Calreticulin-Mediated Quality Control of the Non-Classical MHC-I Molecule MICA: Implications for Immune Surveillance
Abstract
1. Introduction
2. Results
2.1. MICA and MICB Are Predominantly Retained Intracellularly in Malignant Melanoma
2.2. MICA and MICB Colocalize with Endoplasmic Reticulum Chaperones in Melanoma Cells
2.3. Calreticulin Associates with MICA in Melanoma Cells
2.4. Direct Interaction Between MICA and Calreticulin May Be Influenced by the Glycosylation State of MICA
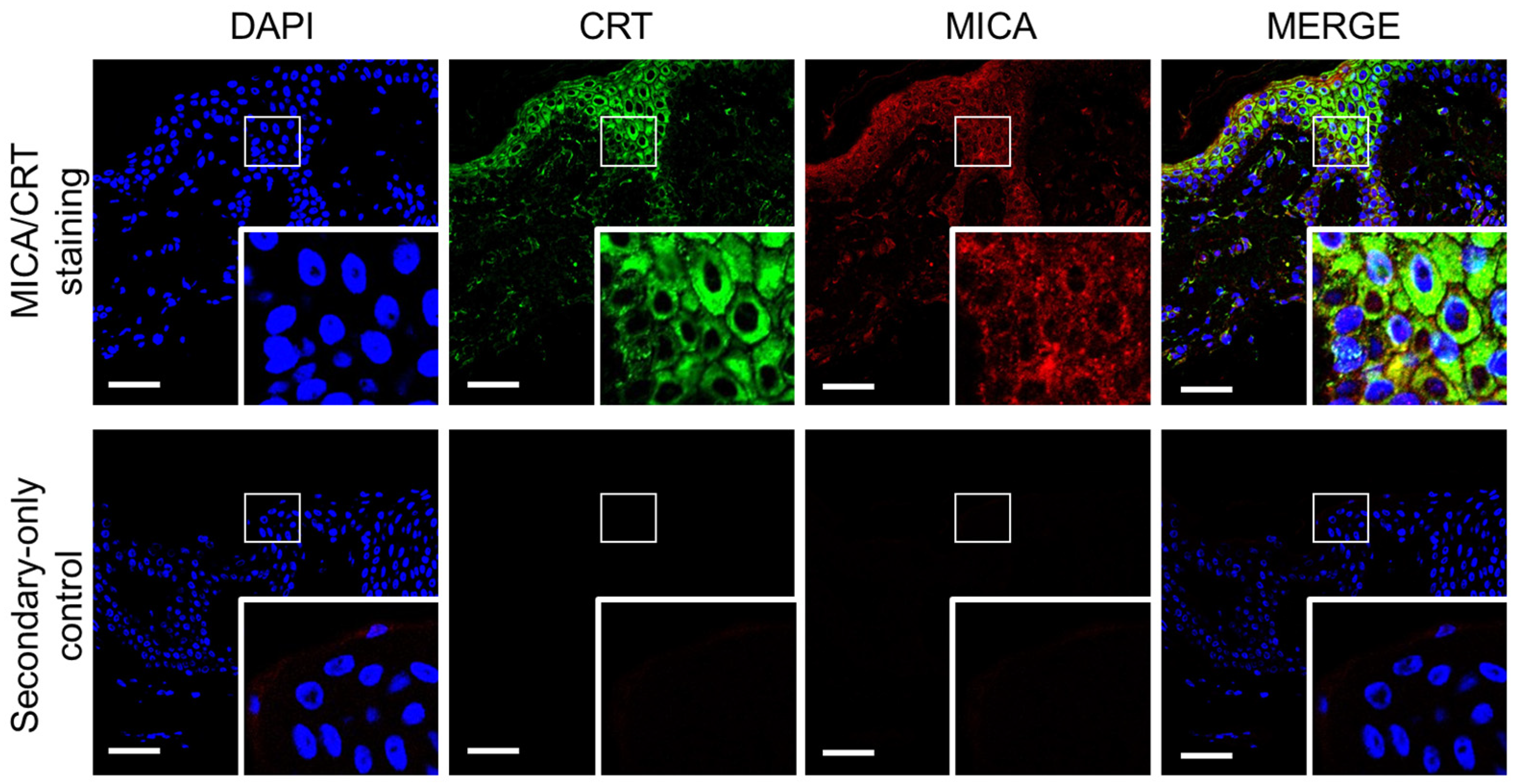
2.5. MICA and Calreticulin Partially Colocalize in Healthy Skin Tissue
2.6. In Silico Modeling Supports a Plausible Interaction Interface Between MICA and Calreticulin
3. Discussion
4. Materials and Methods
4.1. Patients and Samples
4.2. Cell Lines
4.3. Immunohistochemistry
4.4. Immunofluorescence
4.5. Immunoprecipitation
4.6. Western Blot
4.7. Recombinant Prokaryotic MICA and CRT
4.8. ELISA
4.9. Recombinant Eukaryotic MICA
4.10. Surface Plasmon Resonance (SPR)
4.11. In Silico Binding Prediction
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| MICA | Major histocompatibility complex class I chain-related gene A |
| MICB | Major histocompatibility complex class I chain-related gene B |
| CRT | Calreticulin |
| CNX | Calnexin |
| ER | Endoplasmic reticulum |
| NKG2D | Natural Killer Group 2D |
| MHC-I | Major Histocompatibility Complex class I |
| ERp57 | Endoplasmic Reticulum Protein 57 |
| HLA | Human Leukocyte Antigen |
| HE | Hematoxylin and eosin |
| SPR | Surface Plasmon Resonance |
| MD | Molecular Dynamics |
| MART-1 | Melanoma Antigen Recognized by T-cell 1 |
| NK cell | Natural Killer cell |
| HEK | Human Embryonic Kidney cell |
| BL | Melanoma cell line |
| ELISA | Enzyme—Linked Immunosorbent Assay |
| Endo-H | Endoglycosidase H |
| Rg | Radius of Gyration |
| RMSD | Root Mean Square Deviation |
| RMSF | Root Mean Square Fluctuation |
| DCCM | Dynamic Cross-Correlation Matrix |
| MM/GBSA | Molecular Mechanics/Generalized Born Surface Area |
| HRP | Horseradish Peroxidase |
| PDI | Protein Disulfide Isomerase |
| DAB | 3,3′-Diaminobenzidine |
| DAPI | 4′,6-Diamidino-2-Phenylindole |
| FCS | Fetal Bovine Serum |
| BSA | Bovine Serum Albumin |
| PBS | Phosphate-Buffered Saline |
| SDS-PAGE | Sodium Dodecyl Sulfate-Polyacrylamide Gel Electrophoresis |
| TMB | 3,3′,5,5′-Tetramethylbenzidine |
| Ni-NTA | Nickel-Nitrilotriacetic Acid |
| IgG | Immunoglobulin G |
| TIP3P | Transferable Intermolecular Potential 3-point water model |
| NPT | Constant Number of particles, Pressure and Temperature ensemble |
| PNGaseF | Peptide-N-Glycosidase F |
| PMSF | Phenylmethylsulphonyl fluoride |
| SNC | Normal Rabbit Serum |
| RS | Preimmune Rabbit serum |
| ECL | Enhanced Chemiluminescence |
References
- Neefjes, J.; Jongsma, M.L.; Paul, P.; Bakke, O. Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nat. Rev. Immunol. 2011, 11, 823–836. [Google Scholar] [CrossRef]
- Wearsch, P.A.; Cresswell, P. The quality control of MHC class I peptide loading. Curr. Opin. Cell Biol. 2008, 20, 624–631. [Google Scholar] [CrossRef]
- Caramelo, J.J.; Parodi, A.J. Getting in and out from calnexin/calreticulin cycles. J. Biol. Chem. 2008, 283, 10221–10225. [Google Scholar] [CrossRef]
- Groh, V.; Bahram, S.; Bauer, S.; Herman, A.; Beauchamp, M.; Spies, T. Cell stress-regulated human major histocompatibility complex class I gene expressed in gastrointestinal epithelium. Proc. Natl. Acad. Sci. USA 1996, 93, 12445–12450. [Google Scholar] [CrossRef]
- Eagle, R.A.; Trowsdale, J. Promiscuity and the single receptor: NKG2D. Nat. Rev. Immunol. 2007, 7, 737–744. [Google Scholar] [CrossRef]
- Li, P.; Willie, S.T.; Bauer, S.; Morris, D.L.; Spies, T.; Strong, R.K. Crystal structure of the MHC class I homolog MIC-A, a gammadelta T cell ligand. Immunity 1999, 10, 577–584. [Google Scholar] [CrossRef]
- Fernández-Messina, L.; Reyburn, H.T.; Valés-Gómez, M. Human NKG2D-ligands: Cell biology strategies to ensure immune recognition. Front. Immunol. 2012, 3, 299. [Google Scholar] [CrossRef]
- Toledo-Stuardo, K.; Ribeiro, C.H.; Canals, A.; Morales, M.; Gárate, V.; Rodríguez-Siza, J.; Tello, S.; Bustamante, M.; Armisen, R.; Matthies, D.J.; et al. Major Histocompatibility Complex Class I-Related Chain A (MICA) Allelic Variants Associate with Susceptibility and Prognosis of Gastric Cancer. Front. Immunol. 2021, 12, 645528. [Google Scholar] [CrossRef]
- Fuertes, M.B.; Girart, M.V.; Molinero, L.L.; Domaica, C.I.; Rossi, L.E.; Barrio, M.M.; Mordoh, J.; Rabinovich, G.A.; Zwirner, N.W. Intracellular retention of the NKG2D ligand MHC class I chain-related gene A in human melanomas confers immune privilege and prevents NK cell-mediated cytotoxicity. J. Immunol. 2008, 180, 4606–4614. [Google Scholar] [CrossRef]
- Andresen, L.; Skovbakke, S.L.; Persson, G.; Hagemann-Jensen, M.; Hansen, K.A.; Jensen, H.; Skov, S. 2-deoxy D-glucose prevents cell surface expression of NKG2D ligands through inhibition of N-linked glycosylation. J. Immunol. 2012, 188, 1847–1855. [Google Scholar] [CrossRef]
- Mellergaard, M.; Skovbakke, S.L.; Schneider, C.L.; Lauridsen, F.; Andresen, L.; Jensen, H.; Skov, S. N-glycosylation of asparagine 8 regulates surface expression of major histocompatibility complex class I chain-related protein A (MICA) alleles dependent on threonine 24. J. Biol. Chem. 2014, 289, 20078–20091. [Google Scholar] [CrossRef][Green Version]
- Jiang, Y.; Dey, S.; Matsunami, H. Calreticulin: Roles in cell-surface protein expression. Membranes 2014, 4, 630–641. [Google Scholar] [CrossRef]
- Wang, M.; Kaufman, R.J. Protein misfolding in the endoplasmic reticulum as a conduit to human disease. Nature 2016, 529, 326–335. [Google Scholar] [CrossRef]
- Wang, W.Y.; Tian, W.; Zhu, F.M.; Liu, X.X.; Li, L.X.; Wang, F. MICA, MICB Polymorphisms and Linkage Disequilibrium with HLA-B in a Chinese Mongolian Population. Scand. J. Immunol. 2016, 83, 456–462. [Google Scholar] [CrossRef]
- Wijeyesakere, S.J.; Gagnon, J.K.; Arora, K.; Brooks, C.L., 3rd; Raghavan, M. Regulation of calreticulin-major histocompatibility complex (MHC) class I interactions by ATP. Proc. Natl. Acad. Sci. USA 2015, 112, E5608–E5617. [Google Scholar] [CrossRef]
- Salih, H.R.; Rammensee, H.G.; Steinle, A. Cutting edge: Down-regulation of MICA on human tumors by proteolytic shedding. J. Immunol. 2002, 169, 4098–4102. [Google Scholar] [CrossRef]
- Ashiru, O.; Boutet, P.; Fernández-Messina, L.; Agüera-González, S.; Skepper, J.N.; Valés-Gómez, M.; Reyburn, H.T. Natural killer cell cytotoxicity is suppressed by exposure to the human NKG2D ligand MICA*008 that is shed by tumor cells in exosomes. Cancer Res. 2010, 70, 481–489. [Google Scholar] [CrossRef]
- Wang, B.; Wang, Q.; Wang, Z.; Jiang, J.; Yu, S.C.; Ping, Y.F.; Yang, J.; Xu, S.L.; Ye, X.Z.; Xu, C.; et al. Metastatic consequences of immune escape from NK cell cytotoxicity by human breast cancer stem cells. Cancer Res. 2014, 74, 5746–5757. [Google Scholar] [CrossRef]
- Yang, Y.; Ma, F.; Liu, Z.; Su, Q.; Liu, Y.; Liu, Z.; Li, Y. The ER-localized Ca2+-binding protein calreticulin couples ER stress to autophagy by associating with microtubule-associated protein 1A/1B light chain 3. J. Biol. Chem. 2019, 294, 772–782. [Google Scholar] [CrossRef]
- Halenius, A.; Gerke, C.; Hengel, H. Classical and non-classical MHC I molecule manipulation by human cytomegalovirus: So many targets—But how many arrows in the quiver? Cell. Mol. Immunol. 2015, 12, 139–153. [Google Scholar] [CrossRef]
- Chouquet, A.; Païdassi, H.; Ling, W.L.; Frachet, P.; Houen, G.; Arlaud, G.J.; Gaboriaud, C. X-ray structure of the human calreticulin globular domain reveals a peptide-binding area and suggests a multi-molecular mechanism. PLoS ONE 2011, 6, e17886. [Google Scholar] [CrossRef] [PubMed]
- Ghiran, I.; Klickstein, L.B.; Nicholson-Weller, A. Calreticulin is at the surface of circulating neutrophils and uses CD59 as an adaptor molecule. J. Biol. Chem. 2003, 278, 21024–21031. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Thangam, R.; You, S.H.; Sultonova, R.D.; Venu, A.; Min, J.J.; Hong, Y. Engineering Calreticulin-Targeting Monobodies to Detect Immunogenic Cell Death in Cancer Chemotherapy. Cancers 2021, 13, 2801. [Google Scholar] [CrossRef] [PubMed]
- Cosman, D.; Müllberg, J.; Sutherland, C.L.; Chin, W.; Armitage, R.; Fanslow, W.; Kubin, M.; Chalupny, N.J. ULBPs, novel MHC class I-related molecules, bind to CMV glycoprotein UL16 and stimulate NK cytotoxicity through the NKG2D receptor. Immunity 2001, 14, 123–133. [Google Scholar] [CrossRef]
- Duan, S.; Guo, W.; Xu, Z.; He, Y.; Liang, C.; Mo, Y.; Wang, Y.; Xiong, F.; Guo, C.; Li, Y.; et al. Natural killer group 2D receptor and its ligands in cancer immune escape. Mol. Cancer 2019, 18, 29. [Google Scholar] [CrossRef]
- Zuo, J.; Willcox, C.R.; Mohammed, F.; Davey, M.; Hunter, S.; Khan, K.; Antoun, A.; Katakia, P.; Croudace, J.; Inman, C.; et al. A disease-linked ULBP6 polymorphism inhibits NKG2D-mediated target cell killing by enhancing the stability of NKG2D ligand binding. Sci. Signal. 2017, 10, eaai8904. [Google Scholar] [CrossRef]
- Eagle, R.A.; Flack, G.; Warford, A.; Martínez-Borra, J.; Jafferji, I.; Traherne, J.A.; Ohashi, M.; Boyle, L.H.; Barrow, A.D.; Caillat-Zucman, S.; et al. Cellular expression, trafficking, and function of two isoforms of human ULBP5/RAET1G. PLoS ONE 2009, 4, e4503, Erratum in PLoS ONE 2009, 4. https://doi.org/10.1371/annotation/8501cd33-6c9e-437c-9ea1-f96fbb4140d6. [Google Scholar] [CrossRef]
- Serrano, A.E.; Menares-Castillo, E.; Garrido-Tapia, M.; Ribeiro, C.H.; Hernández, C.J.; Mendoza-Naranjo, A.; Gatica-Andrades, M.; Valenzuela-Diaz, R.; Zúñiga, R.; López, M.N.; et al. Interleukin 10 decreases MICA expression on melanoma cell surface. Immunol. Cell Biol. 2011, 89, 447–457. [Google Scholar] [CrossRef]
- Yamane, K.; Yamaguchi, K.; Teruya, Y.; Miyake, N.; Nakayama, Y.; Nonaka, T.; Chikumi, H.; Yamasaki, A. ULBP2 Promotes Tumor Progression by Suppressing NKG2D-Mediated Anti-Tumor Immunity. Int. J. Mol. Sci. 2025, 26, 2950. [Google Scholar] [CrossRef]
- Campos-Silva, C.; Kramer, M.K.; Valés-Gómez, M. NKG2D-ligands: Putting everything under the same umbrella can be misleading. Hla 2018, 91, 489–500. [Google Scholar] [CrossRef]
- Kobata, A.; Amano, J. Altered glycosylation of proteins produced by malignant cells, and application for the diagnosis and immunotherapy of tumours. Immunol. Cell Biol. 2005, 83, 429–439. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Wu, Z.; Zhao, S.; Xiang, R. Chemical chaperones reduce ER stress and adipose tissue inflammation in high fat diet-induced mouse model of obesity. Sci. Rep. 2016, 6, 27486. [Google Scholar] [CrossRef] [PubMed]
- Salazar-Onfray, F.; Nakazawa, T.; Chhajlani, V.; Petersson, M.; Kärre, K.; Masucci, G.; Celis, E.; Sette, A.; Southwood, S.; Appella, E.; et al. Synthetic peptides derived from the melanocyte-stimulating hormone receptor MC1R can stimulate HLA-A2-restricted cytotoxic T lymphocytes that recognize naturally processed peptides on human melanoma cells. Cancer Res. 1997, 57, 4348–4355. [Google Scholar]
- Salazar-Onfray, F.; López, M.; Lundqvist, A.; Aguirre, A.; Escobar, A.; Serrano, A.; Korenblit, C.; Petersson, M.; Chhajlani, V.; Larsson, O.; et al. Tissue distribution and differential expression of melanocortin 1 receptor, a malignant melanoma marker. Br. J. Cancer 2002, 87, 414–422. [Google Scholar] [CrossRef]
- Sotelo, P.; Collazo, N.; Zuñiga, R.; Gutiérrez-González, M.; Catalán, D.; Ribeiro, C.H.; Aguillón, J.C.; Molina, M.C. An efficient method for variable region assembly in the construction of scFv phage display libraries using independent strand amplification. mAbs 2012, 4, 542–550. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ribeiro, C.H.; López, N.C.; Ramírez, G.A.; Valck, C.E.; Molina, M.C.; Aguilar, L.; Rodríguez, M.; Maldonado, I.; Martínez, R.; González, C.; et al. Trypanosoma cruzi calreticulin: A possible role in Chagas’ disease autoimmunity. Mol. Immunol. 2009, 46, 1092–1099. [Google Scholar] [CrossRef]
- Aricescu, A.R.; Lu, W.; Jones, E.Y. A time- and cost-efficient system for high-level protein production in mammalian cells. Acta Crystallogr. Sect. D Biol. Crystallogr. 2006, 62, 1243–1250. [Google Scholar] [CrossRef]
- Schwede, T.; Kopp, J.; Guex, N.; Peitsch, M.C. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res. 2003, 31, 3381–3385. [Google Scholar] [CrossRef]
- Domnick, A.; Winter, C.; Sušac, L.; Hennecke, L.; Hensen, M.; Zitzmann, N.; Trowitzsch, S.; Thomas, C.; Tampé, R. Molecular basis of MHC I quality control in the peptide loading complex. Nat. Commun. 2022, 13, 4701. [Google Scholar] [CrossRef]
- Li, P.; Morris, D.L.; Willcox, B.E.; Steinle, A.; Spies, T.; Strong, R.K. Complex structure of the activating immunoreceptor NKG2D and its MHC class I-like ligand MICA. Nat. Immunol. 2001, 2, 443–451. [Google Scholar] [CrossRef]
- Brenke, R.; Hall, D.R.; Chuang, G.Y.; Comeau, S.R.; Bohnuud, T.; Beglov, D.; Schueler-Furman, O.; Vajda, S.; Kozakov, D. Application of asymmetric statistical potentials to antibody-protein docking. Bioinformatics 2012, 28, 2608–2614. [Google Scholar] [CrossRef]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef]
- Tian, C.; Kasavajhala, K.; Belfon, K.A.A.; Raguette, L.; Huang, H.; Migues, A.N.; Bickel, J.; Wang, Y.; Pincay, J.; Wu, Q.; et al. ff19SB: Amino-Acid-Specific Protein Backbone Parameters Trained against Quantum Mechanics Energy Surfaces in Solution. J. Chem. Theory Comput. 2020, 16, 528–552. [Google Scholar] [CrossRef]
- Case, D.A.; Cheatham, T.E., 3rd; Darden, T.; Gohlke, H.; Luo, R.; Merz, K.M., Jr.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R.J. The Amber biomolecular simulation programs. J. Comput. Chem. 2005, 26, 1668–1688. [Google Scholar] [CrossRef]
- Miller, B.R., 3rd; McGee, T.D., Jr.; Swails, J.M.; Homeyer, N.; Gohlke, H.; Roitberg, A.E. MMPBSA.py: An Efficient Program for End-State Free Energy Calculations. J. Chem. Theory Comput. 2012, 8, 3314–3321. [Google Scholar] [CrossRef]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
González-Herrera, F.; Toledo-Stuardo, K.; Serrano, A.E.; Gatica-Andrades, M.; Matthies, D.J.; López, V.; Aguayo, I.; Indo, S.; Garrido, M.J.; Guerra, Y.; et al. Calreticulin-Mediated Quality Control of the Non-Classical MHC-I Molecule MICA: Implications for Immune Surveillance. Int. J. Mol. Sci. 2026, 27, 1310. https://doi.org/10.3390/ijms27031310
González-Herrera F, Toledo-Stuardo K, Serrano AE, Gatica-Andrades M, Matthies DJ, López V, Aguayo I, Indo S, Garrido MJ, Guerra Y, et al. Calreticulin-Mediated Quality Control of the Non-Classical MHC-I Molecule MICA: Implications for Immune Surveillance. International Journal of Molecular Sciences. 2026; 27(3):1310. https://doi.org/10.3390/ijms27031310
Chicago/Turabian StyleGonzález-Herrera, Fabiola, Karen Toledo-Stuardo, Antonio E. Serrano, Marcela Gatica-Andrades, Douglas J. Matthies, Valentina López, Ignacio Aguayo, Sebastián Indo, María José Garrido, Yuneisy Guerra, and et al. 2026. "Calreticulin-Mediated Quality Control of the Non-Classical MHC-I Molecule MICA: Implications for Immune Surveillance" International Journal of Molecular Sciences 27, no. 3: 1310. https://doi.org/10.3390/ijms27031310
APA StyleGonzález-Herrera, F., Toledo-Stuardo, K., Serrano, A. E., Gatica-Andrades, M., Matthies, D. J., López, V., Aguayo, I., Indo, S., Garrido, M. J., Guerra, Y., Tello, S., Campos, I., Salazar-Onfray, F., Zapata-Torres, G., Ribeiro, C. H., & Molina, M. C. (2026). Calreticulin-Mediated Quality Control of the Non-Classical MHC-I Molecule MICA: Implications for Immune Surveillance. International Journal of Molecular Sciences, 27(3), 1310. https://doi.org/10.3390/ijms27031310






