Abstract
Comprehensive biodiversity assessment of moths in Nigeria rely greatly on accurate species identification. While most of the Nigerian moths are identified effortlessly using their morphological traits, some taxa are morphologically indistinguishable, which makes it difficult for taxon diagnosis. We investigated the efficiency of the DNA barcode, a fragment of the mitochondrial Cytochrome C oxidase subunit I, as a tool for the identification of Nigerian moths. We barcoded 152 individuals comprising 18 morphospecies collected from one of the remaining and threatened rainforest blocks of Nigeria – the Cross River National Park. Phenetic neighbor-joining tree and phylogenetic Maximum Likelihood approach were employed for the molecular-based species identification. Results showed that DNA barcodes enabled species-level identification of most of the individuals collected from the Park. Additionally, DNA barcoding unraveled the presence of at least six potential new and yet undescribed species—Amnemopsyche sp., Arctia sp., Deinypena sp., Hodebertia sp., Otroeda sp., and Palpita sp. The phylogenetic Maximum Likelihood using the combined dataset of all the newly assembled sequences from Nigeria showed that all species formed unique clades. The phylogenetic analyses provided evidence of population divergence in Euchromia lethe, Nyctemera leuconoe, and Deinypena lacista. This study thus illustrates the efficacy of DNA barcoding for species identification and discovery of potential new species, which demonstrates its relevance in biodiversity documentation of Nigerian moths. Future work should, therefore, extend to the creation of an exhaustive DNA barcode reference library comprising all species of moths from Nigeria to have a comprehensive insight on the diversity of moths in the country. Finally, we propose integrated taxonomic methods that would combine morphological, ecological, and molecular data in the identification and diversity studies of moths in Nigeria.
1. Introduction
The order Lepidoptera, which includes butterflies and moths, is one of the most diverse and widely recognized insect groups in the world [1]. Today, more than 180000 species belonging to over 15000 genera have been widely reported [2]. Within the order Lepidoptera, moths make up the vast majority with over 160000 species [3]. Moths have great economic significance in our ecosystem. For instance, some moths, e.g., Bombyx mori, Samia cynthia, Antheraea pernyi, Antheraea assamensis etc., are domesticated for their economic value in silk production [4]. In Africa, the caterpillar of many species of moths (e.g., Gonimbrasia belina and Usta terpsichore) are used as an essential source of nutrition for food. Conversely, in some tropical and temperate regions of the world, the larval stage of some moth species constitutes severe agricultural pests to plants. For example, the African sugarcane borer (Eldana saccharina) is a significant pest of sugarcane, maize, and sorghum in Africa [5].
In Africa, moths inhabit diverse terrestrial ecosystems. To date, more than 28000 recognized species of moths have been reported to occur in continental Africa [6]. Among African countries, Nigeria holds an impressive diversity of moths with about 2400 species and likely many more unrecorded species [6]. The southern part of Nigeria, notably the Cross River National Park, has been reported to hold remarkable diversity of the country’s lepidopteran species, and also rank the top in terms of butterfly species richness and diversity in Africa [7]. Although the previous study of Larson [8] has focused on the documentation of the diversity of butterfly species in the park, fewer to no studies exist on the diversity of moths in this protected area. Consisting of one of the remaining rainforests in Africa and a part of the Guineo–Congolian biodiversity hotspot, the forest ecosystem of the Cross River National Park is threatened daily by human activities [9]. The threats undoubtedly would affect the associated flora and fauna, including lepidopteran species, which live within its forest habitat. This, therefore, calls for coordinated strategies to describe the diversity of moths. Documentation of the diversity and distribution of moths would aid in prioritizing conservation actions for Nigerian moths.
Recently, an increase in the effort to document the diversity of moths in Africa has led to a tremendous rate of discovery of new species and first country records annually [10,11,12,13,14,15,16]. This suggests that the diversity of African moths remain little known. Comprehensive documentation of the continental diversity would rely greatly on the use of precise taxonomic tools that would aid in quick and reliable identification of species. However, the use of morphological characters alone presents a challenge to taxon diagnosis, particularly for species with extremely complex morphological traits, hybridization, and mimicry [17]. Molecular-based species identification, a method referred to as DNA barcoding, has emerged as a complementary tool for taxon diagnosis [18]. This involves the use of a short fragment of the mitochondrial cytochrome c oxidase I (COI) to identify specimens to species-level [18]. This approach has become increasingly popular as an efficient tool for studying and categorizing Afrotropical moth species to prioritize conservation efforts [19,20]. In lepidopteran studies, DNA barcoding has proven effective in investigating biodiversity [21,22,23,24,25,26,27,28], resolving taxonomic problems [29,30], and unraveling cryptic diversities [27,30,31,32,33,34]. Apart from this, the large-scale availability of reliable COI sequence data in the global databases has proven useful for quick and accurate identification of lepidopterans [1,24,25,29].
Although several first records and new species of moths are reported annually from different countries in Africa, a comprehensive DNA barcode reference library is still lacking for correct species identification and addressing taxonomic issues. This is particularly true for Nigeria. In this article, we collected 152 specimens comprising 18 morphospecies of moths from Cross River National Park to investigate the efficiency of the DNA barcoding (COI) for the species-level identification that would enable rapid biodiversity assessments of moth species from Nigeria. We examined the DNA barcodes of Nigerian moths alongside other closely related individuals in the global databases to assess its phylogenetic statuses and confirm its identities. Besides, we make recommendations on the potential use of DNA barcodes to enhance the identification of moth species in Nigeria. Finally, the DNA barcode records generated through this study will be available to researchers and biodiversity managers for accurate taxon diagnosis and development of appropriate conservation measures for Nigerian moths.
2. Materials and Methods
2.1. Field Collection and Preliminary Identification of Specimens
Field collections of moths from Cross River National Park in Southern Nigeria (Figure 1) followed ethical approval from the Nigerian National Park Service. The Park was visited on three occasions: the first was from 1–5 August 2017, second from 8–10 February 2018, and third from 10–12 November 2018. Specimens were collected with the help of LMN, ACA, OA and AOA, and 17 park rangers at the Cross River National Park, Nigeria. To ensure rapid specimen collection, we further divided into seven groups comprising three team members each in a group. As our study is a preliminary step towards establishing the effectiveness of DNA barcoding in rapid biodiversity assessments of Nigerian moths, we focused our sampling in pristine humid forests of Erokut Camp and Nsan communities of Cross River National Park. Sampling was conducted in the night using UV light following the methods of Fry and Waring [35] with slight modifications. In sum, one hundred and fifty-two (152) individuals were collected (Supplementary Table S1). The captured specimens were euthanized humanely with ethyl acetate vapor and preserved for reference purposes. Preliminary identification of individuals based on wing and genitalia patterns were carried out with the help of online literature and a trained taxonomist – Dr. O. Anyaele of the Department of Zoology, University of Ibadan (DZ-UI), Ibadan, Nigeria. The preserved voucher specimens were stored in the museum collection at the DZ-UI, Nigeria. Further, we excised the hind legs of each specimen and subsequently preserved them in ethanol at −80 °C for molecular laboratory analyses at the State Key Laboratory of Genetic Resources and Evolution, Kunming Institute of Zoology, Chinese Academy of Sciences, China.
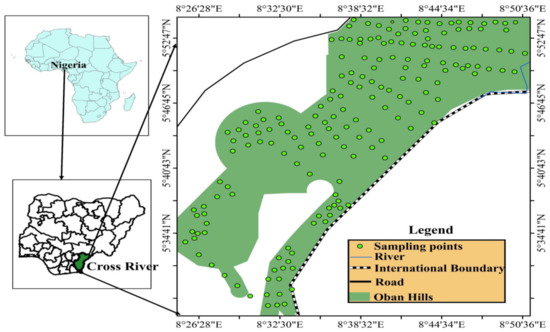
Figure 1.
Map of Nigeria showing our study locality- Cross River National Park, Southern Nigeria.
2.2. Molecular Laboratory Protocols
The total genomic DNA was extracted from the ethanol-preserved hind legs using the Phenol-Chloroform method [36]. The COI gene was amplified using the primer pairs designed by Folmer et al. [37]: (LCO1490: 5′-GGTCAACAAATCATAAAGATATTGG-3′, HCO2198: 5′-TAAACTTCAGGGTGACCAAAAAATCA-3′) and Hebert et al. [38]: (LepF1: 5′-ATTCAACCAATCATAAAGATATTGG-3′, LepR1: 5′-TAAACTTCTGGATGTCCAAAAAATCA-3′). The Polymerase Chain Reaction (PCR) conditions followed standard protocols. The amplified PCR products were checked in 1.2% agarose gel. Using the automated DNA sequencer (ABI 3730), purified PCR products were sequenced directly in both forward and reverse directions. Sequences were aligned in MEGA 7.0 [39] using ClustalW [40] with default parameters. The aligned sequences were translated into amino acids using the invertebrate mitochondrial code, and no premature stop codons observed, indicating that an open reading frame was preserved in the protein-coding locus.
2.3. Molecular-Based Species Identification and Analyses
To test the effectiveness of the DNA barcode library for the identification of Nigerian moths, each of the amplified sequences was regarded as an unknown specimen and submitted to the BLAST searches in the National Center for Biotechnology Information (NCBI) and Barcode of Life Database (BOLD). The neighbor-joining (NJ) analyses of our queried sequences alongside COI sequences of other closely related species were performed using the BOLD platform [41]. The Barcode Index Number, the mean and maximum intra-specific distance, the mean genetic distance to the nearest neighbor, and the nearest neighbor for each species were determined using the BOLD database.
Further, we combined all our newly sequenced datasets to form a combined dataset. We constructed a Maximum Likelihood (ML) phylogeny for the complete data set. The best partition strategy and substitution model were determined using the Akaike information criterion as set out in PartitionFinder 1.0.1 [42]. The ML was performed under model GTR + G as evaluated in PartitionFinder with 100 random addition replicates and per partition branch lengths [43] as implemented in RAXML v. 7.0.3 [44]. Bootstrap analysis, including 1000 replications, assessed how much of the data support the topology of the optimal likelihood tree. The phylogeny was viewed using FigTree v1.4.2 [45]. We further estimated the inter- and intraspecific sequence divergences of between and within all the newly assembled datasets from Nigeria using the K2P distance.
3. Results and Discussion
To the best of our knowledge, this study is the first to examine the effectiveness of DNA barcoding in biodiversity assessment and species-level identification of moths from the Cross River National Park. Identical to Onah [46], our study illustrates the utility of DNA barcodes for the identification of moths from Nigeria. However, our study benefits from having wide taxon sampling compared to the previous study [46], thus, represents a step towards establishing an exhaustive DNA barcode reference library for Nigerian moths. In this current study, one hundred and fifty-two (152) COI sequences were obtained for 18 morphospecies representing the families Brahmaeidae, Crambidae, Erebidae, Geometridae, and Uraniidae (Supplementary Table S1). Our specimen collection also included two species of moths whose caterpillars constitute agricultural pests: Euchromia lethe that feeds on sweet potatoes (Ipomoea batatas) and Polydesma umbricola known to feed on plants like Albizia lebbeck, Albizia saman, and Pithecollobium dulce, and some plants within the genera Acacia, Rosa, and Salix. After trimming the ambiguous bases, a total of 625 base pairs of COI sequence data were aligned. The sequences contained 398 conserved sites and 227 parsimony informative sites. The total nucleotide base content was as follows: T = 40.50%, A = 29.30%, C = 15.30%, and G = 14.90%. The result consequently indicated that the A + T content was 69.80%, evidently higher than the G + C content, which corresponds to the nucleotide composition in the mitochondrial DNA previously reported for insects [47]. All the newly amplified sequences were deposited in the GenBank (MN883673 - MN883824) and also made publicly accessible via the BOLD system website within ‘Identifying species of Moths from Nigeria using DNA barcodes’ project. As a consequence, the production of DNA barcodes through our current study is an important contribution to the rapid identification of Nigerian moths.
As a way of determining the use of DNA barcodes to identify moth species in Nigeria rapidly, the newly amplified sequences of the 18 morphospecies were queried against other COI sequences in the BOLD and NCBI databases. The query search resulted to the species-level placement of 12 out of the 18 morphospecies (Acropteris costinigrata, Archichlora viridimacula, Cadarena sinuata, Dactyloceras lucina, Deinypena lacista, Dissoprumna erycinaria, Erastria albosignata, Euchromia lethe, Mocis mayeri, Nyctemera leuconoe, Polydesma umbricola, and Scopula transsecta), with sequence percentage similarity of ≥97%. The results of the NJ tree analyses using the BOLD v4 platform also showed these species formed DNA barcode (COI) clusters with individuals of the same species, which allowed for their unambiguous species-level identification (Supplementary Figure S1). Here we describe a “diagnostic” barcode as one that identifies a single species to a barcode cluster comprising predominantly of individuals of the same species. In sum, our tree-based analyses showed that the DNA barcode was diagnostic for most of the species of moths collected from the Cross River National Park, Nigeria.
Further, our study illustrates the usefulness of DNA barcoding in unveiling possible new species, which may not be identified using morphology alone. The molecular-based identification recovered unique COI clusters of Amnemopsyche sp., Arctia sp., Deinypena sp., Hodebertia sp., Otroeda sp., and Palpita sp. Besides, these species could not also be classified into species using their morphological traits. From our preliminary results, these six morphospecies (Amnemopsyche sp., Arctia sp., Deinypena sp., Hodebertia sp., Otroeda sp., and Palpita sp.) are regarded as potentially new species pending further taxonomic identification. Integrating morphological, ecological, and genetic data could aid in describing new species and also in the assessment of its conservation status. Studies are currently underway to describe these possible new species using molecular (mitochondrial and nuclear), ecological and morphological data.
Following the species-level classification of sampled individuals, we built an ML tree consisting of the combined dataset (Figure 2). All the 18 species of moths barcoded in our current study were resolved as monophyletic taxa (Figure 2). Sequences from the same species clustered together with high bootstrap supports. Closely related species from the same family were segregated in the analysis. This result thus supports the use of DNA barcodes in the biodiversity assessment and documentation of moths in Nigeria. Hence, DNA barcoding could be used either as a complement to morphology-based identification and/or as the preferred diagnostic tool in situations where the necessary morphological keys are inaccessible or damaged.
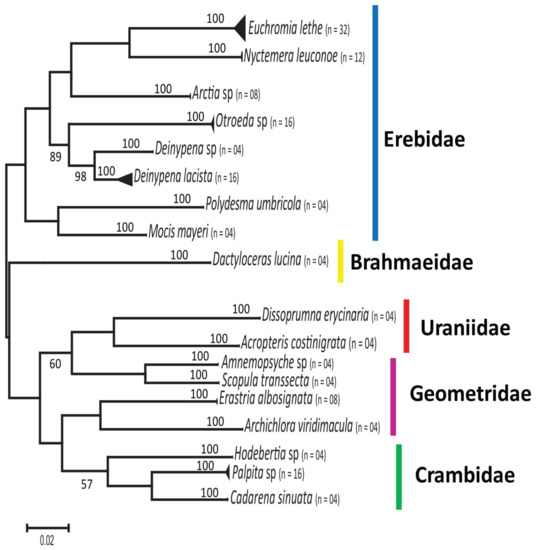
Figure 2.
The unrooted Maximum Likelihood tree based on 152 mitochondrial Cytochrome C oxidase I (COI) sequences of moths from Nigeria.
The mean intraspecific genetic distances between the newly barcoded moth species from Nigeria ranged from 0.00 to 0.0903 (Table 1), with an average of 0.00674. Highest intraspecific difference was seen in Euchromia lethe (Mean = 0.0903), Otroeda sp (Mean = 0.0144), Nyctemera leuconoe (mean = 0.00985) and Deinypena lacista (Mean = 0.00503) (Table 1). Such high levels are likely to be due to population divergence arising from cryptic speciation. In such a case of cryptic speciation, previous studies [48,49,50] have shown the existence of mtDNA clusters (putative cryptic species) separated by geography in case of allopatric speciation or niche partitioning occurring for sympatric speciation. Our NJ tree-based analyses of newly sampled sequences and others in BOLD revealed the presence of at least two distinct clusters in E. lethe, N. leuconoe, and D. lacista corresponding to a geographic pattern of separation (Appendix). These discrete clusters and the existence of high levels of intraspecific difference may indicate the existence of cryptic diversification. Hence, further studies involving detailed field survey and inclusion of more mitochondrial and nuclear genetic markers are required to unravel the processes governing population divergence within this species. The mean evolutionary COI sequence divergence between species of the newly barcoded Nigerian moths ranged from 0.875 to 1.396 (Table 1), while the individual divergence between species ranged from 4.80% to 15.20% (Table 2). The low level of interspecific distance was detected between D. lacista and Deinypena sp (4.80%), while the highest pairwise distance occurred between Dissoprumna erycinaria and Amnemopsyche sp (15.20%).

Table 1.
Mean Intra and interspecific genetic distances and Barcode Index Number details for species of moths from Nigeria.

Table 2.
Estimates of Evolutionary COI Divergence over Sequence Pairs (%) between species of moths from Nigeria using Kimura-2 parameter.
Although our study gives reliance on the efficacy of molecular data in the taxon diagnosis of moth from Nigeria, this would depend on high levels of data integrity in the global databases. Although the vast accumulation of DNA barcode data asserts that BOLD and GenBank could be a useful tool for the identification of specimens, it is not without errors. This often arises either from incorrectly classified specimens and/or cross-contamination of samples. Previous studies, for example, Mitchell and Gopurenko [51], have encountered publically released sequence data that has been incorrectly assigned to species. This ultimately undermines the usefulness of DNA barcodes for taxon diagnosis. Although global databases, particularly BOLD, has a facility for the detection of misidentified specimens, however, these mechanisms are not sufficient enough to ensure a high standard of species identification. Instead, we suggest a multifaceted approach in the diversity studies of moths in Nigeria. The first step could involve rigorous morphological identification of species through the examination of the genitalia and other necessary diagnostic characters and doubly cross-checked by having identification vetted by independent taxonomic experts. Establishing a comprehensive museum collection comprising all the moth species in Nigeria would also go a long way in species identification as well as the comparison of voucher specimens. The second step could involve molecular-based identification using the standard COI barcode.
In some cases, due to the issues arising from the use of a single locus for species identification, additional molecular markers, including nuclear gene markers, could be used to provide stronger evidence for the identification of morphologically similar species. The final step could include a re-examination of specimens that could not be identified using a single gene locus by integrating multiple gene loci and also conducting further genetic analyses. Thus, as previously noted by other researchers, we note that it is highly essential to combine all genetic, morphological, ecological, and behavioral data to conclude the taxon diagnosis [52,53,54].
4. Conclusions
Our study shows the efficacy of DNA barcodes in biodiversity assessments of moth species from Nigeria. This study adds DNA barcodes for quick identification of specimens, which, in turn, will help to identify and document the diversity of moths in Nigeria accurately. Additionally, this study emphasizes the importance of integrating morphological, ecological, behavioral, and molecular (DNA barcoding) data for both the identification and diversity studies of Nigerian moths. We also recommend more exhaustive sampling from broad geographical areas to get a clear insight into the diversity of moths in Nigeria. Finally, we hope this current study will stimulate further synergistic projects to characterize the biodiversity of moths in Nigeria and encourage experts to complement our recommended approach in documenting the diversity of moths in Nigeria.
Supplementary Materials
The following are available online at https://www.mdpi.com/1424-2818/12/2/85/s1, Table S1: List of species including voucher’s specimen number, species name, locality information, and GenBank accession number; Figure S1: Parts of the Neighbor-Joining tree used in the identification of the 18 species of moths collected from Cross River National Park, Nigeria, Note = queried sequences are highlighted in red.
Author Contributions
Conceptualization, L.M.N., and A.C.A.; methodology and sampling, L.M.N., A.C.A., O.A., Y.M., O.O., I.C.N., and A.O.A.; project administration, A.M.A., A.V.A., A.O., and C.S.O.; data curation; L.M.N., A.C.A., and Y.-Y.W.; formal analysis, L.M.N., A.C.A., and M.M.R.; writing—initial draft preparation; L.M.N.; writing—review and editing, A.C.A., I.C.N. and A.O.A. All authors read and approved the final manuscript.
Funding
This work was supported by the Whitley Wildlife Conservation Trust, Sino-Africa Joint Research Centre, Chinese Academy of Sciences (SAJC201611), National Natural Science Foundation of China (31750110480) and the Animal Branch of the Germplasm Bank of Wild Species, Chinese Academy of Sciences (the Large Research Infrastructure Funding).
Acknowledgments
We are grateful to field assistants at the Cross River National Park Nigeria, for their help during sampling.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Kumar, V.; Kundu, S.; Chakraborty, R.; Sanyal, A.; Raha, A.; Sanyal, O.; Ranjan, R.; Pakrashi, A.; Tyagi, K.; Chandra, K. DNA barcoding of Geometridae moths (Insecta: Lepidoptera): A preliminary effort from Namdapha National Park, Eastern Himalaya. Mitochondrial DNA Part B 2018, 4, 309–315. [Google Scholar] [CrossRef]
- Mallet, J. Taxonomy of Lepidoptera: The Scale of the Problem; The Lepidoptera Taxome Project: London, UK, 2007. [Google Scholar]
- Kristensen, N.P.; Scoble, M.J.; Karsholt, O.L.E. Lepidoptera phylogeny and systematics: The state of inventorying moth and butterfly diversity. Zootaxa 2007, 1668, 699–747. [Google Scholar] [CrossRef]
- Zethner, O. South Asian Ways of Silk—A Patchwork of Biology, Manufacture, Culture and History. Entomol. Ornithol. Herpetol. Curr. Res. 2016, 5. [Google Scholar] [CrossRef]
- Conlong, D.E. A review and perspectives for the biological control of the African sugarcane stalkborer Eldana saccharina Walker (Lepidoptera: Pyralidae). Agric. Ecosyst. Environ. 1994, 48, 9–17. [Google Scholar] [CrossRef]
- De Prins, J.; De Prins, W. Afromoths, Online Database of Afrotropical Moth Species (Lepidoptera). World Wide Web Electronic Publication. Available online: www.afromoths.net (accessed on 26 December 2019).
- Obot, E.A.; Ogar, G.; Edet, C.A.; Olory, C.S.; Ayuk, J.; Akongke, C. Biological Inventory in the Okwangwo Division; Progress Report; CRNP: Springboro, OH, USA, 1996; pp. 1994–1996. [Google Scholar]
- Larson, T. Butterflies of the Cross River National Park – diversity writ large. In Proceedings of the workshop: Essential Partnership—the Forest and the People, Cross River National Park, Calabar, Nigeria, 23–28 October 1997; pp. 229–235. [Google Scholar]
- Olory, C.S. Contributions of Cross River National Park to National Development: Prospects and Challenges. In Proceedings of the 6th NSCB Biodiversity Conference, University of Uyo, Uyo, Nigerya, 6–12 May 2018; pp. 309–315. [Google Scholar]
- Li, H.; Zhen, H.; Mey, W. Notes on Dichomeris Hubner, 1818 from Southern Africa and Kenya, with description of seven new species (Lepidoptera, Gelechiidae, Dichomeridinae). Zootaxa 2013, 3608, 561–574. [Google Scholar] [CrossRef]
- Dubatolov, V.V. A new genus of African tiger moths, with a review of the Amsacta melanogastra Holland species group (Lepidoptera, Erebidae). Zootaxa 2013, 3682, 579–583. [Google Scholar] [CrossRef]
- Kovtunovich, V.N.; Ustjuzhanin, P.Y. New Species of Plume Moths of the GenusAgdistisHübner, 1825 (Lepidoptera: Pterophoridae: Agdistinae) from southern Africa. Afr. Invertebr. 2015, 56, 137–145. [Google Scholar] [CrossRef]
- Agassiz, D.J.; Aarvik, L. New Tortricidae (Lepidoptera) from East Africa with an account of the tortricid fauna of acacia in the Kenyan Rift Valley. Zootaxa 2014, 3861, 369–397. [Google Scholar] [CrossRef]
- Maicher, V.; Sáfián, S.; Ishmeal, K.N.; Murkwe, M.; Kimbeng, T.J.; Janeček, Š.; Tropek, R. Two Genera and Nineteen Species of Fruit-Feeding Erebid Moths (Lepidoptera: Erebidae) Recorded in Cameroon for the First Time. Entomol. News. 2016, 126, 64–70. [Google Scholar] [CrossRef]
- Visser, D.; Uys, V.; Nieuwenhuis, R.; Pieterse, W. First records of the tomato leaf miner Tuta absoluta (Meyrick, 1917) (Lepidoptera: Gelechiidae) in South Africa. BioInvasions Rec. 2017, 6, 301–305. [Google Scholar] [CrossRef]
- Ustjuzhanin, P.; Kovtunovich, V.; Safian, S.; Maicher, V.; Tropek, R. A newly discovered biodiversity hotspot of many-plumed moths in the Mount Cameroon area: First report on species diversity, with description of nine new species (Lepidoptera, Alucitidae). Zookeys 2018, 119–139. [Google Scholar] [CrossRef] [PubMed]
- Janzen, D.H.; Hajibabaei, M.; Burns, J.M.; Hallwachs, W.; Remigio, E.; Hebert, P.D. Wedding biodiversity inventory of a large and complex Lepidoptera fauna with DNA barcoding. Philos. Trans. R. Soc. B Biol. Sci. 2005, 360, 1835–1845. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; DeWaard, J.R. Biological identifications through DNA barcodes. Proc. R. Soc. Lond. Ser. B Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Delabye, S.; Rougerie, R.; Bayendi, S.; Andeime-Eyene, M.; Ayala, D.; deWaard, J.R.; Hebert, P.D.N.; Kamgang, R.; Le Gall, P.; Lopez-Vaamonde, C.; et al. Characterization and comparison of poorly known moth communities through DNA barcoding in two Afrotropical environments. Genome 2019, 62, 96–107. [Google Scholar] [CrossRef] [PubMed]
- Lopez, V.C.; Sire, L.; Rasmussen, B.; Rougerie, R.; Wieser, C.; Ahamadi, A.; Minet, J.; deWaard, J.J.R.; Decaëns, T.; Lees, D. DNA barcodes reveal deeply neglected diversity and numerous invasions of micromoths in Madagascar. Genome 2019, 62, 108–121. [Google Scholar] [CrossRef] [PubMed]
- Lukhtanov, V.A.; Sourakov, A.; Zakharov, E.V.; Hebert, P.D. DNA barcoding Central Asian butterflies: Increasing geographical dimension does not significantly reduce the success of species identification. Mol. Ecol. Resour. 2009, 9, 1302–1310. [Google Scholar] [CrossRef]
- Dinca, V.; Zakharov, E.V.; Hebert, P.D.; Vila, R. Complete DNA barcode reference library for a country’s butterfly fauna reveals high performance for temperate Europe. Proc. R. Soc. B Biol. Sci. 2011, 278, 347–355. [Google Scholar] [CrossRef]
- Dewaard, J.R.; Hebert, P.D.N.; Humble, L.M. A Comprehensive DNA Barcode Library for the Looper Moths (Lepidoptera: Geometridae) of British Columbia, Canada. PLoS ONE 2011, 6, e18290. [Google Scholar] [CrossRef]
- Hausmann, A.; Haszprunar, G.; Hebert, P.D.N. DNA Barcoding the Geometrid Fauna of Bavaria (Lepidoptera): Successes, Surprises, and Questions. PLoS ONE 2011, 6, e17134. [Google Scholar] [CrossRef]
- Hausmann, A.; Godfray, H.C.J.; Huemer, P.; Mutanen, M.; Rougerie, R.; van Nieukerken, E.J.; Ratnasingham, S.; Hebert, P.D.N. Genetic Patterns in European Geometrid Moths Revealed by the Barcode Index Number (BIN) System. PLoS ONE 2013, 8, e84518. [Google Scholar] [CrossRef]
- Wilson, J.J.; Sing, K.W.; Sofian-Azirun, M. Building a DNA barcode reference library for the true butterflies (Lepidoptera) of Peninsula Malaysia: What about the subspecies? PLoS ONE 2013, 8, e79969. [Google Scholar] [CrossRef] [PubMed]
- Huemer, P.; Karsholt, O.; Mutanen, M. DNA barcoding as a screening tool for cryptic diversity: An example from Caryocolum, with description of a new species (Lepidoptera, Gelechiidae). Zookeys 2014, 91–111. [Google Scholar] [CrossRef] [PubMed]
- Huemer, P.; Wieser, C.; Stark, W.; Hebert, P.D.N.; Wiesmair, B. DNA barcode library of megadiverse Austrian Noctuoidea (Lepidoptera)—a nearly perfect match of Linnean taxonomy. Biodivers. Data J. 2019, 7, e37734. [Google Scholar] [CrossRef] [PubMed]
- Hajibabaei, M.; Janzen, D.H.; Burns, J.M.; Hallwachs, W.; Hebert, P.D. DNA barcodes distinguish species of tropical Lepidoptera. Proc. Natl. Acad. Sci. USA 2006, 103, 968–971. [Google Scholar] [CrossRef]
- Burns, J.M.; Janzen, D.H.; Hajibabaei, M.; Hallwachs, W.; Hebert, P.D. DNA barcodes and cryptic species of skipper butterflies in the genus Perichares in Area de Conservacion Guanacaste, Costa Rica. Proc. Natl. Acad. Sci. USA 2008, 105, 6350–6355. [Google Scholar] [CrossRef]
- Lavinia, P.D.; Nunez Bustos, E.O.; Kopuchian, C.; Lijtmaer, D.A.; Garcia, N.C.; Hebert, P.D.N.; Tubaro, P.L. Barcoding the butterflies of southern South America: Species delimitation efficacy, cryptic diversity and geographic patterns of divergence. PLoS ONE 2017, 12, e0186845. [Google Scholar] [CrossRef]
- Huemer, P.; Hebert, P.D.N. Cryptic diversity and phylogeography of high alpine Sattleria—A case study combining DNA barcodes and morphology (Lepidoptera: Gelechiidae). Zootaxa 2011, 2981, 1. [Google Scholar] [CrossRef]
- Dinca, V.; Montagud, S.; Talavera, G.; Hernandez-Roldan, J.; Munguira, M.L.; Garcia-Barros, E.; Hebert, P.D.; Vila, R. DNA barcode reference library for Iberian butterflies enables a continental-scale preview of potential cryptic diversity. Sci. Rep. 2015, 5, 12395. [Google Scholar] [CrossRef]
- Chan-Chable, R.J.; Martinez-Arce, A.; Mis-Avila, P.C.; Ortega-Morales, A.I. DNA barcodes and evidence of cryptic diversity of anthropophagous mosquitoes in Quintana Roo, Mexico. Ecol. Evol. 2019, 9, 4692–4705. [Google Scholar] [CrossRef]
- Fry, R.; Waring, P. A guide to moth traps and their use. Amat. Entomol. 1996, 24, 60. [Google Scholar]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1989. [Google Scholar]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar] [PubMed]
- Hebert, P.D.; Penton, E.H.; Burns, J.M.; Janzen, D.H.; Hallwachs, W. Ten species in one: DNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. Proc. Natl. Acad. Sci. USA 2004, 101, 14812–14817. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D. A DNA-based registry for all animal species: The barcode index number (BIN) system. PLoS ONE 2013, 8, e66213. [Google Scholar] [CrossRef]
- Lanfear, R.; Calcott, B.; Ho, S.Y.; Guindon, S. Partitionfinder: Combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol. 2012, 29, 1695–1701. [Google Scholar] [CrossRef]
- Townsend, T.M.; Mulcahy, D.G.; Noonan, B.P.; Sites, J.W., Jr.; Kuczynski, C.A.; Wiens, J.J.; Reeder, T.W. Phylogeny of iguanian lizards inferred from 29 nuclear loci, and a comparison of concatenated and species-tree approaches for an ancient, rapid radiation. Mol. Phylogenet. Evol. 2011, 61, 363–380. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 2006, 22, 2688–2690. [Google Scholar] [CrossRef]
- Rambaut, A. FigTree v1.4.4. Available online: http://tree.bio.ed.ac.uk/software/figtree (accessed on 20 December 2019).
- Onah, E.I.; Taylor, D.; Eyo, J.E.; Ubachukwu, P.O. Identification of the false codling moth, Thaumatotibia leucotreta (meyrick) (Lepidoptera: Tortricidae), infesting sweet oranges in nigeria, by DNA barcoding. Proc. Entomol. Soc. Wash. 2016, 118, 574–581. [Google Scholar] [CrossRef]
- Simon, C.; Frati, F.; Beckenbach, A.; Crespi, B.; Liu, H.; Flook, P. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 1994, 87, 651–701. [Google Scholar] [CrossRef]
- Friberg, M.; Bergman, M.; Kullberg, J.; Wahlberg, N.; Wiklund, C. Niche separation in space and time between two sympatric sister species—A case of ecological pleiotropy. Evol. Ecol. 2007, 22, 1–18. [Google Scholar] [CrossRef]
- Friberg, M.; Leimar, O.; Wiklund, C. Heterospecific courtship, minority effects and niche separation between cryptic butterfly species. J. Evol. Biol. 2013, 26, 971–979. [Google Scholar] [CrossRef] [PubMed]
- Pazhenkova, E.A.; Zakharov, E.V.; Lukhtanov, V.A. DNA barcoding reveals twelve lineages with properties of phylogenetic and biological species within Melitaea didyma sensu lato (Lepidoptera, Nymphalidae). Zookeys 2015, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, A.; Gopurenko, D. DNA Barcoding the Heliothinae (Lepidoptera: Noctuidae) of Australia and Utility of DNA Barcodes for Pest Identification in Helicoverpa and Relatives. PLoS ONE 2016, 11, e0160895. [Google Scholar] [CrossRef] [PubMed]
- Hebert, P.D.N.; Ratnasingham, S.; Zakharov, E.V.; Telfer, A.C.; Levesque-Beaudin, V.; Milton, M.A.; Pedersen, S.; Jannetta, P.; deWaard, J.R. Counting animal species with DNA barcodes: Canadian insects. Philos. Trans. R. Soc. B 2016, 371, 20150333. [Google Scholar] [CrossRef] [PubMed]
- Janzen, D.H.; Burns, J.M.; Cong, Q.; Hallwachs, W.; Dapkey, T.; Manjunath, R.; Hajibabaei, M.; Hebert, P.D.N.; Grishin, N.V. Nuclear genomes distinguish cryptic species suggested by their DNA barcodes and ecology. Proc. Natl. Acad. Sci. USA 2017, 114, 8313–8318. [Google Scholar] [CrossRef]
- Carstens, B.C.; Pelletier, T.A.; Reid, N.M.; Satler, J.D. How to fail at species delimitation. Mol. Ecol. 2013, 22, 4369–4383. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).