Aerial Mapping of Forests Affected by Pathogens Using UAVs, Hyperspectral Sensors, and Artificial Intelligence
Abstract
1. Introduction
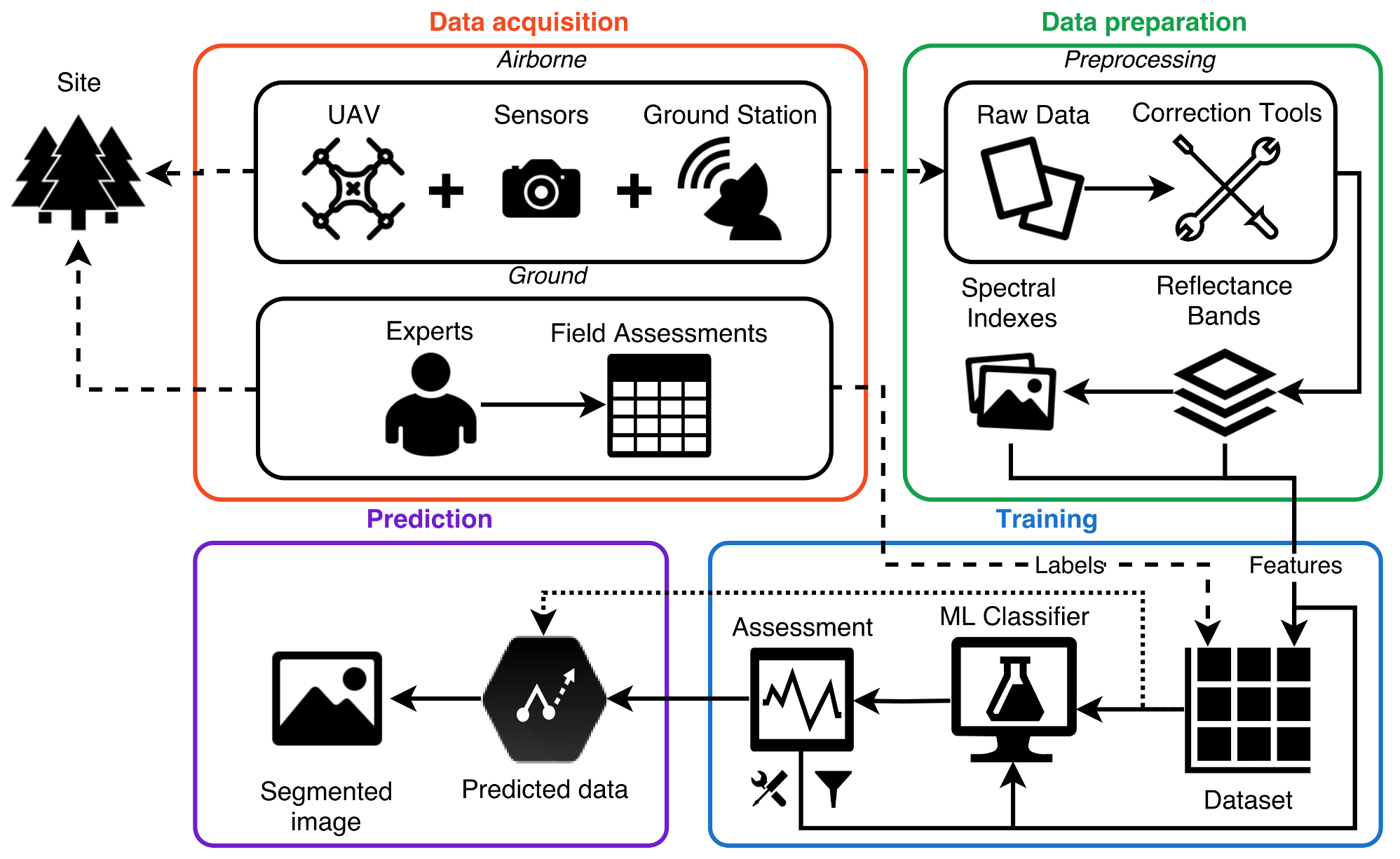
2. System Framework
2.1. Data Acquisition
2.1.1. UAV and Ground Station
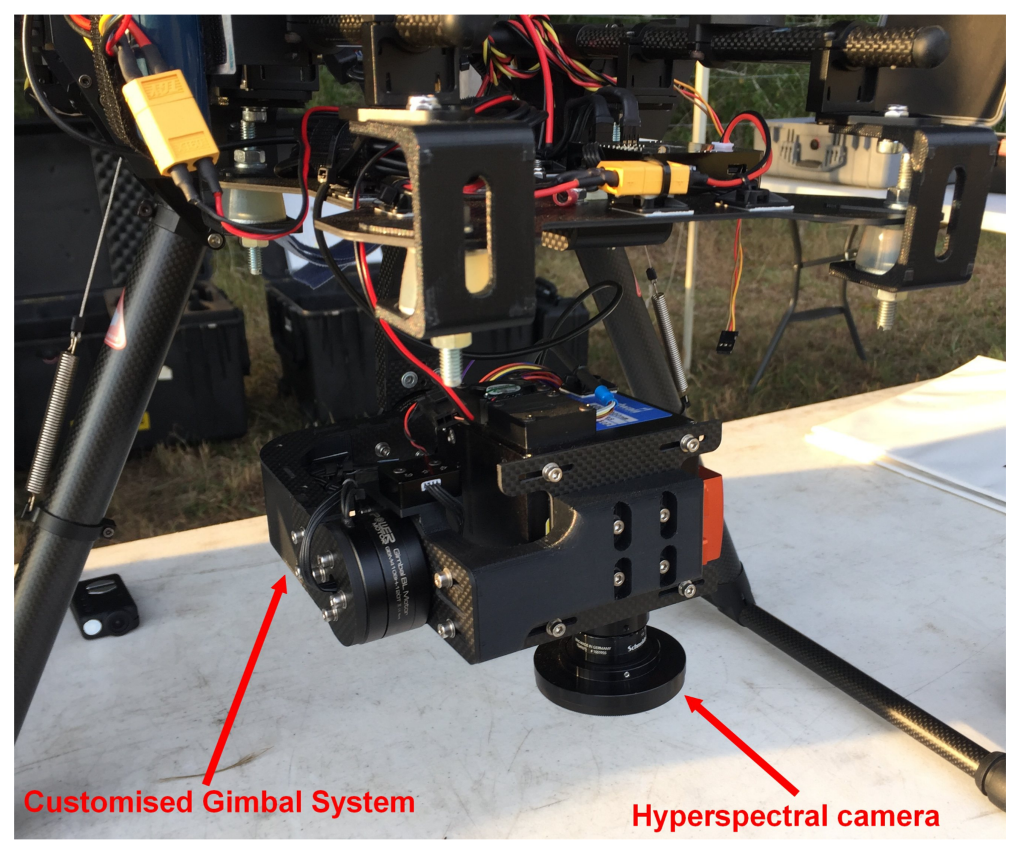
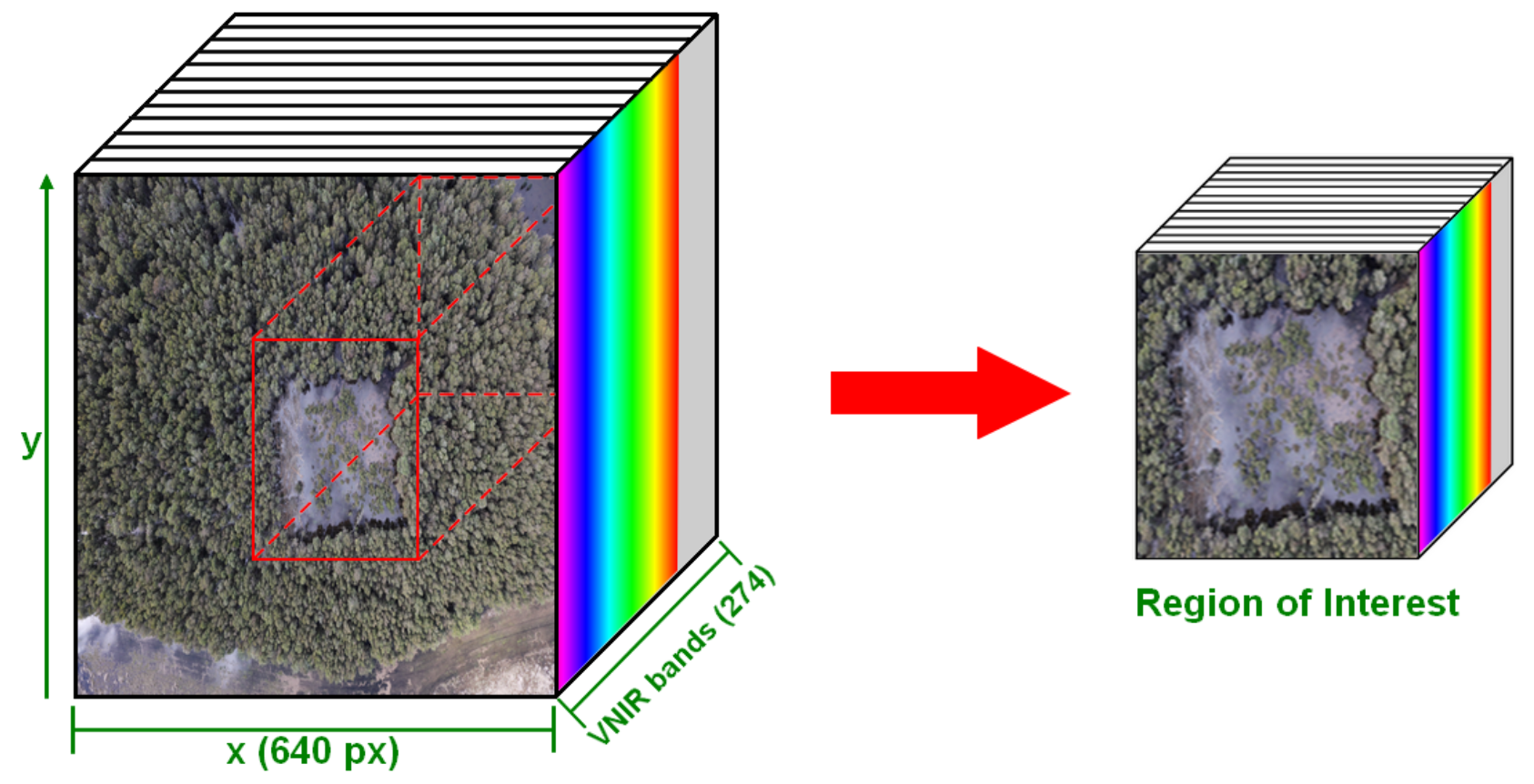
2.1.2. Sensors
2.2. Data Preparation
| Algorithm 1 Detection and mapping of vegetation alterations using spectral imagery and sets of features. | |
| Required: orthorectified layers (bands) in reflectance I. Labelled regions from field assessments L. | |
| Data Preparation | |
| 1: | Load I data. |
| 2: | Spectral indexes array from I. |
| 3: | Features array [I, S]. |
| Training | |
| 4: | Labels array from dataset L. |
| 5: | filtered dataset of features X with corresponding labelled pixel from Y. |
| 6: | Split D into training data and testing data . |
| 7: | Fit an XGBoost classifier C using . |
| 8: | List of unique relevance values of processed features X from C. |
| 9: | for all values in R do |
| 10: | Filtered underscored features from . |
| 11: | Fit C using . |
| 12: | Append accuracy values from C into T. |
| 13: | end for |
| 14: | Fit C using the best features threshold from T. |
| 15: | Validate C with k-fold cross-validation from ▹number of folds = 10 |
| Prediction | |
| 16: | Predicted values for each sample in X. |
| 17: | Convert P array into a 2D orthorectified image. |
| 18: | Displayed/overlayed image. |
| 19: | return O |
2.3. Training and Prediction
3. Experimentation Setup
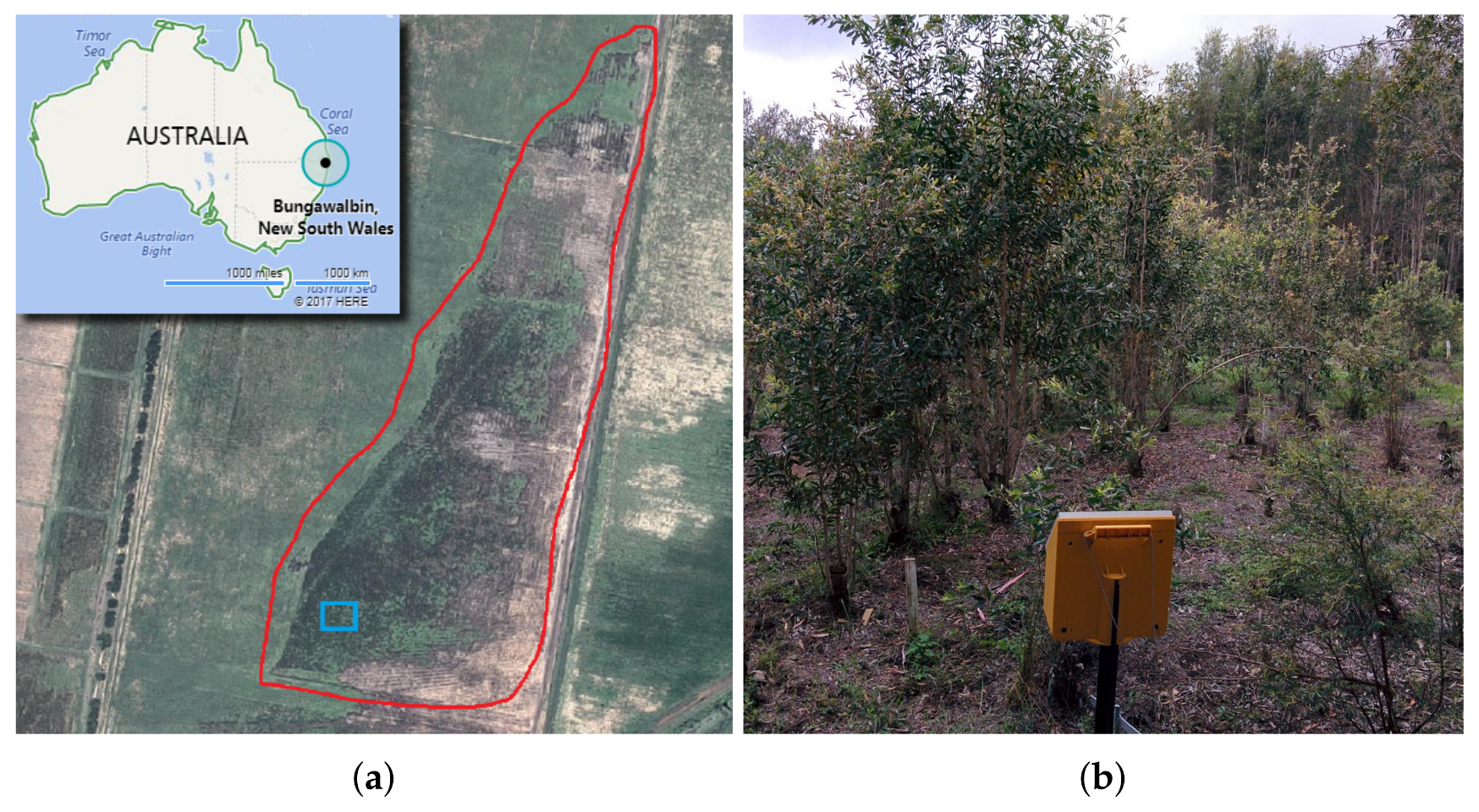
3.1. Site
3.2. Flight Campaign
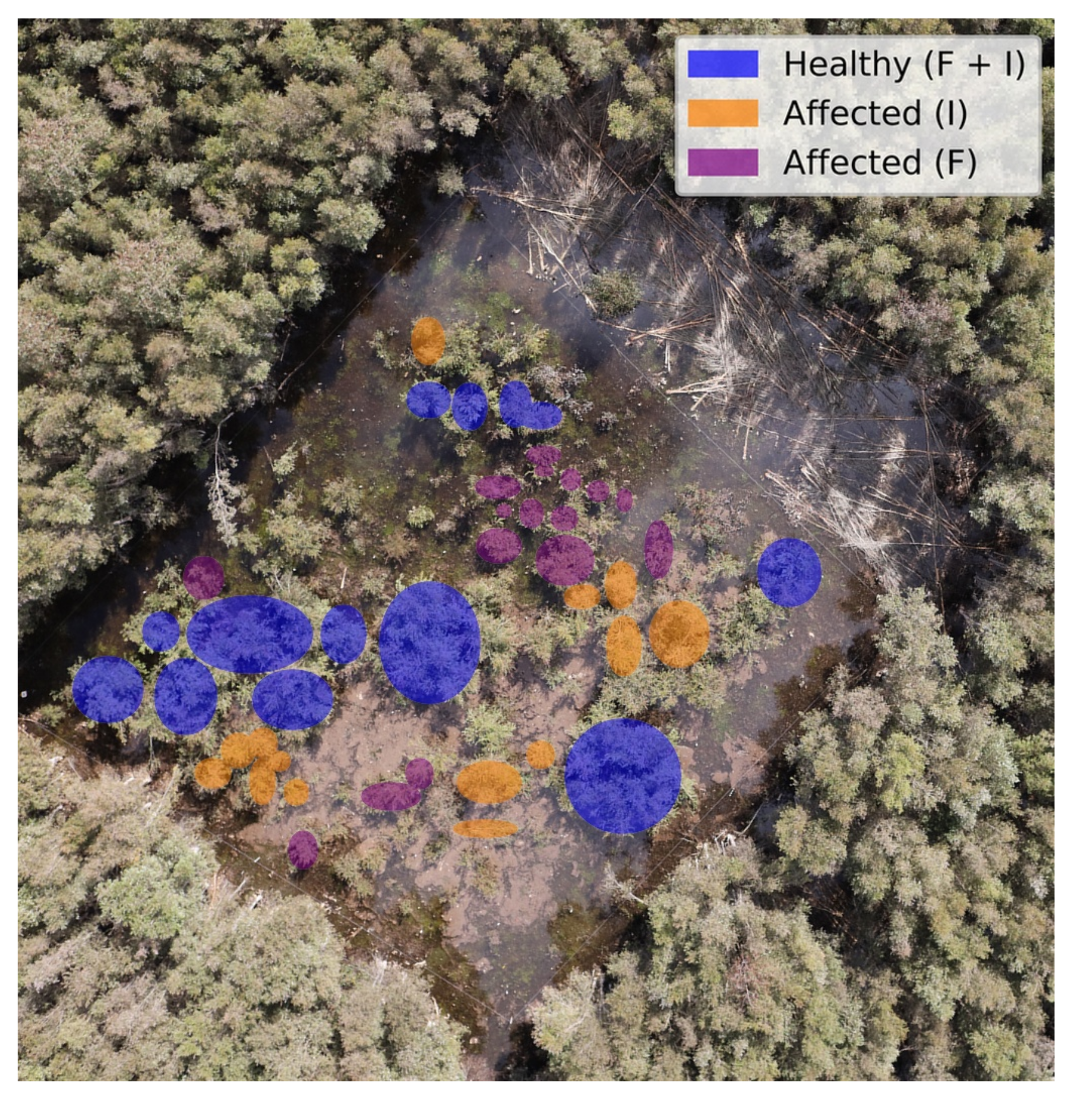
3.3. Field Assessments
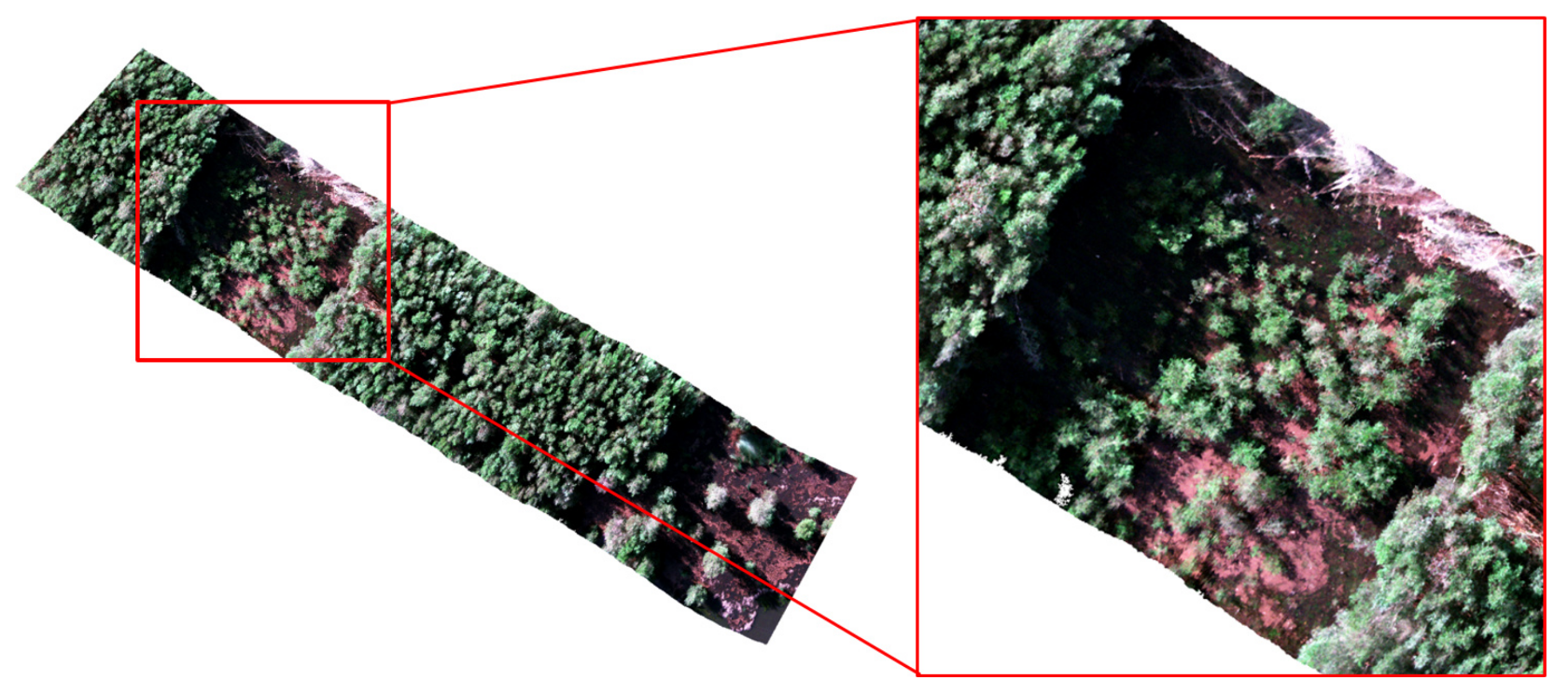
3.4. Preprocessing
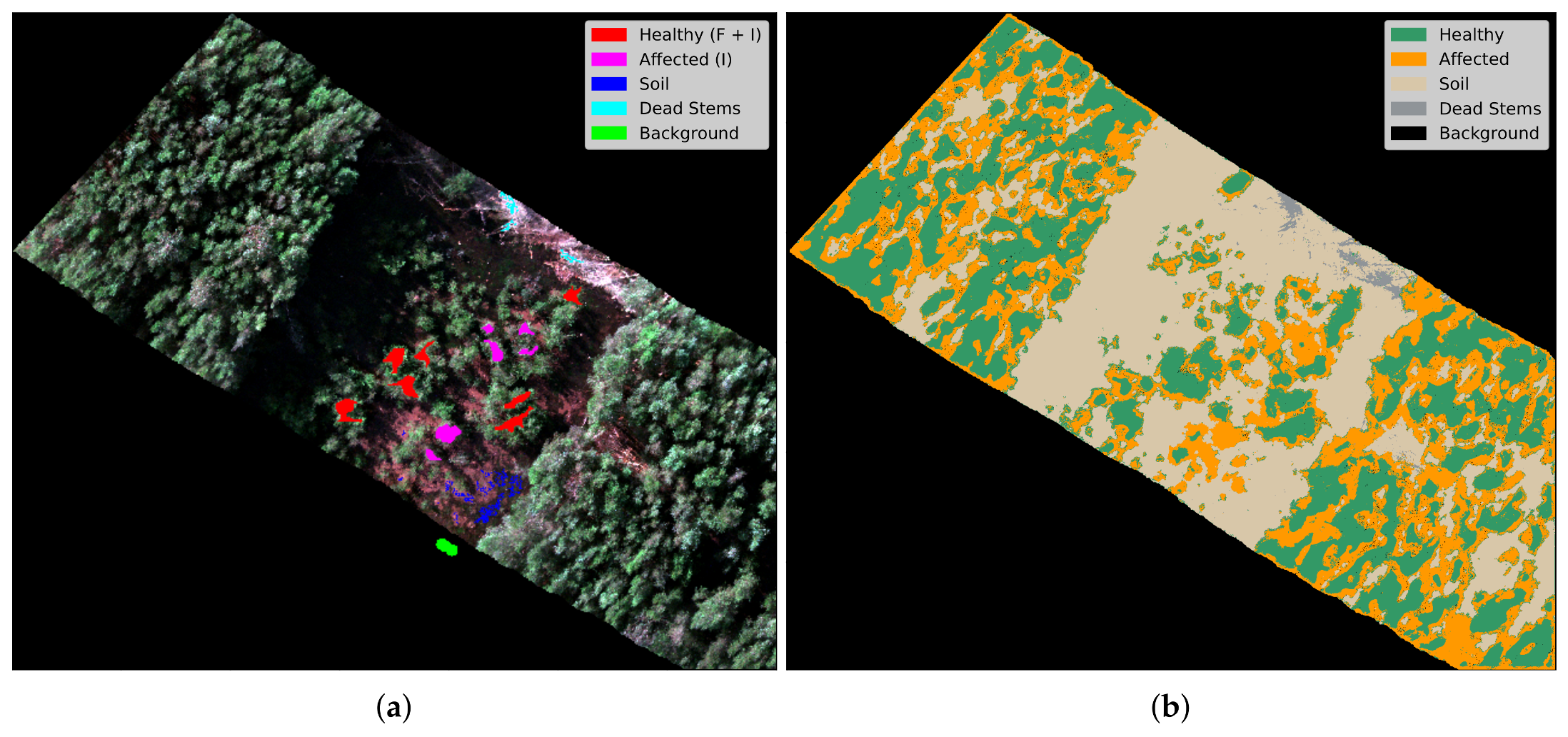
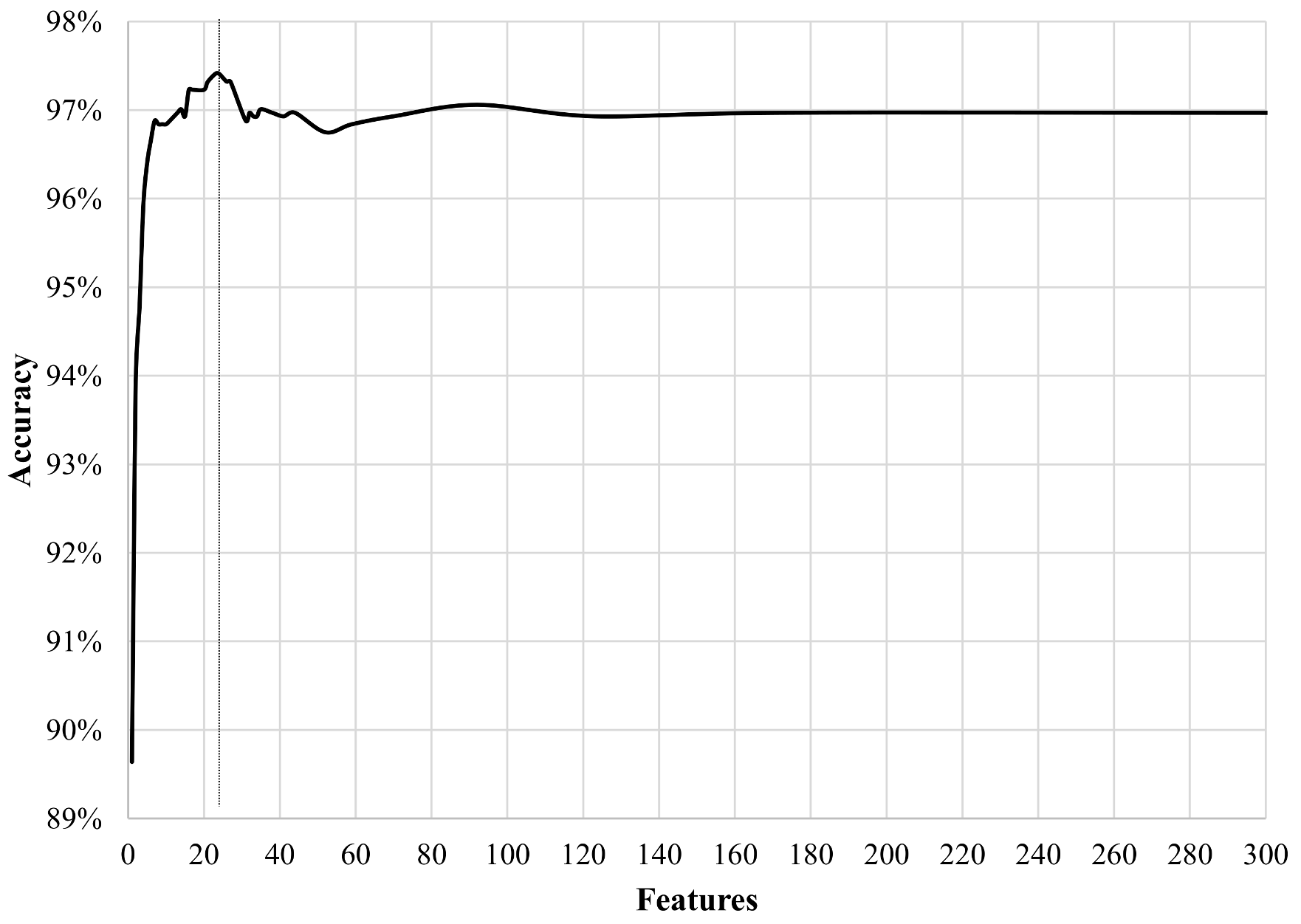
3.5. Training and Prediction
4. Results and Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| 2D | Two-dimensional |
| 3D | Three-dimensional |
| F | Fungicides |
| F + I | Fungicides and Insecticides |
| GDAL | Geospatial data abstraction library |
| GPS | Global positioning system |
| GNDVI | Green normalised difference vegetation index |
| GSD | Ground sampling distance |
| I | Insecticides |
| k-NN | k-nearest neighbours |
| LDA | Linear discriminant analysis |
| LiDAR | Light detection and ranging |
| MDPI | Multidisciplinary digital publishing institute |
| MSAVI2 | Second modified soil-adjusted vegetation index |
| NDVI | Normalised difference vegetation index |
| NSW | New South Wales |
| RGB | Red–green–blue colour model |
| SAVI | Soil-adjusted Vegetation Index |
| SVM | Support Vector Machines |
| TIF | Tagged Image File |
| UAS | Unmanned Aerial System |
| UAV | Unmanned Aerial Vehicle |
| VNIR | Visible Near Infrared |
| WRIS | White Reference Illumination Spectrum |
| XGBoost | eXtreme Gradient Boosting |
References
- Smock, L.A.; MacGregor, C.M. Impact of the American Chestnut Blight on Aquatic Shredding Macroinvertebrates. J. N. Am. Benthol. Soc. 1988, 7, 212–221. [Google Scholar] [CrossRef]
- Anagnostakis, S.L. Chestnut Blight: The classical problem of an introduced pathogen. Mycologia 1987, 79, 23–37. [Google Scholar] [CrossRef]
- Burke, K.L. The effects of logging and disease on American chestnut. For. Ecol. Manag. 2011, 261, 1027–1033. [Google Scholar] [CrossRef]
- Rizzo, D.M.; Garbelotto, M.; Hansen, E.M. Phytophthora ramorum: Integrative research and management of an emerging pathogen in California and oregon forests. Annu. Rev. Phytopathol. 2005, 43, 309–335. [Google Scholar] [CrossRef] [PubMed]
- Frankel, S.J. Sudden oak death and Phytophthora ramorum in the USA: A management challenge. Australas. Plant Pathol. 2008, 37, 19–25. [Google Scholar] [CrossRef]
- Grünwald, N.J.; Garbelotto, M.; Goss, E.M.; Heungens, K.; Prospero, S. Emergence of the sudden oak death pathogen Phytophthora ramorum. Trends Microbiol. 2012, 20, 131–138. [Google Scholar] [CrossRef] [PubMed]
- Hardham, A.R. Phytophthora cinnamomi. Mol. Plant Pathol. 2005, 6, 589–604. [Google Scholar] [CrossRef] [PubMed]
- Shearer, B.L.; Crane, C.E.; Barrett, S.; Cochrane, A. Phytophthora cinnamomi invasion, a major threatening process to conservation of flora diversity in the South-West Botanical Province of Western Australia. Aust. J. Bot. 2007, 55, 225–238. [Google Scholar] [CrossRef]
- Burgess, T.I.; Scott, J.K.; Mcdougall, K.L.; Stukely, M.J.; Crane, C.; Dunstan, W.A.; Brigg, F.; Andjic, V.; White, D.; Rudman, T.; et al. Current and projected global distribution of Phytophthora cinnamomi, one of the world’s worst plant pathogens. Glob. Chang. Biol. 2017, 23, 1661–1674. [Google Scholar] [CrossRef] [PubMed]
- Pegg, G.S.; Giblin, F.R.; McTaggart, A.R.; Guymer, G.P.; Taylor, H.; Ireland, K.B.; Shivas, R.G.; Perry, S. Puccinia psidii in Queensland, Australia: Disease symptoms, distribution and impact. Plant Pathol. 2014, 63, 1005–1021. [Google Scholar] [CrossRef]
- Carnegie, A.J.; Kathuria, A.; Pegg, G.S.; Entwistle, P.; Nagel, M.; Giblin, F.R. Impact of the invasive rust Puccinia psidii (myrtle rust) on native Myrtaceae in natural ecosystems in Australia. Biol. Invasions 2016, 18, 127–144. [Google Scholar] [CrossRef]
- Howard, C.; Findlay, V.; Grant, C. Australia’s transition to management of myrtle rust. J. For. Sci. 2016, 61, 138–139. [Google Scholar] [CrossRef]
- Fernandez Winzer, L.; Carnegie, A.J.; Pegg, G.S.; Leishman, M.R. Impacts of the invasive fungus Austropuccinia psidii (myrtle rust) on three Australian Myrtaceae species of coastal swamp woodland. Austral Ecol. 2017, 43. [Google Scholar] [CrossRef]
- Dayton, L.; Higgins, E. Myrtle rust ‘biggest threat to ecosystem’. Available online: http://www.webcitation.org/6y61T6sI6 (accessed on 19 February 2018).
- Carnegie, A.J.; Cooper, K. Emergency response to the incursion of an exotic myrtaceous rust in Australia. Australas. Plant Pathol. 2011, 40, 346–359. [Google Scholar] [CrossRef]
- Carnegie, A.J. First Report of Puccinia psidii (Myrtle Rust) in Eucalyptus Plantations in Australia. Plant Dis. 2015, 99, 161. [Google Scholar] [CrossRef]
- Pegg, G.; Taylor, T.; Entwistle, P.; Guymer, G.; Giblin, F.; Carnegie, A. Impact of Austropuccinia psidii (myrtle rust) on Myrtaceae-rich wet sclerophyll forests in south east Queensland. PLoS ONE 2017, 12, e0188058. [Google Scholar] [CrossRef] [PubMed]
- Government of Western Australia. Phytophthora Dieback—Parks and Wildlife Service. Available online: http://www.webcitation.org/6xLA86qjW (accessed on 19 February 2018).
- U.S. Forest Service. Sudden Oak Death (SOD)|Partnerships|PSW Research Station|Forest Service. Available online: http://www.webcitation.org/6xLDwPURd (accessed on 19 February 2018).
- State of Hawaii. Department of Agriculture|How to Report Suspected Ohia Wilt/Rapid Ohia Death. Available online: http://www.webcitation.org/6xLCVG70h (accessed on 19 February 2018).
- Lawley, V.; Lewis, M.; Clarke, K.; Ostendorf, B. Site-based and remote sensing methods for monitoring indicators of vegetation condition: An Australian review. Ecol. Indic. 2016, 60, 1273–1283. [Google Scholar] [CrossRef]
- Oliver, I.; Smith, P.L.; Lunt, I.; Parkes, D. Pre-1750 vegetation, naturalness and vegetation condition: What are the implications for biodiversity conservation? Ecol. Manag. Restor. 2002, 3, 176–178. [Google Scholar] [CrossRef]
- Lawley, V.; Parrott, L.; Lewis, M.; Sinclair, R.; Ostendorf, B. Self-organization and complex dynamics of regenerating vegetation in an arid ecosystem: 82 years of recovery after grazing. J. Arid Environ. 2013, 88, 156–164. [Google Scholar] [CrossRef]
- Ostendorf, B. Overview: Spatial information and indicators for sustainable management of natural resources. Ecol. Indic. 2011, 11, 97–102. [Google Scholar] [CrossRef]
- Roux, J.; Germishuizen, I.; Nadel, R.; Lee, D.J.; Wingfield, M.J.; Pegg, G.S. Risk assessment for Puccinia psidii becoming established in South Africa. Plant Pathol. 2015, 64, 1326–1335. [Google Scholar] [CrossRef]
- Berthon, K.; Esperon-Rodriguez, M.; Beaumont, L.; Carnegie, A.; Leishman, M. Assessment and prioritisation of plant species at risk from myrtle rust (Austropuccinia psidii) under current and future climates in Australia. Biol. Conserv. 2018, 218, 154–162. [Google Scholar] [CrossRef]
- Lausch, A.; Erasmi, S.; King, D.J.; Magdon, P.; Heurich, M. Understanding Forest Health with Remote Sensing -Part I –A Review of Spectral Traits, Processes and Remote-Sensing Characteristics. Remote Sens. 2016, 8, 1029. [Google Scholar] [CrossRef]
- Tuominen, J.; Lipping, T.; Kuosmanen, V.; Haapanen, R. Remote sensing of forest health. In Geoscience and Remote Sensing; Ho, P.G.P., Ed.; InTech: Rijeka, Croatia, 2009; Chapter 02. [Google Scholar]
- Lausch, A.; Erasmi, S.; King, D.J.; Magdon, P.; Heurich, M. Understanding forest health with remote sensing-Part II–A review of approaches and data models. Remote Sens. 2017, 9, 129. [Google Scholar] [CrossRef]
- Cui, D.; Zhang, Q.; Li, M.; Zhao, Y.; Hartman, G.L. Detection of soybean rust using a multispectral image sensor. Sens. Instrum. Food Qual. Saf. 2009, 3, 49–56. [Google Scholar] [CrossRef]
- Candiago, S.; Remondino, F.; De Giglio, M.; Dubbini, M.; Gattelli, M. Evaluating multispectral images and vegetation indices for precision farming applications from UAV images. Remote Sens. 2015, 7, 4026–4047. [Google Scholar] [CrossRef]
- Lowe, A.; Harrison, N.; French, A.P. Hyperspectral image analysis techniques for the detection and classification of the early onset of plant disease and stress. Plant Methods 2017, 13, 80. [Google Scholar] [CrossRef] [PubMed]
- Khanal, S.; Fulton, J.; Shearer, S. An overview of current and potential applications of thermal remote sensing in precision agriculture. Comput. Electron. Agric. 2017, 139, 22–32. [Google Scholar] [CrossRef]
- Devadas, R.; Lamb, D.W.; Simpfendorfer, S.; Backhouse, D. Evaluating ten spectral vegetation indices for identifying rust infection in individual wheat leaves. Precis. Agric. 2009, 10, 459–470. [Google Scholar] [CrossRef]
- Ashourloo, D.; Mobasheri, M.; Huete, A. Developing two spectral disease indices for detection of wheat leaf rust (Pucciniatriticina). Remote Sens. 2014, 6, 4723–4740. [Google Scholar] [CrossRef]
- Wang, H.; Qin, F.; Liu, Q.; Ruan, L.; Wang, R.; Ma, Z.; Li, X.; Cheng, P.; Wang, H. Identification and disease index inversion of wheat stripe rust and wheat leaf rust based on hyperspectral data at canopy level. J. Spectrosc. 2015, 2015, 1–10. [Google Scholar] [CrossRef]
- Heim, R.H.J.; Wright, I.J.; Chang, H.C.; Carnegie, A.J.; Pegg, G.S.; Lancaster, E.K.; Falster, D.S.; Oldeland, J. Detecting myrtle rust (Austropuccinia psidii) on lemon myrtle trees using spectral signatures and machine learning. Plant Pathol. 2018. [Google Scholar] [CrossRef]
- Booth, T.H.; Jovanovic, T. Assessing vulnerable areas for Puccinia psidii (eucalyptus rust) in Australia. Australas. Plant Pathol. 2012, 41, 425–429. [Google Scholar] [CrossRef]
- Elith, J.; Simpson, J.; Hirsch, M.; Burgman, M.A. Taxonomic uncertainty and decision making for biosecurity: spatial models for myrtle/guava rust. Australas. Plant Pathol. 2013, 42, 43–51. [Google Scholar] [CrossRef]
- Salami, E.; Barrado, C.; Pastor, E. UAV flight experiments applied to the remote sensing of vegetated areas. Remote Sens. 2014, 6, 11051–11081. [Google Scholar] [CrossRef]
- Glassock, R.; Hung, J.Y.; Gonzalez, L.F.; Walker, R.A. Design, modelling and measurement of a hybrid powerplant for unmanned aerial systems. Aust. J. Mech. Eng. 2008, 6, 69–78. [Google Scholar] [CrossRef]
- Whitney, E.; Gonzalez, L.; Periaux, J.; Sefrioui, M.; Srinivas, K. A robust evolutionary technique for inverse aerodynamic design. In Proceedings of the European Congress on Computational Methods in Applied Sciences and Engineering, Jyvaskyla, Finland, 24–28 July 2004; Volume 2, pp. 1–2. [Google Scholar]
- Gonzalez, L.; Whitney, E.; Srinivas, K.; Periaux, J. Multidisciplinary aircraft design and optimisation using a robust evolutionary technique with variable fidelity models. In Proceedings of the 10th AIAA/ISSMO Multidisciplinary Analysis and Optimization Conference, Albany, NY, USA, 30 August–1 September 2004; Volume 6, pp. 3610–3624. [Google Scholar]
- Ken, W.; Chris, H.C. Remote sensing of the environment with small unmanned aircraft systems (UASs), part 1: A review of progress and challenges. J. Unmanned Veh. Syst. 2014, 2, 69–85. [Google Scholar]
- Gonzalez, L.; Montes, G.; Puig, E.; Johnson, S.; Mengersen, K.; Gaston, K. Unmanned Aerial Vehicles (UAVs) and artificial intelligence revolutionizing wildlife monitoring and conservation. Sensors 2016, 16, 97. [Google Scholar] [CrossRef] [PubMed]
- Sandino, J.; Wooler, A.; Gonzalez, F. Towards the automatic detection of pre-existing termite mounds through UAS and hyperspectral imagery. Sensors 2017, 17, 2196. [Google Scholar] [CrossRef] [PubMed]
- Vanegas, F.; Bratanov, D.; Powell, K.; Weiss, J.; Gonzalez, F. A novel methodology for improving plant pest surveillance in vineyards and crops using UAV-based hyperspectral and spatial data. Sensors 2018, 18, 260. [Google Scholar] [CrossRef] [PubMed]
- Vanegas, F.; Gonzalez, F. Enabling UAV navigation with sensor and environmental uncertainty in cluttered and GPS-denied environments. Sensors 2016, 16, 666. [Google Scholar] [CrossRef] [PubMed]
- Aasen, H.; Burkart, A.; Bolten, A.; Bareth, G. Generating 3D hyperspectral information with lightweight UAV snapshot cameras for vegetation monitoring: From camera calibration to quality assurance. ISPRS J. Photogramm. Remote Sens. 2015, 108, 245–259. [Google Scholar] [CrossRef]
- Nasi, R.; Honkavaara, E.; Lyytikainen-Saarenmaa, P.; Blomqvist, M.; Litkey, P.; Hakala, T.; Viljanen, N.; Kantola, T.; Tanhuanpaa, T.; Holopainen, M. Using UAV-Based photogrammetry and hyperspectral imaging for mapping bark beetle damage at tree-level. Remote Sens. 2015, 7, 15467–15493. [Google Scholar] [CrossRef]
- Calderon, R.; Navas-Cortes, J.; Lucena, C.; Zarco-Tejada, P. High-resolution airborne hyperspectral and thermal imagery for early detection of Verticillium wilt of olive using fluorescence, temperature and narrow-band spectral indices. Remote Sens. Environ. 2013, 139, 231–245. [Google Scholar] [CrossRef]
- Calderon, R.; Navas-Cortes, J.A.; Zarco-Tejada, P.J. Early detection and quantification of verticillium wilt in olive using hyperspectral and thermal imagery over large areas. Remote Sens. 2015, 7, 5584–5610. [Google Scholar] [CrossRef]
- Albetis, J.; Duthoit, S.; Guttler, F.; Jacquin, A.; Goulard, M.; Poilvé, H.; Féret, J.B.; Dedieu, G. Detection of Flavescence dorée Grapevine Disease using Unmanned Aerial Vehicle (UAV) multispectral imagery. Remote Sens. 2017, 9, 308. [Google Scholar] [CrossRef]
- Pause, M.; Schweitzer, C.; Rosenthal, M.; Keuck, V.; Bumberger, J.; Dietrich, P.; Heurich, M.; Jung, A.; Lausch, A. In situ/remote sensing integration to assess forest health–A review. Remote Sens. 2016, 8, 471. [Google Scholar] [CrossRef]
- Stone, C.; Mohammed, C. Application of remote sensing technologies for assessing planted forests damaged by insect pests and fungal pathogens: A review. Curr. For. Rep. 2017, 3, 75–92. [Google Scholar] [CrossRef]
- Habili, N.; Oorloff, J. Scyllarus™: From Research to Commercial Software. In Proceedings of the ASWEC 24th Australasian Software Engineering Conference, Adelaide, SA, Australia, 28 September–1 October 2015; ACM Press: New York, NY, USA, 2015; Volume II, pp. 119–122. [Google Scholar]
- Gu, L.; Robles-Kelly, A.A.; Zhou, J. Efficient estimation of reflectance parameters from imaging spectroscopy. IEEE Trans. Image Process. 2013, 22, 3648–3663. [Google Scholar] [PubMed]
- GDAL Development Team. GDAL—Geospatial Data Abstraction Library, Version 2.1.0; Open Source Geospatial Foundation: Beaverton, OR, USA, 2017. [Google Scholar]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (KDD’16), San Francisco, CA, USA, 13–17 August 2016; ACM Press: New York, NY, USA, 2016; pp. 785–794. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Bradski, G. The OpenCV library. Dr. Dobb’s J. Softw. Tools 2000, 25, 120, 122–125. [Google Scholar]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Rouse, J.W., Jr.; Haas, R.H.; Schell, J.A.; Deering, D.W. Monitoring vegetation systems in the great plains with Erts. NASA Spec. Publ. 1974, 351, 309–317. [Google Scholar]
- Gitelson, A.A.; Kaufman, Y.J.; Merzlyak, M.N. Use of a green channel in remote sensing of global vegetation from EOS-MODIS. Remote Sens. Environ. 1996, 58, 289–298. [Google Scholar] [CrossRef]
- Huete, A. A soil-adjusted vegetation index (SAVI). Remote Sens. Environ. 1988, 25, 295–309. [Google Scholar] [CrossRef]
- Laosuwan, T.; Uttaruk, P. Estimating tree biomass via remote sensing, MSAVI 2, and fractional cover model. IETE Tech. Rev. 2014, 31, 362–368. [Google Scholar] [CrossRef]
- Australian Government. Evans Head, NSW–August 2016–Daily Weather Observations; Bureau of Meteorology: Evans Head, NSW, Australia, 2016.



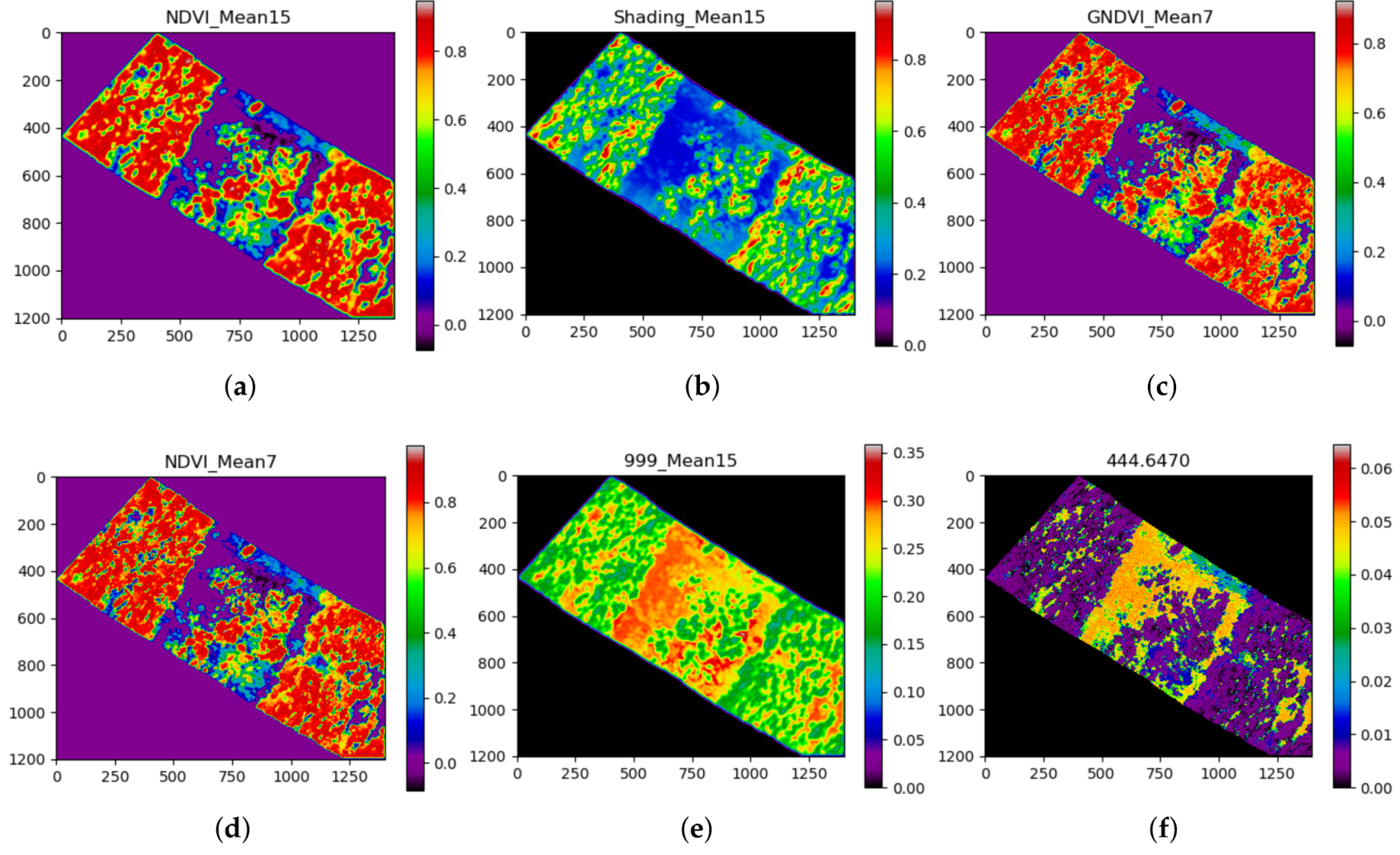

| # | Feature | Score | # | Feature | Score | # | Feature | Score |
|---|---|---|---|---|---|---|---|---|
| 1 | NDVI_Mean15 | 0.0933 | 11 | NDVI_Mean3 | 0.0247 | 21 | 975.3710 | 0.0119 |
| 2 | Shading_Mean15 | 0.0780 | 12 | 759.9730 | 0.0212 | 22 | 671.1490 | 0.0109 |
| 3 | GNDVI_Mean7 | 0.0563 | 13 | 999_Mean3 | 0.0212 | 23 | 893.2090 | 0.0109 |
| 4 | NDVI_Mean7 | 0.0558 | 14 | 999_Mean7 | 0.0202 | 24 | 990.9150 | 0.0099 |
| 5 | 999_Mean15 | 0.0504 | 15 | 997.5770 | 0.0188 | 25 | 877.6650 | 0.0094 |
| 6 | 444.6470 | 0.0494 | 16 | 764.4140 | 0.0148 | 26 | 966.4890 | 0.0094 |
| 7 | Specularity_Mean15 | 0.0380 | 17 | 444_Mean15 | 0.0143 | 27 | 766.6350 | 0.0084 |
| 8 | 999.7980 | 0.0341 | 18 | 462.4120 | 0.0133 | 28 | 853.2380 | 0.0079 |
| 9 | GNDVI_Mean15 | 0.0286 | 19 | NDVI | 0.0133 | 29 | 935.4000 | 0.0079 |
| 10 | 444_Mean7 | 0.0267 | 20 | Shading_Mean7 | 0.0133 | 30 | GNDVI_Mean3 | 0.0079 |
| Predicted | Healthy | Affected | Background | Soil | Stems | |
|---|---|---|---|---|---|---|
| Labelled | Healthy | 1049 | 15 | 0 | 0 | 0 |
| Affected | 45 | 531 | 0 | 0 | 0 | |
| Background | 0 | 0 | 158 | 0 | 0 | |
| Soil | 0 | 0 | 0 | 321 | 0 | |
| Stems | 0 | 0 | 0 | 1 | 157 |
| Class | Precision (%) | Recall (%) | F-Score (%) | Support |
|---|---|---|---|---|
| Healthy | 95.89 | 98.59 | 97.24 | 1064 |
| Affected | 97.25 | 92.19 | 94.72 | 576 |
| Background | 100.00 | 100.00 | 100.00 | 158 |
| Soil | 99.69 | 100.00 | 99.68 | 321 |
| Stems | 100.00 | 99.37 | 99.68 | 158 |
| Mean | 97.32 | 97.32 | 97.35 | ∑ = 2277 |
| Sub-Section | Instance 1 | Instance 2 | Instance 3 | Instance 4 | Instance 5 | Mean | Std. Dev. |
|---|---|---|---|---|---|---|---|
| Data preparation | |||||||
| Loading Hypercube | 11.927 | 10.944 | 11.954 | 11.766 | 11.521 | 11.622 | 0.417 |
| Calculating indexes | 46.864 | 51.860 | 51.901 | 52.622 | 47.322 | 50.114 | 2.779 |
| Training | |||||||
| Preprocessing | 0.152 | 0.141 | 0.149 | 0.148 | 0.140 | 0.146 | 0.005 |
| Fitting XGBoost | 8.948 | 8.654 | 8.758 | 8.679 | 8.692 | 8.746 | 0.119 |
| Features Filtering | 53.236 | 55.433 | 60.364 | 57.253 | 53.446 | 55.946 | 2.962 |
| Re-Fitting XGBoost | 0.964 | 1.023 | 1.010 | 0.998 | 0.965 | 0.992 | 0.026 |
| Prediction | |||||||
| Predicting results | 29.738 | 40.749 | 42.131 | 34.473 | 66.477 | 42.714 | 14.188 |
| Display | 0.776 | 0.705 | 1.043 | 0.917 | 0.612 | 0.811 | 0.171 |
| Total | 152.607 | 169.508 | 177.309 | 166.857 | 189.175 | 171.091 | 13.489 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sandino, J.; Pegg, G.; Gonzalez, F.; Smith, G. Aerial Mapping of Forests Affected by Pathogens Using UAVs, Hyperspectral Sensors, and Artificial Intelligence. Sensors 2018, 18, 944. https://doi.org/10.3390/s18040944
Sandino J, Pegg G, Gonzalez F, Smith G. Aerial Mapping of Forests Affected by Pathogens Using UAVs, Hyperspectral Sensors, and Artificial Intelligence. Sensors. 2018; 18(4):944. https://doi.org/10.3390/s18040944
Chicago/Turabian StyleSandino, Juan, Geoff Pegg, Felipe Gonzalez, and Grant Smith. 2018. "Aerial Mapping of Forests Affected by Pathogens Using UAVs, Hyperspectral Sensors, and Artificial Intelligence" Sensors 18, no. 4: 944. https://doi.org/10.3390/s18040944
APA StyleSandino, J., Pegg, G., Gonzalez, F., & Smith, G. (2018). Aerial Mapping of Forests Affected by Pathogens Using UAVs, Hyperspectral Sensors, and Artificial Intelligence. Sensors, 18(4), 944. https://doi.org/10.3390/s18040944





