Estimated Density, Population Size and Distribution of the Endangered Western Hoolock Gibbon (Hoolock hoolock) in Forest Remnants in Bangladesh
Abstract
:1. Introduction
2. Materials and Methods
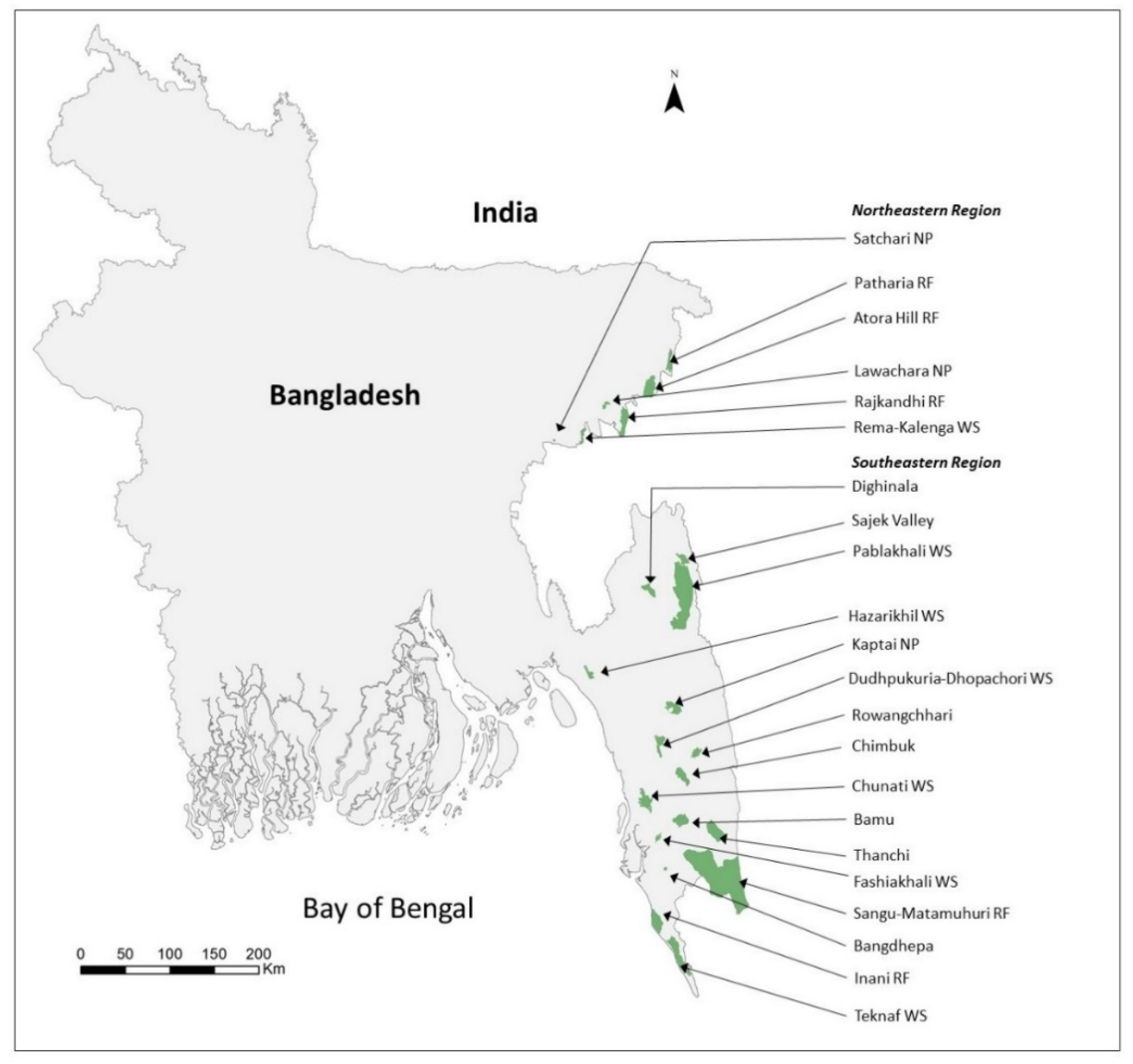
2.1. Study Sites
2.2. Population Surveys
2.3. Modelling Group Density
2.4. Habitat Suitability and Population Size
2.5. Preprocessing of Variables for Habitat Assessment
2.6. Modeling Procedures
2.7. Evaluation of Model Performance
2.8. Habitat Suitability and Spatial Analysis
3. Results
3.1. Group Density and Population Estimates
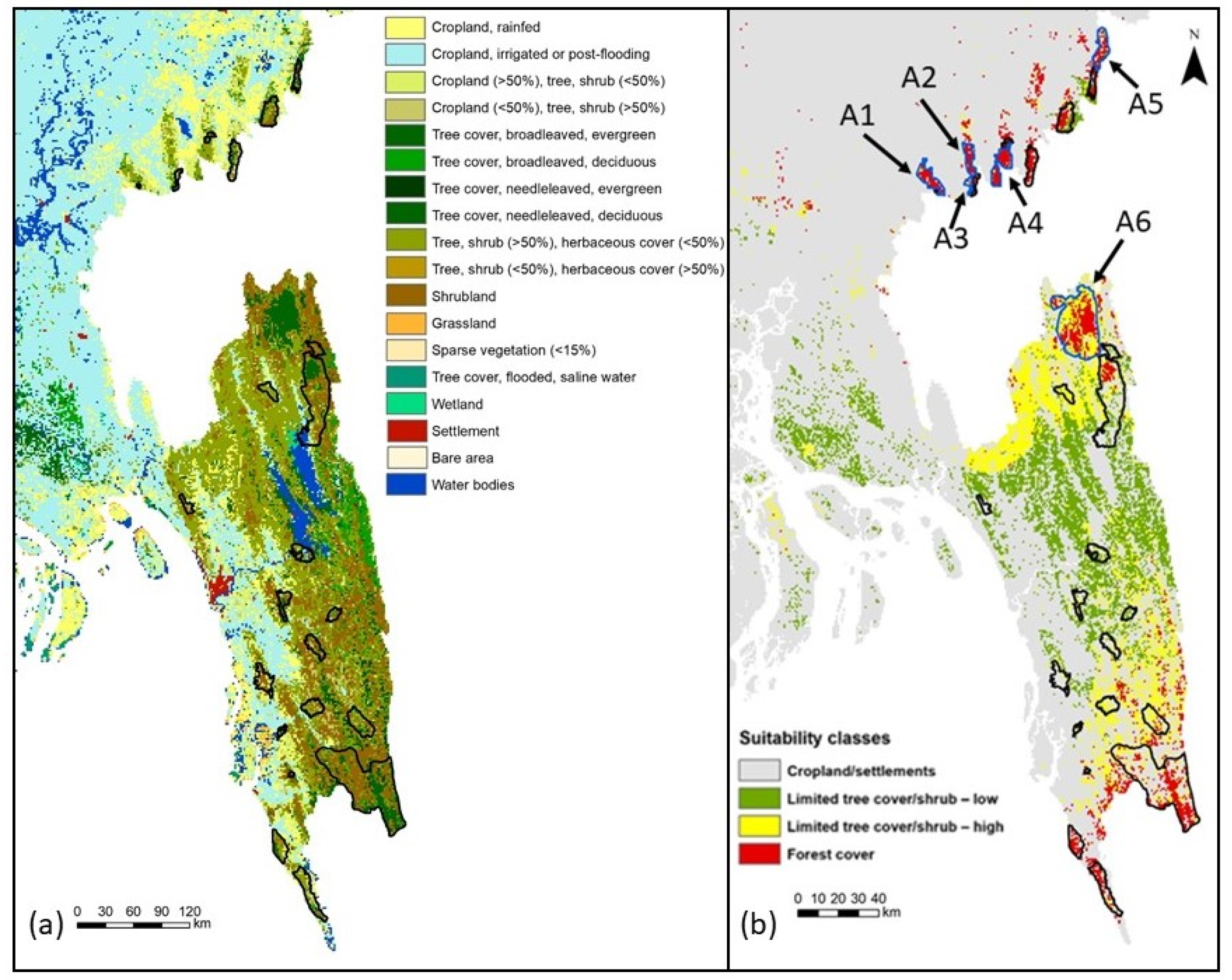
3.2. Model Evaluation and Habitat Quality
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- FAO. State of the World’s Forests 2016. Forests and Agriculture: Land-Use Challenges and Opportunities; Food and Agriculture Organization of the United Nations: Rome, Italy, 2016. [Google Scholar]
- Slik, J.F.; Franklin, J.; Arroyo-Rodríguez, V.; Field, R.; Aguilar, S.; Aguirre, N.; Ahumada, J.; Aiba, S.I.; Alves, L.F.; Anitha, K.; et al. Phylogenetic classification of the world’s tropical forests. Proc. Natl. Acad. Sci. USA 2018, 115, 1837–1842. [Google Scholar] [CrossRef] [Green Version]
- Ahsan, M.M.; Aziz, N.; Morshed, H.M. Assessment of Management Effectiveness of Protected Areas of Bangladesh. SRCWP Project; Bangladesh Forest Department: Dhaka, Bangladesh, 2016. [Google Scholar]
- Potapov, P.; Siddiqui, B.N.; Iqbal, Z.; Aziz, T.; Zzaman, B.; Islam, A.; Pickens, A.; Talero, Y.; Tyukavina, A.; Turubanova, S.; et al. Comprehensive monitoring of Bangladesh tree cover inside and outside of forests, 2000–2014. Environ. Res. Lett. 2017, 12, 104015. [Google Scholar] [CrossRef]
- Champion, H.G.; Seth, S.K. A Revised Survey of the Forest Types of India; Manager of Publications: Delhi, India, 1968. [Google Scholar]
- Muzaffar, S.B.; Islam, M.A.; Feeroz, M.M.; Kabir, M.; Begum, S.; Mahmud, M.S.; Hasan, M.K. Habitat characteristics of the endangered hoolock gibbons of Bangladesh: The role of plant species richness. Biotropica 2007, 39, 539–545. [Google Scholar] [CrossRef]
- Islam, M.A.; Feeroz, M.M.; Muzaffar, S.B.; Kabir, M.; Begum, S.; Hasan, K.; Mahmud, S.; Chakma, S. Population status and conservation of Hoolock Gibbons (Hoolock hoolock Harlan 1834) in Bangladesh. J. Bombay Nat. Hist. Soc. 2008, 105, 19–23. [Google Scholar]
- Muzaffar, S.B.; Islam, M.A.; Kabir, D.S.; Khan, M.H.; Ahmed, F.U.; Chowdhury, G.W.; Aziz, M.A.; Chakma, S.; Jahan, I. The endangered forests of Bangladesh: Why the process of implementation of the Convention on Biological Diversity is not working. Biod. Cons. 2011, 20, 1587–1601. [Google Scholar] [CrossRef]
- Chivers, D.J. The swinging singing apes: Fighting for food and family in far-east forests. In The Apes: Challenges for the 21st Century, Conference Proceedings; Sodaro, V., Sodaro, C., Eds.; Brookfield Zoo: Brookfield, IL, USA, 2001; pp. 1–28. [Google Scholar]
- Mittermeier, R.A.; Ratsimbazafy, J.; Rylands, A.B.; Williamson, L.; Oates, J.F.; Mbora, D.; Ganzhorn, J.U.; Rodríguez-Luna, E.; Palacios, E.; Heymann, E.W.; et al. Primates in peril: The world’s 25 most endangered primates, 2006–2008. Primate Cons. 2007, 22, 1–40. [Google Scholar] [CrossRef]
- Thinh, V.N.; Mootnick, A.R.; Geissmann, T.; Li, M.; Ziegler, T.; Agil, M.; Moisson, P.; Nadler, T.; Walter, L.; Roos, C. Mitochondrial evidence for multiple radiations in the evolutionary history of small apes. BMC Evol. Biol. 2010, 10, 1–13. Available online: http://www.biomedcentral.com/1471-2148/10/74 (accessed on 26 June 2021). [CrossRef] [Green Version]
- Schwitzer, C.; Mittermeier, R.A.; Rylands, A.B.; Chiozza, F.; Williamson, E.A.; Wallis, J.; Cotton, A. Primates in Peril: The World’s 25 Most Endangered Primates 2014–2016; IUCN SSC Primate Specialist Group (PSG): London, UK; International Primatological Society (IPS): Chicago, IL, USA; Conservation International (CI): Virginia, VA, USA; Bristol Zoological Society: Bristol, UK, 2015. [Google Scholar]
- McConkey, K.R.; O’Farrill, G. Loss of seed dispersal before the loss of seed dispersers. Biol. Conserv. 2016, 201, 38–49. [Google Scholar] [CrossRef]
- McConkey, K.R. Seed dispersal by primates in Asian habitats: From species, to communities, to conservation. Int. J. Primatol. 2018, 39, 466–492. [Google Scholar] [CrossRef]
- Geissmann, T.; Grindley, M.E.; Lwin, N.; Aung, S.S.; Aung, T.N.; Htoo, S.B.; Momberg, F. The Conservation Status of Hoolock Gibbons in Myanmar; Gibbon Conservation Alliance: Zurich, Switzerland, 2013. [Google Scholar]
- Fan, P.F.; He, K.; Chen, X.; Ortiz, A.; Zhang, B.; Zhao, C.; Li, Y.Q.; Zhang, H.B.; Kimock, C.; Wang, W.Z.; et al. Description of a new species of Hoolock gibbon (Primates: Hylobatidae) based on integrative taxonomy. Am. J. Primatol. 2017, 79, e22631. [Google Scholar] [CrossRef] [PubMed]
- Fan, P.F.; Turvey, S.T.; Bryant, J.V. Hoolock tianxing (amended version of 2019 assessment). IUCN Red List. Threat. Species 2020, e.T118355648A166597159. Available online: https://dx.doi.org/10.2305/IUCN.UK.2020-1.RLTS.T118355648A166597159.en (accessed on 21 June 2021). [CrossRef] [Green Version]
- Molur, S.; Walker, S.; Islam, A.; Miller, P.; Srinivasulu, C.; Nameer, P.O.; Daniel, B.A.; Ravikumar, L. Conservation of Western Hoolock Gibbon (Hoolock hoolock hoolock) in India and Bangladesh: Population and Habitat Viability Assessment (PHVA) Workshop Report; Coimbatore, Zoo Outreach Organization/CBSG-South Asia: Coimbatore, India, 2005. [Google Scholar]
- IUCN Bangladesh. Red List of Bangladesh Volume 1: Summary; IUCN (International Union for Conservation of Nature), Bangladesh Country Office: Dhaka, Bangladesh, 2015. [Google Scholar]
- Molur, S.; Brandon–Jones, D.; Dittus, W.; Eudey, A.; Kumar, A.; Singh, M.; Feeroz, M.M.; Chalise, M.; Priya, P.; Walker, S. Status of South. Asian Primates: Conservation Assessment and Management Plan. (C.A.M.P.) Workshop Report; Zoo Outreach Organization: Tamil Nadu, India; CBSG-South Asia: Coimbatore, India, 2003. [Google Scholar]
- Alliance, C.C. A Preliminary Wildlife Survey in Sangu-Matamuhuri Reserve Forest, Chittagong Hill Tracts, Bangladesh; Unpublished report submitted to Bangladesh Forest Department: Dhaka, Bangladesh, 2016; p. 52. [Google Scholar]
- Gittins, S.P.; Akonda, A.W. What survives in Bangladesh? Oryx 1982, 16, 275–282. [Google Scholar] [CrossRef]
- Feeroz, M.M.; Islam, M.A. Ecology and Behaviour of Hoolock Gibbons of Bangladesh; Multidisciplinary Action Research Centre (MARC): Dhaka, Bangladesh, 1992; p. 76. [Google Scholar]
- Das, J.; Feeroz, M.M.; Islam, M.A.; Biswas, J.; Bujarborua, P.; Chetry, D.; Medhi, R.; Bose, J. Distribution of hoolock gibbon (Bunopithecus hoolock hoolock) in India and Bangladesh. Zoos’ Print J. 2003, 18, 969–976. [Google Scholar] [CrossRef]
- Rawson, B.M.; Clements, T.; Hor, N.M. Status and conservation of yellow-cheeked crested gibbons (Nomascusgabriellae) in the Seima Biodiversity Conservation Area, Mondulkiri Province, Cambodia. In The Gibbons; Springer: New York, NY, USA, 2009; pp. 387–408. [Google Scholar] [CrossRef]
- Cheyne, S.M.; Gilhooly, L.J.; Hamard, M.C.; Höing, A.; Houlihan, P.R.; Loken, B.; Phillips, A.; Rayadin, Y.; Capilla, B.R.; Rowland, D.; et al. Population mapping of gibbons in Kalimantan, Indonesia: Correlates of gibbon density and vegetation across the species’ range. Endang. Spec. Res. 2016, 30, 133–143. [Google Scholar] [CrossRef] [Green Version]
- Phillips, S.J.; Dudík, M. Modeling of species distributions with Maxent: New extensions and a comprehensive evaluation. Ecography 2008, 31, 161–175. [Google Scholar] [CrossRef]
- Bradie, J.; Leung, B. A quantitative synthesis of the importance of variables used in MaxEnt species distribution models. J. Biogeog. 2017, 44, 1344–1361. [Google Scholar] [CrossRef]
- Rabelo, R.M.; Goncalves, J.R.; Silva, F.E.; Rocha, D.G.; Canale, G.R.; Bernardo, C.S.; Boubli, J.P. Predicted distribution and habitat loss for the endangered black-faced black spider monkey Ateles chamek in the Amazon. Oryx 2020, 54, 699–705. [Google Scholar] [CrossRef]
- Sales, L.P.; Ribeiro, B.R.; Pires, M.M.; Chapman, C.A.; Loyola, R. Recalculating route: Dispersal constraints will drive the redistribution of Amazon primates in the Anthropocene. Ecography 2019, 42, 1789–1801. [Google Scholar] [CrossRef] [Green Version]
- Campos, F.A.; Jack, K.M. A potential distribution model and conservation plan for the critically endangered Ecuadorian capuchin, Cebus albifronsaequatorialis. Int. J. Primatol. 2013, 34, 899–916. [Google Scholar] [CrossRef]
- Cavalcante, T.; de Souza Jesus, A.; Rabelo, R.M.; Messias, M.R.; Valsecchi, J.; Ferraz, D.; Gusmao, A.C.; da Silva, O.D.; Faria, L.; Barnett, A.A. Niche overlap between two sympatric frugivorous Neotropical primates: Improving ecological niche models using closely-related taxa. Biod. Cons. 2020, 29, 2749–2763. [Google Scholar] [CrossRef]
- Moraes, B.; Razgour, O.; Souza-Alves, J.P.; Boubli, J.P.; Bezerra, B. Habitat suitability for primate conservation in north-east Brazil. Oryx 2020, 54, 803–813. [Google Scholar] [CrossRef]
- Sarma, K.; Kumar, A.; Krishna, M.; Medhi, M.; Tripathi, O.P. Predicting Suitable Habitats for the Vulnerable Eastern Hoolock Gibbon, Hoolock leuconedys, in India Using the MaxEnt Model. Folia Primatol. 2015, 86, 387–397. [Google Scholar] [CrossRef]
- Alamgir, M.; Mukul, S.A.; Turton, S.M. Modelling spatial distribution of critically endangered Asian elephant and Hoolock gibbon in Bangladesh forest ecosystems under a changing climate. Appl. Geogr. 2015, 60, 10–19. [Google Scholar] [CrossRef]
- Sarma, K.; Saikia, M.K.; Sarania, B.; Basumatary, H.; Baruah, S.S.; Saikia, B.P.; Kumar, A.; Saikia, P.K. Habitat monitoring and conservation prioritization of Western Hoolock Gibbon in upper Brahmaputra Valley, Assam, India. Sci. Rep. 2021, 11, 15427. [Google Scholar] [CrossRef]
- Stan Development Team. RStan: The R Interface to Stan. R Package Version 2.21.2. 2020. Available online: http://mc-stan.org/ (accessed on 11 August 2021).
- Jaynes, E.T. Probability Theory: The Logic. of Science; Cambridge University Press: Cambridge, UK, 2003. [Google Scholar]
- Sollmann, R.; Gardner, B.; Williams, K.A.; Gilbert, A.T.; Veit, R.R. A hierarchical distance sampling model to estimate abundance and covariate associations of species and communities. Methods Ecol. Evol. 2003, 7, 529–537. [Google Scholar] [CrossRef]
- ESA (European Space Agency). Oxfordshire. Available online: http://maps.elie.ucl.ac.be/CCI/viewer/download.php (accessed on 9 January 2021).
- ESRI (Environmental Systems Research Institute). ArcGIS Desktop; Release 10.8.1; Environmental Systems Research Institute: Redlands, CA, USA, 2020. [Google Scholar]
- Google Earth Pro 2017. Google Earth Pro Version 7.3. Available online: https://www.google.com/earth/download/gep/agree.html?hl=en-GB (accessed on 1 April 2021).
- Earth Point 2015. Earth point—Tools for Google Earth. Available online: http://www.earthpoint.us/Shapes.aspx (accessed on 10 September 2020).
- Hijmans, R.J.; Cameron, S.E.; Parra, J.L.; Jones, P.G.; Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Climatol. A J. R. Meteorol. Soc. 2005, 25, 1965–1978. [Google Scholar] [CrossRef]
- NOAA National Centers for Environmental Information, U.S. Available online: https://www.ngdc.noaa.gov/mgg/topo/gltiles.html (accessed on 10 September 2020).
- Duque-Lazo, J.; Van Gils, H.; Groen, T.A.; Navarro-Cerrillo, R.M. Transferability of species distribution models: The case of Phytophthora cinnamomi in Southwest Spain and Southwest Australia. Ecol. Model. 2016, 320, 62–70. [Google Scholar] [CrossRef]
- Elith, J.; Graham, H.C.; Anderson, R.P.; Dudík, M.; Ferrier, S.; Guisan, A.; Hijmans, R.J.; Huettmann, F.; Leathwick, J.R.; Lehmann, A.; et al. Novel methods improve prediction of species’ distributions from occurrence data. Ecography 2006, 29, 129–151. [Google Scholar] [CrossRef] [Green Version]
- Guralnick, R.; Hill, A. Biodiversity informatics: Automated approaches for documenting global biodiversity patterns and processes. Bioinformatics 2009, 25, 421–428. [Google Scholar] [CrossRef] [PubMed]
- Robertson, M.P.; Cumming, G.S.; Erasmus, B.F.N. Getting the most out of atlas data. Divers. Distrib. 2010, 16, 363–375. [Google Scholar] [CrossRef]
- Hernandez, P.A.; Graham, C.H.; Master, L.L.; Albert, D.L. The effect of sample size and species characteristics on performance of different species distribution modeling methods. Ecography 2006, 29, 773–785. [Google Scholar] [CrossRef]
- Wisz, M.S.; Hijmans, R.J.; Elith, J.; Peterson, A.T.; Graham, C.H.; Guisan, A.; NCEAS Predicting Species Distributions Working Group. Effects of sample size on the performance of species distribution models. Divers. Distrib. 2008, 14, 763–773. [Google Scholar] [CrossRef]
- Raes, N.; terSteege, H. A null-model for significance testing of presence-only species distribution models. Ecography 2007, 30, 727–736. [Google Scholar] [CrossRef]
- Merckx, B.; Steyaert, M.; Vanreusel, A.; Vincx, M.; Vanaverbeke, J. Null models reveal preferential sampling, spatial autocorrelation and overfitting in habitat suitability modelling. Ecol. Model. 2011, 222, 588–597. [Google Scholar] [CrossRef] [Green Version]
- Fielding, A.H.; Bell, J.F. A review of methods for the assessment of prediction errors in conservation presence/absence models. Environ. Conserv. 1997, 24, 38–49. [Google Scholar] [CrossRef]
- Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- Landis, J.R.; Koch, G.G. An application of hierarchical kappa-type statistics in the assessment of majority agreement among multiple observers. Biometrics 1977, 33, 363–374. [Google Scholar] [CrossRef] [PubMed]
- Allouche, O.; Tsoar, A.; Kadmon, R. Assessing the accuracy of species distribution models: Prevalence, kappa and the true skill statistic (TSS). J. Appl. Ecol. 2006, 43, 1223–1232. [Google Scholar] [CrossRef]
- Lantz, C.A.; Nebenzahl, E. Behavior and interpretation of the κ statistic: Resolution of the two paradoxes. J. Clin. Epidemiol. 1996, 49, 431–434. [Google Scholar] [CrossRef]
- West, A.M.; Kumar, S.; Brown, C.S.; Stohlgren, T.J.; Bromberg, J. Field validation of an invasive species Maxent model. Ecol. Inf. 2016, 36, 126–134. [Google Scholar] [CrossRef] [Green Version]
- Chen, W.; Pourghasemi, H.R.; Kornejady, A.; Zhang, N. Landslide spatial modeling: Introducing new ensembles of ANN, MaxEnt, and SVM machine learning techniques. Geoderma 2017, 305, 314–327. [Google Scholar] [CrossRef]
- Jenks, G.F. The data model concept in statistical mapping. Int. Yearb. Cartogr. 1967, 7, 186–190. [Google Scholar]
- Gavazzi, G.M.; Madricardo, F.; Janowski, L.; Kruss, A.; Blondel, P.; Sigovini, M.; Foglini, F. Evaluation of seabed mapping methods for fine-scale classification of extremely shallow benthic habitats–application to the Venice Lagoon, Italy. Estuar. Coast. Shelf Sci. 2016, 170, 45–60. [Google Scholar] [CrossRef] [Green Version]
- Khan, F. An initial seed selection algorithm for k-means clustering of georeferenced data to improve replicability of cluster assignments for mapping application. Appl. Soft Comp. 2012, 12, 3698–3700. [Google Scholar] [CrossRef] [Green Version]
- Choudhury, A. The distribution, status and conservation of hoolock gibbon, Hoolock hoolock, in Karbi Anglong district, Assam, Northeast India. Primate Conserv. 2009, 24, 117–126. [Google Scholar] [CrossRef] [Green Version]
- Das, J.; Biswas, J.; Bhattacherjee, P.C.; Mohnot, S.M. The distribution and abundance of hoolock gibbons in India. In The Gibbons, Developments in Primatology: Progress and Prospects; Lappan, S., Whittaker, D.J., Eds.; Springer: New York, NY, USA, 2006. [Google Scholar] [CrossRef]
- Chetry, D.; Medhi, R.; Biswas, J.; Das, D.; Bhattacharjee, P.C. Nonhuman primates in the Namdapha national park, Arunachal Pradesh, India. Int. J. Primatol. 2003, 24, 383–388. [Google Scholar] [CrossRef]
- Ray, P.C.; Kumar, A.; Devi, A.; Krishna, M.C.; Khan, M.L.; Brockelman, W.Y. Habitat characteristics and their effects on the density of groups of western hoolock gibbon (Hoolock hoolock) in Namdapha National Park, Arunachal Pradesh, India. Int. J. Primatol. 2015, 36, 445–459. [Google Scholar] [CrossRef]
- Syxaiyakhamthor, K.; Ngoprasert, D.; Asensio, N.; Savini, T. Identifying priority areas for the conservation of the Critically Endangered northern white-cheeked gibbon Nomascusleucogenys in northern Lao. Oryx 2020, 54, 767–775. [Google Scholar] [CrossRef]
- Brockelman, W.Y.; Srikosamatara, S. Estimation of density of gibbon groups by use of loud songs. Am. J. Primatol. 1993, 29, 93–108. [Google Scholar] [CrossRef]
- De Milde, R.A.J.; Shaheduzzaman, M.; Chowdhury, J.A. The Kassalong and Rankhiang Reserved Forests in the Chittagong Hill Tracts; Bangladesh Forest Department: Dhaka, Bangladesh, 1985. [Google Scholar]
- Kabir, M.T.; Ahsan, M.F.; Cheyne, S.M.; Sah, S.A.M.; Lappan, S.; Bartlett, T.Q.; Ruppert, N. Population assessment of the endangered Western Hoolock Gibbon Hoolock hoolock Harlan, 1834 at Sheikh Jamal Inani National Park, Bangladesh, and conservation significance of this site for threatened wildlife species. J. Threat. Taxa 2021, 13, 18687–18694. [Google Scholar] [CrossRef]
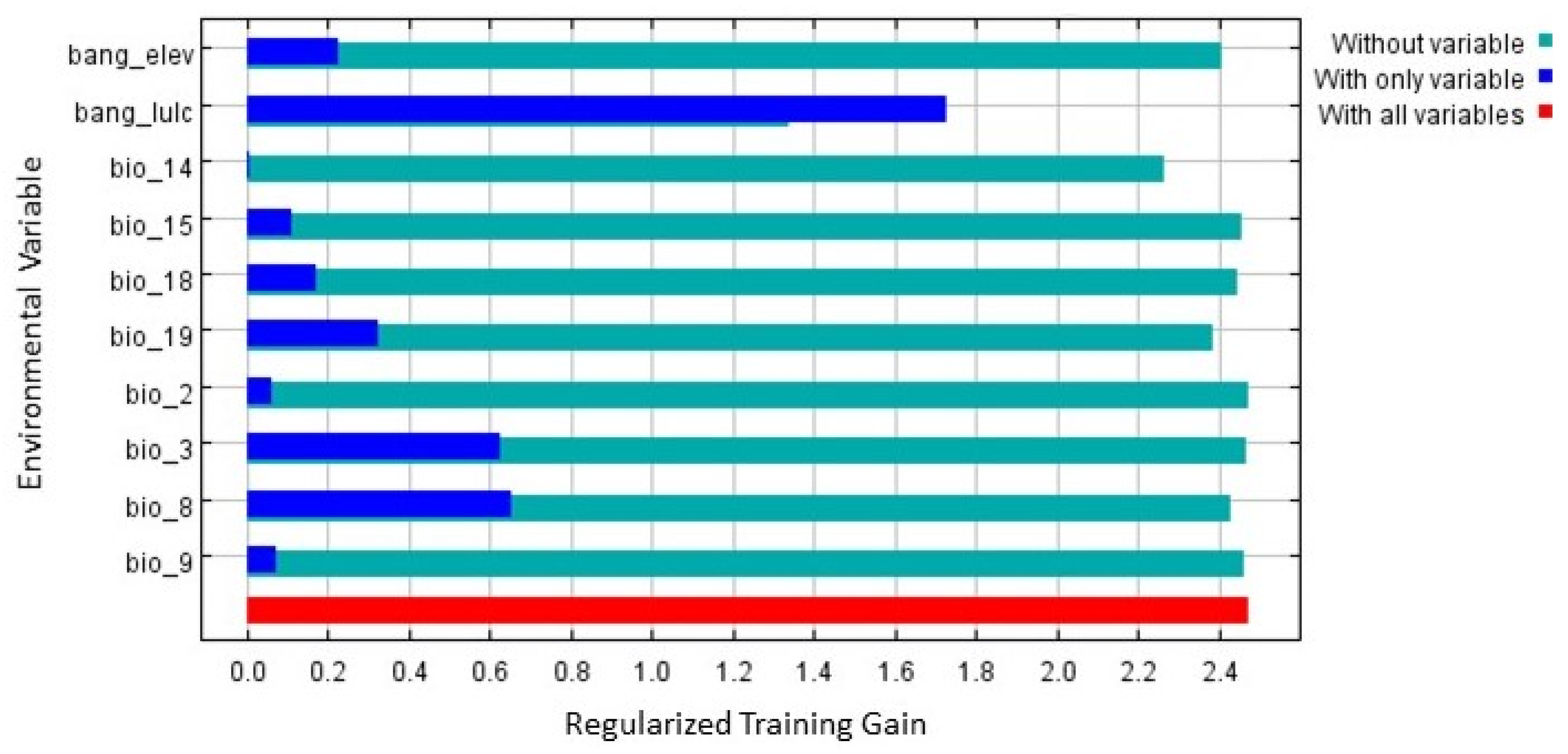
| Symbol | Environmental Variables | Final Model |
|---|---|---|
| Elevation | √ | |
| LULC | Land-use landcover | √ |
| BIO1 | Annual Mean Temperature | |
| BIO2 | Mean Diurnal Range | √ |
| BIO3 | Isothermality | √ |
| BIO4 | Temperature Seasonality | |
| BIO5 | Max Temperature of Warmest Month | |
| BIO6 | Min Temperature of Coldest Month | |
| BIO7 | Temperature Annual Range | |
| BIO8 | Mean Temperature of Wettest Quarter | √ |
| BIO9 | Mean Temperature of Driest Quarter | √ |
| BIO10 | Mean Temperature of Warmest Quarter | |
| BIO11 | Mean Temperature of Coldest Quarter | |
| BIO12 | Annual Precipitation | |
| BIO13 | Precipitation of Wettest Month | |
| BIO14 | Precipitation of Driest Month | √ |
| BIO15 | Precipitation Seasonality | √ |
| BIO16 | Precipitation of Wettest Quarter | |
| BIO17 | Precipitation of Driest Quarter | |
| BIO18 | Precipitation of Warmest Quarter | √ |
| BIO19 | Precipitation of Coldest Quarter | √ |
| Site Name | Total Area of Sites (km2) | Area LULC (Forest Area) | MaxEnt (LULC + Elevation + Bioclim) | Suitable Forest Cover (km2) | Percentage Suitable Area (%) | Number of Transects | Total Transect Length (km) | Number of Observed Groups | Density of Groups (Groups/km2 ± 95% C.I.) | Total Number of Groups | Total Individuals | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| (±SE) d | (±SE) d | |||||||||||
| 1 | Satchari NP a | 2.43 | 2 | 1.61 | 2.35 | 96.71 | 3 | 5.4 | 1 | 1.12 ± 0.98 | 2.62 ± 0.01 | 9.37 ± 1.00 |
| 2 | Lawachara NP | 12.6 | 12 | 11.25 | 10.07 | 79.92 | 7 | 11.73 | 8 | 1.69 ± 0.46 | 13 | 40 |
| 3 | Rema-KalengaWS b | 17.95 | 12 | 14.47 | 16.26 | 90.58 | 5 | 16.79 | 0 | 0.06 | 1 | 5 |
| 4 | RajkandhiRF c | 71.93 | 44 | 49.03 | 27.14 | 37.73 | 7 | 21.35 | 4 | 1.27 ± 0.48 | 34.33 ± 0.19 | 122.57 ± 12.83 |
| 5 | Patharia RF | 36.82 | 23 | 19.29 | 21.66 | 58.83 | 5 | 18.53 | 3 | 1.19 ± 0.55 | 25.73 ± 0.14 | 91.86 ± 9.55 |
| 6 | Atora Hill RF | 86.45 | 46 | 41.8 | 8 | 9.25 | 1 | 6.51 | 0 | 0.13 | 1 | 2 |
| 7 | Hazarikhil WS | 24.62 | 12 | 0 | 18.19 | 73.88 | 3 | 13.22 | 0 | 0 | 0 | 0 |
| 8 | Dighinala | 40.21 | 34 | 0 | 1.44 | 3.58 | 3 | 6.17 | 0 | 0 | 0 | 0 |
| 9 | Sajek Valley | 34.1 | 9 | 0 | 1.48 | 4.34 | 4 | 10.34 | 0 | 0 | 0 | 0 |
| 10 | Pablakhali WS | 420.87 | 188 | 47.43 | 69.03 | 16.40 | 3 | 6.9 | 0 | 0 | 0 | 0 |
| 11 | Kaptai NP | 54.64 | 29 | 1.61 | 21.89 | 40.06 | 3 | 11.54 | 2 | 1.11 ± 0.65 | 24.23 ± 0.13 | 86.51 ± 8.61 |
| 12 | Dhopachori-Dudhpukuria WS | 48.9 | 29 | 0 | 15.59 | 31.88 | 4 | 15.7 | 0 | 0 | 0 | 0 |
| 13 | Chimbuk | 55.53 | 15 | 0.8 | 0 | 0.00 | 4 | 8.8 | 0 | 0 | 0 | 0 |
| 14 | Chunati WS | 77.15 | 6 | 0 | 11.38 | 14.75 | 2 | 8.33 | 0 | 0 | 0 | 0 |
| 15 | Bangdhepa | 4.62 | 3 | 0.8 | 3.77 | 81.53 | 2 | 3.16 | 0 | 0 | 0 | 0 |
| 16 | Bamu | 61.52 | 30 | 0 | 1.94 | 3.15 | 2 | 5 | 0 | 0 | 0 | 0 |
| 17 | Fashiakhali WS | 12.93 | 6 | 0 | 3.31 | 25.59 | 4 | 4.99 | 2 | 1.19 ± 0.74 | 3.94 ± 0.02 | 14.07 ± 1.59 |
| 18 | Innani RF | 74.29 | 44 | 38.58 | 39.7 | 53.44 | 3 | 8.35 | 0 | 0.05 | 7 | 18 |
| 19 | Teknaf WS | 113.75 | 53 | 45.02 | 8.58 | 7.54 | 4 | 7.66 | 0 | 0.12 | 1 | 3 |
| 20 | Rowangchhari | 29.39 | 15 | 0 | 5.52 | 18.78 | 3 | 6.86 | 0 | 0 | 0 | 0 |
| 21 | Sangu-Matamuhuri RF | 803.67 | 385 | 247.58 | 30 | 3.73 | 4 | 7.32 | 2 | 0.72 ± 0.57 | 21.45 ± 0.11 | 76.58 ± 6.72 |
| 22 | Thanchi | 102.71 | 57 | 12.06 | 9.66 | 9.41 | 0 | 0 | 0 | 0 | 0 | 0 |
| TOTAL | 294.54 | 14.95 | 135.31 ± 2.23 | 468.96 ± 45.56 |
| Threshold Independent | Threshold Dependent | ||||
|---|---|---|---|---|---|
| Study Model AUC | Standard Deviation | Null Models AUC | One Sided 95% C.I. | TSS | Cohen’s Kappa (k) Max |
| 0.989 | 0.003 | 0.822 | 0.879 | 0.907 | 0.454 |
| Region | Forest Cover (km2) |
|---|---|
| Northeast | |
| A1 (north of Satchari) | 61.09 |
| A2 (north of Rema) | 49.03 |
| A3 (east of Rema) | 33.76 |
| A4 (south of Lawachara) | 52.25 |
| A5 (north of Patharia) | 52.25 |
| Southeast | |
| A6 (northwest of Pablakhali) | 244.37 |
| Total potential area | 492.76 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Naher, H.; Al-Razi, H.; Ahmed, T.; Hasan, S.; Jaradat, A.; Muzaffar, S.B. Estimated Density, Population Size and Distribution of the Endangered Western Hoolock Gibbon (Hoolock hoolock) in Forest Remnants in Bangladesh. Diversity 2021, 13, 490. https://doi.org/10.3390/d13100490
Naher H, Al-Razi H, Ahmed T, Hasan S, Jaradat A, Muzaffar SB. Estimated Density, Population Size and Distribution of the Endangered Western Hoolock Gibbon (Hoolock hoolock) in Forest Remnants in Bangladesh. Diversity. 2021; 13(10):490. https://doi.org/10.3390/d13100490
Chicago/Turabian StyleNaher, Habibon, Hassan Al-Razi, Tanvir Ahmed, Sabit Hasan, Areej Jaradat, and Sabir Bin Muzaffar. 2021. "Estimated Density, Population Size and Distribution of the Endangered Western Hoolock Gibbon (Hoolock hoolock) in Forest Remnants in Bangladesh" Diversity 13, no. 10: 490. https://doi.org/10.3390/d13100490
APA StyleNaher, H., Al-Razi, H., Ahmed, T., Hasan, S., Jaradat, A., & Muzaffar, S. B. (2021). Estimated Density, Population Size and Distribution of the Endangered Western Hoolock Gibbon (Hoolock hoolock) in Forest Remnants in Bangladesh. Diversity, 13(10), 490. https://doi.org/10.3390/d13100490










