Hyperdiverse Macrofauna Communities Associated with a Common Sponge, Stylissa carteri, Shift across Ecological Gradients in the Central Red Sea
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. DNA Barcoding
2.3. Sequence Analysis
2.4. Statistical Analysis
3. Results
3.1. Sample Summary
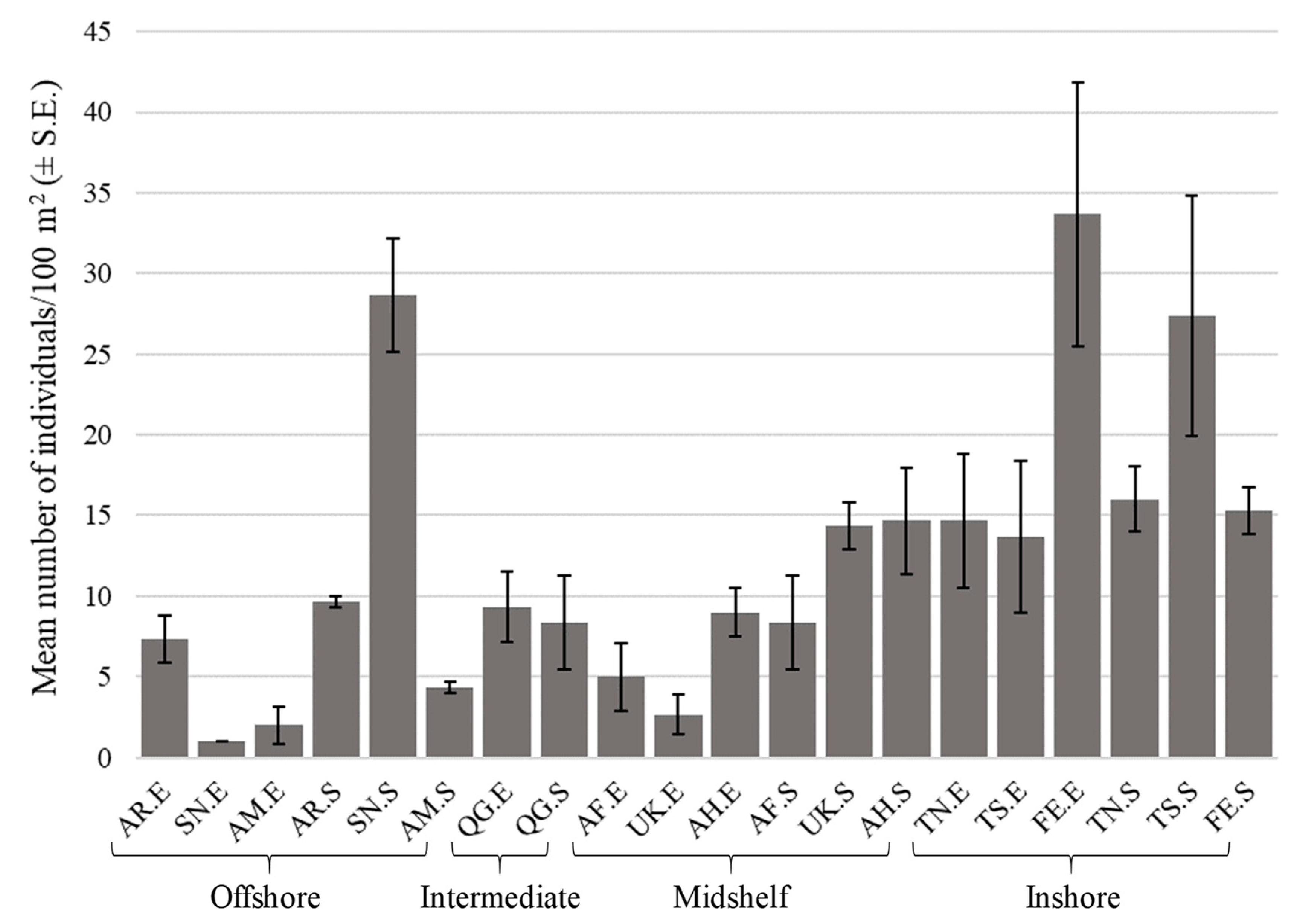
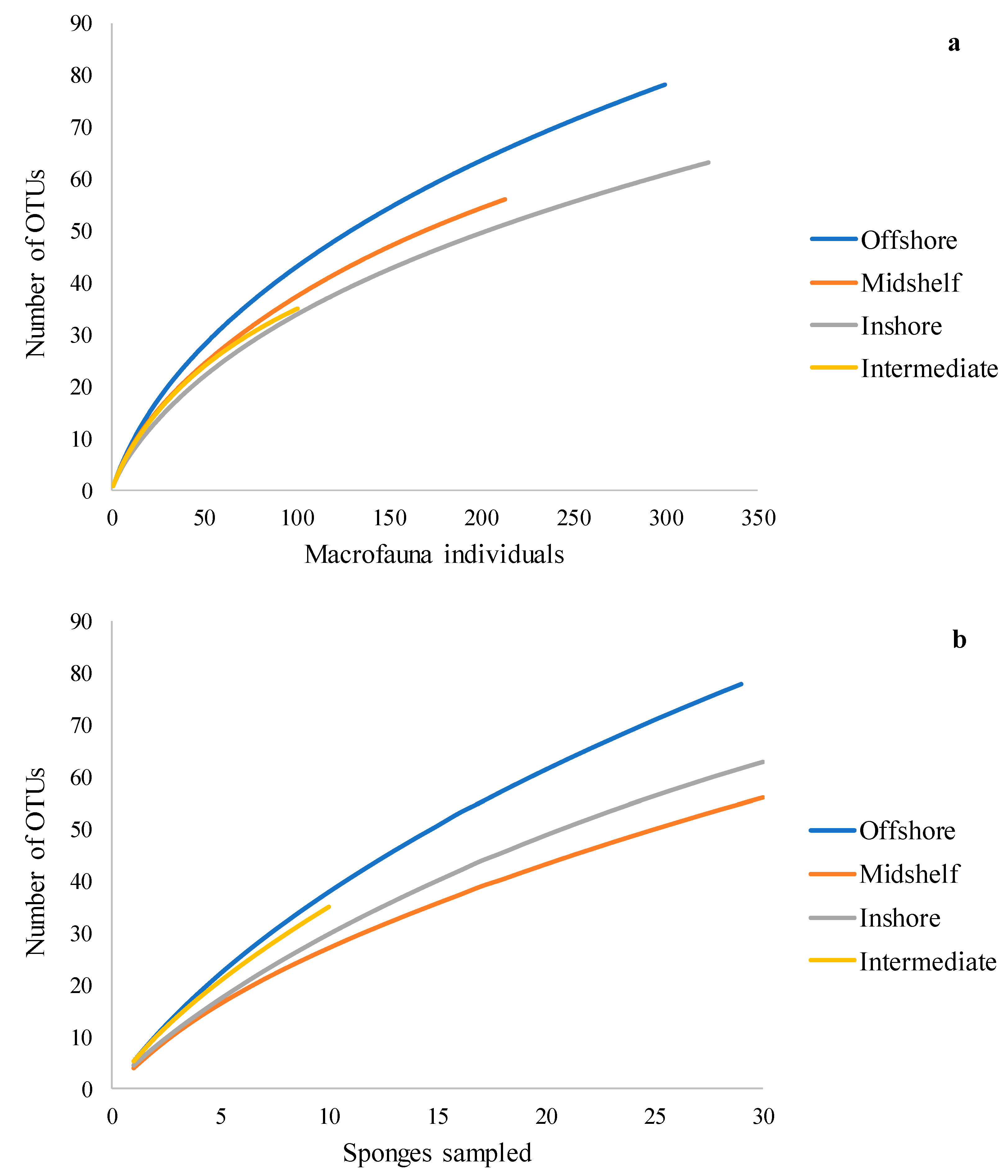
3.2. Summary Statistics
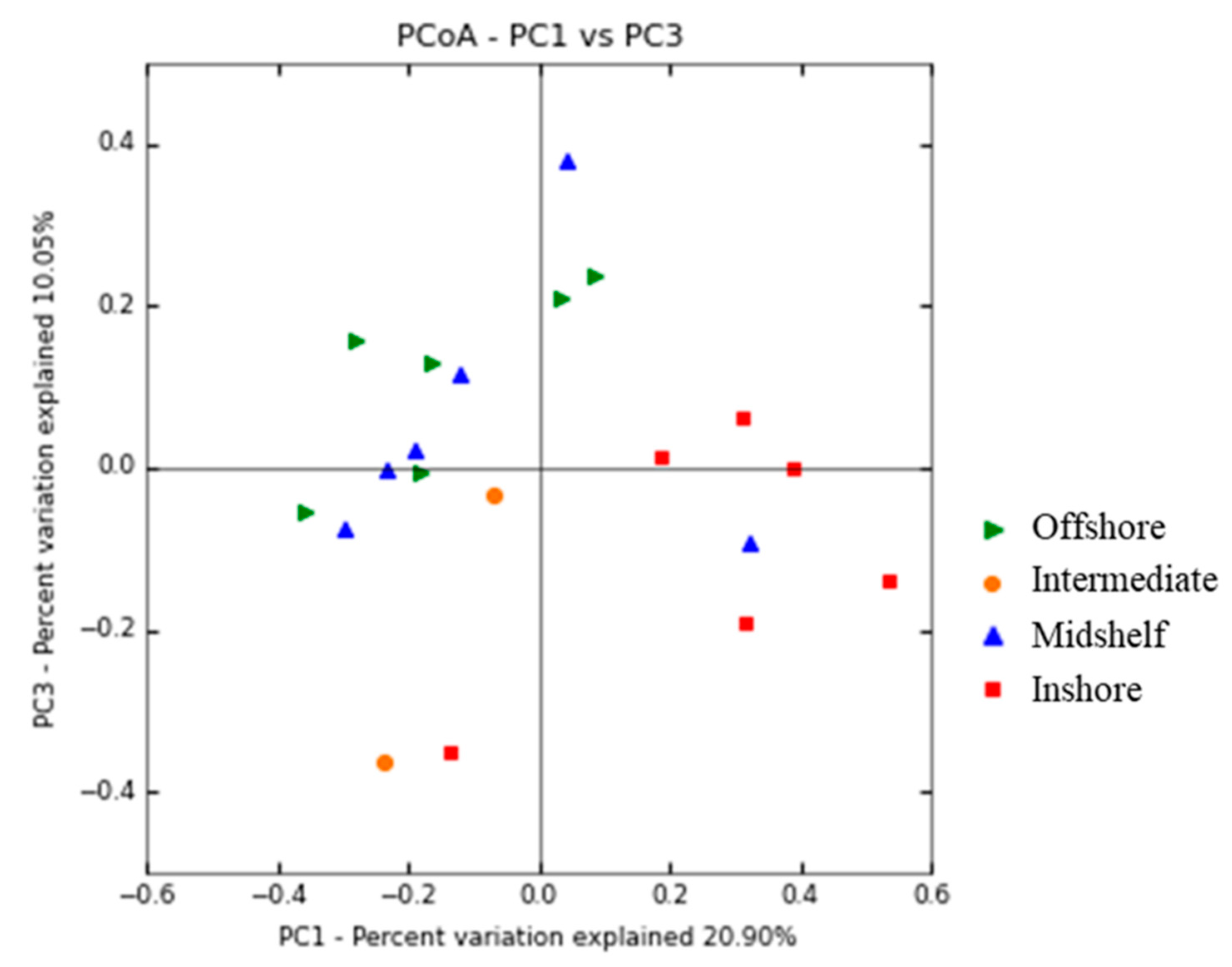
3.3. Spatial Patterns
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mora, C.; Tittensor, D.P.; Adl, S.; Simpson, A.G.B.; Worm, B. How Many Species Are There on Earth and in the Ocean? PLoS Biol. 2011, 9, e1001127. [Google Scholar] [CrossRef] [PubMed]
- Appeltans, W.; Ahyong, S.T.; Anderson, G.; Angel, M.V.; Artois, T.; Bailly, N.; Bamber, R.; Barber, A.; Bartsch, I.; Berta, A. The magnitude of global marine species diversity. Curr. Biol. 2012, 22, 2189–2202. [Google Scholar] [CrossRef]
- Diaz, M.C.; Rützler, K. Sponges: An essential component of Caribbean coral reefs. Bull. Mar. Sci. 2001, 69, 535–546. [Google Scholar]
- Wulff, J. Assessing and monitoring coral reef sponges: Why and how? Bull. Mar. Sci. 2001, 69, 831–846. [Google Scholar]
- Przeslawski, R.; Alvarez, B.; Kool, J.; Bridge, T.; Caley, M.J.; Nichol, S. Implications of sponge biodiversity patterns for the management of a marine reserve in northern Australia. PLoS ONE 2015, 10, e0141813. [Google Scholar] [CrossRef] [PubMed]
- Fromont, J.; Abdul Wahab, M.A.; Gomez, O.; Ekins, M.; Grol, M.; Hooper, J.N.A. Patterns of sponge biodiversity in the Pilbara, northwestern Australia. Diversity 2016, 8, 21. [Google Scholar] [CrossRef]
- Pearse, A. Notes on the inhabitants of certain sponges at Bimini. Ecology 1950, 31, 149–151. [Google Scholar] [CrossRef]
- Westinga, E.; Hoetjes, P. The intrasponge fauna of Spheciospongia vesparia (Porifera, Demospongiae) at Curacao and Bonaire. Mar. Biol. 1981, 62, 139–150. [Google Scholar] [CrossRef]
- Villamizar, E.; Laughlin, R. Fauna associated with the sponges Aplysina archeri and Aplysina lacunosa in a coral reef of the Archipielago de Los Roques, National Park, Venezuela. In Fossil and Recent Sponges; Springer: Berlin/Heidelberg, Germany, 1991; pp. 522–542. [Google Scholar]
- Gherardi, M.; Giangrande, A.; Corriero, G. Epibiontic and endobiontic polychaetes of Geodia cydonium (Porifera, Demospongiae) from the Mediterranean Sea. Hydrobiologia 2001, 443, 87–101. [Google Scholar] [CrossRef]
- Ribeiro, S.M.; Omena, E.P.; Muricy, G. Macrofauna associated to Mycale microsigmatosa (Porifera, Demospongiae) in Rio de Janeiro State, SE Brazil. Estuar. Coast. Shelf Sci. 2003, 57, 951–959. [Google Scholar] [CrossRef]
- Hultgren, K.M.; Duffy, J.E. Sponge host characteristics shape the community structure of their shrimp associates. Mar. Ecol. Prog. Ser. 2010, 407, 1–12. [Google Scholar] [CrossRef]
- Padua, A.; Lanna, E.; Klautau, M. Macrofauna inhabiting the sponge Paraleucilla magna (Porifera: Calcarea) in Rio de Janeiro, Brazil. J. Mar. Biol. Assoc. UK 2013, 93, 889–898. [Google Scholar] [CrossRef]
- Klitgaard, A.B. The fauna associated with outer shelf and upper slope sponges (Porifera, Demospongiae) at the Faroe Islands, northeastern Atlantic. Sarsia 1995, 80, 1–22. [Google Scholar] [CrossRef]
- Duffy, J.E. Eusociality in a coral-reef shrimp. Nature 1996, 381, 512–514. [Google Scholar] [CrossRef]
- Koukouras, A.; Russo, A.; Voultsiadou-Koukoura, E.; Arvanitidis, C.; Stefanidou, D. Macrofauna Associated with Sponge Species of Different Morphology. Mar. Ecol. 1996, 17, 569–582. [Google Scholar] [CrossRef]
- Kersken, D.; Göcke, C.; Brandt, A.; Lejzerowicz, F.; Schwabe, E.; Seefeldt, M.A.; Veit-Köhler, G.; Janussen, D. The infauna of three widely distributed sponge species (Hexactinellida and Demospongiae) from the deep Ekström Shelf in the Weddell Sea, Antarctica. Deep Sea Res. Part II Top. Stud. Oceanogr. 2014, 108, 101–112. [Google Scholar] [CrossRef]
- Bell, J.J. The functional roles of marine sponges. Estuar. Coast. Shelf Sci. 2008, 79, 341–353. [Google Scholar] [CrossRef]
- Magnino, G.; Sarà, A.; Lancioni, T.; Gaino, E. Endobionts of the Coral Reef Sponge Theonella swinhoei (Porifera, Demospongiae). Invertebr. Biol. 1999, 118, 213–220. [Google Scholar] [CrossRef]
- Abdo, D.A. Endofauna differences between two temperate marine sponges (Demospongiae; Haplosclerida; Chalinidae) from southwest Australia. Mar. Biol. 2007, 152, 845–854. [Google Scholar] [CrossRef]
- Wendt, P.; Van Dolah, R.; O’Rourke, C. A comparative study of the invertebrate macrofauna associated with seven sponge and coral species collected from the South Atlantic Bight. J. Elisha Mitchell Sci. Soc. 1985, 101, 187–203. [Google Scholar]
- Tyler, J.C.; Böhlke, J.E. Records of Sponge-Dwelling Fishes, Primarily of the Caribbean. Bull. Mar. Sci. 1972, 22, 601–642. [Google Scholar]
- Proksch, P. Defensive roles for secondary metabolites from marine sponges and sponge-feeding nudibranchs. Toxicon 1994, 32, 639–655. [Google Scholar] [CrossRef]
- Kohn, A. Feeding biology of gastropods. Mollus. Physiol. Part 1983, 2, 2–64. [Google Scholar]
- Wulff, L. Sponge-feeding by the Caribbean starfish Oreaster reticulatus. Mar. Biol. 1995, 123, 313–325. [Google Scholar] [CrossRef]
- Ďuriš, Z.; Horka, I.; Juračka, P.J.; Petrusek, A.; Sandford, F. These squatters are not innocent: The evidence of parasitism in sponge-inhabiting shrimps. PLoS ONE 2011, 6, e21987. [Google Scholar] [CrossRef] [PubMed]
- Swain, T.D. Context-dependent effects of symbiosis: Zoanthidea colonization generally improves Demospongiae condition in native habitats. Mar. Biol. 2012, 159, 1429–1438. [Google Scholar] [CrossRef]
- Tsubaki, R.; Kato, M. A novel filtering mutualism between a sponge host and its endosymbiotic bivalves. PLoS ONE 2014, 9, e108885. [Google Scholar] [CrossRef] [PubMed]
- Bornbusch, S.L.; Lefcheck, J.S.; Duffy, J.E. Allometry of individual reproduction and defense in eusocial colonies: A comparative approach to trade-offs in social sponge-dwelling Synalpheus shrimps. PLoS ONE 2018, 13, e0193305. [Google Scholar] [CrossRef] [PubMed]
- Voultsiadou-Koukoura, H.; Koukouras, A.; Eleftheriou, A. Macrofauna associated with the sponge Verongia aerophoba in the North Aegean Sea. Estuar. Coast. Shelf Sci. 1987, 24, 265–278. [Google Scholar] [CrossRef]
- Rützler, K. Sponges in coral reefs. In Coral Reefs: Res. Methods; Stoddart, D.R., Johannes, R.E., Eds.; UNESCO (Monogr. Oceanogr. Methodol. 5): Paris, France, 1978; pp. 299–313. Available online: https://repository.si.edu/bitstream/handle/10088/7849/iz_Ruetzler_1978b.pdf (accessed on 15 November 2015).
- Bell, J.J.; Davy, S.K.; Jones, T.; Taylor, M.W.; Webster, N.S. Could some coral reefs become sponge reefs as our climate changes? Glob. Chang. Biol. 2013, 19, 2613–2624. [Google Scholar] [CrossRef]
- Ruzicka, R.; Colella, M.; Porter, J.; Morrison, J.; Kidney, J.; Brinkhuis, V.; Lunz, K.; Macaulay, K.; Bartlett, L.; Meyers, M. Temporal changes in benthic assemblages on Florida Keys reefs 11 years after the 1997/1998 El Niño. Mar. Ecol. Prog. Ser. 2013, 489, 125–141. [Google Scholar] [CrossRef]
- McMurray, S.E.; Finelli, C.M.; Pawlik, J.R. Population dynamics of giant barrel sponges on Florida coral reefs. J. Exp. Mar. Biol. Ecol. 2015, 473, 73–80. [Google Scholar] [CrossRef]
- Loh, T.-L.; McMurray, S.E.; Henkel, T.P.; Vicente, J.; Pawlik, J.R. Indirect effects of overfishing on Caribbean reefs: Sponges overgrow reef-building corals. PeerJ 2015, 3, e901. [Google Scholar] [CrossRef] [PubMed]
- DiBattista, J.D.; Berumen, M.L.; Gaither, M.R.; Rocha, L.A.; Eble, J.A.; Choat, J.H.; Craig, M.T.; Skillings, D.J.; Bowen, B.W. After continents divide: Comparative phylogeography of reef fishes from the Red Sea and Indian Ocean. J. Biogeogr. 2013, 40, 1170–1181. [Google Scholar] [CrossRef]
- DiBattista, J.D.; Howard Choat, J.; Gaither, M.R.; Hobbs, J.P.A.; Lozano-Cortés, D.F.; Myers, R.F.; Paulay, G.; Rocha, L.A.; Toonen, R.J.; Westneat, M.W. On the origin of endemic species in the Red Sea. J. Biogeogr. 2016, 43, 13–30. [Google Scholar] [CrossRef]
- Raitsos, D.; Hoteit, I.; Prihartato, P.; Chronis, T.; Triantafyllou, G.; Abualnaja, Y. Abrupt warming of the Red Sea. Geophys. Res. Lett. 2011, 38. [Google Scholar] [CrossRef]
- Berumen, M.L.; Hoey, A.; Bass, W.; Bouwmeester, J.; Catania, D.; Cochran, J.E.; Khalil, M.; Miyake, S.; Mughal, M.; Spaet, J. The status of coral reef ecology research in the Red Sea. Coral Reefs 2013, 32, 737–748. [Google Scholar] [CrossRef]
- Ilan, M.; Loya, Y.; Kolbasov, G.; Brickner, I. Sponge-inhabiting barnacles on Red Sea coral reefs. Mar. Biol. 1999, 133, 709–716. [Google Scholar] [CrossRef]
- Pawlik, J.R.; Burkepile, D.E.; Thurber, R.V. A Vicious Circle? Altered Carbon and Nutrient Cycling May Explain the Low Resilience of Caribbean Coral Reefs. Bioscience 2016, 66, 470–476. [Google Scholar] [CrossRef]
- Dendy, A. IX—Report on a second collection of sponges from the Gulf of Manaar. J. Nat. Hist. 1889, 3, 73–99. [Google Scholar] [CrossRef]
- Khalil, M.T.; Bouwmeester, J.; Berumen, M.L. Spatial variation in coral reef fish and benthic communities in the central Saudi Arabian Red Sea. PeerJ 2017, 5, e3410. [Google Scholar] [CrossRef] [PubMed]
- Roberts, M.B.; Jones, G.P.; McCormick, M.I.; Munday, P.L.; Neale, S.; Thorrold, S.; Robitzch, V.S.; Berumen, M.L. Homogeneity of coral reef communities across 8 degrees of latitude in the Saudi Arabian Red Sea. Mar. Pollut. Bull. 2016, 105, 558–565. [Google Scholar] [CrossRef] [PubMed]
- Geller, J.; Meyer, C.; Parker, M.; Hawk, H. Redesign of PCR primers for mitochondrial cytochrome c oxidase subunit I for marine invertebrates and application in all-taxa biotic surveys. Mol. Ecol. Resour. 2013, 13, 851–861. [Google Scholar] [CrossRef] [PubMed]
- Leray, M.; Knowlton, N. DNA barcoding and metabarcoding of standardized samples reveal patterns of marine benthic diversity. Proc. Natl. Acad. Sci. USA 2015, 112, 2076–2081. [Google Scholar] [CrossRef] [PubMed]
- Leray, M.; Yang, J.Y.; Meyer, C.P.; Mills, S.C.; Agudelo, N.; Ranwez, V.; Boehm, J.T.; Machida, R.J. A new versatile primer set targeting a short fragment of the mitochondrial COI region for metabarcoding metazoan diversity: Application for characterizing coral reef fish gut contents. Front. Zool. 2013, 10, 34. [Google Scholar] [CrossRef] [PubMed]
- Hao, X.; Jiang, R.; Chen, T. Clustering 16S rRNA for OTU prediction: A method of unsupervised Bayesian clustering. Bioinformatics 2011, 27, 611–618. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Schaefer, D.A.; Liu, W.; Popescu, V.D.; Yang, C.; Wang, X.; Wu, C.; Douglas, W.Y. Higher fungal diversity is correlated with lower CO2 emissions from dead wood in a natural forest. Sci. Rep. 2016, 6, 31066. [Google Scholar] [CrossRef] [PubMed]
- Ratnasingham, S.; Hebert, P.D. BOLD: The Barcode of Life Data System (http://www.barcodinglife.org). Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Ratnasingham, S.; Hebert, P.D. A DNA-based registry for all animal species: The Barcode Index Number (BIN) system. PLoS ONE 2013, 8, e66213. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Plaisance, L.; Caley, M.J.; Brainard, R.E.; Knowlton, N. The diversity of coral reefs: What are we missing? PLoS ONE 2011, 6, e25026. [Google Scholar] [CrossRef] [PubMed]
- Leray, M.; Agudelo, N.; Mills, S.C.; Meyer, C.P. Effectiveness of annealing blocking primers versus restriction enzymes for characterization of generalist diets: Unexpected prey revealed in the gut contents of two coral reef fish species. PLoS ONE 2013, 8, e58076. [Google Scholar] [CrossRef] [PubMed]
- Colwell, R.K. EstimateS: Statistical Estimation of Species Richness and Shared Species from Samples. Version 7.5. User’s Guide and Application. 2005. Available online: http://purl.oclc.org/estimates (accessed on 15 October 2015).
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Anderson, M.J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 2001, 26, 32–46. [Google Scholar]
- Al-Rshaidat, M.M.; Snider, A.; Rosebraugh, S.; Devine, A.M.; Devine, T.D.; Plaisance, L.; Knowlton, N.; Leray, M. Deep COI sequencing of standardized benthic samples unveils overlooked diversity of Jordanian coral reefs in the northern Red Sea. Genome 2016, 59, 724–737. [Google Scholar] [CrossRef] [PubMed]
- Pearman, J.K.; Leray, M.; Villalobos, R.; Machida, R.; Berumen, M.L.; Knowlton, N.; Carvalho, S. Cross-shelf investigation of coral reef cryptic benthic organisms reveals diversity patterns of the hidden majority. Sci. Rep. 2018, 8, 8090. [Google Scholar] [CrossRef]
- Wooster, M.K.; McMurray, S.E.; Pawlik, J.R.; Moran, X.A.G.; Berumen, M.L. Feeding and respiration by giant barrel sponges across a gradient of food abundance in the Red Sea. Limnol. Oceanogr. 2019. under review. [Google Scholar]
- Wilkinson, C.R.; Cheshire, A.C. Patterns in the distribution of sponge populations across the central Great Barrier Reef. Coral Reefs 1989, 8, 127–134. [Google Scholar] [CrossRef]
- Mastaller, M. The marine molluscan assemblages of Port Sudan, Red Sea. Zool. Meded. 1978, 53, 117–144. [Google Scholar]
- Nanninga, G.B.; Saenz-Agudelo, P.; Zhan, P.; Hoteit, I.; Berumen, M.L. Not finding Nemo: Limited reef-scale retention in a coral reef fish. Coral Reefs 2015, 34, 383–392. [Google Scholar] [CrossRef]
- McClanahan, T.; Obura, D. Sedimentation effects on shallow coral communities in Kenya. J. Exp. Mar. Biol. Ecol. 1997, 209, 103–122. [Google Scholar] [CrossRef]
- Wilkinson, C.R. Interocean differences in size and nutrition of coral reef sponge populations. Science 1987, 236, 1654–1657. [Google Scholar] [CrossRef] [PubMed]
- Giles, E.C.; Saenz-Agudelo, P.; Hussey, N.E.; Ravasi, T.; Berumen, M.L. Exploring seascape genetics and kinship in the reef sponge Stylissa carteri in the Red Sea. Ecol. Evol. 2015, 5, 2487–2502. [Google Scholar] [CrossRef] [PubMed]
- Villamizar, E.; Díaz, M.C.; Rützler, K.; Nóbrega, R. Biodiversity, ecological structure, and change in the sponge community of different geomorphological zones of the barrier fore reef at Carrie Bow Cay, Belize. Mar. Ecol. 2014, 35, 425–435. [Google Scholar] [CrossRef]
- Wahab, M.A.A.; Radford, B.; Cappo, M.; Colquhoun, J.; Stowar, M.; Depczynski, M.; Miller, K.; Heyward, A. Biodiversity and spatial patterns of benthic habitat and associated demersal fish communities at two tropical submerged reef ecosystems. Coral Reefs 2018, 37, 327–343. [Google Scholar] [CrossRef]
- Furby, K.A.; Bouwmeester, J.; Berumen, M.L. Susceptibility of central Red Sea corals during a major bleaching event. Coral Reefs 2013, 32, 505–513. [Google Scholar] [CrossRef]
- Norström, A.V.; Nyström, M.; Lokrantz, J.; Folke, C. Alternative states on coral reefs: Beyond coral–macroalgal phase shifts. Mar. Ecol. Prog. Ser. 2009, 376, 295–306. [Google Scholar] [CrossRef]
- Hughes, T.P.; Anderson, K.D.; Connolly, S.R.; Heron, S.F.; Kerry, J.T.; Lough, J.M.; Baird, A.H.; Baum, J.K.; Berumen, M.L.; Bridge, T.C. Spatial and temporal patterns of mass bleaching of corals in the Anthropocene. Science 2018, 359, 80–83. [Google Scholar] [CrossRef]
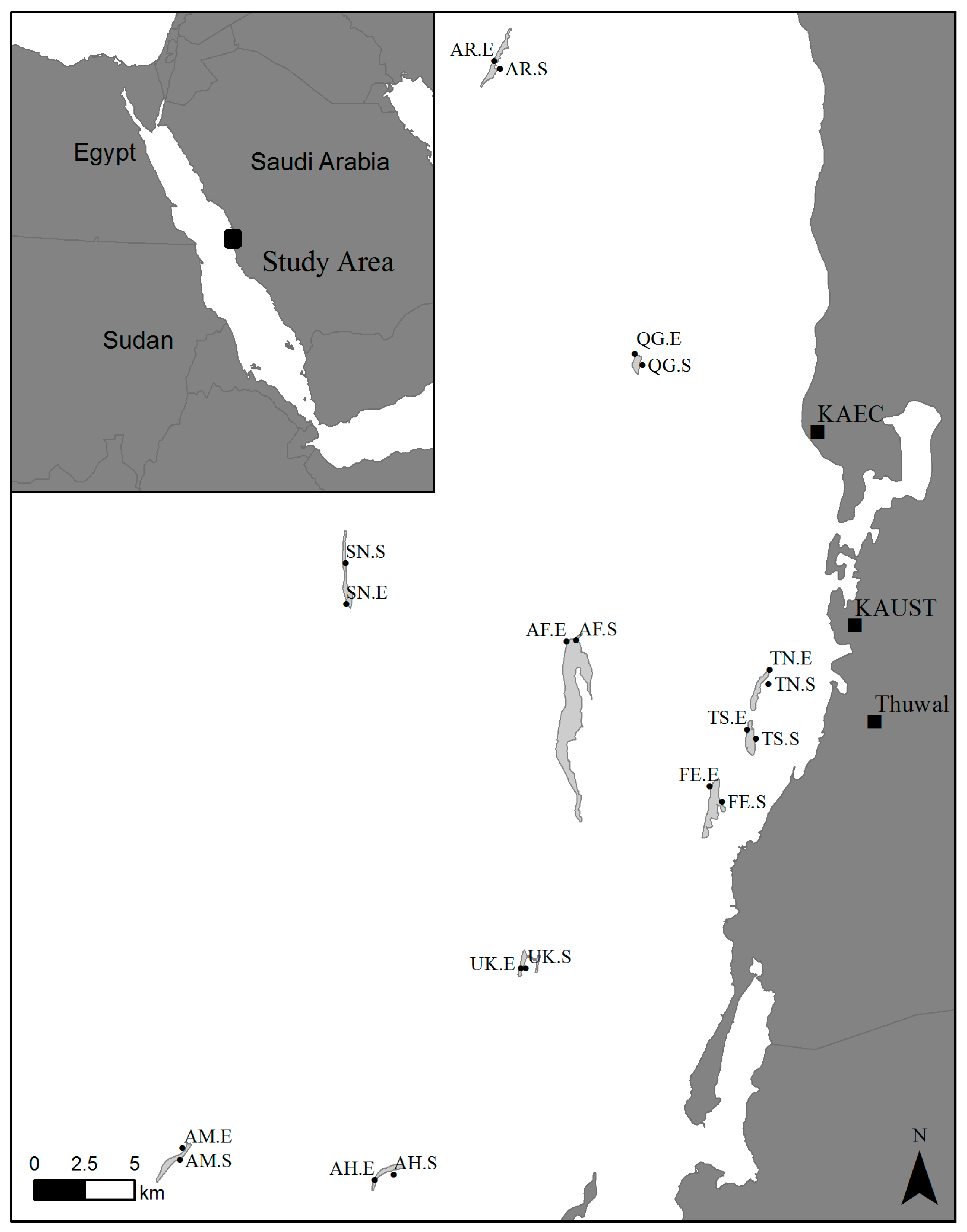

| Label | Reef | Classification | Exposure | Latitude (N) | Longitude (E) |
|---|---|---|---|---|---|
| AR.E | Abu Romah | Offshore | Exposed | 22°34.034′ | 38°55.541′ |
| SN.E | Shi’b Nazar | Offshore | Exposed | 22°19.317′ | 38°51.252′ |
| AM.E | Abu Madafi | Offshore | Exposed | 22°04.594′ | 38°46.505′ |
| AR.S | Abu Romah | Offshore | Sheltered | 22°33.833′ | 38°55.717′ |
| SN.S | Shi’b Nazar | Offshore | Sheltered | 22°20.434′ | 38°51.227′ |
| AM.S | Abu Madafi | Offshore | Sheltered | 22°04.258′ | 38°46.433′ |
| QG.E | Qita al-Girsh | Intermediate | Exposed | 22°26.105′ | 38°59.638′ |
| QG.S | Qita al-Girsh | Intermediate | Sheltered | 22°25.796′ | 38°59.855′ |
| AF.E | Al-Fahal | Midshelf | Exposed | 22°18.316′ | 38°57.648′ |
| UK.E | Umm al-Kiethl | Midshelf | Exposed | 22°09.467′ | 38°56.336′ |
| AH.E | Abu Henshan | Midshelf | Exposed | 22°03.723′ | 38°52.085′ |
| AF.S | Al-Fahal | Midshelf | Sheltered | 22°18.341′ | 38°57.930′ |
| UK.S | Umm al-Kiethl | Midshelf | Sheltered | 22°09.471′ | 38°56.470′ |
| AH.S | Abu Henshan | Midshelf | Sheltered | 22°03.863′ | 38°52.636′ |
| TN.E | Tahla North | Inshore | Exposed | 22°17.545′ | 39°03.562′ |
| TS.E | Tahla South | Inshore | Exposed | 22°15.927′ | 39°02.899′ |
| FE.E | Fsar East | Inshore | Exposed | 22°14.390′ | 39°01.817′ |
| TN.S | Tahla North | Inshore | Sheltered | 22°17.151′ | 39°03.530′ |
| TS.S | Tahla South | Inshore | Sheltered | 22°15.679′ | 39°03.154′ |
| FE.S | Fsar East | Inshore | Sheltered | 22°13.980′ | 39°02.181′ |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kandler, N.M.; Wooster, M.K.; Leray, M.; Knowlton, N.; de Voogd, N.J.; Paulay, G.; Berumen, M.L. Hyperdiverse Macrofauna Communities Associated with a Common Sponge, Stylissa carteri, Shift across Ecological Gradients in the Central Red Sea. Diversity 2019, 11, 18. https://doi.org/10.3390/d11020018
Kandler NM, Wooster MK, Leray M, Knowlton N, de Voogd NJ, Paulay G, Berumen ML. Hyperdiverse Macrofauna Communities Associated with a Common Sponge, Stylissa carteri, Shift across Ecological Gradients in the Central Red Sea. Diversity. 2019; 11(2):18. https://doi.org/10.3390/d11020018
Chicago/Turabian StyleKandler, Nora M., Michael K. Wooster, Matthieu Leray, Nancy Knowlton, Nicole J. de Voogd, Gustav Paulay, and Michael L. Berumen. 2019. "Hyperdiverse Macrofauna Communities Associated with a Common Sponge, Stylissa carteri, Shift across Ecological Gradients in the Central Red Sea" Diversity 11, no. 2: 18. https://doi.org/10.3390/d11020018
APA StyleKandler, N. M., Wooster, M. K., Leray, M., Knowlton, N., de Voogd, N. J., Paulay, G., & Berumen, M. L. (2019). Hyperdiverse Macrofauna Communities Associated with a Common Sponge, Stylissa carteri, Shift across Ecological Gradients in the Central Red Sea. Diversity, 11(2), 18. https://doi.org/10.3390/d11020018







