In or Out of the Checklist? DNA Barcoding and Distribution Modelling Unveil a New Species of Crocidura Shrew for Italy
Abstract
1. Introduction
2. Materials and Methods
2.1. DNA Barcoding
2.2. Species Distribution Modelling
3. Results
3.1. DNA Barcoding Identification
3.2. Species Distribution Modelling
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Castiglia, R.; Annesi, F.; Aloise, G.; Amori, G. Systematics of the Microtus savii complex (Rodentia, Cricetidae) via mitochondrial DNA analyses: Paraphyly and pattern of sex chromosome evolution. Mol. Phylogenet. Evol. 2008, 46, 1157–1164. [Google Scholar] [CrossRef] [PubMed]
- Abiadh, A.; Chetoui, M.; Lamine-Cheniti, T.; Capanna, E.; Colangelo, P. Molecular phylogenetics of the genus Gerbillus (Rodentia, Gerbillinae): Implications for systematics, taxonomy and chromosomal evolution. Mol. Phylogenet. Evol. 2010, 56, 513–518. [Google Scholar] [CrossRef] [PubMed]
- Ancillotto, L.; Mori, E.; Sozio, G.; Solano, E.; Bertolino, S.; Russo, D. A novel approach to field identification of cryptic Apodemus wood mice: Calls differ more than morphology. Mamm. Rev. 2016, 47, 6–10. [Google Scholar] [CrossRef]
- Halley, D.J.; Saveljev, A.P.; Rosell, F. Population and distribution of beavers Castor fiber and Castor canadensis in Eurasia. Mamm. Rev. 2020. [Google Scholar] [CrossRef]
- Castiglia, R.; Aloise, G.; Amori, G.; Annesi, F.; Bertolino, S.; Capizzi, D.; Mori, E.; Colangelo, P. The Italian peninsula hosts a divergent mtDNA lineage of the water vole, Arvicola amphibius s.l., including fossorial and aquatic ecotypes. Hystrix 2016, 27, 99–103. [Google Scholar]
- Ancillotto, L.; Mori, E.; Bosso, L.; Agnelli, P.; Russo, D. The Balkan long-eared bat (Plecotus kolombatovici) occurs in Italy—First confirmed record and potential distribution. Mamm. Biol. 2019, 96, 61–67. [Google Scholar] [CrossRef]
- Mori, E.; Nerva, L.; Lovari, S. Reclassification of the serows and the gorals: The end of a neverending story? Mamm. Rev. 2019, 49, 256–262. [Google Scholar] [CrossRef]
- Li, G.; Sun, N.; Swa, K.; Zhang, M.; Lwin, Y.H.; Quan, R.C. Phylogenetic reassessment of gorals with new evidence from northern Myanmar reveals five distinct species. Mamm. Rev. 2020. [Google Scholar] [CrossRef]
- Galimberti, A.; Spada, M.; Russo, D.; Mucedda, M.; Agnelli, P.; Crottini, A.; Ferri, E.; Martinoli, A.; Casiraghi, M. Integrated operational taxonomic units (IOTUs) in echolocating bats: A bridge between molecular and traditional taxonomy. PLoS ONE 2012, 7, e40122. [Google Scholar] [CrossRef]
- Galimberti, A.; Sandionigi, A.; Bruno, A.; Bellati, A.; Casiraghi, M. DNA barcoding in mammals: What’s new and where next? Hystrix 2015, 26, 13–24. [Google Scholar]
- Sales, N.G.; Kaizer, M.D.C.; Coscia, I.; Perkins, J.C.; Highlands, A.; Boubli, J.P.; Magnusson, W.E.; Da Silva, M.N.F.; Benvenuto, C.; Mcdevitt, A.D. Assessing the potential of environmental DNA metabarcoding for monitoring Neotropical mammals: A case study in the Amazon and Atlantic Forest, Brazil. Mamm. Rev. 2020. [Google Scholar] [CrossRef]
- Loy, A.; Aloise, G.; Ancillotto, L.; Angelici, F.M.; Bertolino, S.; Capizzi, D.; Castiglia, R.; Colangelo, P.; Contoli, L.; Cozzi, B.; et al. Mammals of Italy: An annotated checklist. Hystrix 2019, 30, 87–106. [Google Scholar]
- Ancillotto, L.; Bosso, L.; Smeraldo, S.; Mori, E.; Mazza, G.; Herkt, M.; Galimberti, A.; Ramazzotti, F.; Russo, D. An African bat in Europe, Plecotus gaisleri: Biogeographic and ecological insights from molecular taxonomy and species distribution models. Ecol. Evol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Wilson, D.E.; Mittermeier, R.A. Handbook of the Mammals of the World (Vol. 8), Insectivores, Sloths and Colugos; Lynx Editions: Barcelona, Spain, 2018. [Google Scholar]
- Miller, G.S. Catalogue of the Mammals of Western Europe; Thrustees of British Museum Editions: London, UK, 1912. [Google Scholar]
- Meylan, A.; Hausser, J. Position cytotaxonomique de quelques musaraignes du genre Crocidura au Tessin (Mammalia, Insectivora). Rev. Suisse Zool. 1974, 81, 701–710. [Google Scholar] [CrossRef]
- Rzebik-Kowalska, B. Studies on the genus Crocidura (Insectivora, Mammalia) in Algeria. Acta Zool. Cracovia 1988, 31, 167–192. [Google Scholar]
- Cosson, J.F.; Hutterer, R.; Libois, R.; Sarà, M.; Taberlet, P.; Vogel, P. Phylogeographic footprints of the Strait of Gibraltar and Quaternary climatic fluctuations in the Western Mediterranean: A case study with the greater white-toothed shrew Crocidura russula (Mammalia: Soricidae). Mol. Ecol. 2005, 14, 1151–1162. [Google Scholar] [CrossRef]
- Nicolas, V.; Hamani, A.; Amrouche, L.; Bensidhoum, M.; Boukhemza, M.; Doumandji, S.; Denys, C. First molecular evidence for the presence of Crocidura pachyura (Mammalia, Soricidae) in Kabylie (Algeria). Mammalia 2014, 78, 245–249. [Google Scholar] [CrossRef]
- Aulagnier, S.; Haffner, P.; Mitchell-Jones, A.J.; Moutou, F.; Zima, J. Guide des Mammifères d’Europe, d’Afrique du Nord et du Moyen-Orient; Delachaux & Niestlè SA Editions: Paris, France, 2010. [Google Scholar]
- Amori, G.; Contoli, L.; Nappi, A. Mammalia II: Erinaceomorpha, Soricomorpha, Lagomorpha, Rodentia; Il Sole 24 Ore; Edagricole, Calderini Editions: Bologna, Italy, 2008. [Google Scholar]
- Amori, G.; Rizzo Pinna, V.; Sammuri, G.; Luiselli, L. Diversity of small mammal communities of the Tuscan Archipelago: Testing the effects of island size, distance from mainland and human density. Folia Zool. 2015, 64, 161–166. [Google Scholar] [CrossRef]
- Angelici, F.M.; Cattaneo, C.; Grano, M.; Nappi, A. About the presence of Crocidura suaveolens group (Soricomorpha, Soricidae) on Astipalaia Island (Dodecanese, Greece). Nat. Hist. Sci. 2018, 5, 3–6. [Google Scholar] [CrossRef]
- Catalan, J.; Poitevin, F. Les Crocidures du midi de la France: Leurs characteristiques gènètiques et morphologiques: La place dea populations corses. C.R. Acad. Sci. Paris (III) 1981, 292, 1017–1020. [Google Scholar]
- Catzeflis, F. Relations gènètiques entre trois espèces du genre Crocidura (Soricidae, Mammalia) en Europe. Mammalia 1983, 47, 387–400. [Google Scholar] [CrossRef]
- Castiglia, R.; Annesi, F.; Amori, G.; Solano, E.; Aloise, G. The phylogeography of Crocidura suaveolens from Southern Italy reveals the absence of an endemic lineage and supports a Trans-Adriatic connection with the Balkanic refugium. Hystrix 2017, 28, 104–106. [Google Scholar]
- Amori, G.; Angelici, F.M.; Boitani, L. Mammals of Italy: A revised checklist of species and subspecies. Senckenberg. Biol. 1999, 79, 271–286. [Google Scholar]
- Gippoliti, S. Checklist delle specie di mammiferi italiani (esclusi Mysticeti e Odontoceti): Contributo per la conservazione della biodiversità. Boll. Mus. Civ. Stor. Nat. Verona 2013, 37, 7–28. [Google Scholar]
- Sigillo, P.M. Micromammiferi Nelle Borre di Barbagianni (Tyto alba) Nella Liguria Occidentale. Master’s Thesis, Biological Sciences, University of Rome “La Sapienza”, Rome, Italy, 1983. [Google Scholar]
- Rosi, R.E. Analisi e confronto fra alcuni caratteri morfologici e morfometrici di Crocidura sp. della Lunigiana e di Crocidura leucodon (Hermann, 1780) e Crocidura russula (Hermann, 1780) della Francia. Quad. Stor. Nat. Cent. “Malaspina” 2006, 1, 35–36. [Google Scholar]
- Biedma, L.; Román, J.; Godoy, J.A.; Calzada, J. Using owl pellets to infer habitat associations and clarify the regional distribution of a cryptic shrew. J. Zool. 2019, 308, 139–148. [Google Scholar] [CrossRef]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome-c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar]
- Bellati, A.; Tiberti, R.; Cocca, W.; Galimberti, A.; Casiraghi, M.; Bogliani, G.; Galeotti, P. A dark shell hiding great variability: A molecular insight into the evolution and conservation of melanic Daphnia populations in the Alps. Zool. J. Linn. Soc. 2014, 171, 697–715. [Google Scholar] [CrossRef]
- Mazzamuto, M.V.; Galimberti, A.; Cremonesi, G.; Pisanu, B.; Chapuis, J.L.; Stuyck, J.; Amori, G.; Su, H.J.; Aloise, G.; Preatoni, D.G.; et al. Preventing species invasion: A role for integrative taxonomy? Integr. Zool. 2016, 11, 214–228. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D. BOLD: The Barcode of Life Data System (http://www.barcodinglife.org). Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Vogel, P.; Cosson, J.F.; Jurado, L.F.L. Taxonomic status and origin of the shrews (Soricidae) from the Canary islands inferred from a mtDNA comparison with the European Crocidura species. Mol. Phylogenet. Evol. 2003, 27, 271–282. [Google Scholar] [CrossRef]
- Karger, D.N.; Conrad, O.; Böhner, J.; Kawohl, T.; Kreft, H.; Soria-Auza, R.W.; Zimmermann, N.E.; Linder, H.P.; Kessler, M. Climatologies at high resolution for the earth’s land surface areas. Sci. Data 2017, 4, 170122. [Google Scholar] [CrossRef]
- Neteler, M.; Bowman, M.H.; Landa, M.; Metz, M. GRASS GIS: A multi-purpose open source GIS. Environ. Model. Softw. 2012, 31, 124–130. [Google Scholar] [CrossRef]
- European Environment Agency. Corine Land Cover 2012. Available online: https://www.eea.europa.eu/data-and-maps/data/external/corine-land-cover-2012 (accessed on 17 August 2020).
- Elith, J.; Phillips, S.J.; Hastie, T.; Dudík, M.; Chee, Y.E.; Yates, C.J. A statistical explanation of MaxEnt for ecologists. Divers. Distrib. 2011, 17, 43–57. [Google Scholar] [CrossRef]
- Merow, C.; Smith, M.J.; Silander, J.A. A practical guide to MaxEnt for modeling species’ distributions: What it does, and why inputs and settings matter. Ecography 2013, 36, 1058–1069. [Google Scholar] [CrossRef]
- Brambilla, M.; Resano-Mayor, J.; Arlettaz, R.; Bettega, C.; Binggeli, A.; Bogliani, G.; Braunisch, V.; Celada, C.; Chamberlain, D.; Chiffard Carricaburu, J.; et al. Potential distribution of a climate sensitive species, the White-winged Snowfinch Montifringilla nivalis in Europe. Bird Conserv. Int. 2020. [Google Scholar] [CrossRef]
- Muscarella, R.; Galante, P.J.; Soley-Guardia, M.; Boria, R.A.; Kass, J.M.; Uriarte, M.; Anderson, R.P. ENMeval: An R package for conducting spatially independent evaluations and estimating optimal model complexity for Maxent ecological niche models. Methods Ecol. Evol. 2014, 5, 1198–1205. [Google Scholar] [CrossRef]
- Vignali, S.; Barras, A.; Braunisch, V. SDMtune: Species Distribution Model Selection. R Package Version 1.1.1. Available online: https://github.com/ConsBiol-unibern/SDMtune (accessed on 17 August 2020).
- Phillips, S.J.; Anderson, R.P.; Schapire, R.E. Maximum entropy modeling of species geographic distributions. Ecol. Model. 2006, 190, 231–259. [Google Scholar] [CrossRef]
- Toschi, A. Insectivora Gray, 1827. In Mammalia. Generalità, Insectivora, Chitopteera. Collana “Fauna d’Italia”; Toschi, A., Lanza, B., Eds.; Edizioni Calderini: Bologna, Italy, 1959; Volume IV, pp. 65–186. [Google Scholar]
- Richter, H. Zur Taxonomie und Verbreitung der palaearktischen Crociduren (Mammalia Insectivora, Soricidae). Zool. Abh. Staatl. Mus. Tierk. Dresd. 1970, 31, 294–304. [Google Scholar]
- Aebischer, A.; Nyffeler, P.; Arlettaz, R. Wide-range dispersal in juvenile Eagle Owls (Bubo bubo) across the European Alps calls for transnational conservation programmes. J. Ornithol. 2010, 151, 1–9. [Google Scholar] [CrossRef]
- Massa, C.; Gabelli, F.M.; Cueto, G.R. Using GPS to determine movement patterns and foraging habitat selection of the common barn owl (Tyto alba). Hornero 2015, 30, 7–12. [Google Scholar]
- Mori, E.; Mazzetto, F.; Menchetti, M.; Bodino, N.; Grasso, E.; Sposimo, P. Feeding ecology of the scops owl, Otus scops (Aves: Strigiformes), in the island of Pianosa (Tuscan Archipelago, Central Italy) outside the breeding period. Ital. J. Zool. 2016, 83, 417–422. [Google Scholar] [CrossRef]
- Jaquiery, J.; Guèlat, J.; Broquet, T.; Berset-Brandli, L.; Pellegrini, E.; Moresi, R.; Hirzel, A.H.; Perrin, N. Habitat-quality effects on metapopulation dynamics in greater white-toothed shrews, Crocidura russula. Ecology 2008, 89, 2777–2785. [Google Scholar] [CrossRef]
- Aulagnier, S.; Hutterer, R.; Amori, G.; Kryštufek, B.; Yigit, N.; Mitsain, G.; Palomo, L.J. Crocidura russula. The IUCN Red List of Threatened Species 2016: E.T29652A115169607; IUCN Editors: Gland, Switzerland, 2016. [Google Scholar]
- Gagan, L.M.; Cornette, R.; Yearsley, J.M.; Montgomery, W.I.; Paupèrio, J.; Alves, P.C.; Butler, F.; Pascal, M.; Tresset, A.; Herrel, A.; et al. Molecular and morphological insights into the origin of the invasive greater white-toothed shrew (Crocidura russula) in Ireland. Biol. Invasions 2016, 18, 857–871. [Google Scholar] [CrossRef]
- Amori, G.; Castiglia, R. Mammal endemism in Italy: A review. Biogeographia 2018, 33, 19–31. [Google Scholar] [CrossRef]
- Brambilla, M.; Vitulano, S.; Spina, F.; Baccetti, N.; Gargallo, G.; Fabbri, E.; Guidali, F.; Randi, E. A molecular phylogeny of the Sylvia cantillans complex: Cryptic species within the Mediterranean basin. Mol. Phylogenet. Evol. 2008, 48, 461–472. [Google Scholar] [CrossRef]
- Gvoždík, V.; Canestrelli, D.; García-París, M.; Moravec, J.; Nascetti, G.; Recuero, E.; Teixeira, J.; Kotlík, P. Speciation history and widespread introgression in the European short-call tree frogs (Hyla arborea sensu lato, H. intermedia and H. sarda). Mol. Phylogenet. Evol. 2015, 83, 143–155. [Google Scholar] [CrossRef]
- Mulargia, M.; Corti, C.; Lunghi, E. The herpetofauna of the Monte Albo, Sardinia (Italy). Rus. J. Herpetol. 2018, 25, 172–176. [Google Scholar] [CrossRef]
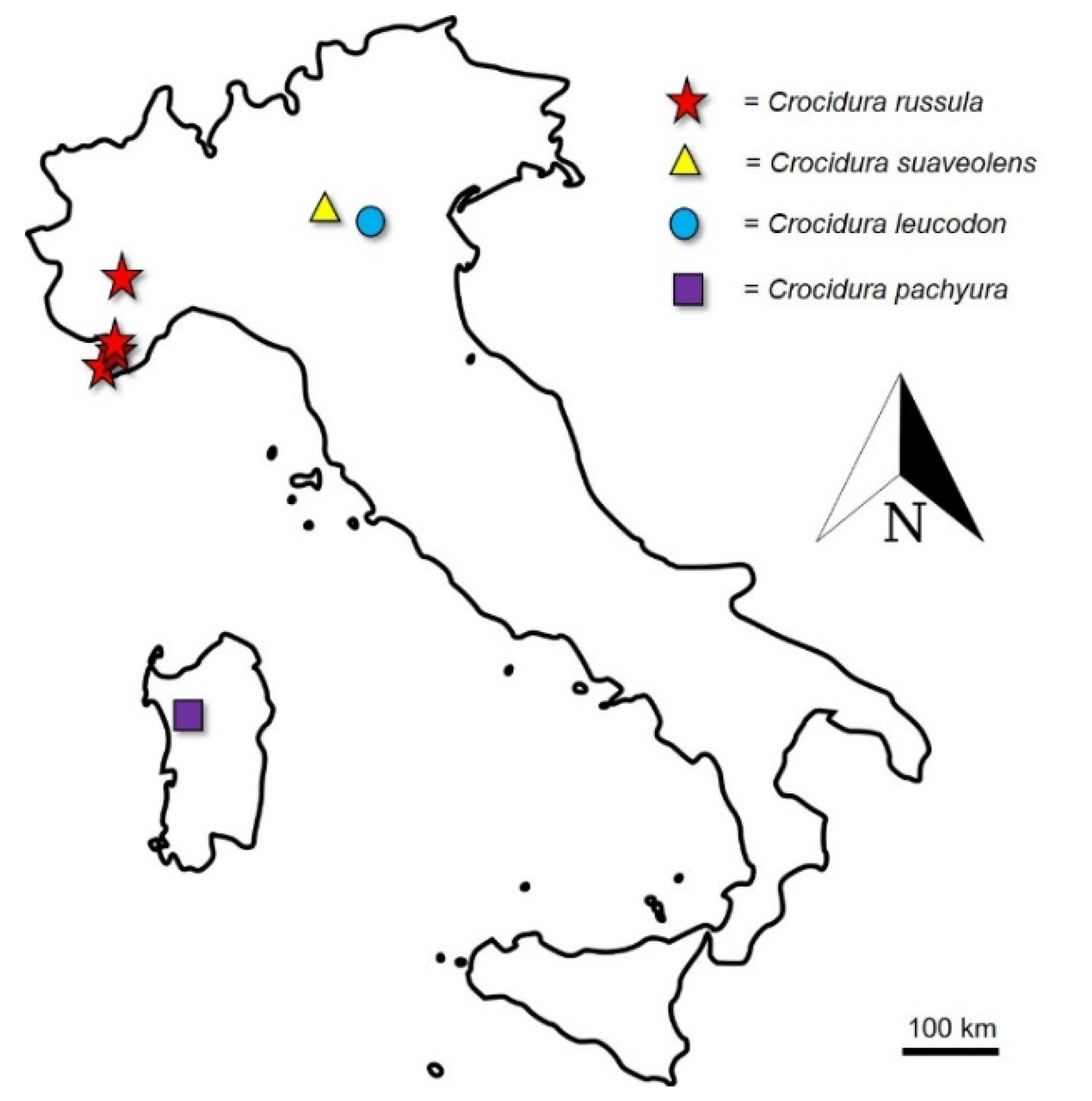


| Taxon | Voucher Sample | COI a.n. | Sampling Locality | Coordinates |
|---|---|---|---|---|
| C. cfr. russula | MIB:ZPL:08649 | LR877375 | Levaldigi (CN), Piedmont | 44.554216° N–7.633327° E |
| C. cfr. russula | MIB:ZPL:08650 | LR877376 | Dolceacqua (IM), Liguria | 43.860976° N–7.620309° E |
| C. cfr. russula | MIB:ZPL:08651 | LR877377 | Rocchetta Nervina (IM), Liguria | 43.890148° N–7.598244° E |
| C. cfr. russula | MIB:ZPL:08652 | LR877378 | Rocchetta Nervina (IM), Liguria | 43.890927° N–7.597591° E |
| C. suaveolens | MIB:ZPL:08653 | LR877379 | Ghedi (BS), Lombardy | 45.404473° N–10.267402° E |
| C. leucodon | MIB:ZPL:08654 | LR877380 | Oasi “Le Bine” (MN), Lombardy | 45.146581° N–10.445089° E |
| C. pachyura | MIB:ZPL:08655 | LR877381 | Thiesi (SS), Sardinia | 40.526080° N–8.663751° E |
| Taxon I | Taxon II | Distance | Standard Error |
|---|---|---|---|
| C. cfr. russula (Italy) | C. russula | 0.0000 | 0.0000 |
| C. cfr. russula (Italy) | C. suaveolens | 0.1592 | 0.0162 |
| C. cfr. russula (Italy) | C. leucodon | 0.1823 | 0.0205 |
| C. cfr. russula (Italy) | C. pachyura | 0.0892 | 0.0129 |
| C. cfr. russula (Italy) | C. olivieri | 0.1472 | 0.0168 |
| C. cfr. russula (Italy) | C. viaria | 0.1541 | 0.0177 |
| C. russula | C. suaveolens | 0.1592 | 0.0162 |
| C. russula | C. leucodon | 0.1823 | 0.0205 |
| C. russula | C. pachyura | 0.0892 | 0.0129 |
| C. russula | C. olivieri | 0.1472 | 0.0168 |
| C. russula | C. viaria | 0.1541 | 0.0177 |
| C. suaveolens | C. leucodon | 0.1376 | 0.0151 |
| C. suaveolens | C. pachyura | 0.1570 | 0.0160 |
| C. suaveolens | C. olivieri | 0.1466 | 0.0148 |
| C. suaveolens | C. viaria | 0.1364 | 0.0147 |
| C. leucodon | C. pachyura | 0.1632 | 0.0189 |
| C. leucodon | C. olivieri | 0.1676 | 0.0183 |
| C. leucodon | C. viaria | 0.1683 | 0.0186 |
| C. pachyura | C. olivieri | 0.1581 | 0.0172 |
| C. pachyura | C. viaria | 0.1607 | 0.0178 |
| C. olivieri | C. viaria | 0.0302 | 0.0048 |
| Variable | Permutation Importance | SD | Effect |
|---|---|---|---|
| C. russula | |||
| urban | 29 | 0.01 | positive/quadratic |
| temperature annual range (BIO7) | 16.8 | 0.02 | negative/quadratic |
| mean temp. of warmest quarter (BIO10) | 15.3 | 0.01 | quadratic |
| broad-leaved forest | 12.7 | 0.02 | negative/quadratic |
| precipitation of warmest quarter (BIO18) | 6.6 | 0.01 | negative |
| non-irrigated arable land | 6.1 | 0.01 | negative |
| bare ground and sparse vegetation | 4.0 | 0.01 | negative |
| sclerophyllous vegetation | 2.7 | 0.01 | positive/quadratic |
| complex crops | 2.7 | 0.00 | negative |
| vineyards | 2.5 | 0.00 | positive |
| fruit trees and berry plantations | 1.8 | 0.00 | positive |
| C. suaveolens | |||
| temperature seasonality (BIO4) | 53.7 | 0.02 | positive |
| min. temp. of the coldest month (BIO6) | 21.6 | 0.01 | positive |
| urban | 7.5 | 0.01 | positive/quadratic |
| broad-leaved forest | 6.9 | 0.01 | quadratic |
| mixed forest | 5.4 | 0.02 | quadratic/negative |
| slope | 3.9 | 0.01 | positive |
| inland marshes | 0.9 | 0.00 | quadratic |
| non-irrigated arable land | 0.0 | 0.00 | negative |
| Species | Body-Tail Length | Tail Length | Hind Foot | Condiloincisive Length | Weight |
|---|---|---|---|---|---|
| Crocidura pachyura | 62–78 mm | 30–40 mm | 11–13 mm | 18–22 mm | 8–16 g |
| Crocidura russula | 44–86 mm | 24–47 mm | 10–14 mm | 18–20 mm | 5–16 g |
| Crocidura sicula | 50–77 mm | 28–42 mm | 11–14 mm | 18–20 mm | 4–10 g |
| Crocidura suaveolens | 50–92 mm | 25–58 mm | 10–16 mm | 16–21 mm | 3–16 g |
| Crocidura leucodon | 50–90 mm | 27–46 mm | 11–15 mm | 17–24 mm | 7–20 g |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mori, E.; Brambilla, M.; Ramazzotti, F.; Ancillotto, L.; Mazza, G.; Russo, D.; Amori, G.; Galimberti, A. In or Out of the Checklist? DNA Barcoding and Distribution Modelling Unveil a New Species of Crocidura Shrew for Italy. Diversity 2020, 12, 380. https://doi.org/10.3390/d12100380
Mori E, Brambilla M, Ramazzotti F, Ancillotto L, Mazza G, Russo D, Amori G, Galimberti A. In or Out of the Checklist? DNA Barcoding and Distribution Modelling Unveil a New Species of Crocidura Shrew for Italy. Diversity. 2020; 12(10):380. https://doi.org/10.3390/d12100380
Chicago/Turabian StyleMori, Emiliano, Mattia Brambilla, Fausto Ramazzotti, Leonardo Ancillotto, Giuseppe Mazza, Danilo Russo, Giovanni Amori, and Andrea Galimberti. 2020. "In or Out of the Checklist? DNA Barcoding and Distribution Modelling Unveil a New Species of Crocidura Shrew for Italy" Diversity 12, no. 10: 380. https://doi.org/10.3390/d12100380
APA StyleMori, E., Brambilla, M., Ramazzotti, F., Ancillotto, L., Mazza, G., Russo, D., Amori, G., & Galimberti, A. (2020). In or Out of the Checklist? DNA Barcoding and Distribution Modelling Unveil a New Species of Crocidura Shrew for Italy. Diversity, 12(10), 380. https://doi.org/10.3390/d12100380










