5,7-Dihydroxy-3,6-Dimethoxy-3′,4′-Methylendioxyflavone
Abstract
:1. Introduction
2. Results and Discussion
2.1. General
2.2. Plant Material
2.3. Extraction and Isolation
2.4. Cytotoxic Assay
2.5. DPPH Radical Scavenging
3. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Tanjung, M.; Saputri, R.D.; Wahjoedi, R.A.; Tjahjandarie, T.S. 4-Methoxy-3-(3-methylbut-2-en-1-yl)-7-((3-methylbut-2-en-1-yl)oxy) quinolin-2(1H)-one from Melicope moluccana T.G. Hartley. Molbank 2017, 2017, M939. [Google Scholar] [CrossRef]
- Nakashima, K.; Oyama, M.; Ito, T.; Akao, Y.; Witono, J.R.; Darnaedi, D.; Tanaka, T.; Murata, T.; Iinuma, M. Novel quinolinone alkaloids bearing a lignoid moiety and related constituents in the leaves of Melicope denhamii. Tetrahedron 2012, 68, 2421–2428. [Google Scholar] [CrossRef]
- Kassim, N.K.; Rahmani, M.; Ismail, A.; Sukari, M.A.; Ee, G.C.L.; Nasir, N.M.; Awang, K. Antioxidant activity-guided separation of coumarins and lignin from Melicope glabra (Rutaceae). Food Chem. 2013, 139, 87–92. [Google Scholar] [CrossRef] [PubMed]
- Simonsen, H.T. Four novel geminaly dialkylated, non-aromatic acetophenone derivatives from Melicope coodeana. Phytochem. Lett. 2012, 5, 371–375. [Google Scholar] [CrossRef]
- Sultana, N.; Hartley, T.G.; Waterman, P.G. Two novel prenylated flavanones from the aerial parts of Melicope micrococca. Phytochemistry 1999, 50, 1249–1253. [Google Scholar] [CrossRef]
- Tanjung, M.; Juliawaty, L.D.; Hakim, E.H.; Syah, Y.M. Flavonoid and stilben derivatives from Macaranga trichocarpa. Fitoterapia 2018, 126, 74–77. [Google Scholar] [CrossRef] [PubMed]
- Marliana, E.; Astuti, W.; Kosala, K.; Hairani, R.; Tjahjandarie, T.S.; Tanjung, M. Chemical composition and anticancer activity of Macaranga hosei leaves. Asian J. Chem. 2018, 30, 795–798. [Google Scholar] [CrossRef]
- Tanjung, M.; Rachmadiarti, F.; Saputri, R.D.; Tjahjandarie, T.S. Mesuacalophylloidin, a new isoprenylated 4-phenylcoumarin from Mesua calophylloides (Ridl.) Kosterm. Nat. Prod. Res. 2018, 32, 1062–1067. [Google Scholar] [CrossRef] [PubMed]
- Tanjung, M.; Hakim, E.H.; Syah, Y.M. Prenylated dihydrostilbenes from Macaranga rubiginosa. Chem. Nat. Compd. 2017, 53, 215–218. [Google Scholar] [CrossRef]
- Tanjung, M.; Saputri, R.D.; Tjahjandarie, T.S. 5,9,11-Trihydroxy-2,2-dimethyl-10-(3′-methyl-2′-butenyl)-3-(2″-methyl-3″-butenyl)pyrano[2,3-a] xanthen-12(2H)-one from the stem bark of Calophyllum pseudomole. Molbank 2016, 2016, M906. [Google Scholar] [CrossRef]
- Tanjung, M.; Saputri, R.D.; Tjahjandarie, T.S. Antioxidant activity of two isomeric benzoxepin derivatives from the stem bark of Bauhinia acuelata L. J. Chem. Pharm. Res. 2014, 6, 705–708. [Google Scholar]
- Tjahjandarie, T.S.; Saputri, R.D.; Tanjung, M. Methyl 2,5-Dihydroxy-4-(3′-methyl-2′-butenyl)benzoate. Molbank 2016, 2016, M892. [Google Scholar] [CrossRef]
- Tjahjandarie, T.S.; Pudjiastuti, P.; Saputri, R.D.; Tanjung, M. Antimalaria and antioxidant activity of phenolic compounds isolated from Erythrina crysta-galli L. J. Chem. Pharm. Res. 2014, 6, 786–790. [Google Scholar]
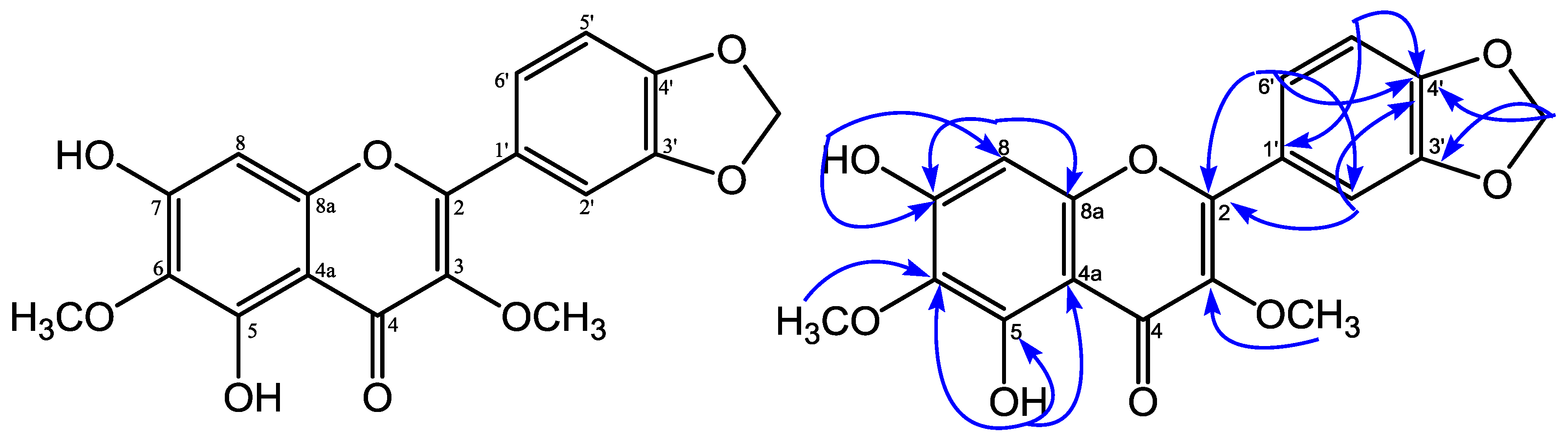
| No.C | δH (Mult. J in Hz) | δC | HMBC |
|---|---|---|---|
| 2 | - | 155.8 | - |
| 3 | - | 138.2 | - |
| 4 | - | 179.3 | - |
| 4a | - | 106.3 | - |
| 5 | - | 151.8 | - |
| 6 | - | 130.1 | - |
| 7 | - | 155.1 | - |
| 8 | 6.54 (s, 1H) | 93.2 | C-4a, C-6, C-7, C-8a |
| 8a | - | 153.9 | - |
| 1′ | - | 124.2 | - |
| 2′ | 7.59 (d, 1.8, 1H) | 108.7 | C-2, C-4′, C-6′ |
| 3′ | - | 149.7 | - |
| 4′ | - | 150.0 | - |
| 5′ | 6.95 (d, 8.4, 1H) | 108.6 | C-1′, C-3′ |
| 6′ | 7.68 (dd, 8.4; 1.8, 1H) | 123.8 | C-2, C-2′, C-4′ |
| 5-OH | 12.88 (s, 1H) | - | C-4a, C-5, C-6 |
| 7-OH | 6.50 (s, 1H) | - | C-7, C-8 |
| 3-OCH3 | 3.85 (s, 3H) | 61.0 | C-3 |
| 6-OCH3 | 4.04 (s, 3H) | 60.3 | C-6 |
| 3′,4′-OCH2-O- | 6.08 (s, 2H) | 101.8 | C-3′, C-4′ |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tjahjandarie, T.S.; Saputri, R.D.; Hasanah, U.; Rachmadiarti, F.; Tanjung, M. 5,7-Dihydroxy-3,6-Dimethoxy-3′,4′-Methylendioxyflavone. Molbank 2018, 2018, M1007. https://doi.org/10.3390/M1007
Tjahjandarie TS, Saputri RD, Hasanah U, Rachmadiarti F, Tanjung M. 5,7-Dihydroxy-3,6-Dimethoxy-3′,4′-Methylendioxyflavone. Molbank. 2018; 2018(3):M1007. https://doi.org/10.3390/M1007
Chicago/Turabian StyleTjahjandarie, Tjitjik Srie, Ratih Dewi Saputri, Ulfatun Hasanah, Fida Rachmadiarti, and Mulyadi Tanjung. 2018. "5,7-Dihydroxy-3,6-Dimethoxy-3′,4′-Methylendioxyflavone" Molbank 2018, no. 3: M1007. https://doi.org/10.3390/M1007
APA StyleTjahjandarie, T. S., Saputri, R. D., Hasanah, U., Rachmadiarti, F., & Tanjung, M. (2018). 5,7-Dihydroxy-3,6-Dimethoxy-3′,4′-Methylendioxyflavone. Molbank, 2018(3), M1007. https://doi.org/10.3390/M1007





