic cell transplantation: A challenging task. Bone Marrow Transplant. 2016, 51, 1342–1349. [Google Scholar] [CrossRef]
- ArabYarmohammadi, S.; Zhang, Z.; Leo, P.; Firouznia, M.; Janowczyk, A.; Li, H.; Xu, J. Computationally derived cytological image markers for predicting risk of relapse in acute myeloid leukemia patients following bone marrow transplantation. In Proceedings of the Medical Imaging 2020: Digital Pathology, International Society for Optics and Photonics, Houston, TX, USA, 15–20 February 2020; p. 1132004. [Google Scholar]
- Krakow, E.F.; Hemmer, M.; Wang, T.; Logan, B.; Arora, M.; Spellman, S.; Couriel, D.; Alousi, A.; Pidala, J.; Last, M.; et al. Tools for the Precision Medicine Era: How to Develop Highly Personalized Treatment Recommendations From Cohort and Registry Data Using Q-Learning. Am. J. Epidemiol. 2017, 186, 160–172. [Google Scholar] [CrossRef]
- Liu, Y.; Logan, B.; Liu, N.; Xu, Z.; Tang, J.; Wang, Y. Deep Reinforcement Learning for Dynamic Treatment Regimes on Medical Registry Data. In Proceedings of the 2017 IEEE International Conference on Healthcare Informatics (ICHI), Park City, UT, USA, 23–26 August 2017; Volume 2017, pp. 380–385. [Google Scholar]
- Shouval, R.; Labopin, M.; Unger, R.; Giebel, S.; Ciceri, F.; Schmid, C.; Esteve, J.; Baron, F.; Gorin, N.C.; Savani, B.; et al. Prediction of Hematopoietic Stem Cell Transplantation Related Mortality- Lessons Learned from the In-Silico Approach: A European Society for Blood and Marrow Transplantation Acute Leukemia Working Party Data Mining Study. PLoS ONE 2016, 11, e0150637. [Google Scholar] [CrossRef]
- Shouval, R.; Labopin, M.; Bondi, O.; Mishan-Shamay, H.; Shimoni, A.; Ciceri, F.; Esteve, J.; Giebel, S.; Gorin, N.C.; Schmid, C.; et al. Prediction of Allogeneic Hematopoietic Stem-Cell Transplantation Mortality 100 Days After Transplantation Using a Machine Learning Algorithm: A European Group for Blood and Marrow Transplantation Acute Leukemia Working Party Retrospective Data Mining Study. J. Clin. Oncol. 2015, 33, 3144–3151. [Google Scholar] [CrossRef]
- Tang, S.; Chappell, G.T.; Mazzoli, A.; Tewari, M.; Choi, S.W.; Wiens, J. Predicting Acute Graft-Versus-Host Disease Using Machine Learning and Longitudinal Vital Sign Data From Electronic Health Records. JCO Clin. Cancer Inform. 2020, 4, 128–135. [Google Scholar] [CrossRef]
- Arai, Y.; Kondo, T.; Fuse, K.; Shibasaki, Y.; Masuko, M.; Sugita, J.; Teshima, T.; Uchida, N.; Fukuda, T.; Kakihana, K.; et al. Using a machine learning algorithm to predict acute graft-versus-host disease following allogeneic transplantation. Blood Adv. 2019, 3, 3626–3634. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Wu, Z.; Fujiwara, H.; Whitesall, S.; Zajac, C.K.; Choi, S.W.; Reddy, P.; Tewari, M. Computational analysis of continuous body temperature provides early discrimination of graft-versus-host disease in mice. Blood Adv. 2019, 3, 3977–3981. [Google Scholar] [CrossRef] [PubMed]
- Serrano-López, J.; Fernández, J.L.; Lumbreras, E.; Serrano, J.; Martínez-Losada, C.; Martín, C.; Hernández-Rivas, J.M.; Sánchez-García, J. Machine learning applied to gene expression analysis of T-lymphocytes in patients with cGVHD. Bone Marrow Transplant. 2020, 55, 1668–1670. [Google Scholar] [CrossRef]
- Sharifi, H.; Lai, Y.K.; Guo, H.; Hoppenfeld, M.; Guenther, Z.D.; Johnston, L.; Brondstetter, T.; Chhatwani, L.; Nicolls, M.R.; Hsu, J.L. Machine Learning Algorithms to Differentiate Among Pulmonary Complications After Hematopoietic Cell Transplant. Chest 2020. [Google Scholar] [CrossRef]
- Gandelman, J.S.; Byrne, M.; Mistry, A.M.; Polikowsky, H.G.; Diggins, K.E.; Chen, H.; Lee, S.J.; Arora, M.; Cutler, C.; Flowers, M.; et al. Machine learning reveals chronic graft-versus-host disease phenotypes and stratifies survival after stem cell transplant for hematologic malignancies. Haematologica 2018, 104, 189–196. [Google Scholar] [CrossRef]
- Sharafeldin, N.; Richman, J.; Bosworth, A.; Chen, Y.; Singh, P.; Patel, S.K.; Wang, X.; Francisco, L.; Forman, S.J.; Wong, F.L.; et al. Clinical and Genetic Risk Prediction of Cognitive Impairment After Blood or Marrow Transplantation for Hematologic Malignancy. J. Clin. Oncol. 2020, 38, 1312–1321. [Google Scholar] [CrossRef]
- Cocho, L.; Fernández, I.; Calonge, M.; Martínez, V.; González-García, M.J.; Caballero, L.; López-Corral, L.; García-Vázquez, C.; Vazquez, L.; Stern, M.E.; et al. Gene Expression–Based Predictive Models of Graft Versus Host Disease–Associated Dry Eye. Investig. Opthalmol. Vis. Sci. 2015, 56, 4570. [Google Scholar] [CrossRef] [PubMed]
- Leclerc, V.; Ducher, M.; Bleyzac, N. Bayesian Networks: A New Approach to Predict Therapeutic Range Achievement of Initial Cyclosporine Blood Concentration After Pediatric Hematopoietic Stem Cell Transplantation. Drugs R D 2018, 18, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Masiliune, A.; Winstone, D.; Gasieniec, L.; Wong, P.; Lin, H.; Pawson, R.; Parkes, G.; Hadley, A. Predicting the Availability of Hematopoietic Stem Cell Donors Using Machine Learning. Biol. Blood Marrow Transplant. 2020, 26, 1406–1413. [Google Scholar] [CrossRef]
- Sivasankaran, A.; Williams, E.; Albrecht, M.; Switzer, G.E.; Cherkassky, V.; Maiers, M. Machine Learning Approach to Predicting Stem Cell Donor Availability. Biol. Blood Marrow Transplant. 2018, 24, 2425–2432. [Google Scholar] [CrossRef]
- Buturovic, L.; Shelton, J.; Spellman, S.R.; Wang, T.; Friedman, L.; Loftus, D.; Hesterberg, L.; Woodring, T.; Fleischhauer, K.; Hsu, K.C.; et al. Evaluation of a Machine Learning-Based Prognostic Model for Unrelated Hematopoietic Cell Transplantation Donor Selection. Biol. Blood Marrow Transplant. 2018, 24, 1299–1306. [Google Scholar] [CrossRef]
- Sivasankaran, A.; Cherkassky, V.; Albrecht, M.; Williams, E.; Maiers, M. Donor Selection for Hematopoietic Stem Cell Transplant Using Cost-Sensitive SVM. In Proceedings of the 2015 IEEE 14th International Conference on Machine Learning and Applications (ICMLA), Miami, FL, USA, 9–11 December 2015; pp. 831–836. [Google Scholar]
- Brasier, A.R.; Zhao, Y.; Spratt, H.M.; Wiktorowicz, J.E.; Ju, H.; Wheat, L.J.; Baden, L.; Stafford, S.; Wu, Z.; Issa, N.; et al. Improved Detection of Invasive Pulmonary Aspergillosis Arising during Leukemia Treatment Using a Panel of Host Response Proteins and Fungal Antigens. PLoS ONE 2015, 10, e0143165. [Google Scholar] [CrossRef]
- Lee, C.; Haneuse, S.; Wang, H.-L.; Rose, S.; Spellman, S.R.; Verneris, M.; Hsu, K.C.; Fleischhauer, K.; Lee, S.J.; Abdi, R. Prediction of absolute risk of acute graft-versus-host disease following hematopoietic cell transplantation. PLoS ONE 2018, 13, e0190610. [Google Scholar] [CrossRef] [PubMed]
- Okamura, H.; Nakamae, M.; Koh, S.; Nanno, S.; Nakashima, Y.; Koh, H.; Nakane, T.; Hirose, A.; Hino, M.; Nakamae, H. Interactive web application for plotting personalized prognosis prediction curves in allogeneic hematopoietic cell transplantation using machine learning. medRxiv 2019. [Google Scholar] [CrossRef]
- Leclerc, V.; Bleyzac, N.; Ceraulo, A.; Bertrand, Y.; Ducher, M. A decision support tool to find the best cyclosporine dose when switching from intravenous to oral route in pediatric stem cell transplant patients. Eur. J. Clin. Pharmacol. 2020, 76, 1409–1416. [Google Scholar] [CrossRef] [PubMed]
- Armand, P.; Kim, H.T.; Logan, B.R.; Wang, Z.; Alyea, E.P.; Kalaycio, M.E.; Rizzo, J.D. Validation and refinement of the Disease Risk Index for allogeneic stem cell transplantation. Blood 2014, 123, 3664–3671. [Google Scholar] [CrossRef] [PubMed]
- Ng, A.Y.; Jordan, M.I.; Weiss, Y. On Spectral Clustering: Analysis and an Algorithm. In Proceedings of the Advances in Neural Information Processing Systems, Vancouver, BC, Canada, 9–14 December 2002; pp. 849–856. [Google Scholar]
- Gigerenzer, G.; Brighton, G.G.H.; Heuristicus, H. RT list: Message to Robyn: Simple heuristics that make us smart. Behav. Brain Sci. 2000, 23, 727–780. [Google Scholar]
- Gragert, L.; Eapen, M.; Williams, E.; Freeman, J.; Spellman, S.; Baitty, R.; Hartzman, R.; Rizzo, J.D.; Horowitz, M.; Confer, D.; et al. HLA Match Likelihoods for Hematopoietic Stem-Cell Grafts in the U.S. Registry. N. Engl. J. Med. 2014, 371, 339–348. [Google Scholar] [CrossRef]
- Xu, J.; Wang, F. Federated Learning for Healthcare Informatics. arXiv 2019, arXiv:1911.06270. [Google Scholar]
- Lundberg, S.; Lee, S.-I. A unified approach to interpreting model predictions. In Proceedings of the Advances in Neural Information Processing Systems, Long Beach, CA, USA, 4–9 December 2017; pp. 4765–4774. [Google Scholar]
- Ribeiro, M.; Singh, S.; Guestrin, C. “Why Should I Trust You?”: Explaining the Predictions of Any Classifier. In Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining, San Francisco, CA, USA, 13–17 August 2016; pp. 1135–1144. [Google Scholar]
- Rozwadowski, M.; Dittakavi, M.; Mazzoli, A.; Hassett, A.L.; Braun, T.; Barton, D.L.; Carlozzi, N.; Tewari, M.; Hanauer, D.A.; Choi, S.W. Promoting Health and Well-Being through Mobile Health Technology (Roadmap 2.0) in Family Caregivers and Patients Undergoing Hematopoietic Stem Cell Transplantation: Protocol Development of a Mobile Randomized Trial (Preprint). JMIR Res. Protoc. 2020, 9, 19288. [Google Scholar] [CrossRef]
- Kaplan, E.L.; Meier, P. Nonparametric-Estimation from Incomplete Observations. J. Am. Stat. Assoc. 1958, 53, 457–481. [Google Scholar] [CrossRef]
- Fine, J.P.; Gray, R.J. A proportional hazards model for the subdistribution of a competing risk. J. Am. Stat. Assoc. 1999, 94, 496–509. [Google Scholar] [CrossRef]
- Domingos, P.; Hulten, G. Mining high-speed data streams. In Proceedings of the Sixth ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Boston, MA, USA, 20–23 August 2000; pp. 71–80. [Google Scholar]
- Manogaran, G.; Lopez, D. Health data analytics using scalable logistic regression with stochastic gradient descent. Int. J. Adv. Intell. Paradig. 2018, 10, 118–132. [Google Scholar] [CrossRef]
- Oza, N.C. Online Bagging and Boosting. In Proceedings of the 2005 IEEE International Conference on Systems, Man and Cybernetics, Waikoloa, HI, USA, 10–12 October 2006; pp. 2340–2345. [Google Scholar]
- Zaharia, M.; Chowdhury, M.; Franklin, M.J.; Shenker, S.; Stoica, I. Spark: Cluster computing with working sets. HotCloud 2010, 10, 95. [Google Scholar]
- Toshniwal, A.; Donham, J.; Bhagat, N.; Mittal, S.; Ryaboy, D.; Taneja, S.; Shukla, A.; Ramasamy, K.; Patel, J.M.; Kulkarni, S.; et al. Storm@twitter. In Proceedings of the 2014 ACM SIGMOD International Conference on Management of Data, Snowbird, UT, USA, 22–27 June 2014; pp. 147–156. [Google Scholar]
- Carbone, P.; Katsifodimos, A.; Ewen, S.; Markl, V.; Haridi, S.; Tzoumas, K. Apache flink: Stream and batch processing in a single engine. Bull. IEEE Comput. Soc. Tech. Comm. Data Eng. 2015, 38, 28–38. [Google Scholar]
| (HSCT OR HCT OR GVHD OR acute GVHD OR aGVHD OR leukemia OR lymphoma OR autologous HCT OR allogeneic HCT OR Hematopoietic Cell Transplantation OR Bone marrow transplant OR Hematopoietic cell transplant OR Hematopoietic stem cell transplantation OR Graft-versus-host disease) AND (Machine Learning OR Artificial Intelligence). |
| Reference | No. of Participants | Data Streams Used | Outcomes | Best ML Technique | Compared ML Techniques | Major Theme Identified |
|---|---|---|---|---|---|---|
| Lu et al., 2019 [16] | 637 | Clinical, genomic & demographics | AML 2-years survival and relapse, mortality | Att-BLSTM | SVM, LR | Post-HSCT complications |
| Fuse et al., 2019 [17] | 217 | Clinical | Risk of Leukemia relapse after 1 year of allo-HSCT | - | ADT | Post-HSCT complications |
| Goswami et al., 2019 [18] | 347 | Clinical | Relapse risk within 36 months of autologous-HSCT | - | Stacked ML | Post-HSCT complications |
| Ritari et al., 2018 [19] | 161 | Clinical & genomic | Genomic biomarkers for relapse risk of various hematological malignancies for allo-HSCT recipient | - | RF | Post-HSCT complications |
| Marino et al., 2016 [20] | 2107 | Clinical | High-risk amino acid substitutions and position types for grade III-IV acute-GVHD, TRM, disease free survival | - | RF, LR | Post-HSCT complications |
| ArabYarmohammadi et al., 2020 [21] | 39 | Images | Relapse risk in AML patients post-HSCT | - | Deep learning, LDA | Post-HSCT complications |
| Krakow et al., 2017 [22] | 9563 | Clinical | Adaptive treatment strategies | - | RL | Post-HSCT complications |
| Liu et al., 2017 [23] | 6021 | Clinical | Optimal Dynamic treatment regimes | - | Deep RL | Post-HSCT complications |
| Shouval et al., 2016 [24] | 26,266 | Clinical | NRM 100 days post HCT in acute leukemia | - | NB, ADT, LR, MLP, RF, AdaBoost | Post-HSCT complications |
| Shouval et al., 2015 [25] | 28,236 | Clinical | Overall Mortality 100 days post-HSCT | - | ADT | Post-HSCT complications |
| Tang et al., 2020 [26] | 324 | Clinical | Grade II-IV acute-GVHD risk | - | L2 regularized LR | Post-HSCT complications |
| Arai et al., 2019 [27] | 26,695 | Clinical | grade II-IV & III-IV aGVHD risk | ADT | NB, MLP, RF, Ada- boost | Post-HSCT complications |
| Kuang et al., 2019 [28] | 28 | Clinical & sensor | Non-invasive biomarkers for acute-GVHD diagnosis in mice | - | PCA, k-means | Post-HSCT complications |
| Serrano-López et al., 2020 [29] | 29 | Genomic | Gene biomarkers for chronic-GVHD diagnosis | - | RF | Post-HSCT complications |
| Sharifi et al., 2020 [30] | 66 | Images | Differentiate among pulmonary complications post-HSCT | - | k-means + SVM | Post-HSCT complications |
| Gandelman et al., 2019 [31] | 339 | Clinical | Classify patients with chronic-GVHD according to organ scores | - | k-means | Post-HSCT complications |
| Sharafeldin et al., 2020 [32] | 277 | Clinical, genomic & demographics | post-BMT cognitive impairment | - | ENR | Post-HSCT complications |
| Cocho et al., 2015 [33] | 36 | Clinical & genomic | Genomic biomarkers for GVHD associated Dry eye | SVM | k-NN, SDA | Post-HSCT complications |
| Leclerc et al., 2018 [34] | 155 | Clinical & biological | initial cyclosporine dose blood concentrations Post-HSCT | BN | NB, SVM, RF | Others |
| Li et al., 2020 [35] | 10,258 | Clinical & Demographics | Donor availability | BDT | LR, SVM | Pre-HSCT factors |
| Sivasankaran et al., 2018 [36] | Not clear | Demographics & member related factors | Donor availability | GBM | SVM, LR | Pre-HSCT factors |
| Buturovic et al., 2018 [37] | 1255 | Clinical | Selecting appropriate unrelated donor for patients undergoing HSCT | - | SVM | Pre-HSCT factors |
| Sivasankaran et al., 2015 [38] | 3035 | Clinical | Selecting appropriate unrelated donor for patients undergoing HSCT | SVM | k-NN, CART | Pre-HSCT factors |
| Brasier et al., 2015 [39] | 68 | Clinical | Detection of pre-HSCT infection in patients undergoing chemotherapy | GPS | RF, CART, MARS | Post-HSCT complications |
| Lee et al., 2018 [40] | 9651 | Clinical | Grade II-IV agvhd risk or death within 100 days post-HSCT | SL | LR, BRT, MARS, BART, RR, ENR, ANN | Predictive Tools Development |
| Okamura, et al. 2020 [41] | 363 | Clinical | 1-year overall survival, PFS, relapse, and NRM | - | RSF | Predictive Tools Development |
| Leclerc et al., 2020 [42] | 211 | Clinical & biological | Best first cyclosporine dose | - | BN | Predictive Tools Development |
| Challenges | Reasons | Potential Solution |
|---|---|---|
| Limited Data Capture |
|
|
| Data Quality Issues |
|
|
| High Dimensional Data |
|
|
| Data Privacy Issues |
|
|
| Obsolete Predictive Models |
|
|
| Diverse Data Types |
|
|
| Data Integration issues |
|
|
| Limitations | Consequences | Potential Solution |
|---|---|---|
| Lack of interpretable predictive models |
|
|
| Lack of model validation |
|
|
| Smaller sample size |
|
|
| Lack of multi-center studies |
|
|
| Lack of diverse data streams used |
|
|
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gupta, V.; Braun, T.M.; Chowdhury, M.; Tewari, M.; Choi, S.W. A Systematic Review of Machine Learning Techniques in Hematopoietic Stem Cell Transplantation (HSCT). Sensors 2020, 20, 6100. https://doi.org/10.3390/s20216100
Gupta V, Braun TM, Chowdhury M, Tewari M, Choi SW. A Systematic Review of Machine Learning Techniques in Hematopoietic Stem Cell Transplantation (HSCT). Sensors. 2020; 20(21):6100. https://doi.org/10.3390/s20216100
Chicago/Turabian StyleGupta, Vibhuti, Thomas M. Braun, Mosharaf Chowdhury, Muneesh Tewari, and Sung Won Choi. 2020. "A Systematic Review of Machine Learning Techniques in Hematopoietic Stem Cell Transplantation (HSCT)" Sensors 20, no. 21: 6100. https://doi.org/10.3390/s20216100
APA StyleGupta, V., Braun, T. M., Chowdhury, M., Tewari, M., & Choi, S. W. (2020). A Systematic Review of Machine Learning Techniques in Hematopoietic Stem Cell Transplantation (HSCT). Sensors, 20(21), 6100. https://doi.org/10.3390/s20216100





