Olive Pomace Extract Contains Low Molecular Weight Peptides and Possesses ACE Inhibitory Activity
Abstract
1. Introduction
2. Results and Discussion
2.1. Preparation of a Water-Soluble LMW Fraction from Olive Pomace Extract
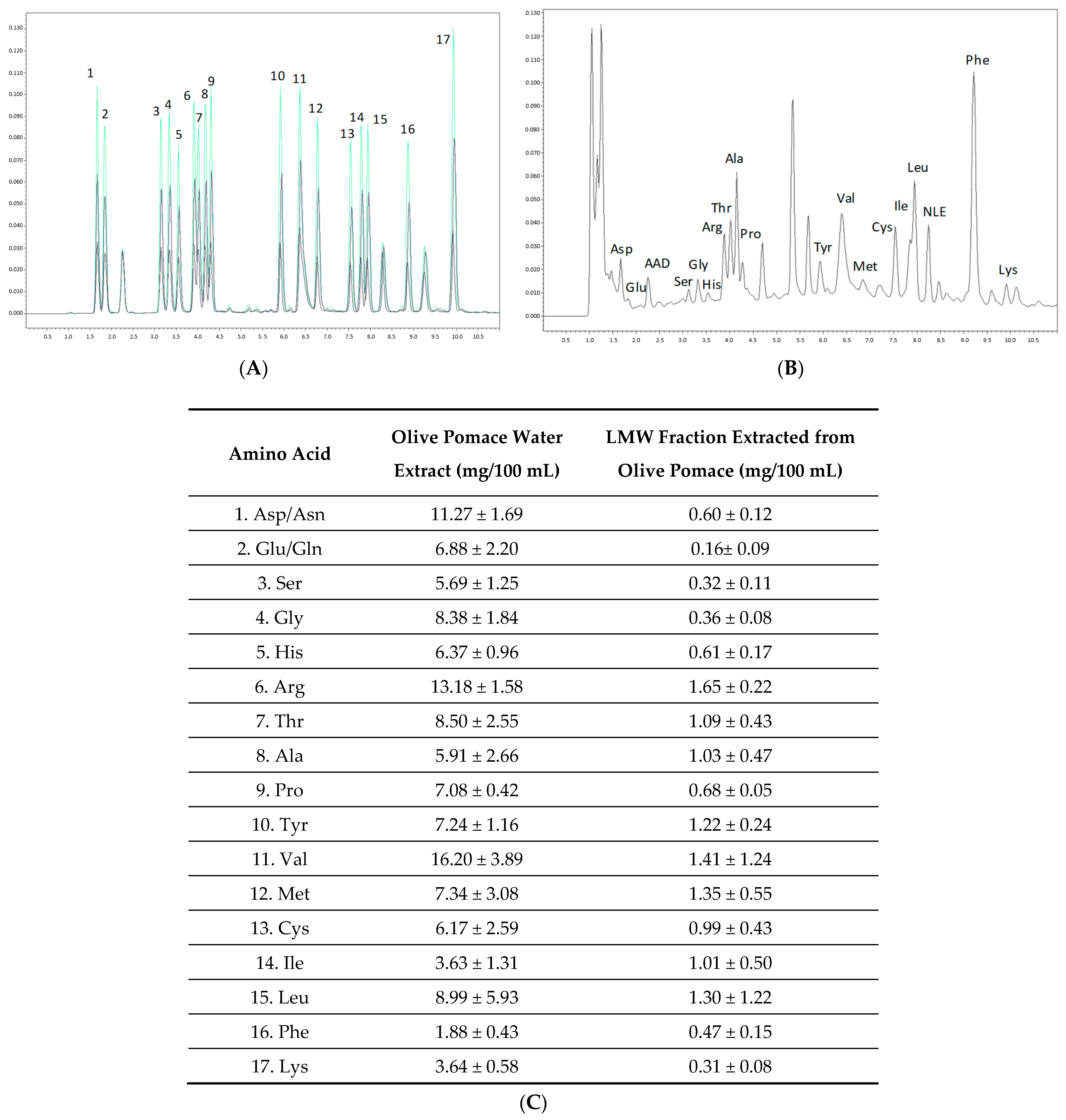
2.2. Amino Acid Analysis and Native PAGE of Olive Pomace LMW Extract
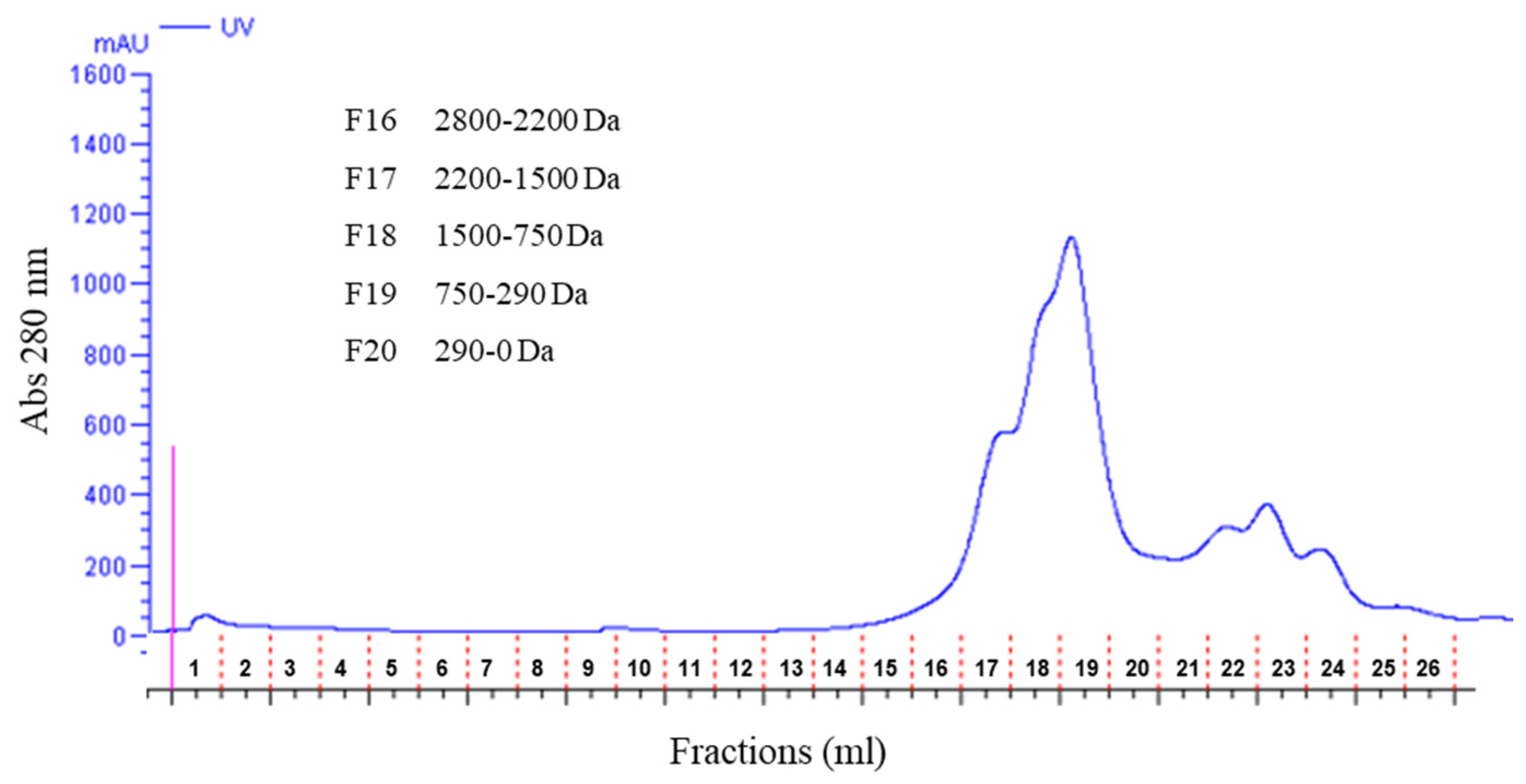
2.3. Fractionation of Peptides by Size-Exclusion Chromatography
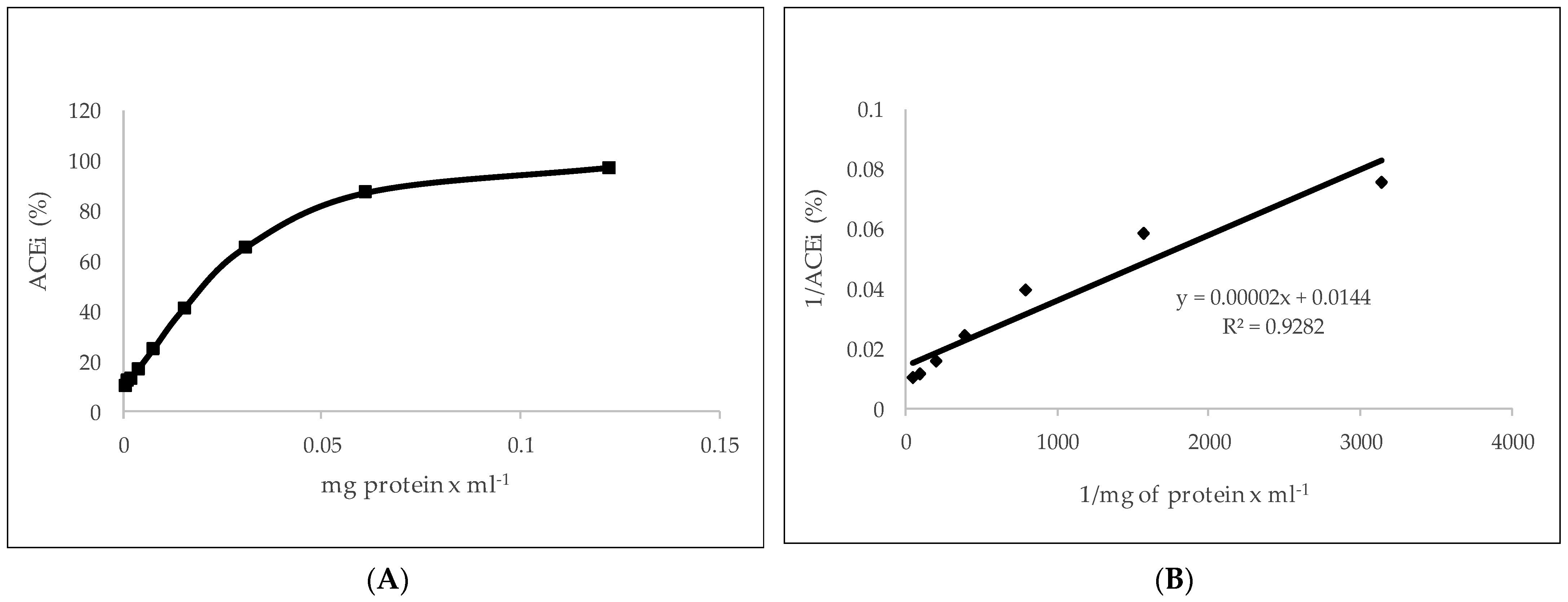
2.4. ACE Inhibitory Activity Determinations
2.5. Endogenous Peptides and Proteins Identified by Peaks DB and De Novo Sequencing in Olive Pomace
2.6. Predicted Activities of Olive Pomace Endogenous Peptides
3. Materials and Methods
3.1. Plant Material and Extraction of Olive Oil
3.2. Preparation of Olive Pomace Extract Containing Low-Molecular Weight Peptides
3.3. Analysis and Quantification of Amino Acids
3.4. Detection of Peptides and Antioxidant Activity of Olive Pomace LMW Peptides in Native Polyacrylamide Gels
3.5. Extraction of Polyphenols from Olive Pomace Extracts
3.6. Analysis of Minerals by Inductively Coupled Plasma-Optical Emission Spectrometry (ICP-OES)
3.7. Fractionation of Olive Pomace Peptides by Size-Exclusion Chromatography
3.8. Determination of Angiotensin-Converting Enzyme Inhibitory Activity
3.9. LC-Orbitrap MS/MS Analysis of Olive Pomace LMW Peptide Fraction
3.10. Peptide and Protein Identification by Database Search and De Novo Sequencing
3.11. Biological Activity Prediction of Identified Peptides from Olive Pomace Peptides
3.12. Determination of Total Nitrogen, Fat, Carbohydrates, and Polyphenols
3.13. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rivas-Garcia, L.; Navarro-Hortal, M.D.; Romero-Marquez, J.M.; Llopis, J.; Forbes-Hernández, T.Y.; Xiao, J.; Quiles, J.L.; Sanchez-Gonzalez, C. Valorization of Olea europaea and olive oil processing by-products/wastes. Adv. Food Nutr. Res. 2023, 107, 193–212. [Google Scholar] [PubMed]
- Alburquerque, J.A.; Gonzálvez, J.; García, D.; Cegarra, J. Agrochemical characterisation of ‘‘alperujo’’, a solid by-product of the two-phase centrifugation method for olive oil extraction. Bioresour. Technol. 2004, 91, 195–200. [Google Scholar]
- Ravindran, R.; Jaiswal, A.K. Exploitation of Food Industry Waste for High-Value Products. Trends Biotechnol. 2016, 34, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Clemente, A.; Sánchez-Vioque, R.; Vioque, J.; Bautista, J.; Millán, F. Chemical composition of extracted dried olive pomaces containing two and three phases. Food Biotechnol. 1997, 11, 273–291. [Google Scholar] [CrossRef]
- Karpouzas, D.G.; Ntougias, S.; Iskidou, E.; Rousidou, C.; Papadopoulou, K.K.; Zervakis, G.; Ehaliotis, C. Olive mill wastewater affects the structure of soil bacterial communities. Appl. Soil Ecol. 2010, 45, 101–111. [Google Scholar] [CrossRef]
- Rana, G.; Rinaldi, M.; Introna, M. Volatilisation of substances after spreading olive oil waste water on the soil in a Mediterranean environment. Agric. Ecosyst. Environ. 2003, 96, 49–58. [Google Scholar] [CrossRef]
- Dermechea, S.; Nadour, M.; Larroche, C.; Moulti-Mati, F.; Michaudb, P. Olive mill wastes: Biochemical characterizations and valorization strategies. Process Biochem. 2013, 48, 1532–1552. [Google Scholar] [CrossRef]
- Roselló-Soto, E.; Moubarik, K.; Lopez, R.P.; Saraiva, J.A.; Boussetta, N.; Barba, F.J. Emerging opportunities for the effective valorization of wastes and by-products generated during olive oil production process: Non-conventional methods for the recovery of high-added value compounds. Trends Food Sci. Technol. 2015, 45, 296–310. [Google Scholar] [CrossRef]
- Nunes, M.A.; Pimentel, F.B.; Costa, A.S.G.; Alves, R.C.; Oliveira, M.B.P.P. Olive by-products for functional and food applications: Challenging opportunities to face environmental constraints. Innov. Food Sci. Emerg. 2016, 35, 139–148. [Google Scholar] [CrossRef]
- Luján, R.J.; Capote, F.P.; Marinas, A.; Luque de Castro, M.D. Liquid chromatography/triple quadrupole tandem mass spectrometry with multiple reaction monitoring for optimal selection of transitions to evaluate nutraceuticals from olive-tree materials. Rapid Commun. Mass Spectrom. 2008, 22, 855–864. [Google Scholar] [CrossRef]
- Parra, A.; López, P.E.; García-Granados, A. Bioactive compounds with added value prepared from terpenes contained in solid wastes from the olive oil industry. Chem. Biodivers. 2010, 7, 421–439. [Google Scholar] [CrossRef] [PubMed]
- Karantonis, H.C.; Tsantila, N.; Stamatakis, G.; Samiotaki, M.; Panayotou, G.; Antonopoulou, S.; Demopoulos, C.A. Bioactive polar lipids in olive oil, pomace and waste byproducts. J. Food Biochem. 2008, 32, 443–459. [Google Scholar] [CrossRef]
- Bartolomei, M.; Li, J.; Capriotti, A.L.; Fanzaga, M.; d’Adduzio, L.; Laganà, A.; Cerrato, A.; Mulinacci, N.; Cecchi, L.; Bollati, C.; et al. Olive (Olea europaea L.) seed as new source of cholesterol-lowering bioactive peptides: Elucidation of their mechanism of action in HepG2 cells and their trans-epithelial transport in differentiated caco-2 cells. Nutrients 2024, 16, 371. [Google Scholar] [CrossRef] [PubMed]
- Esteve, C.; Marina, M.L.; García, M.C. Novel strategy for the revalorization of olive (Olea europaea) residues based on the extraction of bioactive peptides. Food Chem. 2015, 167, 272–280. [Google Scholar] [CrossRef] [PubMed]
- Difonzo, G.; Troilo, M.; Squeo, G.; Pasqualone, A.; Caponio, F. Functional compounds from olive pomace to obtain high-added value foods—A review. J. Sci. Food Agric. 2021, 101, 15–26. [Google Scholar] [CrossRef] [PubMed]
- Delgado, A.; Cholevas, C.; Theoharides, T.C. Neuroinflammation in Alzheimer’s disease and beneficial action of luteolin. Biofactors 2021, 47, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, C.S.; Adedara, I.A.; Farombi, E.O.; Emanuelli, T. Nutraceutical potential of olive pomace: Insights from cell-based and clinical studies. J. Sci. Food Agric. 2024; Published online ahead of print. [Google Scholar] [CrossRef]
- World Health Organisation. A Global Brief on Hypertension. Silent Killer, Global Public Health Crisis; Document number WHO/DCO/WHD/2013.2; WHO Press: Geneva, Switzerland, 2013. [Google Scholar]
- Chobanian, A.V.; Bakris, G.L.; Black, H.R.; Cushman, W.C.; Green, L.A.; Izzo, J.L., Jr.; Jones, D.W.; Materson, B.J.; Oparil, S.; Wright, J.T., Jr.; et al. The Seventh Report of the Joint National Committee on Prevention, Detection, Evaluation, and Treatment of High Blood Pressure: The JNC 7 report. JAMA 2003, 289, 2560–2572. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Ledesma, B.; del Mar Contreras, M.; Recio, I. Antihypertensive peptides: Production, bioavailability and incorporation into foods. Adv. Colloid Interface Sci. 2011, 165, 23–35. [Google Scholar] [CrossRef] [PubMed]
- Saadi, S.; Saari, N.; Anwar, F.; Hamid, A.A.; Mohd Ghazali, H. Recent advances in food biopeptides: Production, biological functionalities and therapeutic applications. Biotechnol. Adv. 2015, 33, 80–116. [Google Scholar] [CrossRef]
- Escudero, E.; Mora, L.; Fraser, P.D.; Aristoy, M.C.; Arihara, K.; Toldrá, F. Purification and Identification of antihypertensive peptides in Spanish dry-cured ham. J. Proteom. 2013, 78, 499–507. [Google Scholar] [CrossRef]
- Mandal, S.M.; Bharti, R.; Porto, W.F.; Gauri, S.S.; Mandal, M.; Franco, O.L.; Ghosh, A.K. Identification of multifunctional peptides from human milk. Peptides 2014, 56, 84–93. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Huertas, E.; del Río, L.A. Characterization of antioxidant enzymes and peroxisomes of olive (Olea europaea L.) fruits. J. Plant Physiol. 2014, 171, 1463–1471. [Google Scholar] [CrossRef] [PubMed]
- Alcaide-Hidalgo, J.M.; Margalef, M.; Bravo, F.I.; Muguerza, B.; López-Huertas, E. Virgin olive oil (unfiltered) extract contains peptides and possesses ACE inhibitory and antihypertensive activity. Clin. Nutr. 2020, 39, 1242–1249. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Huertas, E.; Alcaide-Hidalgo, J.M. Characterisation of Endogenous Peptides Present in Virgin Olive Oil. Int. J. Mol. Sci. 2022, 23, 1712. [Google Scholar] [CrossRef] [PubMed]
- Alcaide-Hidalgo, J.M.; Romero, M.; Duarte, J.; López-Huertas, E. Antihypertensive Effects of Virgin Olive Oil (Unfiltered) Low Molecular Weight Peptides with ACE Inhibitory Activity in Spontaneously Hypertensive Rats. Nutrients 2020, 12, 271. [Google Scholar] [CrossRef] [PubMed]
- Prandi, B.; Faccini, A.; Lambertini, F.; Bencivenni, M.; Jorba, M.; Van Droogenbroek, B.; Bruggeman, G.; Schöber, J.; Petrusan, J.; Elst, K.; et al. Food wastes from agrifood industry as possible sources of proteins: A detailed molecular view on the composition of the nitrogen fraction, amino acid profile and racemisation degree of 39 food waste streams. Food Chem. 2019, 286, 567–575. [Google Scholar] [CrossRef] [PubMed]
- Sinrod, A.J.G.; Avena-Bustillos, R.J.; Olson, D.A.; Crawford, L.M.; Wang, S.C.; McHugh, T.H. Phenolics and antioxidant capacity of pitted olive pomace affected by three drying technologies. J. Food Sci. 2019, 84, 412–420. [Google Scholar] [CrossRef] [PubMed]
- Allouche, N.; Fki, I.; Sayadi, S. Toward a high yield recovery of antioxidants and purified hydroxytyrosol from olive mill wastewaters. J. Agric. Food Chem. 2004, 52, 267–273. [Google Scholar] [CrossRef] [PubMed]
- Cecchi, L.; Migliorini, M.; Giambanelli, E.; Canuti, V.; Bellumori, M.; Mulinacci, N.; Zanoni, B. Exploitation of virgin olive oil by-products (Olea europaea L.): Phenolic and volatile compounds transformations phenomena in fresh two-phase olive pomace (‘alperujo’) under different storage conditions. J. Sci. Food Agric. 2022, 102, 2515–2525. [Google Scholar] [CrossRef]
- Aludatt, M.H.; Alli, I.; Ereifej, K.; Alhamad, M.; Al-Tawaha, A.R.; Rababah, T. Optimisation, characterisation and quantification of phenolic compounds in olive cake. Food Chem. 2010, 123, 117–122. [Google Scholar] [CrossRef]
- Herregods, G.; Van Camp, J.; Morel, N.; Ghesquiere, B.; Gevaert, K.; Vercruysse, L.; Dierckx, S.; Quanten, E.; Smagghe, G. Angiotensin I-Converting Enzyme Inhibitory Activity of Gelatin Hydrolysates and Identification of Bioactive Peptides. J. Agric. Food Chem. 2011, 59, 552–558. [Google Scholar] [CrossRef] [PubMed]
- Hansen, K.; Adsersen, A.; Christensen, S.B.; Jensen, S.R.; Nyman, U.; Smitt, U.W. Isolation of an angiotensin converting enzyme (ACE) inhibitor from Olea europaea and Olea lancea. Phytomedicine 1996, 2, 319–325. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Aluko, R.E.; Nakai, S. Structural requirements of Angiotensin I-converting enzyme inhibitory peptides: Quantitative structure-activity relationship study of di- and tripeptides. J. Agric. Food Chem. 2006, 54, 732–738. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Zhang, K.; Hendrie, C.; Liang, C.; Li, M.; Doherty-Kirby, A.; Lajoie, G. PEAKS: Powerful Software for Peptide De Novo Sequencing by MS/MS. Rapid Commun. Mass Spectrom. 2003, 17, 2337–2342. [Google Scholar] [CrossRef] [PubMed]
- Natesh, R.; Schwager, S.L.U.; Sturrock, E.D.; Acharya, K.R. Crystal structure of the human angiotensin-converting enzymelisinopril complex. Nature 2003, 421, 551–554. [Google Scholar] [CrossRef] [PubMed]
- Elias, R.J.; Kellerby, S.S.; Decker, E.A. Antioxidant Activity of Proteins. Crit. Rev. Food Sci. Nutr. 2008, 48, 430–441. [Google Scholar] [CrossRef] [PubMed]
- Ryan, J.T.; Ross, R.P.; Bolton, D.; Fitzgerald, G.F.; Stanton, C. Bioactive peptides from muscle sources: Meat and fish. Nutrients 2011, 3, 765–791. [Google Scholar] [CrossRef] [PubMed]
- Maestri, E.; Marmiroli, M.; Marmiroli, N. Bioactive peptides in plant-derived foodstuffs. J. Proteom. 2016, 147, 140–155. [Google Scholar] [CrossRef] [PubMed]
- Erdmann, K.; Cheung, B.W.; Schröder, H. The possible roles of food-derived bioactive peptides in reducing the risk of cardiovascular disease. J. Nutr. Biochem. 2008, 19, 643–654. [Google Scholar] [CrossRef]
- Chen, H.M.; Muramoto, K.; Yamauchi, F.; Fujimoto, K.; Nokihara, K. Antioxidative properties of histidine-containing peptides designed from peptide fragments found in the digests of a soybean protein. J. Agric. Food. Chem. 1998, 46, 49–53. [Google Scholar] [CrossRef]
- Heres, A.; Yokoyama, I.; Gallego, M.; Toldrá, F.; Arihara, K.; Mora, L. Antihypertensive potential of sweet Ala-Ala dipeptide and its quantitation in dry-cured ham at different processing conditions. J. Funct. Foods 2021, 87, 104818. [Google Scholar] [CrossRef]
- Jha, S.; Taschler, U.; Domenig, O.; Poglitsch, M.; Bourgeois, B.; Pollheimer, M.; Pusch, L.M.; Malovan, G.; Frank, S.; Madl, T.; et al. Dipeptidyl peptidase 3 modulates the renin-angiotensin system in mice. J. Biol. Chem. 2020, 295, 13711–13723. [Google Scholar] [CrossRef] [PubMed]
- Deacon, C.F. Dipeptidyl peptidase 4 inhibitors in the treatment of type 2 diabetes mellitus. Nat. Rev. Endocrinol. 2020, 16, 642–653. [Google Scholar] [CrossRef]
- López-Huertas, E.; Lozano-Sánchez, J.; Segura-Carretero, A. Olive oil varieties and ripening stages containing the antioxidants hydroxytyrosol and derivatives in compliance with EFSA health claim. Food Chem. 2021, 342, 128291. [Google Scholar] [CrossRef]
- Cohen, S.A.; Meys, M.; Tarvin, T.L. The Pico-Tag Method. A Manual of Advanced Techniques for Amino Acid Analysis; Millipore Corporation: Bedford, MA, USA, 1989. [Google Scholar]
- Palma-Granados, P.; Lara, L.; Seiquer, I.; Aguilera, J.F.; Nieto, R. Genotype and dietary lysine deficiency affect carcass and muscle amino acid composition of pigs growing from 10 to 25 kg body weight. J. Anim. Physiol. Anim. Nutr. 2019, 103, 1857–1865. [Google Scholar] [CrossRef] [PubMed]
- Moore, S. On the determination of cystine as cysteic acid. J. Biol. Chem. 1963, 238, 235–237. [Google Scholar] [CrossRef]
- Ladner, C.L.; Yang, J.; Turner, R.J.; Edwards, R.A. Visible fluorescent detection of proteins in polyacrylamide gels without staining. Anal. Biochem. 2004, 326, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Beauchamp, C.O.; Fridovich, I. Superoxide dismutase improved assays and an assayapplicable to acrylamide gels. Anal. Biochem. 1971, 44, 276–287. [Google Scholar] [CrossRef]
- Bakircioglu, D.; Kurtulus, Y.B.; Ucar, G. Determination of some traces metal levels in cheese samples packaged in plastic and tin containers by ICP-OES after dry, wet and microwave digestion. Food Chem. Toxicol. 2011, 49, 202–207. [Google Scholar] [CrossRef]
- Sentandreu, M.A.; Toldrá, F. A fluorescence-based protocol for quantifying angiotensin-converting enzyme activity. Nat. Protoc. 2006, 1, 2423–2427. [Google Scholar] [CrossRef]
- Cruz, F.; Julca, I.; Gómez-Garrido, J.; Loska, D.; Marcet-Houben, M.; Cano, E.; Galán, B.; Frias, L.; Ribeca, P.; Derdak, S.; et al. Genome sequence of the olive tree, Olea europaea. GigaScience 2016, 5, 29. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Xin, L.; Shan, B.; Chen, W.; Xie, M.; Yuen, D.; Zhang, W.; Zhang, Z.; Gilles, A.; Lajoie, G.A.; et al. Peaks DB: De novo sequencing assisted database search for sensitive and accurate peptide identification. Mol. Cell. Proteom. 2012, 11, M111.010587. [Google Scholar] [CrossRef] [PubMed]
- Peaks Team. Peaks X Pro User Manual. 2020. Available online: https://www.bioinfor.com/user-manual/ (accessed on 27 March 2024).
- Stephen, F.; Altschul, T.L.; Madden, A.A.; Schäffer, J.Z.; Zheng, Z.; Webb, M.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar]
- Minkiewicz, P.; Dziuba, J.; Iwaniak, A.; Dziuba, M.; Darewicz, M. BIOPEP database and other programs for processing bioactive peptide sequences. J. AOAC Int. 2008, 91, 965–980. [Google Scholar] [CrossRef] [PubMed]
- BIOPEP-UWM: Bioactive Peptides. Available online: https://biochemia.uwm.edu.pl/en/biopep-uwm-2/ (accessed on 3 February 2024).
- Dziuba, J.; Iwaniak, A.; Minkiewicz, P. Computer-aided characteristics of proteins as potential precursors of bioactive peptides. Polimery 2003, 48, 50–53. [Google Scholar] [CrossRef]
- AOAC. AOAC official method 990.03, protein (crude) in animal feed, combustion method. In Official Methods of Analysis of AOAC International, 18th ed.; ASA-SSA Inc.: Gaithersburg, ML, USA, 2006. [Google Scholar]
- Spanos, G.A.; Wrolstad, R.E. Influence of processing and storage on the phenolic composition of thompson seedless grape juice. J. Agric. Food Chem. 1990, 38, 1565–1571. [Google Scholar] [CrossRef]
- Matissek, R.; Schnepel, F.M.; Steiner, G. Análisis de los Alimentos. Fundamentos, Métodos, Aplicaciones; Editorial Acribia, S.A.: Zaragoza, Spain, 1998. [Google Scholar]
- The Commission of the European Communities. Commission Regulation (EC) No 121/2008 of 11 February 2008 laying down the method of analysis for the determination of starch content in preparations of a kind used in animal feeding (CN code 2309). Off. J. Eur. Union 2008, L 37/3, 3–8. [Google Scholar]

| Sample | IC50 (µg prot/mL) |
|---|---|
| Olive pomace LMW fraction | 3.57 ± 0.22 |
| F16 | 365.8 ± 32.3 |
| F17 | 125.1 ± 33.1 |
| F18 | 75.6 ± 6.9 |
| F19 | 67.4 ± 14.5 |
| F20 | 68.52 ± 7.49 |
| No. | Peptide Sequence | −10lgP | N | m/z | RT | ppm | Area | Protein Source (BLAST) | Accession |
|---|---|---|---|---|---|---|---|---|---|
| 1 | NSNALACCSA | 34.52 | 10 | 477.1920 | 51.20 | 7.1 | 3.16 × 107 | anthocyanidin 3-O-glucosyltransferase | OE9A062433P1 |
| CAA2981081.1 | |||||||||
| 2 | GGDNGMNH | 31.98 | 8 | 401.1449 | 51.36 | −4.5 | 4.85 × 105 | phospholipase D gamma 1-like | OE9A085147P3 OE9A085147P2 |
| CAA2951337.1 | |||||||||
| 3 | RSCQHRY | 30.05 | 7 | 475.2191 | 21.09 | 0.4 | ND | putative RING-H2 finger protein ATL21A | OE9A113191P1 |
| XP_022855964.1 | |||||||||
| 4 | LPPDNSH | 29.26 | 7 | 390.1841 | 52.42 | 2.7 | 3.26 × 106 | Hypothetical predicted protein | OE9A032896P1 |
| CAA2978878.1 | |||||||||
| 5 | SLIPCCSS | 27.21 | 8 | 809.3359 | 54.15 | −6.6 | 6.69 × 106 | casein kinase 1 HD16 | OE9A059381P2 OE9A059381P1 OE9A059381P3 |
| CAA2934848.1 | |||||||||
| 6 | SGDPTM(+15.99)YE | 26.79 | 8 | 458.1616 | 50.95 | −11.8 | 2.68 × 105 | tryptophan aminotransferase-related 2 isoform X1 | OE9A048079P1 |
| CAA2969551.1 | |||||||||
| 7 | IDHGCAKH | 25.50 | 8 | 440.7025 | 42.72 | −1.2 | 1.65 × 106 | anthocyanidin 3-O-glucosyltransferase | OE9A062433P1 |
| CAA2981081.1 | |||||||||
| 8 | IPATDSQG | 25.47 | 8 | 394.6899 | 36.41 | 4.6 | 1.42 × 106 | auxin response factor 6 isoform X1 | OE9A083314P2 OE9A083314P3 OE9A083314P1 |
| CAA2976018.1 | |||||||||
| 9 | MCTAAEM(+15.99)K | 25.21 | 8 | 450.6838 | 48.32 | 9.9 | 4.66 × 105 | white-brown complex homolog 30 | OE9A053685P3 |
| CAA2969019.1 | |||||||||
| 10 | GGGPGGGLGGPS | 25.13 | 12 | 435.2055 | 45.44 | 3.6 | 3.10 × 106 | dihydrolipoyllysine-residue succinyl transferase component of 2-oxoglutarate de-hydrogenase | OE9A068558P1 |
| CAA3014561.1 | |||||||||
| 11 | QANACHH | 25.10 | 7 | 390.6582 | 33.30 | −2.5 | 2.59 × 105 | DUF946 domain-containing DUF1162 domain-containing Chorein | OE9A008961P3 |
| CAA2957578.1 | |||||||||
| 12 | NSCCVCMVR | 25.00 | 9 | 507.6946 | 42.64 | −4.6 | 1.31 × 107 | probable E3 ubiquitin-ligase LOG2 | OE9A009309P1 |
| CAA3023827.1 |
| No. | Peptide | ALC (%) | N | Z | m/z | RT | Area | ppm | Local Confidence (%) |
|---|---|---|---|---|---|---|---|---|---|
| 1 | LFYHHH | 90 | 6 | 2 | 427.2079 | 30.18 | 1.96 × 106 | −2.2 | 88 85 88 95 88 96 |
| 2 | GDHSGPPR | 89 | 8 | 2 | 411.6919 | 31.88 | 5.36 × 106 | −10.6 | 56 92 97 97 94 97 95 90 |
| 3 | YCFHSAA | 89 | 7 | 2 | 399.6636 | 32.80 | 4.63 × 105 | −5.0 | 96 95 91 88 84 85 88 |
| 4 | YTM(+15.99)M(+15.99)LF | 89 | 6 | 2 | 419.1794 | 23.63 | 4.06 × 106 | −0.8 | 92 90 92 76 93 93 |
| 5 | AEFENFVAFVDK | 88 | 12 | 2 | 708.3472 | 48.76 | 1.88 × 106 | 2.2 | 40 72 98 99 95 96 93 95 99 98 96 82 |
| 6 | RCYHEE | 87 | 6 | 2 | 418.6729 | 49.92 | ND | 3.6 | 84 93 90 85 85 86 |
| 7 | YVM(+15.99)FFL | 87 | 6 | 2 | 418.2032 | 29.26 | 1.31 × 106 | −8.0 | 98 94 92 87 71 80 |
| 8 | FFTYAY | 87 | 6 | 2 | 406.1843 | 36.79 | 2.90 × 105 | −5.9 | 85 85 90 90 85 86 |
| 9 | YEM(+15.99)YPY | 87 | 6 | 2 | 441.1741 | 25.78 | 3.42 × 107 | 2.7 | 82 86 80 92 82 97 |
| 10 | LSVSAFNC | 86 | 8 | 2 | 420.7043 | 28.04 | 6.54 × 105 | 11.2 | 93 94 94 90 92 93 69 69 |
| No. | Peptide Sequence | Predicted Activity | No. Seq. with Activity | A | B |
|---|---|---|---|---|---|
| 1 | NSNALACCSA | ACE inhibitor | 1 | 0.1000 | 0.0003 |
| Antioxidative | 1 | 0.1000 | |||
| DPP III inhibitor | 1 | 0.1000 | |||
| DPP IV inhibitor | 3 | 0.1000 | |||
| Activating ubiquitin-mediated proteolysis | 1 | 0.1000 | 0.0001 | ||
| 2 | GGDNGMNH | ACE inhibitor | 4 | 0.5 | 0.0001 |
| DPP IV inhibitor | 5 | 0.6250 | 0.0018 | ||
| 3 | RSCQHRY | ACE inhibitor | 1 | 0.1429 | 0.0136 |
| Antioxidative | 1 | 0.1429 | 9.72 × 10−6 | ||
| DPP IV inhibitor | 2 | 0.2857 | 8.47 × 10−5 | ||
| 4 | LPPDNSH | ACE inhibitor | 3 | 0.4286 | 0.0161 |
| alpha-glucosidase inhibitor | 1 | 0.1429 | |||
| DPP IV inhibitor | 4 | 0.5714 | 0.0018 | ||
| 5 | SLIPCCSS | ACE inhibitor (IP) | 1 | 0.125 | 0.001 |
| DPP IV inhibitor | 3 | 0.375 | 0.0003 | ||
| Regulator of phosphoglycerate kinase activity | 1 | 0.125 | |||
| Glucose uptake stimulating peptide | 1 | 0.125 | |||
| 6 | SGDPTM(+15.99)YE | ACE inhibitor | 4 | 0.2500 | 0.0001 |
| DPP IV inhibitor | 4 | 0.2500 | |||
| 7 | IDHGCAKH | ACE inhibitor | 1 | 0.125 | 1.98 × 10−5 |
| DPP IV inhibitor | 1 | 0.125 | |||
| antioxidative | 1 | 0.125 | |||
| 8 | IPATDSQG | ACE inhibitor | 3 | 0.375 | 0.0019 |
| DPP IV inhibitor | 6 | 0.75 | 0.0029 | ||
| 9 | MCTAAEM(+15.99)K | ACE inhibitor | 1 | 0.0625 | 0.0001 |
| hypotensive | 1 | 0.0625 | |||
| DPP IV inhibitor | 3 | 0.1875 | 6.64 × 10−6 | ||
| 10 | GGGPGGGLGGPS | ACE inhibitor | 5 | 0.8333 | 0.0008 |
| DPP IV inhibitor | 5 | 0.8333 | 4.91 × 10−5 | ||
| PAM inhibitor | 1 | 0.0833 | |||
| peptide regulating the stomach mucosal membrane activity | 3 | 0.3333 | |||
| Antithrombotic peptide | 3 | 0.3333 | |||
| Prolyl endopeptidase inhibitor (antiamnestic) | 3 | 0.3333 | |||
| 11 | QANACHH | antioxidative | 1 | 0.1429 | 3.11 × 10−2 |
| DPP IV inhibitor | 3 | 0.4286 | |||
| 12 | NSCCVCMVR | ACE inhibitor | 1 | 0.1111 | 0.0021 |
| DPP IV inhibitor | 2 | 0.2222 | 0.0001 |
| No. | De Novo Peptide | Predicted Activity | No. Seq. with Activity | A | B |
|---|---|---|---|---|---|
| 1 | LFYHHH | ACE inhibitor | 3 | 0.5 | 0.0396 |
| Antioxidative | 3 | 0.667 | |||
| DPP III inhibitor | 2 | 0.1667 | |||
| DPP IV inhibitor | 1 | 0.5000 | |||
| 2 | GDHSGPPR | ACE inhibitor | 7 | 0.875 | 0.0380 |
| Antioxidative | 1 | 0.1250 | |||
| DPP III inhibitor | 1 | 0.1250 | |||
| DPP IV inhibitor | 1 | 0.375 | |||
| antiamnestic | 1 | 0.1250 | |||
| antithrombotic | 1 | 0.1250 | |||
| regulating | 1 | 0.1250 | |||
| alpha-glucosidase inhibitor | 1 | 0.1250 | |||
| 3 | YCFHSAA | ACE inhibitor | 3 | 0.2857 | 0.0731 |
| hypotensive | 1 | 0.2857 | 1.519 × 10−5 | ||
| DPP IV inhibitor | 2 | 0.1429 | |||
| 4 | YTM(+15.99)M(+15.99)LF | ACE inhibitor | 2 | 0.3333 | 0.0008 |
| Antioxidative | 1 | 0.1667 | 0.0003 | ||
| DPP IV inhibitor | 4 | 0.6667 | 0.0036 | ||
| 5 | AEFENFVAFVDK | ACE inhibitor | 4 | 0.3333 | 0.0046 |
| DPP IV inhibitor | 5 | 0.0833 | 0.0005 | ||
| CaMPDE inhibitor | 1 | 0.4167 | |||
| hypolipidemic | 2 | 0.0833 | |||
| renin inhibitor | 1 | 0.0833 | |||
| 6 | RCYHEE | ACE inhibitor | 1 | 0.1667 | 0.0325 |
| DPP III inhibitor | 1 | 0.1667 | |||
| DPP IV inhibitor | 1 | 0.3333 | |||
| stimulating | 1 | 0.1667 | |||
| 7 | YVM(+15.99)FFL | ACE inhibitor | 5 | 0.8333 | 0.0113 |
| DPP III inhibitor | 5 | 0.1667 | |||
| DPP IV inhibitor | 1 | 0.8333 | 0.0006 | ||
| 8 | FFTYAY | ACE inhibitor | 4 | 0.5000 | 0.0119 |
| Antioxidative | 4 | 0.6667 | |||
| DPP IV inhibitor | 4 | 0.6667 | 0.0002 | ||
| renin inhibitor | 2 | 0.333 | |||
| 9 | YEM(+15.99)YPY | ACE inhibitor | 3 | 0.5000 | 0.0014 |
| Antioxidative | 1 | 0.1667 | |||
| DPP IV inhibitor | 5 | 0.8333 | 0.0007 | ||
| anti inflammatory | 1 | 0.1667 | |||
| alpha-glucosidase inhibitor | 2 | 0.3333 | 0.0001 | ||
| 10 | LSVSAFNC | ACE inhibitor | 1 | 0.1250 | 0.0007 |
| DPP IV inhibitor | 4 | 0.5000 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
López-Huertas, E.; Rubí-Villegas, J.; Sánchez-Moreno, L.; Nieto, R. Olive Pomace Extract Contains Low Molecular Weight Peptides and Possesses ACE Inhibitory Activity. Int. J. Mol. Sci. 2024, 25, 3962. https://doi.org/10.3390/ijms25073962
López-Huertas E, Rubí-Villegas J, Sánchez-Moreno L, Nieto R. Olive Pomace Extract Contains Low Molecular Weight Peptides and Possesses ACE Inhibitory Activity. International Journal of Molecular Sciences. 2024; 25(7):3962. https://doi.org/10.3390/ijms25073962
Chicago/Turabian StyleLópez-Huertas, Eduardo, Jose Rubí-Villegas, Lourdes Sánchez-Moreno, and Rosa Nieto. 2024. "Olive Pomace Extract Contains Low Molecular Weight Peptides and Possesses ACE Inhibitory Activity" International Journal of Molecular Sciences 25, no. 7: 3962. https://doi.org/10.3390/ijms25073962
APA StyleLópez-Huertas, E., Rubí-Villegas, J., Sánchez-Moreno, L., & Nieto, R. (2024). Olive Pomace Extract Contains Low Molecular Weight Peptides and Possesses ACE Inhibitory Activity. International Journal of Molecular Sciences, 25(7), 3962. https://doi.org/10.3390/ijms25073962







