Genetic Polymorphisms of ALDH2 and ADH1B in Alcohol-Induced Liver Injury: Molecular Mechanisms of Inflammation and Disease Progression in East Asian Populations
Abstract
1. Introduction
2. Methods
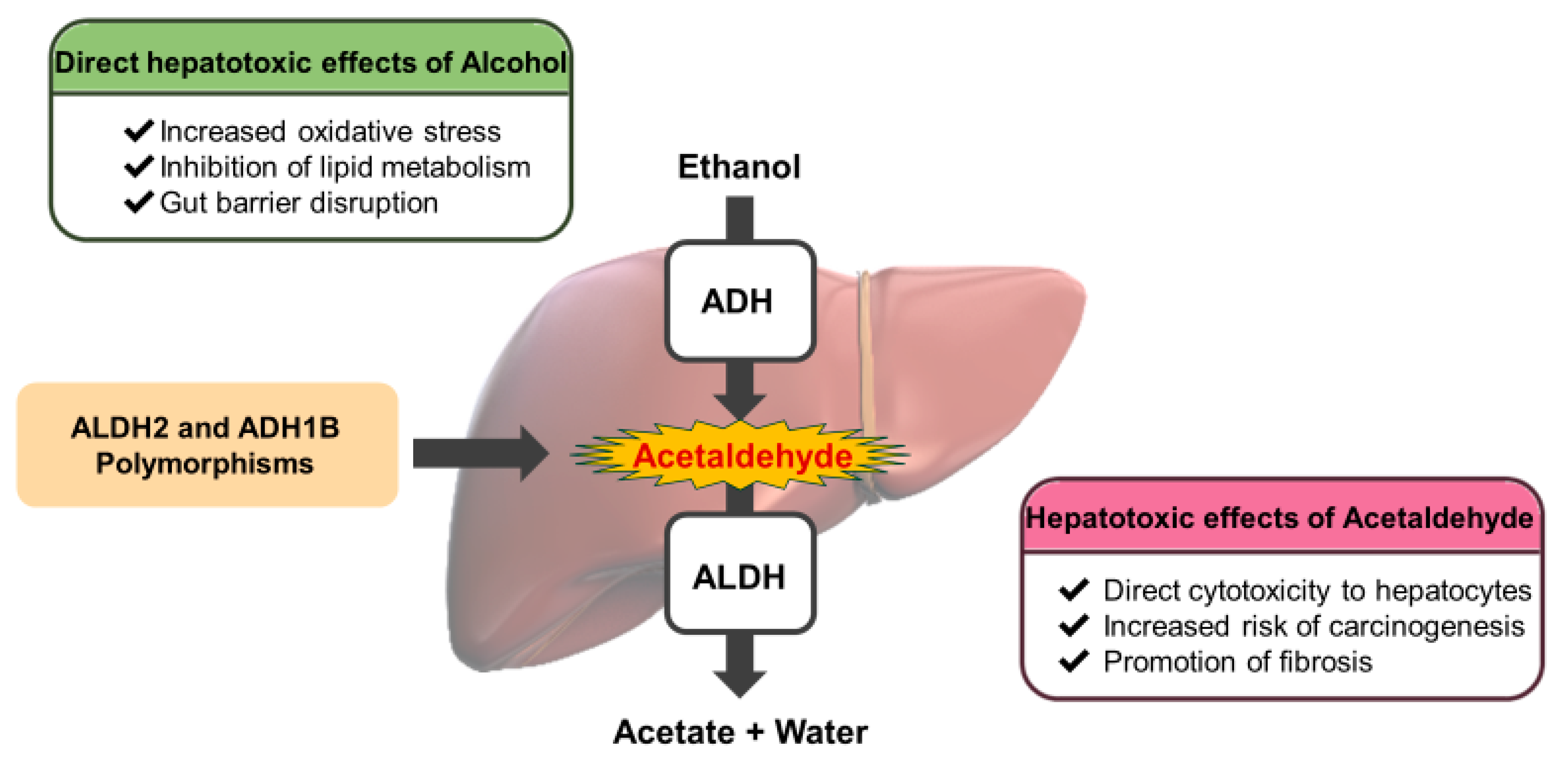
3. Ethanol Metabolism in the Liver
3.1. The Direct Hepatotoxic Effects of Alcohol
3.1.1. Increased Oxidative Stress
3.1.2. Inhibition of Lipid Metabolism
3.1.3. Disruption of the Intestinal Barrier and Translocation of Lipopolysaccharide to the Liver
3.2. Hepatotoxic Effects of Acetaldehyde
3.2.1. Direct Cytotoxicity to Hepatocytes
3.2.2. Increased Risk of Carcinogenesis
3.2.3. Promotion of Fibrosis
4. Impact of ALDH2 and ADH1B Polymorphisms on Liver Disease Progression
4.1. Association Between ADH1B and Liver Disease
4.2. Association Between ALDH2 and Liver Disease
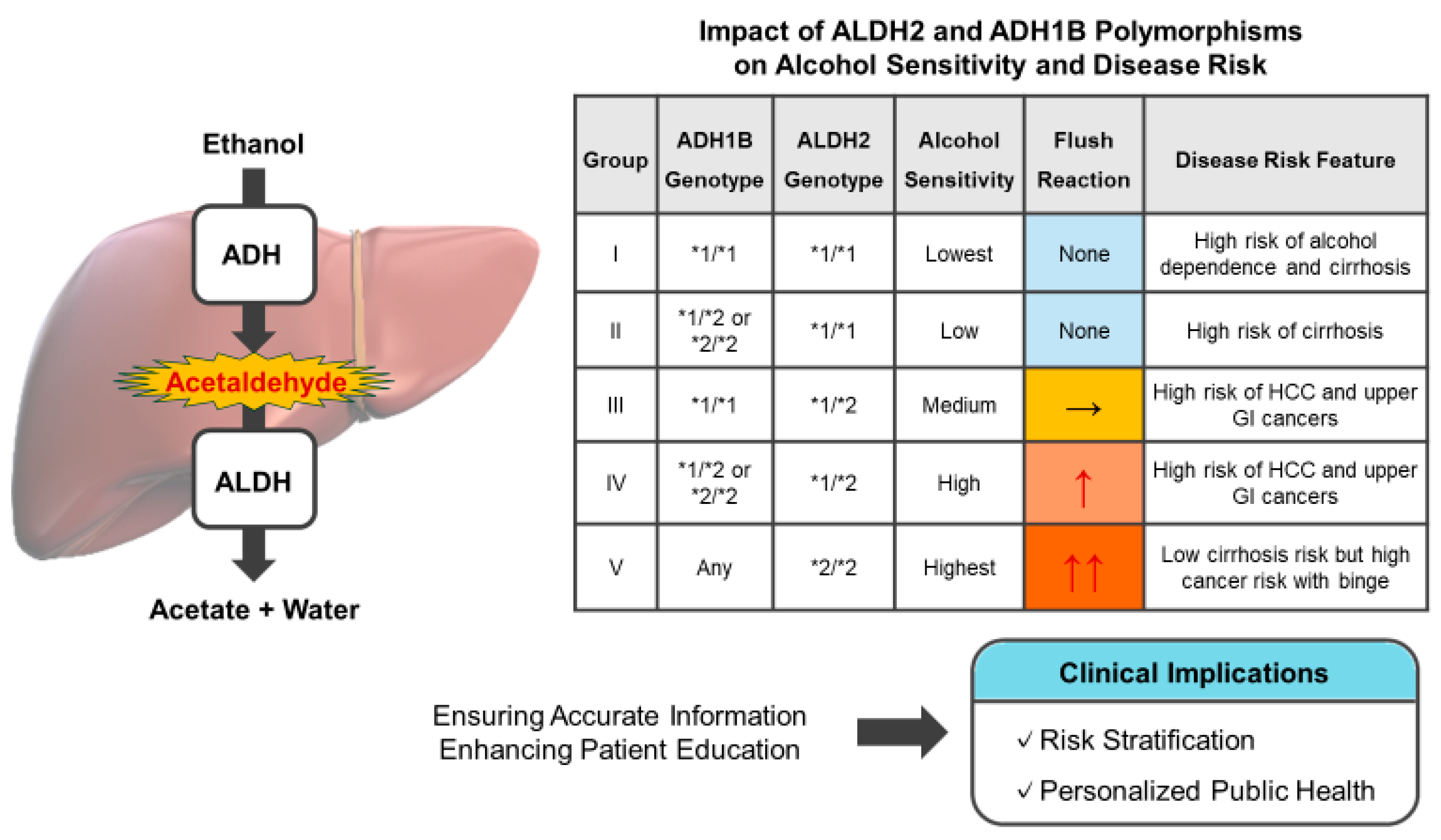
4.3. Classification into Five Groups Based on Alcohol Sensitivity
- Group I: ALDH2*1/*1 and ADH1B*1/*1;
- Group II: ALDH2*1/*1 and ADH1B*1/*2 or *2/*2;
- Group III: ALDH2*1/*2 and ADH1B*1/*1;
- Group IV: ALDH2*1/*2 and ADH1B*1/*2 or *2/*2;
- Group V: ALDH2*2/*2 and ADH1B*1/*1, *1/*2, or *2/*2.
4.3.1. Group I: ALDH2*1/*1 (Active Type) + ADH1B*1/*1 (Low-Activity Type)
4.3.2. Group II: ALDH2*1/*1 (Active Type) + ADH1B*1/*2 or *2/*2 (High-Activity Type)
4.3.3. Group III: ALDH2*1/*2 (Heterozygous) + ADH1B*1/*1 (Low-Activity Type)
4.3.4. Group IV: ALDH2*1/*2 (Heterozygous) + ADH1B*1/*2 or *2/*2 (High-Activity Type)
4.3.5. Group V: ALDH2*2/*2 (Homozygous) + Any ADH1B Genotype (*1/*1, *1/*2, or *2/*2)
5. Risk Stratification and Preventive Guidance Based on Genetic Polymorphisms
6. Conclusions and Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ADH1B | Alcohol dehydrogenase 1B |
| ALDH2 | Aldehyde dehydrogenase 2 |
| ALD | Alcohol-associated liver disease |
| DTC | Direct-to-consumer (genetic testing) |
| HCC | Hepatocellular carcinoma |
| GI | Gastrointestinal |
| ROS | Reactive oxygen species |
| TNF-α | Tumor necrosis factor-alpha |
| IL-1β | Interleukin-1 beta |
| NAD+ | Nicotinamide adenine dinucleotide (oxidized form) |
| NADH | Nicotinamide adenine dinucleotide (reduced form) |
| PPARα | Peroxisome proliferator-activated receptor alpha |
| AMPK | AMP-activated protein kinase |
| SREBP1c | Sterol regulatory element-binding protein 1c |
| VLDL | Very-low-density lipoprotein |
| LPS | Lipopolysaccharide |
| TLR4 | Toll-like receptor 4 |
| DNA | Deoxyribonucleic acid |
| CI | Confidence interval |
| OR | Odds ratio |
References
- Hernandez-Evole, H.; Jimenez-Esquivel, N.; Pose, E.; Bataller, R. Alcohol-associated liver disease: Epidemiology and management. Ann. Hepatol. 2024, 29, 101162. [Google Scholar] [CrossRef]
- Huang, D.Q.; Terrault, N.A.; Tacke, F.; Gluud, L.L.; Arrese, M.; Bugianesi, E.; Loomba, R. Global epidemiology of cirrhosis—Aetiology, trends and predictions. Nat. Rev. Gastroenterol. Hepatol. 2023, 20, 388–398. [Google Scholar] [CrossRef]
- Yoshiji, H.; Nagoshi, S.; Akahane, T.; Asaoka, Y.; Ueno, Y.; Ogawa, K.; Kawaguchi, T.; Kurosaki, M.; Sakaida, I.; Shimizu, M.; et al. Evidence-based clinical practice guidelines for Liver Cirrhosis 2020. J. Gastroenterol. 2021, 56, 593–619. [Google Scholar] [CrossRef]
- Mezzano, G.; Juanola, A.; Cardenas, A.; Mezey, E.; Hamilton, J.P.; Pose, E.; Graupera, I.; Gines, P.; Sola, E.; Hernaez, R. Global burden of disease: Acute-on-chronic liver failure, a systematic review and meta-analysis. Gut 2022, 71, 148–155. [Google Scholar] [CrossRef]
- Zhang, N.; Xue, F.; Wu, X.N.; Zhang, W.; Hou, J.J.; Xiang, J.X.; Lv, Y.; Zhang, X.F. The global burden of alcoholic liver disease: A systematic analysis of the global burden of disease study 2019. Alcohol Alcohol. 2023, 58, 485–496. [Google Scholar] [CrossRef] [PubMed]
- Mackowiak, B.; Fu, Y.; Maccioni, L.; Gao, B. Alcohol-associated liver disease. J. Clin. Investig. 2024, 134, e176345. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Fan, X.; Miyata, T.; Kim, A.; Cajigas-Du Ross, C.K.; Ray, S.; Huang, E.; Taiwo, M.; Arya, R.; Wu, J.; et al. Recent Advances in Understanding of Pathogenesis of Alcohol-Associated Liver Disease. Annu. Rev. Pathol. Mech. Dis. 2023, 18, 411–438. [Google Scholar] [CrossRef] [PubMed]
- Gao, B.; Bataller, R. Alcoholic liver disease: Pathogenesis and new therapeutic targets. Gastroenterology 2011, 141, 1572–1585. [Google Scholar] [CrossRef]
- Eng, M.Y.; Luczak, S.E.; Wall, T.L. ALDH2, ADH1B, and ADH1C genotypes in Asians: A literature review. Alcohol. Res. Health 2007, 30, 22–27. [Google Scholar]
- Verster, J.C.; Vermeulen, S.A.; Loo, A.; Balikji, S.; Kraneveld, A.D.; Garssen, J.; Scholey, A. Dietary Nutrient Intake, Alcohol Metabolism, and Hangover Severity. J. Clin. Med. 2019, 8, 1316. [Google Scholar] [CrossRef]
- Cho, Y.; Lin, K.; Lee, S.H.; Yu, C.; Valle, D.S.; Avery, D.; Lv, J.; Jung, K.; Li, L.; Smith, G.D.; et al. Genetic influences on alcohol flushing in East Asian populations. BMC Genom. 2023, 24, 638. [Google Scholar] [CrossRef] [PubMed]
- Setshedi, M.; Wands, J.R.; Monte, S.M. Acetaldehyde adducts in alcoholic liver disease. Oxid. Med. Cell. Longev. 2010, 3, 178–185. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.H.; Melis, M.; Mai, K.; Gudas, L.J.; Trasino, S.E. Fenretinide Improves Intestinal Barrier Function and Mitigates Alcohol Liver Disease. Front. Pharmacol. 2021, 12, 630557. [Google Scholar] [CrossRef] [PubMed]
- Lamb, R.J.; Griffiths, K.; Lip, G.Y.H.; Sorokin, V.; Frenneaux, M.P.; Feelisch, M.; Madhani, M. ALDH2 polymorphism and myocardial infarction: From alcohol metabolism to redox regulation. Pharmacol. Ther. 2024, 259, 108666. [Google Scholar] [CrossRef]
- Chang, Y.C.; Lee, H.L.; Yang, W.; Hsieh, M.L.; Liu, C.C.; Lee, T.Y.; Huang, J.Y.; Nong, J.Y.; Li, F.A.; Chuang, H.L.; et al. A common East-Asian ALDH2 mutation causes metabolic disorders and the therapeutic effect of ALDH2 activators. Nat. Commun. 2023, 14, 5971. [Google Scholar] [CrossRef] [PubMed]
- Seike, T.; Chen, C.H.; Mochly-Rosen, D. Impact of common ALDH2 inactivating mutation and alcohol consumption on Alzheimer’s disease. Front. Aging Neurosci. 2023, 15, 1223977. [Google Scholar] [CrossRef]
- Chen, C.H.; Kraemer, B.R.; Mochly-Rosen, D. ALDH2 variance in disease and populations. Dis. Model. Mech. 2022, 15, dmm049601. [Google Scholar] [CrossRef]
- Zhai, Z.; Yamauchi, T.; Shangraw, S.; Hou, V.; Matsumoto, A.; Fujita, M. Ethanol Metabolism and Melanoma. Cancers 2023, 15, 1258. [Google Scholar] [CrossRef]
- Chen, C.H.; Wang, W.L.; Hsu, M.H.; Mochly-Rosen, D. Alcohol Consumption, ALDH2 Polymorphism as Risk Factors for Upper Aerodigestive Tract Cancer Progression and Prognosis. Life 2022, 12, 348. [Google Scholar] [CrossRef]
- Contreras-Zentella, M.L.; Villalobos-Garcia, D.; Hernandez-Munoz, R. Ethanol Metabolism in the Liver, the Induction of Oxidant Stress, and the Antioxidant Defense System. Antioxidants 2022, 11, 1258. [Google Scholar] [CrossRef]
- Lu, Y.; Cederbaum, A.I. CYP2E1 and oxidative liver injury by alcohol. Free Radic. Biol. Med. 2008, 44, 723–738. [Google Scholar] [CrossRef]
- Harjumaki, R.; Pridgeon, C.S.; Ingelman-Sundberg, M. CYP2E1 in Alcoholic and Non-Alcoholic Liver Injury. Roles of ROS, Reactive Intermediates and Lipid Overload. Int. J. Mol. Sci. 2021, 22, 8221. [Google Scholar] [CrossRef]
- Seitz, H.K.; Moreira, B.; Neuman, M.G. Pathogenesis of Alcoholic Fatty Liver a Narrative Review. Life 2023, 13, 1662. [Google Scholar] [CrossRef]
- You, M.; Arteel, G.E. Effect of ethanol on lipid metabolism. J. Hepatol. 2019, 70, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Steiner, J.L.; Lang, C.H. Alcohol, Adipose Tissue and Lipid Dysregulation. Biomolecules 2017, 7, 16. [Google Scholar] [CrossRef] [PubMed]
- Dukic, M.; Radonjic, T.; Jovanovic, I.; Zdravkovic, M.; Todorovic, Z.; Kraisnik, N.; Arandelovic, B.; Mandic, O.; Popadic, V.; Nikolic, N.; et al. Alcohol, Inflammation, and Microbiota in Alcoholic Liver Disease. Int. J. Mol. Sci. 2023, 24, 3735. [Google Scholar] [CrossRef]
- An, L.; Wirth, U.; Koch, D.; Schirren, M.; Drefs, M.; Koliogiannis, D.; Niess, H.; Andrassy, J.; Guba, M.; Bazhin, A.V.; et al. The Role of Gut-Derived Lipopolysaccharides and the Intestinal Barrier in Fatty Liver Diseases. J. Gastrointest. Surg. 2022, 26, 671–683. [Google Scholar] [CrossRef]
- Eom, J.A.; Jeong, J.J.; Han, S.H.; Kwon, G.H.; Lee, K.J.; Gupta, H.; Sharma, S.P.; Won, S.M.; Oh, K.K.; Yoon, S.J.; et al. Gut-microbiota prompt activation of natural killer cell on alcoholic liver disease. Gut Microbes 2023, 15, 2281014. [Google Scholar] [CrossRef]
- Tadokoro, T.; Morishita, A.; Himoto, T.; Masaki, T. Nutritional Support for Alcoholic Liver Disease. Nutrients 2023, 15, 1360. [Google Scholar] [CrossRef] [PubMed]
- Rao, R. Endotoxemia and gut barrier dysfunction in alcoholic liver disease. Hepatology 2009, 50, 638–644. [Google Scholar] [CrossRef]
- Dubinkina, V.B.; Tyakht, A.V.; Odintsova, V.Y.; Yarygin, K.S.; Kovarsky, B.A.; Pavlenko, A.V.; Ischenko, D.S.; Popenko, A.S.; Alexeev, D.G.; Taraskina, A.Y.; et al. Links of gut microbiota composition with alcohol dependence syndrome and alcoholic liver disease. Microbiome 2017, 5, 141. [Google Scholar] [CrossRef]
- Swanson, G.R.; Garg, K.; Shaikh, M.; Keshavarzian, A. Increased Intestinal Permeability and Decreased Resiliency of the Intestinal Barrier in Alcoholic Liver Disease. Clin. Transl. Gastroenterol. 2024, 15, e00689. [Google Scholar] [CrossRef]
- Kolios, G.; Valatas, V.; Kouroumalis, E. Role of Kupffer cells in the pathogenesis of liver disease. World J. Gastroenterol. 2006, 12, 7413–7420. [Google Scholar] [CrossRef] [PubMed]
- Khomich, O.; Ivanov, A.V.; Bartosch, B. Metabolic Hallmarks of Hepatic Stellate Cells in Liver Fibrosis. Cells 2019, 9, 24. [Google Scholar] [CrossRef]
- Tuma, D.J. Role of malondialdehyde-acetaldehyde adducts in liver injury. Free Radic. Biol. Med. 2002, 32, 303–308. [Google Scholar] [CrossRef]
- Subramaniyan, V.; Chakravarthi, S.; Jegasothy, R.; Seng, W.Y.; Fuloria, N.K.; Fuloria, S.; Hazarika, I.; Das, A. Alcohol-associated liver disease: A review on its pathophysiology, diagnosis and drug therapy. Toxicol. Rep. 2021, 8, 376–385. [Google Scholar] [CrossRef]
- Visapaa, J.P.; Gotte, K.; Benesova, M.; Li, J.; Homann, N.; Conradt, C.; Inoue, H.; Tisch, M.; Horrmann, K.; Vakevainen, S.; et al. Increased cancer risk in heavy drinkers with the alcohol dehydrogenase 1C*1 allele, possibly due to salivary acetaldehyde. Gut 2004, 53, 871–876. [Google Scholar] [CrossRef] [PubMed]
- Seitz, H.K.; Becker, P. Alcohol metabolism and cancer risk. Alcohol. Res. Health 2007, 30, 38–47. [Google Scholar]
- Seitz, H.K.; Stickel, F. Acetaldehyde as an underestimated risk factor for cancer development: Role of genetics in ethanol metabolism. Genes. Nutr. 2010, 5, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Vijayraghavan, S.; Porcher, L.; Mieczkowski, P.A.; Saini, N. Acetaldehyde makes a distinct mutation signature in single-stranded DNA. Nucleic Acids Res. 2022, 50, 7451–7464. [Google Scholar] [CrossRef]
- Smedra, A.; Berent, J. The Influence of the Oral Microbiome on Oral Cancer: A Literature Review and a New Approach. Biomolecules 2023, 13, 815. [Google Scholar] [CrossRef]
- Hoes, L.; Dok, R.; Verstrepen, K.J.; Nuyts, S. Ethanol-Induced Cell Damage Can Result in the Development of Oral Tumors. Cancers 2021, 13, 1492–1513. [Google Scholar] [CrossRef]
- Osna, N.A.; Rasineni, K.; Ganesan, M.; Donohue, T.M., Jr.; Kharbanda, K.K. Pathogenesis of Alcohol-Associated Liver Disease. J. Clin. Exp. Hepatol. 2022, 12, 1492–1513. [Google Scholar] [CrossRef] [PubMed]
- IARC Working Group on the Evaluation of Carcinogenic Risks to Humans. Personal habits and indoor combustions. In IARC Monographs on the Evaluation of Carcinogenic Risks to Humans; International Agency for Research on Cancer: Lyon, France, 2012; Volume 100, pp. 1–538. [Google Scholar]
- Svegliati-Baroni, G.; Inagaki, Y.; Rincon-Sanchez, A.R.; Else, C.; Saccomanno, S.; Benedetti, A.; Ramirez, F.; Rojkind, M. Early response of alpha2(I) collagen to acetaldehyde in human hepatic stellate cells is TGF-beta independent. Hepatology 2005, 42, 343–352. [Google Scholar] [CrossRef]
- Ceni, E.; Crabb, D.W.; Foschi, M.; Mello, T.; Tarocchi, M.; Patussi, V.; Moraldi, L.; Moretti, R.; Milani, S.; Surrenti, C.; et al. Acetaldehyde inhibits PPARgamma via H2O2-mediated c-Abl activation in human hepatic stellate cells. Gastroenterology 2006, 131, 1235–1252. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.H.; Batey, R.G.; George, J. Role of ethanol in the regulation of hepatic stellate cell function. World J. Gastroenterol. 2006, 12, 6926–6932. [Google Scholar] [CrossRef]
- Vilar-Gomez, E.; Sookoian, S.; Pirola, C.J.; Liang, T.; Gawrieh, S.; Cummings, O.; Liu, W.; Chalasani, N.P. ADH1B*2 Is Associated With Reduced Severity of Nonalcoholic Fatty Liver Disease in Adults, Independent of Alcohol Consumption. Gastroenterology 2020, 159, 929–943. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Deng, T.; Luo, H. Aldehyde Dehydrogenase 2 (ALDH2) Polymorphism and the Risk of Alcoholic Liver Cirrhosis among East Asians: A Meta-Analysis. Yonsei Med. J. 2016, 57, 879–884. [Google Scholar] [CrossRef]
- Li, D.; Zhao, H.; Gelernter, J. Strong association of the alcohol dehydrogenase 1B gene (ADH1B) with alcohol dependence and alcohol-induced medical diseases. Biol. Psychiatry 2011, 70, 504–512. [Google Scholar] [CrossRef]
- Yokoyama, A.; Mizukami, T.; Matsui, T.; Yokoyama, T.; Kimura, M.; Matsushita, S.; Higuchi, S.; Maruyama, K. Genetic polymorphisms of alcohol dehydrogenase-1B and aldehyde dehydrogenase-2 and liver cirrhosis, chronic calcific pancreatitis, diabetes mellitus, and hypertension among Japanese alcoholic men. Alcohol. Clin. Exp. Res. 2013, 37, 1391–1401. [Google Scholar] [CrossRef]
- Li, H.; Borinskaya, S.; Yoshimura, K.; Kal’ina, N.; Marusin, A.; Stepanov, V.A.; Qin, Z.; Khaliq, S.; Lee, M.Y.; Yang, Y.; et al. Refined geographic distribution of the oriental ALDH2*504Lys (nee 487Lys) variant. Ann. Hum. Genet. 2009, 73, 335–345. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Wang, C.; Xu, H.; Gao, Y. Aldehyde Dehydrogenase, Liver Disease and Cancer. Int. J. Biol. Sci. 2020, 16, 921–934. [Google Scholar] [CrossRef]
- Chang, B.; Hao, S.; Zhang, L.; Gao, M.; Sun, Y.; Huang, A.; Teng, G.; Li, B.; Crabb, D.W.; Kusumanchi, P.; et al. Association Between Aldehyde Dehydrogenase 2 Glu504Lys Polymorphism and Alcoholic Liver Disease. Am. J. Med. Sci. 2018, 356, 10–14. [Google Scholar] [CrossRef]
- Yao, P.; Zhang, Z.; Liu, H.; Jiang, P.; Li, W.; Du, W. p53 protects against alcoholic fatty liver disease via ALDH2 inhibition. EMBO J. 2023, 42, e112304. [Google Scholar] [CrossRef]
- Rungratanawanich, W.; Lin, Y.; Wang, X.; Kawamoto, T.; Chidambaram, S.B.; Song, B.J. ALDH2 deficiency increases susceptibility to binge alcohol-induced gut leakiness, endotoxemia, and acute liver injury in mice through the gut-liver axis. Redox Biol. 2023, 59, 102577. [Google Scholar] [CrossRef]
- Montel, R.A.; Munoz-Zuluaga, C.; Stiles, K.M.; Crystal, R.G. Can gene therapy be used to prevent cancer? Gene therapy for aldehyde dehydrogenase 2 deficiency. Cancer Gene Ther. 2022, 29, 889–896. [Google Scholar] [CrossRef]
- Yoshimasu, K.; Mure, K.; Hashimoto, M.; Takemura, S.; Tsuno, K.; Hayashida, M.; Kinoshita, K.; Takeshita, T.; Miyashita, K. Genetic alcohol sensitivity regulated by ALDH2 and ADH1B polymorphisms is strongly associated with depression and anxiety in Japanese employees. Drug Alcohol Depend. 2015, 147, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.J.; Yokoyama, A.; Yokoyama, T.; Huang, Y.C.; Wu, S.Y.; Shao, Y.; Niu, J.; Wang, J.; Liu, Y.; Zhou, X.Q.; et al. Relationship between genetic polymorphisms of ALDH2 and ADH1B and esophageal cancer risk: A meta-analysis. World J. Gastroenterol. 2010, 16, 4210–4220. [Google Scholar] [CrossRef]
- Hishimoto, A.; Fukutake, M.; Mouri, K.; Nagasaki, Y.; Asano, M.; Ueno, Y.; Nishiguchi, N.; Shirakawa, O. Alcohol and aldehyde dehydrogenase polymorphisms and risk for suicide: A preliminary observation in the Japanese male population. Genes Brain Behav. 2010, 9, 498–502. [Google Scholar] [CrossRef]
- Hoang, Y.T.T.; Nguyen, Y.T.; Vu, L.T.; Bui, H.T.T.; Nguyen, Q.V.; Vu, N.P.; Nguyen, T.D.; Nguyen, H.H. Association of ADH1B rs1229984, ADH1C rs698, and ALDH2 rs671 with Alcohol abuse and Alcoholic Cirrhosis in People Living in Northeast Vietnam. Asian Pac. J. Cancer Prev. 2023, 24, 2073–2082. [Google Scholar] [CrossRef] [PubMed]
- Choi, C.K.; Shin, M.H.; Cho, S.H.; Kim, H.Y.; Zheng, W.; Long, J.; Kweon, S.S. Association between ALDH2 and ADH1B Polymorphisms and the Risk for Colorectal Cancer in Koreans. Cancer Res. Treat. 2021, 53, 754–762. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.G.; Yen, T.T.; Wei, C.Y.; Hsiao, T.H.; Chen, I.C. Impacts of ADH1B rs1229984 and ALDH2 rs671 polymorphisms on risks of alcohol-related disorder and cancer. Cancer Med. 2023, 12, 747–759. [Google Scholar] [CrossRef]
- Masaoka, H.; Ito, H.; Soga, N.; Hosono, S.; Oze, I.; Watanabe, M.; Tanaka, H.; Yokomizo, A.; Hayashi, N.; Eto, M.; et al. Aldehyde dehydrogenase 2 (ALDH2) and alcohol dehydrogenase 1B (ADH1B) polymorphisms exacerbate bladder cancer risk associated with alcohol drinking: Gene-environment interaction. Carcinogenesis 2016, 37, 583–588. [Google Scholar] [CrossRef][Green Version]
- Masaoka, H.; Ito, H.; Gallus, S.; Watanabe, M.; Yokomizo, A.; Eto, M.; Matsuo, K. Combination of ALDH2 and ADH1B polymorphisms is associated with smoking initiation: A large-scale cross-sectional study in a Japanese population. Drug Alcohol Depend. 2017, 173, 85–91. [Google Scholar] [CrossRef]
- Liu, Z.Y.; Lin, X.H.; Guo, H.Y.; Shi, X.; Zhang, D.Y.; Sun, J.L.; Zhang, G.C.; Xu, R.C.; Wang, F.; Yu, X.N.; et al. Multi-Omics profiling identifies aldehyde dehydrogenase 2 as a critical mediator in the crosstalk between Treg-mediated immunosuppression microenvironment and hepatocellular carcinoma. Int. J. Biol. Sci. 2024, 20, 2763–2778. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.C.; Lee, H.S.; Jung, S.H.; Yi, S.Y.; Jung, H.K.; Yoon, J.H.; Kim, C.Y. Association between polymorphisms of ethanol-metabolizing enzymes and susceptibility to alcoholic cirrhosis in a Korean male population. J. Korean Med. Sci. 2001, 16, 745–750. [Google Scholar] [CrossRef] [PubMed]
- Chao, Y.C.; Liou, S.R.; Chung, Y.Y.; Tang, H.S.; Hsu, C.T.; Li, T.K.; Yin, S.J. Polymorphism of alcohol and aldehyde dehydrogenase genes and alcoholic cirrhosis in Chinese patients. Hepatology 1994, 19, 360–366. [Google Scholar] [CrossRef]
- Seo, W.; Gao, Y.; He, Y.; Sun, J.; Xu, H.; Feng, D.; Park, S.H.; Cho, Y.E.; Guillot, A.; Ren, T.; et al. ALDH2 deficiency promotes alcohol-associated liver cancer by activating oncogenic pathways via oxidized DNA-enriched extracellular vesicles. J. Hepatol. 2019, 71, 1000–1011. [Google Scholar] [CrossRef]
- Tsai, M.C.; Yang, S.S.; Lin, C.C.; Wang, W.L.; Hsu, Y.C.; Chen, Y.S.; Hu, J.T.; Lin, J.Y.; Yu, M.L.; Lin, C.W. Association of Heavy Alcohol Intake and ALDH2 rs671 Polymorphism with Hepatocellular Carcinoma and Mortality in Patients with Hepatitis B Virus-Related Cirrhosis. JAMA Netw. Open 2022, 5, e2223511. [Google Scholar] [CrossRef]
- Ishioka, K.; Masaoka, H.; Ito, H.; Oze, I.; Ito, S.; Tajika, M.; Shimizu, Y.; Niwa, Y.; Nakamura, S.; Matsuo, K. Association between ALDH2 and ADH1B polymorphisms, alcohol drinking and gastric cancer: A replication and mediation analysis. Gastric Cancer 2018, 21, 936–945. [Google Scholar] [CrossRef]
- Zhong, Z.; Hou, J.; Li, B.; Zhang, Q.; Li, C.; Liu, Z.; Yang, M.; Zhong, W.; Zhao, P. Genetic Polymorphisms of the Mitochondrial Aldehyde Dehydrogenase ALDH2 Gene in a Large Ethnic Hakka Population in Southern China. Med. Sci. Monit. 2018, 24, 2038–2044. [Google Scholar] [CrossRef]
- Polimanti, R.; Gelernter, J. ADH1B: From alcoholism, natural selection, and cancer to the human phenome. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2018, 177, 113–125. [Google Scholar] [CrossRef]
- Mathews, R.; Hall, W.; Carter, A. Direct-to-consumer genetic testing for addiction susceptibility: A premature commercialisation of doubtful validity and value. Addiction 2012, 107, 2069–2074. [Google Scholar] [CrossRef] [PubMed]
- Owaki, Y.; Yoshimoto, H.; Saito, G.; Dobashi, S.; Kushio, S.; Nakamura, A.; Goto, T.; Togo, Y.; Mori, K.; Hokazono, H. Effectiveness of genetic feedback on alcohol metabolism to reduce alcohol consumption in young adults: An open-label randomized controlled trial. BMC Med. 2024, 22, 205. [Google Scholar] [CrossRef] [PubMed]
- Luczak, S.E.; Yarnell, L.M.; Prescott, C.A.; Myers, M.G.; Liang, T.; Wall, T.L. Effects of ALDH2*2 on alcohol problem trajectories of Asian American college students. J. Abnorm. Psychol. 2014, 123, 130–140. [Google Scholar] [CrossRef] [PubMed]
- Mullins, V.A.; Bresette, W.; Johnstone, L.; Hallmark, B.; Chilton, F.H. Genomics in Personalized Nutrition: Can You “Eat for Your Genes”? Nutrients 2020, 12, 3118. [Google Scholar] [CrossRef]
- Roberts, J.S.; Ostergren, J. Direct-to-Consumer Genetic Testing and Personal Genomics Services: A Review of Recent Empirical Studies. Curr. Genet. Med. Rep. 2013, 1, 182–200. [Google Scholar] [CrossRef]
- Orth, M. Direct to Consumer Laboratory Testing (DTCT)—Opportunities and Concerns. EJIFCC 2021, 32, 209–215. [Google Scholar]
- Onstwedder, S.M.; Jansen, M.E.; Cornel, M.C.; Rigter, T. Policy Guidance for Direct-to-Consumer Genetic Testing Services: Framework Development Study. J. Med. Internet Res. 2024, 26, e47389. [Google Scholar] [CrossRef]
- Driver, M.N.; Kuo, S.I.; Dick, D.M.; on behalf of the Spit For Science Working Group. Interest in Genetic Feedback for Alcohol Use Disorder and Related Substance Use and Psychiatric Outcomes among Young Adults. Brain Sci. 2020, 10, 1007. [Google Scholar] [CrossRef]
- The All of Us Research Program Investigators; Denny, J.C.; Rutter, J.L.; Goldstein, D.B.; Philippakis, A.; Smoller, J.W.; Jenkins, G.; Dishman, E. The “All of Us” Research Program. N. Engl. J. Med. 2019, 381, 668–676. [Google Scholar] [CrossRef] [PubMed]
| Gene | SNP (Amino Acid Change) | Genotype | Enzymatic Activity | Characteristics |
|---|---|---|---|---|
| ADH1B | rs1229984 (Arg47His) | *1/*1 (Arg/Arg) | Low | Slow conversion of ethanol to acetaldehyde |
| *1/*2 (Arg/His) | Intermediate–High | Increased enzymatic activity | ||
| *2/*2 (His/His) | High | Rapid production of acetaldehyde | ||
| ALDH2 | rs671 (Glu504Lys) | *1/*1 (Glu/Glu) | Active (normal) | Normal conversion of acetaldehyde to acetate |
| *1/*2 (Glu/Lys) | Low | Approximately 10–20% activity; associated with facial flushing, etc. | ||
| *2/*2 (Lys/Lys) | Inactive (deficient) | Near-zero activity; strong alcohol intolerance |
| Group | ADH1B Genotype | ALDH2 Genotype | Frequency | Alcohol Sensitivity | Flush Reaction | Disease Risk Feature |
|---|---|---|---|---|---|---|
| I | *1/*1 | *1/*1 | 5–10% | Lowest | None | High risk of alcohol dependence and cirrhosis |
| II | *1/*2 or *2/*2 | *1/*1 | 50% | Low | None | High risk of cirrhosis |
| III | *1/*1 | *1/*2 | 3–5% | Medium | Mild | High risk of HCC and upper GI cancers |
| IV | *1/*2 or *2/*2 | *1/*2 | 30–50% | High | Strong | High risk of HCC and upper GI cancers |
| V | Any | *2/*2 | 1–8% | Highest | Very Strong | Low cirrhosis risk but high cancer risk with binge |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tadokoro, T.; Oura, K.; Nakahara, M.; Fujita, K.; Tani, J.; Morishita, A.; Kobara, H. Genetic Polymorphisms of ALDH2 and ADH1B in Alcohol-Induced Liver Injury: Molecular Mechanisms of Inflammation and Disease Progression in East Asian Populations. Int. J. Mol. Sci. 2025, 26, 8328. https://doi.org/10.3390/ijms26178328
Tadokoro T, Oura K, Nakahara M, Fujita K, Tani J, Morishita A, Kobara H. Genetic Polymorphisms of ALDH2 and ADH1B in Alcohol-Induced Liver Injury: Molecular Mechanisms of Inflammation and Disease Progression in East Asian Populations. International Journal of Molecular Sciences. 2025; 26(17):8328. https://doi.org/10.3390/ijms26178328
Chicago/Turabian StyleTadokoro, Tomoko, Kyoko Oura, Mai Nakahara, Koji Fujita, Joji Tani, Asahiro Morishita, and Hideki Kobara. 2025. "Genetic Polymorphisms of ALDH2 and ADH1B in Alcohol-Induced Liver Injury: Molecular Mechanisms of Inflammation and Disease Progression in East Asian Populations" International Journal of Molecular Sciences 26, no. 17: 8328. https://doi.org/10.3390/ijms26178328
APA StyleTadokoro, T., Oura, K., Nakahara, M., Fujita, K., Tani, J., Morishita, A., & Kobara, H. (2025). Genetic Polymorphisms of ALDH2 and ADH1B in Alcohol-Induced Liver Injury: Molecular Mechanisms of Inflammation and Disease Progression in East Asian Populations. International Journal of Molecular Sciences, 26(17), 8328. https://doi.org/10.3390/ijms26178328






