Identification of a 13-Gene Immune Signature in Liver Fibrosis Reveals GABRE as a Novel Candidate Biomarker
Abstract
1. Introduction
2. Results
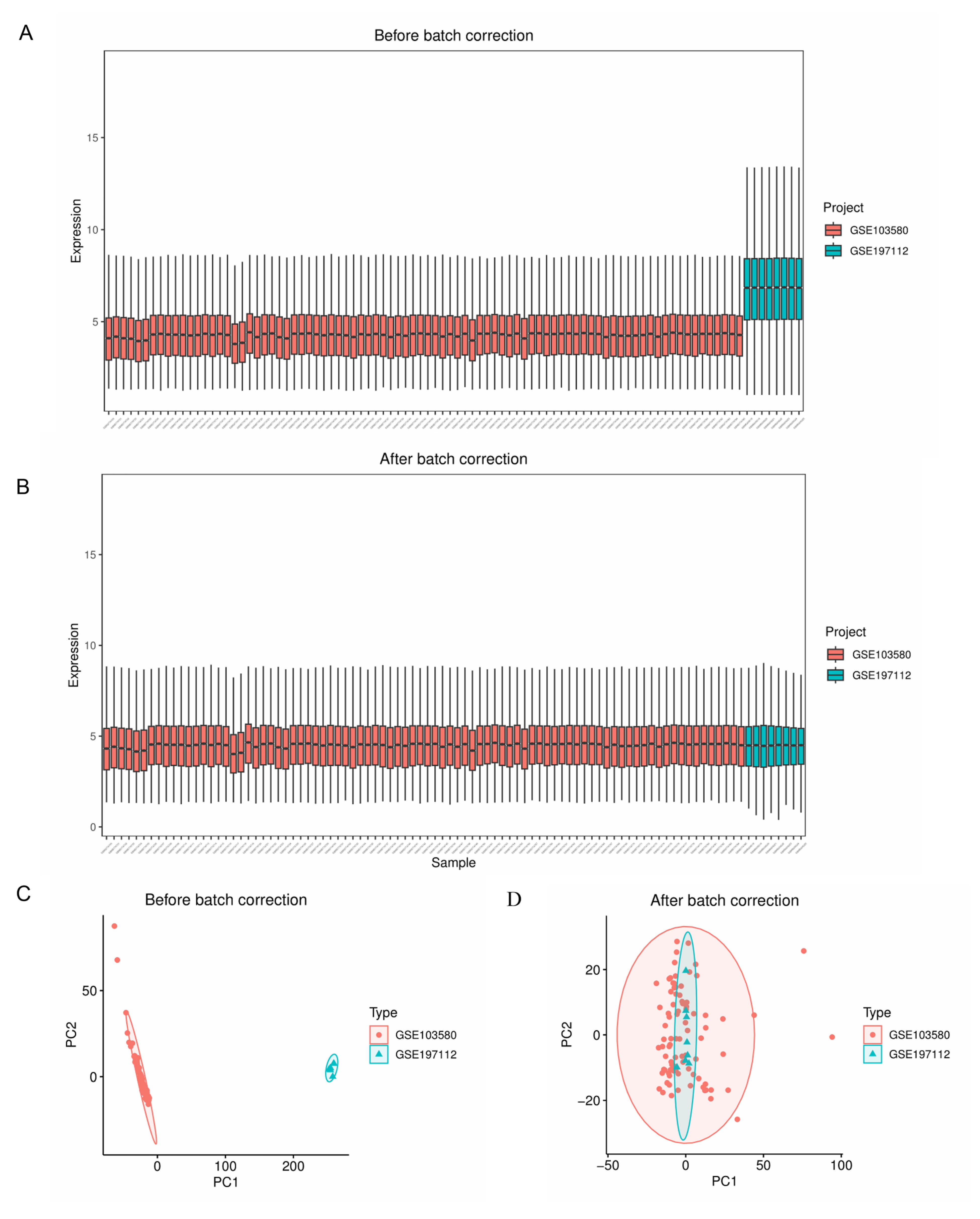
2.1. Data Preprocessing
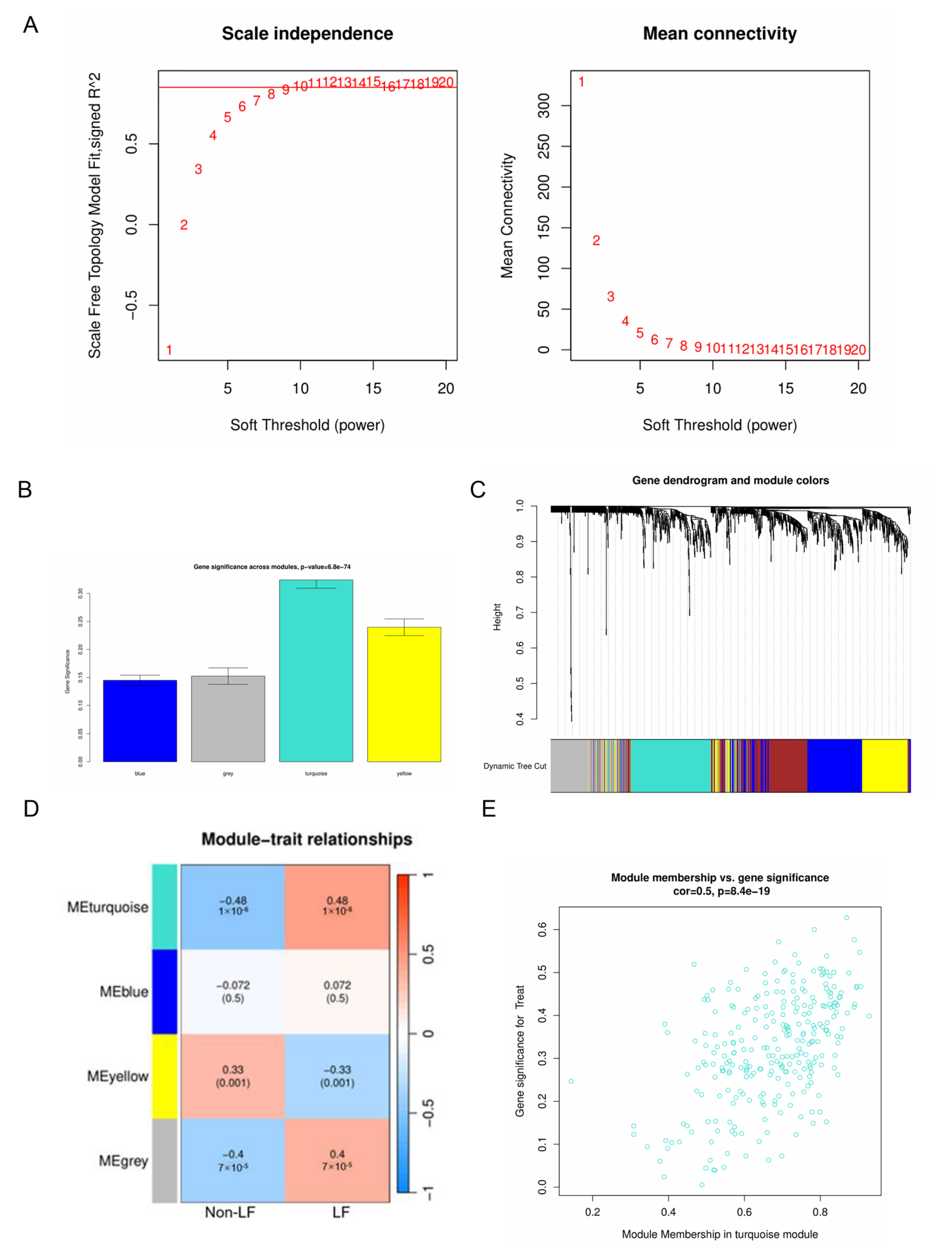
2.2. Hub Gene Screening Using Weighted Gene Co-Expression Network Analysis (WGCNA)
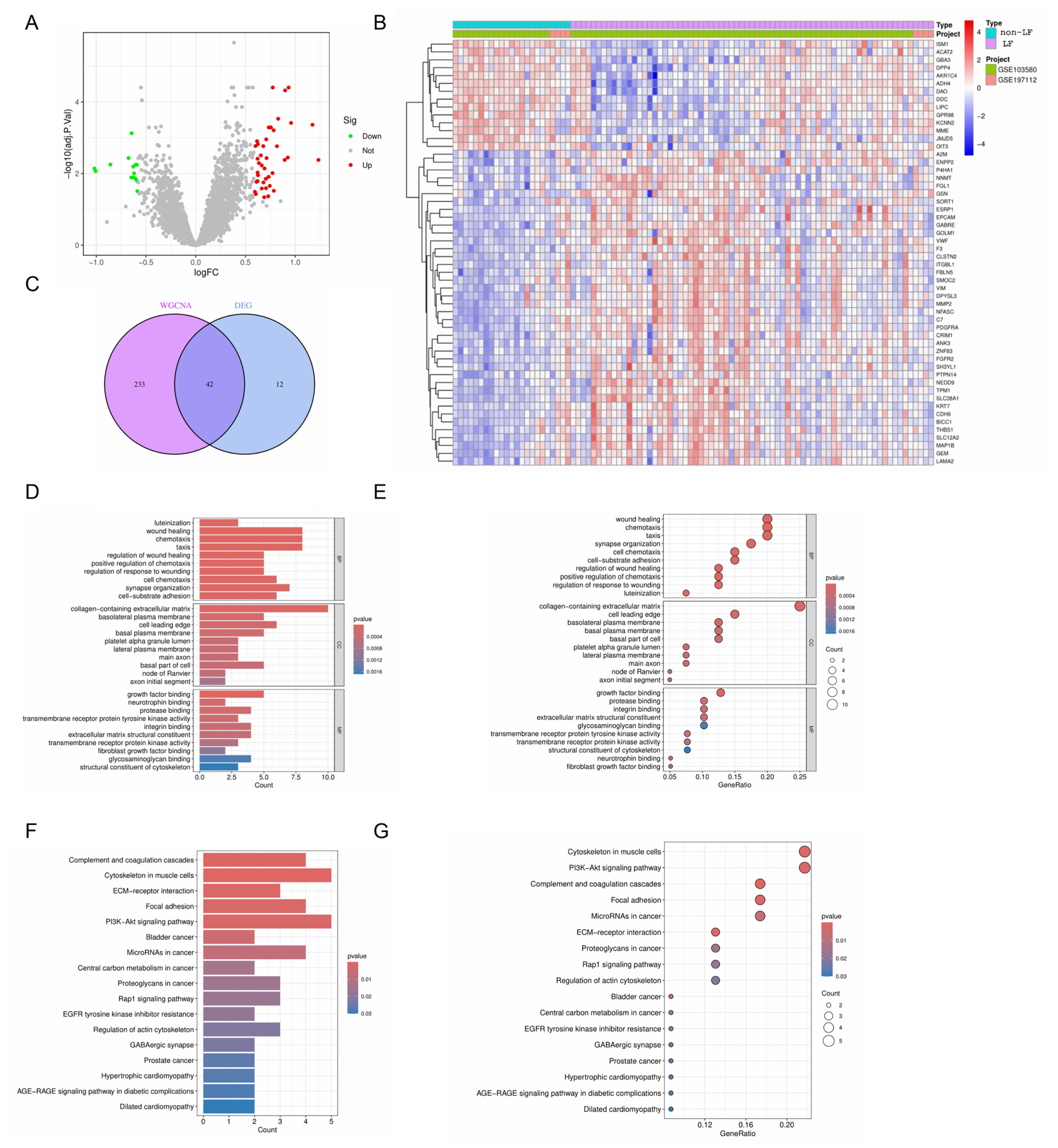
2.3. Identification and Integrative Analysis of DEGs and Intersecting Genes
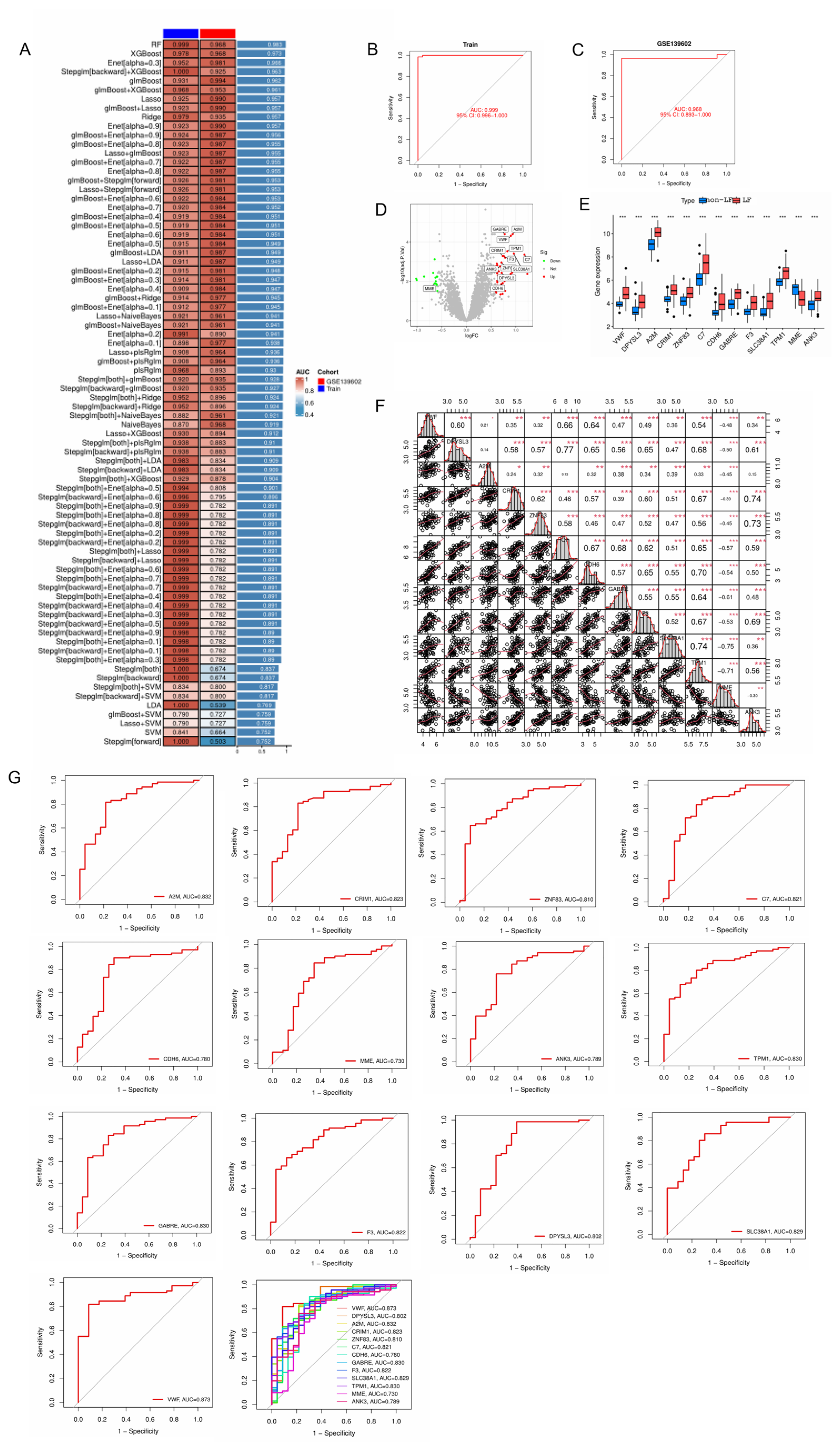
2.4. Screening the Signature Genes of LF Using Machine Learning
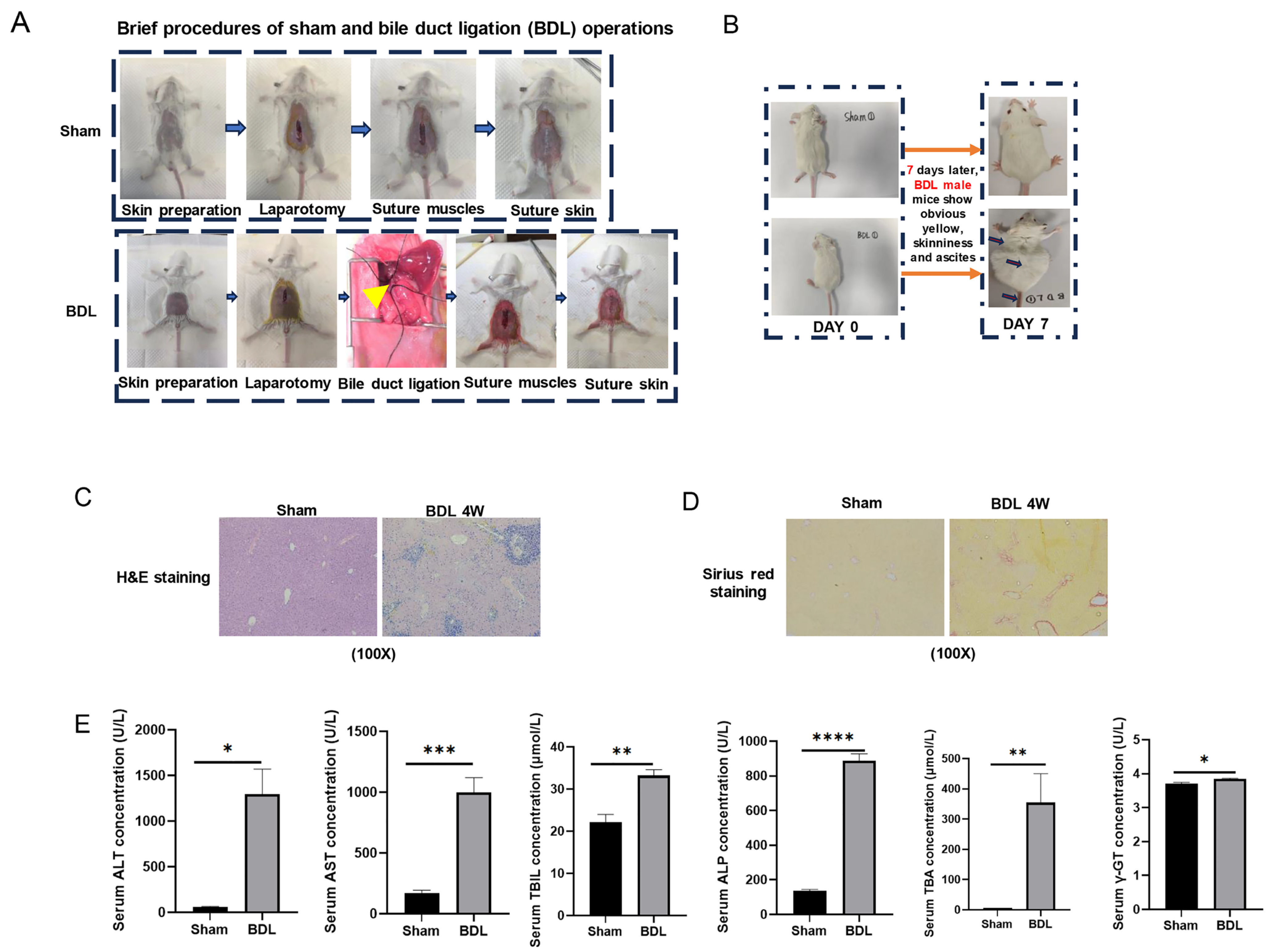
2.5. Establishment of an External Liver Fibrosis Model
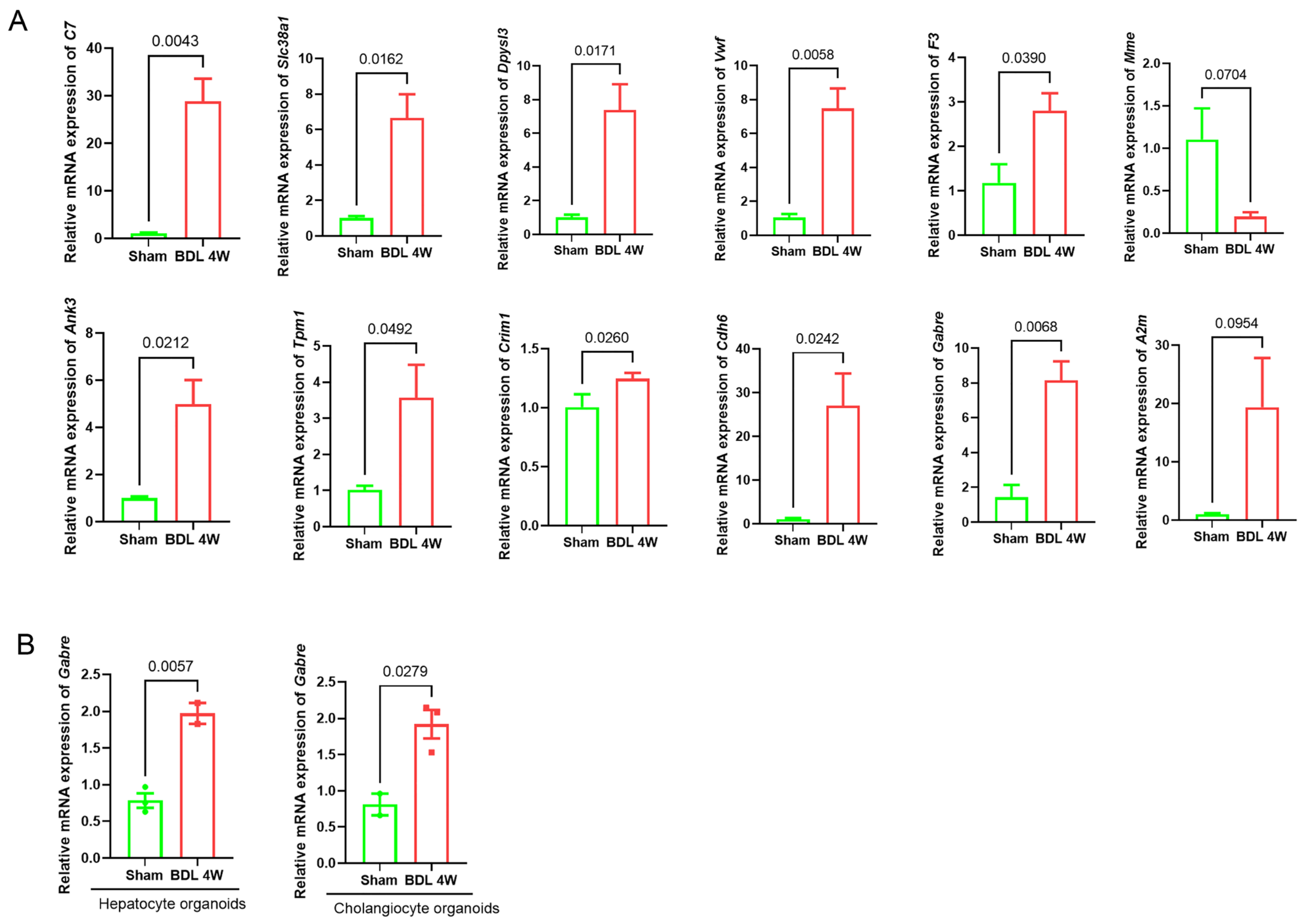
2.6. Validation of Signature Genes in an LF Mouse Model and Gabre mRNA Expression in Hepatocyte Organoids and Cholangiocyte Organoids
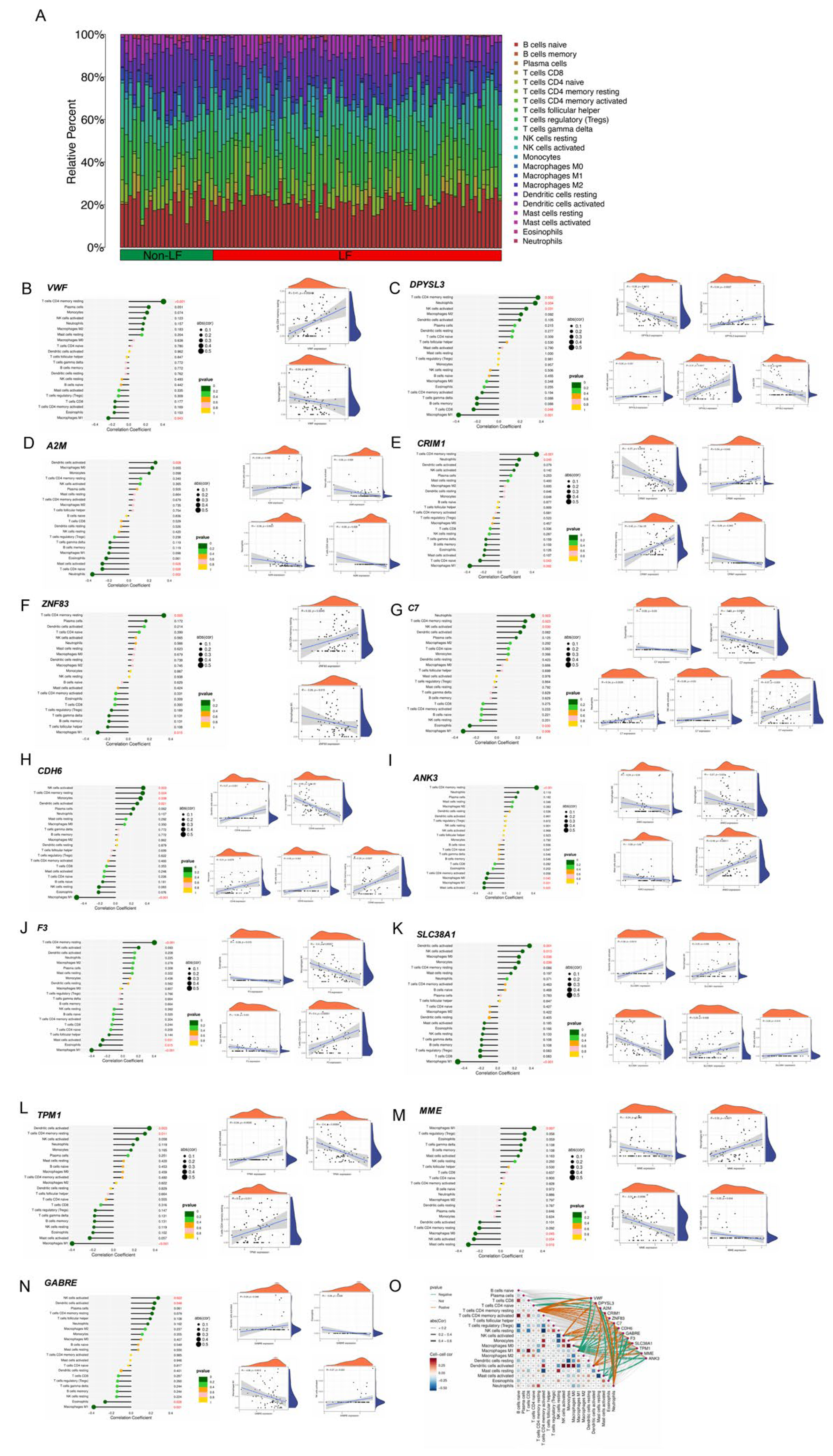
2.7. Immune Landscape and Correlation
3. Discussion
4. Materials and Methods
4.1. Data Collection
4.2. Source of Data
4.3. Identification of Differentially Expressed Genes (DEGs)
4.4. Construction of the Co-Expression Network and Hub Module Identification Using WGCNA
4.5. Identification and Enrichment Analysis of Intersecting Genes
4.6. Screening of Candidate Diagnostic Biomarkers Using Machine Learning
4.7. Development of the Bile Duct Ligation (BDL) Mouse Model
4.8. Serological Testing
4.9. Hematoxylin/Eosin (H&E) and Sirius Red Staining
4.10. Extraction, Culture, and Identification of Hepatocyte Organoids and Cholangiocyte Organoids from BDL Mice
4.11. RNA Isolation and Quantitative Real-Time Reverse Transcriptase Polymerase Chain Reaction (qRT-PCR)
4.12. Analysis of the Immune Landscape and Gene Correlation
4.13. Statistical Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| A2M | Alpha-2-Macroglobulin |
| ANK3 | Ankyrin 3 |
| AUC | Area Under the Curve |
| BDL | Bile Duct Ligation |
| CDH6 | Cadherin 6 |
| CRIM1 | Cysteine-Rich Transmembrane BMP Regulator 1 |
| C7 | Complement Component 7 |
| DEGs | Differentially Expressed Genes |
| DPYSL3 | Dihydropyrimidinase-Like 3 |
| ECM | Extracellular Matrix |
| F3 | Coagulation Factor III (Tissue Factor) |
| GABA | Gamma-Aminobutyric Acid |
| GABRE | Gamma-Aminobutyric Acid Type A Receptor Subunit Epsilon |
| GEO | Gene Expression Omnibus |
| GO | Gene Ontology |
| H&E | Hematoxylin/Eosin |
| HCC | Hepatocellular Carcinoma |
| HSCs | Hepatic Stellate Cells |
| IFN-γ | Interferon Gamma |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| LASSO | Least Absolute Shrinkage and Selection Operator |
| LF | Liver Fibrosis |
| MME | Membrane Metalloendopeptidase |
| NK | Natural Killer |
| NASH | Non-alcoholic Steatohepatitis |
| NAFLD | Non-alcoholic Fatty Liver Disease |
| PPI | Protein–Protein Interaction |
| PI3K-Akt | Phosphoinositide 3-Kinase-Protein Kinase B |
| qRT-PCR | Quantitative Real-Time Reverse Transcriptase Polymerase Chain Reaction |
| RF | Random Forest |
| ROC | Receiver Operating Characteristic |
| SLC38A1 | Solute Carrier Family 38 Member 1 |
| SEM | Standard Error of the Mean; Tregs: Regulatory T Cells |
| TNF-α | Tumor Necrosis Factor Alpha |
| TPM1 | Tropomyosin 1 |
| VWF | Von Willebrand Factor |
| WGCNA | Weighted Gene Co-expression Network Analysis |
| XGBoost | eXtreme Gradient Boosting |
| ZNF83 | Zinc Finger Protein 83 |
References
- Roehlen, N.; Crouchet, E.; Baumert, T.F. Liver Fibrosis: Mechanistic Concepts and Therapeutic Perspectives. Cells 2020, 9, 875. [Google Scholar] [CrossRef] [PubMed]
- Cogliati, B.; Yashaswini, C.N.; Wang, S.; Sia, D.; Friedman, S.L. Friend or foe? The elusive role of hepatic stellate cells in liver cancer. Nat. Rev. Gastroenterol. Hepatol. 2023, 20, 647–661. [Google Scholar] [CrossRef]
- Younossi, Z.M.; Wong, G.; Anstee, Q.M.; Henry, L. The Global Burden of Liver Disease. Clin. Gastroenterol. Hepatol. 2023, 21, 1978–1991. [Google Scholar] [CrossRef]
- Huang, D.Q.; El-Serag, H.B.; Loomba, R. Global epidemiology of NAFLD-related HCC: Trends, predictions, risk factors and prevention. Nat. Rev. Gastroenterol. Hepatol. 2021, 18, 223–238. [Google Scholar] [CrossRef]
- Harrison, S.A.; Ratziu, V.; Boursier, J.; Francque, S.; Bedossa, P.; Majd, Z.; Cordonnier, G.; Sudrik, F.B.; Darteil, R.; Liebe, R.; et al. A blood-based biomarker panel (NIS4) for non-invasive diagnosis of non-alcoholic steatohepatitis and liver fibrosis: A prospective derivation and global validation study. Lancet Gastroenterol. Hepatol. 2020, 5, 970–985. [Google Scholar] [CrossRef]
- Elhence, A.; Shalimar. Von Willebrand Factor as a Biomarker for Liver Disease—An Update. J. Clin. Exp. Hepatol. 2023, 13, 1047–1060. [Google Scholar] [CrossRef]
- Yang, Y.; Deng, X.; Chen, X.; Chen, S.; Song, L.; Meng, M.; Han, Q.; Imani, S.; Li, S.; Zhong, Z.; et al. Landscape of active enhancers developed de novo in cirrhosis and conserved in hepatocellular carcinoma. Am. J. Cancer Res. 2020, 10, 3157–3178. [Google Scholar]
- Alves-Ferreira, M.; Quintas, M.; Sequeiros, J.; Sousa, A.; Pereira-Monteiro, J.; Alonso, I.; Neto, J.L.; Lemos, C. A genetic interaction of NRXN2 with GABRE, SYT1 and CASK in migraine patients: A case-control study. J. Headache Pain. 2021, 22, 57. [Google Scholar] [CrossRef]
- Chen, X.; Zhou, Y.N.; Lu, X.Z.; Li, R.J.; Xiong, Y.F.; Sheng, X.; Zhu, W.W. Cognitive dysfunction in schizophrenia patients caused by down-regulation of gamma-aminobutyric acid receptor subunits. World J. Psychiatry 2024, 14, 784–793. [Google Scholar] [CrossRef]
- Bai, Y.M.; Liang, S.; Zhou, B. Revealing immune infiltrate characteristics and potential immune-related genes in hepatic fibrosis: Based on bioinformatics, transcriptomics and q-PCR experiments. Front. Immunol. 2023, 14, 1133543. [Google Scholar] [CrossRef]
- Yang, Y.; Hua, Y.; Zheng, H.; Jia, R.; Ye, Z.; Su, G.; Gu, Y.; Zhan, K.; Tang, K.; Qi, S.; et al. Biomarkers prediction and immune landscape in ulcerative colitis: Findings based on bioinformatics and machine learning. Comput. Biol. Med. 2024, 168, 107778. [Google Scholar] [CrossRef]
- Zhang, S.; Jiang, C.; Jiang, L.; Chen, H.; Huang, J.; Zhang, J.; Wang, R.; Chi, H.; Yang, G.; Tian, G. Uncovering the immune microenvironment and molecular subtypes of hepatitis B-related liver cirrhosis and developing stable a diagnostic differential model by machine learning and artificial neural networks. Front. Mol. Biosci. 2023, 10, 1275897. [Google Scholar] [CrossRef]
- Xia, P.; Ma, H.; Chen, J.; Liu, Y.; Cui, X.; Wang, C.; Zong, S.; Wang, L.; Liu, Y.; Lu, Z. Differential expression of pyroptosis-related genes in the hippocampus of patients with Alzheimer’s disease. BMC Med. Genom. 2023, 16, 56. [Google Scholar] [CrossRef] [PubMed]
- Zhuo, S.; Jiang, L.; Cai, X. How to define setup channels for an electrophysiological recording system in left bundle branch pacing. Kardiol. Pol. 2023, 81, 284–286. [Google Scholar] [CrossRef] [PubMed]
- Joshi, N.; Kopec, A.K.; Ray, J.L.; Cline-Fedewa, H.; Groeneveld, D.J.; Lisman, T.; Luyendyk, J.P. Von Willebrand factor deficiency reduces liver fibrosis in mice. Toxicol. Appl. Pharm. 2017, 328, 54–59. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Takaya, H.; Kawaratani, H.; Tsuji, Y.; Nakanishi, K.; Saikawa, S.; Sato, S.; Sawada, Y.; Kaji, K.; Okura, Y.; Shimozato, N.; et al. von Willebrand factor is a useful biomarker for liver fibrosis and prediction of hepatocellular carcinoma development in patients with hepatitis B and C. United Eur. Gastroenterol. J. 2018, 6, 1401–1409. [Google Scholar] [CrossRef]
- Pitekova, B.; Kupcova, V.; Uhlikova, E.; Mojto, V.; Turecky, L. Alpha-2-macroglobulin and hyaluronic acid as fibromarkers in patients with chronic hepatitis C. Bratisl. Med. J. 2017, 118, 658–661. [Google Scholar] [CrossRef]
- Tan, Z.; Sun, H.B.; Xue, T.X.; Gan, C.L.; Liu, H.Y.; Xie, Y.T.; Yao, Y.Q.; Ye, T.H. Liver Fibrosis: Therapeutic Targets and Advances in Drug Therapy. Front. Cell Dev. Biol. 2021, 9, 730176. [Google Scholar] [CrossRef]
- Sanchez, J.I.; Parra, E.R.; Jiao, J.J.; Soto, L.M.S.; Ledesma, D.A.; Saldarriaga, O.A.; Stevenson, H.L.; Beretta, L. Cellular and Molecular Mechanisms of Liver Fibrosis in Patients with NAFLD. Cancers 2023, 15, 2871. [Google Scholar] [CrossRef]
- Li, J.; Yuan, Y.; Fu, Q.; Chen, M.; Liang, H.; Chen, X.; Long, X.; Zhang, B.; Zhao, J.; Chen, Q. Novel insights into the role of immunomodulatory extracellular vesicles in the pathogenesis of liver fibrosis. Biomark. Res. 2024, 12, 119. [Google Scholar] [CrossRef]
- Oya, H.; Kanda, M.; Sugimoto, H.; Shimizu, D.; Takami, H.; Hibino, S.; Hashimoto, R.; Okamura, Y.; Yamada, S.; Fujii, T.; et al. Dihydropyrimidinase-like 3 is a putative hepatocellular carcinoma tumor suppressor. J. Gastroenterol. 2015, 50, 590–600. [Google Scholar] [CrossRef]
- Hotta, K.; Kikuchi, M.; Kitamoto, T.; Kitamoto, A.; Ogawa, Y.; Honda, Y.; Kessoku, T.; Kobayashi, K.; Yoneda, M.; Imajo, K.; et al. Identification of core gene networks and hub genes associated with progression of non-alcoholic fatty liver disease by RNA sequencing. Hepatol. Res. 2017, 47, 1445–1458. [Google Scholar] [CrossRef]
- Yang, H.; Atak, D.; Yuan, M.; Li, M.; Altay, O.; Demirtas, E.; Peltek, I.B.; Ulukan, B.; Yigit, B.; Sipahioglu, T.; et al. Integrative proteo-transcriptomic characterization of advanced fibrosis in chronic liver disease across etiologies. Cell Rep. Med. 2025, 6, 101935. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Zhou, J. Identification of key genes and functional enrichment analysis of liver fibrosis in nonalcoholic fatty liver disease through weighted gene co-expression network analysis. Genom. Inform. 2023, 21, e45. [Google Scholar] [CrossRef] [PubMed]
- Hou, W.; Janech, M.G.; Sobolesky, P.M.; Bland, A.M.; Samsuddin, S.; Alazawi, W.; Syn, W.K. Proteomic screening of plasma identifies potential noninvasive biomarkers associated with significant/advanced fibrosis in patients with nonalcoholic fatty liver disease. Biosci. Rep. 2020, 40, BSR20190395. [Google Scholar] [CrossRef] [PubMed]
- de Lima, R.E.; de Holanda Martins, C.M.; do Carmo, R.F.; Aroucha, D.; Pereira, L.; Vasconcelos, L.R.S.; Moura, P. Two sides of a coin: GG genotype of C7 provides protection against fibrosis severity while showing a higher risk for hepatocellular carcinoma in patients with hepatitis C. Hum. Immunol. 2018, 79, 702–707. [Google Scholar] [CrossRef]
- Park, S.R.; Kim, H.J.; Yang, S.R.; Park, C.H.; Lee, H.Y.; Hong, I.S. A novel endogenous damage signal, glycyl tRNA synthetase, activates multiple beneficial functions of mesenchymal stem cells. Cell Death Differ. 2018, 25, 2023–2036. [Google Scholar] [CrossRef]
- MacLeod, M.K.; Kappler, J.W.; Marrack, P. Memory CD4 T cells: Generation, reactivation and re-assignment. Immunology 2010, 130, 10–15. [Google Scholar] [CrossRef]
- Chowdhury, M.I.H.; Nishioka, T.; Mishima, N.; Ohtsuka, T.; Kaibuchi, K.; Tsuboi, D. Prickle2 and Igsf9b Coordinately Regulate the Cytoarchitecture of the Axon Initial Segment. Cell Struct. Funct. 2020, 45, 143–154. [Google Scholar] [CrossRef]
- Garza, J.C.; Qi, X.; Gjeluci, K.; Leussis, M.P.; Basu, H.; Reis, S.A.; Zhao, W.N.; Piguel, N.H.; Penzes, P.; Haggarty, S.J.; et al. Disruption of the psychiatric risk gene Ankyrin 3 enhances microtubule dynamics through GSK3/CRMP2 signaling. Transl. Psychiatry 2018, 8, 135. [Google Scholar] [CrossRef]
- Di, Z.; Muyun, W.; Luan, C.; Hao, W.U.; Ting, W.; Zhiruo, Z.; Ying, Z.; Juan, Y.U.; Jinming, H.; Jinhang, Z.; et al. Drug response biomarkers of Pien Tze Huang treatment for hepatic fibrosis induced by carbon tetrachloride. J. Tradit. Chin. Med. 2022, 42, 530–538. [Google Scholar] [CrossRef] [PubMed]
- Yan, R.; Cai, H.; Zhou, X.; Bao, G.; Bai, Z.; Ge, R.L. Hypoxia-inducible factor-2alpha promotes fibrosis in non-alcoholic fatty liver disease by enhancing glutamine catabolism and inhibiting yes-associated protein phosphorylation in hepatic stellate cells. Front. Endocrinol. 2024, 15, 1344971. [Google Scholar] [CrossRef] [PubMed]
- Tian, Z.; Zhao, J.; Wang, Y. The prognostic value of TPM1-4 in hepatocellular carcinoma. Cancer Med. 2022, 11, 433–446. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Martin, E.; Esguevillas, G.; Serrador, M.; Alonso-Navarro, H.; Navacerrada, F.; Amo, G.; Garcia-Albea, E.; Agundez, J.A.G.; Jimenez-Jimenez, F.J. Gamma-aminobutyric acid (GABA) receptors GABRA4, GABRE, and GABRQ gene polymorphisms and risk for migraine. J. Neural Transm. 2018, 125, 689–698. [Google Scholar] [CrossRef]
- Wang, Z.; Li, L.; Li, M.; Lu, Z.; Qin, L.; Naumann, R.K.; Wang, H. Chemogenetic Modulation of Preoptic Gabre Neurons Decreases Body Temperature and Heart Rate. Int. J. Mol. Sci. 2024, 25, 13061. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, H.; Li, G.; Liu, Y.; Wang, C.; He, D. Exploring the genetic basis of fatty liver development in geese. Sci. Rep. 2020, 10, 14279. [Google Scholar] [CrossRef]
- Pan, J.; Tian, Y.; Hu, F.; Xu, J.; Tan, N.; Han, Y.; Kang, Q.; Chen, H.; Yang, Y.; Xu, X. Exploration of immune infiltration and feature genes in viral hepatitis-associated liver fibrosis using transcriptome data. Ann. Transl. Med. 2022, 10, 1051. [Google Scholar] [CrossRef]
- Wallace, H.L.; Russell, R.S. Inflammatory Consequences: Hepatitis C Virus-Induced Inflammasome Activation and Pyroptosis. Viral Immunol. 2024, 37, 126–138. [Google Scholar] [CrossRef]
- Ha, Y.; Jeong, I.; Kim, T.H. Alcohol-Related Liver Disease: An Overview on Pathophysiology, Diagnosis and Therapeutic Perspectives. Biomedicines 2022, 10, 2530. [Google Scholar] [CrossRef]
- Badmus, O.O.; Hillhouse, S.A.; Anderson, C.D.; Hinds, T.D.; Stec, D.E. Molecular mechanisms of metabolic associated fatty liver disease (MAFLD): Functional analysis of lipid metabolism pathways. Clin. Sci. 2022, 136, 1347–1366. [Google Scholar] [CrossRef]
- Cheng, F.; Li, Q.; Wang, J.; Zeng, F.; Wang, K.; Zhang, Y. Identification of Differential Intestinal Mucosa Transcriptomic Biomarkers for Ulcerative Colitis by Bioinformatics Analysis. Dis. Markers 2020, 2020, 8876565. [Google Scholar] [CrossRef]
- AmeliMojarad, M.; AmeliMojarad, M.; Cui, X. Weighted gene co-expression network analysis identified GBP2 connected to PPARalpha activity and liver cancer. Sci. Rep. 2024, 14, 20745. [Google Scholar] [CrossRef]
- Abudereheman, M.; Lian, Z.; Ainitu, B. Weighted gene co-expression network analysis and whole genome sequencing identify potential lung cancer biomarkers. Front. Oncol. 2024, 14, 1355527. [Google Scholar] [CrossRef]
- Handelman, G.S.; Kok, H.K.; Chandra, R.V.; Razavi, A.H.; Lee, M.J.; Asadi, H. eDoctor: Machine learning and the future of medicine. J. Intern. Med. 2018, 284, 603–619. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Liu, Z.; Liang, R.; Wang, W.; Zhu, R.; Li, J.; Xing, Z.; Weng, S.; Han, X.; Sun, Y.L. Comprehensive machine-learning survival framework develops a consensus model in large-scale multicenter cohorts for pancreatic cancer. Elife 2022, 11, e80150. [Google Scholar] [CrossRef] [PubMed]
- Tag, C.G.; Sauer-Lehnen, S.; Weiskirchen, S.; Borkham-Kamphorst, E.; Tolba, R.H.; Tacke, F.; Weiskirchen, R. Bile duct ligation in mice: Induction of inflammatory liver injury and fibrosis by obstructive cholestasis. J. Vis. Exp. 2015, 10, 52438. [Google Scholar] [CrossRef]
- Feldman, A.T.; Wolfe, D. Tissue processing and hematoxylin and eosin staining. Methods Mol. Biol. 2014, 1180, 31–43. [Google Scholar] [CrossRef]
- Sorensen, M.D.; Thiele, M.; Krag, A.; Daniels, S.J.; Leeming, D.J.; Karsdal, M.; Detlefsen, S. Stage-dependent expression of fibrogenic markers in alcohol-related liver disease. Pathol. Res. Pract. 2022, 231, 153798. [Google Scholar] [CrossRef]
- Babu, R.O.; Lui, V.C.H.; Chen, Y.; Yiu, R.S.W.; Ye, Y.; Niu, B.; Wu, Z.; Zhang, R.; Yu, M.O.N.; Chung, P.H.Y.; et al. Beta-amyloid deposition around hepatic bile ducts is a novel pathobiological and diagnostic feature of biliary atresia. J. Hepatol. 2020, 73, 1391–1403. [Google Scholar] [CrossRef]
- Broutier, L.; Andersson-Rolf, A.; Hindley, C.J.; Boj, S.F.; Clevers, H.; Koo, B.K.; Huch, M. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protoc. 2016, 11, 1724–1743. [Google Scholar] [CrossRef]
- Hu, H.; Gehart, H.; Artegiani, B.; Löpez-Iglesias, C.; Dekkers, F.; Basak, O.; van Es, J.; Chuva de Sousa Lopes, S.M.; Begthel, H.; Korving, J.; et al. Long-Term Expansion of Functional Mouse and Human Hepatocytes as 3D Organoids. Cell 2018, 175, 1591–1606.E19. [Google Scholar] [CrossRef]
- Yuan, S.; Wei, C.; Liu, G.; Zhang, L.; Li, J.; Li, L.; Cai, S.; Fang, L. Sorafenib attenuates liver fibrosis by triggering hepatic stellate cell ferroptosis via HIF-1alpha/SLC7A11 pathway. Cell Prolif. 2022, 55, e13158. [Google Scholar] [CrossRef]
- Pan, P.H.; Wang, Y.Y.; Lin, S.Y.; Liao, S.L.; Chen, Y.F.; Huang, W.C.; Chen, C.J.; Chen, W.Y. Plumbagin ameliorates bile duct ligation-induced cholestatic liver injury in rats. Biomed. Pharmacother. 2022, 151, 113133. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, W.-L.; Lian, H.; Chen, Y.; Song, Z.; Tam, P.K.H.; Chen, Y. Identification of a 13-Gene Immune Signature in Liver Fibrosis Reveals GABRE as a Novel Candidate Biomarker. Int. J. Mol. Sci. 2025, 26, 8387. https://doi.org/10.3390/ijms26178387
Wang W-L, Lian H, Chen Y, Song Z, Tam PKH, Chen Y. Identification of a 13-Gene Immune Signature in Liver Fibrosis Reveals GABRE as a Novel Candidate Biomarker. International Journal of Molecular Sciences. 2025; 26(17):8387. https://doi.org/10.3390/ijms26178387
Chicago/Turabian StyleWang, Wei-Lu, Haoran Lian, Yiling Chen, Zhejun Song, Paul Kwong Hang Tam, and Yan Chen. 2025. "Identification of a 13-Gene Immune Signature in Liver Fibrosis Reveals GABRE as a Novel Candidate Biomarker" International Journal of Molecular Sciences 26, no. 17: 8387. https://doi.org/10.3390/ijms26178387
APA StyleWang, W.-L., Lian, H., Chen, Y., Song, Z., Tam, P. K. H., & Chen, Y. (2025). Identification of a 13-Gene Immune Signature in Liver Fibrosis Reveals GABRE as a Novel Candidate Biomarker. International Journal of Molecular Sciences, 26(17), 8387. https://doi.org/10.3390/ijms26178387





