Metagenomic Analysis Reveals the Anti-Inflammatory Properties of Mare Milk
Abstract
1. Introduction
2. Results
2.1. Comparison of Immune Markers
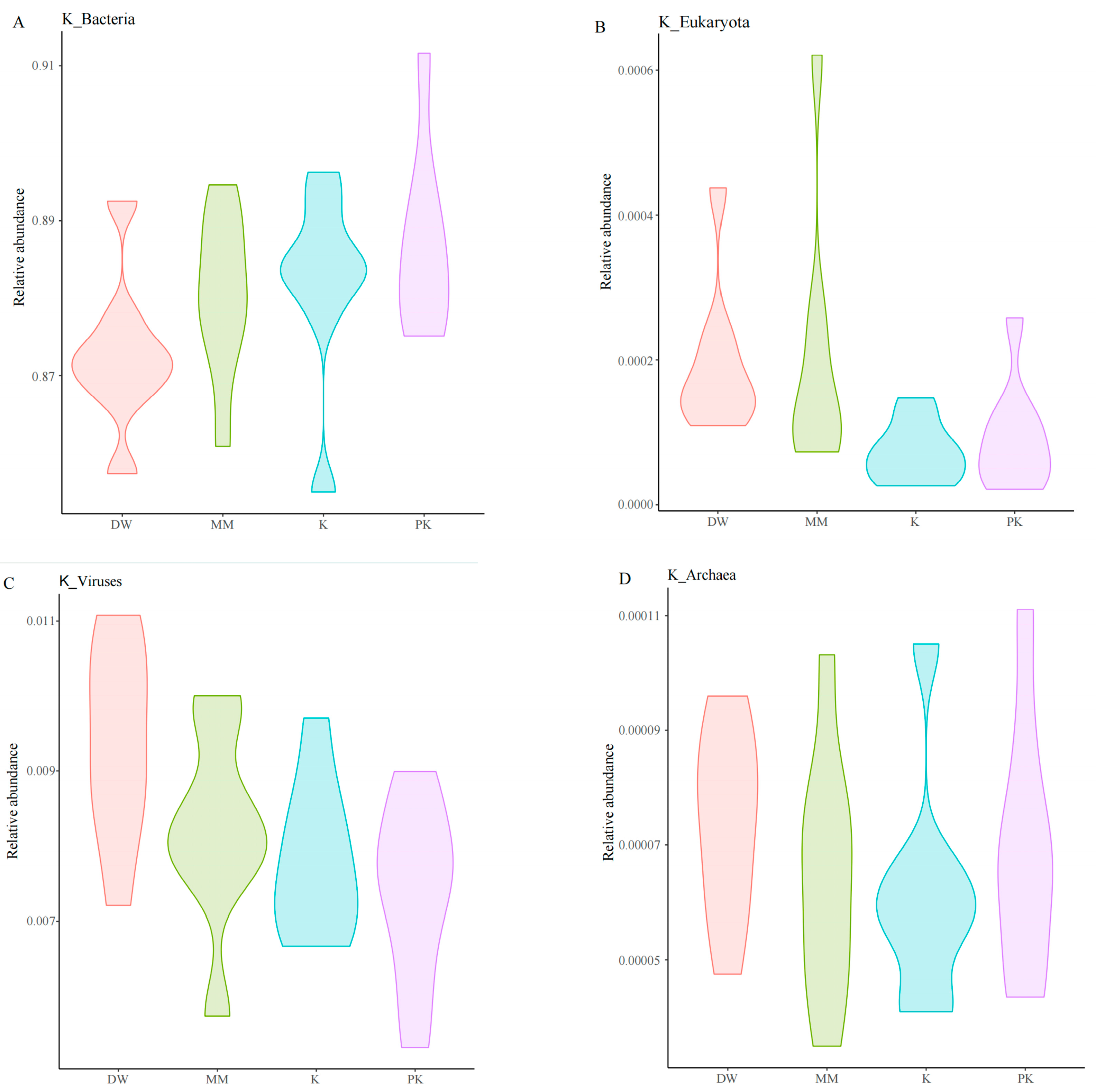
2.2. Analysis of Fecal Genome Characteristics
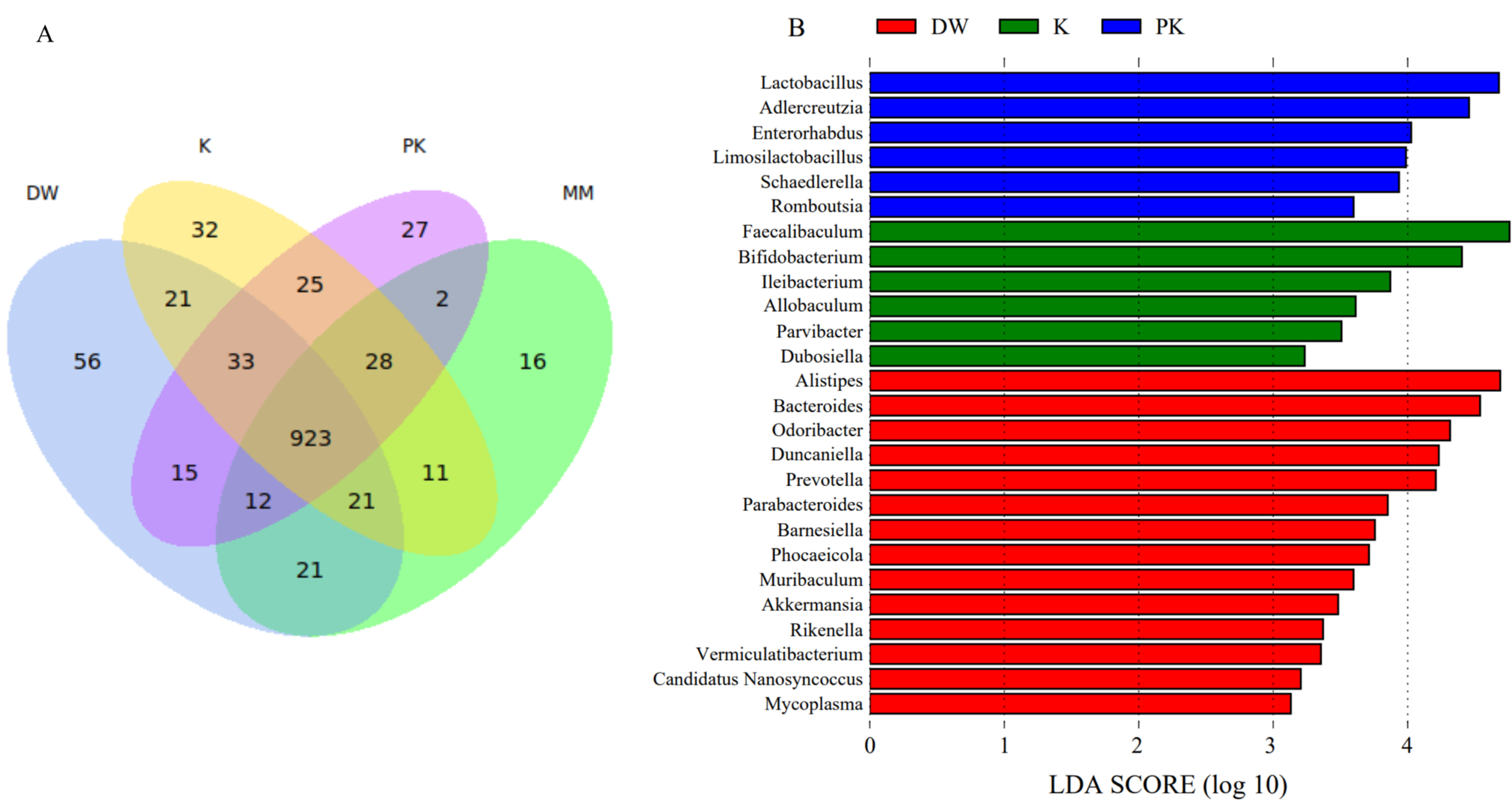
2.3. Comparison of Fecal Microbiota Among Different Groups
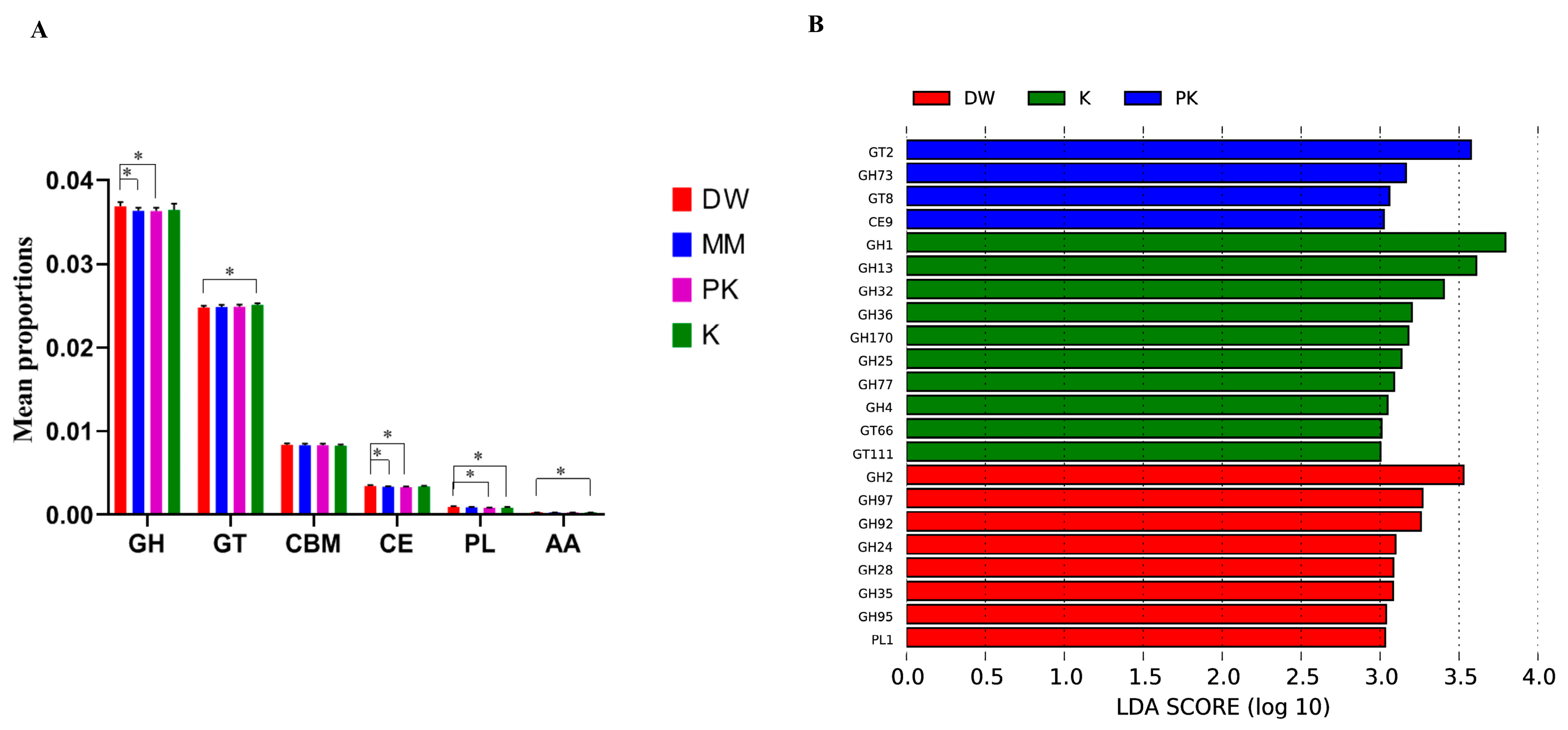
2.4. Functional Annotation of Fecal Microorganisms
2.4.1. Functional Annotation of CAZyme
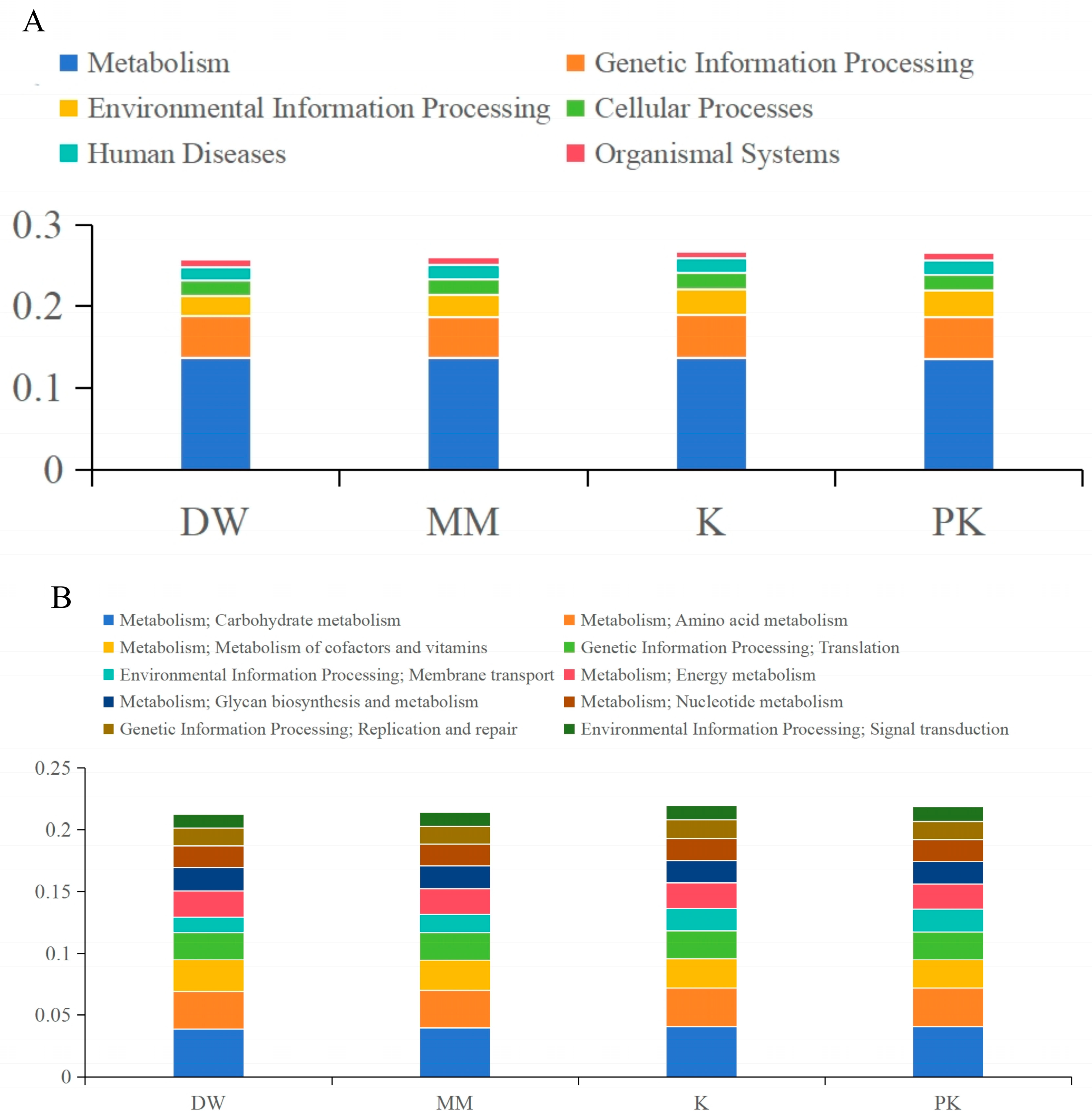
2.4.2. KEGG Functional Annotation
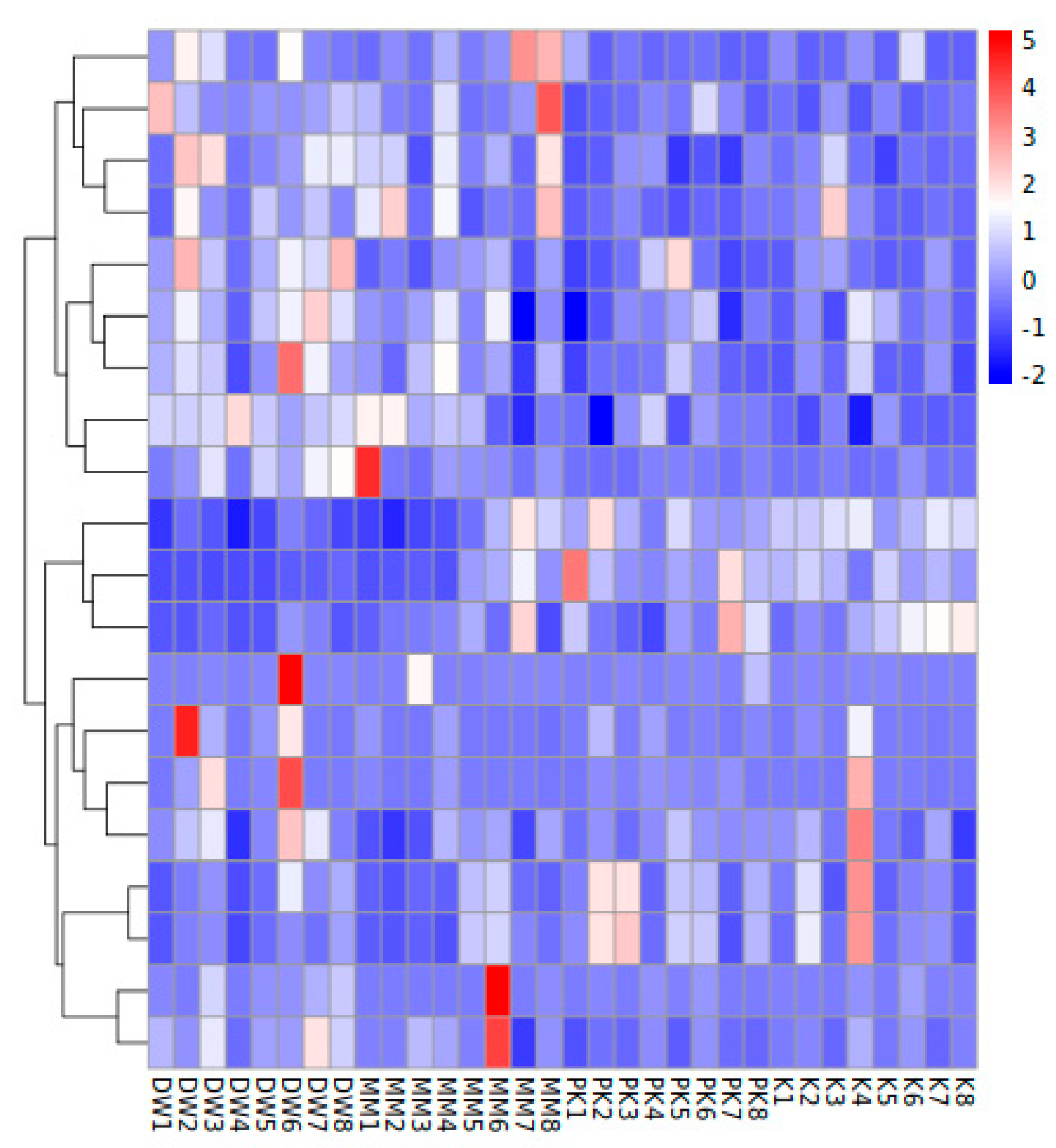
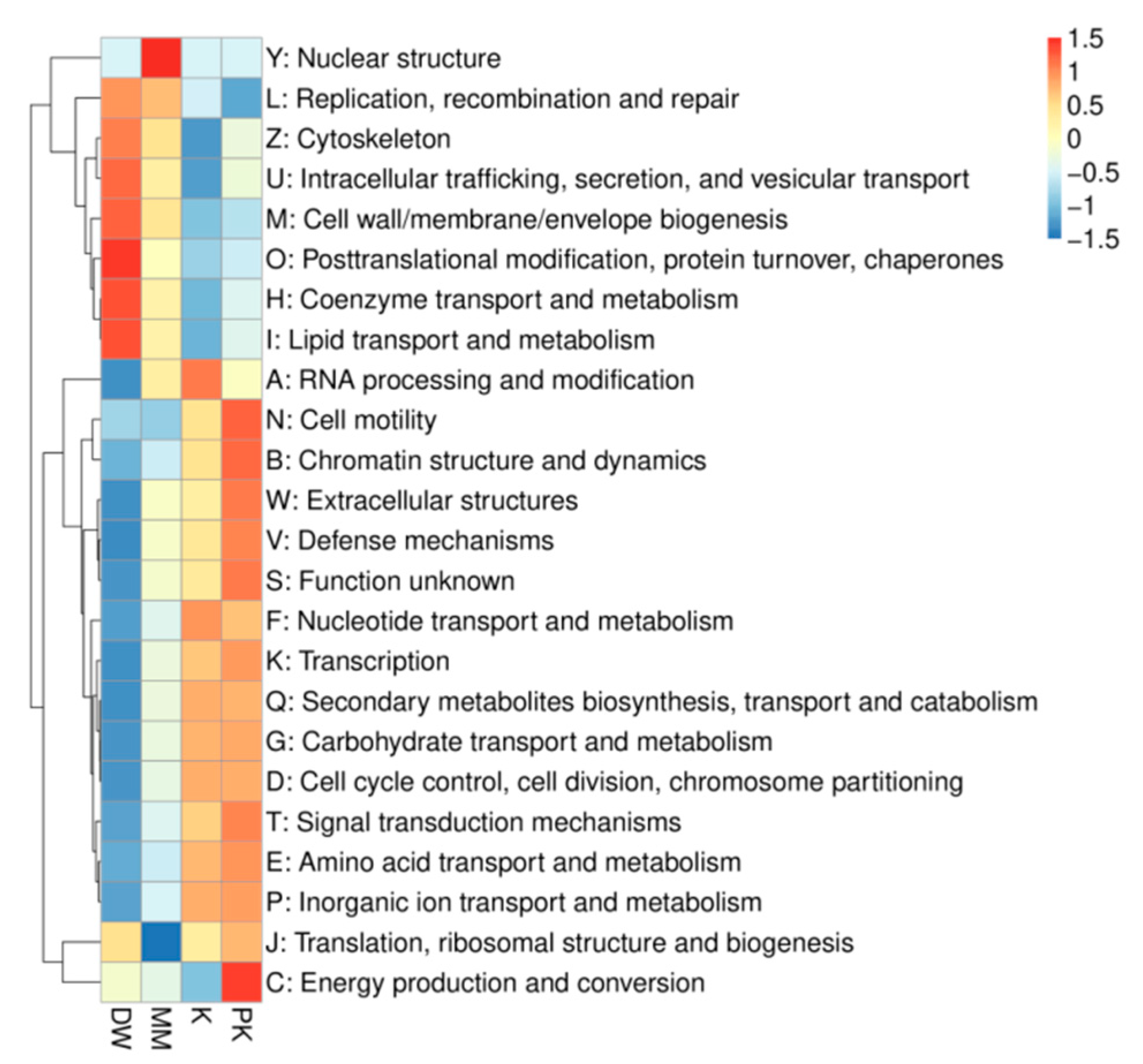
2.4.3. Cluster Analysis of Functional Relative Abundance
3. Discussion
4. Materials and Methods
4.1. Mare Milk Sampling
4.2. Animal Feeding and Sample Collection
4.3. Detection of Immune Markers
4.4. Fecal Microbial DNA Extraction and Sequencing
4.5. Bioinformatics Analysis
4.6. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kocyigit, E.; Abdurakhmanov, R.; Kocyigit, B.F. Potential role of camel, mare milk, and their products in inflammatory rheumatic diseases. Rheumatol. Int. 2024, 44, 425–434. [Google Scholar] [CrossRef]
- Shaikh, A.; Mehta, B.M.; Jana, A.H. Chemistry nutritional properties and application of Mare’s milk: A review. Agric. Rev. 2022, 43, 355–361. [Google Scholar]
- Musaev, A.; Sadykova, S.; Anambayeva, A.; Saizhanova, M.; Balkanay, G.; Kolbaev, M. Mare’s milk: Composition, properties, and application in medicine. Rchives Razi Inst. 2021, 76, 1125. [Google Scholar]
- Xia, Y.; Yu, J.; Miao, W.; Shuang, Q. A UPLC-Q-TOF-MS-based metabolomics approach for the evaluation of fermented mare’s milk to koumiss. Food Chem. 2020, 320, 126619. [Google Scholar] [CrossRef]
- He, Q.; Hou, Q.; Wang, Y.J.; Shen, L.; Sun, Z.; Zhang, H.; Liong, M.-T.; Kwok, L.-Y. Longterm administration of Lactobacillus casei Zhang stabilized aut microbiota of adults and reduced gut microbiota age index of older adults. J. Funct. Foods 2020, 64, 103682. [Google Scholar] [CrossRef]
- Gao, J.; Xu, K.; Liu, H.; Liu, G.; Bai, M.; Peng, C.; Li, T.; Yin, Y. Impact of the gut microbiota on intestinal immunity mediated by tryptophan metabolism. Front. Cell. Infect. Microbiol. 2018, 8, 13. [Google Scholar] [CrossRef]
- Zheng, D.; Liwinski, T.; Elinav, E. Interaction between microbiota and immunity in health and disease. Cell Res. 2020, 30, 492–506. [Google Scholar] [CrossRef] [PubMed]
- Clemente, J.C.; Ursell, L.K.; Parfrey, L.W.; Knight, R. The impact of the gut microbiota on human health: An integrative view. Cell 2012, 148, 1258–1270. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ridaura, V.K.; Faith, J.J.; Rey, F.E.; Knight, R.; Gorden, J.I. The effect of diet on the human gut microbiome: A metagenomic analysis in humanized gnotobiotic mice. Sci. Transl. Med. 2009, 1, 6ra14. [Google Scholar] [CrossRef]
- Cao, Y.; Liu, H.; Qin, N.; Ren, X.; Zhu, B.; Xia, X. Impact of food additives on the composition and function of gut microbiota: A review. Trends Food Sci. Technol. 2020, 99, 295–310. [Google Scholar] [CrossRef]
- Zhang, N.; Ju, Z.; Zuo, T. Time for food: The impact of diet on gut microbiota and human health. Nutrition 2018, 51, 80–85. [Google Scholar] [CrossRef]
- Dömling, A.; Li, X. TNF-α: The shape of small molecules to come? Drug Discov. Today 2022, 27, 3–7. [Google Scholar] [CrossRef]
- Yang, L.; Xie, X.; Tu, Z.; Fu, J.; Xu, D.; Zhou, Y. The signal pathways and treatment of cytokine storm in COVID-19. Signal Transduct. Target. Ther. 2021, 6, 255. [Google Scholar] [CrossRef]
- Andrews, C.; McLean, M.H.; Durum, S.K. Cytokine tuning of intestinal epithelial function. Front. Immunol. 2018, 9, 1270. [Google Scholar] [CrossRef]
- Marcone, S.; Haughton, K.; Simpson, P.J.; Belton, O.; Fitzgerald, D.J. Milk-derived bioactive peptides inhibit human endothelial-monocyte interactions via PPAR-γ dependent regulation of NF-κB. J. Inflamm. 2015, 12, 1–13. [Google Scholar] [CrossRef]
- Tak, P.P.; Firestein, G.S. NF-κB: A key role in inflammatory diseases. J. Clin. Investig. 2001, 107, 7–11. [Google Scholar] [CrossRef] [PubMed]
- Losada Mendez, J.; Palomares, F.; Gómez, F.; Ramírez-López, P.; Ramos-Soriano, J.; Torres, M.J.; Mayorga, C.; Rojo, J. Immunomodulatory Response of Toll-like Receptor Ligand–Peptide Conjugates in Food Allergy. ACS Chem. Biol. 2021, 16, 2651–2664. [Google Scholar] [CrossRef] [PubMed]
- Rong, J.; Zheng, H.; Liu, M.; Hu, X.; Wang, T.; Zhang, X.; Jin, F.; Wang, L. Probiotic and anti-inflammatory attributes of an isolate Lactobacillus helveticus NS8 from Mongolian fermented koumiss. BMC Microbiol. 2015, 15, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Saraiva, M.; O’garra, A. The regulation of IL-10 production by immune cells. Nat. Rev. Immunol. 2010, 10, 170–181. [Google Scholar] [CrossRef]
- Istanbullugil, F.R.; Risvanli, A.; Salikov, R.; Bayraktar, M.; Zhunushova, A.; Acaroz, U.; Acaroz, D.A.; Yilmaz, O.; Yuksel, B.F.; Turanli, M.; et al. Koumiss and immunity: A thorough investigation of fermentation parameters and their impact on health benefits. J. Dairy Sci. 2024, 107, 6451–6459. [Google Scholar] [CrossRef]
- Simonenko, E.S.; Simonenko, S.V.; Gmoshinski, I.V.; Riger, N.A.; Shumakova, A.A.; Zorin, S.N. Influence of lactoferrin and enzymatic hydrolysates of cow’s and mare’s milk proteins on anaphylactic sensitivity and cytokine profile of rats. Vopr. Pitan. 2024, 93, 31–40. [Google Scholar] [CrossRef]
- Vendrig, J.C.; Coffeng, L.E.; Fink-Gremmels, J. Equine colostral carbohydrates reduce lipopolysaccharide-induced inflammatory responses in equine peripheral blood mononuclear cells. Equine Vet. J. 2012, 44, 68–72. [Google Scholar] [CrossRef]
- Li, N.; Xie, Q.; Chen, Q.; Evivie, S.E.; Liu, D.; Dong, J.; Huo, G.; Li, B. Cow, goat, and mare milk diets differentially modulated the immune system and gut microbiota of mice colonized by healthy infant feces. J. Agric. Food Chem. 2020, 68, 15345–15357. [Google Scholar] [CrossRef]
- Donovan, S.M. The role of lactoferrin in gastrointestinal and immune development and function: A preclinical perspective. J. Pediatr. 2016, 173, S16–S28. [Google Scholar] [CrossRef]
- Actor, J.K.; Hwang, S.A.; Kruzel, M.L. Lactoferrin as a natural immune modulator. Curr. Pharm. Des. 2009, 15, 1956–1973. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Jin, S.; Chen, H.; Cao, Y.; Dong, X.; Li, H.; Zhou, Z.; Liu, C. Dose effect of bovine lactoferrin fortification on diarrhea and respiratory tract infections in weaned infants with anemia: A randomized, controlled trial. Nutrition 2021, 90, 111288. [Google Scholar] [CrossRef]
- Mao, B.; He, Z.; Chen, Y.; Stanton, C.; Ross, R.P.; Zhao, J.; Chen, W.; Yang, B. Effects of Bifidobacterium with the Ability of 2′-Fucosyllactose Utilization on Intestinal Microecology of Mice. Nutrients 2022, 14, 5392. [Google Scholar] [CrossRef]
- Tan, H.; O’Toole, P.W. Impact of diet on the human intestinal microbiota. Curr. Opin. Food Sci. 2015, 2, 71–77. [Google Scholar] [CrossRef]
- Panwar, D.; Shubhashini, A.; Kapoor, M. Complex alpha and beta mannan foraging by the human gut bacteria. Biotechnol. Adv. 2023, 66, 108166. [Google Scholar] [CrossRef]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, T.; Takahashi, M.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef]
- Arpaia, N.; Campbell, C.; Fan, X.; Dikiy, S.; van der Veeken, J.; deRoos, P.; Liu, H.; Cross, J.R.; Pfeffer, K.; Coffer, P.J.; et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 2013, 504, 451–455. [Google Scholar] [CrossRef] [PubMed]
- Zeng, W.; Wu, J.; Xie, H.; Xu, H.; Liang, D.; He, Q.; Yang, X.; Liu, C.; Gong, J.; Zhang, Q.; et al. Enteral nutrition promotes the remission of colitis by gut bacteria-mediated histidine biosynthesis. eBioMedicine 2024, 100, 2352–3964. [Google Scholar] [CrossRef]
- Yue, Y.; Wang, Y.; Yang, R.; Liu, D.; Cheng, Y.; Li, S.; Sun, L.; Li, B.; Huo, G. Exopolysaccharides of Bifidobacterium longum subsp. infantis E4 on the immune and anti-inflammatory effects in vitro. J. Funct. Foods 2023, 107, 105699. [Google Scholar] [CrossRef]
- Wang, H.; Fan, C.; Zhao, Z.; Zhai, Z.; Hao, Y. Anti-inflammatory effect of Bifidobacterium animalis subsp. lactis A6 on DSS-induced colitis in mice. J. Appl. Microbiol. 2022, 133, 2063–2073. [Google Scholar] [CrossRef]
- Nakajima, A.; Sasaki, T.; Itoh, K.; Kitahara, T.; Takema, Y.; Hiramatsu, K.; Ishikawa, D.; Shibuya, T.; Kobayashi, O.; Osada, T.; et al. A soluble fiber diet increases Bacteroides fragilis group abundance and immunoglobulin A production in the gut. Appl. Environ. Microbiol. 2020, 86, e00405-20. [Google Scholar] [CrossRef]
- Zheng, C.; Zhong, Y.; Xie, J.; Wang, Z.; Zhang, W.; Pi, Y.; Zhang, W.; Liu, L.; Luo, J.; Xu, W. Bacteroides acidifaciens and its derived extracellular vesicles improve DSS-induced colitis. Front. Microbiol. 2023, 14, 1304232. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.Y.; Xu, X.; Zhou, Y. Research progress on carbohydrate active enzymes of human microbiome. West China J. Stomatol. 2019, 37, 666–670. [Google Scholar]
- Baba, Y.; Tsuge, D.; Aoki, R. Enhancement of carbohydrate metabolism by probiotic and prebiotic intake promotes short-chain fatty acid production in the gut microbiome: A randomized, double-blind, placebo-controlled crossover trial. Biosci. Biotechnol. Biochem. 2025, 89, 1191–1202. [Google Scholar] [CrossRef]
- He, J.; Zhang, P.; Shen, L.; Niu, L.; Tan, Y.; Chen, L.; Zhao, Y.; Bai, L.; Hao, X.; Li, X.; et al. Short-chain fatty acids and their association with signalling pathways in inflammation, glucose and lipid metabolism. Int. J. Mol. Sci. 2020, 21, 6356. [Google Scholar] [CrossRef]
- Tedelind, S.; Westberg, F.; Kjerrulf, M.; Vidal, A. Anti-inflammatory properties of the short-chain fatty acids acetate and propionate: A study with relevance to inflammatory bowel disease. World J. Gastroenterol. 2007, 13, 2826. [Google Scholar] [CrossRef]
- Chambers, E.S.; Preston, T.; Frost, G.; Morrison, D.J. Role of gut microbiota-generated short-chain fatty acids in metabolic and cardiovascular health. Curr. Nutr. Rep. 2018, 7, 198–206. [Google Scholar] [CrossRef]
- Zhou, D.; Wu, Y.; Yan, H.; Shen, T.; Li, S.; Gong, J.; Li, G.; Mai, H.; Wang, D.; Tan, X. Gallic acid ameliorates calcium oxalate crystal-induced renal injury via upregulation of Nrf2/HO-1 in the mouse model of stone formation. Phytomedicine 2022, 106, 154429. [Google Scholar] [CrossRef] [PubMed]
- Lavelle, A.; Sokol, H. Gut microbiota-derived metabolites as key actors in inflammatory bowel disease. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 223–237. [Google Scholar] [CrossRef]
- Wan, Y.; Yang, L.; Jiang, S.; Qian, D.; Duan, J. Excessive apoptosis in ulcerative colitis: Crosstalk between apoptosis, ROS, ER stress, and intestinal homeostasis. Inflamm. Bowel Dis. 2022, 28, 639–648. [Google Scholar] [CrossRef] [PubMed]
- Ma, N.; Ma, X. Dietary amino acids and the gut-microbiome-immune axis: Physiological metabolism and therapeutic prospects. Compr. Rev. Food Sci. Food Saf. 2019, 18, 221–242. [Google Scholar] [CrossRef]
- Seo, S.K.; Kwon, B. Immune regulation through tryptophan metabolism. Exp. Mol. Med. 2023, 55, 1371–1379. [Google Scholar] [CrossRef]
- Qazi, A. The Role of the Serotonin Transporter-Aryl Hydrocarbon Receptor Axis in Intestinal Inflammation; University of Illinois: Chicago, IL, USA, 2024. [Google Scholar]
- Zhu, J.J.; Van den Eynde, B. AHR and tryptophan metabolism: A collaborative dynamics of immune regulation. Genes Immun. 2024, 25, 170–171. [Google Scholar] [CrossRef]



| Nutritional Composition | Protein (%) | Fat (%) | Lactose (%) | Mineral (%) |
|---|---|---|---|---|
| mare milk (MM) | 2.47 | 1.11 | 6.52 | 0.66 |
| fermented mare milk (K) | 2.39 | 1.35 | 1.13 | 0.49 |
| pasteurized fermented mare milk (PK) | 2.32 | 1.02 | 1.24 | 0.46 |
| Group | Administration |
|---|---|
| sterile distilled water (DW) | Sterile Distilled Water |
| mare milk (MM) | Mare Milk |
| fermented mare milk (K) | Fermented mare milk |
| pasteurized fermented mare milk (PK) | Pasteurized Fermented mare milk |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, R.; Ren, W.; Liu, S.; Li, Z.; Li, L.; Ma, S.; Yao, X.; Meng, J.; Zeng, Y.; Wang, J. Metagenomic Analysis Reveals the Anti-Inflammatory Properties of Mare Milk. Int. J. Mol. Sci. 2025, 26, 8239. https://doi.org/10.3390/ijms26178239
Wang R, Ren W, Liu S, Li Z, Li L, Ma S, Yao X, Meng J, Zeng Y, Wang J. Metagenomic Analysis Reveals the Anti-Inflammatory Properties of Mare Milk. International Journal of Molecular Sciences. 2025; 26(17):8239. https://doi.org/10.3390/ijms26178239
Chicago/Turabian StyleWang, Ran, Wanlu Ren, Shibo Liu, Zexu Li, Luling Li, Shikun Ma, Xinkui Yao, Jun Meng, Yaqi Zeng, and Jianwen Wang. 2025. "Metagenomic Analysis Reveals the Anti-Inflammatory Properties of Mare Milk" International Journal of Molecular Sciences 26, no. 17: 8239. https://doi.org/10.3390/ijms26178239
APA StyleWang, R., Ren, W., Liu, S., Li, Z., Li, L., Ma, S., Yao, X., Meng, J., Zeng, Y., & Wang, J. (2025). Metagenomic Analysis Reveals the Anti-Inflammatory Properties of Mare Milk. International Journal of Molecular Sciences, 26(17), 8239. https://doi.org/10.3390/ijms26178239






