Abstract
Angel sharks are distributed worldwide in tropical to subtropical waters. Across the Eastern Pacific Ocean (EPO), two valid species are reported: The Pacific angelshark Squatina californica and the Chilean angelshark Squatina armata; however, there is still uncertainty about their geographic distribution, mainly along the northern Peru coast where the species have been reported to be sympatric. The aim of this study is to describe the genetic differences between the genus Squatina from the EPO, including samples from northern Peru, and using DNA barcoding and three species delimitation models: Poisson tree processes (PTP) model, Bayesian implementation of the PTP (bPTP) model and the general mixed Yule coalescent (GMYC) model. The three approaches summarized 19 nominal Squatina species in 23 consensus Molecular Taxonomic Units (MOTU). Only 16 of them were in accordance with taxonomic identifications. From the EPO, four Squatina MOTUs were identified, one from North America (S. californica USA/Mexico) and three sampled in northern Peru, S. californica Peru, S. armata and Squatina sp. (a potential new species). This study contributes to the management and conservation policies of angel sharks in Peru, suggesting the presence of an undescribed species inhabiting the northern Peruvian coast. The use of molecular approaches, such as DNA barcoding, has the potential to quickly flag undescribed species in poorly studied regions, including the Southeast Pacific, within groups of ecologically and economically important groups like angel sharks.
1. Introduction
One of the most diverse groups of marine predators is the subclass Elasmobranchii (i.e., sharks, skates, and rays). Among them, there is a small but highly distinctive group of bizarrely-shaped benthic sharks, commonly called angel sharks. This group of ray-like sharks belongs to the monophyletic family Squatinidae (Bonaparte, 1838) [1,2,3] encompassing a unique genus, Squatina (Dumeril, 1805), with 22 extant species described based on morphological characters or molecular information [4,5,6,7]. Although, there are two additional species from the Gulf of Mexico described [8], Squatina mexicana Castro–Aguirre, Espinosa–Pérez and Huidobro–Campos, 2007, and Squatina heteroptera Castro–Aguirre, Espinosa–Pérez and Huidobro–Campos, 2007, these two species are considered as not valid [9] because of uncertainty about the validity of their morphological description [6,7,10,11,12]. Currently, S. mexicana and S. heteroptera are considered junior synonyms of Squatina dumeril Lesueur, 1818 [7,10].
Angel sharks are distributed worldwide in tropical to subtropical shelf waters, although most of the species have restricted distribution in regional seas (e.g., S. david distributed within the Southern Caribbean Sea) [10]. Across the Eastern Pacific Ocean (EPO), two valid and sympatric species occur: the Pacific angelshark Squatina californica Ayres, 1859 and the Chilean angelshark Squatina armata (Philippi, 1887) [4]. Until recently it was considered that S. californica was distributed off the coast of North America, from Alaska to the Gulf of California [13]. However, some studies reported its presence in Ecuador [14,15] and Peru [15]. On the other hand, S. armata has been reported to inhabit waters from northern Peru to the central coast of Chile [4,16,17]. Nevertheless, the northern limit of its distribution range is not clear since some studies have suggested its presence in Ecuador [18], Colombia [19], and Costa Rica [20]. Current information such as the range of its geographic extent and abundance has been used to determine the extinction risk by the International Union for Conservation of Nature (IUCN) Red List of threatened species, classifying S. californica as ‘Near Threatened’ [21] and S. armata as ‘Critically Endangered’ [22]. Nonetheless, to establish management measures also at a national level, data concerning its biology, ecology, as well as its taxonomic status, need to be resolved.
Several studies support the validity of both species based on morphological taxonomic characters [10,16,23]; however, other studies consider these species as synonymous [1,23,24]. This conclusion that S. californica and S. armata were identical was made after the comparison of various specimens from the northern and southern hemisphere. Nonetheless, neither information about the exact number of specimens nor how this comparison was made is provided by the authors [25]. Subsequent publications [1,13,14] also do not provide more detail to explain this suggested synonymy. The first species described in the EPO was S. californica based on the revision of one single specimen (around 96 cm, unidentified sex) collected in San Francisco Bay, United States, in September 1857 [26]. The specimen were compared to Squatina dumeril and differed in qualitative characteristic and meristic traits (i.e., form of the orbits, form of teeth, size of pectorals, form of pectorals, form of the ventrals, form of the dorsal and number of teeth) [26]. S. armata, was described as Rhina armata, based also on a single male individual (103 cm) collected at Iquique, Chile (unknown sampling date) [27]. The reference species used to compare external characteristics was a specimen identified by the author as ‘Rhina squatina’ collected in Rio de Janeiro, Brazil, although it might be one of the four known species of the Western Atlantic (i.e., S. dumeril, S. argentina, S. guggenheim, S. occulta, or S. punctata) but not the extant S. squatina. This latter species is excluded because its presence has been only reported at the Baltic Sea, North Sea, Black Sea, Mediterranean Sea, and Canary Islands [9]. In this case, seven taxonomic characteristics were reported to differ between S. armata and the specimen from Brazil (i.e., form of the pectoral fins, size of the pectoral fins, width of the head, shape and size of the two spiracles, form of the tail, and presence of enlarged ‘denticles’ on the pectoral fin) [27].
Around the world, delimitation and the identification of angel shark species have become important due to their biological characteristics (e.g., large size, reproductive cycle, demersal nature) and because this group is susceptible and vulnerable to fisheries and human activities [28,29]. Therefore, to achieve effective conservation and fishery management strategies, correctly delimiting sampling units (i.e., taxon ‘species’) is urgently needed to generate species-specific data to support these strategies [30]. Over the last decades, classical taxonomy (i.e., description and identification of species through taxonomic morphological characters) has been the main approach used to describe angel sharks [6,8,24,26,27,31,32,33,34]. Nonetheless, the use of molecular tools, such as genomic data, is increasingly being considered to support the process of species delimitation [5,30]. For instance, instead of the use of morphological characters, the molecular approach uses DNA sequences which are grouped into Molecular Taxonomic Units (MOTU). A MOTU is considered as an operational definition that groups individual DNA sequences (i.e., cluster of sequences) based on an explicit algorithm and is used to estimate diversity at the species level. A powerful tool that uses standardized gene regions (e.g., cytochrome c oxidase—COI) to delimit and identify species is DNA barcoding. This molecular tool couples a comprehensive dataset of COI DNA sequences together with validated identified voucher specimens to support taxonomic studies [35,36]. Among their benefits, DNA barcoding can assist in defining phylogenetic relationships and species geographic boundaries. Furthermore, DNA barcoding together with single locus species delimitation methods have recently shown to be an effective tool for validating elasmobranch identification and describing new species in a number of fisheries [37,38,39,40,41,42,43].
Stelbrink et al. [4] employed COI and 16S rRNA markers to provide a comprehensive global phylogeny of 17 Squatina species, including the two species found along the EPO, S. armata and S. californica. In that study, S. armata is the first species to branch off the clade that includes the North and South American species, indicating the existence of two different species. However, in that study, only samples from the ends of both distributions, in California and Chile, were used. Thus, the detailed distribution of both species along the EPO remains uncertain, especially in the areas of North of South America where both species have been reported [14,15,18,44]. Additionally, angel sharks are target species and are captured by small-scale fisheries [28]. However, reports of landings are rather general and likely include several species of angel sharks, thus molecular tools could serve well for the accurate identification of the species landed, their distributions, and the fisheries with which they interact [28].
In this regard, the aim of the present study is to describe the genetic differences within the genus Squatina from the EPO, including samples of angel sharks from an area not previously covered (i.e., northern Peru), and integrate them with mtDNA COI sequences data from Barcode of Life Data (BOLD) System to evaluate species boundaries using species delimitation methods and determine MOTUs.
2. Materials and Methods
2.1. Morphological Identification and Sample Collection
As part of the sampling campaign of the consortium for DNA barcoding of Peruvian marine species (PeMar), a total of eight specimens of angel sharks were sampled between 2017 and 2018 from fish markets and landing sites in northern Peru (Figure 1 and Figure 2). All specimens collected were identified using external morphological characteristics following a literature review [1,10,23]. Each specimen was photographed, and one specimen was fixed and deposited in the fish collection of the San Marcos–Natural History Museum (UNMSM) for further analyses. Muscle tissue samples were extracted from all specimens and preserved in 96% ethanol at room temperature (17–20 °C). Sequences, sample records, and voucher numbers can be viewed in Table 1.
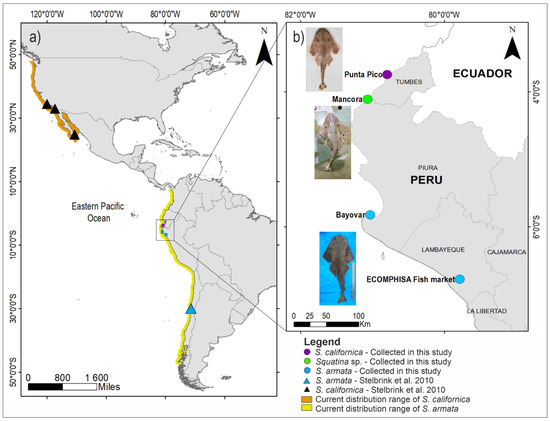
Figure 1.
Geographic distribution map and sampling sites of DNA sequences of Squatina species along Eastern Pacific Ocean. (a) Current known distribution range adapted from Fricke et al., (2020) and sampling sites of Squatina californica (black triangle) and Squatina armata (light blue triangle) reported by Stelbrik et al., 2010. (b) Sampling sites of Squatina californica (purple circle), Squatina sp. (green circle), and Squatina armata (light blue circle) selected by PeMar Project in northern Peru.
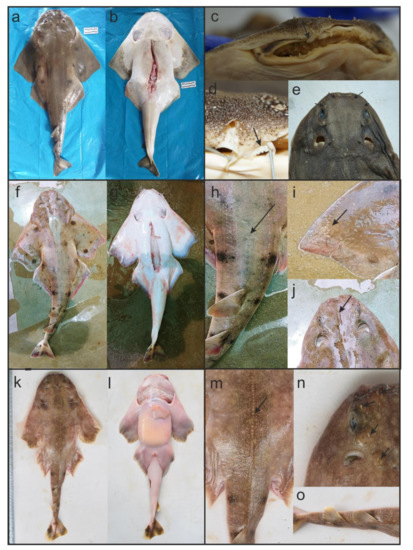
Figure 2.
Specimens of angel sharks collected for this study: Dorsal (a) and ventral (b) images of one fresh specimen of Squatina armata (Pemar_V0173). Detailed images of barbels (c), anterior nasal flaps (d) and thorns on snout, between eyes and spiracles (e) observed on preserved specimen of S. armata (Pemar_V0174). Dorsal (f) and ventral (g) images of one fresh specimen observed on Squatina sp. (Pemar_V0209). Detailed images of thorns along the middle line of the back (h), denticles covering the edges of the pectoral fin (i) and concave between eyes (j) of specimens of Squatina sp. (Pemar_V0209 and Pemar_V0211). Dorsal (k) and ventral (l) images of one fresh pup of S. californica (LCT_2160). Detailed images of thorns (m,n) and pale dorsal fins (o) of S. californica pup (LCT_2160).

Table 1.
Angel shark species sampled along the northern Peruvian coast, PeMar project code, BOLD code, voucher code, sampling site (region and exact site), sex, and total length (TL).
2.2. DNA Extraction, Amplification and Sequencing
Total genomic DNA was isolated using Tissue Kit (Thermo Scientific) following the manufacturer’s instructions. A partial fragment (~655 base-pair) of the mitochondrial Cytochrome Oxidase subunit I (COI) gene was amplified through Polymerase Chain Reaction (PCR) using primers FishF1-FishR1 or FishF2-FishR2 [35], that amplify an overlapping region from the 5’ region of the COI gene. The PCR was performed with a final volume of 25 µL containing 16.35 µL distilled water, 2.5 µL dNTP (8 nM), 0.6 µL of each primer (5 µM), using just one pair of primers (i.e., F1/R1 or F2/R2) and 0.6 µL of Taq polymerase (5 µ/µL). PCR conditions consisted of an initial denaturation at 95 °C for 2 min, followed by 30 cycles including denaturation at 95 °C for 45 s, annealing at 52 °C for 45 s, and extension at 72 °C for 60 s, and a final extension at 72 °C for 5 min. Amplified products were checked on 1% agarose gel and both strands per amplicon were sent to Macrogen (Rockville, MD, USA) for Sanger sequencing. The sequencing was carried out using the same set of primers that was used in the PCR, however a greater number of samples were amplified and sequenced using the Fish F1 and Fish R1 primers, since these had better efficiency for our samples. Sequences were cleaned and contigs were assembled using the software CodonCode Aligner (CodonCode Corporation, Dedlham, MA, USA). Multiple alignments were done using a ClustalW algorithm [45], implemented in the software MEGA 7 [46] and were checked manually for misalignments and trimmed to the shortest common sequence length.
2.3. Species Delimitation Methods
To apply several species delimitation methods to infer MOTUs, our samples were combined with COI sequences from 19 Squatina nominal species (Table 2, Table S1) obtained from the public repository BOLD (www.boldsystems.org, accessed on 5 February 2021), GenBank (www.ncbi.nlm.nih.gov/genbank/, accessed on 5 February 2021), and from the literature [5,35,36,43,47,48,49,50,51,52,53,54,55,56]. The set of DNA fragments were chosen following two criteria: (1) the location of sampling mentioned in the public database (e.g., country, province, region, sector, exact site, coordinates or FAO fishing zones) and (2) length of the DNA fragment (i.e., ~610 bp). To reduce computational time, only 10 COI sequences were chosen per species, when it was possible. The two sequences of Squalus used as an outgroup by Stelbrink et al. [4] were also included in the molecular analysis.

Table 2.
List of extant species, their current distribution (Fricke et al., 2020), number of COI sequences used in this study per species, and sampling sites reported in BOLD System per species. See Table S1 for detailed list.
We performed and compared three molecular species delimitation methods using the pipeline SPdel (https://github.com/jolobito/SPdel, accessed on 8 March 2021) added to a delimitation species method based on classical taxonomy. The first method implemented was the general mixed Yule coalescent (GMYC) model [57] with a single threshold. The GMYC model identifies the threshold value at the transition of branching patterns that are characteristics of the speciation process [58] versus coalescence process [59], and identifies significant changes in the branching rates in a time-calibrated ultrametric tree. The other methods used were the Poisson tree processes (PTP), and the Bayesian implementation of the PTP model (bPTP) [60]. These two methods directly use the number of substitutions as opposed to the GMYC method that uses time. For all models, an ultrametric tree was used as an input file which was generated in BEAUti v2.1 [61], with a normal relaxed clock and a birth and death model, and a HKY+G substitution model suggested by the Bayesian Information Criterion in jModeltest 2 [62]. The analysis was implemented with 60,000,000 million MCMC generations and with a burn-in of 10%. To run the analysis, we used BEAST v2.5 [61] implemented in the Cyber Infrastructure for phylogenetic Research (CIPRES; https://www.phylo.org, accessed on 8 March 2021) [63]. Convergence and adequate sample size (greater than 130) were evaluated in Tracer v1.6.0. The pipeline SPdel used for our analysis performs a comparison between the four chosen methods and ultimately generates a consensus species delimitation considering three molecular approaches. Only MOTUs supported by at least two of the applied delimitation methods were considered consensus MOTUs. Furthermore, we quantified the degree of genetic divergences for nominal species and for each one of the species delimitation methods used, calculating the values of intragroup and intergroup Kimura 2–parameter (K2-P) genetic distances in SPdel.
3. Results
3.1. Taxonomic Identification
Most of our sampled specimens were not retained because they were part of fishers’ daily catch. For this reason, identification was done mainly in the field (i.e., with fresh specimens) and through photographs allowing for diagnostic taxonomic characters to be assessed. The four specimens collected at the ECOMPHISA fish market and close to the Bayovar Port were identified as S. armata (Table 1, Figure 1). For the identification we used an illustrated guide [10] and a taxonomic key [15], distinguishing the following combination of characteristics: reddish-brown to blackish dorsal surface (Figure 2a), white ventral surface with dark brown edged of pectoral and pelvic fins (Figure 2b), narrow and simple barbels (Figure 2c), anterior nasal flaps fringed (Figure 2d), and thorns on snout, between eyes and spiracles (Figure 2e). One angel shark pup was collected off the coast of Punta Pico (Table 1, Figure 1). It was identified as S. californica following the aforementioned illustrated guide, distinguishing the following characteristics: reddish-brown dorsal surface with scattered light spots (Figure 2k), white edged pectoral and pelvic fins (Figure 2l), presence of thorns (Figure 2m,n), and pale dorsal fins (Figure 2o). Finally, three specimens collected in Mancora (Table 1, Figure 1) were identified as S. californica in the field, however after the molecular analysis, they were allocated to the taxon Squatina sp. (Figure 2f–j).
3.2. MOTU Delimitation Analyses
We obtained eight sequences (610 base-pair) from two nominal species, S. armata (n = 4), S. californica (n = 1), and Squatina sp. (n = 3), from northern Peru with 30 parsimony informative sites. The final alignment of mtDNA COI sequences resulted in 591 bp (shortest common sequence length) comprised of a total of 19 nominal species (Result S1).
The species delimitation analyses showed that PTP and the bPTP method delimited the same 23 Squatina MOTUs (Figure 3), with a maximum intra-MOTU divergence of 0.99% (for the MOTU of S. squatina) and a minimum inter-MOTU divergence of 0.49% (between the MOTUs of S. californica collected in Peru and USA/Mex). On the other hand, the GMYC method delimited Squatina 25 MOTUs with a maximum intra-MOTU divergence of 0.99% (for the MOTU of S. squatina) and a minimum inter-MOTU divergence of 0.16% (between the MOTUs of S. squatina collected in Ireland and Turkey). Single-locus species delimitation results from PTP, bPTP, and GMYC approaches were summarized by using the pipeline SPdel, in 25 consensus MOTUs. The maximum intra-MOTU distance was 0.99% (for the MOTU of S. squatina) (Table 3) and the minimum inter-MOTU distance was 0.49% (between the MOTUs of S. californica collected in Peru and USA/Mexico) (Table 3). In contrast, if considering species delimited through traditional taxonomy, the maximum intraspecific distance was 2.51% (between specimens morphologically identified as S. africana) (Table 3) and the minimum interspecific distance was 0 between specimens of S. formosa (collected in Thailand) and S. nebulosa (collected in China and Southern China Sea) (Table 3).
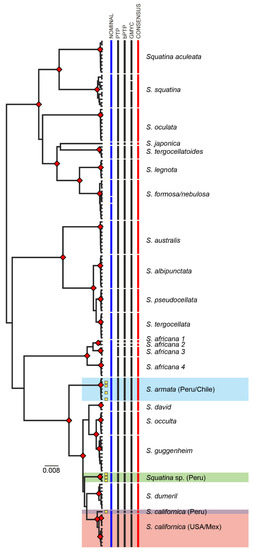
Figure 3.
Bayesian tree showing the clustering of the MOTUs obtained by the species delimitation analyses (PTP, bPTP, and GYMC) and the consensus analysis. The red diamonds indicate nodes with supports higher than 0.9 Bayesian posterior probability. The scale bar indicates nucleotide substitutions per site. Samples from the Eastern Pacific Ocean are delimited by squares. Yellow squares indicate samples from northern Peru collected in this study.

Table 3.
Genetic K2-P distances of MOTUs and nominal species of angel sharks: mean intra-MOTU divergence (mean intra-), maximum intra-MOTU divergence (maximum intra-), distance to the nearest neighbor (distance to NN) and the nearest neighbor (NN).
From the 23 consensus Squatina MOTUs, 16 are in accordance with taxonomic identification: S. aculeata, S. albipunctata, S. armata, S. australis, Squatina sp., S. david, S. dumeril, S. guggenheim, S. japonica, S. legnota, S. occulta, S. oculata, S. pseudocellata, S. tergocellata, S. tergocellatoides, and S. squatina. While seven consensus MOTUs do not match with the current taxonomy: S. californica USA/Mexico, S. californica Per, S. africana 1, S. africana 2, S. africana 3, S. africana 4, and S. formosa/nebulosa. Both nominal species, S. africana and S. californica, were split into four and two MOTUs, respectively (Table 3). The S. californica samples formed a polyphyletic group, with two MOTUs closely related, one from the Northeast Pacific (samples collected within the Gulf of California, Mexico, and in California, USA), and the other composed by samples collected in northern Peru (Figure 3). The minimum intra-MOTU divergence calculated for S. californica from Northeast Pacific was 0% and the maximum intra-MOTU value was 0.83%. Furthermore, the minimum and the maximum inter-MOTU divergences between the MOTU of S. californica from the northern hemisphere and the southern hemisphere were 0.5 and 0.83%, respectively. Samples identified initially as S. californica (Pemar_V0209, Pemar_V0210, and Pemar_V0211) from northern Peru, were grouped into a clade, separately from S. californica and S. armata and for this reason they were renamed as Squatina sp. The minimum and maximum intra-MOTU divergences for Squatina sp. were 0 and 0.33%, respectively.
4. Discussion
Only two valid species had been previously described for the EPO: Squatina californica and Squatina armata [7,10,28], and both species have been reported as sympatric for the northern Peruvian coast [15]. Nonetheless, there is controversy about the validity of these species due to the lack of taxonomic studies confirming their presence across the whole range of their geographic extents [10]. Our results show a new scenario, reporting the existence of four MOTUs along the EPO (Figure 3), revealing a hidden diversity that may include at least one new species for the genus Squatina in the Southeast Pacific.
The Bayesian phylogenetic tree obtained shows a similar topology compared with the comprehensive phylogenetic analysis carried out by Stelbrink et al. [4], and all species delimitation analyses performed (i.e., PTP, bPTP, and GMYC) yielded mostly the same result. The four MOTUs found in the EPO were grouped within the clade of North and South American species described by Stelbrink et al. [4]. However, the four MOTUs are not phylogenetically closely related with the exception of the MOTU of S. californica from North America and the MOTU of S. californica from northern Peru that are sister MOTUs (Figure 3).
All the algorithms used split the cluster of S. californica into two MOTUs. One group includes specimens collected along the Pacific coast of the United States and specimens from the Gulf of California, the other group includes sequences from Punta Pico located in the region of Tumbes, Peru (Figure 3). In some previous studies, two different population lineages originated in the Northern Pacific and in the Gulf of California were found [4,13,64], but this division was not supported by the coalescent species delimitation methods herein used, instead of reciprocal monophyly. Nonetheless, the minimum K2-P genetic distances within the MOTUs of S. californica from the northern hemisphere (0.5%) are comparable to the lowest genetic distances found between Squatina nominal species (0.5% for S. guggenheim and S. occulta). In regard to S. californica from the southern hemisphere, it was discriminated as a different MOTU from the clade of the northern hemisphere, even though the maximum K2-P genetic distances between both clades is 0.5%, similar to the values observed between the population from North America and other Squatina nominal species. This significant heterogeneity between S. californica populations may be promoted by their ecological behavior (e.g., limited ability for sustained swimming due to their morphological and anatomical characteristics) [13], their reproductive behavior (e.g., philopatric behavior) [28] but also due to geographic barriers (e.g., deep marine basins as barriers to dispersal of these populations) [13]. To elucidate if the degree of separation of these MOTUs of S. californica corresponds to a recent divergent species or a strong population structure, more studies including specimens of different ontogenetic stages collected along the Eastern Pacific, their morphological, anatomical, and ecological traits and nuclear markers to evaluate the degree of separation of these MOTUs, assessing introgression and gene flow, are necessary.
A third EPO MOTU corresponds to samples matching the original S. armata description (e.g., the presence of narrow and simple barbels and anterior nasal flaps fringed, and thorns on the head). This MOTU clade is the first to branch off from the American clade and includes samples from Chile and Peru. The last EPO MOTU corresponds to Squatina sp. from northern Peru, a new lineage that represents a potential undescribed new species. The MOTU was related to the clade formed by S. dumeril and the two MOTUs of S. californica, but with a low posterior probability support and their exact evolutionary relationships remain unknown (Figure 3). Additionally, the nearest species to Squatina sp. using K2-P distance was S. guggenheim, with a higher value (1.5%) compared to other nominal Squatina species. An integrative taxonomic study to evaluate if this MOTU corresponds to a new species is imperative. In addition, since there are no taxonomic studies providing a direct comparison of both EPO species, besides the two original descriptions, a revision is necessary of S. armata and S. californica to clarify characteristics and dichotomous identification keys that can help with field identification along the distribution.
The exact distributions of S. californica and S. armata or even of the four MOTUs remain uncertain. Published studies reporting the presence of angel sharks off the coasts of South America are based on literature reviews [16,17] or governmental reports that do not clearly differentiate between both species, reporting angel sharks as Squatina spp. [65]. Similarly, previous distributions reports could be erroneous due to the presence of the cryptic species herein described or by the difficulties during species assignment due to the lack of clear morphological taxonomic characters to distinguish these species.
The limited movement, site fidelity, and preference for coastal areas, as well as other characteristics, such as their large body size and low reproductive output, make angel sharks susceptible to overexploitation by fisheries [28]. In Peru, both S. californica and S. armata have been reported as caught or landed by small scale fisheries by the Institute of Peruvian Sea (Instituto del Mar del Peru in Spanish; IMARPE) [19,65,66]. Before 1996, landings of angel sharks were reported under their common name in Spanish “angelotes” [65]. Nonetheless, some fishing expedition reports, catalogues and identification guides mentioned only the presence of S. armata [19,66,67].
From 1996 to 2010, all fishery landings of angel sharks along the coast were reported by IMARPE as S. californica [65]. These reports showed a marked decline over the period 1996 to 2010 [65]. Currently, the fishery continues and there is still a lack of detailed landing reports of angel sharks in Peru [68]. Managing angel sharks in groups (e.g., genus level) can mask population declines and can represent an impediment to fisheries research but also it may hamper the national enforcement regulation for conservation and management [69]. Landings information along the Peruvian coast (2010–2019) is still reported as Squatina sp. by the Ministry of Production (Ministerio de Producción in Spanish; PRODUCE, Nro Registro 00090925-2020) indicating, at least for the northern region of Peru (i.e., Tumbes, Piura, Lambayeque, and La Libertad regions) a decline of caught specimens reported in tons. Due to the complicated taxonomy of angel sharks added to a potential new, undescribed species, landings reports might be underestimating the population decline of all the species distributed in Peruvian waters. As well as in Peru, in the United States and Mexico, a decline has also been observed [28,70,71,72], for instance, reported landings from US fisheries showed a large decline influenced by fisheries regulation (e.g., minimum landing size and ban on gillnets and trammel nets) [28].
In our species delimitation analysis, that included 19 from the 22 nominal Squatina species, some differences were observed between MOTUs and nominal data that deserve further discussion. All species delimitations split S. africana into four MOTUs. A recent study which included sequences from South Africa and Indian Ocean specimens, found a high number of haplotypes within S. africana [73] using DNA fragments of COI gene, and with a high K2-P genetic distances (up to 2.5%), when compared with values reported in this study between other pairs of nominal Squatina species (e.g., 0.99% between specimens of S. albipunctata and S. pseudocellata). Similar to S. californica, the split of S. africana into 4 MOTUs may be explained due to the life-traits of angel sharks and the geographical barriers. Along the South African coast several barriers to gene flow have been detected, which often coincide with patterns of ocean currents [74]. For example, the South-East and East coasts include three marine ecoregions, the Agulhas Ecoregion, Natal Ecoregion, and part of the Delagoa Ecoregion. Biogeographic forcing agents defining these ecoregions include bathymetric or coastal complexity and currents [75]. This is evident along the east side of South Africa, where a very narrow extension of the continental shelf and wide, deep areas are observed between the northern part of the Agulhas Ecoregion and the Southern part of the Natal Ecoregion. The South-East and East coasts appear to have several population genetic boundaries for several rocky shore and estuarine species (e.g., the fishes Clinus cottoides, Clinus superciliosus, and Muraenoclinus dorsalis) [74,76]. Although previous studies showed a genetic break in these areas, the effect of biogeographic barriers on demersal shark species is still largely unknown and needs further investigation [77]. Finally, the MOTU S. africana 1 was collected in the Indian Ocean close to the Maldives, far from the sampling area of MOTUs S. africana 3 and S. africana 4 which is characterized by different oceanographic and geomorphologic features.
Samples of S. nebulosa (sampling location in Japan and Thailand) and S. formosa (sampling location in China and Sea of South China) from BOLD System were clustered in the same MOTU sharing the same haplotype. Diagnosing both species is particularly challenging, their subtle external morphological differences may cause an overlap among many characteristics used [31]. Due to the difficulties in the correct identification of these species, the source from where we retrieved the sequences (i.e., GenBank which lacks physical voucher information) and the fact the taxonomy for these individuals were not revised, our result needs to be tested in an integrative taxonomic study including morphology and genetic analyses.
Finally, our results have far-reaching implications for the management and conservation policies of angel sharks in EPO, suggesting the presence of an undescribed species inhabiting the northern Peruvian coast, living in sympatry with the two species already reported. Many species of angel sharks are affected by artisanal and industrial fisheries and their populations have declined around the world [28]. All efforts to quantify the impact of fisheries in South American angel sharks will be unsuccessful without a precise identification of the species. The use of molecular approaches, such as DNA barcoding, has a potential to quickly identify undescribed species in poorly studied regions like the Southeast Pacific [78]; and in ecologically and economically important groups like Elasmobranchii that hold a high level of taxonomic uncertainties [78].
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/d13050177/s1, Table S1: Complete list of COI sequences of angel sharks retrieved from BOLD System and used in this study, including collection location and access numbers Bold System and GenBank Result S1: Final matrix containing all the DNA sequences worked on this paper.
Author Contributions
Conceptualization, R.M.C.-A. and J.L.R.; methodology R.M.C.-A.; J.L.R.; formal analysis, R.M.C.-A. and J.L.R.; investigation, R.M.C.-A.; resources, R.S.-R. and J.C.M.; data curation, J.A.-S.; writing—original draft preparation, R.M.C.-A., C.O.-A., R.S.-R., E.A.-C., and J.L.R.; writing—review and editing, J.A.-S., J.C.M., C.O.-A., R.S.-R., E.A.-C., X.V.-Z., C.Y., and J.L.R. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by the Consejo Nacional de Ciencia, Tecnología e Innovación Tecnológica (CONCYTEC)–Peru (Círculo de Investigación 023-2016-FONDECYT) and by the Vicerrectorado de Investigación de la Universidad Nacional Mayor de San Marcos (Número de proceso B18100094).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
We are grateful to the Department of Ichthyology of at the Natural History Museum of San Marcos University for housing the specimen of angel shark collected for this study.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Compagno, L.J.V. FAO Species Catalogue—Volume 4, Sharks of the World: An Annotated and Illustrated Catalogue of Shark Species Known to Date. Part 1—Hexanchiformes to Lamniformes; FAO: Rome, Italy, 1984; pp. 503–512. [Google Scholar]
- Compagno, L.J.V. Sharks of the World: An Annotated and Illustrated Catalogue of Shark Species Known to Date—Volume 2, Bullhead, Mackerel and Carpet Sharks (Heterodontiformes, Lamniformes and Orectolobiformes); FAO Species Catalogue for Fishery Purposes: Rome, Italy, 2001; Volume 2, p. 278. [Google Scholar]
- Vélez-Zuazo, X.; Agnarsson, I. Molecular Phylogenetics and Evolution Shark tales: A molecular species-level phylogeny of sharks (Selachimorpha, Chondrichthyes). Mol. Phylogenet. Evol. 2011, 58, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Stelbrink, B.; Von Rintelen, T.; Cliff, G.; Kriwet, J. Molecular systematics and global phylogeography of angel sharks (genus Squatina). Mol. Phylogenet. Evol. 2010, 54, 395–404. [Google Scholar] [CrossRef] [PubMed]
- Acero, P.A.; Tavera, J.J.; Anguila, R.; Hernández, L. A New Southern Caribbean Species of Angel Shark (Chondrichthyes, Squaliformes, Squatinidae), Including Phylogeny and Tempo of Diversification of American Species. Copeia 2016, 104, 577–585. [Google Scholar] [CrossRef]
- Vaz, D.F.B.; De Carvalho, M.R. New Species of Squatina (Squatiniformes: Squatinidae) from Brazil, with Comments on the Taxonomy of Angel Sharks from the Central and Northwestern Atlantic. Copeia 2018, 106, 144–160. [Google Scholar] [CrossRef]
- Weigmann, S. Annotated checklist of the living sharks, batoids and chimaeras (Chondrichthyes) of the world, with a focus on biogeographical diversity. J. Fish Biol. 2016, 88, 837–1037. [Google Scholar] [CrossRef]
- Castro-Aguirre, J.L.; Pérez, H.E.; Campos, L.H. Dos nuevas especies del género Squatina (Chondrichthyes: Squatinidae) del Golfo de México. Rev. Biol. Trop. 2006, 54, 1031–1040. [Google Scholar] [PubMed]
- Fricke, R.; Eschmeyer, W.N.; Van Der Laan, R. (Eds.) Eschmeyer’s Catalog of Fishes: Genera, Species, References. Available online: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp (accessed on 5 January 2021).
- Ebert, D.A.; Fowler, S.L.; Compagno, L.J. Sharks of the World: A Fully Illustrated Guide; Wild Nature Press: Plymouth, UK, 2013. [Google Scholar]
- Del Moral-Flores, L.F.; Morrone, J.J.; Durand, J.A.; Espinosa-Pérez, H.; De León Pérez-Ponce, G. Lista patrón de los tiburones, rayas y quimeras de México. Arx. Miscellánia Zoológica 2015, 13, 47–163. [Google Scholar] [CrossRef]
- Ehemann, N.R.; del González-González, L.V.; Chollet-Villalpando, J.G.; De La Cruz-Agüero, J. Updated checklist of the extant Chondrichthyes within the Exclusive Economic Zone of Mexico. ZooKeys 2018, 774, 17–39. [Google Scholar] [CrossRef] [PubMed]
- Ramírez-Amaro, S.; Ramírez-Macías, D.; Vázquez-Juárez, R.; Flores-Ramírez, S.; Galván-Magaña, F.; Gutiérrez-Rivera, J.N. Estructura poblacional del tiburón ángel del Pacífico (Squatina californica) a lo largo de la costa noroccidental de México con base en la regíon control del ADN mitochondrial. Ciencias Mar. 2017, 43, 69–80. [Google Scholar] [CrossRef]
- Jacquet, J.; Alava, J.J.; Pramod, G.; Henderson, S.; Zeller, D. In hot soup: Sharks captured in Ecuador’s waters. Environ. Sci. 2008, 5, 269–283. [Google Scholar] [CrossRef]
- Chirichigno, N.; Cornejo, R. Catalogo Comentado Peces Marinos del Perú; Instituto del Mar del Peru: Callao, Peru, 2001; p. 217. [Google Scholar]
- Bustamante, C.; Vargas-Caro, C.; Bennett, M.B. Not all fish are equal: Functional biodiversity of cartilaginous fishes (Elasmobranchii and Holocephali) in Chile. J. Fish Biol. 2014, 85, 1617–1633. [Google Scholar] [CrossRef] [PubMed]
- Cornejo, R.; Vélez-Zuazo, X.; González-Pestana, A.; Kouri, C.; Mucientes, G. An updated checklist of Chondrichthyes from the southeast Pacific off Peru. Check List 2015, 17. [Google Scholar] [CrossRef]
- Calle-Morán, M.D.; Béarez, P. Updated checklist of marine cartilaginous fishes from continental and insular Ecuador (Tropical Eastern Pacific Ocean). Cybium Rev. Int. d’Ichtyologie. 2020, 44, 239–250. [Google Scholar]
- Mejía-Gallegos, J.; Flores-Portugal, L.A.; Segura, G. Exploración sobre Recursos Costeros y Recursos Demersales; Crucero 7104 B/I SNP 1; Instituto del Mar del Peru: Callao, Peru, 1971; p. 16. [Google Scholar]
- Espinoza, M.; Diaz, E.; Angulo, A.; Hernández, S.; Clarke, T.M. Chondrichthyan Diversity, Conservation Status, and Management Challenges in Costa Rica. Front. Mar. Sci. 2018, 5, 1–15. [Google Scholar] [CrossRef]
- Cailliet, G.M.; Chabot, C.L.; Nehmens, M.C.; Carlsle, A.B. Squatina californica (amended version of 2016 assessment). Available online: https://www.iucnredlist.org/species/39328/177163701 (accessed on 20 February 2021).
- Dulvy, N.K.; Acuña, E.; Bustamante, C.; Cavallos, A.; Herman, K.; Navia, A.F.; Pardo, S.A.; Velez-Zuazo, X. Squatina armata. Available online: https://www.iucnredlist.org/species/44571/116831653 (accessed on 20 February 2021).
- Chirichigno, N.; Velez, J. Clave para Identificar los Peces Marinos del Perú; Instituto del Mar del Perú: Callao, Peru, 1998; p. 31. [Google Scholar]
- Last, P.R.; White, W.T. Three new angel sharks (Chondrichthyes: Squatinidae) from the Indo-Australian region. Zootaxa 2008, 1734, 1–26. [Google Scholar] [CrossRef]
- Kato, S.; Springer, S.; Wagner, M.H. Field Guide to Eastern Pacific and Hawaiian Sharks; United States Department of the Interior: Washington, DC, USA, 1967; Volume 271, 47p.
- Ayres, W.O. On new fishes of the California coast. Proc. Calif. Acad. Sci. 1859, 2, 25–32. [Google Scholar]
- Philippi, R.A. Sobre los tiburones y algunos otros peces de Chile. An. Univ. Chile 1887, 71, 535–574. [Google Scholar]
- Ellis, J.R.; Barker, J.; McCully Phillips, S.R.; Meyers, E.K.M.; Heupel, M. Angel sharks (Squatinidae): A review of biological knowledge and exploitation. J. Fish Biol. 2021, 98, 592–621. [Google Scholar] [CrossRef]
- Domingues, R.R.; Hilsdorf, A.W.S.; Gadig, O.B.F. The importance of considering genetic diversity in shark and ray conservation policies. Conserv. Genet. 2017, 19, 501–525. [Google Scholar] [CrossRef]
- Hosegood, J.; Humble, E.; Ogden, R.; De Bruyn, M.; Creer, S.; Stevens, G.M.W.; Abudaya, M.; Bassos-Hull, K.; Bonfil, R.; Fernando, D.; et al. Phylogenomics and species delimitation for effective conservation of manta and devil rays. Mol. Ecol. 2020, 29, 4783–4796. [Google Scholar] [CrossRef]
- Walsh, J.H.; Ebert, D.A. A review of the systematics of western North Pacific angel sharks, genus Squatina, with redescriptions of Squatina formosa, S. japonica, and S. nebulosa (Chondrichthyes: Squatiniformes, Squatinidae). Zootaxa 2007, 1551, 31–47. [Google Scholar] [CrossRef]
- Walsh, J.H.; Ebert, D.A.; Compagno, L.J.V. Squatina caillieti sp. nov., a new species of angel shark (Chondrichthyes: Squat-iniformes: Squatinidae) from the Philippine Islands. Zootaxa 2011, 59, 49–59. [Google Scholar] [CrossRef]
- Theiss, S.M.; Ebert, D.A. Lost and found: Recovery of the holotype of the ocellated angelshark, Squatina tergocellatoides Chen, 1963 (Squatinidae), with comments on western Pacific squatinids. Zootaxa 2013, 3752, 73–85. [Google Scholar] [CrossRef]
- Vaz, D.F.B.; De Carvalho, M.R. Morphological and taxonomic revision of species of Squatina from the Southwestern Atlantic Ocean (Chondrichthyes: Squatiniformes: Squatinidae). Zootaxa 2013, 3695, 1–81. [Google Scholar] [CrossRef] [PubMed]
- Ward, R.D.; Zemlak, T.S.; Innes, B.H.; Last, P.R.; Hebert, P.D. DNA barcoding Australia’s fish species. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2005, 360, 1847–1857. [Google Scholar] [CrossRef]
- Ward, R.D.; Holmes, B.H.; White, W.T.; Last, P.R. DNA barcoding Australasian chondrichthyans: Results and potential uses in conservation. Mar. Freshw. Res. 2008, 59, 57–71. [Google Scholar] [CrossRef]
- Velez-Zuazo, X.; Alfaro-Shigueto, J.; Mangel, J.; Papa, R.; Agnarsson, I. What barcode sequencing reveals about the shark fishery in Peru. Fish. Res. 2015, 161, 34–41. [Google Scholar] [CrossRef]
- Borsa, P.; Arlyza, I.S.; Hoareau, T.B.; Shen, K.-N. Diagnostic description and geographic distribution of four new cryptic species of the blue-spotted maskray species complex (Myliobatoidei: Dasyatidae; Neotrygon spp.) based on DNA sequences. J. Oceanol. Limnol. 2017, 36, 827–841. [Google Scholar] [CrossRef]
- Smart, J.J.; Chin, A.; Baje, L.; Green, M.E.; Appleyard, S.A.; Tobin, A.J.; Simpfendorfer, C.A.; White, W.T. Effects of Including Misidentified Sharks in Life History Analyses: A Case Study on the Grey Reef Shark Carcharhinus amblyrhynchos from Papua New Guinea. PLoS ONE 2016, 11, e0153116. [Google Scholar] [CrossRef]
- Cariani, A.; Messinetti, S.; Ferrari, A.; Arculeo, M.; Bonello, J.J.; Bonnici, L.; Cannas, R.; Carbonara, P.; Cau, A.; Charilaou, C.; et al. Improving the Conservation of Mediterranean Chondrichthyans: The ELASMOMED DNA Barcode Reference Library. PLoS ONE 2017, 12, e0170244. [Google Scholar] [CrossRef]
- Bernardo, C.; Corrêa de Lima Adachi, A.M.; Paes da Cruz, V.; Foresti, F.; Loose, R.H.; Bornatowski, H. The label “Cação” is a shark or a ray and can be a threatened species! Elasmobranch trade in Southern Brazil unveiled by DNA barcoding. Mar. Policy 2020, 116, 103920. [Google Scholar] [CrossRef]
- Bunholi, I.V.; da Silva Ferrette, B.L.; De Biasi, J.B.; de Oliveira Magalhães, C.D.; Rotundo, M.M.; Oliveira, C.; Foresti, F.; Mendonça, F.F. The fishing and illegal trade of the angelshark: DNA barcoding against misleading identifications. Fish. Res. 2018, 206, 193–197. [Google Scholar] [CrossRef]
- Vella, A.; Vella, N.; Schembri, S. A molecular approach towards taxonomic identification of elasmobranch species from Maltese fisheries landings. Mar. Genom. 2017, 36, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Navia, A.F.; Mejía-Falla, P. Checklist of marine elasmobranchs of Colombia. Univ. Sci. 2019, 24, 241–276. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets Brief Communication. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Stoeckle, M.Y.; Das Mishu, M.; Charlop-Powers, Z. GoFish: A streamlined environmental DNA presence/absence assay for marine vertebrates. bioRixv 2018, 331322. [Google Scholar] [CrossRef]
- Steinke, D.; Connell, A.D.; Hebert, P.D. Linking adults and immatures of South African marine fishes. Genome 2016, 59, 959–967. [Google Scholar] [CrossRef]
- Almerón-Souza, F.; Sperb, C.; Castilho, C.L.; Figueiredo, P.I.C.C.; Gonçalves, L.T.; Machado, R.; Oliveira, L.R.; Valiati, V.H.; Fagundes, N.J.R. Molecular Identification of Shark Meat from Local Markets in Southern Brazil Based on DNA Barcoding: Evidence for Mislabeling and Trade of Endangered Species. Front. Genet. 2018, 9, 1–12. [Google Scholar] [CrossRef]
- Fitzpatrick, C.K.; Finnegan, K.A.; Osaer, F.; Narváez, K.; Shivji, M.S. The complete mitochondrial genome of the Critically Endangered Angelshark, Squatina squatina. Mitochondrial DNA Part B Resour. 2017, 2, 212–213. [Google Scholar] [CrossRef]
- Moftah, M.; Aziz, S.H.A.; Elramah, S.; Favereaux, A. Classification of Sharks in the Egyptian Mediterranean Waters Using Morphological and DNA Barcoding Approaches. PLoS ONE 2011, 6, e27001. [Google Scholar] [CrossRef]
- Gao, Y.; Liu, T.; Wei, T.; Geng, X.; Wang, J.; Ma, H. Complete mitochondrial genome of Clouded angelshark (Squatina nebulosa). Mitochondrial DNA Part A 2016, 27, 1599–1600. [Google Scholar] [CrossRef]
- De Oliveira Ribeiro, A.; Caires, R.A.; Mariguela, T.C.; Pereira, L.H.G.; Hanner, R.; Oliveira, C. DNA barcodes identify marine fishes of São Paulo State, Brazil. Mol. Ecol. Resour. 2012, 12, 1012–1020. [Google Scholar] [CrossRef]
- Keskin, E.; Atar, H.H. DNA barcoding commercially important fish species of Turkey. Mol. Ecol. Resour. 2013, 13, 788–797. [Google Scholar] [CrossRef]
- Wang, Z.-D.; Guo, Y.-S.; Liu, X.-M.; Fan, Y.-B.; Liu, C.-W. DNA barcoding South China Sea fishes. Mitochondrial DNA 2012, 23, 405–410. [Google Scholar] [CrossRef]
- Sembiring, A.; Pertiwi, N.P.D.; Mahardini, A.; Wulandari, R.; Kurniasih, E.M.; Kuncoro, A.W.; Cahyani, N.D.; Anggoro, A.W.; Ulfa, M.; Madduppa, H.; et al. DNA barcoding reveals targeted fisheries for endangered sharks in Indonesia. Fish. Res. 2015, 164, 130–134. [Google Scholar] [CrossRef]
- Fujisawa, T.; Barraclough, T.G. Delimiting Species Using Single-Locus Data and the Generalized Mixed Yule Coalescent Approach: A Revised Method and Evaluation on Simulated Data Sets. Syst. Biol. 2013, 62, 707–724. [Google Scholar] [CrossRef] [PubMed]
- Yule, G.U. A Mathematical Theory of Evolution, Based on the Conclusions of Dr. J.C. Willis, F.R.S. Philos. Trans. R. Soc. London Ser. B Contain. Pap. Biol. Character 1925, 213, 21–87. [Google Scholar]
- Hudson, R.R. Gene genealogies and the coalescent process. Oxford Surv. Evol. Biol. 1990, 7, 44. [Google Scholar]
- Zhang, J.; Kapli, P.; Pavlidis, P.; Stamatakis, A. A general species delimitation method with applications to phylogenetic placements. Bioinformatics 2013, 29, 2869–2876. [Google Scholar] [CrossRef] [PubMed]
- Bouckaert, R.; Heled, J.; Kühnert, D.; Vaughan, T.; Wu, C.-H.; Xie, D.; Suchard, M.A.; Rambaut, A.; Drummond, A.J. BEAST 2: A Software Platform for Bayesian Evolutionary Analysis. PLoS Comput. Biol. 2014, 10, e1003537. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the 2010 Gateway Computing Environments Workshop (GCE), IEEE, New Orleans, LA, USA, 14 November 2010; pp. 1–8. [Google Scholar] [CrossRef]
- Gaida, I.H. Evolutionary Aspects of Gene Expression in the Pacific Angel Shark, Squatina california (Squatiniformes: Squatinidae). Copeia 1995, 532–554. [Google Scholar] [CrossRef]
- Gonzalez-Pestana, A.; Kouri, C.; Velez-Zuazo, X. Shark fisheries in the Southeast Pacific: A 61-year analysis from Peru. F1000Research 2014, 3, 164. [Google Scholar] [CrossRef] [PubMed]
- IMARPE. Resultados Preliminares del Primer Crucero de Exploración Pesquera del SNP-1 6901; Instituto del Mar del Perú: Callao, Peru, 1969; pp. 11–12. [Google Scholar]
- Chirichigno, N. Clave para identificar los peces marinos del Perú; Instituto del Mar del Perú: Callao, Peru, 1974; p. 30. [Google Scholar]
- Bartholomew, D.C.; Mangel, J.C.; Alfaro-Shigueto, J.; Pingo, S.; Jimenez, A.; Godley, B.J. Remote electronic monitoring as a potential alternative to on-board observers in small-scale fisheries. Biol. Conserv. 2018, 219, 35–45. [Google Scholar] [CrossRef]
- Cashion, M.S.; Bailly, N.; Pauly, D. Official catch data underrepresent shark and ray taxa caught in Mediterranean and Black Sea fisheries. Mar. Policy 2019, 105, 1–9. [Google Scholar] [CrossRef]
- Leet, W.S.; Dewees, C.M.; Klingbeil, R.; Larson, E.J. California’s Living Marine Resources: A Status Report 2001. Available online: https://wildlife.ca.gov/Conservation/Marine/Status/2001 (accessed on 18 January 2021).
- Pondella, D.J., II; Allen, L.G. The nearshore fish assemblage of Santa Catalina Island. In Proceedings of the Fifth California Islands Symposium, Santa Rosa Island, CA, USA, 29 March–1 April 1999; pp. 394–400. [Google Scholar]
- Bizzarro, J.J.; Smith, W.D.; Hueter, R.E.; Villavicencio–Garayzar, C.J. Activities and Catch Composition of Artisanal Elasmobranch Fishing Sites on the Eastern Coast of Baja California Sur, Mexico. Bull. South Calif. Acad. Sci. 2009, 108, 137–151. [Google Scholar] [CrossRef]
- Ambily, M.N.; Zacharia, P.U.; Najmudeen, T.M.; Ambily, L.; Sunil, K.T.S.; Radhakrishnan, M.; Kishor, T.G. First Record of African Angel Shark, Squatina africana (Chondricthyes: Squatinidae) in Indian Waters, Confirmed by DNA Barcoding. J. Ichthyol. 2018, 58, 312–317. [Google Scholar] [CrossRef]
- Von Der Heyden, S. Why do we need to integrate population genetics into South African marine protected area planning? Afr. J. Mar. Sci. 2009, 31, 263–269. [Google Scholar] [CrossRef]
- Spalding, M.D.; Fox, H.E.; Allen, G.R.; Davidson, N.; Ferdaña, Z.A.; Finlayson, M.; Halpern, B.S.; Jorge, M.A.; Lombana, A.; Lourie, S.A.; et al. Marine Ecoregions of the World: A Bioregionalization of Coastal and Shelf Areas. Bioscience 2007, 57, 573–583. [Google Scholar] [CrossRef]
- Von Der Heyden, S.; Prochazka, K.; Bowie, R.C.K.; Prochazka, K. Significant population structure and asymmetric gene flow patterns amidst expanding populations of Clinus cottoides (Perciformes, Clinidae): Application of molecular data to marine conservation planning in South Africa. Mol. Ecol. 2008, 17, 4812–4826. [Google Scholar] [CrossRef] [PubMed]
- Van der Merwe, A.E.B.; Gledhill, K.S. Molecular species identification and population genetics of chondrichthyans in South Africa: Current challenges, priorities and progress. Afr. Zoöl 2015, 50, 205–217. [Google Scholar] [CrossRef]
- Ramirez, J.L.; Rosas-Puchuri, U.; Cañedo, R.M.; Alfaro-Shigueto, J.; Ayon, P.; Zelada-Mázmela, E.; Siccha-Ramirez, R.; Velez-Zuazo, X. DNA barcoding in the Southeast Pacific marine realm: Low coverage and geographic representation despite high diversity. PLoS ONE 2020, 15, e0244323. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).