Prevalence of Selected Polymorphisms of Il7R, CD226, CAPSL, and CLEC16A Genes in Children and Adolescents with Autoimmune Thyroid Diseases
Abstract
1. Introduction
2. Results
3. Discussion
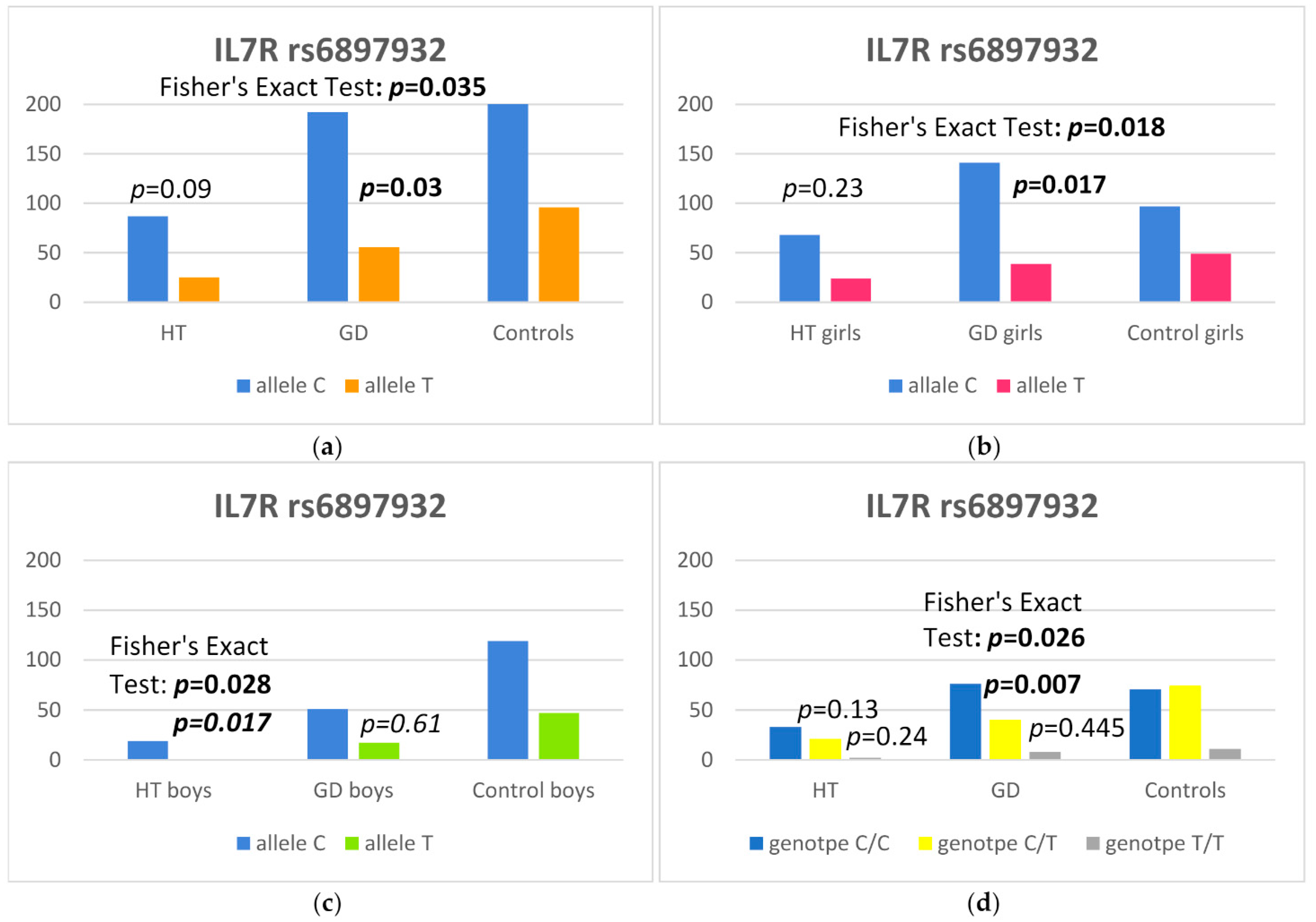
3.1. IL7R
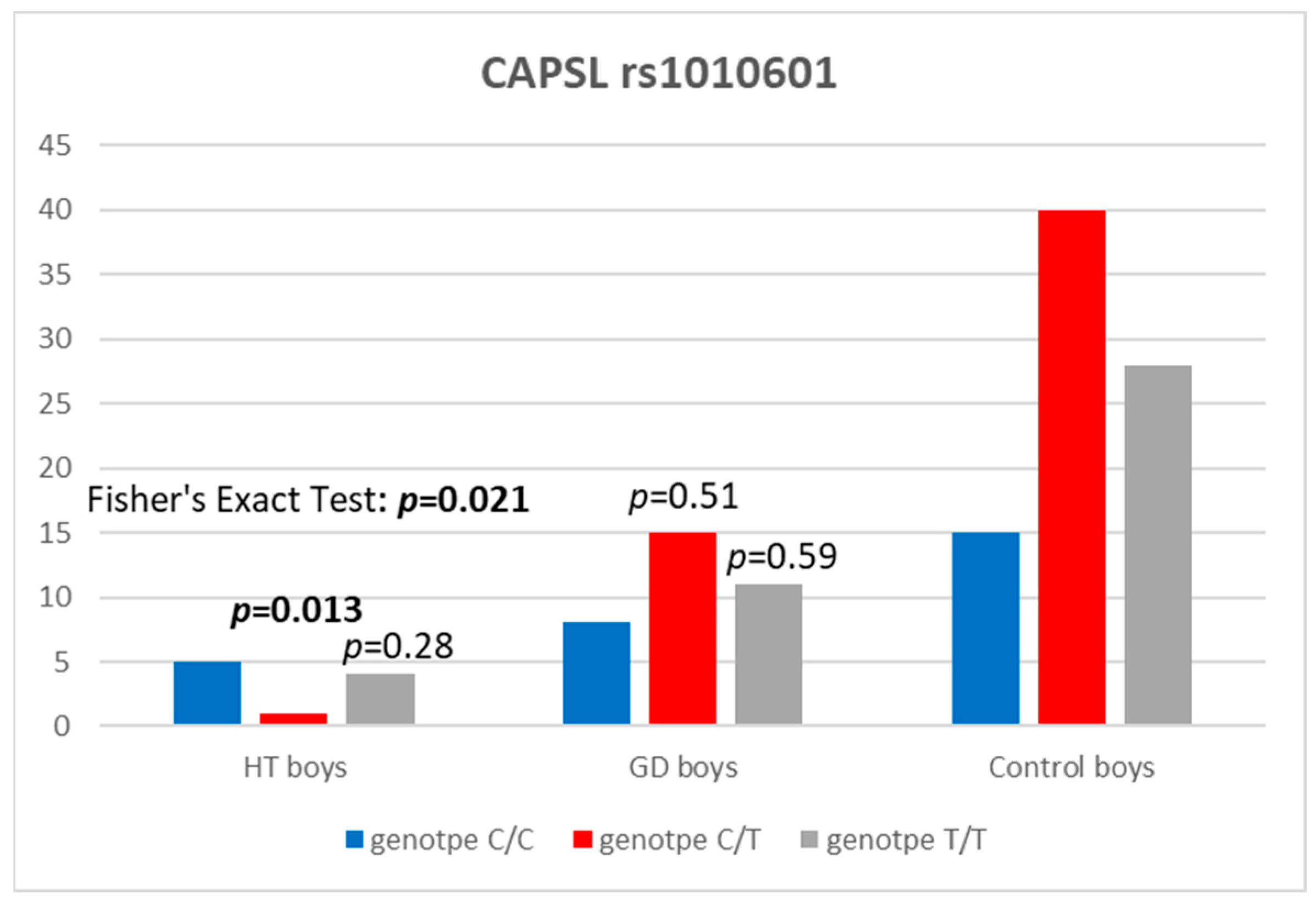
3.2. CAPSL
3.3. CD226
3.4. CLEC16A
4. Materials and Methods
4.1. Patients
4.2. Blood Analysis
4.3. DNA Extraction
4.4. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Rose, N.R. Prediction and Prevention of Autoimmune Disease in the 21st Century: A Review and Preview. Am. J. Epidemiol. 2016, 183, 403–406. [Google Scholar] [CrossRef]
- Krzewska, A.; Ben-Skowronek, I. Effect of Associated Autoimmune Diseases on Type 1 Diabetes Mellitus Incidence and Metabolic Control in Children and Adolescents. Biomed. Res. Int. 2016, 2016, 6219730. [Google Scholar] [CrossRef]
- Brown, R.S. Autoimmune thyroiditis in childhood. J. Clin. Res. Pediatr. Endocrinol. 2013, 5, 45–49. [Google Scholar] [CrossRef]
- Bossowski, A.; Borysewicz-Sańczyk, H.; Wawrusiewicz-Kurylonek, N.; Zasim, A.; Szalecki, M.; Wikiera, B.; Barg, E.; Myśliwiec, M.; Kucharska, A.; Bossowska, A.; et al. Analysis of chosen polymorphisms in FoxP3 gene in children and adolescents with autoimmune thyroid diseases. Autoimmunity 2014, 47, 395–400. [Google Scholar] [CrossRef]
- Wasniewska, M.G.; Bossowski, A. Special Issue: Autoimmune Disease Genetics. Genes 2021, 12, 1937. [Google Scholar] [CrossRef]
- Shastry, B.S. SNPs: Impact on gene function and phenotype. Methods Mol. Biol. 2009, 578, 3–22. [Google Scholar] [CrossRef]
- Fadason, T.; Farrow, S.; Gokuladhas, S.; Golovina, E.; Nyaga, D.; O’Sullivan, J.M.; Schierding, W. Assigning function to SNPs: Considerations when interpreting genetic variation. Semin. Cell Dev. Biol. 2022, 121, 135–142. [Google Scholar] [CrossRef]
- Borysewicz-Sańczyk, H.; Sawicka, B.; Wawrusiewicz-Kurylonek, N.; Głowińska-Olszewska, B.; Kadłubiska, A.; Gościk, J.; Szadkowska, A.; Łosiewicz, A.; Młynarski, W.; Kretowski, A.; et al. Genetic Association Study of IL2RA, IFIH1, and CTLA-4 Polymorphisms With Autoimmune Thyroid Diseases and Type 1 Diabetes. Front. Pediatr. 2020, 8, 481. [Google Scholar] [CrossRef]
- Sawicka, B.; Borysewicz-Sańczyk, H.; Wawrusiewicz-Kurylonek, N.; Aversa, T.; Corica, D.; Gościk, J.; Krętowski, A.; Waśniewska, M.; Bossowski, A. Analysis of Polymorphisms rs7093069-IL-2RA, rs7138803-FAIM2, and rs1748033-PADI4 in the Group of Adolescents with Autoimmune Thyroid Diseases. Front. Endocrinol. 2020, 11, 544658. [Google Scholar] [CrossRef]
- Barata, J.T.; Durum, S.K.; Seddon, B. Flip the coin: IL-7 and IL-7R in health and disease. Nat. Immunol. 2019, 20, 1584–1593. [Google Scholar] [CrossRef]
- Wang, C.; Kong, L.; Kim, S.; Lee, S.; Oh, S.; Jo, S.; Jang, I.; Kim, T.D. The Role of IL-7 and IL-7R in Cancer Pathophysiology and Immunotherapy. Int. J. Mol. Sci. 2022, 23, 10412. [Google Scholar] [CrossRef]
- Yasunaga, M. Antibody therapeutics and immunoregulation in cancer and autoimmune disease. Semin. Cancer Biol. 2020, 64, 1–12. [Google Scholar] [CrossRef]
- Pickens, S.R.; Chamberlain, N.D.; Volin, M.V.; Pope, R.M.; Talarico, N.E.; Mandelin, A.M., 2nd; Shahrara, S. Characterization of interleukin-7 and interleukin-7 receptor in the pathogenesis of rheumatoid arthritis. Arthritis Rheum. 2011, 63, 2884–2893. [Google Scholar] [CrossRef]
- Mazzucchelli, R.; Durum, S.K. Interleukin-7 receptor expression: Intelligent design. Nat. Rev. Immunol. 2007, 7, 144–154. [Google Scholar] [CrossRef]
- Hoffmann, M.; Enczmann, J.; Balz, V.; Kummer, S.; Reinauer, C.; Döing, C.; Förtsch, K.; Welters, A.; Kohns Vasconcelos, M.; Mayatepek, E.; et al. Interleukin-7 and soluble Interleukin-7 receptor levels in type 1 diabetes—Impact of IL7RA polymorphisms, HLA risk genotypes and clinical features. Clin. Immunol. 2022, 235, 108928. [Google Scholar] [CrossRef]
- Qiu, K.; Li, K.; Zeng, T.; Liao, Y.; Min, J.; Zhang, N.; Peng, M.; Kong, W.; Chen, L.L. Integrative Analyses of Genes Associated with Hashimoto’s Thyroiditis. J. Immunol. Res. 2021, 2021, 8263829. [Google Scholar] [CrossRef]
- Dooms, H. Interleukin-7: Fuel for the autoimmune attack. J. Autoimmun. 2013, 45, 40–48. [Google Scholar] [CrossRef]
- Marković, I.; Savvides, S.N. Modulation of Signaling Mediated by TSLP and IL-7 in Inflammation, Autoimmune Diseases, and Cancer. Front. Immunol. 2020, 11, 1557. [Google Scholar] [CrossRef]
- Santiago, J.L.; Alizadeh, B.Z.; Martínez, A.; Espino, L.; de la Calle, H.; Fernández-Arquero, M.; Figueredo, M.A.; de la Concha, E.G.; Roep, B.O.; Koeleman, B.P.; et al. Study of the association between the CAPSL-IL7R locus and type 1 diabetes. Diabetologia 2008, 51, 1653–1658. [Google Scholar] [CrossRef]
- Xi, J.S.; Nie, C.L.; Wang, J.; Ma, Y.; Ma, A.H. Association of CD226 polymorphisms with the susceptibility to type 1 diabetes in Chinese children. Genet. Mol. Res. 2015, 14, 15249–15255. [Google Scholar] [CrossRef]
- Todd, J.A.; Walker, N.M.; Cooper, J.D.; Smyth, D.J.; Downes, K.; Plagnol, V.; Bailey, R.; Nejentsev, S.; Field, S.F.; Payne, F.; et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat. Genet. 2007, 39, 857–864. [Google Scholar] [CrossRef]
- Shapiro, M.R.; Yeh, W.I.; Longfield, J.R.; Gallagher, J.; Infante, C.M.; Wellford, S.; Posgai, A.L.; Atkinson, M.A.; Campbell-Thompson, M.; Lieberman, S.M.; et al. CD226 Deletion Reduces Type 1 Diabetes in the NOD Mouse by Impairing Thymocyte Development and Peripheral T Cell Activation. Front. Immunol. 2020, 11, 2180. [Google Scholar] [CrossRef]
- Bai, L.; Jiang, J.; Li, H.; Zhang, R. Role of CD226 Rs763361 Polymorphism in Susceptibility to Multiple Autoimmune Diseases. Immunol. Investig. 2020, 49, 926–942. [Google Scholar] [CrossRef]
- Qiu, Z.X.; Zhang, K.; Qiu, X.S.; Zhou, M.; Li, W.M. CD226 Gly307Ser association with multiple autoimmune diseases: A meta-analysis. Hum. Immunol. 2013, 74, 249–255. [Google Scholar] [CrossRef]
- Huang, Z.; Qi, G.; Miller, J.S.; Zheng, S.G. CD226: An Emerging Role in Immunologic Diseases. Front. Cell Dev. Biol. 2020, 8, 564. [Google Scholar] [CrossRef]
- Pandey, R.; Bakay, M.; Hakonarson, H. CLEC16A-An Emerging Master Regulator of Autoimmunity and Neurodegeneration. Int. J. Mol. Sci. 2023, 24, 8224. [Google Scholar] [CrossRef]
- Frommer, L.; Kahaly, G.J. Type 1 Diabetes and Autoimmune Thyroid Disease-The Genetic Link. Front. Endocrinol. 2021, 12, 618213. [Google Scholar] [CrossRef]
- Hakonarson, H.; Grant, S.F.; Bradfield, J.P.; Marchand, L.; Kim, C.E.; Glessner, J.T.; Grabs, R.; Casalunovo, T.; Taback, S.P.; Frackelton, E.C.; et al. A genome-wide association study identifies KIAA0350 as a type 1 diabetes gene. Nature 2007, 448, 591–594. [Google Scholar] [CrossRef]
- Muhali, F.; Cai, T.T.; Zhu, J.L.; Qin, Q.; Xu, J.; He, S.T.; Shi, X.H.; Jiang, W.J.; Xiao, L.; Li, D.F.; et al. Polymorphisms of CLEC16A region and autoimmune thyroid diseases. G3 Genes Genomes Genet. 2014, 4, 973–977. [Google Scholar] [CrossRef]
- Mazzucchelli, R.I.; Riva, A.; Durum, S.K. The human IL-7 receptor gene: Deletions, polymorphisms and mutations. Semin. Immunol. 2012, 24, 225–230. [Google Scholar] [CrossRef]
- Lundtoft, C.; Seyfarth, J.; Oberstrass, S.; Rosenbauer, J.; Baechle, C.; Roden, M.; Holl, R.W.; Mayatepek, E.; Kummer, S.; Meissner, T.; et al. Autoimmunity risk- and protection-associated IL7RA genetic variants differentially affect soluble and membrane IL-7Rα expression. J. Autoimmun. 2019, 97, 40–47. [Google Scholar] [CrossRef]
- Barros, P.O.; Berthoud, T.K.; Aloufi, N.; Angel, J.B. Soluble IL-7Rα/sCD127 in Health, Disease, and Its Potential Role as a Therapeutic Agent. Immunotargets Ther. 2021, 10, 47–62. [Google Scholar] [CrossRef]
- Paysan-Lafosse, T.; Blum, M.; Chuguransky, S.; Grego, T.; Pinto, B.L.; Salazar, G.A.; Bileschi, M.L.; Bork, P.; Bridge, A.; Colwell, L.; et al. InterPro in 2022. Nucleic Acids Res. 2023, 51, D418–D427. [Google Scholar] [CrossRef]
- Franz, M.; Rodriguez, H.; Lopes, C.; Zuberi, K.; Montojo, J.; Bader, G.D.; Morris, Q. GeneMANIA update 2018. Nucleic Acids Res. 2018, 46, W60–W64. [Google Scholar] [CrossRef]
- Maiti, A.K.; Kim-Howard, X.; Viswanathan, P.; Guillén, L.; Qian, X.; Rojas-Villarraga, A.; Sun, C.; Cañas, C.; Tobón, G.J.; Matsuda, K.; et al. Non-synonymous variant (Gly307Ser) in CD226 is associated with susceptibility to multiple autoimmune diseases. Rheumatology 2010, 49, 1239–1244. [Google Scholar] [CrossRef]
- Hafler, J.P.; Maier, L.M.; Cooper, J.D.; Plagnol, V.; Hinks, A.; Simmonds, M.J.; Stevens, H.E.; Walker, N.M.; Healy, B.; Howson, J.M.; et al. CD226 Gly307Ser association with multiple autoimmune diseases. Genes Immun. 2009, 10, 5–10. [Google Scholar] [CrossRef]
- Gan, E.H.; Mitchell, A.L.; Macarthur, K.; Pearce, S.H. The role of a nonsynonymous CD226 (DNAX-accessory molecule-1) variant (Gly 307Ser) in isolated Addison’s disease and autoimmune polyendocrinopathy type 2 pathogenesis. Clin. Endocrinol. 2011, 75, 165–168. [Google Scholar] [CrossRef]
- Awata, T.; Kawasaki, E.; Tanaka, S.; Ikegami, H.; Maruyama, T.; Shimada, A.; Nakanishi, K.; Kobayashi, T.; Iizuka, H.; Uga, M.; et al. Association of type 1 diabetes with two Loci on 12q13 and 16p13 and the influence coexisting thyroid autoimmunity in Japanese. J. Clin. Endocrinol. Metab. 2009, 94, 231–235. [Google Scholar] [CrossRef]
- Lydersen, S.; Fagerland, M.W.; Laake, P. Recommended tests for association in 2 × 2 tables. Statist. Med. 2009, 28, 1159–1175. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Lewontin, R.C. The interaction of selection and linkage. I. General considerations; heterotic models. Genetic 1964, 49, 49–67. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2012; Available online: http://www.R-project.org/ (accessed on 4 April 2020).
- Faul, F.; Erdfelder, E.; Buchner, A.; Lang, A.-G. Statistical power analyses using G*Power 3.1: Tests for correlation and regression analyses. Behav. Res. Methods 2009, 41, 1149–1160. [Google Scholar] [CrossRef]
- Warnes G with Contributions from Gorjanc G Leisch F and Man M. Genetics: Population Genetics. R Package Version 1.3.8.1.2 2019. Available online: https://CRAN.R-project.org/package=genetics (accessed on 4 April 2020).
| HT (Mean ± SD) | p * | GD (Mean ± SD) | p ** | Controls (Mean ± SD) | |
|---|---|---|---|---|---|
| n (F/M) | 56 (46/10) | 124 (90/34) | 156 (73/83) | ||
| Age (years) | 15.2 ± 2.2 | ns | 16.5 ± 2 | ns | 16.3 ± 3 |
| Weight (kg) | 58 ± 5.28 | ns | 55.19 ± 2.39 | ns | 60.9 ± 7.8 |
| Height (cm) | 154.26 ± 4.14 | ns | 162.19 ± 2.69 | ns | 160 ± 8 |
| BMI (kg/m2) | 24.45 ± 1.33 | ns | 21.1 ± 2.1 | 0.012 | 23.78 ± 2.5 |
| TSH (mIU/L) | 9.87 ± 4.37 | <0.025 | 0.37 ± 0.1 | <0.01 | 3.04 ± 0.72 |
| fT4 (ng/dL) | 1.21 ± 0.03 | ns | 3.6 ± 1.4 | <0.001 | 1.1 ± 0.17 |
| fT3 (pg/mL) | 3.08 ± 0.5 | ns | 7.19 ± 1.65 | <0.001 | 3.79 ± 0.18 |
| TR-Ab (IU/L) a | 0.5 ± 0.32 | ns | 11.56 ± 2.11 | <0.001 | 0.4 ± 0.2 |
| Anti-TPO Ab (IU/mL) | 329.91 ± 92.93 | <0.001 | 331.97 ± 58.12 | <0.001 | 26.72 ± 6.8 |
| Anti-TG Ab (IU/mL) | 620.98 ± 240.34 | <0.001 | 347.49 ± 86.7 | <0.001 | 41.64 ± 12.1 |
| Treatment | L-thyroxine | Methimazole | - |
| GD, n (%) | Controls, n (%) | OR (95% CI) | p | |
|---|---|---|---|---|
| IL7R rs3194051 | ||||
| A | 161 (65) | 203 (65) | 1 (reference) | |
| G | 87 (35) | 109 (35) | 1.01 (0.70–1.43) | 0.971 |
| AA | 57 (46.0) | 73(46.8) | 1 (reference) | |
| AG | 47 (37.9) | 57 (36.5) | 1.05 (0.63–1.78) | 0.838 |
| GG | 20 (16.1) | 26 (16.6) | 0.98 (0.49–1.95) | 0.969 |
| IL7R rs6897932 | ||||
| C | 192 (77.4) | 216 (69.2) | 1.52 (1.04–2.24) | 0.030 |
| T | 56 (22.6) | 96 (30.8) | 1 (reference) | |
| CC | 76 (61.3) | 71 (45.5) | 1 (reference) | |
| CT | 40 (32.2) | 74 (47.4) | 0.51 (0.30–0.84) | 0.007 |
| TT | 8 (6.5) | 11 (7.1) | 0.68 (0.25–1.80) | 0.445 |
| CD226 rs763361 | ||||
| C | 136 (55.0) | 173 (55.0) | 1 (reference) | |
| T | 112 (45.0) | 139 (45.0) | 1.02 (0.73–1.43) | 0.88 |
| CC | 43 (34.7) | 53 (34.0) | 1 (reference) | |
| CT | 50 (40.3) | 67 (43.0) | 0.92 (0.53–1.59) | 0.765 |
| TT | 31 (25.0) | 36 (23.0) | 1.06 (0.56–1.99) | 0.853 |
| CAPSL rs1010601 | ||||
| C | 111 (45.0) | 123 (39.4) | 1.24 (0.89–1.75) | |
| T | 137 (55.0) | 189 (60.6) | 1 (reference) | 0.205 |
| C | 1 (0.8) | 0 (0) | ||
| CC | 31 (25.0) | 28 (17.9) | 1.1 (0.03–44.50) | 0.950 |
| CT | 47 (37.9) | 67 (42.9) | 0.7 (0.02–27.92) | 0.827 |
| TT | 45 (36.3) | 61 (39.1) | 0.7 (0.02–29.38) | 0.851 |
| CLEC16A rs725613 | ||||
| G | 79 (31.9) | 101 (32.4) | 1 (reference) | |
| T | 169 (68.1) | 211 (67.6) | 1.02 (0.72–1.47) | 0.898 |
| GG | 10 (8.0) | 15 (9.6) | 1 (reference) | |
| GT | 59 (47.6) | 71 (45.5) | 1.24 (0.52–3.07) | 0.632 |
| TT | 55 (44.3) | 70 (44.9) | 1.17 (0.49–2.91) | 0.724 |
| HT, n (%) | Controls, n (%) | OR (95% CI) | p | |
|---|---|---|---|---|
| IL7R rs3194051 | ||||
| A | 72 (64.3) | 203 (65.1) | 1 (reference) | |
| G | 40 (35.7) | 109 (34.9) | 1.03 (0.65–1.62) | 0.88 |
| AA | 24 (42.8) | 73 (46.8) | 1 (reference) | |
| AG | 24 (42.8) | 57 (36.5) | 1.28 (0.65–2.50) | 0.47 |
| GG | 8 (14.3) | 26 (16.7) | 0.94 (0.35–2.32) | 0.90 |
| IL7R rs6897932 | ||||
| C | 87 (77.7) | 216 (69.2) | 1.5 (0.94–2.60) | 0.09 |
| T | 25 (22.3) | 96 (30.8) | 1 (reference) | |
| CC | 33 (59.0) | 71 (45.5) | 1 (reference) | |
| CT | 21 (37.5) | 74 (47.4) | 0.61 (0.32–1.16) | 0.13 |
| TT | 2 (3.5) | 11 (7.1) | 0.42 (0.06–1.70) | 0.24 |
| CD226 rs763361 | ||||
| C | 51 (45.5) | 173 (55.4) | 1 (reference) | |
| T | 6154.5) | 139 44.6) | 1.49 (0.96–2.30) | 0.07 |
| CC | 11 (19.6) | 53 (34.0) | 1 (reference) | |
| CT | 29 (51.8) | 67(42.9) | 2.06 (0.96–4.70) | 0.06 |
| TT | 16 (28.6) | 36 (23.1) | 2.12 (088–5.26) | 0.09 |
| CAPSL rs1010601 | ||||
| C | 45 (40.2) | 123 (39.4) | 1.03 (0.66–1.60) | 0.89 |
| T | 67 (59.8) | 189 60.6) | 1 (reference) | |
| CC | 11 (19.6) | 28 (17.9) | 1 (reference) | |
| CT | 23 (41.1) | 67 (42.9) | 0.87 (0.38–2.09) | 0.75 |
| TT | 22 (39.3) | 61 (39.1) | 0.91 (0.39–2.21) | 0.84 |
| CLEC16A rs725613 | ||||
| G | 31 (27.7) | 101 (32.4) | 1 (reference) | |
| T | 81 (72.3) | 211 (67.6) | 1.25 (0.78–2.03) | 0.36 |
| GG | 3 (5.4) | 15 (9.6) | 1 (reference) | |
| GT | 25 (44.6) | 71 (45.5) | 1.69 (0.50–8.13) | 0.42 |
| TT | 28 (5.0) | 70 (44.9) | 1.92 (0.57–9.19) | 0.31 |
| IL7R rs6897932 | GD Girls, n (%) | Controls, n (%) | OR (95% CI) | p |
|---|---|---|---|---|
| C | 141 (78.3) | 97 (66.4) | 1.82 (1.11–3.00) | 0.017 |
| T | 39 (21.7) | 49 (33.6) | 1 (reference) | |
| CC | 56 (62.3) | 30 (41.1) | 1 (reference) | |
| CT | 29 (32.2) | 37 (50.7) | 0.42 (0.21–0.81) | 0.010 |
| TT | 5 (5.5) | 6 (8.2) | 0.45 (0.12–1.67) | 0.223 |
| IL7R rs6897932 | HT Boys, n (%) | Controls, n (%) | OR (95% CI) | p |
|---|---|---|---|---|
| C | 19 (95) | 119 (71.7) | 6.58 (1.30–160.93) | 0.017 |
| T | 1 (5) | 47 (28.3) | 1 (reference) | |
| CC | 9 (90) | 41 (49.4) | 1 (reference) | |
| CT | 1 (1) | 37 (44.6) | 0.14 (0.005–0.82) | 0.026 |
| TT | 0 (0) | 5 (6) | 0.99 (0.03–7.68) | 0.99 |
| CAPSL rs1010601 | HT Boys, n (%) | Controls, n (%) | OR (95% CI) | p |
|---|---|---|---|---|
| C | 11 (55) | 70 (42.2) | 1.66 (0.65–4.40) | 0.29 |
| T | 9 (45) | 96 (57.8) | 1 (reference) | |
| CC | 5 (50) | 15 (18.1) | 1 (reference) | |
| CT | 1 (10) | 40 (48.2) | 0.086 (0.003–0.623) | 0.013 |
| TT | 4 (40) | 28 (33.7) | 0.44 (0.09–1.97) | 0.279 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Borysewicz-Sańczyk, H.; Wawrusiewicz-Kurylonek, N.; Gościk, J.; Sawicka, B.; Bossowski, F.; Corica, D.; Aversa, T.; Waśniewska, M.; Bossowski, A. Prevalence of Selected Polymorphisms of Il7R, CD226, CAPSL, and CLEC16A Genes in Children and Adolescents with Autoimmune Thyroid Diseases. Int. J. Mol. Sci. 2024, 25, 4028. https://doi.org/10.3390/ijms25074028
Borysewicz-Sańczyk H, Wawrusiewicz-Kurylonek N, Gościk J, Sawicka B, Bossowski F, Corica D, Aversa T, Waśniewska M, Bossowski A. Prevalence of Selected Polymorphisms of Il7R, CD226, CAPSL, and CLEC16A Genes in Children and Adolescents with Autoimmune Thyroid Diseases. International Journal of Molecular Sciences. 2024; 25(7):4028. https://doi.org/10.3390/ijms25074028
Chicago/Turabian StyleBorysewicz-Sańczyk, Hanna, Natalia Wawrusiewicz-Kurylonek, Joanna Gościk, Beata Sawicka, Filip Bossowski, Domenico Corica, Tommaso Aversa, Małgorzata Waśniewska, and Artur Bossowski. 2024. "Prevalence of Selected Polymorphisms of Il7R, CD226, CAPSL, and CLEC16A Genes in Children and Adolescents with Autoimmune Thyroid Diseases" International Journal of Molecular Sciences 25, no. 7: 4028. https://doi.org/10.3390/ijms25074028
APA StyleBorysewicz-Sańczyk, H., Wawrusiewicz-Kurylonek, N., Gościk, J., Sawicka, B., Bossowski, F., Corica, D., Aversa, T., Waśniewska, M., & Bossowski, A. (2024). Prevalence of Selected Polymorphisms of Il7R, CD226, CAPSL, and CLEC16A Genes in Children and Adolescents with Autoimmune Thyroid Diseases. International Journal of Molecular Sciences, 25(7), 4028. https://doi.org/10.3390/ijms25074028









