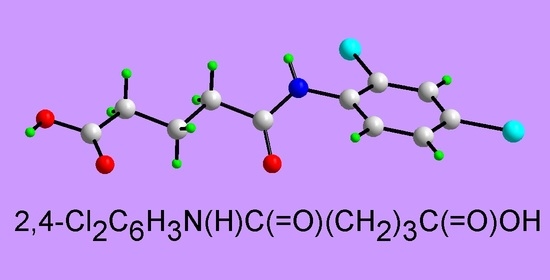
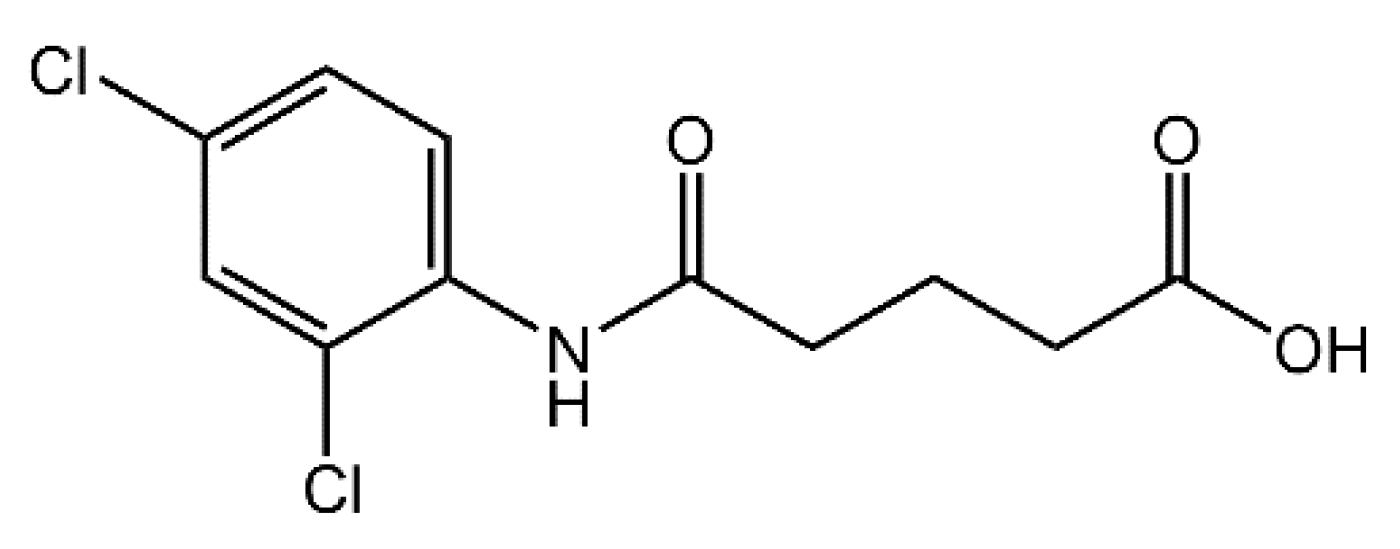
4-[(2,4-Dichlorophenyl)carbamoyl]butanoic Acid
Abstract
1. Introduction
2. Results and Discussion
3. Materials and Methods
3.1. General Information
3.2. Synthesis and Characterization of (1)
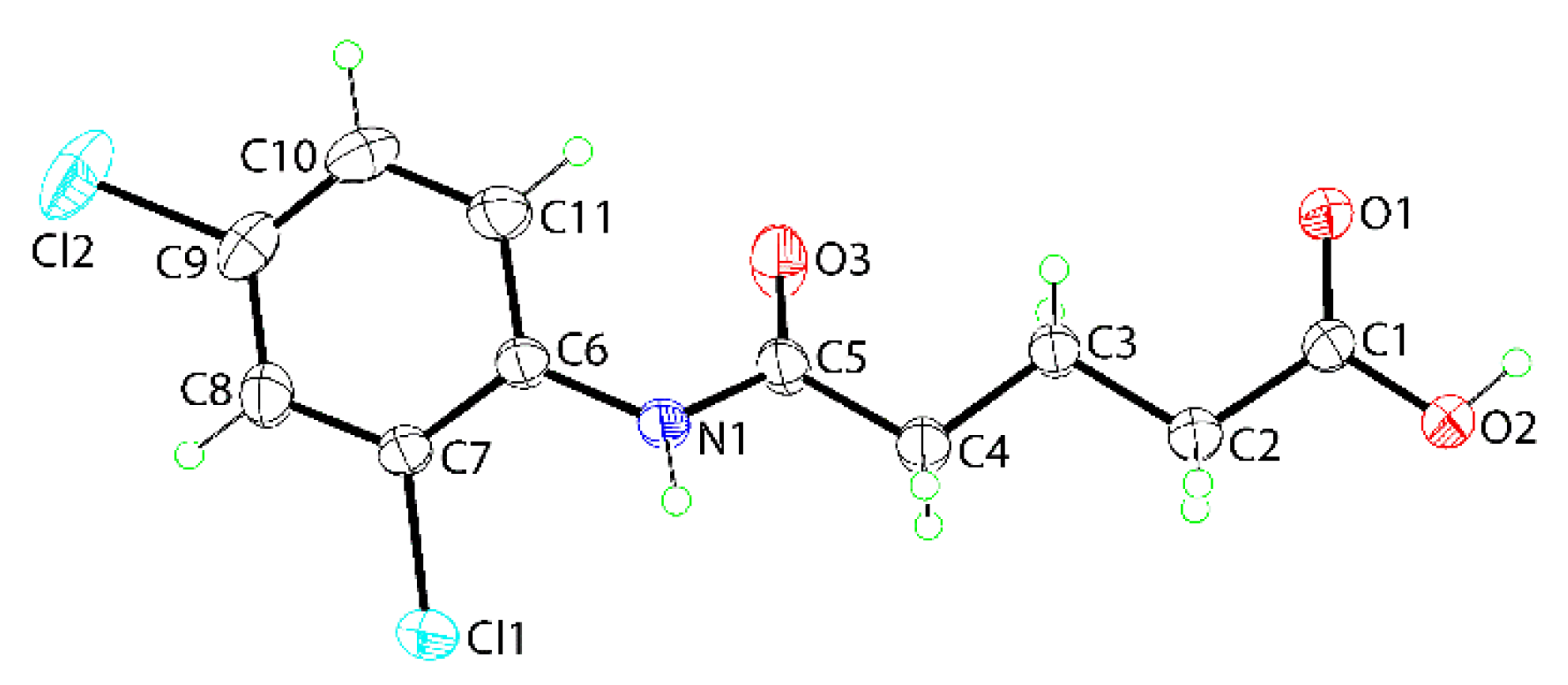
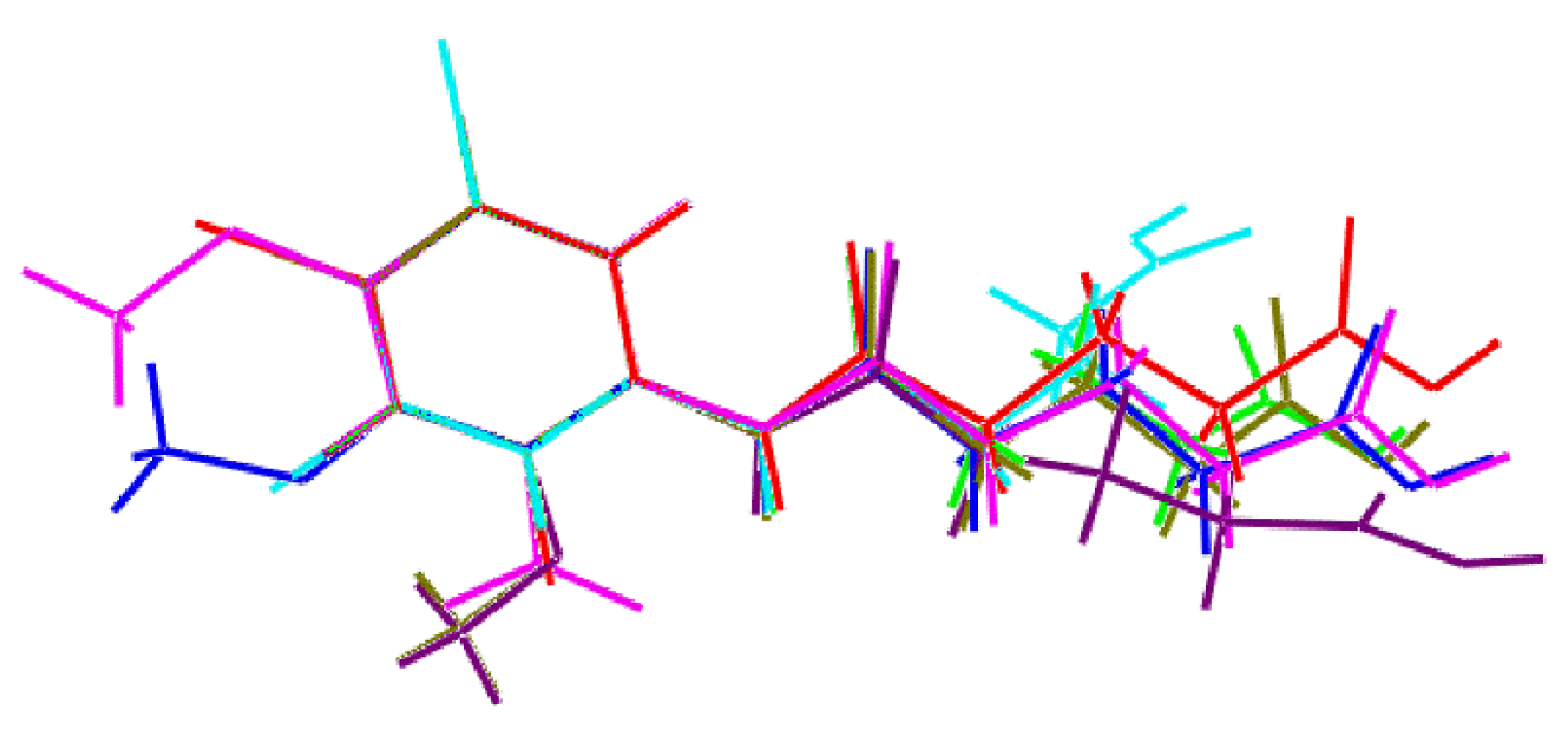
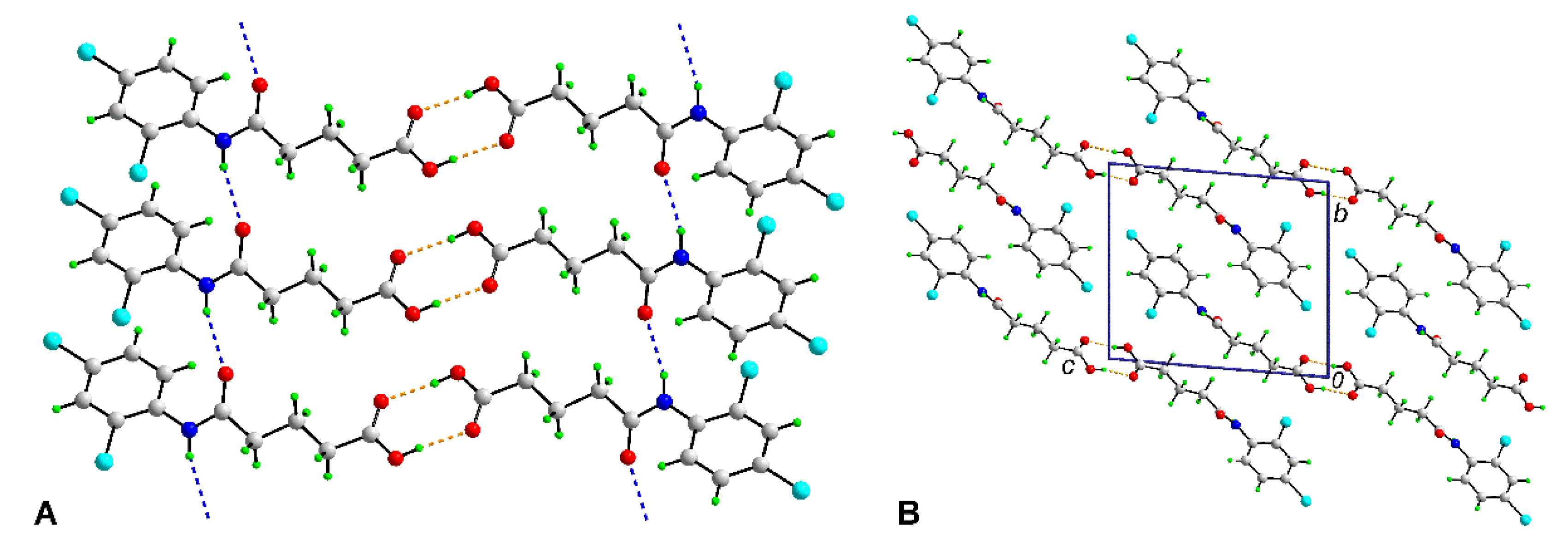
3.3. Crystallography
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sirajuddin, M.; Ali, S.; Shahnawaz, A.; Perveen, F.; Andleeb, S.; Alie, S. Exploration of biological potency of carboxylic acid derivatives: Designing, synthesis, characterizations and molecular docking study. J. Mol. Struct. 2020, 1207, 127809. [Google Scholar] [CrossRef]
- Sirajuddin, M.; Ali, S.; McKee, V.; Matin, A. Synthesis, characterization and biological screenings of 5-coordinated organotin(IV) complexes based on carboxylate ligand. J. Mol. Struct. 2020, 1206, 127683. [Google Scholar] [CrossRef]
- Hanifa, B.; Sirajuddin, M.; Lo, K.M.; Tiekink, E.R.T. Crystal structure of 4-[(4-methoxy-2-nitrophenyl)carbamoyl]butanoic acid, C12H14N2O6. Z. Kristallogr. New Cryst. Struct. 2020, 235, 1435–1437. [Google Scholar] [CrossRef]
- Hanifa, B.; Sirajuddin, M.; Lo, K.M.; Tiekink, E.R.T. Crystal structure of 4-[(2-methoxyphenyl)carbamoyl]butanoic acid, C12H15NO4. Z. Kristallogr. New Cryst. Struct. 2020, 235, 1481–1483. [Google Scholar] [CrossRef]
- Sirajuddin, M.; Hanifa, B.; Ullah, S.; Lo, K.M.; Tiekink, E.R.T. Crystal structure of 4-[(3,5-dichlorophenyl)carbamoyl]butanoic acid, C11H11Cl2NO3. Z. Kristallogr. New Cryst. Struct. 2020, 235, 1495–1497. [Google Scholar] [CrossRef]
- Sirajuddin, M.; Hanifa, B.; Ullah, S.; Lo, K.M.; Tiekink, E.R.T. Crystal structure of 4-[(3-methoxyphenyl)carbamoyl]butanoic acid, C12H15NO4. Z. Kristallogr. New Cryst. Struct. 2020, 235, 1519–1521. [Google Scholar] [CrossRef]
- Hanifa, B.; Sirajuddin, M.; Bari, A.; Lee, S.M.; Lo, K.M.; Tiekink, E.R.T. 4-[(4-Chlorophenyl)carbamoyl]butanoic acid. Molbank 2021, 2021, M1209. [Google Scholar] [CrossRef]
- Skinner, C.G.; Sargent, D.R. Homeosterically related plant growth regulators. I. Synthesis. J. Agric. Food Chem. 1973, 21, 1057–1060. [Google Scholar] [CrossRef]
- ChemDoodle Version 9.1.0, Chemical and Biological Publishing Software; iChemLabs LLC.: Piscataway, VA, USA, 2007–2021. Available online: http://www.chemdoodle.com (accessed on 2 June 2021).
- Spek, A.L. checkCIF validation ALERTS: What they mean and how to respond. Acta Crystallogr. E 2020, 76, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Rigaku Oxford Diffraction. CrysAlis PRO; Rigaku Corporation: Oxford, UK, 2017. [Google Scholar]
- Sheldrick, G.M. A short history of SHELX. Acta Crystallogr. A 2008, 64, 112–122. [Google Scholar] [CrossRef] [PubMed]
- Sheldrick, G.M. Crystal structure refinement with SHELXL. Acta Crystallogr. C 2015, 71, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Farrugia, L.J. WinGX and ORTEP for Windows: An update. J. Appl. Crystallogr. 2012, 45, 849–854. [Google Scholar] [CrossRef]
- Brandenburg, K.; Putz, H. DIAMOND; Crystal Impact GbR: Bonn, Germany, 2006. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hanifa, B.; Sirajuddin, M.; Ullah, Z.; Mahboob, S.; Lee, S.M.; Lo, K.M.; Tiekink, E.R.T. 4-[(2,4-Dichlorophenyl)carbamoyl]butanoic Acid. Molbank 2021, 2021, M1227. https://doi.org/10.3390/M1227
Hanifa B, Sirajuddin M, Ullah Z, Mahboob S, Lee SM, Lo KM, Tiekink ERT. 4-[(2,4-Dichlorophenyl)carbamoyl]butanoic Acid. Molbank. 2021; 2021(2):M1227. https://doi.org/10.3390/M1227
Chicago/Turabian StyleHanifa, Bibi, Muhammad Sirajuddin, Zafran Ullah, Sumera Mahboob, See Mun Lee, Kong Mun Lo, and Edward R. T. Tiekink. 2021. "4-[(2,4-Dichlorophenyl)carbamoyl]butanoic Acid" Molbank 2021, no. 2: M1227. https://doi.org/10.3390/M1227
APA StyleHanifa, B., Sirajuddin, M., Ullah, Z., Mahboob, S., Lee, S. M., Lo, K. M., & Tiekink, E. R. T. (2021). 4-[(2,4-Dichlorophenyl)carbamoyl]butanoic Acid. Molbank, 2021(2), M1227. https://doi.org/10.3390/M1227