HCV Detection, Discrimination, and Genotyping Technologies
Abstract
1. Introduction
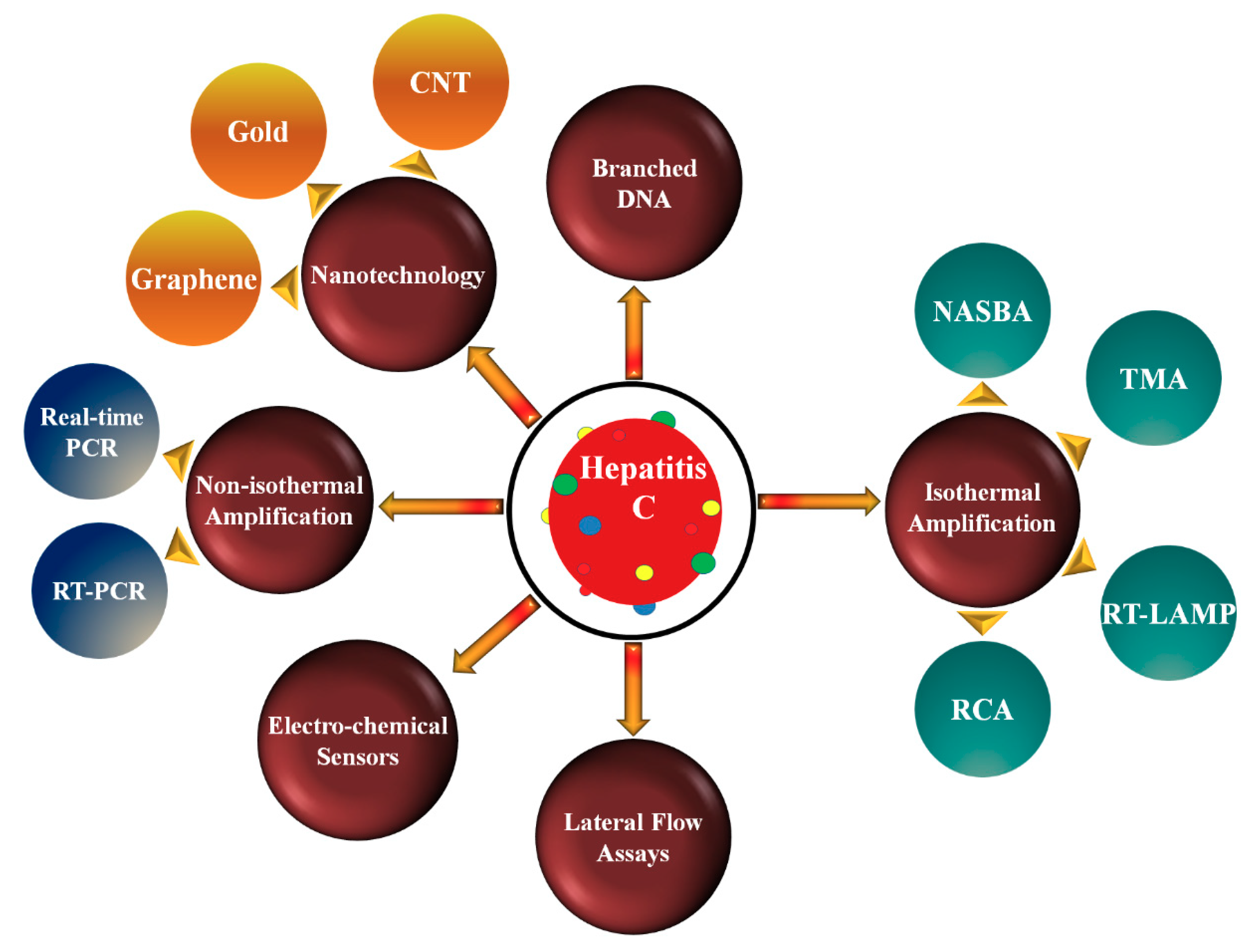
2. Nucleic Acid Amplification Technologies
2.1. Non-Isothermal Amplification Methods
2.1.1. Reverse Transcription Polymerase Chain Reaction (RT-PCR)
2.1.2. Real-Time PCR (RT-PCR)
2.2. Isothermal Amplification Methods (IMA)
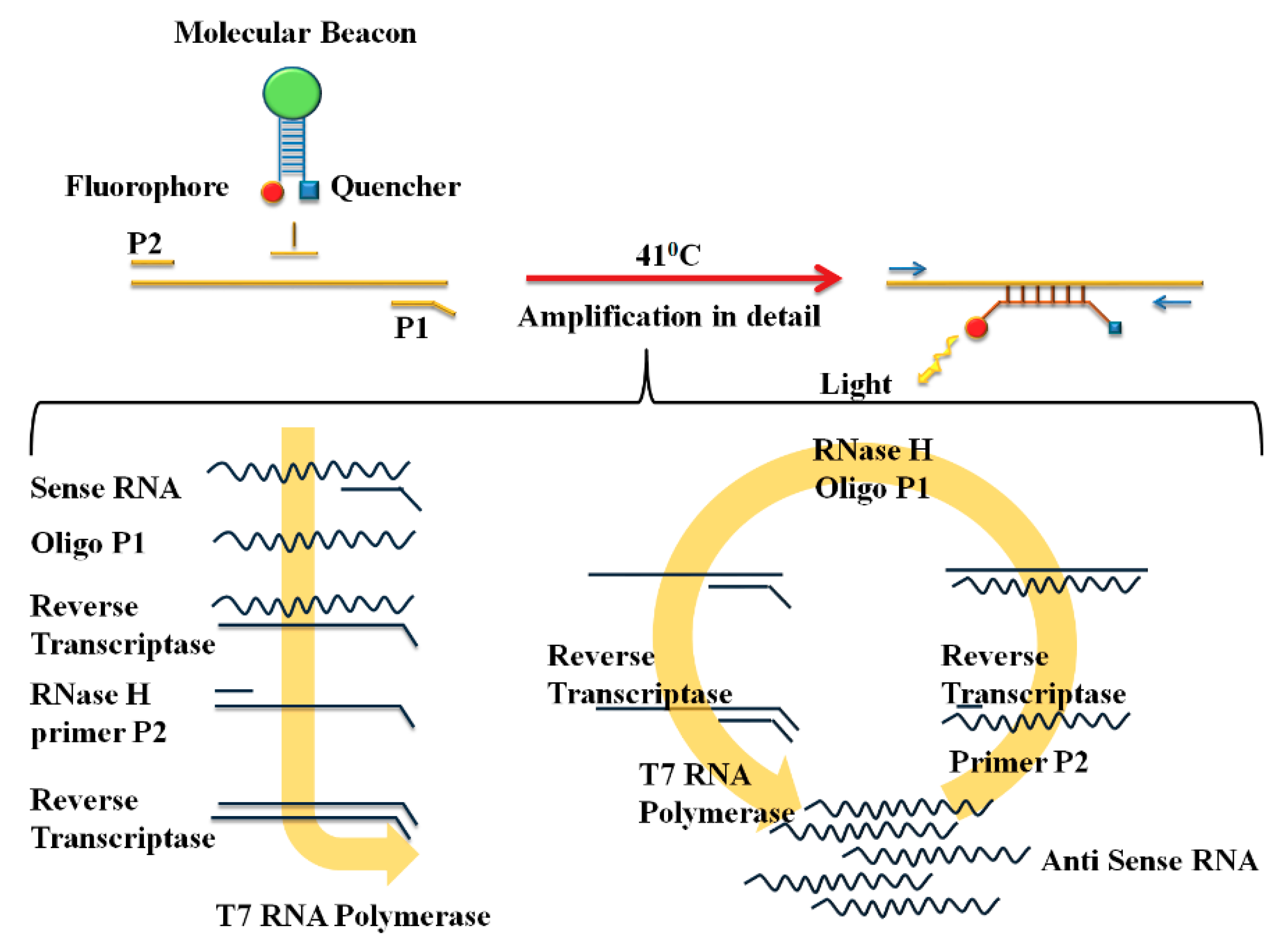
2.2.1. Nucleic Acid Sequence-Based Amplification (NASBA)
2.2.2. Transcription-Mediated Amplification (TMA)
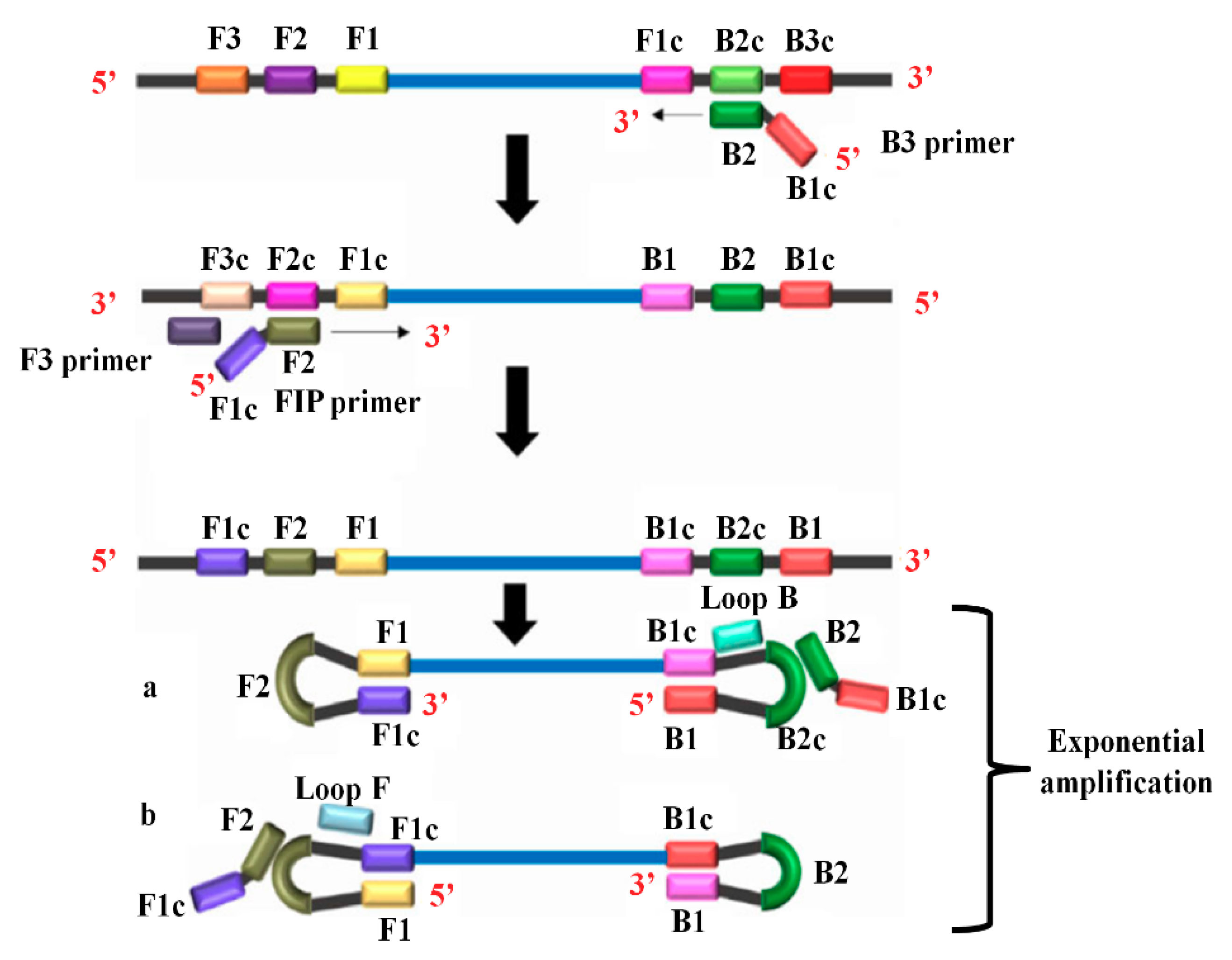
2.2.3. Reverse Transcription Loop-Mediated Isothermal Amplification (RT-LAMP)
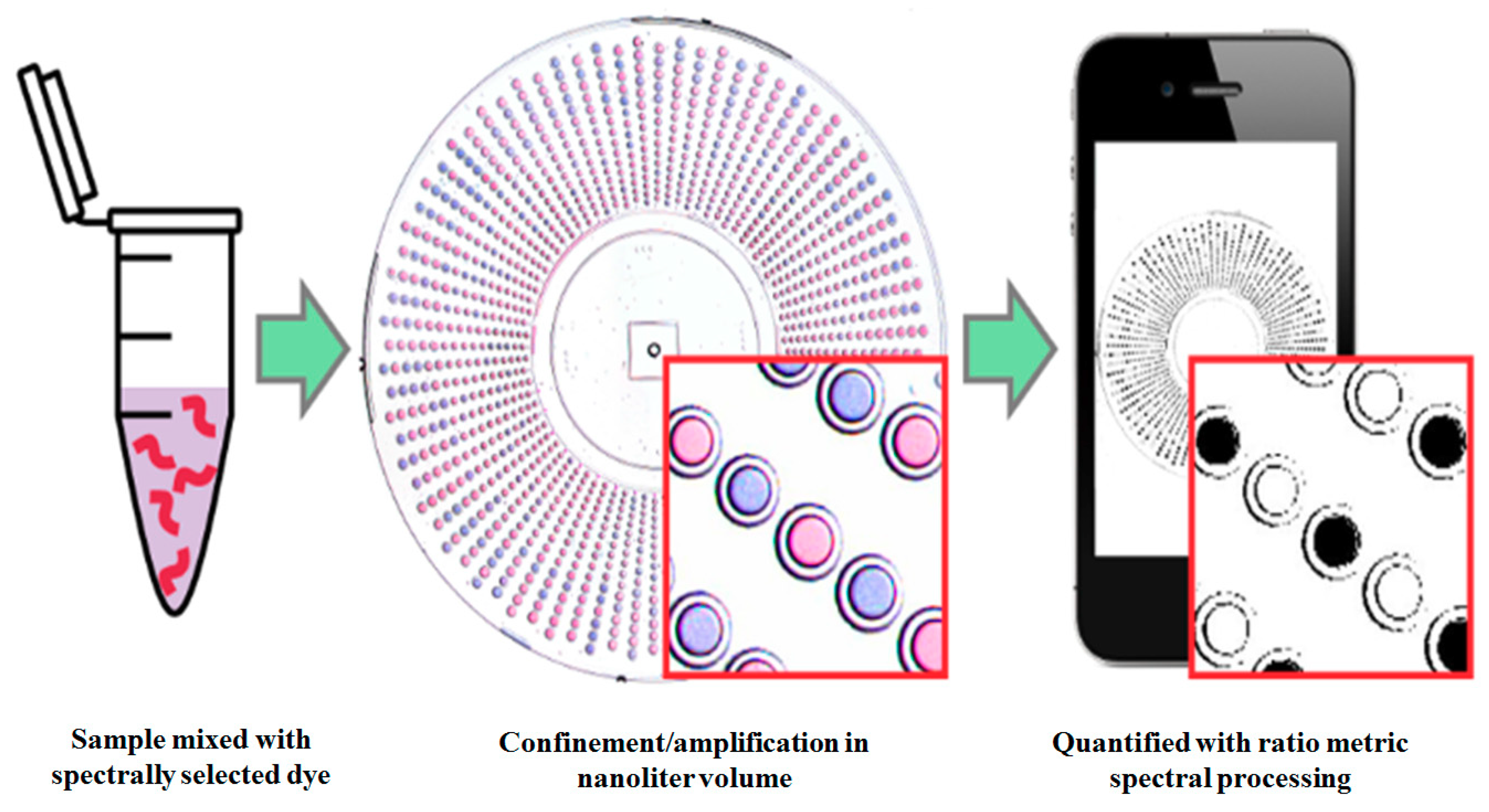
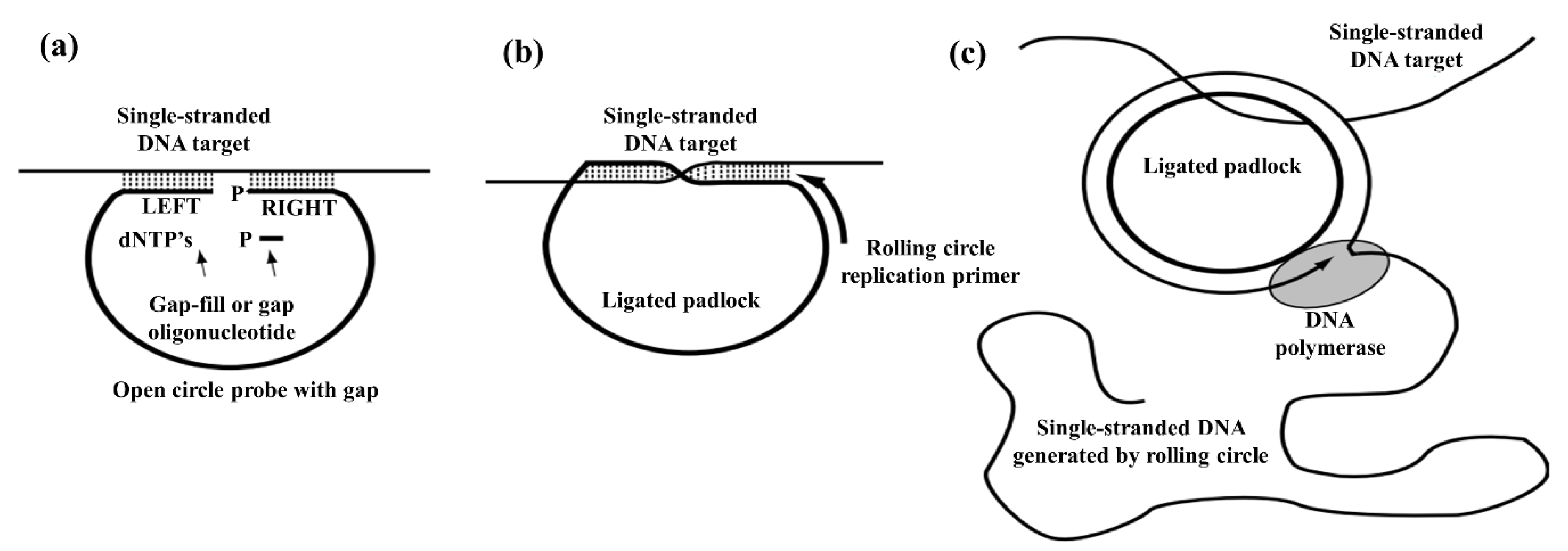
2.2.4. Rolling Circle Amplification (RCA) Method
3. Transducer Technologies
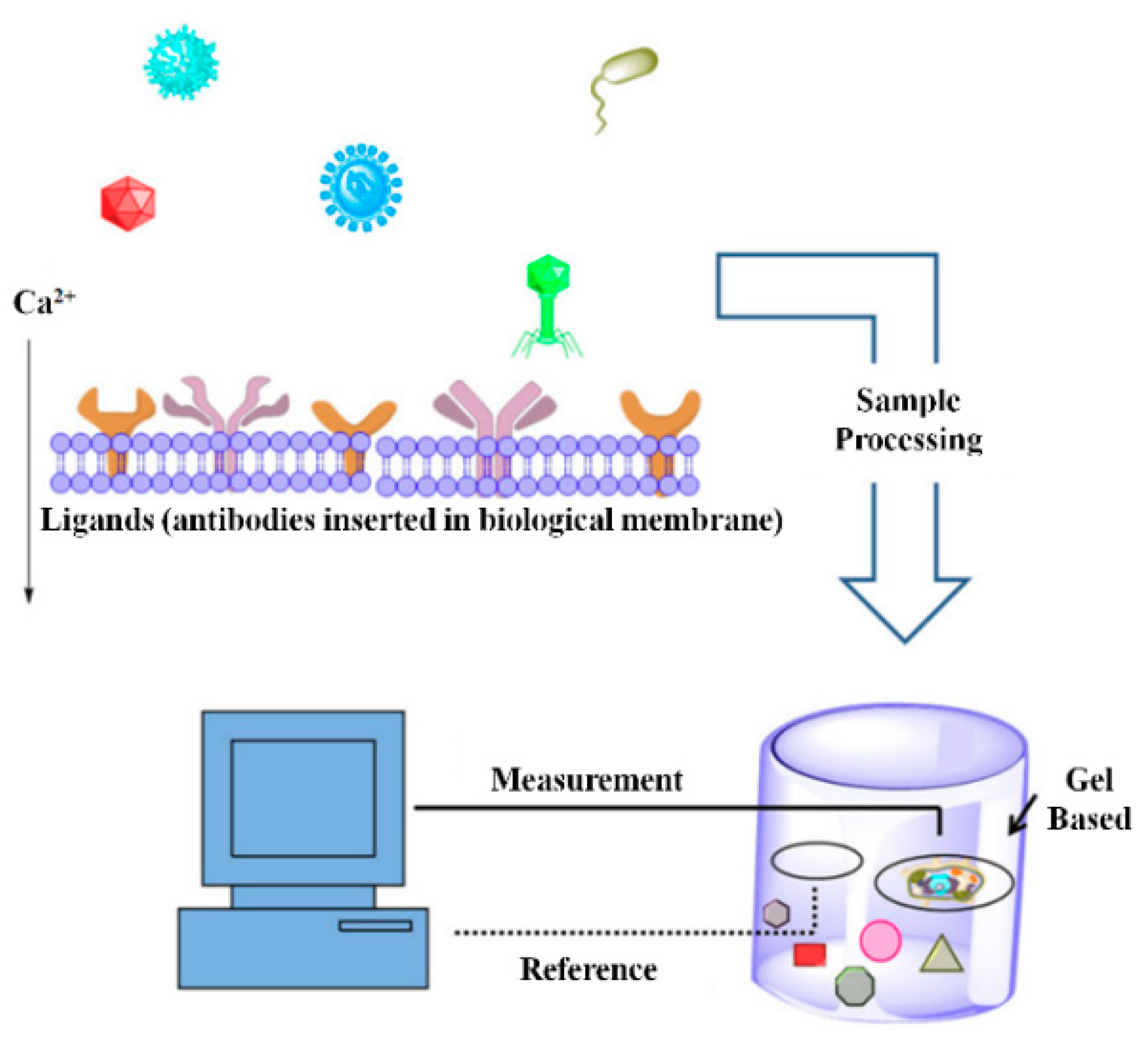
3.1. Bioelectric Recognition Assay (BERA)
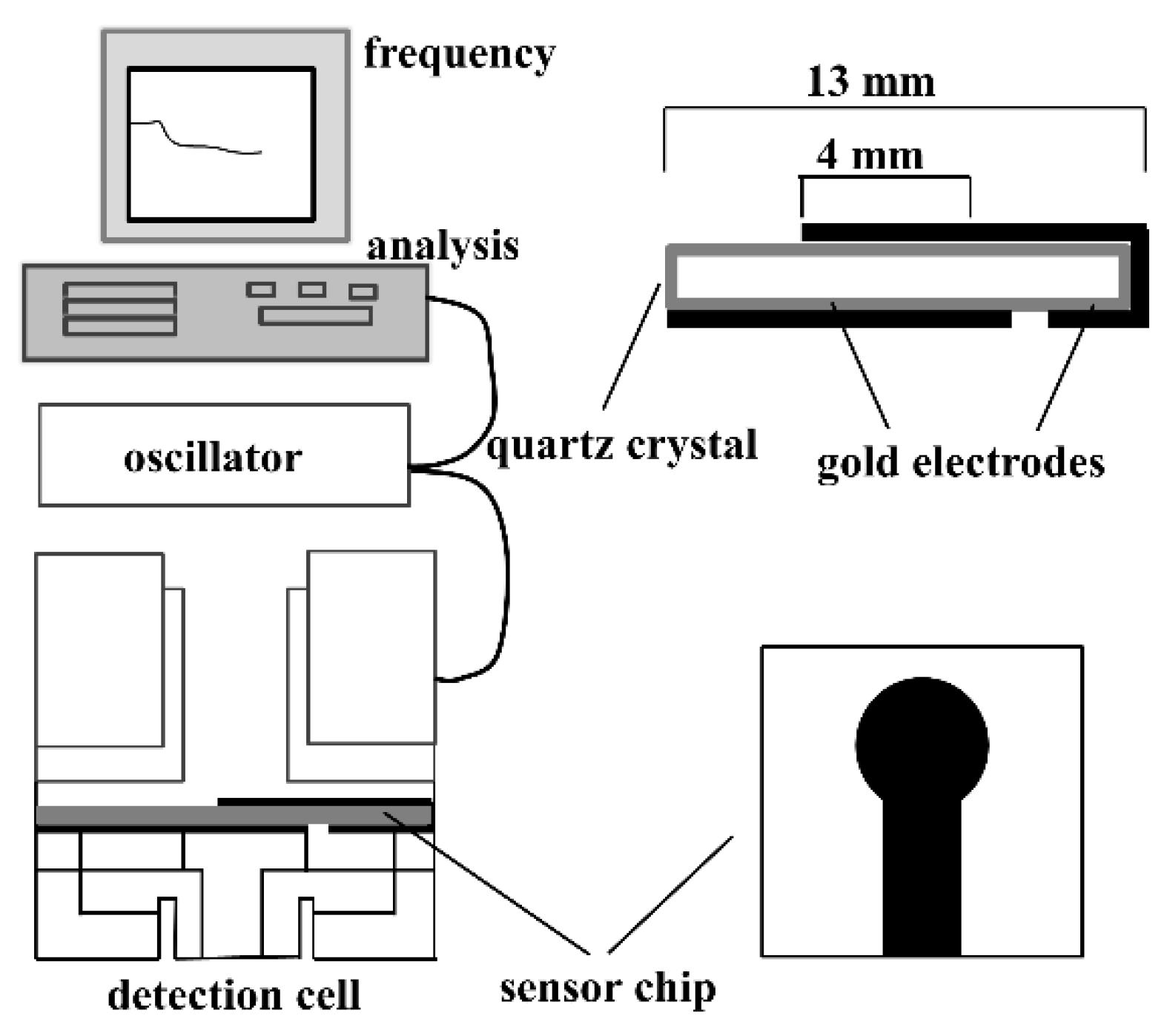
3.2. Piezoelectric Biosensors (PZ)
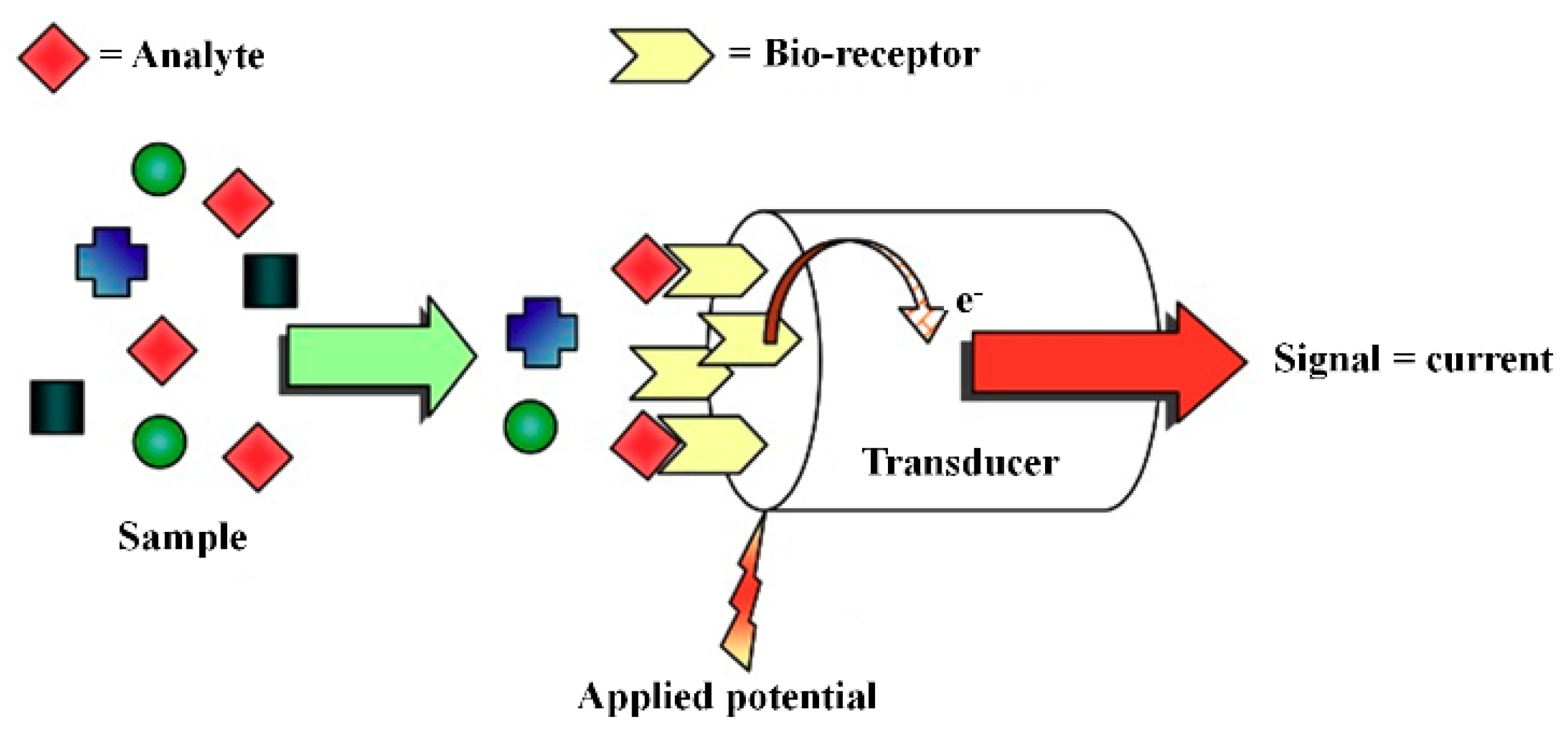
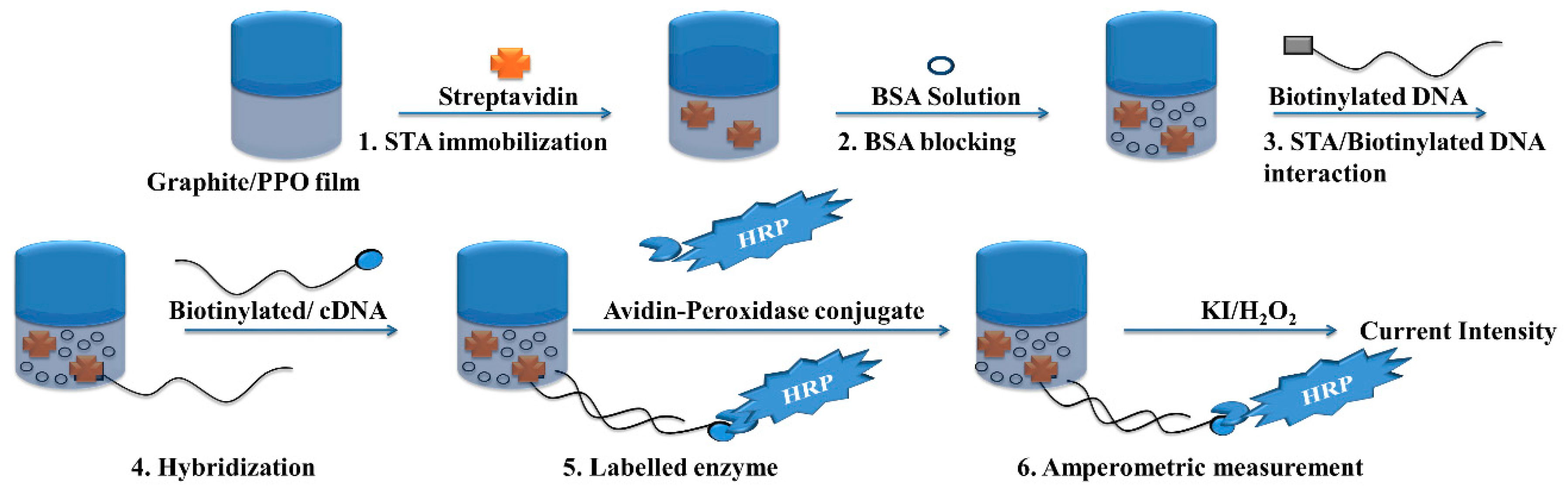
3.3. Amperometric Biosensor
3.4. Nanotechnology
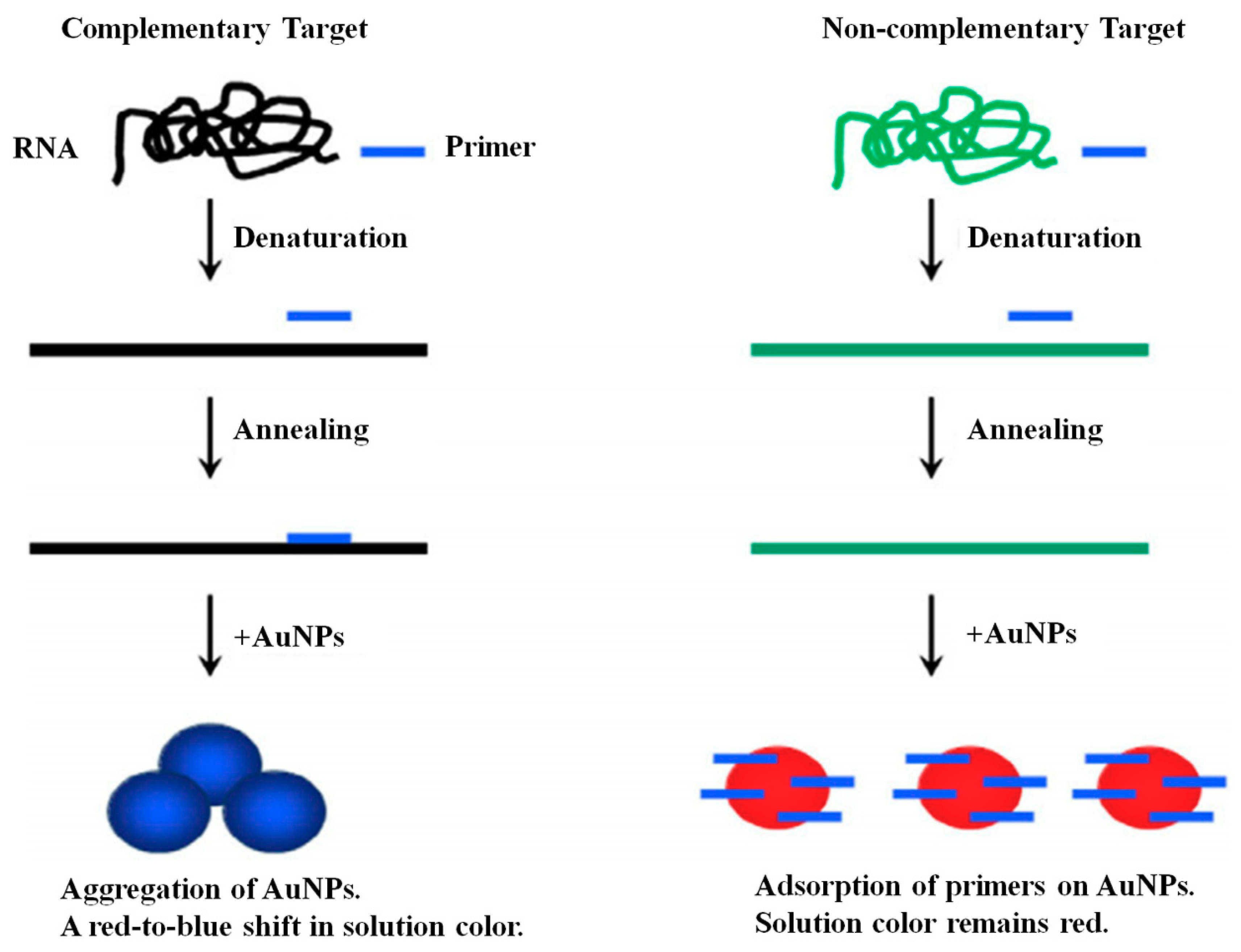
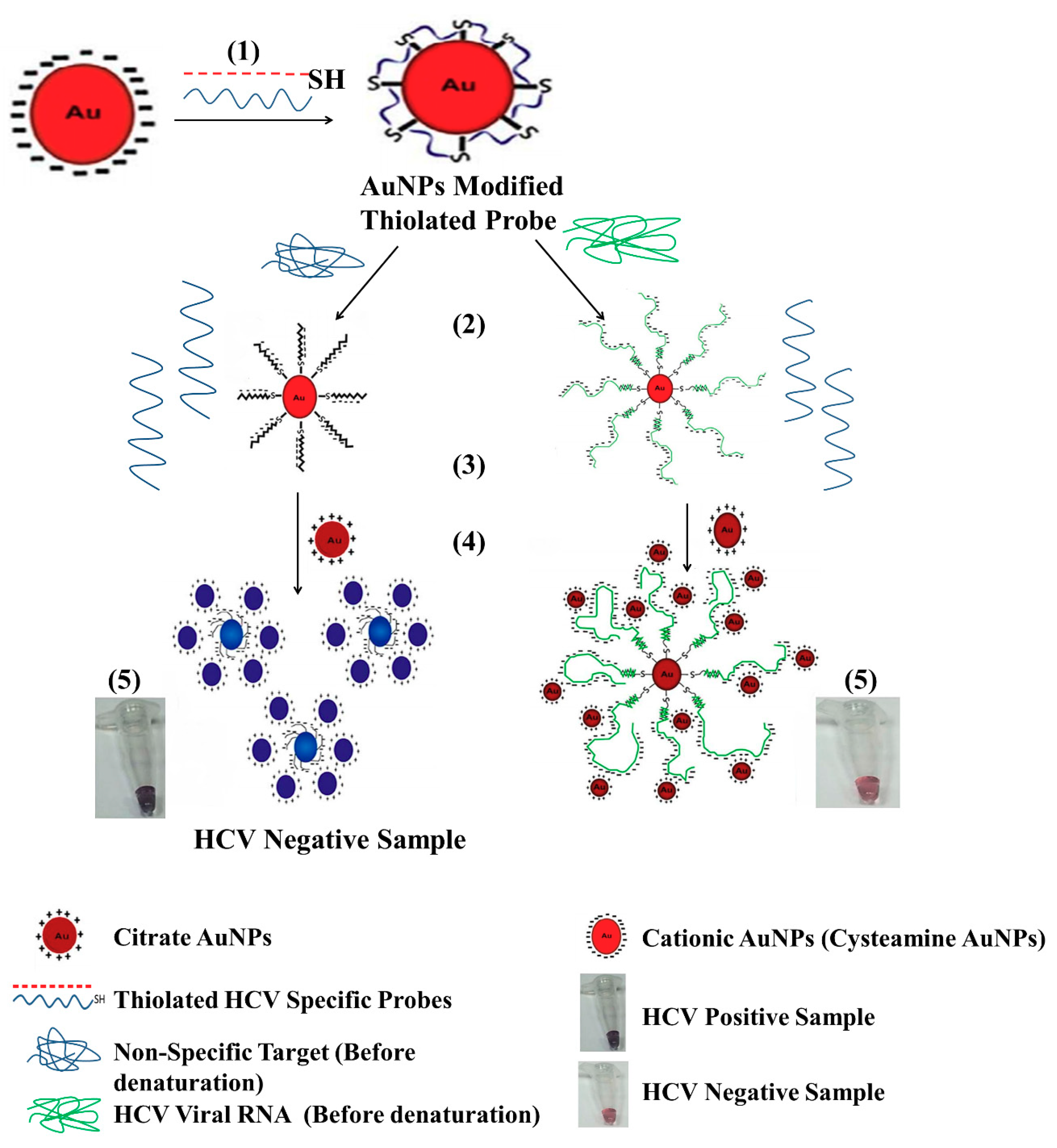
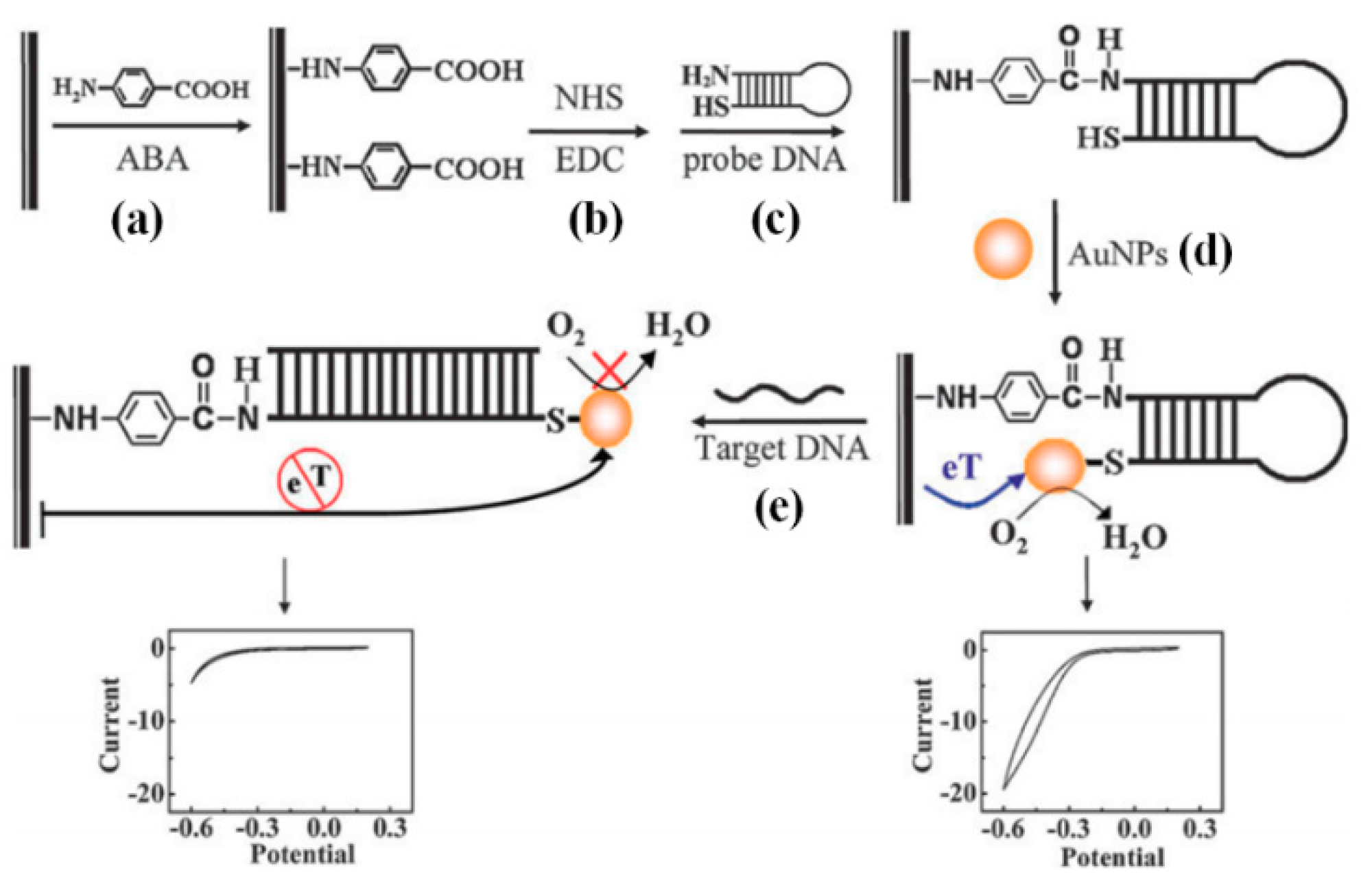
3.4.1. Gold Nanoparticle (GNP)
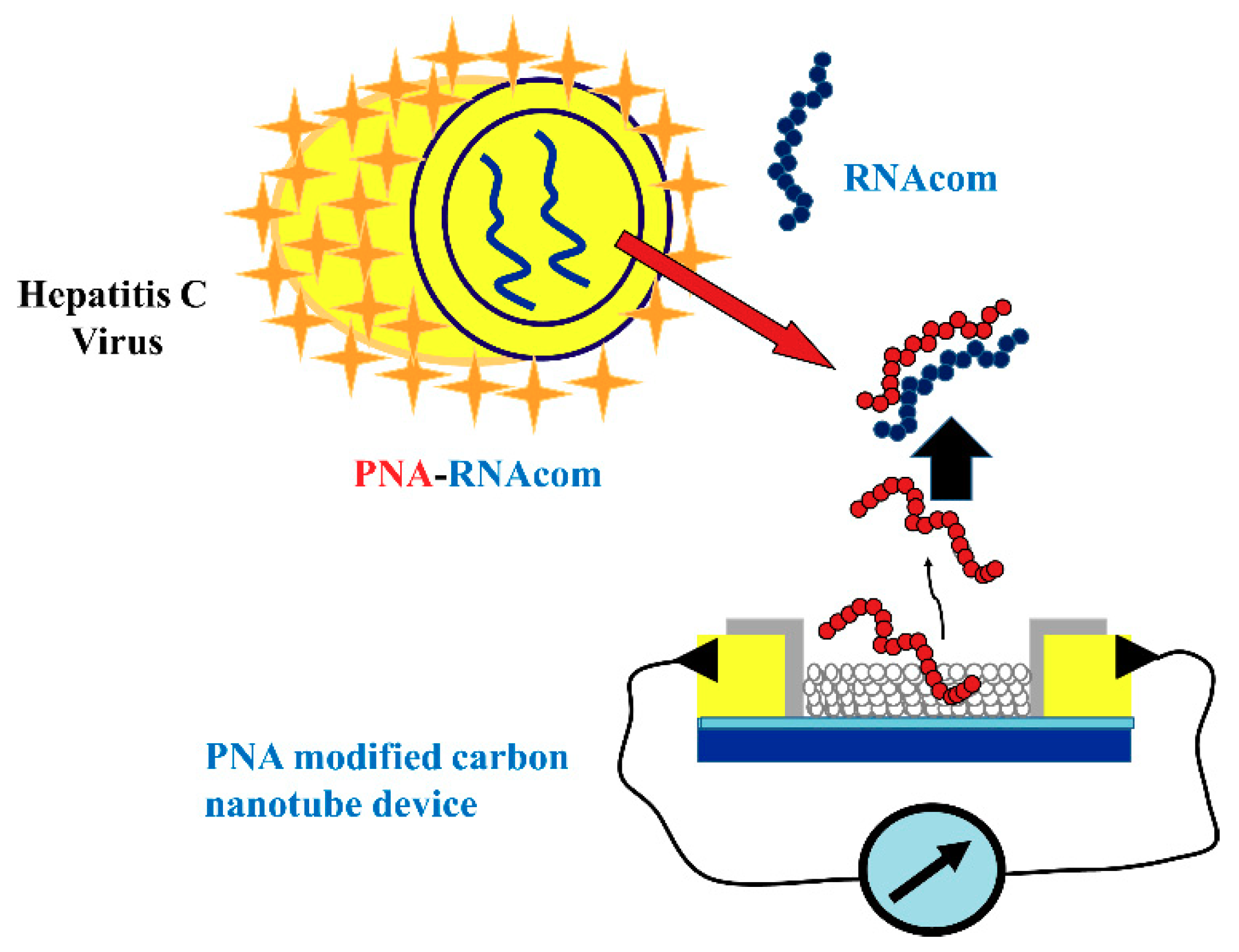
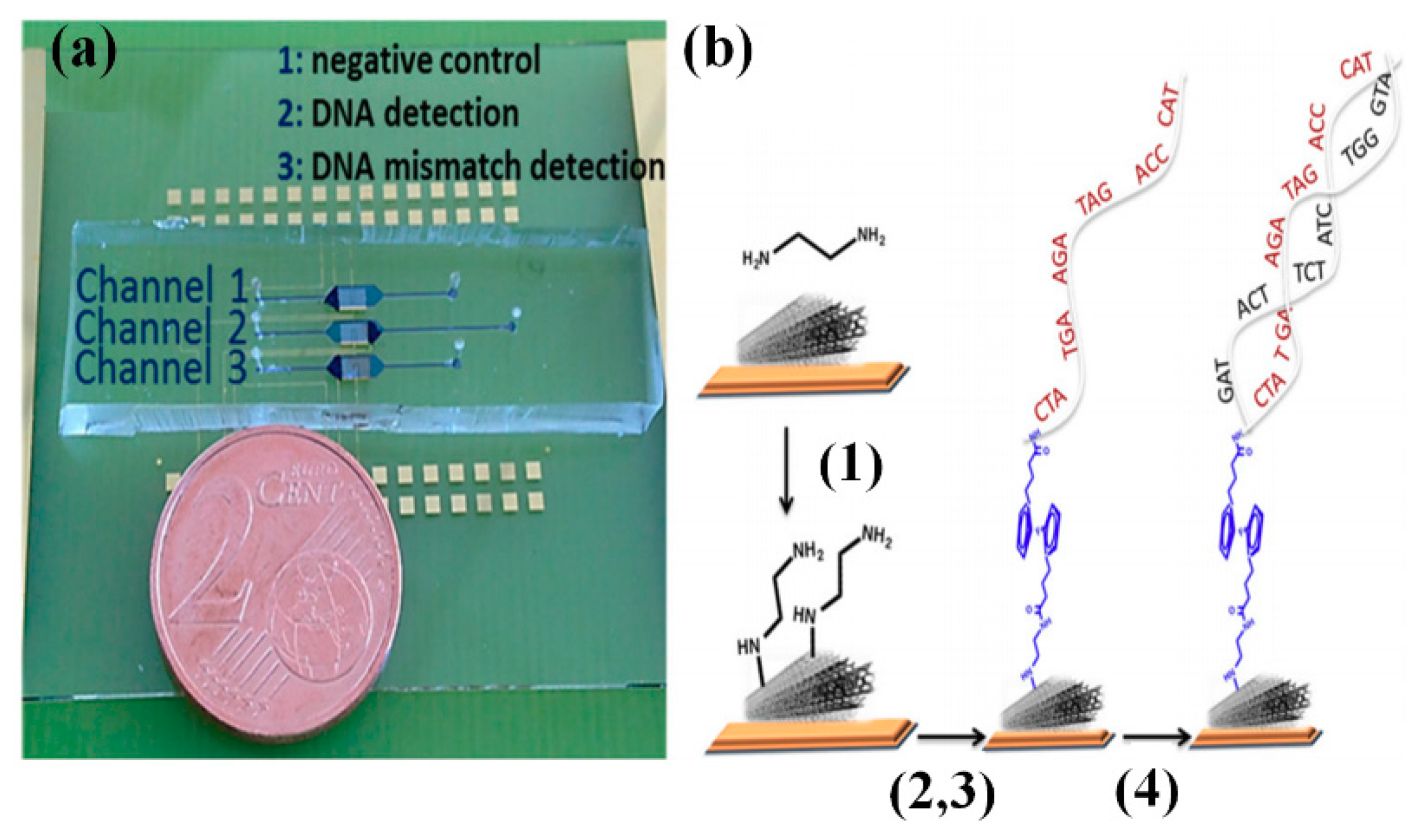
3.4.2. Carbon Nanotube and Quantum Dot
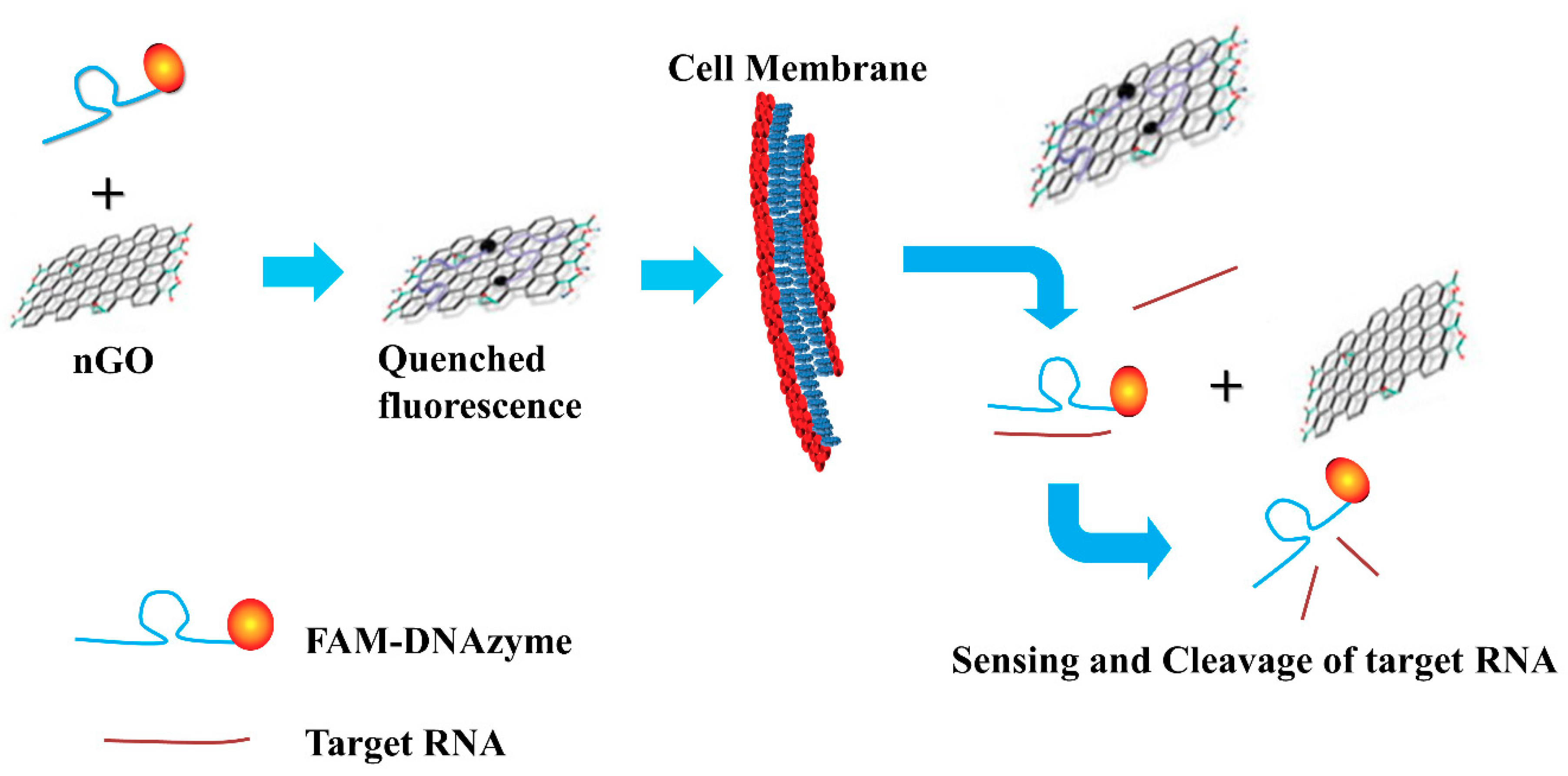
3.4.3. Graphene
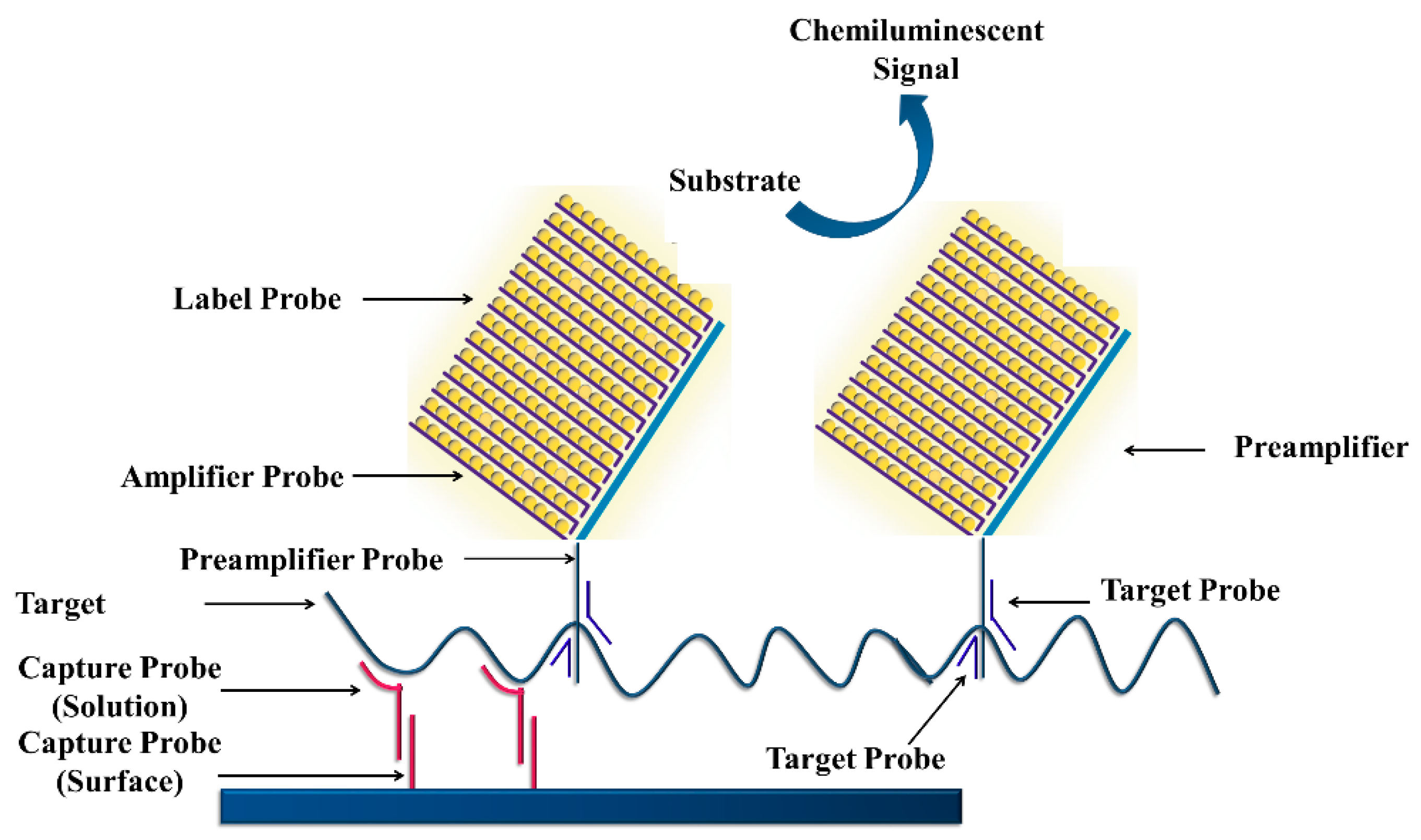
3.4.4. Branched DNA Signal Amplification Technology (bDNA)
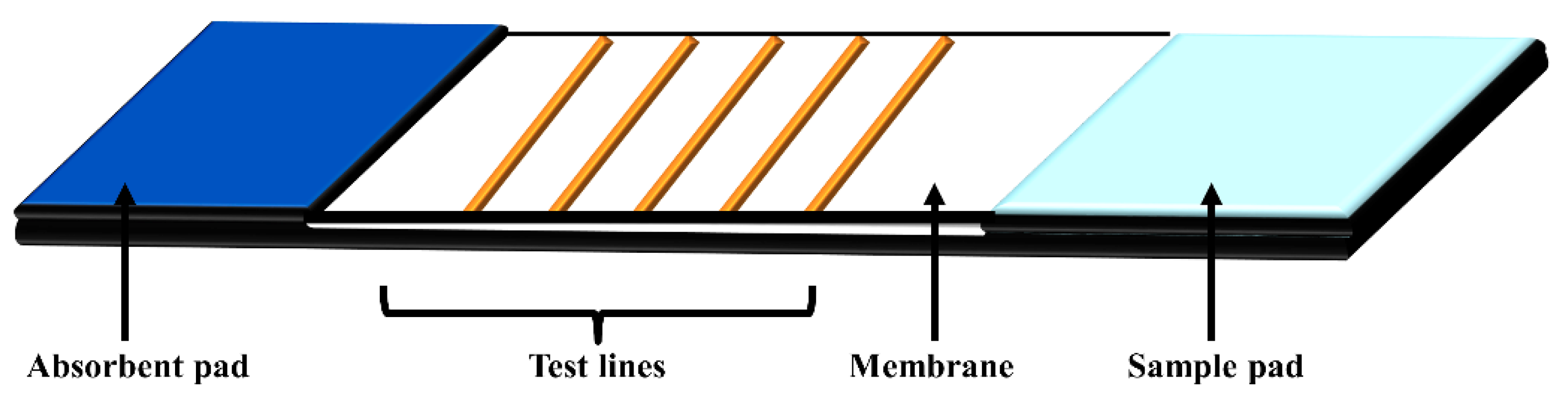
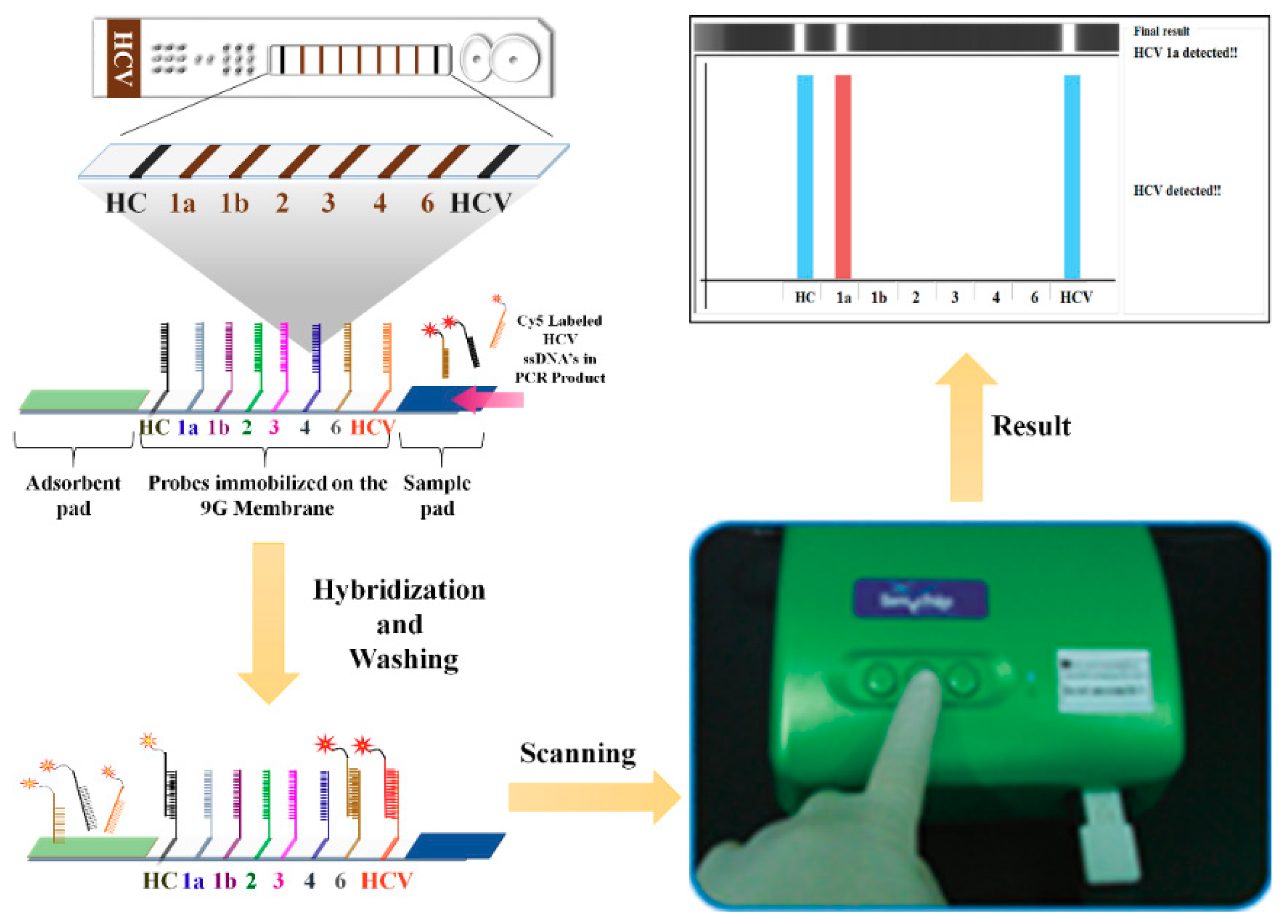
4. Lateral Flow Assays
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Gravitz, L.A. Introduction: A smouldering public-health crisis. Nature 2011, 474, S2–S4. [Google Scholar] [CrossRef] [PubMed]
- WHO. Global Hepatitis Report. 2017. Available online: https://afro.who.int/sites/default/files/2017-06/9789241565455-eng.pdf (accessed on 21 September 2018).
- GBD 2013 Mortality and Causes of Death Collaborators. Global, regional, and national age–sex specific all-cause and cause-specific mortality for 240 causes of death, 1990–2013: A systematic analysis for the Global Burden of Disease Study 2013. Lancet 2015, 385, 117–171. [Google Scholar] [CrossRef]
- Lee, M.H.; Yang, H.I.; Yuan, Y.; L’Italien, G.; Chen, C.J. Epidemiology and natural history of hepatitis C virus Infection. World J. Gastroenterol. 2014, 20, 9270–9280. [Google Scholar] [PubMed]
- WHO. Guidelines for the Care and Treatment of Persons Diagnosed with Chronic Hepatitis C Virus Infection. 2018. Available online: http://apps.who.int/iris/bitstream/handle/10665/273174/9789241550345-eng.pdf?ua=1 (accessed on 21 September 2018).
- Lavanchy, D. Evolving epidemiology of hepatitis C virus. Clin. Microbiol. Infect. 2011, 17, 107–115. [Google Scholar] [CrossRef] [PubMed]
- Pozzetto, B.; Bourlet, T.; Grattard, F.; Bonnevial, L. Structure, genomic organization, replication and variability of hepatitis C virus. Nephrol. Dial. Transplant. 1996, 11, 2–5. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Zhao, Z.; Liu, Y.; Shao, X.; Gao, Z. Comparison of three different HCV genotyping methods: Core, NS5B sequence analysis and line probe assay. Int. J. Mol. Med. 2013, 31, 347–352. [Google Scholar] [CrossRef] [PubMed]
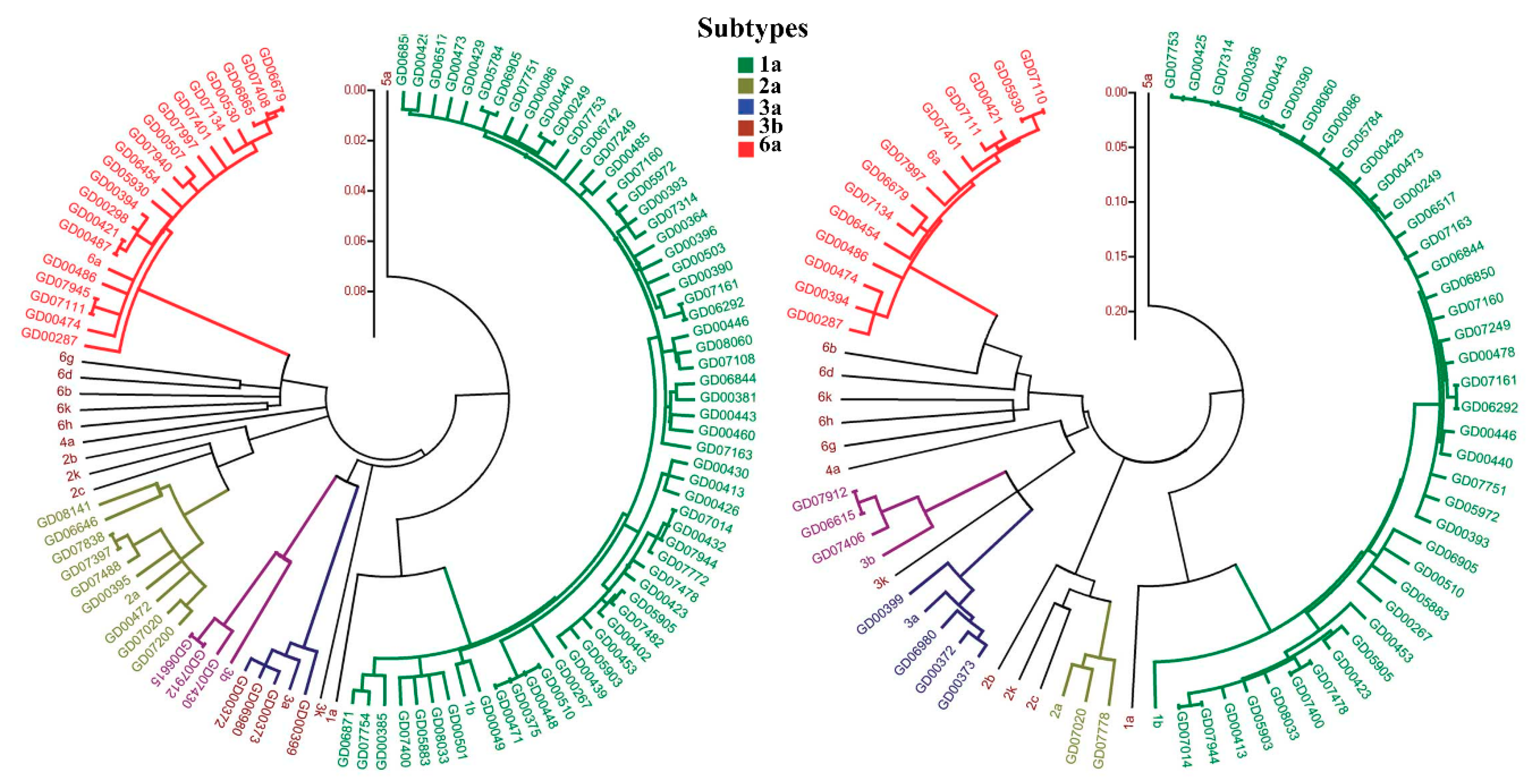
- Smith, D.B.; Bukh, J.; Kuiken, C.; Muerhoff, A.S.; Rice, C.M.; Stapleton, J.T.; Simmonds, P. Expanded Classification of Hepatitis C Virus Into 7 Genotypes and 67 Subtypes: Updated Criteria and Genotype Assignment Web Resource. Hepatology 2014, 59, 318–327. [Google Scholar] [CrossRef] [PubMed]
- Messina, J.P.; Humphreys, I.; Flaxman, A.; Brown, A.; Cooke, G.S.; Pybus, O.G.; Barnes, E. Global Distribution and Prevalence of Hepatitis C Virus Genotypes. Hepatology 2015, 61, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Wasitthankasem, R.; Vongpunsawad, S.; Siripon, N.; Suya, C.; Chulothok, P.; Chaiear, K.; Rujirojindakul, P.; Kanjana, S.; Theamboonlers, A.; Tangkijvanich, P.; et al. Genotypic Distribution of Hepatitis C Virus in Thailand and Southeast Asia. PLoS ONE 2015, 10, e0126764. [Google Scholar] [CrossRef] [PubMed]
- WHO. Guidelines for the Screening, Care and Treatment of Persons with Chronic Hepatitis C Infection. 2016. Available online: http://www.who.int/hepatitis/publications/hepatitis-c-guidelines-2016/en/ (accessed on 21 September 2018).
- European Association for Study of Liver. EASL Recommendations on Treatment of Hepatitis C 2015. J. Hepatol. 2015, 63, 199–236. [Google Scholar] [CrossRef] [PubMed]
- Mishra, P.; Murray, J.; Birnkrant, D. Direct-acting antiviral drug approvals for treatment of chronic hepatitis C virus infection: Scientific and regulatory approaches to clinical trial designs. Hepatology 2015, 62, 1298–1303. [Google Scholar] [CrossRef] [PubMed]
- European Association for Study of Liver. EASL Recommendations on Treatment of Hepatitis C 2018. J. Hepatol. 2018, 69, 461–511. [Google Scholar] [CrossRef] [PubMed]
- Gitto, S.; Gamal, N.; Andreone, P. NS5A inhibitors for the treatment of hepatitis C infection. J. Viral. Hepat. 2017, 24, 180–186. [Google Scholar] [CrossRef] [PubMed]
- LaBarre, K.R.; Hawkins, J.; Gerlach, J.; Wilmoth, A.; Beddoe, J.; Singleton, J.; Boyle, D.; Weigl, B. A simple, inexpensive device for nucleic acid amplification without electricity-toward instrument-free molecular diagnostics in low-resource settings. PLoS ONE 2011, 6, e19738. [Google Scholar] [CrossRef] [PubMed]
- Mabey, D.; Peeling, R.W.; Ustianowski, A.; Perkins, M.D. Diagnostics for the developing world. Nat. Rev. Microbiol. 2004, 2, 231–240. [Google Scholar] [CrossRef] [PubMed]
- Ngom, Y.; Guo, X.; Wang, D.B. Development and application of lateral flow test strip technology for detection of infectious agents and chemical contaminants: A review. Anal. Bioanal. Chem. 2010, 397, 1113–1135. [Google Scholar] [CrossRef] [PubMed]
- Peeling, R.W.; Mabey, D.; Herring, A.; Hook, E.W., 3rd. Why do we need quality-assured diagnostic tests for sexually transmitted infections? Nat. Rev. Microbiol. 2006, 4, 909–921. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, A.; Davidson, F.; Simmonds, P. Comparison of plasma virus loads among individuals infected with hepatitis C virus (HCV) genotypes 1, 2, and 3 by quantiplex HCV RNA assay versions 1 and 2, Roche Monitor assay, and an in-house limiting dilution method. J. Clin. Microbiol. 1997, 35, 187–192. [Google Scholar] [PubMed]
- Beld, M.; Sentjens, R.; Rebers, S.; Weegink, C.; Weel, J.; Sol, C.; Boom, R. Performance of the New Bayer VERSANT HCV RNA 3.0 assay for quantitation of hepatitis C virus RNA in plasma and serum: Conversion to international units and comparison with the Roche COBAS Amplicor HCV Monitor, Version 2.0, assay. J. Clin. Microbiol. 2002, 40, 788–793. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.H.; Liang, C.C.; Liu, C.J.; Lin, C.L.; Su, T.H.; Yang, H.C.; Chen, P.J.; Chen, D.S.; Kao, J.H. Comparison of Abbott RealTime HCV Genotype II with Versant line probe assay 2.0 for hepatitis C virus genotyping. J. Clin. Microbiol. 2015, 53, 1754–1757. [Google Scholar] [CrossRef] [PubMed]
- White, P.A.; Zhai, X.; Carter, I.; Zhao, Y.; Rawlinson, W.D. Simplified hepatitis C virus genotyping by heteroduplex mobility analysis. J. Clin. Microbiol. 2000, 38, 477–482. [Google Scholar] [PubMed]
- De Filippo, D.; Cortes-Mancera, F.; Beltran, M.; Arbelaez, M.P.; Jaramillo, S.; Restrepo, J.C.; Correa, G.; Navas, M. Molecular characterization of hepatitis c virus in multi-transfused Colombian patients. Virol. J. 2012, 9, 242. [Google Scholar] [CrossRef] [PubMed]
- Mazzuti, L.; Lozzi, M.A.; Riva, E.; Maida, P.; Falasca, F.; Antonelli, G.; Turriziani, O. Evaluation of performances of VERSANT HCV RNA 1.0 assay (kPCR) and Roche COBAS AmpliPrep/COBAS TaqMan HCV test v2.0 at low level viremia. New Microbiol. 2016, 39, 224–227. [Google Scholar] [PubMed]
- Kessler, H.H.; Cobb, B.R.; Wedemeyer, H.; Maasoumy, B.; Michel-Treil, V.; Ceccherini-Nelli, L.; Bremer, B.; Hübner, M.; Helander, A.; Khiri, H.; et al. Evaluation of the COBAS(®) AmpliPrep/COBAS(®) TaqMan(®) HCV Test, v2.0 and comparison to assays used in routine clinical practice in an international multicenter clinical trial: The ExPECT study. J. Clin. Virol. 2015, 67, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Firdaus, R.; Saha, K.; Biswas, A.; Sadhukhan, P.C. Current molecular methods for the detection of hepatitis C virus in high risk group population: A systematic review. World J. Virol. 2015, 4, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Angelika, N.; Tanya, M.F.; David, S.B. Point-of-care nucleic acid testing for infectious diseases. Trends Biotechnol. 2011, 29, 240–250. [Google Scholar]
- Freeman, W.M.; Walker, S.J.; Vrana, K.E. Quantitative RT-PCR: Pitfalls and potential. BioTechniques 1999, 26, 112–122. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Zakrajsek, B.A.; Mills, A.G.; Gorn, V.; Singer, M.J.; Reed, M.W. Quantitative reverse transcription-polymerase chain reaction to study mRNA decay: Comparison of endpoint and real-time methods. Anal. Biochem. 2000, 285, 194–204. [Google Scholar] [CrossRef] [PubMed]
- Carter, M.; Shieh, J. Guide to Research Techniques in Neuroscience, 2nd ed.; Academic Press: Cambridge, MA, USA, 2015; pp. 219–237. ISBN 9780128005118. [Google Scholar]
- Young, K.K.; Resnick, R.M.; Myers, T.W. Detection of hepatitis C virus RNA by a combined reverse transcription-polymerase chain reaction assay. J. Clin. Microbiol. 1993, 31, 882–886. [Google Scholar] [PubMed]
- Bustin, S.A. Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): Trends and problems. J. Mol. Endocrinol. 2002, 29, 23–39. [Google Scholar] [CrossRef] [PubMed]
- Meng, S.; Li, J. A novel duplex real-time reverse transcriptase-polymerase chain reaction assay for the detection of hepatitis C viral RNA with armored RNA as internal control. Virol. J. 2010, 7, 117. [Google Scholar] [CrossRef] [PubMed]
- Sung, C.L.; Anisha, A.; Nitta, L.; Jeff, L.; Jian, Q.; Yang, S.S.; Karen, G.; Maurice, R. Improved Version 2.0 Qualitative and Quantitative AMPLICOR Reverse Transcription-PCR Tests for Hepatitis C Virus RNA: Calibration to International Units, Enhanced Genotype Reactivity, and Performance Characteristics. J. Clin. Microbiol. 2000, 38, 4171–4179. [Google Scholar]
- Robbins, D.J.; Pasupuleti, V.; Cuan, J.; Chiang, C.S. Reverse transcriptase PCR quantitation of hepatitis C virus. Clin. Lab. Sci. 2000, 13, 23–30. [Google Scholar] [PubMed]
- Nakatani, S.M.; Santos, C.A.; Riediger, I.N.; Krieger, M.A.; Duarte, C.A.; Lacerda, M.A.; Biondo, A.W.; Carrilho, F.J.; Ono-Nita, S.K. Development of hepatitis C virus genotyping by real-time PCR based on the NS5B region. PLoS ONE 2010, 5, e10150. [Google Scholar] [CrossRef]
- Gianluca, B.L.; Rizzotti, F.D.; Sandra, T. Development of Reverse Transcription (RT)-PCR and Real-Time RT-PCR Assays for Rapid Detection and Quantification of Viable Yeasts and Molds Contaminating Yogurts and Pasteurized Food Products. Appl. Environ. Microbiol. 2003, 69, 4116–4122. [Google Scholar]
- Ramakers, C.; Ruijter, J.M.; Deprez, R.H.; Moorman, A.F. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci. Lett. 2003, 339, 62–66. [Google Scholar] [CrossRef]
- Shiao, Y.H. A new reverse transcription-polymerase chain reaction method for accurate quantification. BMC Biotechnol. 2003, 3, 22. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Deepak, S.A.; Kottapalli, K.R.; Rakwal, R.; Oros, G.; Rangappa, K.S.; Iwahashi, H.; Masuo, Y.; Agrawal, G.K. Real-Time PCR: Revolutionizing detection and expression analysis of genes. Curr. Genom. 2007, 8, 234–251. [Google Scholar] [CrossRef]
- Bassam, B.J.; Allen, T.; Flood, S.; Stevens, J.; Wyatt, P.; Livak, K.J. Nucleic acid sequence detection systems: Revolutionary automation for monitoring and reporting PCR products. Australas. Biotechnol. 1996, 6, 285–294. [Google Scholar]
- Higuchi, R.; Fockler, C.; Dollinger, G.; Watson, R. Kinetic PCR analysis: Real-time monitoring of DNA amplification reactions. Biotechnology 1993, 11, 1026–1030. [Google Scholar] [CrossRef] [PubMed]
- Cobb, B.; Pockros, P.J.; Vilchez, R.A.; Vierling, J.M. HCV RNA viral load assessments in the era of direct-acting antivirals. Am. J. Gastroenterol. 2013, 108, 471–475. [Google Scholar] [CrossRef] [PubMed]
- Vermehren, J.; Kau, A.; Gärtner, B.C.; Göbel, R.; Zeuzem, S.; Sarrazin, C. Differences between Two Real-Time PCR-Based Hepatitis C Virus (HCV) Assays (RealTime HCV and Cobas AmpliPrep/Cobas TaqMan) and One Signal Amplification Assay (Versant HCV RNA 3.0) for RNA Detection and Quantification. J. Clin. Microbiol. 2008, 46, 3880–3891. [Google Scholar] [CrossRef] [PubMed]
- Cobb, B.; Heilek, G.; Vilchez, R.A. Molecular diagnostics in the management of chronic hepatitis C: Key considerations in the era of new antiviral therapies. BMC Infect. Dis. 2014, 14, S8. [Google Scholar] [CrossRef] [PubMed]
- Laperche, S.; Bouchardeau, F.; André-Garnier, E.; Thibault, V.; Roque-Afonso, A.M.; Trimoulet, P.; Colimon, R.; Duverlie, G.; Leguillou-Guillemette, H.; Lunel, F.; et al. Interpretation of Real-Time PCR Results for Hepatitis C Virus RNA When Viral Load Is Below Quantification Limits. J. Clin. Microbiol. 2011, 49, 1113–1115. [Google Scholar] [CrossRef] [PubMed]
- Sonia, V.M.; Pablo, R.B.; Ardizone-Jiménez, D.M.; Jesus, T.; Guillermo, C.; Jorge, V.; María, A.J.; Ana, A.; Salvador, R. Evaluation of dried blood spot samples for screening of hepatitis C and human immunodeficiency virus in a real-world setting. Sci. Rep. 2018, 8, 1858. [Google Scholar] [PubMed]
- Wong, M.L.; Medrano, J.F. Real-time PCR for mRNA quantitation. BioTechniques 2005, 39, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Zhou, J.; Zheng, Y.; Gamson, A.S.; Roembke, B.T.; Nakayama, S.; Sintim, H.O. Isothermal amplified detection of DNA and RNA. Mol. Biosyst. 2014, 10, 970–1003. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Manzano, J.; Karymov, M.A.; Begolo, S.; Selck, D.A.; Zhukov, D.V.; Jue, E.; Ismagilov, R.F. Reading out single-molecule digital RNA and DNA isothermal amplification in nanoliter volumes with unmodified camera phones. ACS Nano 2016, 10, 3102–3113. [Google Scholar] [CrossRef] [PubMed]
- Compton, J. Nucleic acid sequence-based amplification. Nature 1991, 350, 91–92. [Google Scholar] [CrossRef] [PubMed]
- Morré, S.A.; Sillekens, P.; Jacobs, M.V.; van Aarle, P.; de Blok, S.; van Gemen, B.; Walboomers, J.M.; Meijer, C.J.; van den Brule, A.J. RNA amplification by nucleic acid sequence-based amplification with an internal standard enables reliable detection of Chlamydia trachomatis in cervical scrapings and urine samples. J. Clin. Microbiol. 1996, 34, 3108–3214. [Google Scholar] [PubMed]
- Kievits, T.; van Gemen, B.; van Strijp, D.; Schukkink, R.; Dircks, M.; Adriaanse, H.; Malek, L.; Sooknanan, R.; Lens, P. NASBA isothermal enzymatic in vitro nucleic acid amplification optimized for the diagnosis of HIV-1 infection. J. Virol. Methods 1991, 35, 273–286. [Google Scholar] [CrossRef]
- Brink, A.A.; Vervoort, M.B.; Middeldorp, J.M.; Meijer, C.J.; van den Brule, A.J. Nucleic acid sequence-based amplification, a new method for analysis of spliced and unspliced epstein-barr virus latent transcripts, and its comparison with reverse transcriptase PCR. J. Clin. Microbiol. 1998, 36, 3164–3169. [Google Scholar] [PubMed]
- Deiman, B.; van, A.P.; Sillekens, P. Characteristics and applications of nucleic acid sequence-based amplification (NASBA). Mol. Biotechnol. 2002, 20, 163. [Google Scholar] [CrossRef]
- Polstra, A.M.; Goudsmit, J.; Cornelissen, M. Development of real-time NASBA assays with molecular beacon detection to quantify mRNA coding for HHV-8 lytic and latent genes. BMC Infect. Dis. 2002, 2, 18. [Google Scholar] [CrossRef]
- Guichon, A.; Chiparelli, H.; Martinez, A.; Rodriguez, C.; Trento, A.; Russi, J.C.; Carballal, G. Evaluation of a new NASBA assay for the qualitative detection of hepatitis C virus based on the NucliSense Basic Kit reagents. J. Clin. Virol. 2004, 29, 84–91. [Google Scholar] [CrossRef]
- Giachetti, C.J.; Linnen, M.; Kolk, D.P.; Dockter, J.; Gillotte-Taylor, K.; Park, M.; Ho-Sing-Loy, M.; McCormick, M.K.; Mimms, L.T.; McDonough, S.H. Highly Sensitive Multiplex Assay for Detection of Human Immunodeficiency Virus Type 1 and Hepatitis C Virus RNA. J. Clin. Microbiol. 2002, 40, 2408–2419. [Google Scholar] [CrossRef] [PubMed]
- Karami, A.; Gill, P.; Motamedi, M.H.K.; Saghafinia, M. A Review of the Current Isothermal Amplification Techniques: Applications, Advantages and Disadvantages. J. Glob. Infect. Dis. 2011, 3, 293–302. [Google Scholar]
- Comanor, L.; Anderson, F.; Ghany, M.; Perrillo, R.; Heathcote, E.J.; Sherlock, C.; Zitron, I.; Hendricks, D.; Gordon, S.C. Transcription-mediated amplification is more sensitive than conventional PCR-based assays for detecting residual serum HCV RNA at end of treatment. Am. J. Gastroenterol. 2001, 96, 2968–2972. [Google Scholar] [CrossRef] [PubMed]
- Sarrazin, C.; Teuber, G.; Kokka, R.; Rabenau, H.; Zeuzem, S. Detection of residual hepatitis C virus RNA by transcription-mediated amplification in patients with complete virologic response according to polymerase chain reaction-based assays. Hepatology 2000, 32, 818–823. [Google Scholar] [CrossRef] [PubMed]
- Rao, V.; Fabrizi, F.; Pennell, P.; Schiff, E.; de Medina, M.; Lane, J.R.; Martin, P.; Ivor, L. Improved detection of hepatitis C virus infection by transcription-mediated amplification technology in dialysis population. Ren. Fail. 2010, 32, 721–726. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, W.P.; Dries, V.; Herrmann, E.; Gärtner, B.; Zeuzem, S.; Sarrazin, C. Comparison of transcription mediated amplification (TMA) and reverse transcription polymerase chain reaction (RT-PCR) for detection of hepatitis C virus RNA in liver tissue. J. Clin. Virol. 2005, 32, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Langabeer, S.E.; Gale, R.E.; Harvey, R.C.; Cook, R.W.; Mackinnon, S.; Linch, D.C. Transcription-mediated amplification and hybridisation protection assay to determine BCR-ABL transcript levels in patients with chronic myeloid leukaemia. Leukemia 2002, 16, 393–399. [Google Scholar] [CrossRef] [PubMed]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, E63. [Google Scholar] [CrossRef] [PubMed]
- Notomi, T.; Mori, Y.; Tomita, N.; Kanda, H. Loop-mediated isothermal amplification (LAMP): Principle, features, and future prospects. J. Microbiol. 2015, 53, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Kumar, Y.; Bansal, S.; Jaiswal, P. Loop-Mediated Isothermal Amplification (LAMP): A Rapid and Sensitive Tool for Quality Assessment of Meat Products. Compr. Rev. Food Sci. Food Saf. 2017, 16, 1359–1378. [Google Scholar] [CrossRef]
- Wong, Y.P.; Othman, S.; Lau, Y.L.; Radu, S.; Chee, H.Y. Loop-mediated isothermal amplification (LAMP): A versatile technique for detection of micro-organisms. J. Appl. Microbiol. 2018, 124, 626–643. [Google Scholar] [CrossRef] [PubMed]
- Parida, M.M.; Sannarangaiah, S.; Dash, P.K.; Rao, P.V.L.; Morita, K. Loop mediated isothermal amplification (LAMP): A new generation of innovative gene amplification technique; perspectives in clinical diagnosis of infectious diseases. Rev. Med. Virol. 2008, 18, 407–421. [Google Scholar] [CrossRef] [PubMed]
- Kargar, M.; Askari, A.; Doosti, A.; Ghorbani-Dalini, S. Loop-Mediated Isothermal Amplification Assay for Rapid Detection of Hepatitis C virus. Indian J. Virol. 2012, 23, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Nyan, D.C.; Swinson, K.L. A method for rapid detection and genotype identification of hepatitis C virus 1–6 by one-step reverse transcription loop-mediated isothermal amplification. Int. J. Infect. Dis. 2016, 43, 30–36. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Fang, M.X.; Li, J.; Lou, G.Q.; Lu, H.J.; Wu, N.P. Detection of hepatitis C virus by an improved loop-mediated isothermal amplification assay. Arch. Virol. 2011, 156, 1387–1396. [Google Scholar] [CrossRef] [PubMed]
- Gong, P.; Zhang, T.; Chen, F.; Wang, L.; Jin, S.; Bai, X. Advances in loop-mediated isothermal amplification: Integrated with several point-of-care diagnostic methods. Anal. Methods 2014, 6, 7585–7589. [Google Scholar] [CrossRef]
- Lizardi, P.M.; Huang, X.; Zhu, Z.; Bray-Ward, P.; Thomas, D.C.; Ward, D.C. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat Genet. 1998, 19, 225–232. [Google Scholar] [CrossRef] [PubMed]
- Ali, M.M.; Li, F.; Zhang, Z.; Zhang, K.; Kang, D.K.; Ankrum, J.A.; Le, X.C.; Zhao, W. Rolling circle amplification: A versatile tool for chemical biology, materials science and medicine. Chem. Soc. Rev. 2014, 43, 3324. [Google Scholar] [CrossRef] [PubMed]
- Ji, M.; Hu, G.; Zheng, Y.; Gu, D.; Long, J.; Lu, W.; He, J.; Tan, S.; Shi, L.; Liu, C.; et al. Surface plasmon resonance technology combined with rolling circle amplification for detection of hepatitis C virus. J Shanghai Jiaotong Univ. 2012, 32, 693–705. [Google Scholar]
- Kintzios, S.; Bem, F.; Mangana, O.; Nomikou, K.; Markoulatos, P.; Alexandropoulos, N.; Fasseas, C.; Arakelyan, V.; Petrou, A.L.; et al. Study on the mechanism of Bioelectric Recognition Assay: Evidence for immobilized cell membrane interactions with viral fragments. Biosens. Bioelectron. 2004, 20, 907–916. [Google Scholar] [CrossRef] [PubMed]
- Saeui, C.T.; Mathew, M.P.; Liu, L.; Urias, E.; Yarema, K.J. Cell Surface and Membrane Engineering: Emerging Technologies and Applications. J. Funct. Biomater. 2015, 6, 454–485. [Google Scholar] [CrossRef] [PubMed]
- Kintzios, S.; Pistola, E.; Panagiotopoulos, P.; Bomsel, M.; Alexandropoulos, N.; Bem, F.; Ekonomou, G.; Biselis, J.; Levin, R. Bioelectric recognition assay (BERA). Biosens. Bioelectron. 2001, 16, 325–336. [Google Scholar] [CrossRef]
- Yao, C.; Zhu, T.; Tang, J.; Wu, R.; Chen, Q.; Chen, M.; Zhang, B.; Huang, J.; Fu, W. Hybridization assay of hepatitis B virus by QCM peptide nucleic acid biosensor. Biosens. Bioelectron. 2008, 23, 879–885. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Mao, Q.; Zhang, X.; Jiang, T.; Chen, M.; Yu, F.; Fu, W. A novel piezoelectric quartz micro-array immunosensor based on self-assembled monolayer for determination of human chorionic gonadotropin. Biosens. Bioelectron. 2004, 19, 711–720. [Google Scholar] [CrossRef]
- Skládal, P.; dos Santos Riccardi, C.; Yamanaka, H.; da Costa, P.I. Piezoelectric biosensors for real-time monitoring of hybridization and detection of hepatitis C virus. J. Virol. Methods 2004, 117, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Belluzo, M.S.; Ribone, M.E.; Lagier, C.M. Assembling amperometric biosensors for clinical diagnostics. Sensors 2008, 8, 1366–1399. [Google Scholar] [CrossRef] [PubMed]
- Riccardi, C.S.; Dahmouche, K.; Santilli, C.V.; da Costa, P.I.; Yamanaka, H. Immobilization of streptavidin in sol-gel films: Application on the diagnosis of hepatitis C virus. Talanta 2006, 70, 637–643. [Google Scholar] [CrossRef] [PubMed]
- Uliana, C.V.; Riccardi, C.S.; Tognolli, J.O.; Yamanaka, H. Optimization of an amperometric biosensor for the detection of hepatitis C virus using fractional factorial designs. J. Braz. Chem. 2008, 19, 782–787. [Google Scholar] [CrossRef]
- Dos Santos, R.C.; Kranz, C.; Kowalik, J.; Yamanaka, H.; Mizaikoff, B.; Josowicz, M. Label-free DNA detection of hepatitis C virus based on modified conducting polypyrrole films at microelectrodes and atomic force microscopy tip-integrated electrodes. Anal. Chem. 2008, 80, 237–245. [Google Scholar]
- Tripp, R.A.; Alvarez, R.; Anderson, B.; Jones, L.; Weeks, C.; Chen, W. Bioconjugated nanoparticle detection of respiratory syncytial virus infection. Int. J. Nanomed. 2007, 2, 117–124. [Google Scholar] [CrossRef]
- Tang, S.; Zhao, J.; Storhoff, J.J.; Norris, P.J.; Little, R.F.; Yarchoan, R.; Stramer, S.L.; Patno, T.; Domanus, M.; Dhar, A.; et al. Nanoparticle-Based biobarcode amplifi cation assay (BCA) for sensitive and early detection of human immunodefi ciency type 1 capsid (p24) antigen. J. Acquir. Immune Defic. Syndr. 2007, 46, 231–237. [Google Scholar] [CrossRef] [PubMed]
- Wei, F.; Lillehoj, P.B.; Ho, C.M. DNA Diagnostics: Nanotechnology-enhanced Electrochemical Detection of Nucleic Acids. Pediatr Res. 2010, 67, 458–468. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Jain, P.K.; EI-Sayed, I.H.; El-Sayed, M.A. Gold nanoparticles: Interesting optical properties and recent applications in cancer diagnostics and therapy. Nanomedicine 2007, 2, 681–693. [Google Scholar] [CrossRef] [PubMed]
- Liandris, E.; Gazouli, M.; Andreadou, M.; Comor, M.; Abazovic, N.; Sechi, L.A.; Ikonomopoulos, J. Direct detection of unamplified DNA from pathogenic mycobacteria using DNA-derivatized gold nanoparticles. Microbiol. Methods 2009, 78, 260–264. [Google Scholar] [CrossRef] [PubMed]
- Draz, M.S.; Shafiee, H. Applications of gold nanoparticles in virus detection. Theranostics 2018, 8, 1985–2017. [Google Scholar] [CrossRef] [PubMed]
- Elghanian, R.; Storhoff, J.J.; Mucic, R.C.; Letsinger, R.L.; Mirkin, C.A. Selective colorimetric detection of polynucleotides based on the distance-dependent optical properties of gold nanoparticles. Science 1997, 277, 1078–1081. [Google Scholar] [CrossRef] [PubMed]
- Larguinho, M.; Canto, R.; Cordeiro, M.; Pedrosa, P.; Fortuna, A.; Vinhas, R.; Baptista, P.V. Gold nanoprobe-based non-crosslinking hybridization for molecular diagnostics. Expert Rev. Mol. Diagn. 2015, 15, 1355–1368. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Hosokawa, K.; Maeda, M. Rapid Aggregation of gold nanoparticles induced by non-cross-linking DNA hybridization. J. Am. Chem. Soc. 2003, 125, 8102–8103. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.F.; Pang, D.W.; Zhang, Z.L.; Zheng, H.Z.; Cao, J.P.; Shen, J.T. Visual gene diagnosis of HBV and HCV based on nanoparticle probe amplification and silver staining enhancement. J. Med. Virol. 2003, 70, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Shawky, S.M.; Bald, D.; Azzazy, H.M. Direct detection of unamplified hepatitis C virus RNA using unmodified gold nanoparticles. Clin. Biochem. 2010, 43, 1163–1168. [Google Scholar] [CrossRef] [PubMed]
- Shawky, S.M.; Awad, A.M.; Allam, W.; Alkordi, M.H.; El-Khamisy, S.F. Gold aggregating gold: A novel nanoparticle biosensor approach for the direct quantification of hepatitis C virus RNA in clinical samples. Biosens. Bioelectron. 2017, 92, 349–356. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Wu, P.; Zhang, H.; Cai, C. Catalytic signal amplification of gold nanoparticles combining with conformation-switched hairpin DNA probe for hepatitis C virus quantification. Chem. Commun. 2012, 48, 7877–7879. [Google Scholar] [CrossRef] [PubMed]
- Dastagir, T.; Forzani, E.S.; Zhang, R.; Amlani, I.; Nagahara, L.A.; Tsui, R.; Tao, N. Electrical detection of hepatitis C virus RNA on single wall carbon nanotube-field effect transistors. Analyst 2007, 132, 738–740. [Google Scholar] [CrossRef] [PubMed]
- Zribi, B.; Roy, E.; Pallandre, A.; Chebil, S.; Koubaa, M.; Mejri, N.; Magdinier, G.H.; Sola, C.; Korri-Youssoufi, H.; Haghiri-Gosnet, A.M. A microfluidic electrochemical biosensor based on multiwall carbon nanotube/ferrocene for genomic DNA detection of Mycobacterium tuberculosis in clinical isolates. Biomicrofluidics 2016, 10, 014115. [Google Scholar] [CrossRef] [PubMed]
- Zribi, B.; Castro-Arias, J.; Decanini, D.; Gogneau, N.; Dragoe, D.; Cattoni, A.; Ouerghi, A.; Korri-Youssoufib, H.; Haghiri-Gosnet, A. Large area graphene nanomesh: An artificial platform for edge-electrochemical biosensing at the sub-attomolar level. Nanoscale 2016, 8, 15479–15485. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Ryoo, S.R.; Na, H.K.; Kim, Y.K.; Choi, B.S.; Lee, Y.; Kim, D.E.; Min, D.H. Deoxyribozyme-loaded nano-graphene oxide for simultaneous sensing and silencing of the hepatitis C virus gene in liver cells. Chem. Commun. 2013, 49, 8241–8243. [Google Scholar] [CrossRef] [PubMed]
- Nolte, F.S. Branched DNA signal amplification for direct quantitation of nucleic acid sequences in clinical specimens. Adv. Clin. Chem. 1999, 33, 201–232. [Google Scholar]
- Tsongalis, G.J. Branched DNA Technology in Molecular Diagnostics. Am. J. Clin. Pathol. 2006, 126, 448–453. [Google Scholar] [CrossRef] [PubMed]
- Lunel, F.; Cresta, P.; Vitour, D.; Payan, C.; Dumont, B.; Frangeul, L.; Reboul, D.; Brault, C.; Piette, J.C.; Huraux, J.M. Comparative evaluation of Hepatitis C virus RNA quantitation by branched DNA, NASBA, and monitor assays. Hepatology 1999, 29, 528–535. [Google Scholar] [CrossRef] [PubMed]
- Pieretti, M. Molecular Diagnostics: Techniques and Applications for the Clinical Laboratory, 1st ed.; Academic Press: Cambridge, MA, USA, 2010; pp. 15–19. ISBN 978-0-12-369428-7. [Google Scholar]
- Sarrazin, C.; Gärtner, B.C.; Sizmann, D.; Babiel, R.; Mihm, U.; Hofmann, W.P.; von Wagner, M.; Zeuzem, S. Comparison of conventional PCR with real-time PCR and branched DNA-based assays for hepatitis C virus RNA quantification and clinical significance for genotypes 1 to 5. J. Clin. Microbiol. 2006, 44, 729–737. [Google Scholar] [CrossRef] [PubMed]
- Stuyver, L.; Rossau, R.; Wyseur, A.; Duhamel, M.; Vanderborght, B.; Van Heuverswyn, H.; Maertens, G. Typing of hepatitis C virus isolates and characterization of new subtypes using a line probe assay. J. Gen. Virol. 1993, 74, 1093–102. [Google Scholar] [CrossRef] [PubMed]
- Verbeeck, J.; Stanley, M.J.; Shieh, J.; Celis, L.; Huyck, E.; Wollants, E.; Morimoto, J.; Farrior, A.; Sablon, E.; Jankowski-Hennig, M.; et al. Evaluation of Versant Hepatitis C Virus Genotype Assay (LiPA) 2.0. J. Clin. Microbiol. 2008, 46, 1901–1906. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.; Cong, X.; Du, S.; Fei, R.; Rao, H.; Weicorresponding, L. Performance Comparison of the Versant HCV Genotype 2.0 Assay (LiPA) and the Abbott Realtime HCV Genotype II Assay for Detecting Hepatitis C Virus Genotype 6. J. Clin. Microbiol. 2014, 52, 3685–3692. [Google Scholar] [CrossRef] [PubMed]
- Chueca, N.; Rivadulla, I.; Lovatti, R.; Reina, G.; Blanco, A.; Fernandez-Caballero, J.A.; Cardeñoso, L.; Rodriguez-Granjer, J.; Fernandez-Alonso, M.; Aguilera, A.; et al. Using NS5B sequencing for hepatitis C virus genotyping reveals discordances with commercial platforms. PLoS ONE 2016, 11, e0153754. [Google Scholar] [CrossRef] [PubMed]
- Chantratita, W.; Song, K.S.; GunHo, C.; Pongthanapisith, V.; Thongbaiphet, N.; Wongtabtim, G.; Pasomsub, E.; Angkanavin, K.; Nimse, S.B.; Sonawane, M.D.; et al. 6 HCV genotyping 9G test and its comparison with VERSANT HCV genotype 2.0 assay (LiPA) for the hepatitis C virus genotyping. J. Virol. Methods 2017, 239, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Song, K.; Nimse, S.B.; Kim, J.; Sayyed, D.R.; Kim, T. New platform for convenient genotyping system. Chem. Commun. 2013, 49, 2661–2663. [Google Scholar] [CrossRef] [PubMed]
- Nimse, S.B.; Song, K.; Kim, J.; Kim, H.; Nguyen, V.; Ta, V.; Kim, T. Generalized probe selection method for DNA Chips. Chem. Commun. 2011, 47, 12444–12446. [Google Scholar] [CrossRef] [PubMed]
- Chantratita, W.; Song, K.S.; Nimse, S.B.; Pongthanapisith, V.; Thongbaiphet, N.; Wongtabtim, G.; Pasomsub, E.; Angkanavin, K.; Sonawane, M.D.; Warkad, S.D.; et al. 6 HCV Genotyping 9G test for HCV 1a, 1b, 2, 3, 4 and 6 (6a, 6f, 6i and 6n) with high accuracy. J. Virol. Methods. 2017, 246, 95–99. [Google Scholar] [CrossRef] [PubMed]
- Nadarajah, R.; Khan, G.Y.; Miller, S.A.; Brooks, G.F. Evaluation of a new-generation line-probe assay that detects 5′ untranslated and core regions to genotype and subtype hepatitis C virus. Am. J. Clin. Pathol. 2007, 128, 300–304. [Google Scholar] [CrossRef] [PubMed]
- Warkad, S.D.; Nimse, S.B.; Song, K.; Chantratita, W.; Pongthanapisith, V.; Nawale, L.U.; Kim, T. Performance of 6 HCV genotyping 9G test for HCV genotyping in clinical samples. Virol. J. 2018, 15, 107. [Google Scholar] [CrossRef] [PubMed]
- Swellam, M.M.; Mahmoud, M.S.; Ali, A.A. Diagnosis of hepatitis C virus infection by enzyme-linked immunosorbent assay and reverse transcriptase-nested polymerase chain reaction: A comparative evaluation. IUBMB Life 2011, 63, 430–434. [Google Scholar] [CrossRef] [PubMed]
- Mohammadi-Yeganeh, S.; Paryan, M.; Mirab Samiee, S.; Kia, V.; Rezvan, H. Molecular beacon probes–base multiplex NASBA Real-time for detection of HIV-1 and HCV. Iran J. Microbiol. 2012, 4, 47–54. [Google Scholar] [PubMed]
- Loens, K.; Ursi, D.; Goossens, H.; Ieven, M. Medical Biomethods Handbook; Walker, J.M. Medical Biomethods Handbook; Walker, J.M., Repley, R., Eds.; Humana press, Inc.: Totowa, NJ, USA, 2005. [Google Scholar]
- Wang, D.; Huo, G.; Wang, F.; Li, Y.; Ren, D. Drawback of loop-mediated isothermal amplification. Afr. J. Food Sci. 2008, 2, 083–086. [Google Scholar]
- Gu, L.; Yan, W.; Liu, L.; Wang, S.; Zhang, X.; Lyu, M. Research Progress on Rolling Circle Amplification (RCA)-Based Biomedical Sensing. Pharmaceuticals 2018, 11, 35. [Google Scholar] [CrossRef] [PubMed]
- Jeong, Y.J.; Park, K.; Kim, D.E. Isothermal DNA amplification in vitro: The helicase-dependent amplification system. Cell Mol. Life Sci. 2009, 66, 3325–3336. [Google Scholar] [CrossRef] [PubMed]
- Pohanka, M. The Piezoelectric Biosensors: Principles and Applications, a Review. Int. J. Electrochem. Sci. 2017, 12, 496–506. [Google Scholar] [CrossRef]
- Jiang, X.; Kim, K.; Zhang, S.; Johnson, J.; Salazar, G. High-temperature piezoelectric sensing. Sensors 2013, 14, 144–169. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.Q.; Luo, J.K.; Nguyen, N.T.; Waltone, A.J.; Flewitt, A.J.; Zu, X.T.; Li, Y.; McHale, G.; Matthews, A.; Iborra, E.; et al. Advances in piezoelectric thin films for acoustic biosensors, acoustofluidics and lab-on-chip applications. Prog. Mater. Sci. 2017, 89, 31–91. [Google Scholar] [CrossRef]
- Borgmann, S.; Schulte, A.; Neugebauer, S.; Schuhmann, W. Amperometric Biosensors. In Advances in Electrochemical Science and Engineering; Alkire, R.C., Kolb, D.M., Lipkowski, J., Eds.; WILEY-VCH Verlag GmbH & Co. KGaA: Weinheim, Germany, 2011; ISBN 978-3-527-32885-7. [Google Scholar]
- Li, S.; Gu, Y.; Lyu, Y.; Jiang, Y.; Liu, P. Integrated Graphene Oxide Purification-Lateral Flow Test Strips (iGOP-LFTS) for Direct Detection of PCR Products with Enhanced Sensitivity and Specificity. Anal. Chem. 2017, 89, 12137–12144. [Google Scholar] [CrossRef] [PubMed]
| Parameter | TaqMan®-Based HCV detection assay | SYBR®-Green Based HCV Detection Assay |
|---|---|---|
| Chemistry | Fluorogenic probes are used for detection of specific PCR products | PCR product is detected by SYBR Green I dye that binds to double-stranded DNA with high specificity |
| Advantages | Low signal to background ratio reduces the number of false positive results. Multiplex detection in a single tube reaction is possible by using distinguishable reporter dyes No post-PCR processing Detects specific amplification products only |
|
| Disadvantages | Requires use of multiple probes for target specificity in multiplex assay | Detects any double-stranded DNA, including non-specific reaction products |
| Technology | LOD | % Sens. | % Spec. | Advantages | Disadvantages | Ref. |
|---|---|---|---|---|---|---|
| RT-PCR | 125–500 IU/mL | 98.0 | 100.0 | Exponential amplification of template RNA | Inaccurate quantification due to numerous sources of variation | [38,121] |
| Real-Time PCR | 28–5000 copies/mL | 99.1 | 100.0 | Qualitative and quantitative analysis | Requires expensive equipment and reagents | [46,49,50] |
| NASBA | 500 copies/mL | 98.0 | 100.0 | Requires single-step isothermal RNA-specific amplification process | Does not allow HCV genotyping, 120–250 nucleotides sequence >250 or <120 amplifies less efficiently | [57,59,122,123] |
| TMA | 50 copies/mL | 100.0 | 99.5 | Single tube reaction Low-risk of cross contamination | - | [63,66] |
| RT-LAMP | 10 copies/mL | 91.5 | 100.0 | Rapid amplification Simple operation Easy detection | Effective only when the template is pure | [72,73,74,124] |
| RCA | 0.1 nmol/L | 90.0 | 84.8 | Aptamer/DNAzyme can replicate hundreds of times in a short time. Improvement of LOD is possible if combined with other biosensors. | Extremely complicated and it is not capable of amplifying a satisfactory length of nucleic acids | [78,125,126] |
| BERA | - | - | - | High operational speed | Chances of cross-contamination are high (require separate space) | [79] |
| PZ | - | - | - | Can be modified in different assay formats including direct detection, label free detection etc. | High temperature sensitivity | [127,128,129] |
| AB | 1.82 × 10−21 mol/L | - | - | High selectivity for substrates | Used biosensors exhibit limited stability. | [88,130] |
| GNP | 4.57 IU/µL | 93.3 | 100 | Assay is highly applicable for the detection of HCV | Not applicable for genotyping in its current form | [100] |
| CNT | 0.5 pM | - | - | Detection of HCV to picomolar range | HCV genotyping is not possible | [102] |
| Graphene | 0.1 nmol/L | - | - | Can serve as a sensor to locate and monitor the HCV gene in live mammalian cells | Not applicable for HCV genotyping in its current form | [104,131] |
| bDNA | 3.2 × 103 copies/mL | - | 98.2 | Low non-specific interactions. Multiple sample detections at ones | High LOD | [107] |
| LFA | 38 copies/mL | 99.5 | 98.8 | Low-cost, simple, rapid, and portable | Quantification is not possible | [115] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Warkad, S.D.; Nimse, S.B.; Song, K.-S.; Kim, T. HCV Detection, Discrimination, and Genotyping Technologies. Sensors 2018, 18, 3423. https://doi.org/10.3390/s18103423
Warkad SD, Nimse SB, Song K-S, Kim T. HCV Detection, Discrimination, and Genotyping Technologies. Sensors. 2018; 18(10):3423. https://doi.org/10.3390/s18103423
Chicago/Turabian StyleWarkad, Shrikant Dashrath, Satish Balasaheb Nimse, Keum-Soo Song, and Taisun Kim. 2018. "HCV Detection, Discrimination, and Genotyping Technologies" Sensors 18, no. 10: 3423. https://doi.org/10.3390/s18103423
APA StyleWarkad, S. D., Nimse, S. B., Song, K.-S., & Kim, T. (2018). HCV Detection, Discrimination, and Genotyping Technologies. Sensors, 18(10), 3423. https://doi.org/10.3390/s18103423






