Molecular Diversity of Tidal Swamp Rice (Oryza sativa L.) in South Kalimantan, Indonesia
Abstract
1. Introduction
2. Materials and Methods
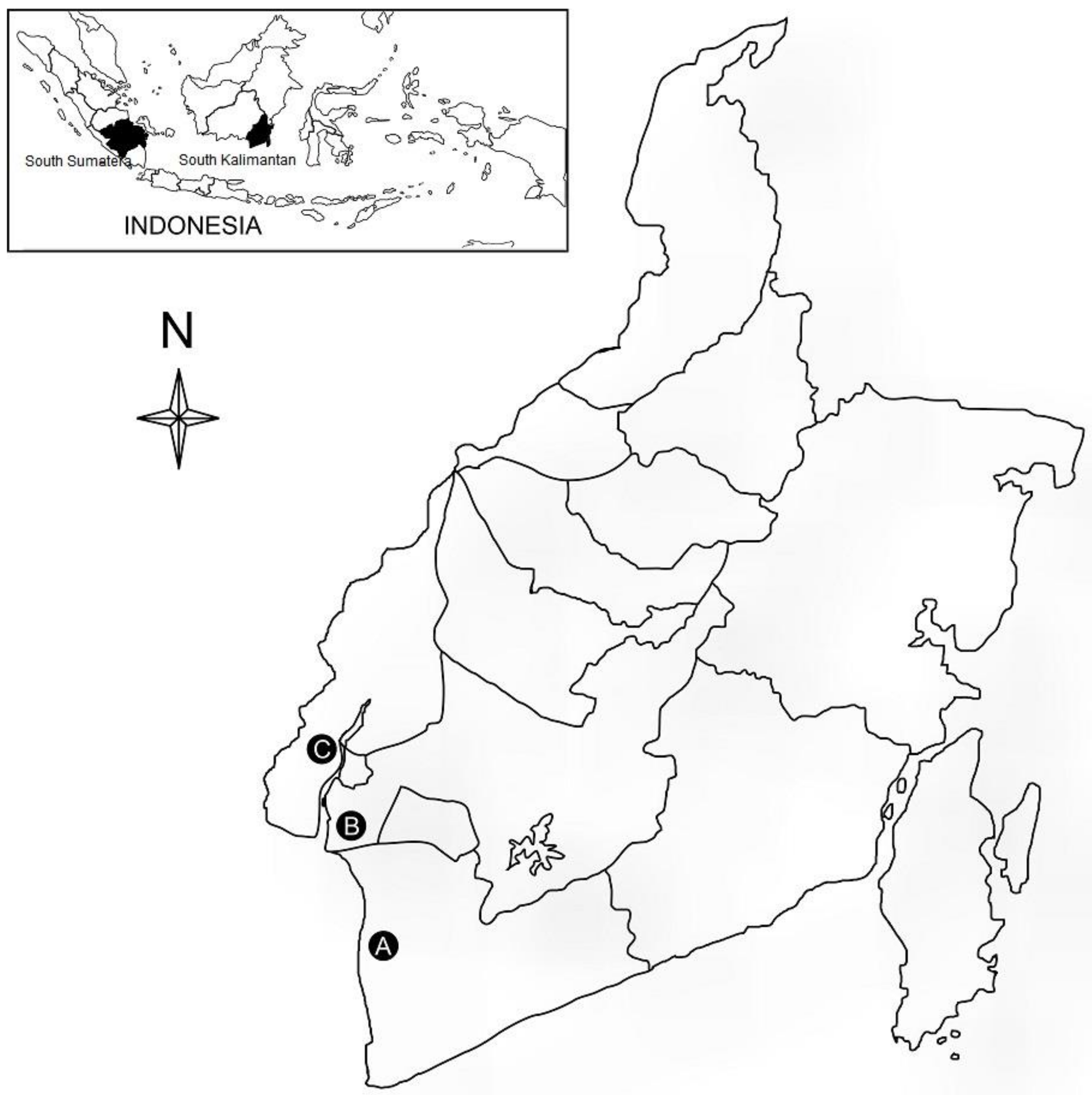
2.1. Plant Materials
2.2. DNA Isolation, Amplification, and Sequencing
2.3. Data Analysis
3. Results
3.1. Sequence Characteristics and Genetic Diversity
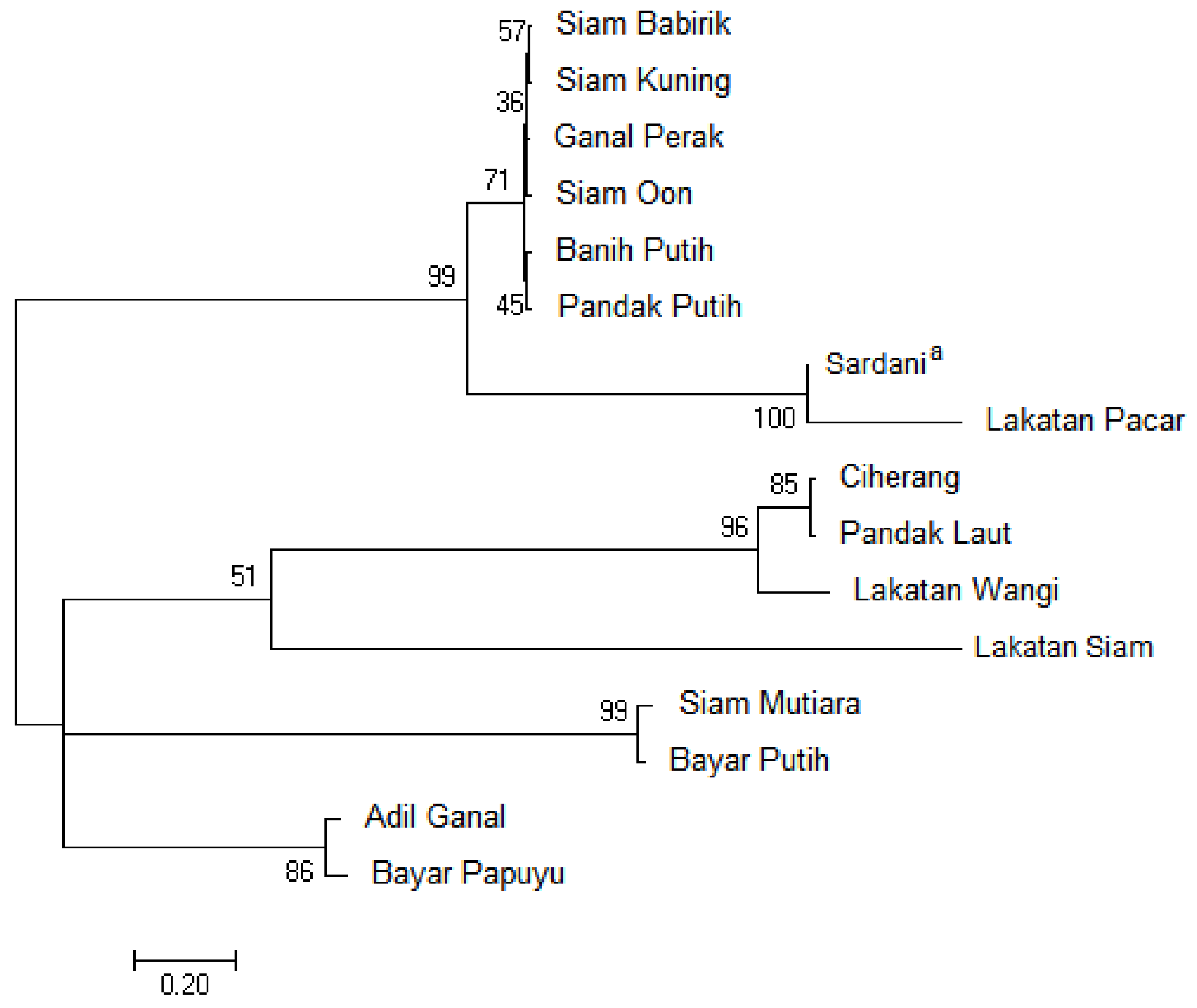
3.2. Phylogenetic Relationship Based on the Chloroplast trnL-F Spacer
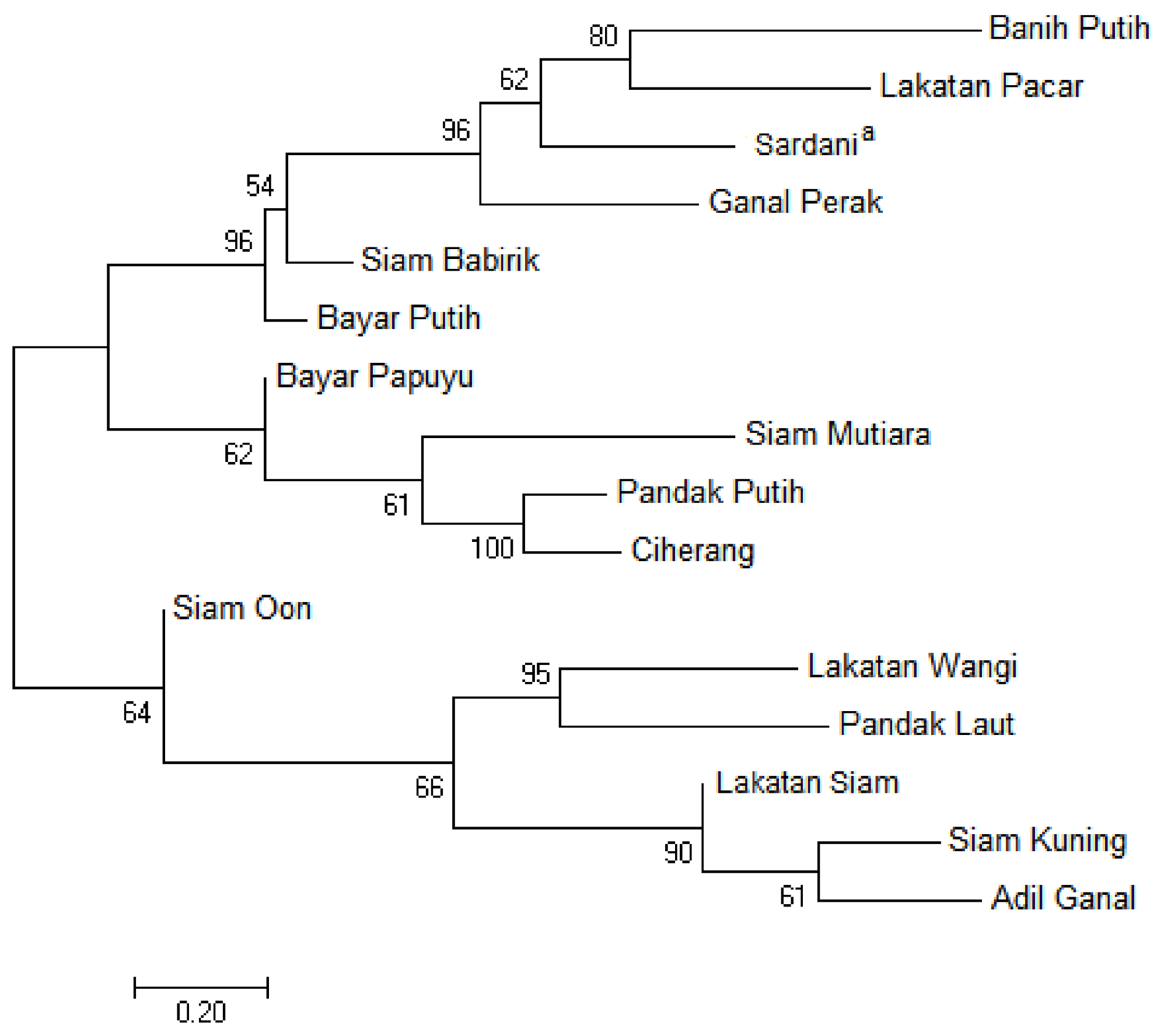
3.3. Phylogenetic Relationship Based on the Nuclear IGS Region
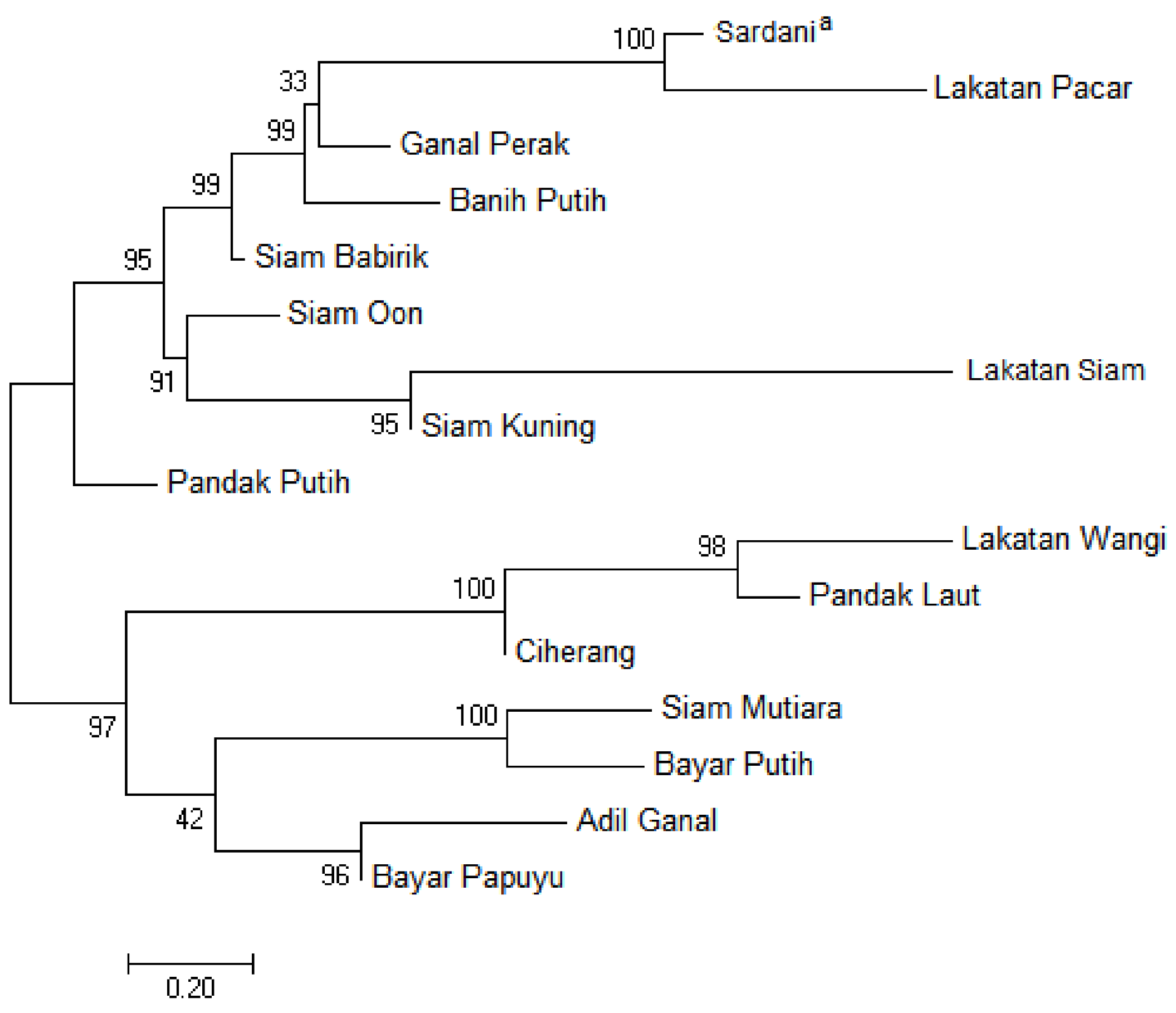
3.4. Phylogenetic Relationship of Combined Chloroplast and Nuclear Datasets
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Sinaga, P.H.; Trikoesoemaningtyas; Sopandie, D.; Aswidinnoor, H. Screening of rice genotypes and evaluation of their ratooning ability in tidal swamp area. Asian J. Agric. Res. 2014, 8, 218–233. [Google Scholar] [CrossRef][Green Version]
- Ray, A.; Deb, D.; Ray, R.; Chattopadhayay, B. Phenotypic characters of rice landraces reveal independent lineages of short-grain aromatic indica rice. AOB Plants 2013, 5, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Chakravorty, A.; Ghosh, P.D.; Sahu, P.K. Multivariate analysis of phenotypic diversity of landraces of rice of West Bengal. Am. J. Exp. Agric. 2013, 3, 110–123. [Google Scholar] [CrossRef]
- Divya, K. Study of genetic diversity in Karnataka rice (Oryza Sativa) landraces using trait specific simple sequence repeat (SSR) markers. Int. J. Thesis Proj. Diss. 2013, 1, 45–70. [Google Scholar]
- Das, B.; Sengupta, S.; Parida, S.K.; Roy, B.; Ghosh, M.; Prasad, M.; Ghose, T.K. Genetic diversity and population structure of rice landraces from Eastern and North Eastern States of India. BMC Genet. 2013, 14, 71. [Google Scholar] [CrossRef] [PubMed]
- Mursyidin, D.H.; Nazari, Y.A.; Daryono, B.S. Tidal swamp rice cultivars of South Kalimantan Province, Indonesia: A case study of diversity and local culture. Biodivers. J. Biol. Divers. 2017, 18, 427–432. [Google Scholar] [CrossRef]
- Thomson, M.J.; Septiningsih, E.M.; Suwardjo, F.; Santoso, T.J.; Silitonga, T.S.; McCouch, S.R. Genetic diversity analysis of traditional and improved Indonesian rice (Oryza sativa L.) germplasm using microsatellite markers. Theor. Appl. Genet. 2007, 114, 559–568. [Google Scholar] [CrossRef] [PubMed]
- Parikh, M.; Sharma, B.; Chaudhari, P.R.; Bhandarkar, S.; Sharma, D. Collection and conservation of extinct land races of rice from Raigarh district of Chhattisgarh. Recent Res. Sci. Technol. 2014, 6, 111–112. [Google Scholar]
- Semwal, D.P.; Pandey, A.; Bhandari, D.C.; Dhariwal, O.P.; Sharma, S.K. Variability study in seed morphology and uses of indigenous rice landraces (Oryza sativa L.) collected from West Bengal, India. Aust. J. Crop Sci. 2014, 8, 460–467. [Google Scholar]
- Anumalla, M.; Roychowdhury, R.; Geda, C.K.; Mazid, M.; Rathoure, A.K. Utilization of plant genetic resources and diversity analysis tools for sustainable crop improvement with special emphasis on rice. Int. J. Adv. Res. 2015, 3, 1155–1175. [Google Scholar]
- Liu, P.; Cai, X.X.; Lu, B.R. Single-seeded InDel fingerprints in rice: An effective tool for indica-japonica rice classification and evolutionary studies. J. Syst. Evol. 2012, 50, 1–11. [Google Scholar] [CrossRef]
- Pervaiz, Z.H.; Ashiq Rabbani, M.; Khaliq, I.; Pearce, S.R.; Malik, S.A. Genetic diversity associated with agronomic traits using microsatellite markers in Pakistani rice landraces. Electron. J. Biotechnol. 2010, 13, 4–5. [Google Scholar] [CrossRef][Green Version]
- Soltis, E.D.; Soltis, P.S. Contributions of plant molecular systematics to studies of molecular evolution. Plant Mol. Biol. 2000, 42, 45–75. [Google Scholar] [CrossRef] [PubMed]
- Duan, S.; Lu, B.; Li, Z.; Tong, J.; Kong, J.; Yao, W.; Li, S.; Zhu, Y. Phylogenetic analysis of AA-genome Oryza species (Poaceae) based on chloroplast, mitochondrial, and nuclear DNA sequences. Biochem. Genet. 2007, 45, 113–129. [Google Scholar] [CrossRef] [PubMed]
- Baldwin, B.G.; Sanderson, M.J.; Porter, J.M.; Wojciechowski, M.F.; Campbell, C.S.; Donoghue, M.J. The ITS region of nuclear ribosomal DNA—A valuable source of evidence on angiosperm phylogeny. Ann. Mo. Bot. Gard. 1995, 82, 247–277. [Google Scholar] [CrossRef]
- Soltis, D.E.; Soltis, P.S. Choosing an approach and an appropriate gene for phylogenetic anlysis. In Molecular Systematics of Plants II; Springer: Boston, MA, USA, 1998; pp. 1–42. ISBN 0412111314. [Google Scholar]
- Mort, M.E.; Archibald, J.K.; Randle, C.P.; Levsen, N.D.; Ryan O’Leary, T.; Topalov, K.; Wiegand, C.M.; Crawford, D.J. Inferring phylogeny at low taxonomic levels: Utility of rapidly evolving cpDNA and nuclear ITS loci. Am. J. Bot. 2007, 94, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Rogers, S.O.; Bendich, A.J. Ribosomal RNA genes in plants: Variability in copy number and in the intergenic spacer. Plant Mol. Biol. 1987, 9, 509–520. [Google Scholar] [CrossRef] [PubMed]
- Poczai, P.; Hyvönen, J. Nuclear ribosomal spacer regions in plant phylogenetics: Problems and prospects. Mol. Biol. Rep. 2010, 37, 1897–1912. [Google Scholar] [CrossRef] [PubMed]
- Taberlet, P.; Gielly, L.; Bouvet, J.; Pautou, G. Universal primer for amplification of three non-coding regions of chloroplast DNA. Plant Mol. Biol. 1991, 17, 1105–1109. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Robin, A.H.K.; Yi, G.E.; Lee, J.; Chung, M.Y.; Yang, T.J.; Nou, I.S. Diversity and inheritance of intergenic spacer sequences of 45s ribosomal DNA among accessions of Brassica oleracea L. var. capitata. Int. J. Mol. Sci. 2015, 16, 28783–28799. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Jeong, E.G.; Ahn, S.-N.; Doyle, J.; Singh, N.; Greenberg, A.J.; Won, Y.J.; McCouch, S.R. Nuclear and chloroplast diversity and phenotypic distribution of rice (Oryza sativa L.) germplasm from the democratic People’s Republic of Korea (DPRK; North Korea). Rice 2014, 7, 7. [Google Scholar] [CrossRef] [PubMed]
- Jo, S.-H.; Koo, D.-H.; Kim, J.F.; Hur, C.-G.; Lee, S.; Yang, T.; Kwon, S.-Y.; Choi, D. Evolution of ribosomal DNA-derived satellite repeat in tomato genome. BMC Plant Biol. 2009, 9, 42. [Google Scholar] [CrossRef] [PubMed]
- Pirie, M.D.; Vargas, M.P.B.; Botermans, M.; Bakker, F.T.; Chatrou, L.W. Ancient paralogy in the cpDNA trnL-F region in Annonaceae: Implications for plant molecular systematics. Am. J. Bot. 2007, 94, 1003–1016. [Google Scholar] [CrossRef] [PubMed]
- Thomson, M.J.; Polato, N.R.; Prasetiyono, J.; Trijatmiko, K.R.; Silitonga, T.S.; McCouch, S.R. Genetic diversity of isolated populations of Indonesian landraces of rice (Oryza sativa L.) collected in East Kalimantan on the Island of Borneo. Rice 2009, 2, 80–92. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1989; ISBN 13 978-0879693091. [Google Scholar]
- Cordesse, F.; Cooke, R.; Tremousaygue, D.; Grellet, F.; Delseny, M. Fine structure and evolution of the rDNA intergenic spacer in rice and other cereals. J. Mol. Evol. 1993, 36, 369–379. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; Mcgettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Petersen, G.; Seberg, O. Molecular evolution and phylogenetic application of DMC1. Mol. Phylogenet. Evol. 2002, 22, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide-sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Confidence-Limits on phylogenies—An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Data sets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Frankham, R.; Ballou, J.D.; Briscoe, D.A. A Primer of Conservation Genetics; Cambridge University Press: New York, NY, USA, 2004; Volume 39, ISBN 0521538270. [Google Scholar]
- Chung, M.C.; Lee, Y.I.; Cheng, Y.Y.; Chou, Y.J.; Lu, C.F. Chromosomal polymorphism of ribosomal genes in the genus Oryza. Theor. Appl. Genet. 2008, 116, 745–753. [Google Scholar] [CrossRef] [PubMed]
- Drábková, L.; Kirschner, J.; Vlcek, C.; Paces, V. TrnL-trnF intergenic spacer and trnL intron define major clades within Luzula and Juncus (Juncaceae): Importance of structural mutations. J. Mol. Evol. 2004, 59, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Bayer, R.; Starr, J. Tribal phylogeny of the Asteraceae based on two non-coding chloroplast sequences, the trnL intron and trnL/trnF intergenic spacer. Ann. Mo. Bot. Gard. 1998, 85, 242–256. [Google Scholar] [CrossRef]
- Guo, L.; Gao, Z.; Qian, Q. Application of resequencing to rice genomics, functional genomics and evolutionary analysis. Rice 2014, 7, 4. [Google Scholar] [CrossRef] [PubMed]
- Vaughan, D.A.; Lu, B.R.; Tomooka, N. Was Asian rice (Oryza sativa) domesticated more than once? Rice 2008, 1, 16–24. [Google Scholar] [CrossRef]
- Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 1989, 123, 585–595. [Google Scholar] [PubMed]
- Chakhonkaen, S.; Pitnjam, K.; Saisuk, W.; Ukoskit, K.; Muangprom, A. Genetic structure of Thai rice and rice accessions obtained from the International Rice Research Institute. Rice 2012, 5, 19. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, D.R.; Singh, N.; Singh, A.K.; Kumar, S.; Srinivasan, K. Analysis of genetic diversity and population structure of rice germplasm from North-Eastern region of India and development of a core germplasm set. PLoS ONE 2014, 9, e113094. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Hu, S.; Zhang, G.; Pan, L.; Zhang, X.; Al-Mssallem, I.S.; Yu, J. The organelle genomes of Hassawi rice (Oryza sativa L.) and its hybrid in Saudi Arabia: Genome variation, rearrangement, and origins. PLoS ONE 2012, 7, e42041. [Google Scholar] [CrossRef] [PubMed]
- Anderberg, A.A.; Rydin, C.; Källersjö, M. Phylogenetic relationships in the order Ericales s.l.: Analyses of molecular data from five genes from the plastid and mitochondrial genomes. Am. J. Bot. 2002, 89, 677–687. [Google Scholar] [CrossRef] [PubMed]
- Hall, J.C.; Sytsma, K.J.; Iltis, H.H. Phylogeny of Capparaceae and Brassicaceae based on chloroplast sequence data. Am. J. Bot. 2002, 89, 1826–1842. [Google Scholar] [CrossRef] [PubMed]
- Bellstedt, D.U.; Linder, H.P.; Harley, E.H. Phylogenetic relationships in Disa based on non-coding Trn L-Trn F chloroplast sequences: Evidence of numerous repeat region. Am. J. Bot. 2001, 88, 2088–2100. [Google Scholar] [CrossRef] [PubMed]
- Wambugu, P.W.; Brozynska, M.; Furtado, A.; Waters, D.L.; Henry, R.J. Relationships of wild and domesticated rices (Oryza AA genome species) based upon whole chloroplast genome sequences. Sci. Rep. 2015, 5, 13957. [Google Scholar] [CrossRef] [PubMed]
| Name of Cultivars | Genetic Status | Grain Type | Sub-Species | Origin |
|---|---|---|---|---|
| Adil Ganal | Landrace | Non-glutinous | Javanica | Banjar, South Kalimantan |
| Banih Putih | Landrace | Non-glutinous | Javanica | Banjar, South Kalimantan |
| Bayar Papuyu | Landrace | Non-glutinous | Close to Japonica | Banjar, South Kalimantan |
| Bayar Putih | Landrace | Non-glutinous | Javanica | Banjar, South Kalimantan |
| Ciherang | Improved | Non-glutinous | Javanica | Tanah Laut, South Kalimantan |
| Ganal Perak | Landrace | Non-glutinous | Javanica | Banjar, South Kalimantan |
| Lakatan Siam | Landrace | Glutinous | Javanica | Banjar, South Kalimantan |
| Lakatan Pacar | Landrace | Glutinous | Javanica | Barito Kuala, South Kalimantan |
| Lakatan Wangi | Landrace | Glutinous | Javanica | Banjar, South Kalimantan |
| Pandak Laut | Landrace | Non-glutinous | Intermediate | Banjar, South Kalimantan |
| Pandak Putih | Landrace | Non-glutinous | Javanica | Banjar, South Kalimantan |
| Siam Babirik | Landrace | Non-glutinous | Intermediate | Tanah Laut, South Kalimantan |
| Siam Kuning | Landrace | Non-glutinous | Javanica | Tanah Laut, South Kalimantan |
| Siam Mutiara | Landrace | Non-glutinous | Indica | Barito Kuala, South Kalimantan |
| Siam Oon | Landrace | Non-glutinous | Indica | Banjar, South Kalimantan |
| Sardani1 | Landrace | Non-glutinous | Close to Indica | South Sumatera |
| Marker | Primer | Sequences (5′ to 3′) | Expected Size (bp) | Ta/Tm (°C) | Reference |
|---|---|---|---|---|---|
| trnL-F | Forward | CGAAATCGGTAGACGCTACG | 950 | 55 | Taberlet et al. [20] |
| Reverse | ATTTGAACTGGTGACACGAG | ||||
| IGS | Forward | TAGGCCTCTCTAAGTCCGCT | 1612 | 53 | Cordesse et al. [27] |
| Reverse | GTCCAAAGCAGCTCATACGC |
| Parameter | Chloroplast trnL-F | Nuclear IGS | Combined |
|---|---|---|---|
| Range of sequence length (bp) | 936–953 | 849–1405 | 1796–2352 |
| Total number of sites (n) | 936 | 849 | 1780 |
| Number of polymorphic sites (S) | 929 | 845 | 266 |
| Parameters | 30 | 30 | 30 |
| Bayesian information criterion (BIC) | 20,221.72 | 29,081.93 | 60,694.01 |
| Akaike information criterion (AICc) | 19,993.42 | 28,856.57 | 60,446.29 |
| Maximum likelihood value (lnL) | −9966.65 | −14,398.22 | −30,193.11 |
| Transition/transversion bias value (R) | 0.58 | 0.56 | 0.68 |
| Ti/Tv ratio | 1.13 | 1.13 | 0.89 |
| GC content (%) | 35.71 | 69.10 | 51.60 |
| Nucleotide diversity (π) | 0.58 | 0.61 | 0.04 |
| Tajima’s test of neutrality (D) | 4.12 | 4.57 | −0.54 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mursyidin, D.H.; Purnomo, P.; Sumardi, I.; Daryono, B.S. Molecular Diversity of Tidal Swamp Rice (Oryza sativa L.) in South Kalimantan, Indonesia. Diversity 2018, 10, 22. https://doi.org/10.3390/d10020022
Mursyidin DH, Purnomo P, Sumardi I, Daryono BS. Molecular Diversity of Tidal Swamp Rice (Oryza sativa L.) in South Kalimantan, Indonesia. Diversity. 2018; 10(2):22. https://doi.org/10.3390/d10020022
Chicago/Turabian StyleMursyidin, Dindin H., Purnomo Purnomo, Issirep Sumardi, and Budi S. Daryono. 2018. "Molecular Diversity of Tidal Swamp Rice (Oryza sativa L.) in South Kalimantan, Indonesia" Diversity 10, no. 2: 22. https://doi.org/10.3390/d10020022
APA StyleMursyidin, D. H., Purnomo, P., Sumardi, I., & Daryono, B. S. (2018). Molecular Diversity of Tidal Swamp Rice (Oryza sativa L.) in South Kalimantan, Indonesia. Diversity, 10(2), 22. https://doi.org/10.3390/d10020022





