A Novel Technique for Fetal ECG Extraction Using Single-Channel Abdominal Recording
Abstract
:1. Introduction
2. Materials and Methods
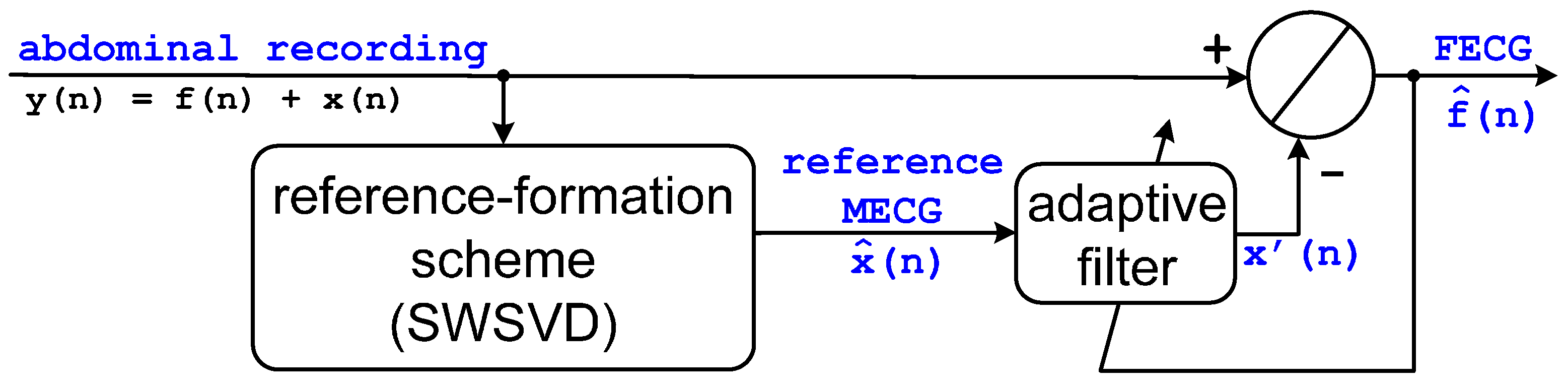
2.1. Basic Structure
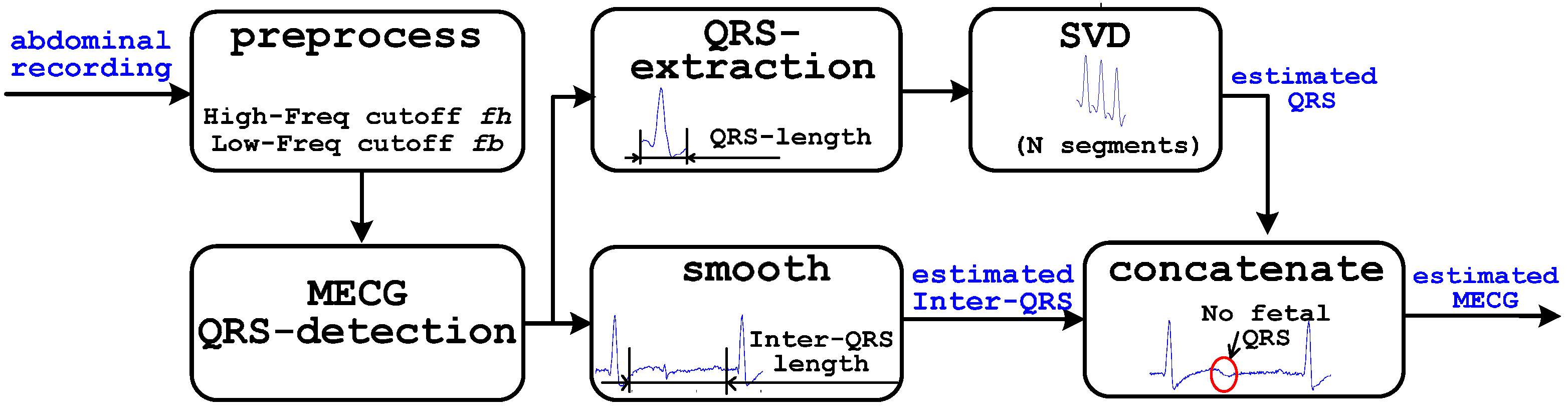
2.2. Reference Formation Scheme SWSVD
2.3. Adaptive Filter
3. Database and Statistical Assessment
3.1. Database
3.2. Statistical Assessment
4. Compared Methods
4.1. Cerutti Method
4.2. Kanjial Method
4.3. Suzanna Method
4.4. Vullings Method
5. Results
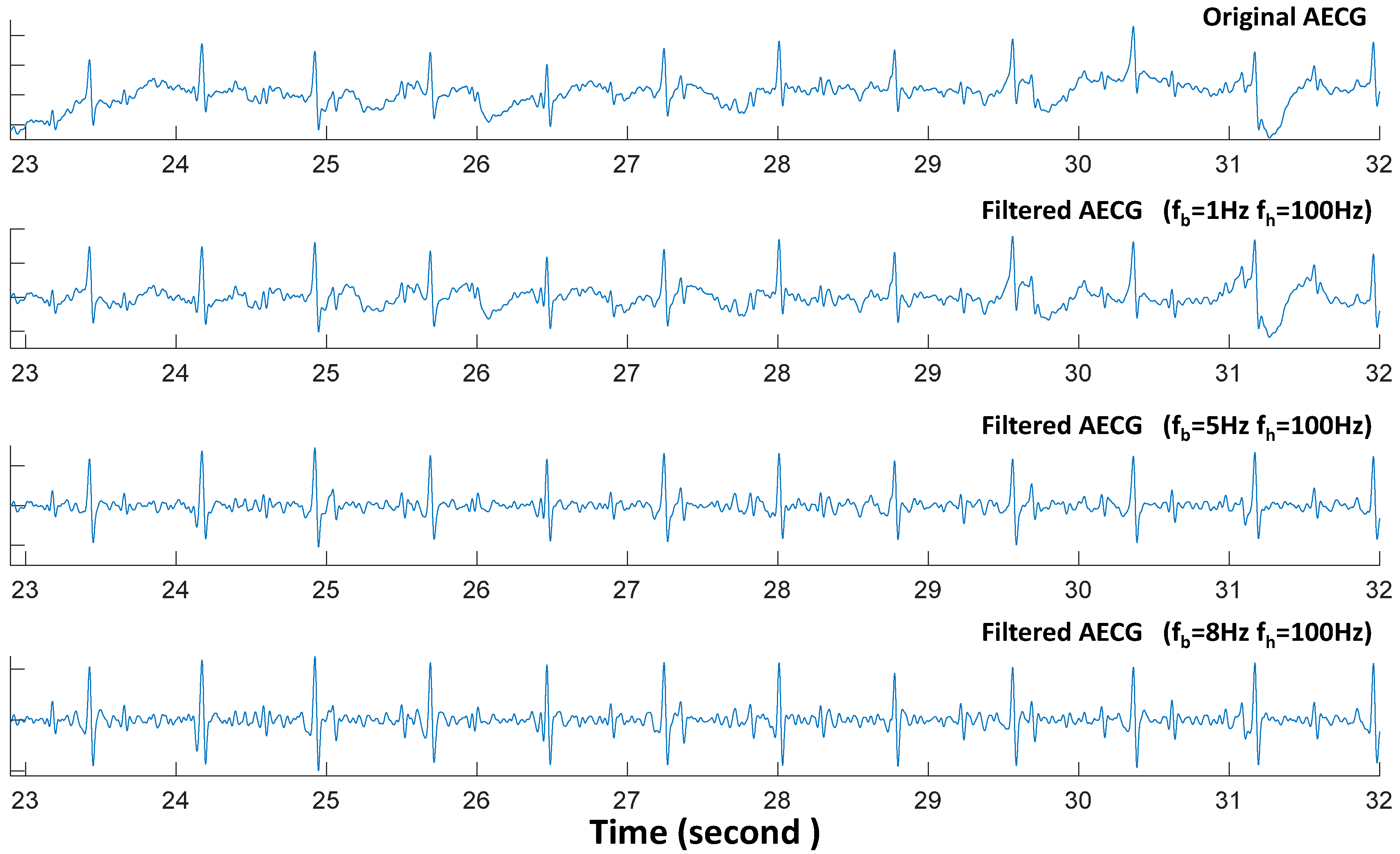
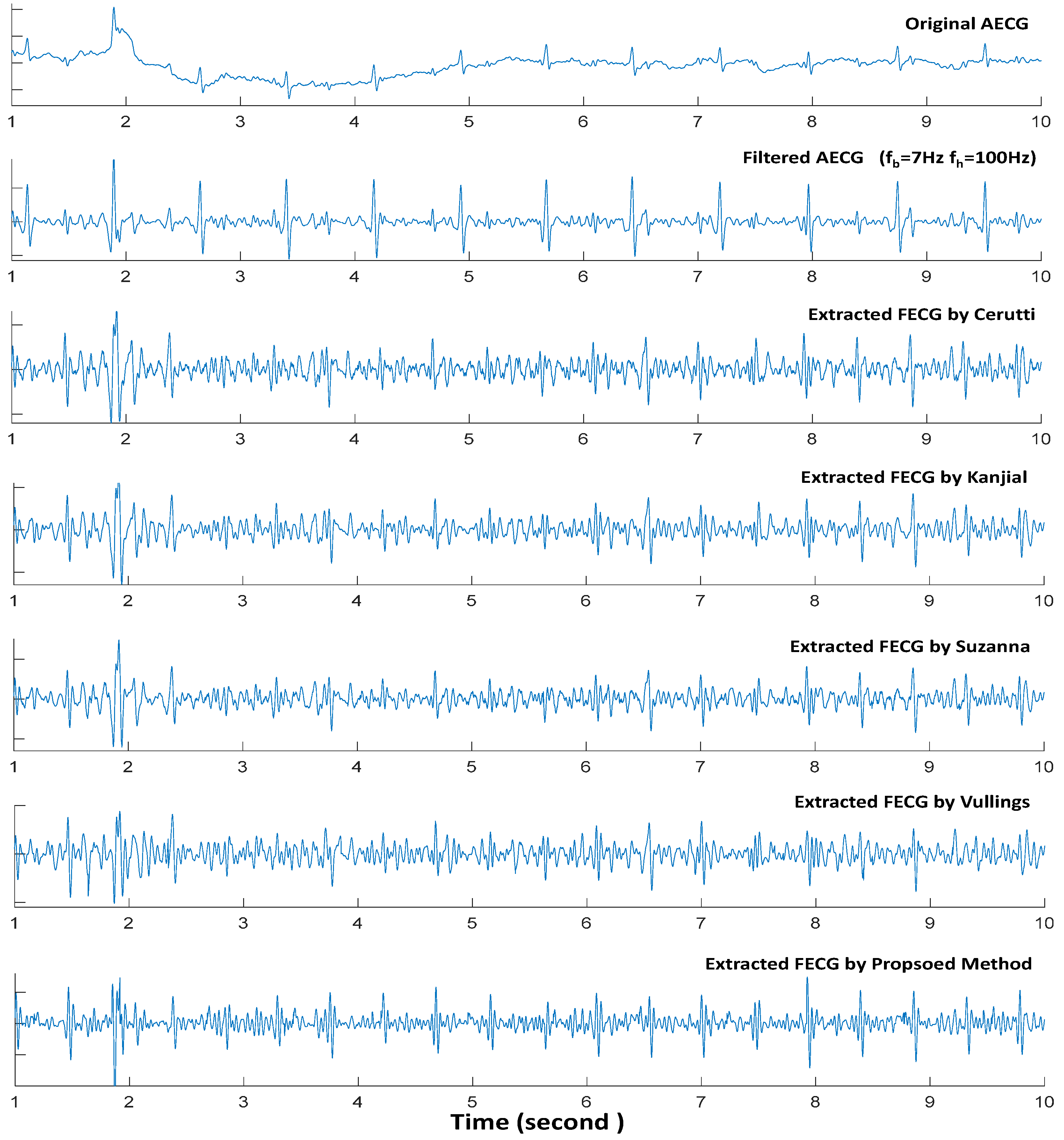
5.1. Preprocess
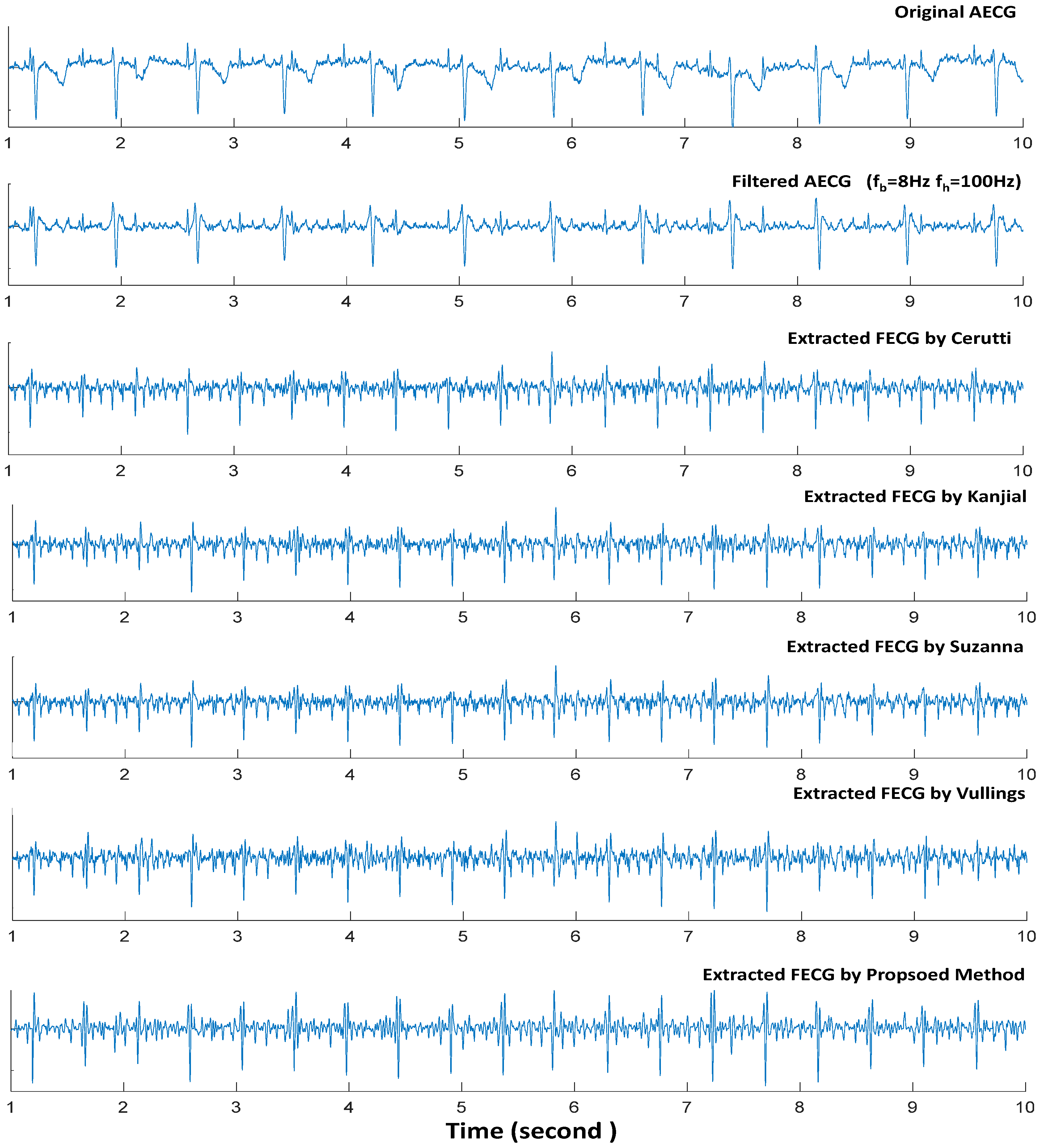
5.2. Performance Based on DB1
5.3. Performance Based on DB2
6. Discussion
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lewis, M.J. Review of electromagnetic source investigations of the fetal heart. Med. Eng. Phys. 2003, 25, 801–810. [Google Scholar]
- Barnett, S.; Maulik, D. Guidelines and recommendations for safe use of Doppler ultrasound in perinatal application. J. Matern-Fetal Neonatal Med. 2001, 10, 75–84. [Google Scholar] [CrossRef]
- Vullings, R.; Peters, C.; Sluijter, R.; Mischi, M.; Oei, S.; Bergmans, J. Dynamic segmentation and linear prediction for maternal ECG removal in antenatal abdominal recordings. Physiol. Meas. 2009, 30, 290–307. [Google Scholar] [CrossRef] [PubMed]
- Cohen, W.R.; Ommani, S.; Hassan, S.; Mirza, F.G.; Solomon, M.; Brown, R.; Schifrin, B.S.; Himsworth, J.M.; Hayes-Gill, B.R. Accuracy and reliability of fetal heart rate monitoring using maternal abdominal surface electrodes. Acta Obstet. Gynecol. Scand. 2012, 91, 1306–1313. [Google Scholar] [CrossRef] [PubMed]
- Naeye, R.L.; Peters, E.C. Antenatal hypoxia and low IQ value. Am. J. Dis. Child. 1987, 141, 50–54. [Google Scholar] [PubMed]
- Blair, E.; Stanley, F.J. Intrapartumasphyxia: A rare cause of cerebral palsy. J. Pediatr. 1988, 112, 515–519. [Google Scholar] [CrossRef]
- Philip, J.S.; Lone, E.H. Scientific and clinical evidence for the use of fetal ECG ST segment analysis (STAN). Acta Obstet. Gynecol. Scand. 2014, 93, 533–538. [Google Scholar]
- Zaunseder, S.; Andreotti, F.; Cruz, M.; Stepan, H.; Schmieder, C.; Wessel, N.; Jank, A.; Malberg, H. Fetal QRS detection by means of Kalman filtering and using the event synchronous canceller. Int. J. Bioelectromagn. 2013, 15, 83–89. [Google Scholar]
- Lipponen, J.A.; Tarvainen, M.P. Principal component model for maternal ECG extraction in fetal QRS detection. Physiol. Meas. 2014, 35, 1637–1648. [Google Scholar] [CrossRef] [PubMed]
- Zarzoso, V.; Nandi, A.K.; Bacharakis, E. Maternal and foetal ECG separation using blind source separation methods. IMA J. Math. Appl. Med. Biol. 1997, 14, 207–225. [Google Scholar] [CrossRef] [PubMed]
- Camargo-Olivarse, J.L.; Martín-Clemente, R.; Hornillo-Mellado, S.; Elena, M.M.; Román, I. The maternal abdominal ECG as input to MICA in the fetal ECG extraction problem. IEEE Signal Process. Lett. 2011, 18, 161–164. [Google Scholar] [CrossRef]
- Lathauwer, L.D.; Moor, B.D.; Vanderwalle, J. Fetal electrocardiogram extraction by blind source subspace separation. IEEE Trans. Biomed. Eng. 2000, 47, 567–572. [Google Scholar] [CrossRef] [PubMed]
- Sameni, R.; Juttern, C.; Shamsollahi, M. Multichannel electrocardiogram decomposition using periodic component analysis. IEEE Trans. Biomed. Eng. 2008, 55, 1935–1940. [Google Scholar] [CrossRef] [PubMed]
- Podziemski, P.; Gieraltowski, J. Fetal heart rate discovery: Algorithm for detection of fetal heart rate from noisy, noninvasive fetal ECG recordings. In Proceedings of the 2013 Computing in Cardiology Conference (CinC), Zaragoza, Spain, 22–25 September 2013; pp. 333–336.
- Behar, J.; Oster, J.; Clifford, G.D. Combining and benchmarking methods of foetal ECG extraction without maternal or scalp electrode data. Physiol. Meas. 2014, 35, 1569–1589. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Zeng, X.; Zhou, X.; Li, G.; Liu, Q. Robust Adaptive Fetal Heart Rate Estimation for Single-channel Abdominal ECG Recording. In Proceedings of the 5th International Conference on BioMedical Engineering and Informatics (BMEI), Chongqing, China, 16–18 October 2012.
- Rao, Y.; Zeng, H.; Li, X.; Li, Y. Estimate MECG from Abdominal ECG Signal Using Extended Kalman RTS Smoother. In Proceedings of the 6th International Conference on Intelligent Control and Information Processing, Wuhan, China, 26–28 November 2015.
- Sameni, R. Extraction of Fetal Cardiac Signals from an Array of Maternal Abdominal Recordings. Ph.D. Thesis, Sharif University of Technology-Institute National Polytechnique de Grenoble, Grenoble, France, 2008. [Google Scholar]
- Lee, K.J.; Lee, B. Sequential Total Variation Denoising for the Extraction of Fetal ECG from Single-Channel Maternal Abdominal ECG. Sensors 2016, 16, 1020. [Google Scholar] [CrossRef] [PubMed]
- Castillo, E.; Morales, D.P.; Botella, G.; Garcia, A.; Parrilla, L.; Palma, A.J. Efficient wavelet-based ECG processing for single-lead FHR extraction. Digit. Signal Process. 2013, 23, 1897–1909. [Google Scholar] [CrossRef]
- Richter, M.; Schreiber, T.; Kaplan, D.T. Fetal ECG extraction with nonlinear statespace projections. IEEE Trans. Biomed. Eng. 1998, 45, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Widrow, B.; Glover, J.R., Jr.; McCool, J.M.; Kaunitz, J.; Williams, C.S.; Hearn, R.H.; Zeidler, J.R.; Dong, E., Jr.; Goodlin, R.C. Adaptive noise cancelling: Principles and applications. Proc. IEEE 1975, 63, 1692–1716. [Google Scholar] [CrossRef]
- Niknazar, M.; Rivet, B.; Jutten, C. Fetal ECG Extraction by Extended State Kalman Filtering Based on Single-Channel Recordings. IEEE Trans. 2013, 60, 1345–1352. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Xiao, Y.; Wei, G.; Sun, J. A multichannel nonlinear adaptive noise canceller based on generalized FLANN for fetal ECG extraction. Meas. Sci. Technol. 2016, 27, 015703. [Google Scholar] [CrossRef]
- Strobach, P.; Fuchs, K.A.; Harer, W. Event-synchronous cancellation of the heart interference in biomedical signals. IEEE Trans. Biomed. Eng. 1994, 41, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Wolf, W.; Schnell, R.; Appel, U. New aspects to event-synchronous cancellation of ECG interference: An application of the method in diaphragmatic EMG signals. IEEE Trans. Biomed. Eng. 2000, 47, 1177–1184. [Google Scholar] [CrossRef] [PubMed]
- Shao, M.; Barner, K.E.; Goodman, M.H. An interference cancellation algorithm for noninvasive extraction of transabdominal fetal electroencephalogram. IEEE Trans. Biomed. Eng. 2004, 51, 471–483. [Google Scholar] [CrossRef] [PubMed]
- Behar, J.; Johnson, A.; Clifford, G.D.; Oster, J. A Comparison of Single Channel Fetal ECG Extraction Methods. Ann. Biomed. Eng. 2014, 42, 1340–1353. [Google Scholar] [CrossRef] [PubMed]
- Guyton, A.; Hall, J. Textbook of Medical Physiology, 10th ed.; WB Saunders: Philadelphia, PA, USA, 2000. [Google Scholar]
- Suzanna, M.; Rabotti, C.; Mischi, M.; Sluijter, R.J. A robust fetal ECG detection methods for abdominal recordings. Physiol. Meas. 2007, 28, 373–388. [Google Scholar]
- Guyton, A.C.; Hall, J.E. Textbook of Medical Physiology, 9th ed.; WB Saunders: Philadelphia, PA, USA, 1996. [Google Scholar]
- Kanjilal, P.P.; Palit, S.; Saha, G. Fetal ECG extraction from single-channel maternal ECG using singular value decomposition. IEEE Trans. Biomed. Eng. 1997, 44, 51–59. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Tompkins, W.J. A real-time QRS detection algorithm. IEEE Trans. Biomed. Eng. 1985, 32, 230–235. [Google Scholar] [CrossRef] [PubMed]
- Brambati, B.; Pardi, G. The intraventricular conduction time of fetal heart in uncomplicated pregnancies. Br. J. Obstet. Gynaecol. 1980, 87, 941–948. [Google Scholar] [CrossRef]
- Abdominal and Direct FECG Database. Available online: https://physionet.org/physiobank/database/adfecgdb/ (accessed on 15 January 2016).
- Available online: http://www.ambu.com/sensor (accessed on 7 January 2016).
- Behar, J.; Andreotti, F.; Oster, J.; Clifford, G.D. A Bayesian filtering framework for accurate extracting of the non-invasive FECG morphology. In Proceedings of the IEEE Conference on Computing in Cardiology, Boston, MA, USA, 7–10 September 2014; pp. 53–56.
- ANSI/AAMI/ISO EC57 (1998/(R) 2008). Testing and Reporting Performance Results of Cardiac Rhythm and ST-Segment Measurement Algorithms; American National Standards Institute: New York, NY, USA, 2008. [Google Scholar]
- Samuel, O.W.; Asogbon, G.M.; Sangaiah, A.K.; Fang, P.; Li, G. Integrated decision support system based on ANN and Fuzzy_AHP for heart failure risk prediction. Expert Syst. Appl. 2016, 68, 163–172. [Google Scholar] [CrossRef]
- Andreotti, F.; Riedl, M.; Himmelsbach, T.; Wedekind, D.; Wessel, N.; Stepan, H.; Schmieder, C.; Jank, A.; Malber, H.; Zaunseder, S. Robust fetal ECG extraction and detection from abdominal leads. Physiol. Meas. 2014, 35, 1551–1567. [Google Scholar] [CrossRef] [PubMed]
- Fernando, A.; Joachim, B.; Sebastian, Z.; Julien, O.; Gari, D.C. An open-source framework for stress-testing non-invasive foetal ECG extraction algorithms. Physiol. Meas. 2014, 37, 627–948. [Google Scholar]
- Extracting NI-FECG signals. Available online: http://fernandoandreotti.github.io/fecgsyn/pages/documentation.html# Extraction (accessed on 18 May 2016).
- Cerutti, S.; Baselli, G.; Civardi, S.; Ferrazzi, E.; Marconi, A.M.; Pagani, M.; Pardi, G. Varibility analysis of fetal heart rate signals as obtained from abdominal electrocardiographic recordings. J. Perinat. Med. 1986, 14, 445–452. [Google Scholar] [CrossRef] [PubMed]
- Pipberger, H.V.; Arzbaecher, R.C.; Berson, A.S.; Briller, S.A.; Brody, D.A.; Flowers, N.C.; Geselowitz, D.B.; Lepeschkin, E.; Oliver, C.; Schmitt, O.H.; et al. Recommendations for standardization of leads and of specifications for instruments in electrocardiography and vectorcardiography. Circulation 1975, 52, 11–31. [Google Scholar]
- Behar, J.; Andreotti, F.; Zaunseder, S.; Oster, J.; Clifford, G.D. A practical guide to non-invasive foetal electrocardiogram extraction and analysis. Physiol. Meas. 2016, 37, R1–R35. [Google Scholar] [PubMed]


| Methods | F1 (%) | PPV (%) | SE (%) | ACC (%) | TP | FP | FN |
|---|---|---|---|---|---|---|---|
| Cerutti | 0.9945 | 0.9953 | 0.9938 | 0.9891 | 638 | 3 | 4 |
| Kanjial | 0.9922 | 0.9938 | 0.9907 | 0.9845 | 636 | 4 | 6 |
| Suzanna | 0.9953 | 0.9953 | 0.9953 | 0.9907 | 639 | 3 | 3 |
| Vullings | 0.9890 | 0.9937 | 0.9844 | 0.9783 | 632 | 4 | 10 |
| Proposed Method | 0.9961 | 0.9969 | 0.9953 | 0.9922 | 639 | 2 | 3 |
| Methods | F1 (%) | PPV (%) | SE (%) | ACC (%) | TP | FP | FN |
|---|---|---|---|---|---|---|---|
| Cerutti | 0.9896 | 0.9873 | 0.9920 | 0.9795 | 620 | 8 | 5 |
| Kanjial | 0.9880 | 0.9872 | 0.9888 | 0.9763 | 618 | 8 | 7 |
| Suzanna | 0.9912 | 0.9889 | 0.9936 | 0.9826 | 621 | 7 | 4 |
| Vullings | 0.9689 | 0.9666 | 0.9712 | 0.9396 | 607 | 21 | 18 |
| Proposed Method | 0.9928 | 0.9936 | 0.9920 | 0.9857 | 620 | 4 | 5 |
| Methods | F1 (%) | PPV (%) | SE (%) | ACC (%) | TP | FP | FN |
|---|---|---|---|---|---|---|---|
| Cerutti | 0.9465 | 0.9438 | 0.9492 | 0.8984 | 168 | 10 | 9 |
| Kanjial | 0.9489 | 0.9543 | 0.9435 | 0.9027 | 167 | 8 | 10 |
| Suzanna | 0.9235 | 0.9261 | 0.9209 | 0.8579 | 163 | 13 | 14 |
| Vullings | 0.8262 | 0.8333 | 0.8192 | 0.7039 | 145 | 29 | 32 |
| Proposed Method | 0.9858 | 0.9886 | 0.9831 | 0.9721 | 174 | 2 | 3 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, N.; Zhang, J.; Li, H.; Mumini, O.O.; Samuel, O.W.; Ivanov, K.; Wang, L. A Novel Technique for Fetal ECG Extraction Using Single-Channel Abdominal Recording. Sensors 2017, 17, 457. https://doi.org/10.3390/s17030457
Zhang N, Zhang J, Li H, Mumini OO, Samuel OW, Ivanov K, Wang L. A Novel Technique for Fetal ECG Extraction Using Single-Channel Abdominal Recording. Sensors. 2017; 17(3):457. https://doi.org/10.3390/s17030457
Chicago/Turabian StyleZhang, Nannan, Jinyong Zhang, Hui Li, Omisore Olatunji Mumini, Oluwarotimi Williams Samuel, Kamen Ivanov, and Lei Wang. 2017. "A Novel Technique for Fetal ECG Extraction Using Single-Channel Abdominal Recording" Sensors 17, no. 3: 457. https://doi.org/10.3390/s17030457
APA StyleZhang, N., Zhang, J., Li, H., Mumini, O. O., Samuel, O. W., Ivanov, K., & Wang, L. (2017). A Novel Technique for Fetal ECG Extraction Using Single-Channel Abdominal Recording. Sensors, 17(3), 457. https://doi.org/10.3390/s17030457








