Abstract
Larks constitute an avian family of exceptional cryptic diversity and striking examples of convergent evolution. Therefore, traditional morphology-based taxonomy has recurrently failed to reflect evolutionary relationships. While taxonomy ideally should integrate morphology, vocalizations, behaviour, ecology, and genetics, this can be challenging for groups that span several continents including areas that are difficult to access. Here, we combine morphometrics and mitochondrial DNA to evaluate the taxonomy of Calandrella larks, with particular focus on the African C. cinerea and the Asian C. acutirostris complexes. We describe a new range-restricted West African taxon, Calandrella cinerea rufipecta ssp. nov. (type locality: Jos, Plateau State, Nigeria), with an isolated relic population 3000 km from its closest relative in the Rift Valley. We performed molecular species delimitation, employing coalescence-based multi-rate Poisson Tree Processes (mPTP) on cytochrome b sequences across 52 currently recognized lark species, including multiple taxa currently treated as subspecies. Three species-level splits were inferred within the genus Calandrella and another 13 across other genera, primarily among fragmented sub-Saharan taxa and taxa distributed from Northwest Africa to Arabia or East Africa. Previously unknown divergences date back as far as to the Miocene, indicating the presence of currently unrecognized species. However, we stress that taxonomic decisions should not be based on single datasets, such as mitochondrial DNA, although analyses of mitochondrial DNA can be a good indicator of taxa in need of further integrative taxonomic assessment.
1. Introduction
Despite being a central entity in biology, the definition of a species is under constant debate, e.g., [1,2,3], and the application of different species definitions—with focus varying from, e.g., reproductive isolation, diagnosable characters, or monophyly [4,5]—result in widely different delineations [6]. This is not only of a conceptual interest but plays an important role in conservation efforts [6,7], as their focus are commonly aimed at the species level, while evolutionarily distinct lineages of lower taxonomic rank receive less attention [8,9]. Over time, the number of lineages recognized at the species level has increased [9,10,11]. Isaac, Mallet and Mace [9] argued that this is taxonomic inflation due to a shift from the traditional Biological Species Concept towards versions of the Phylogenetic Species Concept, and further claimed that this may be counterproductive to conservation efforts in the long run. However, Sangster [11] demonstrated that the increased number of recognized bird species follows taxonomic progress due to new discoveries and knowledge, rather than splits of populations and subspecies solely based on phylogenetic information, in which case the global number of species has been predicted to double [8]. At the core of the issue lies detectability and diagnosability of species. Whereas gene flow could theoretically homogenize the full genomes between interbreeding species, save a few specific genes that maintain striking difference in, e.g., coloration and perception [12,13], other lineages may have been isolated so long that they are—or would be—reproductively isolated upon secondary contact, but differ ever so slightly in morphology or/and behaviour, e.g., [14,15]. Such cryptic taxa are often discovered through DNA sequencing and open a new window to diversity and conservation [16]. In birds, taxonomy has traditionally been based on morphology, including plumage patterns, and convergent as well as divergent evolution has often led to misclassifications at every taxonomic level, e.g., [17,18,19,20,21,22].
Larks (Aves: Alaudidae) are a widespread and large avian family, comprising over 100 species in 21 genera [23] across Africa and Eurasia, with single representatives in Oceania and the Americas [24]. Using multi-locus DNA sequencing, Alström, et al. [25] demonstrated several cases of spectacular morphological convergences and divergences among taxa of different genera, and proposed a revised generic taxonomy of the family. They also revealed multiple cases of deep divergences among taxa considered to be conspecific, as well as some shallow splits between currently recognized species. As an example, the genus Calandrella was found to be paraphyletic, leading to the resurrection of the genus Alaudala for one clade of the traditional genus Calandrella [25]. Moreover, unexpected relationships were found within Alaudala, which were later corroborated by more comprehensive studies [26]. Within Calandrella sensu stricto (s.s.), the Mongolian Short-toed Lark C. dukhunensis was considered to be a subspecies of Greater Short-toed Lark C. brachydactyla, but was suggested to be more closely related to Hume’s Short-toed Lark C. acutirostris than to C. brachydactyla [25], which was later supported by genomic [27] and non-molecular data [28], strongly supporting species status of C. dukhunensis. Stervander, et al. [27] demonstrated multiple cases of paraphyly within the genus Calandrella, and on the one hand showed that many subspecies were parts of large panmictic populations, while on the other hand, identified substantial genetic divergences, including a highly localized taxon on the Jos Plateau in Nigeria.
As an additional challenge to taxonomic classification, larks display a correlation between substrate colour and the plumage coloration of their upperparts [25,29], as well as between bill morphology and habitat [25]. These examples of local adaptation have created morphological variation that has provided ground for the description of a multitude of subspecies [23].
Africa is home to 80% of the lark species, of which 62% are endemic to Africa south of the Sahara. Concordant with Africa receiving relatively little attention to avian taxonomy and biogeography despite housing a substantial part of the global diversity [30], several species and numerous subspecies and discrete populations remain uncharacterized genetically and to some degree morphologically. For the few genera or species that are exceptions, thorough evaluations have often resulted in the elevation of subspecies to species rank [30,31,32,33,34].
In the present study, we aim to (1) evaluate morphological and genetic differentiation within Calandrella s.s.; (2) formally describe a new taxon restricted to the Jos Plateau in Nigeria, (3) perform genetically-based species delimitation analyses with particular regard to (a) Calandrella and (b) other larks distributed across northern Africa, and (4) discuss potential taxonomic implications.
2. Materials and Methods
2.1. Morphology
We recorded morphometrics from 403 Calandrella specimens in five museum collections: Natural History Museum (NHMUK), Tring, UK; Musée Royal de l’Afrique Centrale (MRAC), Tervuren, Belgium; Zoological Museum, University of Copenhagen (ZMUC), Copenhagen, Denmark; University of Michigan Museum of Zoology (UMMZ), Ann Arbor, MI, USA; and American Museum of Natural History (AMNH), New York, NY, USA. Three standard measurements were recorded: wing length (‘maximum chord’, primaries straightened and flattened; Svensson [35]), tail length (measured with a ruler under the undertail-coverts to the nearest 0.5 mm), and bill length (from bill tip to the base of the skull with calipers to the nearest 0.1 mm). Tail/wing ratio was calculated by dividing tail length with wing length. M.S. measured 252 specimens, P.A. 151 specimens. For African species, the formula of the folded wing was recorded by M.S. with a ruler to the nearest 0.5 mm as the distance from the wing tip to individual primaries 2–7 (P2–P7, numbered from the outermost, minute, primary).
While blind tests of bill length in 10 overlapping specimens of C. acutirostris at NHMUK revealed no systematic differences between the two authors, to unveil any hidden bias, we ran three ANOVAs with each of the main measurements as dependent variable for the central populations of C. acutirostris (around Ladakh, India; the population most extensively measured by both M.S. and P.A.), using sex and measurer as factors. There was a significant effect on tail length, and all tail length measurements by P.A. were therefore adjusted with +2.53 mm. We tested the differences between populations by running ANOVAs for each of the main measurements and the individual wing formula measurements, using sex and population as factors (Table 1). Populations were defined as clades according to Stervander, et al. [27], except for C. acutirostris, where all birds from the central population were grouped, since most were not genotyped and clades B3 and B4 in Stervander, et al. [27] therefore could not be distinguished.

Table 1.
Summaries of variable-specific ANOVAs testing for differences between taxa. Independent variables were taxon (clade) and sex.
We performed Principal Component (PC) Analysis (PCA) using the function prcomp from the stats R package. Omitting any individuals with missing data, we did one PCA on 369 individuals across all taxa, using logarithmized measurements of wing length, bill length, tail length, and tail/wing ratio as variables. We replicated this PCA while only including males, to remove sex-dependent variation, and because our material included a majority of males (229 of the 369 individuals above). We also ran a corresponding PCA separately on only the C. acutirostris complex. Further, for (i) all African taxa and (ii) only the Red-capped Lark C. cinerea complex we performed two different PCAs. First, we used the two wing formula measurements providing the strongest separation between taxa, the distance of P6 and P7 to the wing tip (Table 1) divided by wing length (and then logarithmized), as a measurement of wing roundedness (with relatively longer P6 and P7 signifying rounder wing). Second, we included all wing formula measurements that were differentiated among taxa, i.e., the distance of P4–P7 to the wing tip (Table 1). For some individuals, P4 or P5 constituted the wing tip and thus had a measurement of 0. We therefore added a constant (10 mm) to all distances, and then logarithmized them. Finally, we performed Linear Discrimination Analyses (LDA) on C. acutirostris (western vs. eastern birds; logarithmized bill, wing, and adjusted tail length) and C. cinerea (NE African vs. southern African birds; the same measurements, and additionally logarithmized P6 and P7 measurements) using the R packages MASS and ROCR. Using the lda function, we (1) calculated the posterior probability on all data for classification as one or the other taxon, and (2) used prediction and performance functions to train classification using 40% of the datasets, and then predict the remaining 60% of the datasets. We replicated this 1000 times and drew receiver operating characteristic (ROC) curves, and calculated the areas under the ROC curves (AUC) as a measurement of successful classification (0.5 equaling no predictive power and 1.0 signifying perfect classification), averaging AUC values over the replicates. All statistical analyses were done in R v. 3.6.1 [36].
2.2. Phylogeny and Species Delimitation
The core sequence dataset comprised the cytochrome b data from Alström, et al. [25], Stervander, et al. [27], Ghorbani, et al. [37] and Ghorbani, et al. [26]. We further complemented this with additional sequences (U.O., P.A.; Raso Lark Alauda razae from Elisa G. Dierickx, M. de L. Brooke, Yun Li, and Yang Liu) and sequences from other studies available on GenBank (see Table S1 or Figure S1 for accession numbers and taxon information). For phylogenetic analyses, we added outgroups represented by Bearded Reedling Panurus biarmicus (sister to Alaudidae), Himalayan Prinia Prinia crinigera, Great Tit Parus major, Goldcrest Regulus regulus, Song Thrush Turdus philomelos, European Goldfinch Carduelis carduelis, and White-faced Robin Tregellasia leucops. All sequences were aligned with the MAFFT algorithm [38] implemented in Geneious v. 10.2.6 [39], inspected manually, and one obvious codon-violating misalignment was corrected.
We evaluated 24 substitution models with jModelTest v. 2.1.10 [40] and selected the HKY model [41] with rate variation according to a gamma distribution (Γ), and a proportion of invariant sites (I), based on the Bayesian Information Criterion [42]. Employing this substitution model with four Γ rate categories, we specified a phylogenetic analysis for BEAST v. 2.6.1 [43], with the speciation process following a birth–death model, setting a relaxed molecular clock of 0.0105 substitutions/site/million years (MA) [44] following a lognormal distribution. We enforced monophyly of Alaudidae and Alaudidae+Panuridae, which were highly supported as monophyletic clades in multi-locus nuclear studies [45,46,47], since the restricted number and density of outgroups risk inaccurate rooting. We employed default priors and operators, with the exception of using an exponential prior with mean 1 for BDBirthRate.t, doubling the weight (from 3 to 6) for operators ucldStdev.c, Scale_internal_Tree.t, Scale_root_Tree.t, BDBirthRate.t, and BDDeathRate.t, and increasing from 30 to 40 for Uniform_Tree.t, in order to improve performance. We ran two independent runs, sampling every 1000 generation, for 40 million generations.
The output of the replicate runs was inspected for stationarity and convergence in Tracer v. 1.7.1 [48] and checked for effective sample sizes (ESS) of ~200 or higher. We discarded 5% as burn-in fraction and computed maximum clade credibility trees with median node heights in TreeAnnotator [43]. Detecting no significant differences, we used the tree from the run generating overall highest ESS, and imported the tree into R v. 3.6.1 [36], in which the packages phyloch v. 1.5-5 [49], phytools v. 0.6-99 [50], and ape v. 5.3 [51,52] were used for manipulation and visualization. Since the summarizing of trees into a maximum clade credibility tree can produce negative branch lengths if the variance in divergence time is large [53], which is particularly prone to happen with intraspecific divergence, we changed six such instances of small negative intraspecific branch lengths to zero.
We pruned the tree in R from species that were only represented by a single sequence, since species delimitation methods relying on identifying the shifting point between intraspecific coalescence and speciation cannot handle such data [54]. We then performed species delimitation analyses using multi-rate Poisson Tree Processes (mPTP), which is a maximum likelihood-based method employing PTP on a single locus, allowing for variable rates of intraspecific coalescence, coupled with MCMC-based assessment of accuracy [54]. We first inferred the minimum branch length threshold value, below which sequences are considered identical (but having non-zero branch lengths because of different sequence lengths or missing data) using the --minbr_auto command, and then used this value for --minbr in the species delimitation applying ten million generations of MCMC.
2.3. Description of a New Taxon
On 6 February 2004, Mark F. Hulme and Ross McGregor (A. P. Leventis Ornithological Research Institute and University of St Andrews) observed two larks near Gwafan, 7 km east of Jos (9°52′55′′ N, 8°57′3′′ E), Jos Plateau, Nigeria, believed to be Calandrella cinerea saturatior as described by Sharland [55,56], but not observed since 1994 (Mark Hopkins, pers. comm.). However, they noted that the Jos Plateau larks were markedly different in plumage from C. c. saturatior from elsewhere (including D. R. Congo, which is also disjunct from other populations of C. c. saturatior, and geographically closest to the Jos plateau). The first individual ever observed on the Jos Plateau was collected on 8 October 1958 and sent to the Natural History Museum, Tring, UK, where it was classified as C. c. saturatior [55]. Here, we characterized the morphology of the genetically differentiated Nigerian population of C. cinerea [27] based on detailed studies of the above museum specimen, NHMUK 1960.8.4, which is the only one ever collected, and supplemented with observations of birds photographed in the field and three individuals mist-netted for ringing.
3. Results
3.1. Morphology of Calandrella larks
3.1.1. Descriptive Statistics
ANOVAs revealed no measurer bias for wing length (p = 0.65 for effect of measurer; model F2,28 = 12.71, adjusted r2 = 0.44, p < 0.001) or bill length (p = 0.62 for effect of measurer; model F2,28 = 2.27, adjusted R2 = 0.08, p = 0.12); however, tail length differed systematically (effect of measurer 2.53 at t = 2.89, p = 0.007; model F2,28 = 16.62, adjusted R2 = 0.51, p < 0.001). Our preliminary test of differentiation between taxa for the four main variables, as well as wing formula measurements in African taxa, confirmed differences—of varying magnitude—in all variables except distance to wing tip from P2 and P3 (Table 1). The three main measurements are summarized in Table 2.

Table 2.
Summary statistics for three morphometric measurements recorded in 403 museum specimens of Calandrella larks, grouped according to (1) currently recognized species and (2) major clades or groups. S, W, NW, NE, and E represent directions, and C central, referring to the distribution of a taxon.
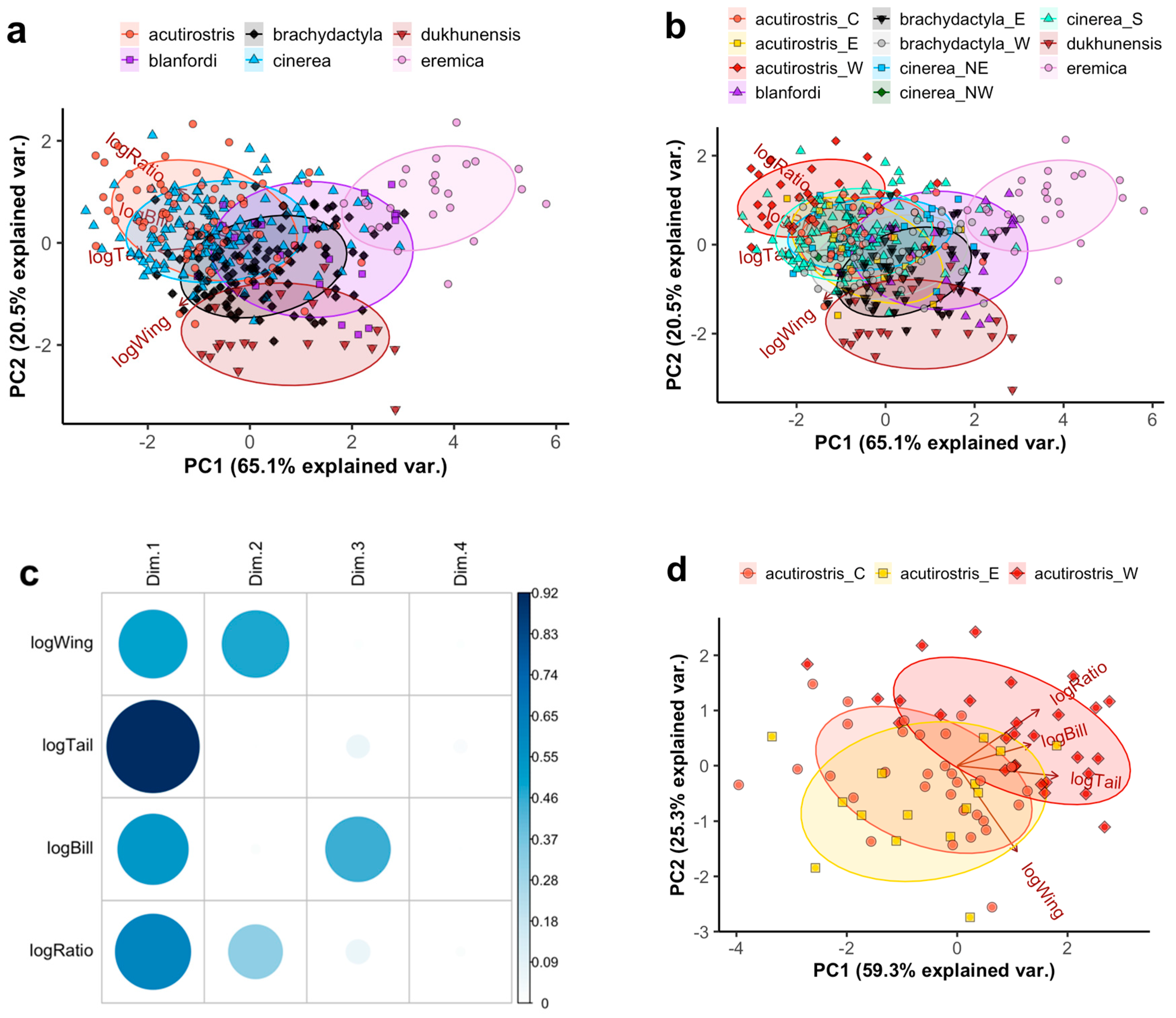
The PCA across 369 individuals of all taxa using the four main variables resulted in relatively little separation (Figure 1a,b), primarily distinguishing C. eremica being short-winged and C. dukhunensis being long-winged with a low tail/wing ratio. The first two principal components account for 84.6% and PC3 for 14.5%. All four variables contributed significantly to PC1, wing length and tail/wing ratio to PC2, and bill length to PC3 (Figure 1c). Restricting the analysis to males only did not result in a significantly different pattern (Figure S2). A PCA on the same variables specifically for the C. acutirostris complex separated western birds (clade B2, all from Afghanistan) from eastern birds (clade B1, all from Tibet, currently defined as C. a. tibetana) to some degree, with the former having longer bills and larger tail/wing ratios (Figure 1d). Birds sampled in the geographically central Ladakh region (“acutirostris_C”), belonging to clades B3 and B4 or of unknown phylogenetic affiliation, placed intermediately in the PCA (Figure 1d).
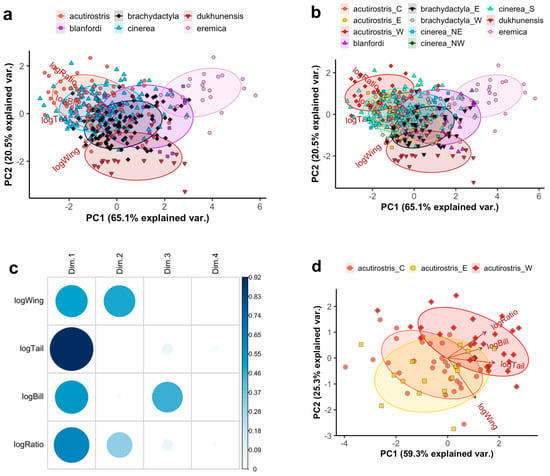
Figure 1.
Principal Component (PC) Analysis (PCA) plots based on logarithmized morphometric measurements of Calandrella larks: wing length (logWing), tail length (logTail), tail/wing length ratio (logRatio), and bill length (logBill). Groups are indicated by colour/shape according to (a) currently recognized species and (b) clades or groups discussed in this study. For the latter, C. acutirostris is divided into a western group (W; corresponding to clade B2 following Stervander, et al. [27]), an eastern group (E; corresponding to clade B1 following Stervander, et al. [27]), and a central group from the Ladakh area (C; corresponding to birds from clades B3 and B4 following Stervander, et al. [27]). Calandrella brachydactyla is divided into a western (W) and eastern (E) clade. Calandrella cinerea is divided into a southern clade (S; all currently recognized subspecies except C. c. williamsi), a northeastern clade (NE; C. c. williamsi) and a northwestern clade (NW; a bird from the Jos Plateau, Nigeria). Ellipses correspond to one standard deviation from the mean (centroid). Note that “cinerea_NW” is only represented by a single point (located approximately at PC1 = −1.5; PC2 = 0) and thus lacks an ellipse. Variables and loadings are shown with brown arrows and labels. This PCA includes males, females, and birds of undetermined sex; for a corresponding PCA restricted to males only, see Figure S2. (c) Quality of representation (cos2) of the variables on different PCs for a–b. (d) A PCA corresponding to panel a–b run exclusively on C. acutirostris. Note that the PC1 axis is mirrored, and that the loading of bill length is highly correlated with that of tail/wing ratio rather than tail length. See Figure S3 for quality of representation of the variables on different PCs.
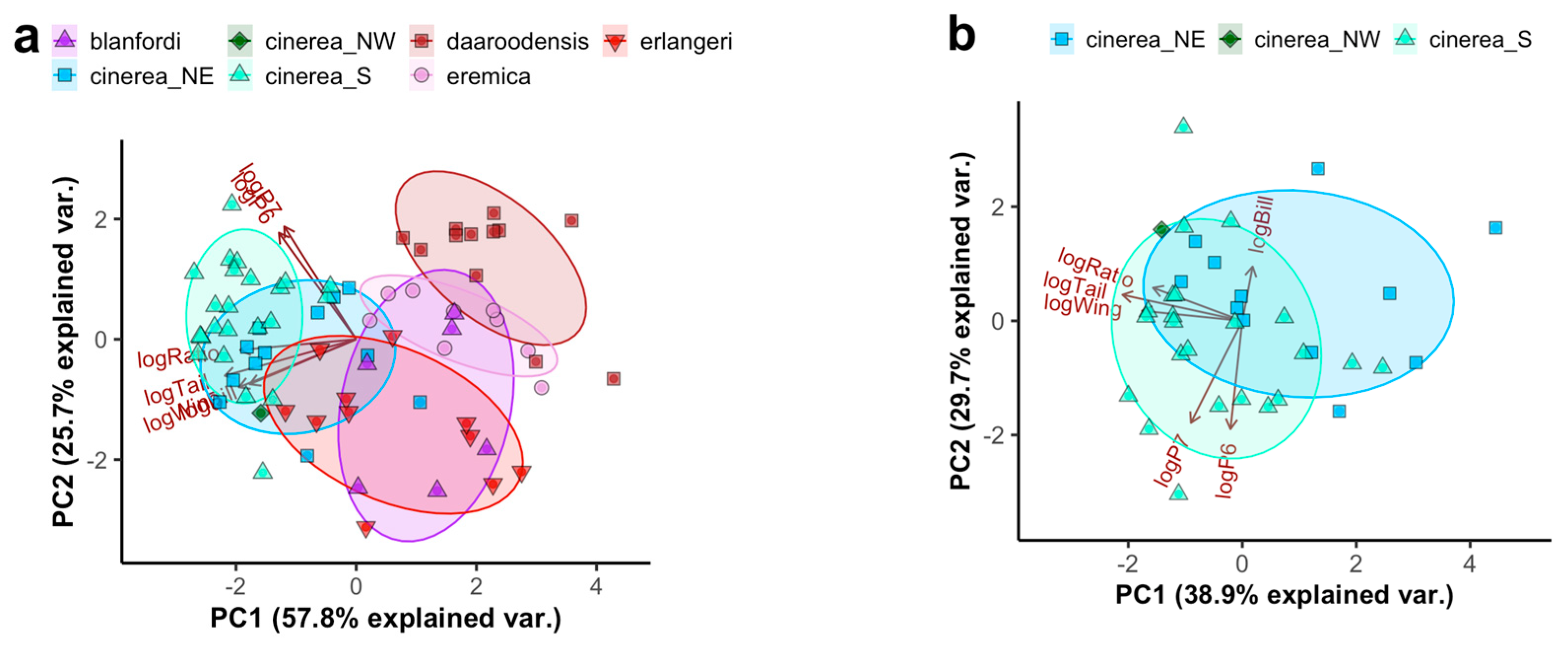
Using the extended morphometrics including wing formula separates the African taxa somewhat further along an axis of more or less rounded wing shape (Figure 2a), with Blanford’s Lark C. blanfordi (C. b. blanfordi and C. b. erlangeri) having the most rounded wing. A separate PCA on the C. cinerea complex shows moderate separation between southern birds (C. cinerea s.s.) and northeastern birds (C. c. williamsi; Figure 2b), the latter being shorter-tailed (estimated average of 56.3 vs. 58.8, t = −3.18, p = 0.003; ANOVA with factors taxon and sex: F2,28 = 17.72, adjusted r2 = 0.59, p < 0.001) and rounder-winged (estimated average of the distance from P7 to wing tip in relation to wing length 18.3% vs. 19.9%, t = −1.42, p = 0.001; ANOVA with factors taxon and sex: F2,28 = 5.78, adjusted r2 = 0.29, p = 0.003).
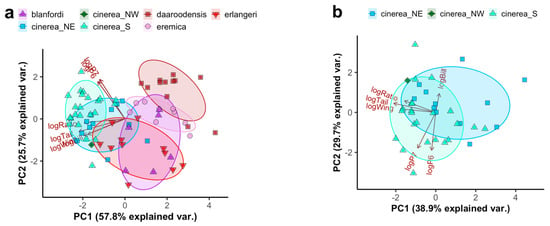
Figure 2.
Principal Component (PC) Analysis (PCA) plot based on logarithmized morphometric measurements of (a) all African Calandrella and (b) the C. cinerea complex: wing length (logWing), tail length (logTail), tail/wing length ratio (logRatio), bill length (logBill), and the wing formula measurements of distance between the wing tip and the tips of primaries 6 and 7 (numbered from the outermost 1st primary) relative to the wing length (logP6 and logP7). Taxa are indicated by colour/shape: C. blanfordi comprising C. b. blanfordi and C. b. erlangeri, C. eremica comprising C. e. eremica and C. e. daaroodensis, C. cinerea comprising a southern clade (S; all currently recognized subspecies except C. c. williamsi), a northeastern clade (NE; C. c. williamsi) and a northwestern clade (NW; a bird from the Jos Plateau, Nigeria). Ellipses correspond to one standard deviation from the mean (centroid) of groups (except “cinerea_NW”). For quality of representation (cos2) plots, see Figure S4. For PCAs corresponding to (a) and (b), using different wing formula measurements, see Figure S5.
3.1.2. Prediction of Taxon Membership
The LDA classifications were largely correct, with average AUC = 0.94 over 1000 replicates of training and prediction for western vs. eastern C. acutirostris (Figure S6a) based on three measurements. Classifications of southern vs. northeastern C. cinerea based on five measurements were less specific, with average AUC = 0.83 (Figure S6b).
3.2. Description of a New Taxon Endemic to the Jos Plateau in Nigeria

Figure 3.
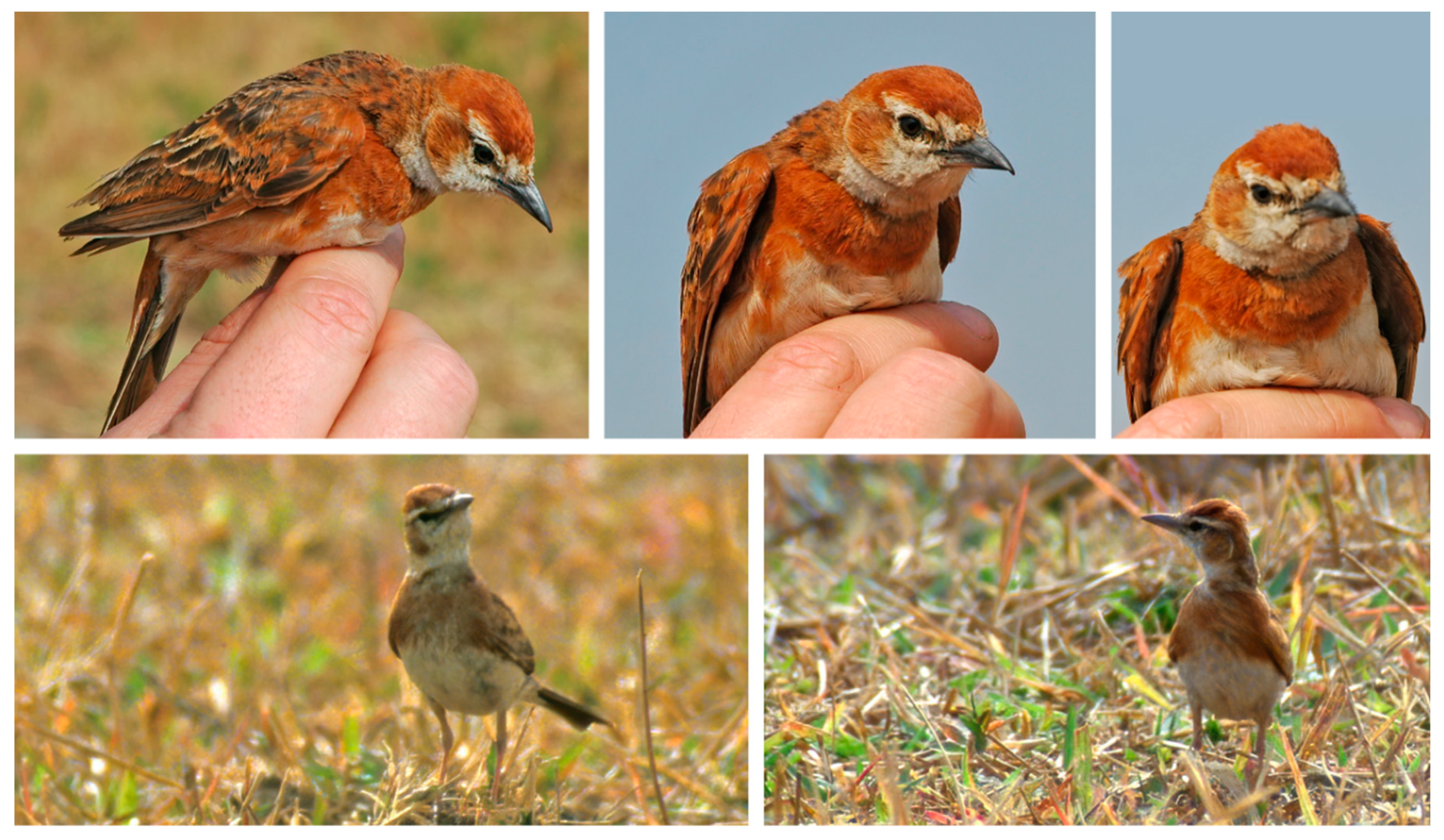
Photos of live Calandrella cinerea rufipecta ssp. nov. from the Jos Plateau, Nigeria. The upper panel is individual FA67266 trapped for ringing at Gwafan, 31 January 2006 (sample cinerea_saturatior_27_NGA of Stervander, et al. [27]; cytochrome b GenBank accession number KX379982; photos © Mark Hulme). The lower panel includes two photos of a bird observed in the field near Gwafan in February 2004 (photos © Ross McGregor, with permission). Note the deep russet crown and unbroken breast band, the latter diagnostic of the taxon.
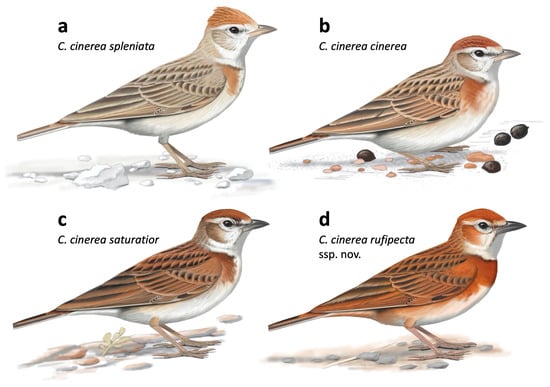
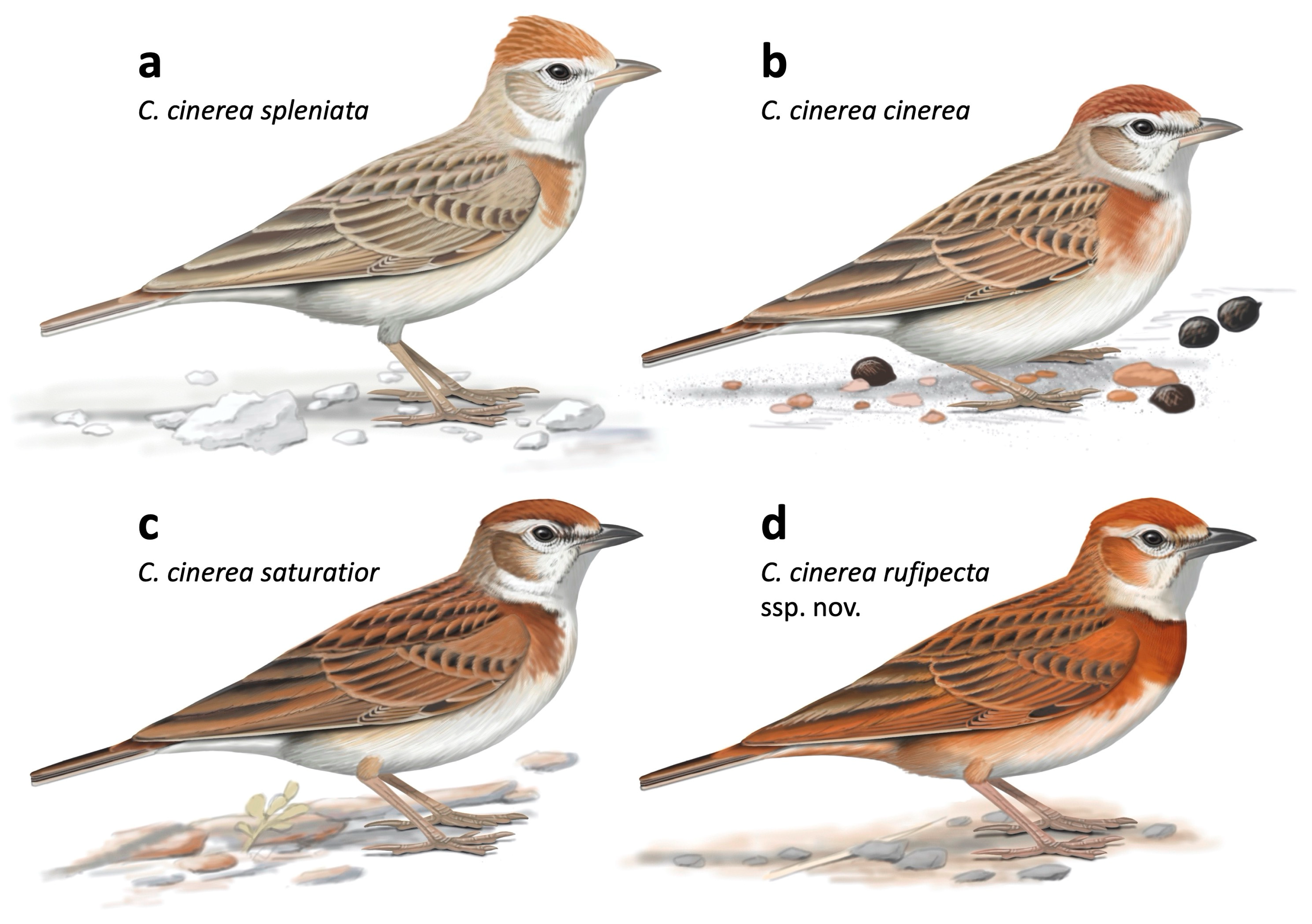
Figure 4.
Illustrations of the Calandrella cinerea complex, reflecting range of plumage variation within the southern clade (a–c), assigned to several subspecies, of which three are depicted here. However, cytochrome b sequencing recovers a panmictic population with no structure following geography or subspecies (Figure S1), and this likely represents clinal variation. We propose the name C. c. rufipecta ssp. nov. for the range-restricted Nigerian population (d). This is most closely related to C. c. williamsi (not depicted), distributed in southern Kenya and northern Tanzania. Note the rich rufous coloration of C. c. rufipecta, and its complete russet breast band, which differentiates it from all other taxa within the complex. Illustration © Faansie Peacock, from Donald and Alström [59], with permission.
Holotype. Adult (no sex determined). The Natural History Museum, Tring, UK, number NHMUK 1960.8.4, collected at Jos, N Nigeria, at 4500 feet, by R. E. Sharland. Stated to be the “first record” (for Nigeria). See Figure S7. Note that the NHMUK specimen label states 28 September 1958 as collection date, whereas Sharland’s own account states the collection was done 8 October 1958 between Laminga and Bukuru on the Jos Plateau [55] in present-day Plateau State.
Description of holotype. Plumage: Moderately worn head, body, wing coverts and tertials, fresh remiges and rectrices, indicating that it had gone through a complete moult fairly recently. Forehead and crown deep rufous, unstreaked, except for a few very fine dark streaks on rearmost crown. Nape marginally paler rufous than crown, with tiny, indistinct, dark grey-brown spots. Feathers on mantle and scapulars have broad dark grey-brown centres and rufous outer and inner edges; rufous tone similar to nape or, at least on extreme edges of some inner webs, paler, tending towards rufous-buff. Back similar to mantle, though with slightly less dark centres. Rump and uppertail-coverts rufous, slightly paler than crown, with narrow, diffuse, pale buffish fringes. Lesser coverts deep rufous, with dark grey-brown centres to lowest row. Median coverts have dark grey-brown centres and broad, rather clear-cut rufous tips. Greater coverts have dark grey-brown centres and broad, rather clear-cut rufous tips to outer webs (narrower on inner webs) and narrow, paler edges, grading towards pale buffish on very edges. Tertials dark grey-brown with narrow, diffuse, pale greyish edges; traces of broader rufous edges are visible at the concealed bases, so presumably the tertials had rufous edges when fresh. Alula dark grey-brown with rather broad rufous edges. Primary coverts and remiges dark grey-brown with narrow, clear-cut paler edges, which are rufous except on outer primaries, especially the outermost long (second) primary, which has a buffish-white outer edge. Underwing-coverts brown (difficult to examine without damaging specimen). Central pair of tail feathers dark grey-brown with narrow rufous edges, grading towards pale buffish on extreme edges and tips (probably due to wear and bleaching). Other rectrices dark grey-brown/blackish-brown, with pale rufous-brown outer web (narrowly extending onto tip of inner web) to distal c. half of outermost feather, grading to buffish-white on extreme edge; inner c. half of outer web of outermost feather dark along shaft, with narrow buffish-white outer edge. Lores, subocular area and distinct supercilium buffish-white, probably with a dark grey-brown loral stripe, at least close to eye (detailed pattern of side of head and throat difficult to determine due to style of preparation; cf. Figure S7). Ear-coverts rufous, a shade browner than the crown. Throat buffish-white, with a few dark speckles in malar region or/and along border between throat and breast. Side of neck and underparts buffish-white, with a broad rufous band across upper breast, slightly paler and with some diffuse buffish fringes in centre, lower edge slightly mottled; a few diffuse rufous-brown streaks on the flanks (concealed under wings). Bare parts: Bill pale brown with diffuse blackish tip to upper and lower mandibles. Tarsi and toes are pale brown, claws dark brown. Colour of pale part surely not identical to when alive; according to label: “Bill base pale brown, tip shading to black. Feet pale brown”. Wing formula (distance from wing tip in mm): Outermost long primary 2 (P2) 2, P3 0, P4 0, P5 1, P6 10.5, P7 17.5, longest tertial 1. The tip of the outermost short primary (P1) –10.5 mm in relation to the tip of the longest primary covers. Photos of this specimen are available in Figures S7 and S8.
Complementary description based on field examination and photos of a live individual with ring number FA67266 caught 31 January 2006 at Gwafan, Nigeria (9°53′ N, 8°56′ E). Plumage: Supercilium, subocular area, anterior ear-coverts and throat whitish; forehead and crown deep rufous; rear ear-coverts rufous-brown. Lores show a dark brown spot, not quite reaching bill. Mantle washed with rufous. Rump deep rufous. Breast-band deep rufous, complete; about 10 mm wide, a little darker on the sides. Belly and undertail-coverts whitish with some rufous mixed in. Flanks rufous, slightly paler than breast. Median and greater wing-coverts dark brown with rufous fringes; lesser wing-coverts deep rufous. Remiges dark brown with rufous-brown edges. Tail dark brown. Bare parts: Bill dark grey with diffuse pale grey base to lower mandible. Flight feathers: Emarginations on outer webs of primaries 3, 4 and 5. Wing length: 94 mm. Tail length: 63 mm. Wing formula (distance to wing tip in mm): P2 1, P3 0, P4 0, P5 2, P6 14, P7 17, P8 21, P9 23, P10 30, longest secondary 31. Photos of this individual, as well as from field observations, are available in Figure 3.
Comparison with other taxa. The main difference from the other taxa in the Calandrella cinerea complex is the complete rufous breast band in C. c. rufipecta, which is not shown by any other taxon. The nape, mantle and scapulars are deeper rufous than in any other taxon in the C. cinerea complex, although they are close to C. c. saturatior (including forms anderssoni and niveni). The rufous colour of the crown and breast is clearly deeper than in at least C. c. spleniata and C. c. millardi. The streaks on the mantle and scapulars are darker than in C. c. cinerea, C. c. niveni, C. c. spleniata, C. c. millardi, and C. c. williamsi. See Figure S8 for a comparison of specimens of the above taxa. As we have only had one specimen of C. c. rufipecta for direct comparison with other taxa, the differences in colour and streaking above need to be evaluated on a larger sample, whereas the breast pattern was obviously consistently different also in live birds. No black breast-side patches have been observed.
Diagnosis. The complete rufous breast band distinguishes this taxon from all other taxa in the Calandrella cinerea complex (Figure 4) as well as C. blanfordi and C. eremica.
Etymology. The name rufipecta means rufous breast and refers to the diagnostic rufous breast band.
Nomenclatural acts. The electronic edition of this article conforms to the requirements of Article 8.5.3 of the amended International Code of Zoological Nomenclature [57], and hence the new name contained herein is available under that Code from the electronic edition of this article. This published work and the nomenclatural act it contains have been registered in ZooBank, the online registration system for the International Commission of Zoological Nomenclature. The ZooBank Life Science Identifiers (LSIDs) can be resolved and the associated information viewed through any standard web browser by appending the LSID to the prefix “http://zoobank.org/”. The LSID for this publication is urn:lsid:zoobank.org:pub:95759AE0-C9F8-4403-9451-BED6770DE210 and the LSID for the nomenclatural act associated with C. c. rufipecta urn:lsid:zoobank.org:act:565ECF77-AEAF-42C0-B099-5C3696A49484. The electronic edition of this work was published in a journal with ISSN 1424-2818, and has been archived and is available from the digital repository http://www.ncbi.nlm.nih.gov/pmc/.
Distribution and status. The Nigerian population of C. cinerea was initially described as an uncommon and very local resident in open grasslands on the Jos Plateau, central Nigeria [55,56], and due to low levels of ornithological activity in that area very little is known about it. The most well-known classic site was around Vom (25 km SSW of Jos), however it has not been observed there since 1994 (Mark Hopkins, pers. comm.). During intensive censusing in 2003–2006 covering 2524 100-m transects at four sites, spanning 600 km2 of the Jos Plateau [58], the species was only observed twice, consisting of the rediscovery by M.F.H. and Ross McGregor of two birds at Gwafan (9°52′55′′ N, 8°57′3″ E; 10 km E of Jos; Figure 3) and of two birds near the village of Bisichi (9°42′33′′ N, 8°53’26” E; 25 km SSE of Jos) in February 2004. A follow-up visit to the Gwafan site in May 2004 resulted in an observation of three birds, suggesting a successful breeding attempt. Subsequently, several pairs were observed until the area was developed for housing, starting in 2008, after which it was observed sporadically at nearby sites until 2015, but not thereafter. During censusing for the Nigerian Bird Atlas Project, a pair was observed close to Bokkos (50 km SSE of Jos) in April 2017. Even after the establishment of A. P. Leventis Ornithological Research Institute 10 km E of Jos in 2001, and the considerably increased ornithological activity which followed, observations of C. cinerea rufipecta remain scarce.
3.3. Molecular Species Delimitation
3.3.1. Focal Group
For our focal genus Calandrella, we recovered the same clades as Stervander, et al. [27], and mPTP delimited them in nine species (Figure 5), compared to the currently recognized six [23]:
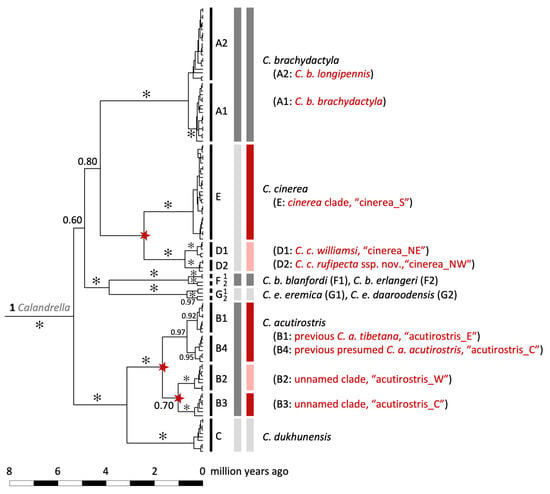
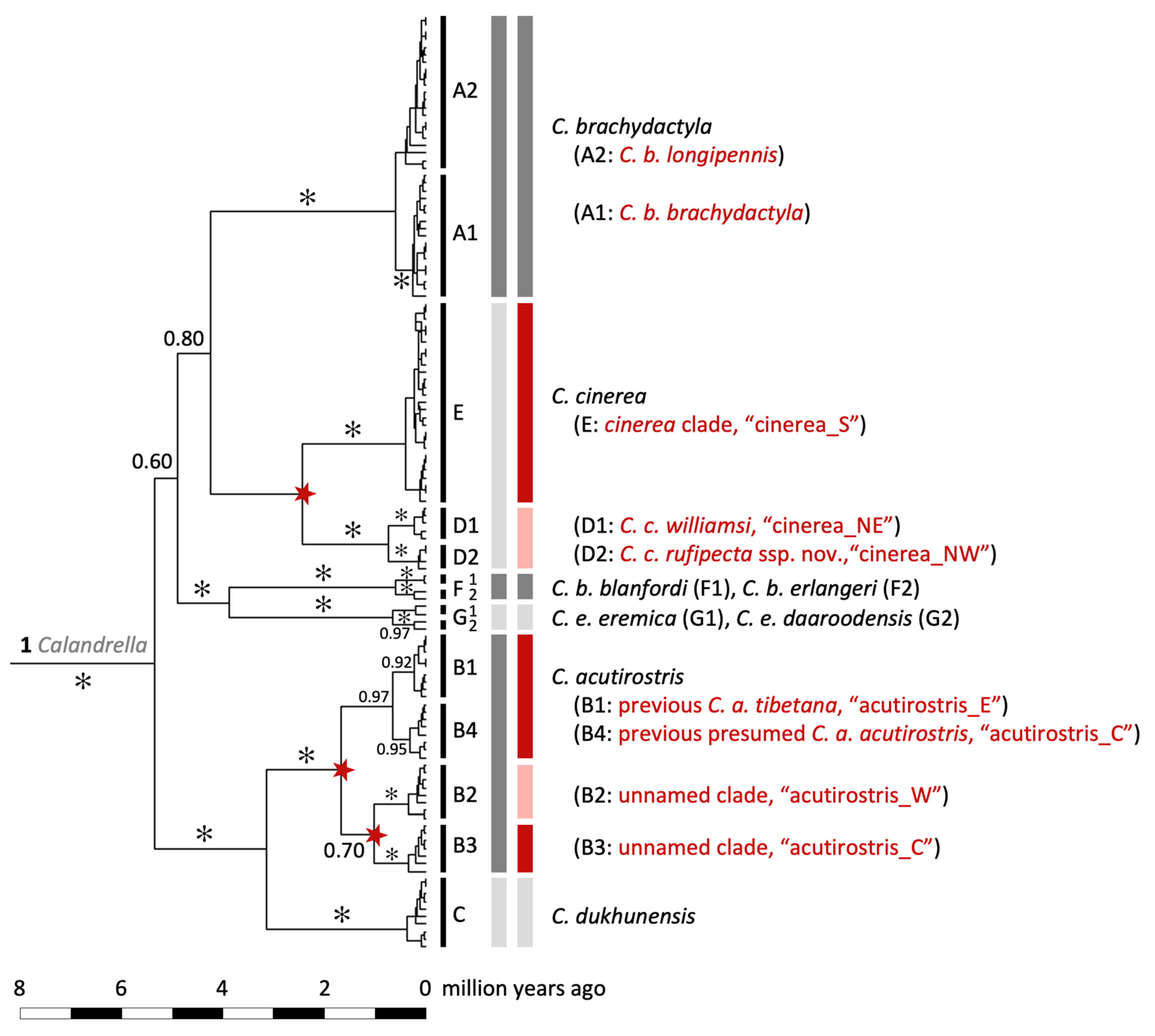
Figure 5.
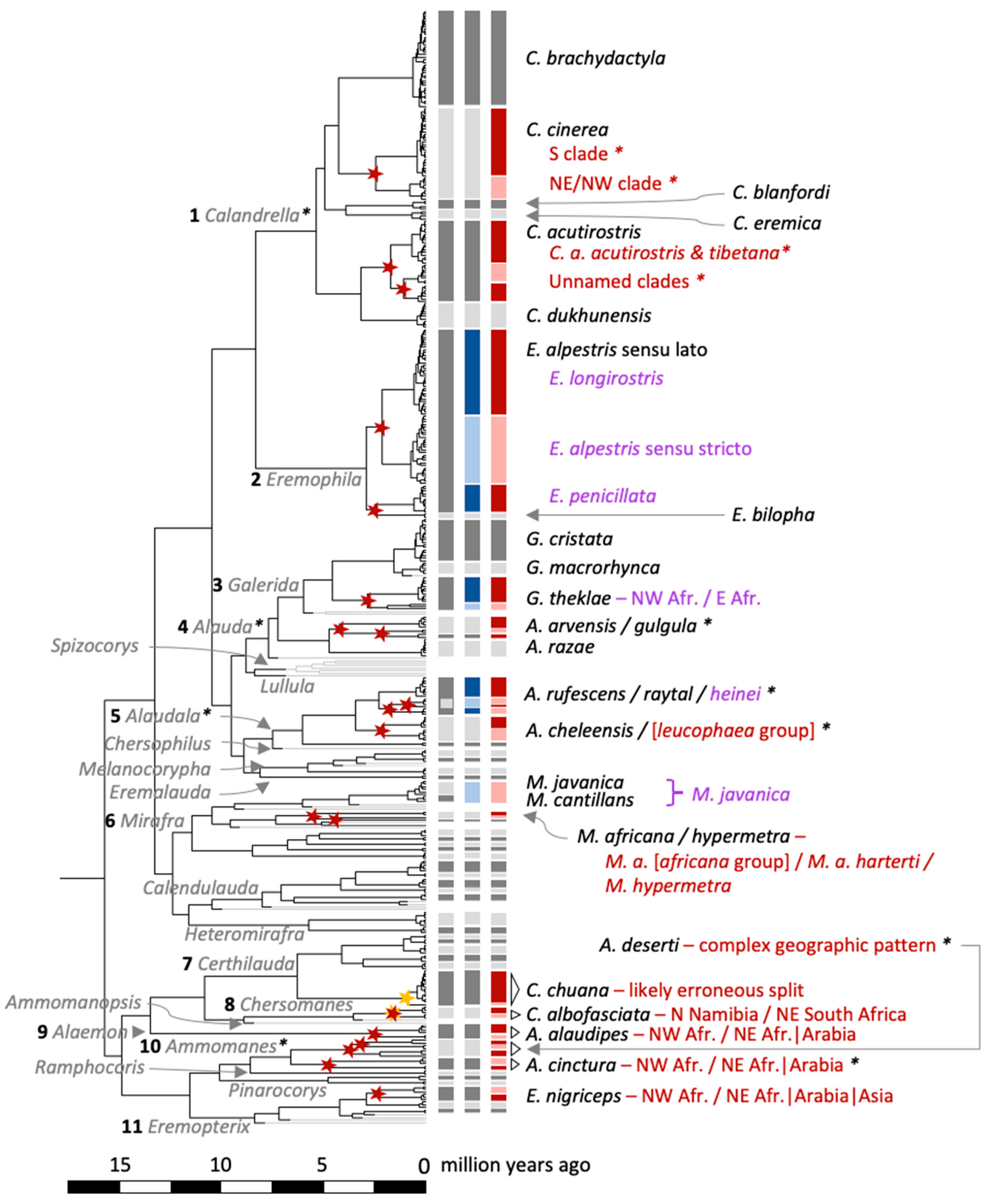
Maximum clade credibility tree for Calandrella larks based on the mitochondrial cytochrome b gene and a molecular clock rate corresponding to 0.0105 substitutions/site/million years. Posterior probabilities (PP) are given for major clades (✻, PP ≥ 0.99), which are delimited by narrow, vertical black bars at the tips. In the following columns to the right are (1) clade codes which are in accordance with Stervander, et al. [27], (2) currently recognized species [23] demarcated by wider grey bars (with alternating dark and pale shades for easier interpretation), and (3) results from the multi-rate Poisson Tree Processes (mPTP) species delimitation (wider grey and coloured bars; with alternating dark and pale shades for easier interpretation). When different from current taxonomy, bars are highlighted in red, and inferred novel speciation nodes are indicated by a red star. The taxon names and clade labels are the ones used in the text (e.g., “cinerea_S”).
- 1.
- Genus Calandrella
- C. brachydactyla, inferred as a single species by mPTP: Only two clear groups that split 0.71 (95% highest posterior density, HPD: 0.35–0.97) million years ago (MYA) (Figure S1), containing western birds (clade A1) and eastern birds (clade A2; Figure 5; Calandrella clade designations in accordance with Stervander, et al. [27]), but with no internal sorting within these clades according to currently recognized subspecies.
- C. cinerea, inferred as two species by mPTP: Split 2.55 (1.64–3.31) MYA between (1) a clade containing the cinerea group with all southern populations, the population in DR Congo and west of Lake Victoria (clade E), and (2) one clade comprising birds sampled in Kenya and Tanzania east of Lake Victoria (C. c. williamsi; clade D1) and Nigeria (described above as C. c. rufipecta ssp. nov.; clade D2).
- C. blanfordi, inferred as a single species by mPTP (clade F, including C. b. blanfordi and C. b. erlangeri).
- Rufous-crowned Lark C. eremica, inferred as a single species by mPTP (clade G, including C. e. eremica and C. e. daaroodensis).
- C. acutirostris, three species delimited by mPTP: Split at 1.78 (1.11–2.35) MYA between (1) a clade containing eastern populations in Tibet (clade B1, corresponding to the population traditionally assigned as C. a. tibetana in eastern Tibet and Qinghai, China) and eastern Ladakh, India (clade B4, birds traditionally assigned to C. a. tibetana); and a clade that, in turn, is split at 1.16 (0.45–1.75) MYA between (2) birds sampled in Afghanistan (clade B2); and (3) birds sampled in Ladakh (clade B3).
- C. dukhunensis, inferred as a single species by mPTP (clade C), separated from C. acutirostris at 3.27 (2.29–4.18) MYA.
3.3.2. Other Larks
Across the 47 currently recognized lark species outside the genus Calandrella [23] for which multiple cytochrome b sequences were available, mPTP delimited 60 species at a MCMC support value > 0.95 (Figure 6). The species that deviated from current taxonomy [23] were:
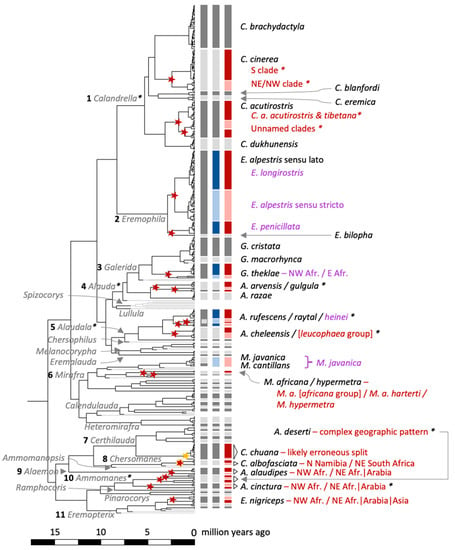
Figure 6.
Maximum clade credibility tree for Calandrella larks based on the mitochondrial cytochrome b gene and a molecular clock rate corresponding to 0.0105 substitutions/site/million years. All genera are labelled with grey font, and those discussed in the text are numbered. Species represented by single individuals (sequences) have grey branches and were not included in the multi-rate Poisson Tree Processes (mPTP) species delimitation, since they violate the underlying presumptions. The three columns with vertical bars to the right of the tree tips are, from left to right, (1) currently recognized species [23] in grey, (2) alternative species delimitation based on multiple sources (see Discussion) highlighted in blue, and (3) the mPTP species delimitation from our analysis highlighted in red; all colours with alternating dark and pale shades for easier interpretation. Inferred novel speciation nodes based on mPTP are indicated by a star (red star, credible delimitation; orange star, incorrect delimitation [see Discussion]). Clades/species in which species delimitation differs from current taxonomy are labelled with taxon names and/or geographic differentiation, with red font referring to our mPTP results (red bars), and purple font referring to when our results agree with alternative sources (blue bars). Clades for which more detailed figures are available are marked with an asterisk: Calandrella (Figure 5), Alauda, Alaudala, and Ammomanes (Figure S9). The full tree, including all taxa and providing the names of all individuals included, is available in Figure S1.
- 2.
- Horned Lark Eremophila alpestris and Temminck’s Lark E. bilopha, mPTP delimiting 4 species: Split at 2.25 (1.47–2.92) MYA between a clade containing (1) E. alpestris elwesi and E. a. deosaiensis (Tibet–Himalaya) and (2) a clade comprising multiple subspecies (e.g., alpestris, flava, brandti) across the North Palearctic and North America. A clade comprising (3) E. a. penicillata/albigula (Iran) and E. a. atlas (Morocco) split at 2.61 (1.75–3.41) MYA from (4) E. bilopha (Morocco and Saudi Arabia). The deepest split within Eremophila was at 3.00 (2.12–3.84) MYA.
- 3.
- Thekla’s Lark Galerida theklae, mPTP delimiting 2 species: Split 2.94 (1.99–3.77) MYA between (1) a clade of NW African birds sampled in Morocco (G. t. theklae) and Tunisia (G. t. superflua) and (2) a clade comprising E African birds sampled in Ethiopia (G. t. praetermissa and G. t. hueii) and Somalia (G. t. ellioti).
- 4.
- Eurasian Skylark Alauda arvensis and Oriental Skylark A. gulgula, mPTP delimiting 3 species: (1) western A. arvensis (sampled across Europe east to Kazakhstan) diverged at 4.33 (3.17–5.41) MYA from a clade comprising (2) A. gulgula (likely A. g. inconspicua) from Kazakhstan and India and (3) eastern birds (A. arvensis japonica from Japan and a bird sampled in Anhui Province in SE China, which split at 2.26 (1.43–3.02) MYA.
- 5.
- Genus Alaudala, mPTP delimiting 6 species in the rufescens/cheleensis/raytal complex that is traditionally divided in two or three species:
- Asian Short-toed Lark A. cheleensis: Split at 2.24 (1.36–2.94) MYA between (1) A. c. cheleensis (sampled in eastern Mongolia and Inner Mongolia, China) and (2) a clade containing A. c. leucophaea (Kazakhstan) and A. c. seebohmi (Xinjiang, China).
- Sand Lark A. raytal: Split at 1.01 (0.52–1.36) MYA between (3) a single sample of A. r. adamsi (Punjab, India) and (4) a clade containing A. r. raytal (Haryana and Delhi, India).
- Lesser Short-toed Lark A. rufescens: A clade comprising (5) A. r. heinei (Kazakhstan) and A. r. aharoni (Turkey) split at 1.90 (1.23–2.44) MYA from A. raytal, whose common ancestor split at 2.39 (1.64–3.05) MYA from (6) a clade comprising A. rufescens rufescens and A. r. polatzeki (Canary Islands), A. r. minor (Morocco and Saudi Arabia), and A. r. apetzii (Spain).
- 6.
- Genus Mirafra
- Rufous-naped Lark M. africana, mPTP delimiting 2 species: Split at 5.69 (3.87–7.36) MYA between M. a. transvaalensis (South Africa) and M. a. harterti (Kenya; the latter grouping with low support with Red-winged Lark M. hypermetra).
- Singing Bush Lark M. cantillans and Horsfield’s Bush Lark M. javanica were inferred by mPTP to be conspecific, with a most recent common ancestor at 0.97 (0.54–1.27) MYA.
- 7.
- Short-clawed Lark Certhilauda chuana, mPTP delimiting 2 species: Split at 0.98 (0.40–1.48) MYA between (1) a clade comprising a dozen samples from the disjunct western and eastern South African populations, and (2) a single bird from the eastern South African population.
- 8.
- Spike-heeled Lark Chersomanes albofasciata, mPTP delimiting 2 species: Split at 1.62 (0.72–2.53) MYA between one clade comprising (1) C. a. boweni (Namibia) and (2) another clade comprising samples from the North West and Limpopo provinces of South Africa, presumably all of C. a. alticola.
- 9.
- Greater Hoopoe-Lark Alaemon alaudipes, mPTP delimiting 2 species: Split at 2.65 (1.56–3.85) MYA between (1) Northwest African A. a. alaudipes (sampled in Morocco) and A. a. boavistae (Cape Verde) and (2) Northeast African/Arabian A. a. desertorum (Egypt and Saudi Arabia).
- 10.
- Genus Ammomanes
- Bar-tailed Lark Ammomanes cinctura, mPTP delimiting 2 species: Split at 4.86 (3.33–6.25) MYA between (1) Northwest African A. c. arenicolor (sampled in Morocco) and A. c. cinctura (Cape Verde) and (2) Northeast African/Arabian A. c. arenicolor (sampled in Saudi Arabia).
- Desert Lark Ammomanes deserti, mPTP delimiting 3 species: Split at 3.28 (2.26–4.22) MYA between (1) Northwest African A. d. payni (sampled in Morocco) and (2) Northeast African/Arabian A. d. isabellina (sampled in Saudi Arabia) and A. d. annae (Jordan). This clade split at 3.80 (2.73–4.75) MYA from a clade comprising (3) A. d. deserti (sampled in Israel) and A. d. phoenicuroides (sampled in Pakistan).
- 11.
- Black-crowned Sparrow-Lark Eremopterix nigriceps, mPTP delimiting 2 species: Split at 2.45 (1.50–3.30) MYA between (1) Northwest African E. n. albifrons (sampled in Mauritania) and E. n. nigriceps (Cape Verde) and (2) Northeast African/Arabian/Asian E. n. melanauchen (in Saudi Arabia) and E. n. affinis (in Pakistan).
4. Discussion
4.1. Divergence within African and African–Arabian Lineages
4.1.1. Fragmented Sub-Saharan Species Distributions and Relictual Lineages
The herein described Calandrella cinerea rufipecta ssp. nov. is highly localized and endemic to the Jos Plateau in Nigeria, which houses two endemic bird species, the Rock Firefinch Lagonosticta sanguinodorsalis and the Jos Plateau Indigobird Vidua maryae [60]. This population is situated over 1500 km from the closest population within the C. cinerea complex (C. c. saturatior in DR Congo) and over 3000 km from its closest relative, C. c. williamsi in the eastern Rift Valley in southern Kenya and northern Tanzania [61]. It seems plausible that C. c. rufipecta and C. c. williamsi are relictual lineages remaining from a once more widely distributed common ancestor in a northern sub-Saharan belt, which became fragmented around 0.86 (95% HPD 0.42–1.15) MYA.
The pattern of a disjunct distribution in savanna habitat across West Africa, East Africa and southern Africa resembles that of Mirafra africana (clade 6a in Figure 6; Figure S10), which—in addition to isolated populations in eastern West Africa, the East African Rift Valley, and southern Africa—also has further disjunct, unsampled, populations, including in westernmost West Africa [62]. The only sequences available represent a southern population (M. a. transvaalensis) and one bird from the Rift Valley population (M. a. harterti [25]). The latter cannot confidently be placed as sister to M. a. transvaalensis, but instead is placed as sister to M. hypermetra (though with low support; PP 0.63). Mirafra hypermetra has disjunct East African populations, the main of which overlaps marginally with M. africana, with which it has sometimes been considered conspecific [63]. The split between M. a. harterti and M. hypermetra dates to mid-Pliocene 4.54 (2.90–6.01) MYA, whereas the split between southern M. africana and M. hypermetra dates to the late Messinian of Miocene 5.69 (3.79–6.79) MYA, both rather deep divergences on par with species level divergence times within Mirafra (Figure S1). The pattern also broadly resembles that found in multiple lineages of birds, e.g., [64,65,66], mammals, e.g., [67,68,69,70,71,72] and plants, e.g., [73].
Similarly disjunct distributions, although with less strong geographical isolation and/or covering fewer major African regions, are shared with several other species within the genus Mirafra: Flappet Lark M. rufocinnamomea (Ryan, et al. [74]; a single lineage sequenced [25]), White-tailed Lark M. albicauda (Ryan, et al. [75]; no genetic data), and M. cantillans (BirdLife International [76]; lineages from Kenya, Saudi Arabia, and India sequenced [25,77], grouping with M. javanica; clade 6b in Figure 6; populations in the Sahel zone not sampled). Kordofan Lark Mirafra cordofanica [78] and Rusty Bush Lark M. rufa [79] both lack genetic data and have disjunct distributions along a west–east gradient along the Sahel zone. Among other genera, Chestnut-backed Sparrow-Lark Eremopterix leucotis (no genetic data) is distributed continuously from West to East Africa, with substantial breaks between that and populations in central–eastern and western regions of southern Africa [80].
4.1.2. Fragmented NW African vs. E African/Arabian Populations
Across or north of the Sahara, several other lark species have fragmented populations reaching from Northwest Africa (sometimes including the Cape Verde archipelago) east to Northeast Africa and Arabia (and sometimes extending eastward further into Asia). Three of the species for which mPTP inferred Pliocene splits are in this category: Alaemon alaudipes (de Juana, et al. [81]; clade 9 in Figure 6; divergence at 2.63 [1.56–3.85] MYA), Ammomanes cinctura (de Juana, et al. [82]; clade 10a; 4.86 [3.33–6.25] MYA), and A. deserti (de Juana, et al. [83]; clade 10b; 3.80 and 3.28 [2.73–4.75 and 2.26–4.22] MYA). A similar distribution is shown by Eremophila bilopha de Juana, et al. [84], which—in contrast—shows no divergence at cytochrome b between birds from Morocco [37] and Saudi Arabia [77]. Thick-billed Lark Ramphocoris clotbey has the most disjunct distribution, with a break between central northern Libya and eastern Israel [85], but only a Moroccan bird has been sequenced [25].
In accordance with detailed studies by Guillaumet, et al. [30], mPTP delimited the disjunct populations of Galerida theklae (clade 3) in Northwest Africa and East Africa as different species, diverged during the Pliocene at 2.94 (1.99–3.77) MYA. Finally, Eremopterix nigriceps supposedly has a continuous distribution from West Africa across the Sahel zone to East Africa, the Horn of Africa, Arabia, and eastward to northwestern India [86]. Nevertheless, there is an Early Pleistocene divergence dated at 2.45 (1.50–3.30) MYA between, on the one hand, West African and Cape Verdean populations [87] and, on the other hand, Arabian and South Asian populations [25,77].
4.1.3. Biogeography of Fragmented African Populations
Africa houses almost a quarter of the global avian diversity but has received relatively little recent attention to taxonomy and biogeography compared to, e.g., South America [88]. Within the lark family, more detailed studies have been made on African species in the genera Certhilauda [33], Calendulauda [34], Galerida [30,31], Heteromirafra [89], and Calandrella [27], this study, but overall there is limited understanding of taxonomic and biogeographic patterns. These can be further obscured by two opposing patterns of phenotypic differentiation. On the one hand, there can be substantial within-species plumage variation not reflected in mitochondrial genetic differentiation [25], e.g., due to the matching of substrate colour [29]. On the other hand, very little morphological differentiation may accrue over time within genera (Figure 1 and Figure 2b; Guillaumet, et al. [30]). Moreover, distantly related clades in different genera may display spectacular convergence in plumage and bill morphology, whereas some more closely related lineages have diverged substantially in morphology [25].
Larks exclusively inhabit open habitats—desert and shrubland, semi-desert, steppe, and savanna—habitats whose distribution have changed dramatically during Pliocene and Pleistocene, both in North Africa [90,91] and sub-Saharan Africa [92,93]. The habitat distributions have been influenced by East African orogeny [94] as well as being largely determined by changes in how the exposure to solar radiation, induced by the Earth’s orbital eccentricity, affected monsoon dynamics [93]. During warmer/wetter periods, forests displaced savanna and arid landscapes, and would have fragmented and isolated open habitat species. During cooler/drier periods, forests retracted and were replaced by grasslands, allowing open habitat species to expand and come into secondary contact. Most oscillations lasted some 20–40 thousand years [92], however there have been prolonged periods of climatic instability during warmer/wetter conditions that lasted several hundred thousand years, which have been suggested to coincide with increased speciation and extinction rates in hominins [95] and other mammals [93]. Several prolonged periods of maximal climatic variability occurred 2–3 MYA, an interval that coincides with five of the ten species splits inferred by our mPTP analyses (genera Alaemon, Calandrella, Galerida, Eremophila, Eremopterix), the other five being older (genera Ammomanes (two splits), Mirafra) or younger (genus Chersomanes) and further deemed invalid (genus Certhilauda; see Section 4.3).
Just like relic populations of African forest birds have been identified as remnants left from large, continuous forests, e.g., [96,97,98], the Calandrella cinerea populations in West (C. c. rufipecta ssp. nov.) and East Africa (C. c. williamsi) are relics in savanna habitat, historically presumably separated from populations of C. c. cinerea by the extension of the Congo Basin rainforest. We also note that the similarities in disjunct distributions across West, East, and southern Africa between several lark species [61,62,63,74,75,76,78,79,80,81,82,83,84,85] to some degree is mirrored by several concordantly distributed mammalian savanna specialists, namely ungulates [68].
4.2. Taxonomy of Calandrella Larks
4.2.1. The Calandrella cinerea Complex
The C. cinerea complex diverged from other Calandrella larks in the early Pliocene (Figure 5), and the molecular species inference based on cytochrome b split it into two species: one comprising C. c. williamsi in the Rift Valley of East Africa and the Nigerian C. c. rufipecta ssp. nov., and one comprising all other populations (Figure 5), which among themselves show no differentiation in cytochrome b (Figure S1). While C. c. rufipecta has distinctly different plumage characters compared to all other taxa in the C. cinerea complex, C. c. williamsi does not show any diagnostic morphological characters. Morphometrics overall distinguishes C. cinerea from other African congeners (Figure 2a), but only partly differentiates C. c. williamsi from the remaining subspecies of C. cinerea (Figure 2b). While it is plausible that C. c. rufipecta and C. c. williamsi should be considered a separate species, as inferred by mPTP, we urge sequencing of nuclear DNA beyond Stervander, et al. [27], and analyses of vocalizations and other non-genetic data prior to any taxonomic revision. We note a lack of differentiation among the different subspecies of C. cinerea in clade E, and that the plumage variation is at least partly clinal (Figure S1; [61]). Accordingly, the validity of these subspecies should be re-evaluated based on independent data.
4.2.2. Calandrella dukhunensis and Calandrella brachydactyla
Until recently, Calandrella dukhunensis was treated as a subspecies of C. brachydactyla, which it closely resembles in appearance. Alström, et al. [25] demonstrated that it was strongly differentiated in mtDNA from C. brachydactyla and instead sister to C. acutirostris. As their analyses were based solely on mtDNA, the pattern could have been caused by historical introgression of the mitochondrion from C. acutirostris [99,100] or “ghost” introgression [101,102,103]. However, subsequent genomic analyses corroborated its placement [27], inferring the same topology for the nuclear genome. In addition, vocal and behavioural data have recently been put forward as further support of the species status of this taxon [28].
At the species level, mPTP agrees with current taxonomy [23]. However, similar to C. cinerea, a multitude of subspecies have been described for C. brachydactyla, receiving little phylogenetic support (Figure S1). We recovered a shallow divergence estimated at 0.71 MYA between a western clade (including subspecies brachydactyla, rubiginosa, and hungarica) and an eastern clade (including subspecies longipennis, artemisiana, hermonensis, and woltersi), separating roughly across the Aegean Sea (with birds of both the western and eastern clades sampled on Crete, Greece, in mid–late May), although the determination of exact bounds, and assessment of a contact zone, would require dense geographic sampling. Thus, mitochondrial data suggest synonymisation into only two subspecies, brachydactyla and longipennis, that do not differentiate structurally (Figure 1b). Studies of plumages suggest mostly clinal variation within these two taxa, supporting recognition of two or perhaps three subspecies [59].
4.2.3. The Calandrella acutirostris Complex
Within C. acutirostris, Stervander, et al. [27] uncovered four distinct mitochondrial clades (designated B1–B4; Figure 5), two of which were also sequenced for nuclear genomic data that reflected this divergence. Here, mPTP delimited clades B1+B4, B2, and B3 as three different species (Figure 5) diverged in the Pleistocene. Clade B1 is the easternmost population, with samples from Qinghai Province, China, and eastern Tibet, reaching west to easternmost Ladakh, and corresponds to the main distribution of the currently recognized subspecies C. a. tibetana [23,104]. However, this clade contains the holotype of C. a. acutirostris (NHMUK 1887.7.1.3739; Figure S1), collected in the upper Karakash Valley of the Sughet Range. Its sister clade B4, with a divergence time estimated at 0.77 MYA, was found exclusively in samples from eastern Ladakh [27]. Clade B2 (Figure 5) represents western birds, traditionally assigned to C. a. acutirostris [23,104], all sampled in Afghanistan [27]. The samples of clade B3, which form the sister clade to B2, are distributed similarly to clade B4, but with a more westerly centre of gravity, with the majority in central/western Ladakh (present day Gilgit–Baltistan). The placement of clades B2 and B3 as sisters diverging at 1.16 MYA is uncertain, with posterior probability of 0.70. Their divergence from clades B1+B4 was estimated at 1.78 MYA.
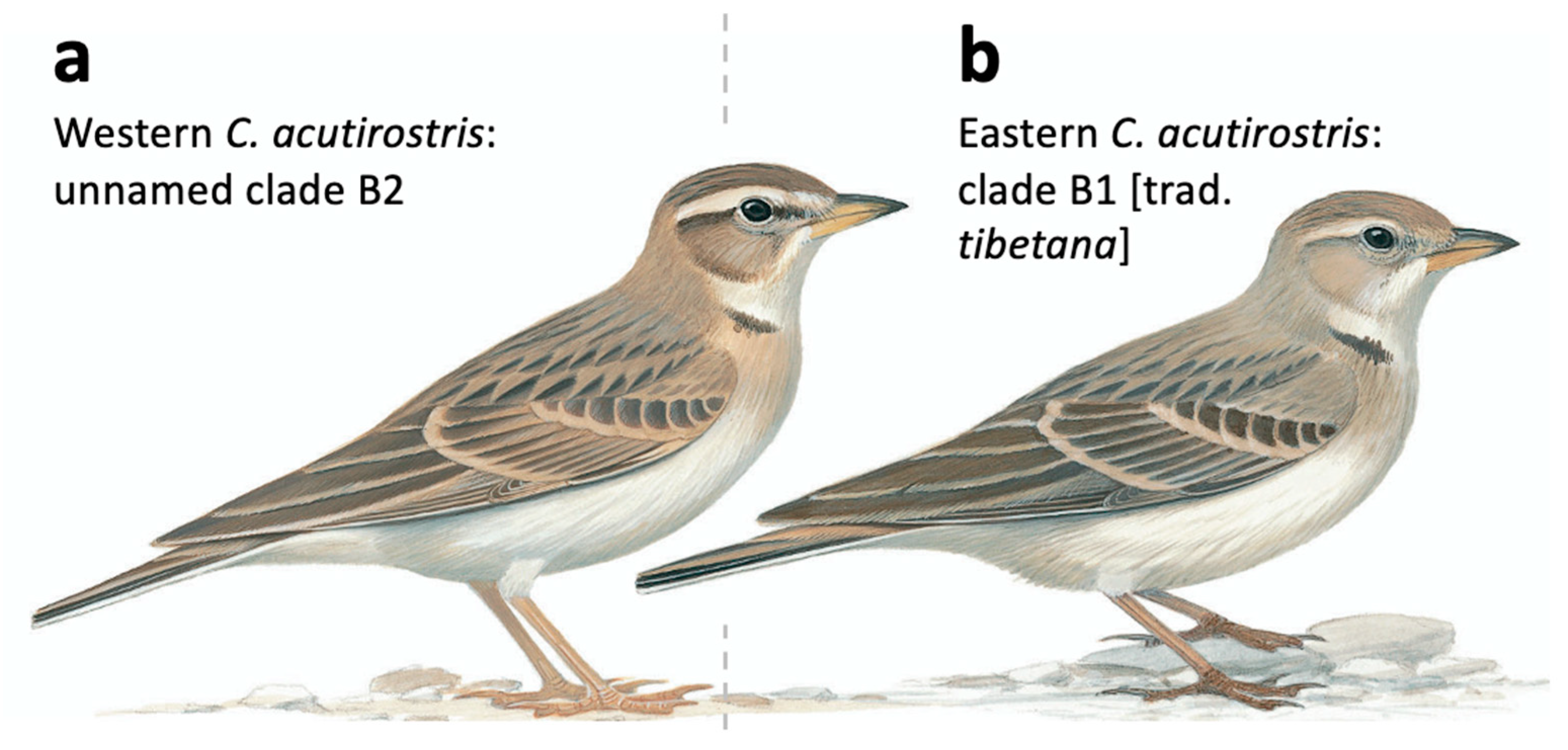
Morphologically, the westernmost (B2) and easternmost (B1) clades differ subtly. While there is no difference in wing length, western birds have on average longer bill and longer tail, and a larger tail/wing ratio (Table 1), separating primarily along an axis of bill length and tail/wing ratio (Figure 1d). Central populations in Ladakh (clades B3 and B4) are intermediate, and since all measured specimens were not genotyped, they were grouped together in the morphometric analyses. Based on studies of birds in the field and museum collections, the head patterns usually differ: western birds have a more contrasting head pattern, with darker crown, loral stripe, stripe behind the eye and patch on rear ear-coverts, and whiter and more prominent supercilium than eastern birds (Figure 7a).
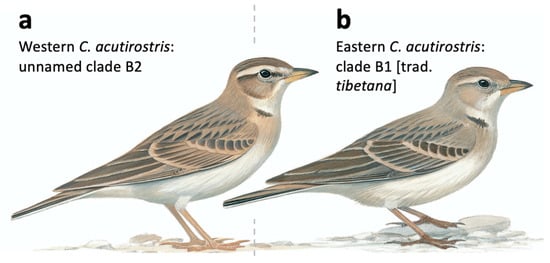
Figure 7.
Illustrations of the Calandrella acutirostris complex. (a) Western birds (Ladakh and westwards), corresponding to clade B2 (and likely B3) in the present study and in Stervander, et al. [27]. (b) Eastern birds (Ladakh and eastwards), corresponding to clade B1 (and likely B4) in the present study and in Stervander, et al. [27]. Note, especially, more contrasting head pattern of birds in the western clade. We conclude that the name C. a. tibetana, which is usually applied to Tibetan birds (clade B1), is a junior synonym of C. a. acutirostris, leaving western birds without a subspecific name. Illustration © Bill Zetterström, from Donald and Alström [59], with permission.
As the holotype of C. a. acutirostris was genetically identified as belonging to clade B1, which corresponds to traditional C. a. tibetana, C. a. tibetana is a junior synonym of C. a. acutirostris (based on priority). We are not aware of any available name for the western populations, which are distinctive based on mtDNA (Figure 5) and plumage (Figure 7). Further study of this complex using more extensive sampling, nuclear markers and/or analyses of vocalizations and detailed geographical distributions are warranted to revise the taxonomy.
4.3. Reliability of Molecular Species Delimitation
While mitochondrial trees have traditionally been used for inference of evolutionary history [105] and thereby taxonomy, it is now well established that mitochondrial markers may reflect a different history than the nuclear genome and present a tree that differs from the overall evolutionary history, e.g., [100,102,106,107]. For our focal clade, Calandrella, we combined a mitochondrial tree with morphometric data obtained in museum collections. However, we acknowledge that the Calandrella larks are poorly differentiated morphometrically (Figure 1), and we stress that other types of data, in combination with dense taxon sampling, are necessary for confident taxonomic revisions. Here, we lack substantive data from nuclear DNA markers, vocalizations, other behaviours, ecology etc., to take the fully integrative taxonomic approach [108,109,110] that is becoming more common, e.g., [111,112,113,114,115,116,117,118,119,120] and which we argue should be the gold standard. Yet, while we emphasize that taxonomic decisions should never rest on mitochondrial trees alone [101,106], we here make use of available morphometric data and evaluate a recent tool for single-locus molecular species delimitation [54], which has been used to propose avian taxonomic revisions based solely on mitochondrial data [121] or in combination with morphometry [65].
The molecular species delimitation with mPTP, based on cytochrome b, inferred several species which seem evolutionarily and biogeographically plausible. In those cases where detailed studies based on phylogenetic analyses in combination with independent data are available, the present mPTP delimitations largely agree: For example, based on mitochondrial and nuclear DNA sequence data, Ghorbani, et al. [26] tentatively suggested the same species delimitations for Alaudala, as did mPTP (clade 5; Figure 6) except that they made a reservation regarding the leucophaea–seebohmi group because of shallower nuclear DNA divergence and non-monophyly. Further, they note that the single sample of A. r. adamsi, that was separated from A. r. raytal by mPTP, requires further study and a larger sample size [26]. Alström, et al. [120] analysed molecular as well as non-molecular data and suggested recognition of four species in the A. rufescens–cheleensis–raytal complex: A. cheleensis (s.s.), A. rufescens (s.s.), A. raytal and A. heinei. As already noted above, the species status of Calandrella dukhunensis suggested by mPTP has been corroborated by genomic [27], as well as non-genetic data [28]. Ghorbani, et al. [37] suggested that the same four clades of Eremophila (clade 2; Figure 6) should be treated as species (whereas Drovetski, et al. [122] proposed to recognise six species based mainly on mtDNA); Guillaumet, et al. [30] suggested that Galerida larks (clade 3; Figure 6) should be delimited in the same way as suggested by mPTP (of which the split between G. cristata and G. macrorhyncha has been implemented [23], but not the split of G. theklae, pending further studies).
Mirafra javanica and M. cantillans (clade 6b) have alternately been treated as conspecific or as separate species [25] and are currently lumped by BirdLife International [123] but not by IOC [23]. Alström, et al. [25] suggested the taxa should be treated as separate species in the early stage of speciation. Our dating of the Alaudidae tree overall resulted in somewhat younger node ages than did Alström, et al. [25], and the estimated divergence between M. javanica and M. cantillans was 0.97 MYA (95% HPD: 0.5–1.3 MYA; vs. 1.2 MYA; 0.7–1.7 MYA, 95% HPD according to Alström, et al. [25]). The split of Chersomanes albofasciata (clade 8) at 1.62 MYA represents the endpoints of the species’ main continuous distribution [124] with one clade containing two independently obtained sequences [25,77] that subsequently turned out to likely originate from the same individual of C. a. boweni in N Namibia [P. Bloomer pers. comm.], and one clade containing NE South African birds of C. a. alticola. Removing one of the presumed duplicates does not affect the mPTP delimitation of two species within C. albofasciata, and the split between distant populations is consistent with patterns based on shorter cytochrome b sequences [125]. Perhaps the split would not have been inferred had we had access to sequences of geographically intermediate populations in the Karoo and Namaqualand region—see [125]—but on the other hand it is comparably deep.
The species within Alauda (clade 4) show a complex phylogenetic pattern, with A. razae sister to the others (this position should be regarded as unresolved, as the posterior probability is low; Figure S1; however, the basal position of A. razae is also recovered based on nuclear DNA [126]), while A. arvensis and A. gulgula are distributed over three clades that are delimited as species by mPTP. The Japanese taxon japonica is currently recognized as a subspecies of A. arvensis [23] but has historically sometimes been treated either as a subspecies of A. gulgula, or as a separate species [127]. Our phylogenetic analyses of cytochrome b confidently place a clade with japonica and a bird from SE China [128]; possibly representing A. g. weigoldi or A. g. coelivox. as sister to A. gulgula sampled in Kazakhstan and India, diverged during Early Pleistocene, and delimited as a separate species by mPTP. These sister species diverged from European and Central Asian A. arvensis in the Early Pliocene. The deep mitochondrial split between western and eastern A. arvensis has been observed previously [129], but nuclear DNA shows a starkly different pattern [126], and a more complex history is suggested in a comprehensive study of mitochondrial and nuclear DNA [130].
Mitochondrial patterns such as those exhibited by the Alauda arvensis–gulgula complex may be in agreement with the species phylogeny, but could represent a simplified or incorrect phylogeographic pattern, both because of sparse geographical sampling and because of potential introgression and/or incomplete lineage sorting. Independent data, such as nuclear genetic markers, morphological and bioacoustic data, are justified to properly resolve species limits. These cautions are also warranted when interpreting the remaining mitochondrial splits within Alaemon, Ammomanes, Mirafra, and Eremopterix suggested by mPTP. However, it has recently been suggested that the songs differ between western and eastern populations in both Ammomanes cinctura and Eremopterix nigriceps [131], in agreement with the mtDNA data. The same has been suggested for different populations of Ammomanes deserti [131], though no detailed studies of the songs have been undertaken.
In contrast to the above examples, one species split inferred by mPTP appears incorrect, as it does not mirror any phylogeographic pattern, and may rather reflect high within-population mitochondrial diversity, cf., e.g., [103]: for Certhilauda chuana (clade 7) a split divided a single eastern bird from sympatric eastern individuals and disjunct western populations.
5. Conclusions
Our study suggests that opposing patterns of phenotypic differentiation and convergence in the open-habitat specialist family Alaudidae have resulted in a taxonomy that often does not reflect the evolutionary history as inferred by mitochondrial markers, and previously by multilocus data [25]. Calandrella is the only genus for which we have analysed both morphometric and molecular data, but we conclude that its Bauplan is highly conserved, as morphometric analyses reveal only moderate differentiation (Figure 2 and Figure 3). On the one hand, panmixia at mitochondrial DNA together with clinal plumage differences suggest that synonymisation of multiple subspecies within C. brachydactyla and C. cinerea might be justified; on the other hand, in two species complexes—C. acutirostris and C. cinerea—we describe genetic, morphometric, and plumage differentiation that may warrant future taxonomic re-evaluation, with molecular species delimitation suggesting species splits. In the C. cinerea complex, we formally describe the population on the Jos Plateau in Nigeria as a new subspecies, C. c. rufipecta ssp. nov., and note that it is highly localized and isolated, with a probably small and threatened population. While subspecific populations generally attract less conservation efforts [6,8,9], and the application of a strictly phylogenetic species concept may have alleviated this potential problem by disassociating C. c. rufipecta from the very large population and wide distribution of southern African C. cinerea [132], we recommend revisiting the C. cinerea species complex for a fully integrative taxonomic assessment. Meanwhile, we stress that, as a genotypically and phenotypically distinct lineage, the conservation of C. c. rufipecta should deserve full priority.
Our application of single-locus molecular species delimitation revealed patterns similar to C. cinerea for other lark species with fragmented sub-Saharan distributions, as well as for several species with Northwest African–Arabian/East African distributions, indicating that—if the mitochondrial patterns indeed reflect the species’ evolutionary history—Africa may harbour much hidden species diversity that has been overlooked within this family of morphologically poorly differentiated birds. Most inferred species splits are within a reasonable age range, and plausibly follow historic climatic and biogeographic patterns. For understudied taxa, we demonstrate, for the first time, deep divergences within the Mirafra africana–hypermetra complex, that date to late Miocene and mid-Pliocene. Deep intraspecific divergences were further found in the genus Ammomanes, Alaemon alaudipes, and Eremopterix nigriceps, leading us to call for extended geographic sampling of multiple individuals per population, sequencing of nuclear DNA, and integration of morphological, behavioural, and ecological data.
Although we do not support species delimitations based on mtDNA alone, as implemented in programs such as mPTP, mtDNA can be a very useful first indicator of lineage divergence that can be further evaluated by independent data, such as nuclear DNA, morphology, bioacoustics, and ecology.
Supplementary Materials
The following are available (as .pdf file) online at https://www.mdpi.com/1424-2818/12/11/428/s1, Figure S1: Full phylogenetic tree, Figure S2: Standard morphometrics PCA for males, Figure S3: Quality of representation of 4 PCA variables for Calandrella acutirostris, Figure S4: Quality of representation of 6 PCA variables for African Calandrella, Figure S5: 9-variable PCA and quality of representation for African Calandrella, Figure S6: ROC curves for LDA of C. acutirostris and C. cinerea. Figure S7: Photos of the type specimen of C. cinerea rufipecta ssp. nov., Figure S8: Comparative photos of C. cinerea specimens, Figure S9: Species delimitation details for clades of genera Alauda, Alaudala, Ammomanes, Figure S10: Distribution map of the C. cinerea complex, Mirafra africana, and M. hypermetra, Table S1 comprises details—including taxonomic affinity, geographic origin, and accession numbers—of the included sequences/samples.
Author Contributions
Conceptualization, M.S. and P.A.; resources, B.H., U.O. (Urban Olsson), M.F.H., U.O. (Ulf Ottosson), and P.A.; methodology and data collection, M.S., P.A., M.F.H, U.O. (Urban Olsson); analysis, M.S.; writing—original draft preparation, M.S.; writing—review and editing, P.A., U.O. (Urban Olsson), B.H.; data visualization, M.S. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Swedish Research Council (2015-04402, P.A.; 2016-00689, B.H.), Jornvall Foundation and Mark and Mo Constantine (P.A.) with support from BirdLife Sweden (M.S.).
Acknowledgments
We could access museum collections for the recording of morphometrics thanks to Michel Louette, Alain Reygel and Gael Carin (RMCA); Mark Adams, Robert Prys-Jones, and Hein van Grouw (NHMUK); Paul Sweet, Peter Capainolo, and Thomas J. Trombone (AMNH); Janet Hinshaw (UMMZ); and Jon Fjeldså and Jan Bolding Kristensen (ZMUK). Yun Li kindly provided unpublished cytochrome b sequences of Alauda razae. Faansie Peacock and Bill Zetterström graciously let us use their Calandrella illustrations from a forthcoming book [59]. Ross McGregor kindly provided field photographs of C. c. rufipecta. Irene Tieleman, Joseph Williams, and Paulette Bloomer provided geographic information for previously sequenced samples. This is contribution no. 162 from the A. P. Leventis Ornithological Research Institute.
Conflicts of Interest
The authors declare no conflict of interest.
Data Accessibility
The unpublished cytochrome b sequences have been deposited at GenBank with the accession numbers MW240686–MW240711. The sequence alignment, and input and output files from the phylogenetic and species delimitation analyses, have deposited at Zenodo: https://doi.org/10.5281/zenodo.4104677.
References
- Dobzhansky, T. A Critique of the Species Concept in Biology. Philos. Sci. 1935, 2, 344–355. [Google Scholar] [CrossRef]
- Mishler, B.D.; Brandon, R.N. Individuality, Pluralism, and the Phylogenetic Species Concept. Biol. Philos. 1987, 2, 397–414. [Google Scholar] [CrossRef]
- Hausdorf, B. Progress toward a General Species Concept. Evolution 2011, 65, 923–931. [Google Scholar] [CrossRef] [PubMed]
- de Queiroz, K. The General Lineage Concept of Species, Species Criteria, and the Process of Speciation: A Conceptual Unification and Terminological Recommendations. In Endless Forms: Species and Speciation; Howard, D.J., Berlocher, S.H., Eds.; Oxford University Press: Oxford, UK, 1998; pp. 57–75. [Google Scholar]
- de Queiroz, K. Species Concepts and Species Delimitation. Syst. Biol. 2007, 56, 879–886. [Google Scholar] [CrossRef] [PubMed]
- Agapow, P.M.; Bininda-Emonds, O.R.P.; Crandall, K.A.; Gittleman, J.L.; Mace, G.M.; Marshall, J.C.; Purvis, A. The Impact of Species Concept on Biodiversity Studies. Q. Rev. Biol. 2004, 79, 161–179. [Google Scholar] [CrossRef]
- Groves, C.P.; Cotterill, F.P.D.; Gippoliti, S.; Robovský, J.; Roos, C.; Taylor, P.J.; Zinner, D. Species Definitions and Conservation: A Review and Case Studies from African Mammals. Conserv. Genet. 2017, 18, 1247–1256. [Google Scholar] [CrossRef]
- Barrowclough, G.F.; Cracraft, J.; Klicka, J.; Zink, R.M. How Many Kinds of Birds Are There and Why Does It Matter? PLoS ONE 2016, 11, e0166307. [Google Scholar] [CrossRef]
- Isaac, N.J.; Mallet, J.; Mace, G.M. Taxonomic Inflation: Its Influence on Macroecology and Conservation. Trends Ecol. Evol. 2004, 19, 464–469. [Google Scholar] [CrossRef]
- Haffer, J. The History of Species Concepts and Species Limits in Ornithology. Bull. Br. Ornithol. Club 1992, 112, 107–158. [Google Scholar]
- Sangster, G. Increasing Numbers of Bird Species Result from Taxonomic Progress, Not Taxonomic Inflation. Proc. R. Soc. Biol. Sci. Ser. B 2009, 276, 3185–3191. [Google Scholar] [CrossRef] [PubMed]
- Poelstra, J.W.; Vijay, N.; Bossu, C.M.; Lantz, H.; Ryll, B.; Muller, I.; Baglione, V.; Unneberg, P.; Wikelski, M.; Grabherr, M.G.; et al. The Genomic Landscape Underlying Phenotypic Integrity in the Face of Gene Flow in Crows. Science 2014, 344, 1410–1414. [Google Scholar] [CrossRef] [PubMed]
- Toews, D.P.L.; Taylor, S.A.; Vallender, R.; Brelsford, A.; Butcher, B.G.; Messer, P.W.; Lovette, I.J. Plumage Genes and Little Else Distinguish the Genomes of Hybridizing Warblers. Curr. Biol. 2016, 26, 2313–2318. [Google Scholar] [CrossRef] [PubMed]
- Irwin, D.E.; Alström, P.; Olsson, U.; Benowitz-Fredericks, Z.M. Cryptic Species in the Genus Phylloscopus (Old World Leaf Warblers). Ibis 2008, 143, 233–247. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Penton, E.H.; Burns, J.M.; Janzen, D.H.; Hallwachs, W. Ten Species in One: DNA Barcoding Reveals Cryptic Species in the Neotropical Skipper Butterfly Astraptes fulgerator. Proc. Natl. Acad. Sci. USA 2004, 101, 14812–14817. [Google Scholar] [CrossRef] [PubMed]
- Bickford, D.; Lohman, D.J.; Sodhi, N.S.; Ng, P.K.L.; Meier, R.; Winker, K.; Ingram, K.K.; Das, I. Cryptic Species as a Window on Diversity and Conservation. Trends Ecol. Evol. 2007, 22, 148–155. [Google Scholar] [CrossRef] [PubMed]
- Jarvis, E.D.; Mirarab, S.; Aberer, A.J.; Li, B.; Houde, P.; Li, C.; Ho, S.Y.W.; Faircloth, B.C.; Nabholz, B.; Howard, J.T.; et al. Whole-Genome Analyses Resolve Early Branches in the Tree of Life of Modern Birds. Science 2014, 346, 1320–1331. [Google Scholar] [CrossRef] [PubMed]
- Van Tuinen, M.; Butvill, D.B.; Kirsch, J.A.W.; Hedges, S.B. Convergence and Divergence in the Evolution of Aquatic Birds. Proc. R. Soc. Biol. Sci. Ser. B 2001, 268, 1345–1350. [Google Scholar] [CrossRef]
- Alström, P.; Hooper, D.M.; Liu, Y.; Olsson, U.; Mohan, D.; Gelang, M.; Le Manh, H.; Zhao, J.; Lei, F.; Price, T.D. Discovery of a Relict Lineage and Monotypic Family of Passerine Birds. Biol. Lett. 2014, 10, 20131067. [Google Scholar] [CrossRef]
- Garcia-R, J.C.; Lemmon, E.M.; Lemmon, A.R.; French, N. Phylogenomic Reconstruction Sheds Light on New Relationships and Timescale of Rails (Aves: Rallidae) Evolution. Diversity 2020, 12, 70. [Google Scholar] [CrossRef]
- Alström, P.; Jønsson, K.A.; Fjeldså, J.; Ödeen, A.; Ericson, P.G.P.; Irestedt, M. Dramatic Niche Shifts and Morphological Change in Two Insular Bird Species. R. Soc. Open Sci. 2015, 2. [Google Scholar] [CrossRef]
- Alström, P.; Olsson, U.; Lei, F. A Review of the Recent Advances in the Systematics of the Avian Superfamily Sylvioidea. Chin. Birds 2013, 4, 99–131. [Google Scholar] [CrossRef]
- IOC World Bird List (v. 10.2). 2020. Available online: https://www.worldbirdnames.org/new/ioc-lists/crossref (accessed on 25 July 2020).
- de Juana, E.; Suárez, F.; Ryan, P.G.; Alström, P.; Donald, P.F. Family Alaudidae (Larks). In Handbook of the Birds of the World; del Hoyo, J., Elliott, A., Christie, D.A., Eds.; Lynx Edicions: Barcelona, Spain, 2004; Volume 9, pp. 496–601. [Google Scholar]
- Alström, P.; Barnes, K.N.; Olsson, U.; Barker, F.K.; Bloomer, P.; Khan, A.A.; Qureshi, M.A.; Guillaumet, A.; Crochet, P.A.; Ryan, P.G. Multilocus Phylogeny of the Avian Family Alaudidae (Larks) Reveals Complex Morphological Evolution, Non-Monophyletic Genera and Hidden Species Diversity. Mol. Phylogenet. Evol. 2013, 69, 1043–1056. [Google Scholar] [CrossRef] [PubMed]
- Ghorbani, F.; Aliabadian, M.; Zhang, R.; Irestedt, M.; Hao, Y.; Sundev, G.; Lei, F.; Ma, M.; Olsson, U.; Alström, P. Densely Sampled Phylogenetic Analyses of the Lesser Short-toed Lark (Alaudala rufescens)—Sand Lark (A. raytal) species Complex (Aves, Passeriformes) Reveal Cryptic Diversity. Zool. Scr. 2020, 49, 427–439. [Google Scholar] [CrossRef]
- Stervander, M.; Alström, P.; Olsson, U.; Ottosson, U.; Hansson, B.; Bensch, S. Multiple Instances of Paraphyletic Species and Cryptic Taxa Revealed by Mitochondrial and Nuclear RAD Data for Calandrella Larks (Aves: Alaudidae). Mol. Phylogenet. Evol. 2016, 102, 233–245. [Google Scholar] [CrossRef] [PubMed]
- Alström, P.; Sundev, G. Mongolian Short-toed Lark Calandrella dukhunensis, an Overlooked East Asian Species. J. Ornithol. 2020. [Google Scholar] [CrossRef]
- Donald, P.F.; Alström, P.; Engelbrecht, D. Possible Mechanisms of Substrate Colour-Matching in Larks (Alaudidae) and Their Taxonomic Implications. Ibis 2017, 159, 699–702. [Google Scholar] [CrossRef]
- Guillaumet, A.; Crochet, P.A.; Pons, J.M. Climate-Driven Diversification in Two Widespread Galerida Larks. BMC Evol. Biol. 2008, 8, 32. [Google Scholar] [CrossRef]
- Guillaumet, A.; Crochet, P.A.; Godelle, B. Phenotypic Variation in Galerida Larks in Morocco: The Role of History and Natural Selection. Mol. Ecol. 2005, 14, 3809–3821. [Google Scholar] [CrossRef]
- Guillaumet, A.; Pons, J.-M.; Godelle, B.; Crochet, P.-A. History of the Crested Lark in the Mediterranean Region as Revealed by MtDNA Sequences and Morphology. Mol. Phylogenet. Evol. 2006, 39, 645–656. [Google Scholar] [CrossRef]
- Ryan, P.G.; Bloomer, P. The Long-billed Lark Complex: A Species Mosaic in Southwestern Africa. Auk 1999, 116, 194–208. [Google Scholar] [CrossRef]
- Ryan, P.G.; Hood, I.; Bloomer, P.; Komen, J.; Crowe, T.M. Barlow’s Lark: A New Species in the Karoo Lark Certhilauda albescens Complex of Southwest Africa. Ibis 1998, 140, 605–619. [Google Scholar] [CrossRef]
- Svensson, L. Identification Guide to European Passerines, 4th ed.; British Trust for Ornithology: Stockholm, Sweden, 1992. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Ghorbani, F.; Aliabadian, M.; Olsson, U.; Donald, P.F.; Khan, A.A.; Alström, P. Mitochondrial Phylogeography of the Genus Eremophila Confirms Underestimated Species Diversity in the Palearctic. J. Ornithol. 2020, 161, 297–312. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An Integrated and Extendable Desktop Software Platform for the Organization and Analysis of Sequence Data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. Jmodeltest 2: More Models, New Heuristics and Parallel Computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef]
- Hasegawa, M.; Kishino, H.; Yano, T.A. Dating of the Human Ape Splitting by a Molecular Clock of Mitochondrial-DNA. J. Mol. Evol. 1985, 22, 160–174. [Google Scholar] [CrossRef]
- Schwarz, G. Estimating the Dimension of a Model. Ann. Stat. 1978, 6, 461–464. [Google Scholar] [CrossRef]
- Bouckaert, R.; Heled, J.; Kuhnert, D.; Vaughan, T.; Wu, C.H.; Xie, D.; Suchard, M.A.; Rambaut, A.; Drummond, A.J. Beast 2: A Software Platform for Bayesian Evolutionary Analysis. PLoS Comput. Biol. 2014, 10, e1003537. [Google Scholar] [CrossRef]
- Weir, J.T.; Schluter, D. Calibrating the Avian Molecular Clock. Mol. Ecol. 2008, 17, 2321–2328. [Google Scholar] [CrossRef]
- Alström, P.; Ericson, P.G.; Olsson, U.; Sundberg, P. Phylogeny and Classification of the Avian Superfamily Sylvioidea. Mol. Phylogenet. Evol. 2006, 38, 381–397. [Google Scholar] [CrossRef]
- Moyle, R.G.; Oliveros, C.H.; Andersen, M.J.; Hosner, P.A.; Benz, B.W.; Manthey, J.D.; Travers, S.L.; Brown, R.M.; Faircloth, B.C. Tectonic Collision and Uplift of Wallacea Triggered the Global Songbird Radiation. Nat. Commun. 2016, 7, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Ericson, P.G.P.; Johansson, U.S. Phylogeny of Passerida (Aves: Passeriformes) Based on Nuclear and Mitochondrial Sequence Data. Mol. Phylogenet. Evol. 2003, 29, 126–138. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G.; Suchard, M.A. Posterior Summarization in Bayesian Phylogenetics Using Tracer 1.7. Syst. Biol. 2018, 67, 901–904. [Google Scholar] [CrossRef] [PubMed]
- Heibl, C. Phyloch: Interfaces and Graphic Tools for Phylogenetic Data in R. Available online: http://www.christophheibl.de/Rpackages.html (accessed on 30 April 2013).
- Revell, L.J. Phytools: An R Package for Phylogenetic Comparative Biology (and Other Things). Methods Ecol. Evol. 2012, 3, 217–223. [Google Scholar] [CrossRef]
- Paradis, E. Analysis of Phylogenetics and Evolution with R. In Use R! 2nd ed.; Springer: New York, NY, USA, 2012; p. 386. [Google Scholar] [CrossRef]
- Paradis, E.; Claude, J.; Strimmer, K. APE: Analyses of Phylogenetics and Evolution in R Language. Bioinformatics 2004, 20, 289–290. [Google Scholar] [CrossRef]
- Heled, J.; Bouckaert, R.R. Looking for Trees in the Forest: Summary Tree from Posterior Samples. BMC Evol. Biol. 2013, 13, 221. [Google Scholar] [CrossRef]
- Kapli, P.; Lutteropp, S.; Zhang, J.; Kobert, K.; Pavlidis, P.; Stamatakis, A.; Flouri, T. Multi-Rate Poisson Tree Processes for Single-Locus Species Delimitation under Maximum Likelihood and Markov Chain Monte Carlo. Bioinformatics 2017. [Google Scholar] [CrossRef]
- Sharland, R.E. Two Interesting Plateau Birds: The Red-capped Lark Calandrella cinerea and the Three-banded Plover Afroxyechus tricollaris. Bull. Niger. Ornithol. Soc. 1964, 1, 4. [Google Scholar]
- Sharland, R.E. Bird-Ringing in West Africa, 1959. Niger. Field 1960, 25, 125–127. [Google Scholar] [CrossRef]
- International Commission on Zoological Nomenclature. Amendment of Articles 8, 9, 10, 21 and 78 of the International Code of Zoological Nomenclature to Expand and Refine Methods of Publication. ZooKeys 2012. [Google Scholar] [CrossRef]
- Hulme, M.F.; Cresswell, W. Density and Behaviour of Whinchats Saxicola rubetra on African Farmland Suggest That Winter Habitat Conditions Do Not Limit European Breeding Populations. Ibis 2012, 154, 680–692. [Google Scholar] [CrossRef]
- Donald, P.F.; Alström, P. Larks of the World. Bloomsbury Publishing: London, UK, forthcoming.
- Payne, R.B. A New Species of Firefinch Lagonosticta from Northern Nigeria and Its Association with the Jos Plateau Indigobird Vidua maryae. Ibis 2008, 140, 369–381. [Google Scholar] [CrossRef]
- Ryan, P.G. Red-capped Lark (Calandrella cinerea). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- BirdLife International. Species Factsheet: Mirafra africana. Available online: http://datazone.birdlife.org/species/factsheet/rufous-naped-lark-mirafra-africana (accessed on 4 May 2020).
- Ryan, P.G. Red-winged Lark (Mirafra hypermetra). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- Fuchs, J.; Alström, P.; Yosef, R.; Olsson, U. Miocene Diversification of an Open-Habitat Predatorial Passerine Radiation, the Shrikes (Aves: Passeriformes: Laniidae). Zool. Scr. 2019, 48, 571–588. [Google Scholar] [CrossRef]
- Fuchs, J.; Douno, M.; Bowie, R.C.K.; Fjeldså, J. Taxonomic Revision of the Square-tailed Drongo Species Complex (Passeriformes: Dicruridae) with Description of a New Species from Western Africa. Zootaxa 2018, 4438, 105–127. [Google Scholar] [CrossRef]
- Olsson, U.; Yosef, R.; Alström, P. Assessment of Species Limits in African ‘Brown Buntings’ (Emberiza, Passeriformes) Based on Mitochondrial and Nuclear Sequence Data. Ibis 2013, 155, 534–543. [Google Scholar] [CrossRef]
- Bertola, L.D.; Jongbloed, H.; van der Gaag, K.J.; de Knijff, P.; Yamaguchi, N.; Hooghiemstra, H.; Bauer, H.; Henschel, P.; White, P.A.; Driscoll, C.A.; et al. Phylogeographic Patterns in Africa and High Resolution Delineation of Genetic Clades in the Lion (Panthera leo). Sci. Rep. 2016, 6, 30807. [Google Scholar] [CrossRef]
- Lorenzen, E.D.; Heller, R.; Siegismund, H.R. Comparative Phylogeography of African Savannah Ungulates. Mol. Ecol. 2012, 21, 3656–3670. [Google Scholar] [CrossRef]
- Flagstad, O.; Syvertsen, P.O.; Stenseth, N.C.; Jakobsen, K.S. Environmental Change and Rates of Evolution: The Phylogeographic Pattern within the Hartebeest Complex as Related to Climatic Variation. Proc. R. Soc. Biol. Sci. Ser. B 2001, 268, 667–677. [Google Scholar] [CrossRef]
- Moodley, Y.; Bruford, M.W. Molecular Biogeography: Towards an Integrated Framework for Conserving Pan-African Biodiversity. PLoS ONE 2007, 2, e454. [Google Scholar] [CrossRef]
- Muwanika, V.B.; Nyakaana, S.; Siegismund, H.R.; Arctander, P. Phylogeography and Population Structure of the Common Warthog (Phacochoerus africanus) Inferred from Variation in Mitochondrial DNA Sequences and Microsatellite Loci. Heredity 2003, 91, 361–372. [Google Scholar] [CrossRef]
- Rohland, N.; Pollack, J.L.; Nagel, D.; Beauval, C.; Airvaux, J.; Paabo, S.; Hofreiter, M. The Population History of Extant and Extinct Hyenas. Mol. Biol. Evol. 2005, 22, 2435–2443. [Google Scholar] [CrossRef] [PubMed]
- Droissart, V.; Dauby, G.; Hardy, O.J.; Deblauwe, V.; Harris, D.J.; Janssens, S.; Mackinder, B.; Blach-Overgaard, A.; Sonke, B.; Sosef, M.S.M.; et al. Beyond Trees: Biogeographical Regionalization of Tropical Africa. J. Biogeogr. 2018, 45, 1153–1167. [Google Scholar] [CrossRef]
- Ryan, P.G. Flappet Lark (Mirafra rufocinnamomea). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- Ryan, P.G. White-tailed Lark (Mirafra albicauda). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- BirdLife International. Species Factsheet: Mirafra javanica. Available online: http://datazone.birdlife.org/species/factsheet/rufous-naped-lark-mirafra-javanica (accessed on 4 May 2020).
- Tieleman, B.I.; Williams, J.B.; Bloomer, P. Adaptation of Metabolism and Evaporative Water Loss Along an Aridity Gradient. Proc. R. Soc. B Biol. Sci. 2003, 270, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Ryan, P.G.; Christie, D. Kordofan Lark (Mirafra cordofanica). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- Ryan, P.G. Rusty Lark (Mirafra rufa). Birds of the World. In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- Ryan, P.G. Chestnut-backed Sparrow-Lark (Eremopterix leucotis). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- de Juana, E.; Suárez, F. Greater Hoopoe-Lark (Alaemon alaudipes). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- de Juana, E.; Suárez, F. Bar-Tailed Lark (Ammomanes cinctura). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- de Juana, E.; Suárez, F. Desert Lark (Ammomanes deserti). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- de Juana, E.; Suárez, F. Temminck’s Lark (Eremophila bilopha). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- de Juana, E.; Suárez, F. Thick-Billed Lark (Ramphocoris clotbey). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- Ryan, P.G. Black-Crowned Sparrow-Lark (Eremopterix nigriceps). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- Valente, L.; Illera, J.C.; Havenstein, K.; Pallien, T.; Etienne, R.S.; Tiedemann, R. Equilibrium Bird Species Diversity in Atlantic Islands. Curr. Biol. 2017, 27, 1660–1666. [Google Scholar] [CrossRef]
- Fuchs, J.; Pons, J.-M.; Bowie, R.C.K. Biogeography and Diversification Dynamics of the African Woodpeckers. Mol. Phylogenet. Evol. 2017, 108, 88–100. [Google Scholar] [CrossRef]
- Spottiswoode, C.N.; Olsson, U.; Mills, M.S.L.; Cohen, C.; Francis, J.E.; Toye, N.; Hoddinott, D.; Dagne, A.; Wood, C.; Donald, P.F.; et al. Rediscovery of a Long-Lost Lark Reveals the Conspecificity of Endangered Heteromirafra Populations in the Horn of Africa. J. Ornithol. 2013, 154, 813–825. [Google Scholar] [CrossRef]
- Larrasoaña, J.C.; Roberts, A.P.; Rohling, E.J. Dynamics of Green Sahara Periods and Their Role in Hominin Evolution. PLoS ONE 2013, 8, e76514. [Google Scholar] [CrossRef]
- Larrasoaña, J.C.; Roberts, A.P.; Rohling, E.J.; Winklhofer, M.; Wehausen, R. Three Million Years of Monsoon Variability over the Northern Sahara. Clim. Dyn. 2003, 21, 689–698. [Google Scholar] [CrossRef]
- deMenocal, P.B. African Climate Change and Faunal Evolution during the Pliocene–Pleistocene. Earth Planet. Sci. Lett. 2004, 220, 3–24. [Google Scholar] [CrossRef]
- Trauth, M.H.; Larrasoaña, J.C.; Mudelsee, M. Trends, Rhythms and Events in Plio–Pleistocene African Climate. Quat. Sci. Rev. 2009, 28, 399–411. [Google Scholar] [CrossRef]
- Linder, H.P. East African Cenozoic Vegetation History. Evol. Anthr. 2017, 26, 300–312. [Google Scholar] [CrossRef] [PubMed]
- Potts, R.; Faith, J.T. Alternating High and Low Climate Variability: The Context of Natural Selection and Speciation in Plio–Pleistocene Hominin Evolution. J. Hum. Evol. 2015, 87, 5–20. [Google Scholar] [CrossRef] [PubMed]
- Päckert, M.; Martens, J.; Hering, J.; Kvist, L.; Illera, J.C. Return Flight to the Canary Island—The Key Role of Peripheral Populations of Afrocanarian Blue Tits (Aves: Cyanistes teneriffae) in Multi-Gene Reconstructions of Colonization Pathways. Mol. Phylogenet. Evol. 2013, 67, 458–467. [Google Scholar] [CrossRef] [PubMed]
- Stervander, M.; Illera, J.C.; Kvist, L.; Barbosa, P.; Keehnen, N.P.; Pruisscher, P.; Bensch, S.; Hansson, B. Disentangling the Complex Evolutionary History of the Western Palearctic Blue Tits (Cyanistes spp.)—Phylogenomic Analyses Suggest Radiation by Multiple Colonization Events and Subsequent Isolation. Mol. Ecol. 2015, 24, 2477–2494. [Google Scholar] [CrossRef]
- Irestedt, M.; Gelang, M.; Sangster, G.; Olsson, U.; Ericson, P.G.P.; Alström, P. Neumann’s Warbler Hemitesia neumanni (Sylvioidea): The Sole African Member of a Palaeotropic Miocene Avifauna. Ibis 2011, 153, 78–86. [Google Scholar] [CrossRef]
- Rheindt, F.E.; Edwards, S.V. Genetic Introgression: An Integral but Neglected Component of Speciation in Birds. Auk 2011, 128, 620–632. [Google Scholar] [CrossRef]
- Toews, D.P.L.; Brelsford, A. The Biogeography of Mitochondrial and Nuclear Discordance in Animals. Mol. Ecol. 2012, 21, 3907–3930. [Google Scholar] [CrossRef]
- Ottenburghs, J. Ghost Introgression: Spooky Gene Flow in the Distant Past. Bioessays 2020. [Google Scholar] [CrossRef]
- Zhang, D.; Tang, L.; Cheng, Y.; Hao, Y.; Xiong, Y.; Song, G.; Qu, Y.; Rheindt, F.E.; Alström, P.; Jia, C.; et al. “Ghost Introgression” as a Cause of Deep Mitochondrial Divergence in a Bird Species Complex. Mol. Biol. Evol. 2019, 36, 2375–2386. [Google Scholar] [CrossRef]
- Hogner, S.; Laskemoen, T.; Lifjeld, J.T.; Porkert, J.; Kleven, O.; Albayrak, T.; Kabasakal, B.; Johnsen, A. Deep Sympatric Mitochondrial Divergence without Reproductive Isolation in the Common Redstart Phoenicurus phoenicurus. Ecol. Evol 2012, 2, 2974–2988. [Google Scholar] [CrossRef]
- Alström, P. Hume’s Lark (Calandrella acutirostris). In Birds of the World; del Hoyo, J., Elliott, A., Sargatal, J., Christie, D., de Juana, E., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- Zink, R.M.; Barrowclough, G.F. Mitochondrial DNA under Siege in Avian Phylogeography. Mol. Ecol. 2008, 17, 2107–2121. [Google Scholar] [CrossRef] [PubMed]
- Funk, D.J.; Omland, K.E. Species-Level Paraphyly and Polyphyly: Frequency, Causes, and Consequences, with Insights from Animal Mitochondrial DNA. Annu. Rev. Ecol. Evol. Syst. 2003, 34, 397–423. [Google Scholar] [CrossRef]
- Harris, R.B.; Alström, P.; Ödeen, A.; Leaché, A.D. Discordance between Genomic Divergence and Phenotypic Variation in a Rapidly Evolving Avian Genus (Motacilla). Mol. Phylogenet. Evol. 2018, 120, 183–195. [Google Scholar] [CrossRef] [PubMed]
- Dayrat, B. Towards Integrative Taxonomy. Biol. J. Linn. Soc. 2005, 85, 407–415. [Google Scholar] [CrossRef]
- Padial, J.M.; Miralles, A.; De la Riva, I.; Vences, M. The Integrative Future of Taxonomy. Front. Zool. 2010, 7. [Google Scholar] [CrossRef]
- Sangster, G. Integrative Taxonomy of Birds: The Nature and Delimitation of Species. In Bird Species: How They Arise, Modify and Vanish; Tietze, D.T., Ed.; Springer Open: Cham, Switzerland, 2018; pp. 9–37. [Google Scholar]
- Alström, P.; Rasmussen, P.C.; Olsson, U.; Sundberg, P. Species Delimitation Based on Multiple Criteria: The Spotted Bush Warbler Bradypterus thoracicus Complex (Aves: Megaluridae). Zool. J. Linn. Soc. 2008, 154, 291–307. [Google Scholar] [CrossRef]
- Alström, P.; Rasmussen, P.C.; Sangster, G.; Dalvi, S.; Round, P.D.; Zhang, R.; Yao, C.T.; Irestedt, M.; Le Manh, H.; Lei, F.; et al. Multiple Species within the Striated Prinia Prinia crinigera–Brown Prinia P. polychroa Complex Revealed through an Integrative Taxonomic Approach. Ibis 2019, 162, 936–967. [Google Scholar] [CrossRef]
- Alström, P.; Rasmussen, P.C.; Zhao, C.; Xu, J.; Dalvi, S.; Cai, T.; Guan, Y.; Zhang, R.; Kalyakin, M.V.; Lei, F.; et al. Integrative Taxonomy of the Plain-backed Thrush (Zoothera mollissima) Complex (Aves, Turdidae) Reveals Cryptic Species, Including a New Species. Avian Res. 2016, 7. [Google Scholar] [CrossRef]
- Alström, P.; Xia, C.; Rasmussen, P.C.; Olsson, U.; Dai, B.; Zhao, J.; Leader, P.J.; Carey, G.J.; Dong, L.; Cai, T.; et al. Integrative Taxonomy of the Russet Bush Warbler Locustella mandelli Complex Reveals a New Species from Central China. Avian Res. 2015, 6. [Google Scholar] [CrossRef]
- Liu, Y.; Chen, G.; Huang, Q.; Jia, C.; Carey, G.; Leader, P.; Li, Y.; Zou, F.; Yang, X.; Olsson, U.; et al. Species Delimitation of the White-tailed Rubythroat Calliope pectoralis Complex (Aves, Muscicapidae) Using an Integrative Taxonomic Approach. J. Avian Biol. 2016, 47, 899–910. [Google Scholar] [CrossRef]
- Sangster, G.; Rodríguez-Godoy, F.; Roselaar, C.S.; Robb, M.S.; Luksenburg, J.A. Integrative Taxonomy Reveals Europe’s Rarest Songbird Species, the Gran Canaria Blue Chaffinch Fringilla polatzeki. J. Avian Biol. 2016, 47, 159–166. [Google Scholar] [CrossRef]
- Gjershaug, J.O.; Diserud, O.H.; Kleven, O.; Rasmussen, P.C.; Espmark, Y. Integrative Taxonomy of the Changeable Hawk-Eagle Nisaetus cirrhatus Complex (Accipitriformes: Accipitridae) in India. Zootaxa 2020, 4789, 554–574. [Google Scholar] [CrossRef] [PubMed]
- Venkatraman, M.X.; Deraad, D.A.; Tsai, W.L.E.; Zarza, E.; Zellmer, A.J.; Maley, J.M.; McCormack, J.E. Cloudy with a Chance of Speciation: Integrative Taxonomy Reveals Extraordinary Divergence within a Mesoamerican Cloud Forest Bird. Biol. J. Linn. Soc. 2019, 126, 1–15. [Google Scholar] [CrossRef]
- Younger, J.L.; Strozier, L.; Maddox, J.D.; Nyári, Á.S.; Bonfitto, M.T.; Raherilalao, M.J.; Goodman, S.M.; Reddy, S. Hidden Diversity of Forest Birds in Madagascar Revealed Using Integrative Taxonomy. Mol. Phylogenet. Evol. 2018, 124, 16–26. [Google Scholar] [CrossRef]
- Alström, P.; van Linschooten, J.; Donald, P.F.; Sundev, G.; Mohammadi, Z.; Ghorbani, F.; Shafaeipour, A.; van den Berg, A.; Robb, M.; Aliabadian, M.; et al. Multiple Species Delimitation Approaches Applied to the Avian Lark Genus Alaudala. Mol. Phylogenet. Evol. 2020. submitted. [Google Scholar] [CrossRef]
- Marki, P.Z.; Fjeldså, J.; Irestedt, M.; Jønsson, K.A. Molecular Phylogenetics and Species Limits in a Cryptically Coloured Radiation of Australo-Papuan Passerine Birds (Pachycephalidae: Colluricincla). Mol. Phylogenet. Evol. 2018, 124, 100–105. [Google Scholar] [CrossRef]
- Drovetski, S.V.; Raković, M.; Semenov, G.; Fadeev, I.V.; Red’kin, Y.A. Limited Phylogeographic Signal in Sex-Linked and Autosomal Loci Despite Geographically, Ecologically, and Phenotypically Concordant Structure of MtDNA Variation in the Holarctic Avian Genus Eremophila. PLoS ONE 2014, 9, e87570. [Google Scholar] [CrossRef]
- del Hoyo, J.; Collar, N.J.; Christie, D.A.; Elliott, A.; Fishpool, L.D.C.; Boesman, P.; Kirwan, G.M. Passerines. In HBW and Birdlife International Illustrated Checklist of the Birds of the World; Lynx Edicions and BirdLife International: Barcelona, Spain; Cambridge, UK, 2016; Volume 2. [Google Scholar]
- Ryan, P.G. Spike-heeled Lark (Chersomanes albofasciata). In Birds of the World; Billerman, S.M., Keeney, B.K., Rodewald, P.G., Schulenberg, T.S., Eds.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- Barnes, K.N. The Phylogenetics and Evolution of Africa’s Larks (Alaudidae). Ph.D. Thesis, University of Cape Town, Cape Town, South Africa, 2007. [Google Scholar]
- Dierickx, E.G.; Shultz, A.J.; de Brooke, M.L.; Alström, P.; Liu, Y. Phylogeography of the Raso Lark, a single-island endemic, and its widespread, continental relatives. Ibis 2020. under review. [Google Scholar]
- Campbell, R.W.; Van Damme, L.M.; Johnson, S.R.; Donald, P.F.; Garcia, E. Eurasian Skylark (Alauda arvensis). In Birds of the World; Billerman, S.M., Ed.; Cornell Lab of Ornithology: Ithaca, NY, USA, 2020. [Google Scholar] [CrossRef]
- Qian, C.; Wang, Y.; Guo, Z.; Yang, J.; Kan, X. Complete Mitochondrial Genome of Skylark, Alauda arvensis (Aves: Passeriformes): The First Representative of the Family Alaudidae with Two Extensive Heteroplasmic Control Regions. Mitochondrial DNA 2013, 24, 246–248. [Google Scholar] [CrossRef]
- Zink, R.M.; Pavlova, A.; Drovetski, S.; Rohwer, S. Mitochondrial Phylogeographies of Five Widespread Eurasian Bird Species. J. Ornithol. 2008, 149, 399–413. [Google Scholar] [CrossRef]
- Li, Y.; Chen, G.; Liu, S.; Dierickx, E.G.; Wang, P.; Birks, S.; Yao, C.; Shiraki, S.; Lei, F.; Brooke, M.d.L.; et al. Complex divergence patterns across the speciation continuum of Alauda skylarks reject a ‘ring species’ origin. Manuscript in preparation.
- van den Berg, A.B. The Sound Approach. In Morocco: Sharing the Birds—A Sound Approach Guide to Birds of the Maghreb; The Sound Approach: Poole, UK, 2020. [Google Scholar]
- BirdLife International. Species Factsheet: Calandrella cinerea. Available online: http://datazone.birdlife.org/species/factsheet/red-capped-lark-calandrella-cinerea (accessed on 4 May 2020).
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).