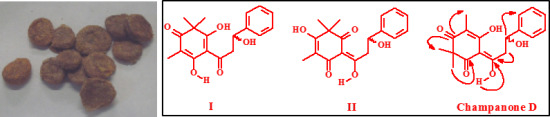
6-(1,3-Dihydroxy-3-phenylpropylidene)-5-hydroxy-2,2,4-trimethylcyclohex-4-ene-1,3-dione
Abstract
:Introduction
Results and Discussion
Experimental Section
Supplementary materials
Supplementary File 1Supplementary File 2Supplementary File 3Supplementary File 4Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bonilla, A.; Duque, C.; Garzon, C.; Takaishi, Y.; Yamaguchi, K.; Hara, N.; Fujimoto, Y. Champanones, yellow pigments from the seeds of champa (Campomanesia lineatifolia). Phytochemistry 2005, 66, 1736–1740. [Google Scholar] [CrossRef] [PubMed]
- Osorio, C.; Alarcon, M.; Moreno, C.; Bonilla, A.; Barrios, J.; Garzon, C.; Duque, C.J. Characterization of odor-active volatiles in Champa (Campomanesia lineatifolia R. & P.). J. Agric. Food Chem. 2006, 54, 509–516. [Google Scholar] [PubMed]
- Ghisalberti, E.L. Bioactive acylphloroglucinol derivatives from eucalyptus species. Phytochemistry 1996, 41, 7–22. [Google Scholar] [CrossRef]
- Liu, L.S.; Liu, M.H.; He, J.Y. Hypericum japonicum Thunb. ex Murray: Phytochemistry, pharmacology, quality control and pharmacokinetics of an important herbal medicine. Molecules 2014, 9, 10733–10754. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.I.; Bharate, S.P. Phloroglucinol compounds of natural origin. Nat. Prod. Rep. 2006, 23, 558–591. [Google Scholar] [CrossRef] [PubMed]
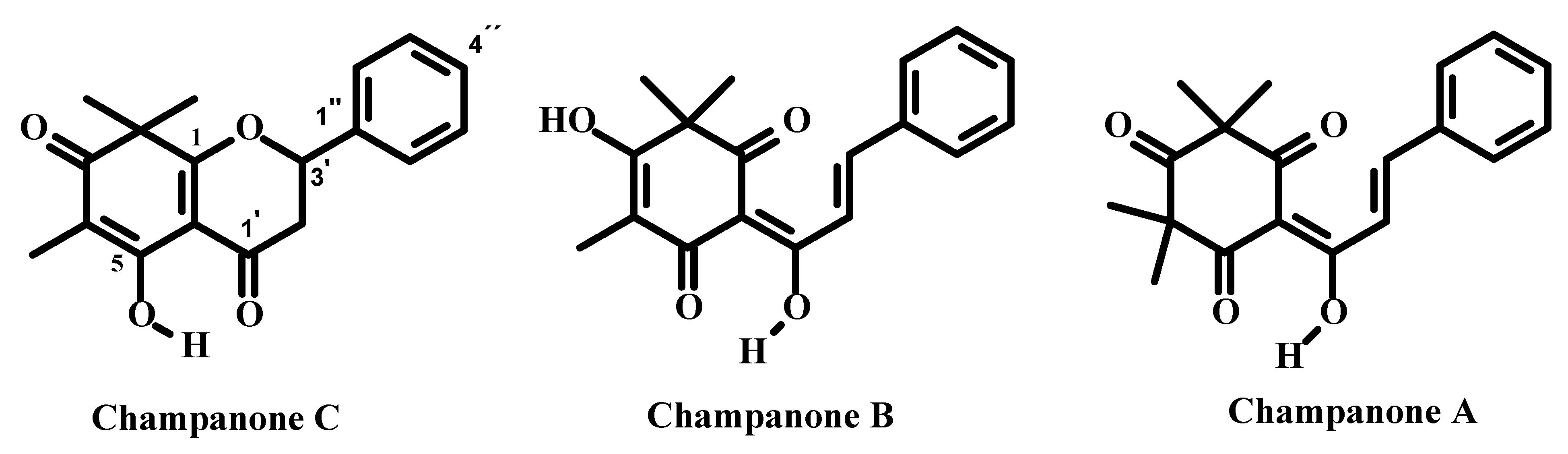
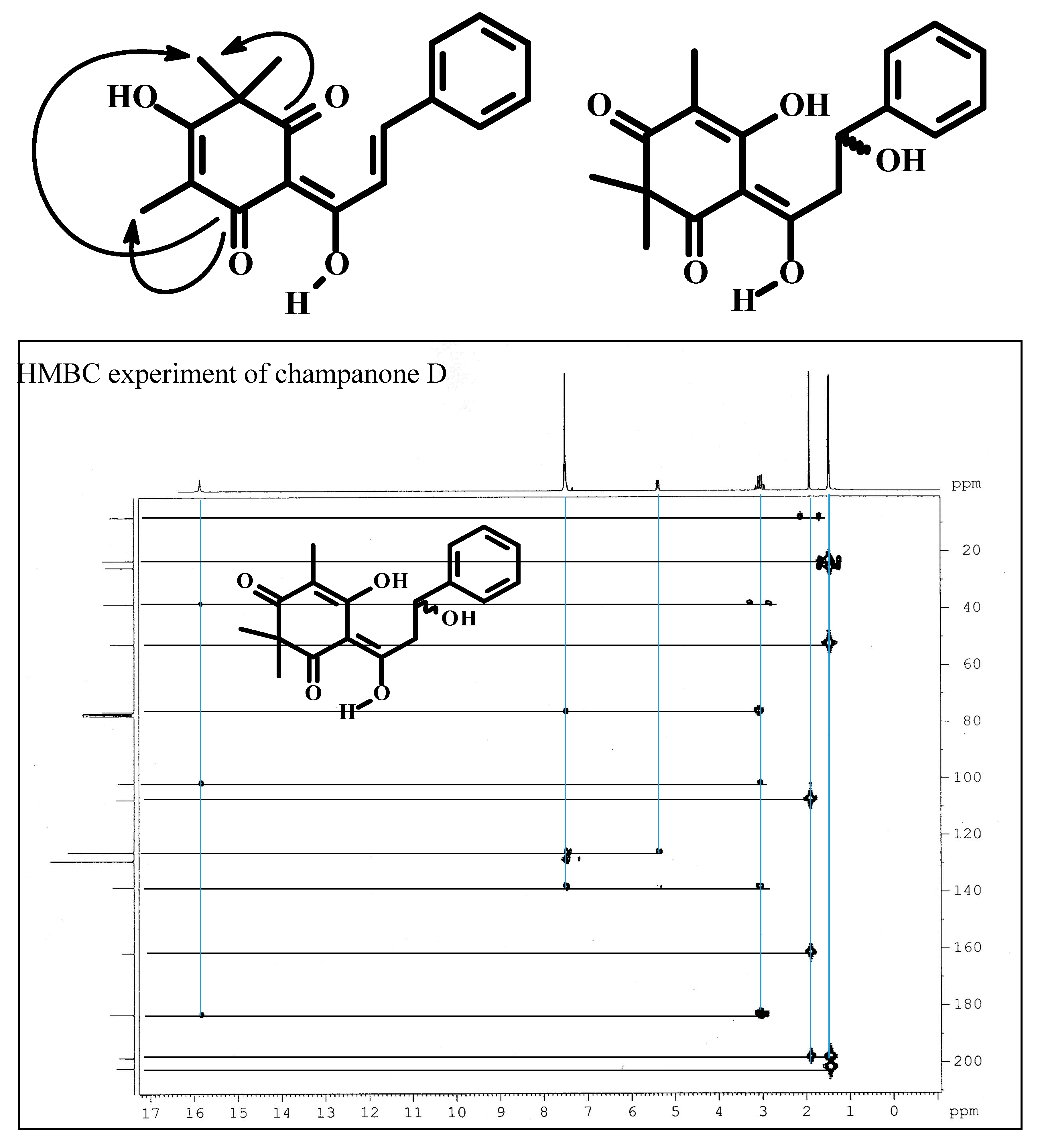
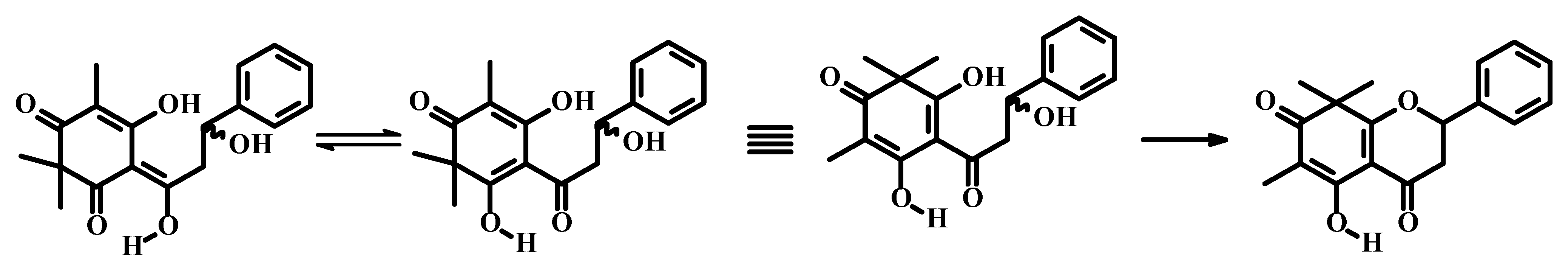
| No. | Champanone C 1 | Champanone C [1] | Champanone D 2 | Champanone B [1] | ||||
|---|---|---|---|---|---|---|---|---|
| δ1H | δ13C | δ1H | δ13C | δ1H | δ13C | δ1H | δ13C | |
| 1 | 186.4 | 186.0 | 161.32 | 197.4 | ||||
| 2 | 48.84 | 48.4 | 52.40 | 48.2 | ||||
| 2-Me | 1.43 s | 25.23 | 1.41 s | 24.8 | 1.41 s | 23.11 | 1.45 s | 24.6 |
| 2-Me′ | 1.46 s | 25.18 | 1.45 s | 24.7 | 1.43 s | 25.53 | 1.45 s | 24.6 |
| 3 | 197.05 | 196.3 | 201.79 | 172.1 | ||||
| 4 | 106.04 | 105.6 | 107.35 | 104.5 | ||||
| 4-Me | 1.82 s | 7.08 | 1.80 s | 6.6 | 1.87 s | 7.94 | 1.92 s | 6.7 |
| 5 | 164.50 | 164.0 | 182.91 | 191.0 | ||||
| 6 | 103.53 | 103.1 | 101.48 | 105.8 | ||||
| 1′ | 194.97 | 194.5 | 198.11 | 186.8 | ||||
| 2′ | 2.85 dd | 42.04 | 2.88 dd | 41.6 | 3.00 m | 38.26 | 7.92 d | 123.3 |
| 3.12 dd | 3.10 dd | |||||||
| 3′ | 5.60 dd | 81.62 | 5.58 dd | 81.2 | 5.35 d | 76.12 | 8.30 d | 144.5 |
| 1″ | 126.45 | 129.6 | 138.13 | 135.3 | ||||
| 2″ y 6″ | 7.44 m | 129.56 | 7.41 m | 126.0 | 7.45 s | 128.97 | 7.66 m | 128.8 |
| 3″ y 5″ | 7.48 m | 130.03 | 7.47 m | 129.1 | 7.45 s | 128.97 | 7.39 m | 129.0 |
| 4″ | 7.48 m | 7.47 m | 129.6 | 7.45 s | 128.97 | 7.39 m | 130.5 | |
| -OH | 11.65 s | 11.61 s | 15.86 s | 19.18 s | ||||
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Echeverri, F.; Gil, J.F.; Quiñones, W.; Correa, E. 6-(1,3-Dihydroxy-3-phenylpropylidene)-5-hydroxy-2,2,4-trimethylcyclohex-4-ene-1,3-dione. Molbank 2015, 2015, M853. https://doi.org/10.3390/M853
Echeverri F, Gil JF, Quiñones W, Correa E. 6-(1,3-Dihydroxy-3-phenylpropylidene)-5-hydroxy-2,2,4-trimethylcyclohex-4-ene-1,3-dione. Molbank. 2015; 2015(2):M853. https://doi.org/10.3390/M853
Chicago/Turabian StyleEcheverri, Fernando, Juan F. Gil, Winston Quiñones, and Edwin Correa. 2015. "6-(1,3-Dihydroxy-3-phenylpropylidene)-5-hydroxy-2,2,4-trimethylcyclohex-4-ene-1,3-dione" Molbank 2015, no. 2: M853. https://doi.org/10.3390/M853
APA StyleEcheverri, F., Gil, J. F., Quiñones, W., & Correa, E. (2015). 6-(1,3-Dihydroxy-3-phenylpropylidene)-5-hydroxy-2,2,4-trimethylcyclohex-4-ene-1,3-dione. Molbank, 2015(2), M853. https://doi.org/10.3390/M853