Pluripotent Stem Cells as a Preclinical Cellular Model for Studying Hereditary Spastic Paraplegias
Abstract
1. Introduction
2. HSP-Related iPS Cell Lines
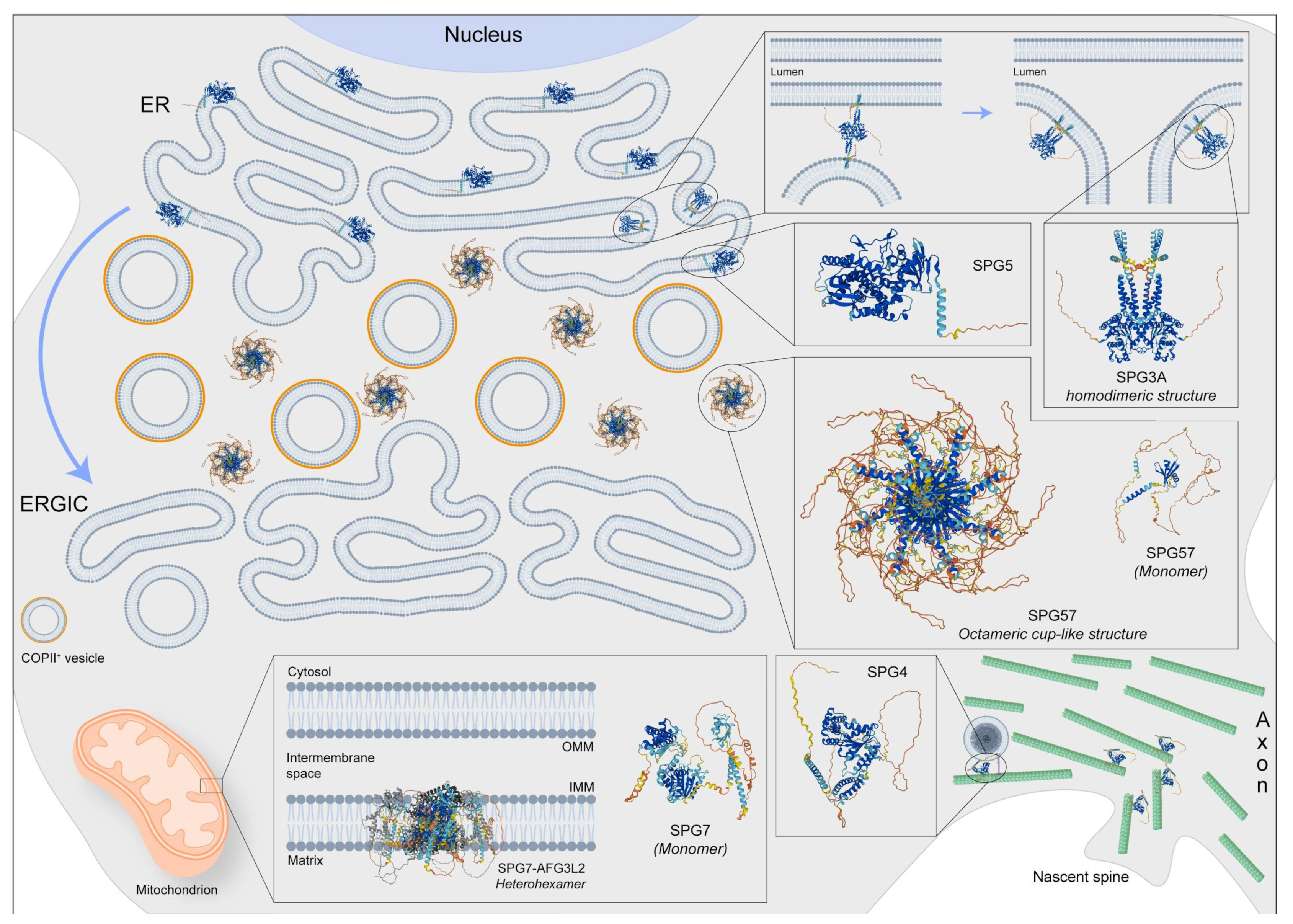
2.1. SPG3A
2.2. SPG4
2.3. SPG5
2.4. SPG7
2.5. SPG57
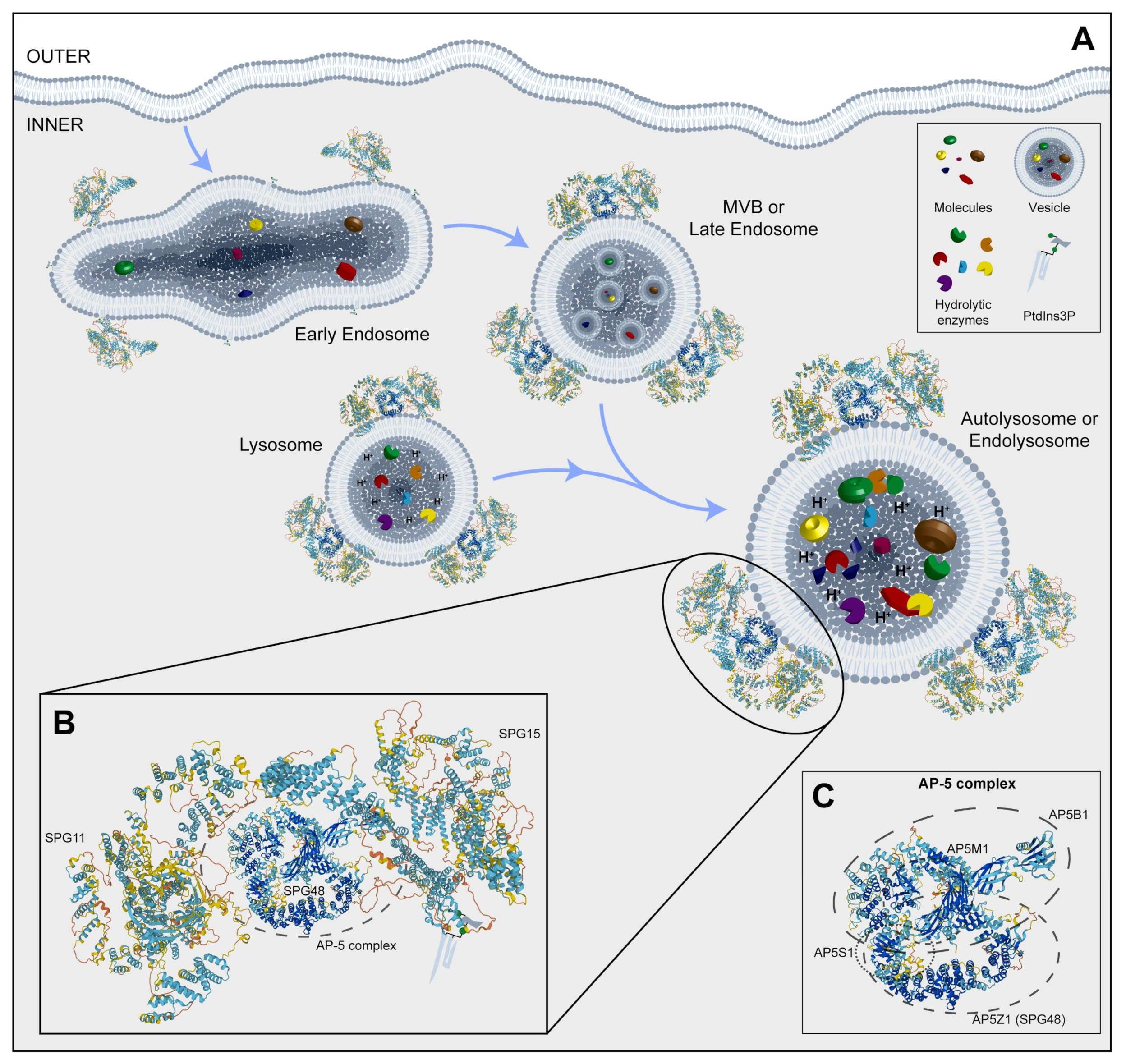
2.6. SPG11
2.7. SPG15 and SPG48
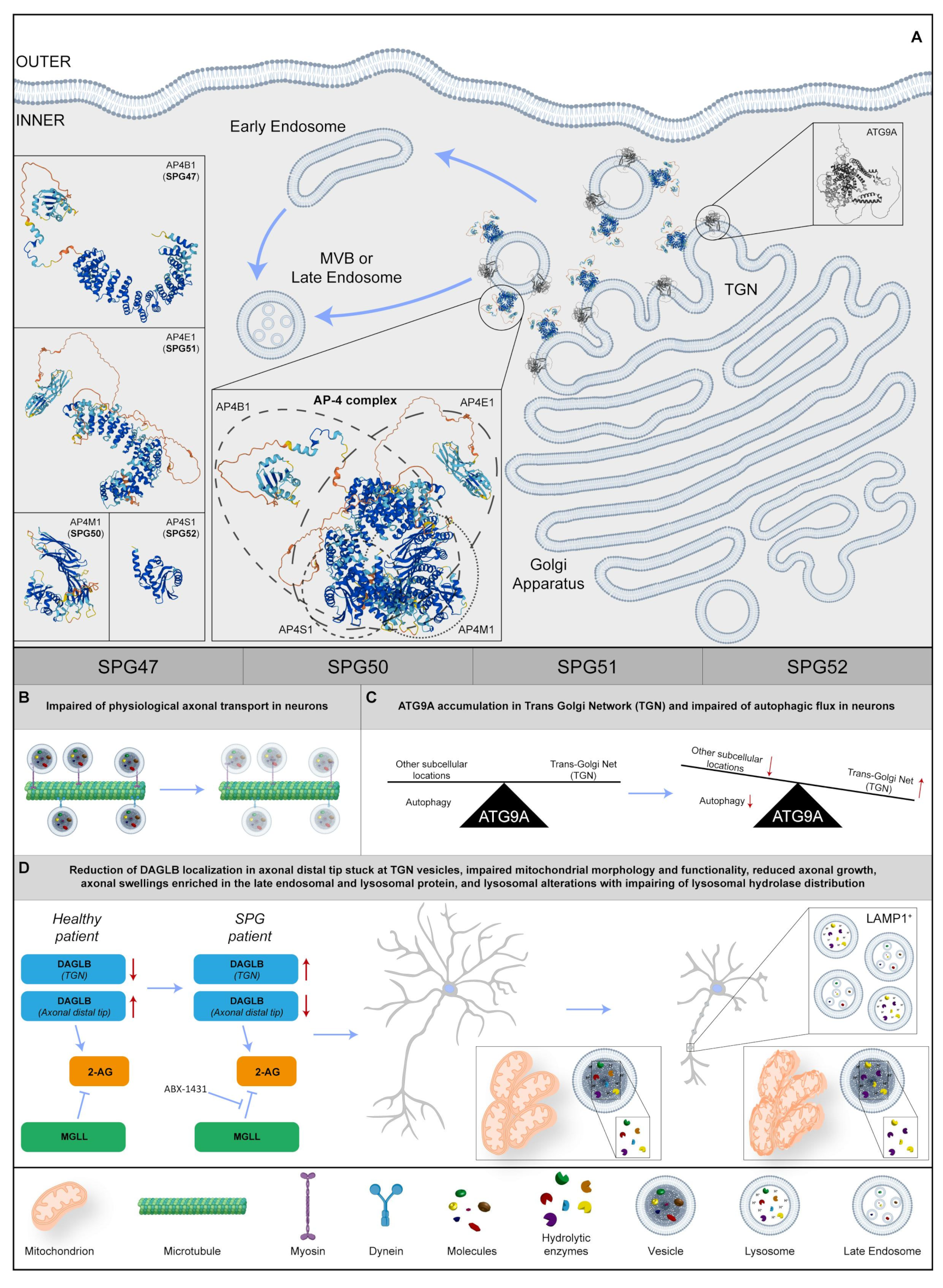
2.8. AP-4 Complex (SPG47, SPG50, SPG51, SPG52)
2.9. Other SPG Genes
3. Conclusions with a Look over the Horizon
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Faber, I.; Pereira, E.R.; Martinez, A.R.M.; França, M., Jr.; Teive, H.A.G. Hereditary Spastic Paraplegia from 1880 to 2017: An Historical Review. Arq. Neuropsiquiatr. 2017, 75, 813–818. [Google Scholar] [CrossRef]
- Elsayed, L.E.O.; Eltazi, I.Z.; Ahmed, A.E.; Stevanin, G. Insights into Clinical, Genetic, and Pathological Aspects of Hereditary Spastic Paraplegias: A Comprehensive Overview. Front. Mol. Biosci. 2021, 8, 690899. [Google Scholar] [CrossRef]
- Lo Giudice, T.; Lombardi, F.; Santorelli, F.M.; Kawarai, T.; Orlacchio, A. Hereditary Spastic Paraplegia: Clinical-Genetic Characteristics and Evolving Molecular Mechanisms. Exp. Neurol. 2014, 261, 518–539. [Google Scholar] [CrossRef]
- Meyyazhagan, A.; Orlacchio, A. Hereditary Spastic Paraplegia: An Update. Int. J. Mol. Sci. 2022, 23, 1697. [Google Scholar] [CrossRef]
- Ishiura, H.; Takahashi, Y.; Hayashi, T.; Saito, K.; Furuya, H.; Watanabe, M.; Murata, M.; Suzuki, M.; Sugiura, A.; Sawai, S.; et al. Molecular Epidemiology and Clinical Spectrum of Hereditary Spastic Paraplegia in the Japanese Population Based on Comprehensive Mutational Analyses. J. Hum. Genet. 2014, 59, 163–172. [Google Scholar] [CrossRef]
- Coutinho, P.; Barros, J.; Zemmouri, R.; Guimarães, J.; Alves, C.; Chorão, R.; Lourenço, E.; Ribeiro, P.; Loureiro, J.L.; Santos, J.V.; et al. Clinical Heterogeneity of Autosomal Recessive Spastic Paraplegias. Arch. Neurol. 1999, 56, 943. [Google Scholar] [CrossRef]
- Behan, W.M.H.; Maia, M. Strumpell’s Familial Spastic Paraplegia: Genetics and Neuropathology. J. Neurol. Neurosurg. Psychiatry 1974, 37, 8–20. [Google Scholar] [CrossRef]
- Bruyn, R.P.M. The Neuropathology of Hereditary Spastic Paraparesis. Clin. Neurol. Neurosurg. 1992, 94, 16–18. [Google Scholar] [CrossRef] [PubMed]
- Sack, G.H.; Huether, C.A.; Garg, N. Familial Spastic Paraplegia-Clinical and Pathologic Studies in a Large Kindred. Johns. Hopkins Med. J. 1978, 143, 117–121. [Google Scholar] [PubMed]
- Harding, A.E. Classification of the Hereditary Ataxias and Paraplegias. Lancet 1983, 1, 1151–1155. [Google Scholar] [CrossRef] [PubMed]
- Ruano, L.; Melo, C.; Silva, M.C.; Coutinho, P. The Global Epidemiology of Hereditary Ataxia and Spastic Paraplegia: A Systematic Review of Prevalence Studies. Neuroepidemiology 2014, 42, 174–183. [Google Scholar] [CrossRef]
- Riso, V.; Rossi, S.; Nicoletti, T.; Tessa, A.; Travaglini, L.; Zanni, G.; Aiello, C.; Perna, A.; Barghigiani, M.; Pomponi, M.; et al. Application of a Clinical Workflow May Lead to Increased Diagnostic Precision in Hereditary Spastic Paraplegias and Cerebellar Ataxias: A Single Center Experience. Brain Sci. 2021, 11, 246. [Google Scholar] [CrossRef] [PubMed]
- Boutry, M.; Morais, S.; Stevanin, G. Update on the Genetics of Spastic Paraplegias. Curr. Neurol. Neurosci. Rep. 2019, 19, 18. [Google Scholar] [CrossRef] [PubMed]
- Martinuzzi, A.; Blackstone, C.; O’Kane, C.J.; Stevanin, G. Editorial: Hereditary Spastic Paraplegias: At the Crossroads of Molecular Pathways and Clinical Options. Front. Neurosci. 2021, 15, 708642. [Google Scholar] [CrossRef] [PubMed]
- Dahlem, T.J.; Hoshijima, K.; Jurynec, M.J.; Gunther, D.; Starker, C.G.; Locke, A.S.; Weis, A.M.; Voytas, D.F.; Grunwald, D.J. Simple Methods for Generating and Detecting Locus-Specific Mutations Induced with TALENs in the Zebrafish Genome. PLoS Genet. 2012, 8, e1002861. [Google Scholar] [CrossRef] [PubMed]
- Hwang, W.Y.; Fu, Y.; Reyon, D.; Maeder, M.L.; Tsai, S.Q.; Sander, J.D.; Peterson, R.T.; Yeh, J.-R.J.; Joung, J.K. Efficient Genome Editing in Zebrafish Using a CRISPR-Cas System. Nat. Biotechnol. 2013, 31, 227–229. [Google Scholar] [CrossRef] [PubMed]
- Babin, P.J.; Goizet, C.; Raldúa, D. Zebrafish Models of Human Motor Neuron Diseases: Advantages and Limitations. Prog. Neurobiol. 2014, 118, 36–58. [Google Scholar] [CrossRef] [PubMed]
- Genc, B.; Gozutok, O.; Ozdinler, P.H. Complexity of Generating Mouse Models to Study the Upper Motor Neurons: Let Us Shift Focus from Mice to Neurons. Int. J. Mol. Sci. 2019, 20, 3848. [Google Scholar] [CrossRef] [PubMed]
- Zhu, P.-P.; Soderblom, C.; Tao-Cheng, J.-H.; Stadler, J.; Blackstone, C. SPG3A Protein Atlastin-1 Is Enriched in Growth Cones and Promotes Axon Elongation during Neuronal Development. Hum. Mol. Genet. 2006, 15, 1343–1353. [Google Scholar] [CrossRef]
- Gao, Y.; Jiang, T.; Qu, C.; Tao, H.; Cao, H.; Zhao, Y.; Wang, Y.; Qu, J.; Chen, J.-G. Atlastin-1 Regulates Dendritic Morphogenesis in Mouse Cerebral Cortex. Neurosci. Res. 2013, 77, 137–142. [Google Scholar] [CrossRef]
- Shih, Y.-T.; Hsueh, Y.-P. VCP and ATL1 Regulate Endoplasmic Reticulum and Protein Synthesis for Dendritic Spine Formation. Nat. Commun. 2016, 7, 11020. [Google Scholar] [CrossRef]
- Fassier, C.; Hutt, J.A.; Scholpp, S.; Lumsden, A.; Giros, B.; Nothias, F.; Schneider-Maunoury, S.; Houart, C.; Hazan, J. Zebrafish Atlastin Controls Motility and Spinal Motor Axon Architecture via Inhibition of the BMP Pathway. Nat. Neurosci. 2010, 13, 1380–1387. [Google Scholar] [CrossRef]
- Jardin, N.; Giudicelli, F.; Ten Martín, D.; Vitrac, A.; De Gois, S.; Allison, R.; Houart, C.; Reid, E.; Hazan, J.; Fassier, C. BMP- and Neuropilin-1-Mediated Motor Axon Navigation Relies on Spastin Alternative Translation. Development 2018, 145, dev162701. [Google Scholar] [CrossRef]
- Lee, Y.; Paik, D.; Bang, S.; Kang, J.; Chun, B.; Lee, S.; Bae, E.; Chung, J.; Kim, J. Loss of Spastic Paraplegia Gene Atlastin Induces Age-Dependent Death of Dopaminergic Neurons in Drosophila. Neurobiol. Aging 2008, 29, 84–94. [Google Scholar] [CrossRef]
- Lee, M.; Paik, S.K.; Lee, M.-J.; Kim, Y.-J.; Kim, S.; Nahm, M.; Oh, S.-J.; Kim, H.-M.; Yim, J.; Lee, C.J.; et al. Drosophila Atlastin Regulates the Stability of Muscle Microtubules and Is Required for Synapse Development. Dev. Biol. 2009, 330, 250–262. [Google Scholar] [CrossRef]
- De Gregorio, C.; Delgado, R.; Ibacache, A.; Sierralta, J.; Couve, A. Drosophila Atlastin in Motor Neurons Is Required for Locomotion and Presynaptic Function. J. Cell Sci. 2017, 130, 3507–3516. [Google Scholar] [CrossRef]
- Montagna, A.; Vajente, N.; Pendin, D.; Daga, A. In Vivo Analysis of CRISPR/Cas9 Induced Atlastin Pathological Mutations in Drosophila. Front. Neurosci. 2020, 14, 547746. [Google Scholar] [CrossRef] [PubMed]
- Orso, G.; Pendin, D.; Liu, S.; Tosetto, J.; Moss, T.J.; Faust, J.E.; Micaroni, M.; Egorova, A.; Martinuzzi, A.; McNew, J.A.; et al. Homotypic Fusion of ER Membranes Requires the Dynamin-like GTPase Atlastin. Nature 2009, 460, 978–983. [Google Scholar] [CrossRef] [PubMed]
- Bertin, F.; Jara-Wilde, J.; Auer, B.; Köhler-Solís, A.; González-Silva, C.; Thomas, U.; Sierralta, J. Drosophila Atlastin Regulates Synaptic Vesicle Mobilization Independent of Bone Morphogenetic Protein Signaling. Biol. Res. 2023, 56, 49. [Google Scholar] [CrossRef] [PubMed]
- Tarrade, A.; Fassier, C.; Courageot, S.; Charvin, D.; Vitte, J.; Peris, L.; Thorel, A.; Mouisel, E.; Fonknechten, N.; Roblot, N.; et al. A Mutation of Spastin Is Responsible for Swellings and Impairment of Transport in a Region of Axon Characterized by Changes in Microtubule Composition. Hum. Mol. Genet. 2006, 15, 3544–3558. [Google Scholar] [CrossRef] [PubMed]
- Kasher, P.R.; De Vos, K.J.; Wharton, S.B.; Manser, C.; Bennett, E.J.; Bingley, M.; Wood, J.D.; Milner, R.; McDermott, C.J.; Miller, C.C.J.; et al. Direct Evidence for Axonal Transport Defects in a Novel Mouse Model of Mutant Spastin-induced Hereditary Spastic Paraplegia (HSP) and Human HSP Patients. J. Neurochem. 2009, 110, 34–44. [Google Scholar] [CrossRef]
- Fassier, C.; Tarrade, A.; Peris, L.; Courageot, S.; Mailly, P.; Dalard, C.; Delga, S.; Roblot, N.; Lefèvre, J.; Job, D.; et al. Microtubule-Targeting Drugs Rescue Axonal Swellings in Cortical Neurons from Spastin Knockout Mice. Dis. Model. Mech. 2013, 6, 72–83. [Google Scholar] [CrossRef]
- Piermarini, E.; Akarsu, S.; Connors, T.; Kneussel, M.; Lane, M.A.; Morfini, G.; Karabay, A.; Baas, P.W.; Qiang, L. Modeling Gain-of-Function and Loss-of-Function Components of SPAST-Based Hereditary Spastic Paraplegia Using Transgenic Mice. Hum. Mol. Genet. 2022, 31, 1844–1859. [Google Scholar] [CrossRef] [PubMed]
- Yu, W.; Qiang, L.; Solowska, J.M.; Karabay, A.; Korulu, S.; Baas, P.W. The Microtubule-Severing Proteins Spastin and Katanin Participate Differently in the Formation of Axonal Branches. Mol. Biol. Cell 2008, 19, 1485–1498. [Google Scholar] [CrossRef] [PubMed]
- Solowska, J.M.; D’Rozario, M.; Jean, D.C.; Davidson, M.W.; Marenda, D.R.; Baas, P.W. Pathogenic Mutation of Spastin Has Gain-of-Function Effects on Microtubule Dynamics. J. Neurosci. 2014, 34, 1856–1867. [Google Scholar] [CrossRef] [PubMed]
- Solowska, J.M.; Rao, A.N.; Baas, P.W. Truncating Mutations of SPAST Associated with Hereditary Spastic Paraplegia Indicate Greater Accumulation and Toxicity of the M1 Isoform of Spastin. Mol. Biol. Cell 2017, 28, 1728–1737. [Google Scholar] [CrossRef] [PubMed]
- Wood, J.D.; Landers, J.A.; Bingley, M.; McDermott, C.J.; Thomas-McArthur, V.; Gleadall, L.J.; Shaw, P.J.; Cunliffe, V.T. The Microtubule-Severing Protein Spastin Is Essential for Axon Outgrowth in the Zebrafish Embryo. Hum. Mol. Genet. 2006, 15, 2763–2771. [Google Scholar] [CrossRef] [PubMed]
- Butler, R.; Wood, J.D.; Landers, J.A.; Cunliffe, V.T. Genetic and Chemical Modulation of Spastin-Dependent Axon Outgrowth in Zebrafish Embryos Indicates a Role for Impaired Microtubule Dynamics in Hereditary Spastic Paraplegia. Dis. Model. Mech. 2010, 3, 743–751. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Li, D.; Ma, Y.; Yan, J.; Yang, B.; Li, P.; Yu, A.; Lu, C.; Ma, X. Role of Spastin and Protrudin in Neurite Outgrowth. J. Cell Biochem. 2012, 113, 2296–2307. [Google Scholar] [CrossRef]
- Allison, R.; Lumb, J.H.; Fassier, C.; Connell, J.W.; Ten Martin, D.; Seaman, M.N.J.; Hazan, J.; Reid, E. An ESCRT–Spastin Interaction Promotes Fission of Recycling Tubules from the Endosome. J. Cell Biol. 2013, 202, 527–543. [Google Scholar] [CrossRef]
- Julien, C.; Lissouba, A.; Madabattula, S.; Fardghassemi, Y.; Rosenfelt, C.; Androschuk, A.; Strautman, J.; Wong, C.; Bysice, A.; O’sullivan, J.; et al. Conserved Pharmacological Rescue of Hereditary Spastic Paraplegia-Related Phenotypes across Model Organisms. Hum. Mol. Genet. 2016, 25, 1088–1099. [Google Scholar] [CrossRef] [PubMed]
- Arribat, Y.; Grepper, D.; Lagarrigue, S.; Qi, T.; Cohen, S.; Amati, F. Spastin Mutations Impair Coordination between Lipid Droplet Dispersion and Reticulum. PLoS Genet. 2020, 16, e1008665. [Google Scholar] [CrossRef]
- Sherwood, N.T.; Sun, Q.; Xue, M.; Zhang, B.; Zinn, K. Drosophila Spastin Regulates Synaptic Microtubule Networks and Is Required for Normal Motor Function. PLoS Biol. 2004, 2, e429. [Google Scholar] [CrossRef] [PubMed]
- Trotta, N.; Orso, G.; Rossetto, M.G.; Daga, A.; Broadie, K. The Hereditary Spastic Paraplegia Gene, Spastin, Regulates Microtubule Stability to Modulate Synaptic Structure and Function. Curr. Biol. 2004, 14, 1135–1147. [Google Scholar] [CrossRef]
- Orso, G.; Martinuzzi, A.; Rossetto, M.G.; Sartori, E.; Feany, M.; Daga, A. Disease-Related Phenotypes in a Drosophila Model of Hereditary Spastic Paraplegia Are Ameliorated by Treatment with Vinblastine. J. Clin. Investig. 2005, 115, 3026–3034. [Google Scholar] [CrossRef]
- Ozdowski, E.F.; Wentzell, J.S.; Engert, S.M.; Abbott, H.; Sherwood, N.T. Suppression of Spastin Mutant Phenotypes by Pak3 Loss Implicates a Role for Reactive Glia in AD-HSP. Front. Neurosci. 2020, 14, 92. [Google Scholar] [CrossRef] [PubMed]
- Ferreirinha, F.; Quattrini, A.; Pirozzi, M.; Valsecchi, V.; Dina, G.; Broccoli, V.; Auricchio, A.; Piemonte, F.; Tozzi, G.; Gaeta, L.; et al. Axonal Degeneration in Paraplegin-Deficient Mice Is Associated with Abnormal Mitochondria and Impairment of Axonal Transport. J. Clin. Investig. 2004, 113, 231–242. [Google Scholar] [CrossRef]
- Pirozzi, M.; Quattrini, A.; Andolfi, G.; Dina, G.; Malaguti, M.C.; Auricchio, A.; Rugarli, E.I. Intramuscular Viral Delivery of Paraplegin Rescues Peripheral Axonopathy in a Model of Hereditary Spastic Paraplegia. J. Clin. Investig. 2006, 116, 202–208. [Google Scholar] [CrossRef]
- Sambri, I.; Massa, F.; Gullo, F.; Meneghini, S.; Cassina, L.; Carraro, M.; Dina, G.; Quattrini, A.; Patanella, L.; Carissimo, A.; et al. Impaired Flickering of the Permeability Transition Pore Causes SPG7 Spastic Paraplegia. EBioMedicine 2020, 61, 103050. [Google Scholar] [CrossRef]
- Martinelli, P.; La Mattina, V.; Bernacchia, A.; Magnoni, R.; Cerri, F.; Cox, G.; Quattrini, A.; Casari, G.; Rugarli, E.I. Genetic Interaction between the m -AAA Protease Isoenzymes Reveals Novel Roles in Cerebellar Degeneration. Hum. Mol. Genet. 2009, 18, 2001–2013. [Google Scholar] [CrossRef]
- Montoro-Gámez, C.; Nolte, H.; Molinié, T.; Evangelista, G.; Tröder, S.E.; Barth, E.; Popovic, M.; Trifunovic, A.; Zevnik, B.; Langer, T.; et al. SARM1 Deletion Delays Cerebellar but Not Spinal Cord Degeneration in an Enhanced Mouse Model of SPG7 Deficiency. Brain 2023, 146, 4117–4131. [Google Scholar] [CrossRef]
- Pareek, G.; Thomas, R.E.; Pallanck, L.J. Loss of the Drosophila M-AAA Mitochondrial Protease Paraplegin Results in Mitochondrial Dysfunction, Shortened Lifespan, and Neuronal and Muscular Degeneration. Cell Death Dis. 2018, 9, 304. [Google Scholar] [CrossRef]
- Meljon, A.; Crick, P.J.; Yutuc, E.; Yau, J.L.; Seckl, J.R.; Theofilopoulos, S.; Arenas, E.; Wang, Y.; Griffiths, W.J. Mining for Oxysterols in Cyp7b1−/− Mouse Brain and Plasma: Relevance to Spastic Paraplegia Type 5. Biomolecules 2019, 9, 149. [Google Scholar] [CrossRef]
- Goikolea, J.; Latorre-Leal, M.; Tsagkogianni, C.; Pikkupeura, S.; Gulyas, B.; Cedazo-Minguez, A.; Loera-Valencia, R.; Björkhem, I.; Rodriguez Rodriguez, P.; Maioli, S. Different Effects of CYP27A1 and CYP7B1 on Cognitive Function: Two Mouse Models in Comparison. J. Steroid Biochem. Mol. Biol. 2023, 234, 106387. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Brangulí, F.; Mishra, H.K.; Prots, I.; Havlicek, S.; Kohl, Z.; Saul, D.; Rummel, C.; Dorca-Arevalo, J.; Regensburger, M.; Graef, D.; et al. Dysfunction of Spatacsin Leads to Axonal Pathology in SPG11-Linked Hereditary Spastic Paraplegia. Hum. Mol. Genet. 2014, 23, 4859–4874. [Google Scholar] [CrossRef] [PubMed]
- Varga, R.-E.; Khundadze, M.; Damme, M.; Nietzsche, S.; Hoffmann, B.; Stauber, T.; Koch, N.; Hennings, J.C.; Franzka, P.; Huebner, A.K.; et al. In Vivo Evidence for Lysosome Depletion and Impaired Autophagic Clearance in Hereditary Spastic Paraplegia Type SPG11. PLoS Genet. 2015, 11, e1005454. [Google Scholar] [CrossRef] [PubMed]
- Branchu, J.; Boutry, M.; Sourd, L.; Depp, M.; Leone, C.; Corriger, A.; Vallucci, M.; Esteves, T.; Matusiak, R.; Dumont, M.; et al. Loss of Spatacsin Function Alters Lysosomal Lipid Clearance Leading to Upper and Lower Motor Neuron Degeneration. Neurobiol. Dis. 2017, 102, 21–37. [Google Scholar] [CrossRef] [PubMed]
- Boutry, M.; Branchu, J.; Lustremant, C.; Pujol, C.; Pernelle, J.; Matusiak, R.; Seyer, A.; Poirel, M.; Chu-Van, E.; Pierga, A.; et al. Inhibition of Lysosome Membrane Recycling Causes Accumulation of Gangliosides That Contribute to Neurodegeneration. Cell Rep. 2018, 23, 3813–3826. [Google Scholar] [CrossRef] [PubMed]
- Pierga, A.; Matusiak, R.; Cauhapé, M.; Branchu, J.; Danglot, L.; Boutry, M.; Darios, F. Spatacsin Regulates Directionality of Lysosome Trafficking by Promoting the Degradation of Its Partner AP5Z1. PLoS Biol. 2023, 21, e3002337. [Google Scholar] [CrossRef]
- Southgate, L.; Dafou, D.; Hoyle, J.; Li, N.; Kinning, E.; Critchley, P.; Németh, A.H.; Talbot, K.; Bindu, P.S.; Sinha, S.; et al. Novel SPG11 Mutations in Asian Kindreds and Disruption of Spatacsin Function in the Zebrafish. Neurogenetics 2010, 11, 379–389. [Google Scholar] [CrossRef]
- Martin, E.; Yanicostas, C.; Rastetter, A.; Naini, S.M.A.; Maouedj, A.; Kabashi, E.; Rivaud-Péchoux, S.; Brice, A.; Stevanin, G.; Soussi-Yanicostas, N. Spatacsin and Spastizin Act in the Same Pathway Required for Proper Spinal Motor Neuron Axon Outgrowth in Zebrafish. Neurobiol. Dis. 2012, 48, 299–308. [Google Scholar] [CrossRef]
- Khundadze, M.; Kollmann, K.; Koch, N.; Biskup, C.; Nietzsche, S.; Zimmer, G.; Hennings, J.C.; Huebner, A.K.; Symmank, J.; Jahic, A.; et al. A Hereditary Spastic Paraplegia Mouse Model Supports a Role of ZFYVE26/SPASTIZIN for the Endolysosomal System. PLoS Genet. 2013, 9, e1003988. [Google Scholar] [CrossRef] [PubMed]
- Khundadze, M.; Ribaudo, F.; Hussain, A.; Stahlberg, H.; Brocke-Ahmadinejad, N.; Franzka, P.; Varga, R.-E.; Zarkovic, M.; Pungsrinont, T.; Kokal, M.; et al. Mouse Models for Hereditary Spastic Paraplegia Uncover a Role of PI4K2A in Autophagic Lysosome Reformation. Autophagy 2021, 17, 3690–3706. [Google Scholar] [CrossRef] [PubMed]
- Marrone, L.; Marchi, P.M.; Webster, C.P.; Marroccella, R.; Coldicott, I.; Reynolds, S.; Alves-Cruzeiro, J.; Yang, Z.-L.; Higginbottom, A.; Khundadze, M.; et al. SPG15 Protein Deficits Are at the Crossroads between Lysosomal Abnormalities, Altered Lipid Metabolism and Synaptic Dysfunction. Hum. Mol. Genet. 2022, 31, 2693–2710. [Google Scholar] [CrossRef] [PubMed]
- Vantaggiato, C.; Orso, G.; Guarato, G.; Brivio, F.; Napoli, B.; Panzeri, E.; Masotti, S.; Santorelli, F.M.; Lamprou, M.; Gumeni, S.; et al. Rescue of Lysosomal Function as Therapeutic Strategy for SPG15 Hereditary Spastic Paraplegia. Brain 2023, 146, 1103–1120. [Google Scholar] [CrossRef] [PubMed]
- Khundadze, M.; Ribaudo, F.; Hussain, A.; Rosentreter, J.; Nietzsche, S.; Thelen, M.; Winter, D.; Hoffmann, B.; Afzal, M.A.; Hermann, T.; et al. A Mouse Model for SPG48 Reveals a Block of Autophagic Flux upon Disruption of Adaptor Protein Complex Five. Neurobiol. Dis. 2019, 127, 419–431. [Google Scholar] [CrossRef]
- Matsuda, S.; Miura, E.; Matsuda, K.; Kakegawa, W.; Kohda, K.; Watanabe, M.; Yuzaki, M. Accumulation of AMPA Receptors in Autophagosomes in Neuronal Axons Lacking Adaptor Protein AP-4. Neuron 2008, 57, 730–745. [Google Scholar] [CrossRef]
- Scarrott, J.M.; Alves-Cruzeiro, J.; Marchi, P.M.; Webster, C.P.; Yang, Z.-L.; Karyka, E.; Marroccella, R.; Coldicott, I.; Thomas, H.; Azzouz, M. Ap4b1-Knockout Mouse Model of Hereditary Spastic Paraplegia Type 47 Displays Motor Dysfunction, Aberrant Brain Morphology and ATG9A Mislocalization. Brain Commun. 2022, 5, fcac335. [Google Scholar] [CrossRef]
- Pembridge, O.G.; Wallace, N.S.; Clements, T.P.; Jackson, L.P. AP-4 Loss in CRISPR-Edited Zebrafish Affects Early Embryo Development. Adv. Biol. Regul. 2023, 87, 100945. [Google Scholar] [CrossRef]
- De Pace, R.; Skirzewski, M.; Damme, M.; Mattera, R.; Mercurio, J.; Foster, A.M.; Cuitino, L.; Jarnik, M.; Hoffmann, V.; Morris, H.D.; et al. Altered Distribution of ATG9A and Accumulation of Axonal Aggregates in Neurons from a Mouse Model of AP-4 Deficiency Syndrome. PLoS Genet. 2018, 14, e1007363. [Google Scholar] [CrossRef] [PubMed]
- Ivankovic, D.; Drew, J.; Lesept, F.; White, I.J.; López Doménech, G.; Tooze, S.A.; Kittler, J.T. Axonal Autophagosome Maturation Defect through Failure of ATG9A Sorting Underpins Pathology in AP-4 Deficiency Syndrome. Autophagy 2020, 16, 391–407. [Google Scholar] [CrossRef] [PubMed]
- D’Amore, A.; Tessa, A.; Naef, V.; Bassi, M.T.; Citterio, A.; Romaniello, R.; Fichi, G.; Galatolo, D.; Mero, S.; Battini, R.; et al. Loss of Ap4s1 in Zebrafish Leads to Neurodevelopmental Defects Resembling Spastic Paraplegia 52. Ann. Clin. Transl. Neurol. 2020, 7, 584–589. [Google Scholar] [CrossRef]
- Witte, K.; Schuh, A.L.; Hegermann, J.; Sarkeshik, A.; Mayers, J.R.; Schwarze, K.; Yates III, J.R.; Eimer, S.; Audhya, A. TFG-1 Function in Protein Secretion and Oncogenesis. Nat. Cell Biol. 2011, 13, 550–558. [Google Scholar] [CrossRef]
- Yamamotoya, T.; Nakatsu, Y.; Kushiyama, A.; Matsunaga, Y.; Ueda, K.; Inoue, Y.; Inoue, M.-K.; Sakoda, H.; Fujishiro, M.; Ono, H.; et al. Trk-Fused Gene (TFG) Regulates Pancreatic β Cell Mass and Insulin Secretory Activity. Sci. Rep. 2017, 7, 13026. [Google Scholar] [CrossRef]
- Yamamotoya, T.; Hasei, S.; Akasaka, Y.; Ohata, Y.; Nakatsu, Y.; Kanna, M.; Fujishiro, M.; Sakoda, H.; Ono, H.; Kushiyama, A.; et al. Involvement of Neuronal and Muscular Trk-Fused Gene (TFG) Defects in the Development of Neurodegenerative Diseases. Sci. Rep. 2022, 12, 1966. [Google Scholar] [CrossRef]
- Chen, X.; Liu, F.; Chen, K.; Wang, Y.; Yin, A.; Kang, X.; Yang, S.; Zhao, H.; Dong, S.; Li, Y.; et al. TFG Mutation Induces Haploinsufficiency and Drives Axonal Charcot–Marie–Tooth Disease by Causing Neurite Degeneration. CNS Neurosci. Ther. 2022, 28, 2076–2089. [Google Scholar] [CrossRef] [PubMed]
- Peotter, J.L.; Pustova, I.; Lettman, M.M.; Shatadal, S.; Bradberry, M.M.; Winter-Reed, A.D.; Charan, M.; Sharkey, E.E.; Alvin, J.R.; Bren, A.M.; et al. TFG Regulates Secretory and Endosomal Sorting Pathways in Neurons to Promote Their Activity and Maintenance. Proc. Natl. Acad. Sci. USA 2022, 119, e2210649119. [Google Scholar] [CrossRef] [PubMed]
- Gaspard, N.; Bouschet, T.; Hourez, R.; Dimidschstein, J.; Naeije, G.; van den Ameele, J.; Espuny-Camacho, I.; Herpoel, A.; Passante, L.; Schiffmann, S.N.; et al. An Intrinsic Mechanism of Corticogenesis from Embryonic Stem Cells. Nature 2008, 455, 351–357. [Google Scholar] [CrossRef]
- Watanabe, K.; Kamiya, D.; Nishiyama, A.; Katayama, T.; Nozaki, S.; Kawasaki, H.; Watanabe, Y.; Mizuseki, K.; Sasai, Y. Directed Differentiation of Telencephalic Precursors from Embryonic Stem Cells. Nat. Neurosci. 2005, 8, 288–296. [Google Scholar] [CrossRef]
- Shiraishi, A.; Muguruma, K.; Sasai, Y. Generation of Thalamic Neurons from Mouse Embryonic Stem Cells. Development 2017, 144, 1211–1220. [Google Scholar] [CrossRef]
- Chambers, S.M.; Qi, Y.; Mica, Y.; Lee, G.; Zhang, X.-J.; Niu, L.; Bilsland, J.; Cao, L.; Stevens, E.; Whiting, P.; et al. Combined Small-Molecule Inhibition Accelerates Developmental Timing and Converts Human Pluripotent Stem Cells into Nociceptors. Nat. Biotechnol. 2012, 30, 715–720. [Google Scholar] [CrossRef]
- Wataya, T.; Ando, S.; Muguruma, K.; Ikeda, H.; Watanabe, K.; Eiraku, M.; Kawada, M.; Takahashi, J.; Hashimoto, N.; Sasai, Y. Minimization of Exogenous Signals in ES Cell Culture Induces Rostral Hypothalamic Differentiation. Proc. Natl. Acad. Sci. USA 2008, 105, 11796–11801. [Google Scholar] [CrossRef]
- Doi, D.; Magotani, H.; Kikuchi, T.; Ikeda, M.; Hiramatsu, S.; Yoshida, K.; Amano, N.; Nomura, M.; Umekage, M.; Morizane, A.; et al. Pre-Clinical Study of Induced Pluripotent Stem Cell-Derived Dopaminergic Progenitor Cells for Parkinson’s Disease. Nat. Commun. 2020, 11, 3369. [Google Scholar] [CrossRef]
- Osakada, F.; Jin, Z.-B.; Hirami, Y.; Ikeda, H.; Danjyo, T.; Watanabe, K.; Sasai, Y.; Takahashi, M. In Vitro Differentiation of Retinal Cells from Human Pluripotent Stem Cells by Small-Molecule Induction. J. Cell Sci. 2009, 122, 3169–3179. [Google Scholar] [CrossRef]
- Lancaster, M.A.; Renner, M.; Martin, C.-A.; Wenzel, D.; Bicknell, L.S.; Hurles, M.E.; Homfray, T.; Penninger, J.M.; Jackson, A.P.; Knoblich, J.A. Cerebral Organoids Model Human Brain Development and Microcephaly. Nature 2013, 501, 373–379. [Google Scholar] [CrossRef] [PubMed]
- Kadoshima, T.; Sakaguchi, H.; Nakano, T.; Soen, M.; Ando, S.; Eiraku, M.; Sasai, Y. Self-Organization of Axial Polarity, inside-out Layer Pattern, and Species-Specific Progenitor Dynamics in Human ES Cell–Derived Neocortex. Proc. Natl. Acad. Sci. USA 2013, 110, 20284–20289. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Okita, K.; Nakagawa, M.; Yamanaka, S. Induction of Pluripotent Stem Cells from Fibroblast Cultures. Nat. Protoc. 2007, 2, 3081–3089. [Google Scholar] [CrossRef]
- Son, Y.S.; Seong, R.H.; Ryu, C.J.; Cho, Y.S.; Bae, K.-H.; Chung, S.J.; Lee, B.; Min, J.-K.; Hong, H.J. Brief Report: L1 Cell Adhesion Molecule, a Novel Surface Molecule of Human Embryonic Stem Cells, Is Essential for Self-Renewal and Pluripotency. Stem Cells 2011, 29, 2094–2099. [Google Scholar] [CrossRef]
- Li, S.; Wang, Y.; Sun, H.; Wang, Z.; Zhang, Q.; Wang, Y.; Yang, J.; Shi, C.; Yuan, Y.; Wang, H.; et al. Establishment of Induced Pluripotent Stem Cell Line (ZZUi033-A) of a Male with a Novel L1CAM Missense Mutation. Stem Cell Res. 2022, 59, 102663. [Google Scholar] [CrossRef] [PubMed]
- Zhu, P.-P.; Denton, K.R.; Pierson, T.M.; Li, X.-J.; Blackstone, C. Pharmacologic Rescue of Axon Growth Defects in a Human IPSC Model of Hereditary Spastic Paraplegia SPG3A. Hum. Mol. Genet. 2014, 23, 5638–5648. [Google Scholar] [CrossRef]
- Costamagna, D.; Casters, V.; Beltrà, M.; Sampaolesi, M.; Van Campenhout, A.; Ortibus, E.; Desloovere, K.; Duelen, R. Autologous IPSC-Derived Human Neuromuscular Junction to Model the Pathophysiology of Hereditary Spastic Paraplegia. Cells 2022, 11, 3351. [Google Scholar] [CrossRef]
- Abrahamsen, G.; Fan, Y.; Matigian, N.; Wali, G.; Bellette, B.; Sutharsan, R.; Raju, J.; Wood, S.A.; Veivers, D.; Sue, C.M.; et al. A Patient-Derived Stem Cell Model of Hereditary Spastic Paraplegia with SPAST Mutations. Dis. Model. Mech. 2013, 6, 489–502. [Google Scholar] [CrossRef] [PubMed]
- Denton, K.R.; Lei, L.; Grenier, J.; Rodionov, V.; Blackstone, C.; Li, X.-J. Loss of Spastin Function Results in Disease-Specific Axonal Defects in Human Pluripotent Stem Cell-Based Models of Hereditary Spastic Paraplegia. Stem Cells 2014, 32, 414–423. [Google Scholar] [CrossRef] [PubMed]
- Havlicek, S.; Kohl, Z.; Mishra, H.K.; Prots, I.; Eberhardt, E.; Denguir, N.; Wend, H.; Plotz, S.; Boyer, L.; Marchetto, M.C.N.; et al. Gene Dosage-Dependent Rescue of HSP Neurite Defects in SPG4 Patients’ Neurons. Hum. Mol. Genet. 2014, 23, 2527–2541. [Google Scholar] [CrossRef] [PubMed]
- Wali, G.; Sutharsan, R.; Fan, Y.; Stewart, R.; Tello Velasquez, J.; Sue, C.M.; Crane, D.I.; Mackay-Sim, A. Mechanism of Impaired Microtubule-Dependent Peroxisome Trafficking and Oxidative Stress in SPAST-Mutated Cells from Patients with Hereditary Spastic Paraplegia. Sci. Rep. 2016, 6, 27004. [Google Scholar] [CrossRef]
- Mou, Y.; Nandi, G.; Mukte, S.; Chai, E.; Chen, Z.; Nielsen, J.E.; Nielsen, T.T.; Criscuolo, C.; Blackstone, C.; Fraidakis, M.J.; et al. Chenodeoxycholic Acid Rescues Axonal Degeneration in Induced Pluripotent Stem Cell-Derived Neurons from Spastic Paraplegia Type 5 and Cerebrotendinous Xanthomatosis Patients. Orphanet J. Rare Dis. 2023, 18, 72. [Google Scholar] [CrossRef] [PubMed]
- Wali, G.; Kumar, K.R.; Liyanage, E.; Davis, R.L.; Mackay-Sim, A.; Sue, C.M. Mitochondrial Function in Hereditary Spastic Paraplegia: Deficits in SPG7 but Not SPAST Patient-Derived Stem Cells. Front. Neurosci. 2020, 14, 820. [Google Scholar] [CrossRef] [PubMed]
- Santangelo, S.; Bossolasco, P.; Magri, S.; Colombrita, C.; Invernizzi, S.; Gellera, C.; Nanetti, L.; Di Bella, D.; Silani, V.; Taroni, F.; et al. Generation of an IPSC Line from a Patient with Spastic Paraplegia Type 10 Carrying a Novel Mutation in KIF5A Gene. Stem Cell Res. 2023, 66, 103008. [Google Scholar] [CrossRef] [PubMed]
- Mishra, H.K.; Prots, I.; Havlicek, S.; Kohl, Z.; Perez-Branguli, F.; Boerstler, T.; Anneser, L.; Minakaki, G.; Wend, H.; Hampl, M.; et al. GSK3ß-dependent Dysregulation of Neurodevelopment in SPG11-patient Induced Pluripotent Stem Cell Model. Ann. Neurol. 2016, 79, 826–840. [Google Scholar] [CrossRef]
- Pérez-Brangulí, F.; Buchsbaum, I.Y.; Pozner, T.; Regensburger, M.; Fan, W.; Schray, A.; Börstler, T.; Mishra, H.; Gräf, D.; Kohl, Z.; et al. Human SPG11 Cerebral Organoids Reveal Cortical Neurogenesis Impairment. Hum. Mol. Genet. 2019, 28, 961–971. [Google Scholar] [CrossRef]
- Güner, F.; Pozner, T.; Krach, F.; Prots, I.; Loskarn, S.; Schlötzer-Schrehardt, U.; Winkler, J.; Winner, B.; Regensburger, M. Axon-Specific Mitochondrial Pathology in SPG11 Alpha Motor Neurons. Front. Neurosci. 2021, 15, 680572. [Google Scholar] [CrossRef]
- Denton, K.; Mou, Y.; Xu, C.-C.; Shah, D.; Chang, J.; Blackstone, C.; Li, X.-J. Impaired Mitochondrial Dynamics Underlie Axonal Defects in Hereditary Spastic Paraplegias. Hum. Mol. Genet. 2018, 27, 2517–2530. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Ma, Y.; Duan, N.; Liu, X.; Zhang, H.; Ma, J.; Liu, Y.; Sun, W.; Liu, Q. Generation of a Human Induced Pluripotent Stem Cell Line (SDUBMSi001-A) from a Hereditary Spastic Paraplegia Patient Carrying Kif1a c.773C>T Missense Mutation. Stem Cell Res. 2020, 43, 101727. [Google Scholar] [CrossRef]
- Lin, H.-Y.; Ou-Yang, C.-H.; Lin, C.-H. Reprogramming of a Human Induced Pluripotent Stem Cell (IPSC) Line from a Patient with Neurodegeneration with Brain Iron Accumulation (NBIA) Harboring a Novel Frameshift Mutation in C19orf12 Gene. Stem Cell Res. 2020, 49, 102032. [Google Scholar] [CrossRef] [PubMed]
- Behne, R.; Teinert, J.; Wimmer, M.; D’Amore, A.; Davies, A.K.; Scarrott, J.M.; Eberhardt, K.; Brechmann, B.; Chen, I.P.-F.; Buttermore, E.D.; et al. Adaptor Protein Complex 4 Deficiency: A Paradigm of Childhood-Onset Hereditary Spastic Paraplegia Caused by Defective Protein Trafficking. Hum. Mol. Genet. 2020, 29, 320–334. [Google Scholar] [CrossRef]
- Leeson, H.C.; Goh, D.; Coman, D.; Wolvetang, E.J. Generation of IPSC Lines from Hereditary Spastic Paraplegia 56 (SPG56) Patients and Family Members Carrying CYP2U1 Mutations. Stem Cell Res. 2022, 64, 102917. [Google Scholar] [CrossRef] [PubMed]
- Slosarek, E.L.; Schuh, A.L.; Pustova, I.; Johnson, A.; Bird, J.; Johnson, M.; Frankel, E.B.; Bhattacharya, N.; Hanna, M.G.; Burke, J.E.; et al. Pathogenic TFG Mutations Underlying Hereditary Spastic Paraplegia Impair Secretory Protein Trafficking and Axon Fasciculation. Cell Rep. 2018, 24, 2248–2260. [Google Scholar] [CrossRef] [PubMed]
- Nagel, M.; Müßig, S.; Höflinger, P.; Schöls, L.; Hauser, S.; Schüle, R. Generation of the CRISPR/Cas9-Mediated KIF1C Knock-out Human IPSC Line HIHRSi003-A-1. Stem Cell Res. 2020, 49, 102059. [Google Scholar] [CrossRef] [PubMed]
- Gu, H.; Shi, X.; Liu, C.; Wang, C.; Sui, N.; Zhao, Y.; Gong, J.; Wang, F.; Zhang, H.; Li, W.; et al. USP8 Maintains Embryonic Stem Cell Stemness via Deubiquitination of EPG5. Nat. Commun. 2019, 10, 1465. [Google Scholar] [CrossRef]
- Lu, Y.; Dong, E.; Yang, W.; Lai, L.; Lin, X.; Ma, L.; Chen, W.; Wang, N.; Lin, X. Generation of an Integration-Free Induced Pluripotent Stem Cell Line, FJMUi001-A, from a Hereditary Spastic Paraplegia Patient Carrying Compound Heterozygous p.P498L and p.R618W Mutations in CAPN1 (SPG76). Stem Cell Res. 2019, 34, 101354. [Google Scholar] [CrossRef]
- Hedera, P.; Fenichel, G.M.; Blair, M.; Haines, J.L. Novel Mutation in the SPG3A Gene in an African American Family with an Early Onset of Hereditary Spastic Paraplegia. Arch. Neurol. 2004, 61, 1600. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, P.A.; Hart, P.E.; Patel, H.; Warner, T.T.; Crosby, A.H. SPG3A Mutation Screening in English Families with Early Onset Autosomal Dominant Hereditary Spastic Paraplegia. J. Neurol. Sci. 2003, 216, 43–45. [Google Scholar] [CrossRef]
- Alecu, J.E.; Saffari, A.; Jordan, C.; Srivastava, S.; Blackstone, C.; Ebrahimi-Fakhari, D. De Novo Variants Cause Complex Symptoms in HSP- ATL1 (SPG3A) and Uncover Genotype–Phenotype Correlations. Hum. Mol. Genet. 2023, 32, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Da Graça, F.F.; de Rezende, T.J.R.; Vasconcellos, L.F.R.; Pedroso, J.L.; Barsottini, O.G.P.; França, M.C. Neuroimaging in Hereditary Spastic Paraplegias: Current Use and Future Perspectives. Front. Neurol. 2019, 9, 1117. [Google Scholar] [CrossRef]
- Orlacchio, A.; Montieri, P.; Babalini, C.; Gaudiello, F.; Bernardi, G.; Kawarai, T. Late-Onset Hereditary Spastic Paraplegia with Thin Corpus Callosum Caused by a New SPG3A Mutation. J. Neurol. 2011, 258, 1361–1363. [Google Scholar] [CrossRef]
- Muriel, M.; Dauphin, A.; Namekawa, M.; Gervais, A.; Brice, A.; Ruberg, M. Atlastin-1, the Dynamin-like GTPase Responsible for Spastic Paraplegia SPG3A, Remodels Lipid Membranes and May Form Tubules and Vesicles in the Endoplasmic Reticulum. J. Neurochem. 2009, 110, 1607–1616. [Google Scholar] [CrossRef] [PubMed]
- Ulengin, I.; Park, J.J.; Lee, T.H. ER Network Formation and Membrane Fusion by Atlastin1/SPG3A Disease Variants. Mol. Biol. Cell 2015, 26, 1616–1628. [Google Scholar] [CrossRef] [PubMed]
- Varadi, M.; Anyango, S.; Deshpande, M.; Nair, S.; Natassia, C.; Yordanova, G.; Yuan, D.; Stroe, O.; Wood, G.; Laydon, A.; et al. AlphaFold Protein Structure Database: Massively Expanding the Structural Coverage of Protein-Sequence Space with High-Accuracy Models. Nucleic Acids Res. 2022, 50, D439–D444. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Namekawa, M.; Muriel, M.-P.; Janer, A.; Latouche, M.; Dauphin, A.; Debeir, T.; Martin, E.; Duyckaerts, C.; Prigent, A.; Depienne, C.; et al. Mutations in the SPG3A Gene Encoding the GTPase Atlastin Interfere with Vesicle Trafficking in the ER/Golgi Interface and Golgi Morphogenesis. Mol. Cell. Neurosci. 2007, 35, 1–13. [Google Scholar] [CrossRef]
- Bianchi, F.; Malboubi, M.; Li, Y.; George, J.H.; Jerusalem, A.; Szele, F.; Thompson, M.S.; Ye, H. Rapid and Efficient Differentiation of Functional Motor Neurons from Human IPSC for Neural Injury Modelling. Stem Cell Res. 2018, 32, 126–134. [Google Scholar] [CrossRef]
- Paranjape, S.R.; Nagendran, T.; Poole, V.; Harris, J.; Taylor, A.M. Compartmentalization of Human Stem Cell-Derived Neurons within Pre-Assembled Plastic Microfluidic Chips. J. Vis. Exp. 2019, e59250. [Google Scholar] [CrossRef]
- Parodi, L.; Fenu, S.; Barbier, M.; Banneau, G.; Duyckaerts, C.; Tezenas du Montcel, S.; Monin, M.-L.; Ait Said, S.; Guegan, J.; Tallaksen, C.M.E.; et al. Spastic Paraplegia Due to SPAST Mutations Is Modified by the Underlying Mutation and Sex. Brain 2018, 141, 3331–3342. [Google Scholar] [CrossRef]
- Kumar, K.R.; Sue, C.M.; Burke, D.; Ng, K. Peripheral Neuropathy in Hereditary Spastic Paraplegia Due to Spastin (SPG4) Mutation—A Neurophysiological Study Using Excitability Techniques. Clin. Neurophysiol. 2012, 123, 1454–1459. [Google Scholar] [CrossRef]
- Akaba, Y.; Takeguchi, R.; Tanaka, R.; Takahashi, S. A Complex Phenotype of a Patient with Spastic Paraplegia Type 4 Caused by a Novel Pathogenic Variant in the SPAST Gene. Case Rep. Neurol. 2021, 13, 763–771. [Google Scholar] [CrossRef]
- Orlacchio, A.; Kawarai, T.; Totaro, A.; Errico, A.; St George-Hyslop, P.H.; Rugarli, E.I.; Bernardi, G. Hereditary Spastic Paraplegia: Clinical Genetic Study of 15 Families. Arch. Neurol. 2004, 61, 849–855. [Google Scholar] [CrossRef]
- Hazan, J.; Fonknechten, N.; Mavel, D.; Paternotte, C.; Samson, D.; Artiguenave, F.; Davoine, C.-S.; Cruaud, C.; Dürr, A.; Wincker, P.; et al. Spastin, a New AAA Protein, Is Altered in the Most Frequent Form of Autosomal Dominant Spastic Paraplegia. Nat. Genet. 1999, 23, 296–303. [Google Scholar] [CrossRef] [PubMed]
- Claudiani, P.; Riano, E.; Errico, A.; Andolfi, G.; Rugarli, E.I. Spastin Subcellular Localization Is Regulated through Usage of Different Translation Start Sites and Active Export from the Nucleus. Exp. Cell Res. 2005, 309, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Errico, A.; Ballabio, A.; Rugarli, E.I. Spastin, the Protein Mutated in Autosomal Dominant Hereditary Spastic Paraplegia, Is Involved in Microtubule Dynamics. Hum. Mol. Genet. 2002, 11, 153–163. [Google Scholar] [CrossRef]
- Salinas, S.; Carazo-Salas, R.E.; Proukakis, C.; Schiavo, G.; Warner, T.T. Spastin and Microtubules: Functions in Health and Disease. J. Neurosci. Res. 2007, 85, 2778–2782. [Google Scholar] [CrossRef] [PubMed]
- Beetz, C.; Nygren, A.O.H.; Schickel, J.; Auer-Grumbach, M.; Bürk, K.; Heide, G.; Kassubek, J.; Klimpe, S.; Klopstock, T.; Kreuz, F.; et al. High Frequency of Partial SPAST Deletions in Autosomal Dominant Hereditary Spastic Paraplegia. Neurology 2006, 67, 1926–1930. [Google Scholar] [CrossRef]
- Crane, D.I. Revisiting the Neuropathogenesis of Zellweger Syndrome. Neurochem. Int. 2014, 69, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Khalik, J.; Yutuc, E.; Crick, P.J.; Gustafsson, J.-Å.; Warner, M.; Roman, G.; Talbot, K.; Gray, E.; Griffiths, W.J.; Turner, M.R.; et al. Defective Cholesterol Metabolism in Amyotrophic Lateral Sclerosis. J. Lipid Res. 2017, 58, 267–278. [Google Scholar] [CrossRef] [PubMed]
- Rickman, O.J.; Baple, E.L.; Crosby, A.H. Lipid Metabolic Pathways Converge in Motor Neuron Degenerative Diseases. Brain 2020, 143, 1073–1087. [Google Scholar] [CrossRef] [PubMed]
- Tsaousidou, M.K.; Ouahchi, K.; Warner, T.T.; Yang, Y.; Simpson, M.A.; Laing, N.G.; Wilkinson, P.A.; Madrid, R.E.; Patel, H.; Hentati, F.; et al. Sequence Alterations within CYP7B1 Implicate Defective Cholesterol Homeostasis in Motor-Neuron Degeneration. Am. J. Hum. Genet. 2008, 82, 510–515. [Google Scholar] [CrossRef] [PubMed]
- Merino-Serrais, P.; Loera-Valencia, R.; Rodriguez-Rodriguez, P.; Parrado-Fernandez, C.; Ismail, M.A.; Maioli, S.; Matute, E.; Jimenez-Mateos, E.M.; Björkhem, I.; DeFelipe, J.; et al. 27-Hydroxycholesterol Induces Aberrant Morphology and Synaptic Dysfunction in Hippocampal Neurons. Cereb. Cortex 2019, 29, 429–446. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; An, Y.; Zhang, D.; Yu, H.; Zhang, X.; Wang, Y.; Tao, L.; Xiao, R. 27-Hydroxycholesterol Alters Synaptic Structural and Functional Plasticity in Hippocampal Neuronal Cultures. J. Neuropathol. Exp. Neurol. 2019, 78, 238–247. [Google Scholar]
- Schüle, R.; Siddique, T.; Deng, H.-X.; Yang, Y.; Donkervoort, S.; Hansson, M.; Madrid, R.E.; Siddique, N.; Schöls, L.; Björkhem, I. Marked Accumulation of 27-Hydroxycholesterol in SPG5 Patients with Hereditary Spastic Paresis. J. Lipid Res. 2010, 51, 819–823. [Google Scholar] [CrossRef] [PubMed]
- Biancheri, R.; Ciccolella, M.; Rossi, A.; Tessa, A.; Cassandrini, D.; Minetti, C.; Santorelli, F.M. White Matter Lesions in Spastic Paraplegia with Mutations in SPG5/CYP7B1. Neuromuscul. Disord. 2009, 19, 62–65. [Google Scholar] [CrossRef]
- Boisvert, E.M.; Denton, K.; Lei, L.; Li, X.-J. The Specification of Telencephalic Glutamatergic Neurons from Human Pluripotent Stem Cells. J. Vis. Exp. 2013, e50321. [Google Scholar] [CrossRef]
- Li, X.-J.; Zhang, X.; Johnson, M.A.; Wang, Z.-B.; LaVaute, T.; Zhang, S.-C. Coordination of Sonic Hedgehog and Wnt Signaling Determines Ventral and Dorsal Telencephalic Neuron Types from Human Embryonic Stem Cells. Development 2009, 136, 4055–4063. [Google Scholar] [CrossRef]
- Bramlett, K.S.; Yao, S.; Burris, T.P. Correlation of Farnesoid X Receptor Coactivator Recruitment and Cholesterol 7α-Hydroxylase Gene Repression by Bile Acids. Mol. Genet. Metab. 2000, 71, 609–615. [Google Scholar] [CrossRef]
- Kallner, M. The Effect of Chenodeoxycholic Acid Feeding on Bile Acid Kinetics and Fecal Neutral Steroid Excretion in Patients with Hyperlipoproteinemia Types II and IV. J. Lab. Clin. Med. 1975, 86, 595–604. [Google Scholar]
- Bonnot, O.; Fraidakis, M.J.; Lucanto, R.; Chauvin, D.; Kelley, N.; Plaza, M.; Dubourg, O.; Lyon-Caen, O.; Sedel, F.; Cohen, D. Cerebrotendinous Xanthomatosis Presenting with Severe Externalized Disorder: Improvement after One Year of Treatment with Chenodeoxycholic Acid. CNS Spectr. 2010, 15, 231–237. [Google Scholar] [CrossRef]
- Dai, D.; Mills, P.B.; Footitt, E.; Gissen, P.; McClean, P.; Stahlschmidt, J.; Coupry, I.; Lavie, J.; Mochel, F.; Goizet, C.; et al. Liver Disease in Infancy Caused by Oxysterol 7α-hydroxylase Deficiency: Successful Treatment with Chenodeoxycholic Acid. J. Inherit. Metab. Dis. 2014, 37, 851–861. [Google Scholar] [CrossRef] [PubMed]
- Schüle, R.; Wiethoff, S.; Martus, P.; Karle, K.N.; Otto, S.; Klebe, S.; Klimpe, S.; Gallenmüller, C.; Kurzwelly, D.; Henkel, D.; et al. Hereditary Spastic Paraplegia: Clinicogenetic Lessons from 608 Patients. Ann. Neurol. 2016, 79, 646–658. [Google Scholar] [CrossRef]
- Casari, G.; De Fusco, M.; Ciarmatori, S.; Zeviani, M.; Mora, M.; Fernandez, P.; De Michele, G.; Filla, A.; Cocozza, S.; Marconi, R.; et al. Spastic Paraplegia and OXPHOS Impairment Caused by Mutations in Paraplegin, a Nuclear-Encoded Mitochondrial Metalloprotease. Cell 1998, 93, 973–983. [Google Scholar] [CrossRef] [PubMed]
- De Michele, G.; De Fusco, M.; Cavalcanti, F.; Filla, A.; Marconi, R.; Volpe, G.; Monticelli, A.; Ballabio, A.; Casari, G.; Cocozza, S. A New Locus for Autosomal Recessive Hereditary Spastic Paraplegia Maps to Chromosome 16q24.3. Am. J. Hum. Genet. 1998, 63, 135–139. [Google Scholar] [CrossRef]
- McDermott, C.J.; Dayaratne, R.K.; Tomkins, J.; Lusher, M.E.; Lindsey, J.C.; Johnson, M.A.; Casari, G.; Turnbull, D.M.; Bushby, K.; Shaw, P.J. Paraplegin Gene Analysis in Hereditary Spastic Paraparesis (HSP) Pedigrees in Northeast England. Neurology 2001, 56, 467–471. [Google Scholar] [CrossRef]
- Warnecke, T.; Duning, T.; Schirmacher, A.; Mohammadi, S.; Schwindt, W.; Lohmann, H.; Dziewas, R.; Deppe, M.; Ringelstein, E.B.; Young, P. A Novel Splice Site Mutation in the SPG7 Gene Causing Widespread Fiber Damage in Homozygous and Heterozygous Subjects. Mov. Disord. 2010, 25, 413–420. [Google Scholar] [CrossRef] [PubMed]
- Tzoulis, C.; Denora, P.S.; Santorelli, F.M.; Bindoff, L.A. Hereditary Spastic Paraplegia Caused by the Novel Mutation 1047insC in the SPG7 Gene. J. Neurol. 2008, 255, 1142–1144. [Google Scholar] [CrossRef]
- Warnecke, T.; Duning, T.; Schwan, A.; Lohmann, H.; Epplen, J.T.; Young, P. A Novel Form of Autosomal Recessive Hereditary Spastic Paraplegia Caused by a New SPG7 Mutation. Neurology 2007, 69, 368–375. [Google Scholar] [CrossRef]
- Nolden, M.; Ehses, S.; Koppen, M.; Bernacchia, A.; Rugarli, E.I.; Langer, T. The M-AAA Protease Defective in Hereditary Spastic Paraplegia Controls Ribosome Assembly in Mitochondria. Cell 2005, 123, 277–289. [Google Scholar] [CrossRef]
- James, A.M.; Wei, Y.H.; Pang, C.Y.; Murphy, M.P. Altered Mitochondrial Function in Fibroblasts Containing MELAS or MERRF Mitochondrial DNA Mutations. Biochem. J. 1996, 318, 401–407. [Google Scholar] [CrossRef]
- Wali, G.; Li, Y.; Liyanage, E.; Kumar, K.R.; Day, M.L.; Sue, C.M. Pharmacological Rescue of Mitochondrial and Neuronal Defects in SPG7 Hereditary Spastic Paraplegia Patient Neurons Using High Throughput Assays. Front. Neurosci. 2023, 17, 1231584. [Google Scholar] [CrossRef] [PubMed]
- Chambers, S.M.; Fasano, C.A.; Papapetrou, E.P.; Tomishima, M.; Sadelain, M.; Studer, L. Highly Efficient Neural Conversion of Human ES and IPS Cells by Dual Inhibition of SMAD Signaling. Nat. Biotechnol. 2009, 27, 275–280. [Google Scholar] [CrossRef] [PubMed]
- Maroof, A.M.; Keros, S.; Tyson, J.A.; Ying, S.-W.; Ganat, Y.M.; Merkle, F.T.; Liu, B.; Goulburn, A.; Stanley, E.G.; Elefanty, A.G.; et al. Directed Differentiation and Functional Maturation of Cortical Interneurons from Human Embryonic Stem Cells. Cell Stem Cell 2013, 12, 559–572. [Google Scholar] [CrossRef] [PubMed]
- Sundberg, T.B.; Ney, G.M.; Subramanian, C.; Opipari, A.W.; Glick, G.D. The Immunomodulatory Benzodiazepine Bz-423 Inhibits B-Cell Proliferation by Targeting c-Myc Protein for Rapid and Specific Degradation. Cancer Res. 2006, 66, 1775–1782. [Google Scholar] [CrossRef] [PubMed]
- Ishiura, H.; Sako, W.; Yoshida, M.; Kawarai, T.; Tanabe, O.; Goto, J.; Takahashi, Y.; Date, H.; Mitsui, J.; Ahsan, B.; et al. The TRK-Fused Gene Is Mutated in Hereditary Motor and Sensory Neuropathy with Proximal Dominant Involvement. Am. J. Hum. Genet. 2012, 91, 320–329. [Google Scholar] [CrossRef] [PubMed]
- Beetz, C.; Johnson, A.; Schuh, A.L.; Thakur, S.; Varga, R.-E.; Fothergill, T.; Hertel, N.; Bomba-Warczak, E.; Thiele, H.; Nürnberg, G.; et al. Inhibition of TFG Function Causes Hereditary Axon Degeneration by Impairing Endoplasmic Reticulum Structure. Proc. Natl. Acad. Sci. USA 2013, 110, 5091–5096. [Google Scholar] [CrossRef] [PubMed]
- Kawarai, T.; Morita, M.; Morigaki, R.; Fujita, K.; Nodera, H.; Izumi, Y.; Goto, S.; Nakano, I.; Kaji, R. Pathomechanisms of Motor Neuron Death by Mutant TFG. Rinsho Shinkeigaku 2013, 53, 1199. [Google Scholar] [CrossRef]
- Yagi, T.; Ito, D.; Suzuki, N. TFG-Related Neurologic Disorders: New Insights into Relationships between Endoplasmic Reticulum and Neurodegeneration. J. Neuropathol. Exp. Neurol. 2016, 75, 299–305. [Google Scholar] [CrossRef]
- Johnson, A.; Bhattacharya, N.; Hanna, M.; Pennington, J.G.; Schuh, A.L.; Wang, L.; Otegui, M.S.; Stagg, S.M.; Audhya, A. TFG Clusters COPII-coated Transport Carriers and Promotes Early Secretory Pathway Organization. EMBO J. 2015, 34, 811–827. [Google Scholar] [CrossRef] [PubMed]
- Harlalka, G.V.; McEntagart, M.E.; Gupta, N.; Skrzypiec, A.E.; Mucha, M.W.; Chioza, B.A.; Simpson, M.A.; Sreekantan-Nair, A.; Pereira, A.; Günther, S.; et al. Novel Genetic, Clinical, and Pathomechanistic Insights into TFG-Associated Hereditary Spastic Paraplegia. Hum. Mutat. 2016, 37, 1157–1161. [Google Scholar] [CrossRef] [PubMed]
- Winther, M.; Walmod, P.S. Neural Cell Adhesion Molecules Belonging to the Family of Leucine-Rich Repeat Proteins. Adv. Neurobiol. 2014, 8, 315–395. [Google Scholar] [PubMed]
- Jouet, M.; Rosenthal, A.; Armstrong, G.; MacFarlane, J.; Stevenson, R.; Paterson, J.; Metzenberg, A.; Ionasescu, V.; Temple, K.; Kenwrick, S. X–Linked Spastic Paraplegia (SPG1), MASA Syndrome and X–Linked Hydrocephalus Result from Mutations in the L1 Gene. Nat. Genet. 1994, 7, 402–407. [Google Scholar] [CrossRef] [PubMed]
- Rosenthal, A.; Jouet, M.; Kenwrick, S. Aberrant Splicing of Neural Cell Adhesion Molecule L1 MRNA in a Family with X-Linked Hydrocephalus. Nat. Genet. 1992, 2, 107–112. [Google Scholar] [CrossRef] [PubMed]
- Stumpel, C.; Vos, Y.J. L1 Syndrome; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Vos, Y.J.; de Walle, H.E.K.; Bos, K.K.; Stegeman, J.A.; ten Berge, A.M.; Bruining, M.; van Maarle, M.C.; Elting, M.W.; den Hollander, N.S.; Hamel, B.; et al. Genotype-Phenotype Correlations in L1 Syndrome: A Guide for Genetic Counselling and Mutation Analysis. J. Med. Genet. 2010, 47, 169–175. [Google Scholar] [CrossRef] [PubMed]
- Weller, S.; Gärtner, J. Genetic and Clinical Aspects of X-Linked Hydrocephalus (L1 Disease): Mutations in the L1CAM Gene. Hum. Mutat. 2001, 18, 1–12. [Google Scholar] [CrossRef]
- Stevanin, G.; Azzedine, H.; Denora, P.; Boukhris, A.; Tazir, M.; Lossos, A.; Rosa, A.L.; Lerer, I.; Hamri, A.; Alegria, P.; et al. Mutations in SPG11 Are Frequent in Autosomal Recessive Spastic Paraplegia with Thin Corpus Callosum, Cognitive Decline and Lower Motor Neuron Degeneration. Brain 2008, 131, 772–784. [Google Scholar] [CrossRef]
- Pensato, V.; Castellotti, B.; Gellera, C.; Pareyson, D.; Ciano, C.; Nanetti, L.; Salsano, E.; Piscosquito, G.; Sarto, E.; Eoli, M.; et al. Overlapping Phenotypes in Complex Spastic Paraplegias SPG11, SPG15, SPG35 and SPG48. Brain 2014, 137, 1907–1920. [Google Scholar] [CrossRef]
- Hehr, U.; Bauer, P.; Winner, B.; Schule, R.; Olmez, A.; Koehler, W.; Uyanik, G.; Engel, A.; Lenz, D.; Seibel, A.; et al. Long-term Course and Mutational Spectrum of Spatacsin-linked Spastic Paraplegia. Ann. Neurol. 2007, 62, 656–665. [Google Scholar] [CrossRef] [PubMed]
- Crimella, C.; Arnoldi, A.; Crippa, F.; Mostacciuolo, M.L.; Boaretto, F.; Sironi, M.; D’Angelo, M.G.; Manzoni, S.; Piccinini, L.; Turconi, A.C.; et al. Point Mutations and a Large Intragenic Deletion in SPG11 in Complicated Spastic Paraplegia without Thin Corpus Callosum. J. Med. Genet. 2009, 46, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Lee, S.; Blackstone, C. Spastic Paraplegia Proteins Spastizin and Spatacsin Mediate Autophagic Lysosome Reformation. J. Clin. Investig. 2014, 124, 5249–5262. [Google Scholar] [CrossRef] [PubMed]
- Renvoisé, B.; Chang, J.; Singh, R.; Yonekawa, S.; FitzGibbon, E.J.; Mankodi, A.; Vanderver, A.; Schindler, A.; Toro, C.; Gahl, W.A.; et al. Lysosomal Abnormalities in Hereditary Spastic Paraplegia Types SPG15 and SPG11. Ann. Clin. Transl. Neurol. 2014, 1, 379–389. [Google Scholar] [CrossRef] [PubMed]
- Pozner, T.; Schray, A.; Regensburger, M.; Lie, D.C.; Schlötzer-Schrehardt, U.; Winkler, J.; Turan, S.; Winner, B. Tideglusib Rescues Neurite Pathology of SPG11 IPSC Derived Cortical Neurons. Front. Neurosci. 2018, 12, 914. [Google Scholar] [CrossRef]
- Hanein, S.; Martin, E.; Boukhris, A.; Byrne, P.; Goizet, C.; Hamri, A.; Benomar, A.; Lossos, A.; Denora, P.; Fernandez, J.; et al. Identification of the SPG15 Gene, Encoding Spastizin, as a Frequent Cause of Complicated Autosomal-Recessive Spastic Paraplegia, Including Kjellin Syndrome. Am. J. Hum. Genet. 2008, 82, 992–1002. [Google Scholar] [CrossRef] [PubMed]
- Goizet, C.; Boukhris, A.; Maltete, D.; Guyant-Maréchal, L.; Truchetto, J.; Mundwiller, E.; Hanein, S.; Jonveaux, P.; Roelens, F.; Loureiro, J.; et al. SPG15 Is the Second Most Common Cause of Hereditary Spastic Paraplegia with Thin Corpus Callosum. Neurology 2009, 73, 1111–1119. [Google Scholar] [CrossRef]
- Pascual, B.; de Bot, S.T.; Daniels, M.R.; França, M.C.; Toro, C.; Riverol, M.; Hedera, P.; Bassi, M.T.; Bresolin, N.; van de Warrenburg, B.P.; et al. “Ears of the Lynx” MRI Sign Is Associated with SPG11 and SPG15 Hereditary Spastic Paraplegia. Am. J. Neuroradiol. 2019, 40, 199–203. [Google Scholar] [CrossRef]
- Vantaggiato, C.; Crimella, C.; Airoldi, G.; Polishchuk, R.; Bonato, S.; Brighina, E.; Scarlato, M.; Musumeci, O.; Toscano, A.; Martinuzzi, A.; et al. Defective Autophagy in Spastizin Mutated Patients with Hereditary Spastic Paraparesis Type 15. Brain 2013, 136, 3119–3139. [Google Scholar] [CrossRef]
- Słabicki, M.; Theis, M.; Krastev, D.B.; Samsonov, S.; Mundwiller, E.; Junqueira, M.; Paszkowski-Rogacz, M.; Teyra, J.; Heninger, A.-K.; Poser, I.; et al. A Genome-Scale DNA Repair RNAi Screen Identifies SPG48 as a Novel Gene Associated with Hereditary Spastic Paraplegia. PLoS Biol. 2010, 8, e1000408. [Google Scholar] [CrossRef]
- Hirst, J.; Barlow, L.D.; Francisco, G.C.; Sahlender, D.A.; Seaman, M.N.J.; Dacks, J.B.; Robinson, M.S. The Fifth Adaptor Protein Complex. PLoS Biol. 2011, 9, e1001170. [Google Scholar] [CrossRef] [PubMed]
- Hirst, J.; Edgar, J.R.; Esteves, T.; Darios, F.; Madeo, M.; Chang, J.; Roda, R.H.; Dürr, A.; Anheim, M.; Gellera, C.; et al. Loss of AP-5 Results in Accumulation of Aberrant Endolysosomes: Defining a New Type of Lysosomal Storage Disease. Hum. Mol. Genet. 2015, 24, 4984–4996. [Google Scholar] [CrossRef]
- Ikeda, K.; Goebel, H.H.; Burck, U.; Kohlschütter, A. Ultrastructural Pathology of Human Lymphocytes in Lysosomal Disorders: A Contribution to Their Morphological Diagnosis. Eur. J. Pediatr. 1982, 138, 179–185. [Google Scholar] [CrossRef] [PubMed]
- Walkley, S.U. Cellular Pathology of Lysosomal Storage Disorders. Brain Pathol. 1998, 8, 175–193. [Google Scholar] [CrossRef] [PubMed]
- Parkinson-Lawrence, E.J.; Shandala, T.; Prodoehl, M.; Plew, R.; Borlace, G.N.; Brooks, D.A. Lysosomal Storage Disease: Revealing Lysosomal Function and Physiology. Physiology 2010, 25, 102–115. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H.; Guo, M.; Martins-Taylor, K.; Wang, X.; Zhang, Z.; Park, J.W.; Zhan, S.; Kronenberg, M.S.; Lichtler, A.; Liu, H.-X.; et al. Specification of Region-Specific Neurons Including Forebrain Glutamatergic Neurons from Human Induced Pluripotent Stem Cells. PLoS ONE 2010, 5, e11853. [Google Scholar] [CrossRef] [PubMed]
- Li, X.-J.; Du, Z.-W.; Zarnowska, E.D.; Pankratz, M.; Hansen, L.O.; Pearce, R.A.; Zhang, S.-C. Specification of Motoneurons from Human Embryonic Stem Cells. Nat. Biotechnol. 2005, 23, 215–221. [Google Scholar] [CrossRef]
- Xi, J.; Liu, Y.; Liu, H.; Chen, H.; Emborg, M.E.; Zhang, S. Specification of Midbrain Dopamine Neurons from Primate Pluripotent Stem Cells. Stem Cells 2012, 30, 1655–1663. [Google Scholar] [CrossRef]
- Murmu, R.P.; Martin, E.; Rastetter, A.; Esteves, T.; Muriel, M.-P.; El Hachimi, K.H.; Denora, P.S.; Dauphin, A.; Fernandez, J.C.; Duyckaerts, C.; et al. Cellular Distribution and Subcellular Localization of Spatacsin and Spastizin, Two Proteins Involved in Hereditary Spastic Paraplegia. Mol. Cell. Neurosci. 2011, 47, 191–202. [Google Scholar] [CrossRef]
- Joshi, D.C.; Bakowska, J.C. Determination of Mitochondrial Membrane Potential and Reactive Oxygen Species in Live Rat Cortical Neurons. J. Vis. Exp. 2011, 51, e2704. [Google Scholar] [CrossRef]
- Gottlieb, E.; Armour, S.M.; Harris, M.H.; Thompson, C.B. Mitochondrial Membrane Potential Regulates Matrix Configuration and Cytochrome c Release during Apoptosis. Cell Death Differ. 2003, 10, 709–717. [Google Scholar] [CrossRef] [PubMed]
- Cui, M.; Tang, X.; Christian, W.V.; Yoon, Y.; Tieu, K. Perturbations in Mitochondrial Dynamics Induced by Human Mutant PINK1 Can Be Rescued by the Mitochondrial Division Inhibitor Mdivi-1. J. Biol. Chem. 2010, 285, 11740–11752. [Google Scholar] [CrossRef] [PubMed]
- Hirst, J.; Bright, N.A.; Rous, B.; Robinson, M.S. Characterization of a Fourth Adaptor-Related Protein Complex. Mol. Biol. Cell 1999, 10, 2787–2802. [Google Scholar] [CrossRef] [PubMed]
- Ebrahimi-Fakhari, D.; Teinert, J.; Behne, R.; Wimmer, M.; D’Amore, A.; Eberhardt, K.; Brechmann, B.; Ziegler, M.; Jensen, D.M.; Nagabhyrava, P.; et al. Defining the Clinical, Molecular and Imaging Spectrum of Adaptor Protein Complex 4-Associated Hereditary Spastic Paraplegia. Brain 2020, 143, 2929–2944. [Google Scholar] [CrossRef] [PubMed]
- Hirst, J.; Irving, C.; Borner, G.H.H. Adaptor Protein Complexes AP-4 and AP-5: New Players in Endosomal Trafficking and Progressive Spastic Paraplegia. Traffic 2013, 14, 153–164. [Google Scholar] [CrossRef] [PubMed]
- Mattera, R.; Park, S.Y.; De Pace, R.; Guardia, C.M.; Bonifacino, J.S. AP-4 Mediates Export of ATG9A from the Trans.-Golgi Network to Promote Autophagosome Formation. Proc. Natl. Acad. Sci. USA 2017, 114, E10697–E10706. [Google Scholar] [CrossRef] [PubMed]
- Davies, A.K.; Itzhak, D.N.; Edgar, J.R.; Archuleta, T.L.; Hirst, J.; Jackson, L.P.; Robinson, M.S.; Borner, G.H.H. AP-4 Vesicles Contribute to Spatial Control of Autophagy via RUSC-Dependent Peripheral Delivery of ATG9A. Nat. Commun. 2018, 9, 3958. [Google Scholar] [CrossRef]
- Zhang, Y.; Pak, C.; Han, Y.; Ahlenius, H.; Zhang, Z.; Chanda, S.; Marro, S.; Patzke, C.; Acuna, C.; Covy, J.; et al. Rapid Single-Step Induction of Functional Neurons from Human Pluripotent Stem Cells. Neuron 2013, 78, 785–798. [Google Scholar] [CrossRef]
- Davies, A.K.; Alecu, J.E.; Ziegler, M.; Vasilopoulou, C.G.; Merciai, F.; Jumo, H.; Afshar-Saber, W.; Sahin, M.; Ebrahimi-Fakhari, D.; Borner, G.H.H. AP-4-Mediated Axonal Transport Controls Endocannabinoid Production in Neurons. Nat. Commun. 2022, 13, 1058. [Google Scholar] [CrossRef]
- Bisogno, T.; Howell, F.; Williams, G.; Minassi, A.; Cascio, M.G.; Ligresti, A.; Matias, I.; Schiano-Moriello, A.; Paul, P.; Williams, E.-J.; et al. Cloning of the First Sn1-DAG Lipases Points to the Spatial and Temporal Regulation of Endocannabinoid Signaling in the Brain. J. Cell Biol. 2003, 163, 463–468. [Google Scholar] [CrossRef]
- Williams, E.-J.; Walsh, F.S.; Doherty, P. The FGF Receptor Uses the Endocannabinoid Signaling System to Couple to an Axonal Growth Response. J. Cell Biol. 2003, 160, 481–486. [Google Scholar] [CrossRef] [PubMed]
- Mulder, J.; Aguado, T.; Keimpema, E.; Barabás, K.; Ballester Rosado, C.J.; Nguyen, L.; Monory, K.; Marsicano, G.; Di Marzo, V.; Hurd, Y.L.; et al. Endocannabinoid Signaling Controls Pyramidal Cell Specification and Long-Range Axon Patterning. Proc. Natl. Acad. Sci. USA 2008, 105, 8760–8765. [Google Scholar] [CrossRef]
- Oudin, M.J.; Hobbs, C.; Doherty, P. DAGL-dependent Endocannabinoid Signalling: Roles in Axonal Pathfinding, Synaptic Plasticity and Adult Neurogenesis. Eur. J. Neurosci. 2011, 34, 1634–1646. [Google Scholar] [CrossRef]
- Blankman, J.L.; Simon, G.M.; Cravatt, B.F. A Comprehensive Profile of Brain Enzymes That Hydrolyze the Endocannabinoid 2-Arachidonoylglycerol. Chem. Biol. 2007, 14, 1347–1356. [Google Scholar] [CrossRef] [PubMed]
- Edmison, D.; Wang, L.; Gowrishankar, S. Lysosome Function and Dysfunction in Hereditary Spastic Paraplegias. Brain Sci. 2021, 11, 152. [Google Scholar] [CrossRef] [PubMed]
- Fernandopulle, M.S.; Prestil, R.; Grunseich, C.; Wang, C.; Gan, L.; Ward, M.E. Transcription Factor–Mediated Differentiation of Human IPSCs into Neurons. Curr. Protoc. Cell Biol. 2018, 79, e51. [Google Scholar] [CrossRef]
- Tian, R.; Gachechiladze, M.A.; Ludwig, C.H.; Laurie, M.T.; Hong, J.Y.; Nathaniel, D.; Prabhu, A.V.; Fernandopulle, M.S.; Patel, R.; Abshari, M.; et al. CRISPR Interference-Based Platform for Multimodal Genetic Screens in Human IPSC-Derived Neurons. Neuron 2019, 104, 239–255.e12. [Google Scholar] [CrossRef]
- Wang, C.; Ward, M.E.; Chen, R.; Liu, K.; Tracy, T.E.; Chen, X.; Xie, M.; Sohn, P.D.; Ludwig, C.; Meyer-Franke, A.; et al. Scalable Production of IPSC-Derived Human Neurons to Identify Tau-Lowering Compounds by High-Content Screening. Stem Cell Rep. 2017, 9, 1221–1233. [Google Scholar] [CrossRef]
- Braulke, T.; Bonifacino, J.S. Sorting of Lysosomal Proteins. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2009, 1793, 605–614. [Google Scholar] [CrossRef]
- Wang, L.; Brown, A. A Hereditary Spastic Paraplegia Mutation in Kinesin-1A/KIF5A Disrupts Neurofilament Transport. Mol. Neurodegener. 2010, 5, 52. [Google Scholar] [CrossRef]
- Okada, Y.; Yamazaki, H.; Sekine-Aizawa, Y.; Hirokawa, N. The Neuron-Specific Kinesin Superfamily Protein KIF1A Is a Uniqye Monomeric Motor for Anterograde Axonal Transport of Synaptic Vesicle Precursors. Cell 1995, 81, 769–780. [Google Scholar] [CrossRef]
- Yonekawa, Y.; Harada, A.; Okada, Y.; Funakoshi, T.; Kanai, Y.; Takei, Y.; Terada, S.; Noda, T.; Hirokawa, N. Defect in Synaptic Vesicle Precursor Transport and Neuronal Cell Death in KIF1A Motor Protein–Deficient Mice. J. Cell Biol. 1998, 141, 431–441. [Google Scholar] [CrossRef] [PubMed]
- Della Vecchia, S.; Tessa, A.; Dosi, C.; Baldacci, J.; Pasquariello, R.; Antenora, A.; Astrea, G.; Bassi, M.T.; Battini, R.; Casali, C.; et al. Correction to: Monoallelic KIF1A-Related Disorders: A Multicenter Cross Sectional Study and Systematic Literature Review. J. Neurol. 2023, 270, 2345–2346. [Google Scholar] [CrossRef] [PubMed]
- Tesson, C.; Nawara, M.; Salih, M.A.M.; Rossignol, R.; Zaki, M.S.; Al Balwi, M.; Schule, R.; Mignot, C.; Obre, E.; Bouhouche, A.; et al. Alteration of Fatty-Acid-Metabolizing Enzymes Affects Mitochondrial Form and Function in Hereditary Spastic Paraplegia. Am. J. Hum. Genet. 2012, 91, 1051–1064. [Google Scholar] [CrossRef] [PubMed]
- Pujol, C.; Legrand, A.; Parodi, L.; Thomas, P.; Mochel, F.; Saracino, D.; Coarelli, G.; Croon, M.; Popovic, M.; Valet, M.; et al. Implication of Folate Deficiency in CYP2U1 Loss of Function. J. Exp. Med. 2021, 218, e20210846. [Google Scholar] [CrossRef] [PubMed]
- Husain, R.A.; Grimmel, M.; Wagner, M.; Hennings, J.C.; Marx, C.; Feichtinger, R.G.; Saadi, A.; Rostásy, K.; Radelfahr, F.; Bevot, A.; et al. Bi-Allelic HPDL Variants Cause a Neurodegenerative Disease Ranging from Neonatal Encephalopathy to Adolescent-Onset Spastic Paraplegia. Am. J. Hum. Genet. 2020, 107, 364–373. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.G.; Lee, S.; Fabunan, R.; Chai, G.; Zaki, M.S.; Abdel-Salam, G.; Sultan, T.; Ben-Omran, T.; Alvi, J.R.; McEvoy-Venneri, J.; et al. Biallelic Variants in HPDL, Encoding 4-Hydroxyphenylpyruvate Dioxygenase-like Protein, Lead to an Infantile Neurodegenerative Condition. Genet. Med. 2021, 23, 524–533. [Google Scholar] [CrossRef] [PubMed]
- Wiessner, M.; Maroofian, R.; Ni, M.-Y.; Pedroni, A.; Müller, J.S.; Stucka, R.; Beetz, C.; Efthymiou, S.; Santorelli, F.M.; Alfares, A.A.; et al. Erratum to: Biallelic Variants in HPDL Cause Pure and Complicated Hereditary Spastic Paraplegia. Brain 2021, 144, e70. [Google Scholar] [CrossRef]
- Banh, R.S.; Kim, E.S.; Spillier, Q.; Biancur, D.E.; Yamamoto, K.; Sohn, A.S.W.; Shi, G.; Jones, D.R.; Kimmelman, A.C.; Pacold, M.E. The Polar Oxy-Metabolome Reveals the 4-Hydroxymandelate CoQ10 Synthesis Pathway. Nature 2021, 597, 420–425. [Google Scholar] [CrossRef]
- Sloan, S.A.; Andersen, J.; Pașca, A.M.; Birey, F.; Pașca, S.P. Generation and Assembly of Human Brain Region–Specific Three-Dimensional Cultures. Nat. Protoc. 2018, 13, 2062–2085. [Google Scholar] [CrossRef]
- Andersen, J.; Revah, O.; Miura, Y.; Thom, N.; Amin, N.D.; Kelley, K.W.; Singh, M.; Chen, X.; Thete, M.V.; Walczak, E.M.; et al. Generation of Functional Human 3D Cortico-Motor Assembloids. Cell 2020, 183, 1913–1929.e26. [Google Scholar] [CrossRef]
- Salter, M.W.; Stevens, B. Microglia Emerge as Central Players in Brain Disease. Nat. Med. 2017, 23, 1018–1027. [Google Scholar] [CrossRef]
- Lendahl, U.; Nilsson, P.; Betsholtz, C. Emerging Links between Cerebrovascular and Neurodegenerative Diseases—A Special Role for Pericytes. EMBO Rep. 2019, 20, e48070. [Google Scholar] [CrossRef] [PubMed]
- Matsui, T.K.; Tsuru, Y.; Hasegawa, K.; Kuwako, K. Vascularization of Human Brain Organoids. Stem Cells 2021, 39, 1017–1024. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Jiang, J.; Xu, Z.; Yan, H.; Tang, B.; Liu, C.; Chen, C.; Meng, Q. Microglia-Containing Human Brain Organoids for the Study of Brain Development and Pathology. Mol. Psychiatry 2023, 28, 96–107. [Google Scholar] [CrossRef] [PubMed]
- Dahan, P.; Lu, V.; Nguyen, R.M.T.; Kennedy, S.A.L.; Teitell, M.A. Metabolism in Pluripotency: Both Driver and Passenger? J. Biol. Chem. 2019, 294, 5420–5429. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Zhang, Y. Genetic and Epigenetic Variations in IPSCs: Potential Causes and Implications for Application. Cell Stem Cell 2013, 13, 149–159. [Google Scholar] [CrossRef] [PubMed]
- Volpato, V.; Webber, C. Addressing Variability in IPSC-Derived Models of Human Disease: Guidelines to Promote Reproducibility. Dis. Model. Mech. 2020, 13, dmm042317. [Google Scholar] [CrossRef] [PubMed]
- Grobarczyk, B.; Franco, B.; Hanon, K.; Malgrange, B. Generation of Isogenic Human IPS Cell Line Precisely Corrected by Genome Editing Using the CRISPR/Cas9 System. Stem Cell Rev. Rep. 2015, 11, 774–787. [Google Scholar] [CrossRef]
- Li, H.; Busquets, O.; Verma, Y.; Syed, K.M.; Kutnowski, N.; Pangilinan, G.R.; Gilbert, L.A.; Bateup, H.S.; Rio, D.C.; Hockemeyer, D.; et al. Highly Efficient Generation of Isogenic Pluripotent Stem Cell Models Using Prime Editing. eLife 2022, 11, e79208. [Google Scholar] [CrossRef]
- Araki, H.; Miura, F.; Watanabe, A.; Morinaga, C.; Kitaoka, F.; Kitano, Y.; Sakai, N.; Shibata, Y.; Terada, M.; Goto, S.; et al. Base-Resolution Methylome of Retinal Pigment Epithelial Cells Used in the First Trial of Human Induced Pluripotent Stem Cell-Based Autologous Transplantation. Stem Cell Rep. 2019, 13, 761–774. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, J. Clinical Trial for Parkinson’s Disease Gets a Green Light in the US. Cell Stem Cell 2021, 28, 182–183. [Google Scholar] [CrossRef]
- McKenna, D.H.; Perlingeiro, R.C.R. Development of Allogeneic iPS Cell-based Therapy: From Bench to Bedside. EMBO Mol. Med. 2023, 15, e15315. [Google Scholar] [CrossRef]
- Handel, A.E.; Chintawar, S.; Lalic, T.; Whiteley, E.; Vowles, J.; Giustacchini, A.; Argoud, K.; Sopp, P.; Nakanishi, M.; Bowden, R.; et al. Assessing Similarity to Primary Tissue and Cortical Layer Identity in Induced Pluripotent Stem Cell-Derived Cortical Neurons through Single-Cell Transcriptomics. Hum. Mol. Genet. 2016, 25, 989–1000. [Google Scholar] [CrossRef] [PubMed]
- Logan, S.; Arzua, T.; Yan, Y.; Jiang, C.; Liu, X.; Yu, L.-K.; Liu, Q.-S.; Bai, X. Dynamic Characterization of Structural, Molecular, and Electrophysiological Phenotypes of Human-Induced Pluripotent Stem Cell-Derived Cerebral Organoids, and Comparison with Fetal and Adult Gene Profiles. Cells 2020, 9, 1301. [Google Scholar] [CrossRef] [PubMed]
- Vadodaria, K.C.; Jones, J.R.; Linker, S.; Gage, F.H. Modeling Brain Disorders Using Induced Pluripotent Stem Cells. Cold Spring Harb. Perspect. Biol. 2020, 12, a035659. [Google Scholar] [CrossRef]
- Vandana, J.J.; Manrique, C.; Lacko, L.A.; Chen, S. Human Pluripotent-Stem-Cell-Derived Organoids for Drug Discovery and Evaluation. Cell Stem Cell 2023, 30, 571–591. [Google Scholar] [CrossRef]


| HSP Designation | Human Gene | Protein Name | Animal Model | Gene Modification | Main Phenotype | References |
|---|---|---|---|---|---|---|
| SPG3A | ATL1 | ATLASTIN1 | Mouse | KD in primary cortical neurons | Impairment of axonal growth and dendritic branching defects | [19] |
| OE in cortical upper layer neurons (IUE) of WT and R217Q Atlastin1 | Increase in the formation and/or maintenance of new segments within existing dendritic trees in neurons by WT but not mutant variant. No effect on radial cortical migration. | [20] | ||||
| Rat | Expression of ATL1 R217Q mutant (Primary hippocampal neurons) | Impaired protein synthesis. Reduced dendritic spine density | [21] | |||
| Danio rerio | Morphant | Significant reduction of larval motility, with an increase of axonal branching in spinal motoneurons. Phenotype specific to motor and cerebellar neurons. | [22,23] | |||
| OE (human and zebrafish mRNA) | Complete loss of ventral structures likely via inhibition of BMP signaling | [22,23] | ||||
| Drosophila melanogaster | KO | High mortality. Small size. Age-dependent locomotor deficits due to degeneration of dopaminergic neurons. Rescued by levo-dopa and SK&F 38393 (dopamine agonists) | [24] | |||
| KO | Reduction of muscle size, increase of synaptic and satellite boutons in motor neurons | [25] | ||||
| KD and OE in motor neurons | Larvae: decreased crawling speed and contraction frequency; Adult flies: age-dependent degeneration of dopaminergic neurons, decline in climbing ability | [26] | ||||
| OE of pathological variants (R214C, C350R, M383T, R192Q) | Lethality at larval stage and eye size reduction. Heavy overfusion of ER membranes in neuronal cytoplasm. Phenotype severity is dependent on the % of activity of the protein variant (R214C > C350R > M383T > R192Q, with the last having almost no phenotype). | [27] | ||||
| CRISPR-Cas9 mediated KI of pathological variants (R214C, C350R, M383T, R192Q) | Decrease of the adult eclosion rate and a reduction in size of all developmental stages. Phenotype severity is dependent on the % of activity of the protein variant (R192Q > M383T > C350R > R214C, with the last having almost no phenotype). | [27] | ||||
| KD | Embryonic lethality, reduced lifespan of surviving animals. Eye size reduction. ER reduction and fragmentation in neurons. No impairment of secretory traffic | [28] | ||||
| Eye/neural ganglia specific OE of WT Atlastin | Eye size reduction | [28] | ||||
| Motoneuron specific OE of WT Atlastin | Expansion of ER membranes, severe impairment of secretory traffic (absence of Golgi complex) | [28] | ||||
| OE of mutant K51A variant (different drivers) | Normal eyes and fly survival | [28] | ||||
| Motoneuron specific KO | Increases synaptic and satellite boutons in the same way that constitutively activating the BMP-receptor Tkv (thick veins). | [29] | ||||
| SPG4 | SPAST | SPASTIN | Mouse | KO (SpΔ/Δ) | Late and mild motor defect for progressive CNS axonal degeneration. Presence of focal axonal swellings, associated with abnormal accumulation of organelles and cytoskeletal components. Focal impairment of retrograde transport | [30] |
| KO (SpastΔE7/ΔE7) | Gait abnormalities.Axonal swellings and impaired anterograde transport in primary cortical neurons. Late and mild motor defect for progressive CNS axonal degeneration. transport and outgrowth deficits accompanied by reduced motor control. Deficits in working and associative memory | [31] | ||||
| KO (SpΔ/Δ; Primary cortical neurons) | Axonal swellings and cargo stalling, likely due to impairment of microtubule dynamics. Phenotype rescued by microtubule-targeting drugs | [32] | ||||
| KI (ROSA26R-hSPAST-C448Y) | Adult-onset gait deficiencies and corticospinal degeneration. No axonal swellings | [33] | ||||
| Spast KO X Spast KI (ROSA26R-hSPAST-C448Y) | Same features as the hSPASTC448Y mouse, but with axonal swellings and stronger and earlier onset | [33] | ||||
| Rat | KD in hippocampal neurons | Reduction of axonal branches and overall length | [34] | |||
| OE of WT Spastin in hippocampal neurons | Increase in axonal branches (mostly minor processes) | [34] | ||||
| OE of human M1-C448Y and M87-C448Y variants in cortical neurons | Decreased ratio of cells with long processes, increased ratio of cells with short processes. M1-C448Y decreased outgrowth | [35] | ||||
| OE of M1 N184X, M1 S245X, or M87 S245X truncated variants in primary cortical neurons | Decreased neuritic outgrowth | [36] | ||||
| Danio rerio | Morphant | Impairment of branchiomotor and spinal mototoneuron axonal outhgrowth. CNS-specific apoptosis. Motility defects. Phenotype is not rescued by nocodazole | [37,38] | |||
| Morphant (Spast X Protrudin) | Severe impairment of spinal and branchiomotor axonal outgrowth. Rescued by OE of human Spastin and Protrudin | [39] | ||||
| Morphants (specific for DrM1 or DrM61 isoforms) | Curved-tail phenotype and yolk tube agenesis in KD of M1 isoform. Smaller eyes in KD of M61 isoform. Isoform-specific motor neuron and locomotion defects, not rescued by the selective expression of the other isoform. | [23] | ||||
| Morphant | Aberrant branching and truncation of spinal motor axons, associated with dysregulated endosomal tubulation | [40] | ||||
| Morphant | Hydrocephalia, perturbed yolk sac extension, arched-back phenotype, and high oxydative stress. Partially rescued by phenazine, methylene blue, guanabenz and salubrinal | [41] | ||||
| KO (CRISPant) | Impairment of metabolic properties: reduction of mitochondrial respiration in larvae; reduction size, weight and BMI in adults. Impairment of sarcoplasmic reticulum in skeletal muscles | [42] | ||||
| C. elegans | KD | Progressive motor defects, reduced lifespan, ER stress and increased oxydative stress. Rescued by guanabenz, salubrinal, phenazine, and methylene blue | [41] | |||
| Drosophila melanogaster | KO (spastin5.75) | Severe movement defects. Larval neuromuscular junction (NMJ) phenotypes (boutons are smaller, more numerous and clustered; synaptic transmission is impaired) | [43] | |||
| Neuronal-specific OE | Embryonic CNS collapse | [43] | ||||
| KD (ubiquitous) | Lethal in embryonic or early larval stage | [44] | ||||
| OE (ubiquitous) | Lethal in embryonic or early larval stage | [44] | ||||
| Neuronal KD | Severely impaired adult locomotor performance, apoptosis, decreased lifespan. Partial rescue by treatment with vinblastine | [44,45] | ||||
| Neuronal OE (Dspastin K467R) | Severely impaired adult locomotor performance, apoptosis, decreased lifespan. Partial rescue by treatment with vinblastine | [44,45] | ||||
| Neuronal OE of human M1-C448Y and M87-C448Y variants | Mutants M1 spast showed a more severe phenotype than mutant M87 with a high axonal degeneration in the corticospinal tracts. | [35] | ||||
| Neuronal-specific KD | Severe climbing defects and ER stress. Rescued by methylene blue, phenazine, and N-acetyl-L-cysteine | [41] | ||||
| KO (spastin5.75) X Glial KD of Pak3 | No phenotype observed (glial involvement in spastin phenotype) | [46] | ||||
| SPG7 | SPG7 | PARAPLEGIN | Mouse | KO | Spastic-ataxic symptoms associated to distal axonopathy of spinal and peripheral axons, characterized by axonal swelling and degeneration due to accumulation of organelles and neurofilaments. Rescued with intramuscular injection of AAV-Paraplegin | [47,48] |
| KO (cortical neurons) | Impaired mitochondrial permeability and altered electrophysiological patterns. Rescued by treatment with Bz-423 | [49] | ||||
| KO | Partial rescue of motor phenotype by Bz-423 administration | [49] | ||||
| Spg7 X Afg3l2 double mutant | Early-onset spastic phenotye, complicated with cerebellar ataxia (closer to human disease) | [50] | ||||
| eSpg7 KO (Spg7 X Afgrl1 double mutant) | Progressive motor phenotype with uncoordinated gait, and degeneration not only of long spinal axons, but also of spinocerebellar and cerebellar granule axons | [51] | ||||
| Drosophila melanogaster | KO | Reduced lifespan. Progressive loss of ability to control movements, greater sensitivity to chemical and environmental stress, muscular and neuronal degeneration. Reduced activity of respiratory chain complexes I and II, severely swollen and aberrantly shaped mitochondria in the synaptic terminals of photoreceptors. | [52] | |||
| SPG5A | CYP7B1 | CYP7B1 | Mouse | KO | Accumulation of 25- and 27-hydroxycholesterol in plasma and tissues, normal levels of brain-derived 24(S)-hydroxycholesterol. No recapitulation of human disease: normal neuronal morphology, no neurological defects, no motor defect, no learning and memory capacities. | [53,54] |
| SPG11 | SPG11/KIAA1840 | SPATACSIN | Mouse | KD (Primary cortical neurons) | Reduction of axonal outgrowth | [55] |
| KO | Spastic paraplegia and progressive loss of cortical motoneurons and Purkinje cells for accumulation of autolysosome-derived material. No motor or cognitive deficits, amyotrophy or alterations of the corpus callosum | [56] | ||||
| KO (stop codons exon 32) | Early-onset motor and cognitive deficit. Cortical, cerebellar, hippocampal and spinal cord atrophy. Corticospinal tract degeneration. Accumulation of gangliosides in lysosomes. Partial rescue in primary neurons by miglustat | [57,58] | ||||
| KO (primary cortical neurons) | Decreased number of tubular lysosomes in axons | [59] | ||||
| Danio rerio | Morphant | Abnormal twisted/bent tail phenotype. Mild hydrocephaly, small eyes, generalized perturbation of neuronal differentiation. Irregular motor neuron outgrowth | [60] | |||
| Morphant | Abnormal twisted/bent tail phenotype. No major brain abnormalities. Reduction of larval motility. | [61] | ||||
| Morphant | Loss of motility or paralysis accompanied by accumulation of GM2. Partial rescue by miglustat | [58] | ||||
| SPG15 | ZFYVE26 | SPASTIZIN | Mouse | KO | Late-onset spastic paraplegia with cerebellar ataxia due to loss of both cortical motoneurons and Purkinje cells. High levels of lysosomal enzymes in brain of aged mice (lysosomal dysfunction) | [62] |
| zfyve26 X spg11 double KO | Simultaneous disruption does not aggravate the phenotype | [63] | ||||
| KO (primary cortical neurons) | Defective anterograde transport, impaired neurite outgrowth, axonal swelling, reduced autophagic flux, lipid accumulation within the lysosomal compartment. Synaptic dysfunction, accompanied by augmented vulnerability to glutamate-induced excitotoxicity | [64] | ||||
| Danio rerio | Morphant | Severely curly tails. Reduction of head and eye volume. Motor deficits. Impaired spinal motor neuron axon outgrowth and neuromuscular junction connections | [61] | |||
| Drosophila melanogaster | KO | Reduced lifespan, locomotor deficits. Autophagosome accumulation, enlarged lysosomes, reduced free lysosomes, autophagic lysosome reformation defects. Small-Molecule Enhancer of Rapamycin 28 rescues both autophagic lysosome reformation defects and locomotor deficit. | [65] | |||
| SPG48 | AP5Z1 | AP5Z1 | Mouse | KO | Age-dependent degeneration of corticospinal axons due to a block of autophagic flux. Late onset progressive gait abnormalities (recapitulates human phenotype) | [66] |
| SPG47 | AP4B1 | AP4B1 | Mouse | KO | Thin corpus callosum, enlarged lateral ventricles, motor co-ordination deficits, hyperactivity, a hindlimb clasping phenotype, and neurodegeneration. Accumulation of AMPA receptors in autophagosomes in axons of Purkinje cells and hippocampal neurons | [67,68] |
| Danio rerio | CRISPant | Microcephaly, neural necrosis. Alteration of expression level and pattern of autophagy genes atg9a and map1lc3b. | [69] | |||
| SPG50 | AP4M1 | AP4M1 | Danio rerio | CRISPant | Microcephaly, neural necrosis. Alteration of expression level and pattern of autophagy genes atg9a and map1lc3b. | [69] |
| SPG51 | AP4E1 | AP4E1 | Mouse | KO | Hindlimb clasping, decreased motor coordination and weak grip strength. Thin corpus callosum, lateral ventricle enlargement. Defective autophagosome maturation in leading to axonal swellings and reduced elongation. in various areas of the brain and spinal cord. | [70,71] |
| Danio rerio | CRISPant | Microcephaly, neural necrosis. Alteration of expression level and pattern of autophagy genes atg9a and map1lc3b. | [69] | |||
| SPG52 | AP4S1 | AP4S1 | Danio rerio | Morphant | Reduction of head size, altered CNS development, locomotor deficits, and abnormal neuronal excitability. Shorter axonal length in spinal cord neurons | [72] |
| CRISPant | Microcephaly, neural necrosis. Alteration of expression level and pattern of autophagy genes atg9a and map1lc3b. | [69] | ||||
| SPG57 | TFG | TFG | C. elegans | KD | Impaired protein secretion for defects in assembly of complexes containing SEC-16 and COPII proteins | [73] |
| Mouse | β-cell specific KO | Marked glucose intolerance with reduced insulin secretion, corrlated with smaller β-cell islets | [74] | |||
| Spinal motoneuron specific KO | Deterioration of motor function and muscle atrophy due to denervation of neuromuscular junctions (NMJs) | [75] | ||||
| Muscle specific KO | Normal motor functions, no muscle atrophy. Possible phenotype in aging | [75] | ||||
| OE of pathological p.G269V or p.P265A variants in primary cortical neurons | Slightly reduced neurite length and an increased apoptotic cell ratio upon OE of p.P265A variant | [76] | ||||
| KD in cortical neurons | Late neurite degeneration and neuronal apoptosis | [76] | ||||
| Rat | KI for p.R106C mutation (CRISPR-Cas9 mediated) | Progressive motor deficits, thinning of the corpus callosum, ventriculomegaly, and hind limb spasticity. Slowed trafficking of integral membrane proteins. No lysosomal defects. | [77] | |||
| Danio rerio | Translation blocking morphant (ATG-MO) and splice-blocking morphant (E3I3-MO) | Malformations with a curved body axis, reduced motor capacity, high mortality, significant increase in neuronal apoptosic cells in both brain and spinal cord | [76] |
| HSP Designation | Gene | Protein Name | Type of HSP | Onset | Distinguishing Clinical Features | Frequency | Inheritance | iPSCs/hES-Experiments Done | References |
|---|---|---|---|---|---|---|---|---|---|
| SPG1 | L1CAM | L1CAM | Complicated | Infancy | ID, Adducted thumb, Corpus callosum hypoplasia, Aphasia, Obstructive hydrocephalus | Rare | X-Linked | Generation of L1CAM shRNA depleted hES cells | [88] |
| Generation of iPS model | [89] | ||||||||
| SPG3A | ATL1 | ATLASTIN1 | Pure | Infantile to childhood (rarely adult onset) | Very slow progression. May present as spastic diplegic cerebral palsy. Complicated phenotype with peripheral neuropathy or autonomic failure reported | 80% of early-onset AD HSP, 10–15% of all AD HSP | AD | Differentiation of forebrain neurons | [90] |
| AR inheritance is very rare | AR | Differentiation of lower motoneurons and in vitro neuromuscular junction | [91] | ||||||
| SPG4 | SPAST | SPASTIN | Pure | Variable from infancy to 7th decade | Cognitive decline & dementia common, Distal amyotrophy variably present, Complicated phenotype with ataxia variably present | 40% of AD HSP | AD | Neurons differentiated from Olfactory neurosphere-derived stem cells taken from patients | [92] |
| Differentiation of cortical neurons; SPAST KD in hES | [93] | ||||||||
| Differentiation of neurons (putative cortical) | [94] | ||||||||
| Neurons differentiated from Olfactory neurosphere-derived stem cells taken from patients | [95] | ||||||||
| Differentiation of lower motoneurons and in vitro neuromuscular junction | [91] | ||||||||
| SPG5 | CYP7B1 | CYP7B1 | Pure or complicated | Juvenile to early adulthood | Ataxia, Polyneuropathy, Extrapyramidal signs, MRI signs of leukodystrophy | 9 of 172 families with AR HSP | AR | Differentiation of forebrain glutamatergic neurons from patient derived and CRISPR-Cas9 mediated KO | [96] |
| SPG7 | SPG7 | PARAPLEGIN | Pure or complicated | Juvenile or adulthood | Dysarthria, Ataxia, Optic atrophy, Supranuclear palsy, Mitochondrial abnormalities on skeletal muscle biopsy | 5–12% of AR HSP. AD inheritance suggested for some pathogenic variants | AR | Olfactory neurosphere-derived stem cells taken from patients | [97] |
| SPG10 | KIF5A | KINESIN5A | Complicated | Juvenile or adulthood | Polyneuropathy; Pes cavus | 1–2% of all AD HSP. 5–8% of all complicated AD HSP | AD | Generation of iPS model | [98] |
| SPG11 | SPG11/KIAA1840 | SPATACSIN | Complicated | Childhood or early adulthood | DD, Optic atrophy, Ataxia, Pseudobulbar signs, Polyneuropathy, Levodopa-responsive parkinsonism, Hypoplastic or absent corpus callosum | 5% of AR HSP; 75% of HSP with developmental delay and hypoplasia of corpus callosum | AR | Neurons differentiated from SPG11 hiPS cells | [55] |
| Cortical neurons differentiated from SPG11 hiPS cells | [99] | ||||||||
| Human SPG11 cerebral organoids | [100] | ||||||||
| Motoneurons differentiated from SPG11 hiPS cells | [101] | ||||||||
| SPG15 | ZFYVE26 | SPASTIZIN | Complicated | Childhood or early adulthood | DD, Optic atrophy, Ataxia, Pseudobulbar signs, Central retinal degeneration, Polyneuropathy | 1–2% of AR HSP | AR | Cortical neurons, spinal MNs and DA neurons differentiated from SPG15 iPS | [102] |
| SPG30 | KIF1A | KINESIN1A | Pure (for AD inheritance) | Juvenile to adulthood | Mild ID in some individuals. Optic nerve atrophy. Epilepsy (rare) | 5–6% of all AD HSP | AD | Generation of iPS model | [103] |
| SPG43 | C19orf12 | Complicated | Childhood | Amyotrophy, Dysarthria, Multiple contractures, Neurodegeneration with brain iron accumulation | Rare | AR | Generation of iPS model | [104] | |
| SPG48 | AP5Z1 | AP5Z1 | Pure | Typically adulthood; rarely infancy | Urinary incontinence, Parkinsonism, Dystonia, Thin corpus callosum, Leukodystrophy, Severe DD in infantile onset | Single family | AR | Cortical neurons, spinal MNs and DA neurons differentiated from SPG48 iPS | [102] |
| SPG47 | AP4B1 | AP4B1 | Complicated | Infancy | Severe ID, Facial dysmorphism, Seizures, Stereotypic laughter w/tongue protrusion | Rare | AR | Cortical neurons differentiated from iPS lines with loss of function of several AP4 complex subunit | [105] |
| SPG50 | AP4M1 | AP4M1 | |||||||
| SPG51 | AP4E1 | AP4E1 | |||||||
| SPG52 | AP4S1 | AP4S1 | |||||||
| SPG56 | CYP2U1 | CYP2U1 | Complicated | Infancy | Severe DD, Dystonia, Polyneuropathy, Calcification of basal ganglia | Rare | AR | Generation of iPS model | [106] |
| SPG57 | TFG | TFG | Complicated | Childhood | Optic atrophy, Severe polyneuropathy | Rare | AR | Knock-in hES cells for mutant form of TFG. Differentiation of cortical neurons | [107] |
| SPG58 | KIF1C | KINESIN1C | Complicated | Childhood | Spastic ataxia, Dystonia | Rare | AR | Generation of iPS model | [108] |
| SPG59 | USP8 | USP8 | Pure | Childhood | Lower limb spasticity, borderline intellectual disability. Nystagmus and pes equinovarus | Rare | AR | USP8 KO murine and human ES cells | [109] |
| SPG76 | CAPN1 | CALPAIN 1 | Complicated | Young adulthood | Spasticity of the lower limbs. Cerebellar ataxia. Upper limb involvement, foot deformities, dysarthria. Cognition unaffected | Extremely rare | AR | Generation of iPS model | [110] |
| HSP Designation | Gene | Gene ID | Genomic Location | RefSeq Transcripts |
|---|---|---|---|---|
| SPG1 | L1CAM | 3897 | chrX:153861516-153888990 | 4 transcript variants (NM_000425.5; NM_024003.3; NM_001143963.2; NM_001278116.2) |
| SPG3A | ATL1 | 51062 | chr14:50560145-50633045 | 3 transcript variants (NM_015915.5; NM_181598.4; NM_001127713.1) |
| SPG4 | SPAST | 6683 | chr2:32063556-32157637 | 5 transcript variants (NM_014946.4; NM_199436.2; NM_001363823.2; NM_001363875.2; NM_001377959.1) |
| SPG5 | CYP7B1 | 9420 | chr8:64590851-64798737 | 2 transcript variants (NM_004820.5; NM_001324112.2) |
| SPG7 | SPG7 | 6687 | chr16:89508403-89557766 | 3 transcript variants (NM_003119.4; NM_199367.3; NM_001363850.1) |
| SPG10 | KIF5A | 3798 | chr12:57550044-57586633 | 2 transcript variants (NM_004984.4; NM_001354705.2) |
| SPG11 | SPG11/KIAA1840 | 80208 | chr15:44562696-44663662 | 3 transcript variants (NM_025137.4; NM_001160227.2; NM_001411132.1) |
| SPG15 | ZFYVE26 | 23503 | chr14:67746522-67816590 | 1 transcript variant (NM_015346.4) |
| SPG30 | KIF1A | 547 | chr2:240713767-240820219 | 23 transcript variants (NM_001244008.2; NM_004321.8; NM_001330289.2; NM_001330290.2; NM_001379631.1; NM_001379632.1; NM_001379633.1; NM_001379634.1; NM_001379635.1; NM_001379636.1; NM_001379637.1; NM_001379638.1; NM_001379639.1; NM_001379640.1; NM_001379641.1; NM_001379642.1; NM_001379645.1; NM_001379646.1; NM_001379648.1; NM_001379649.1; NM_001379650.1; NM_001379651.1; NM_001379653.1) |
| SPG43 | C19orf12 | 83636 | chr19:29698937-29715261 | 7 transcript variants (NM_001031726.4; NM_031448.6; NM_001256046.3; NM_001256047.2; NM_001282929.1; NM_001282930.3; NM_001282931.3) |
| SPG48 | AP5Z1 | 9907 | chr7:4775623-4794397 | 3 transcript variants (NM_014855.3; NM_001364858.1; NR_157345.1) |
| SPG47 | AP4B1 | 10717 | chr1:113894194-113904799 | 4 transcript variants (NM_006594.5; NM_001253852.3; NM_001253853.3; NM_001308312.2) |
| SPG50 | AP4M1 | 9179 | chr7:100101643-100109039 | 1 transcript variant (NM_001363671.2) |
| SPG51 | AP4E1 | 23431 | chr15:50908683-51005895 | 2 transcript variants (NM_007347.5; NM_001252127.2) |
| SPG52 | AP4S1 | 11154 | chr14:31025649-31096450 | 6 transcript variants (NM_007077.5 5; NM_001128126.3; NM_001254726.2; NM_001254727.2; NM_001254728.2; NM_001254729.2) |
| SPG56 | CYP2U1 | 113612 | chr4:107931549-107953461 | 1 transcript variant (NM_183075.3) |
| SPG57 | TFG | 10342 | chr3:100709494-100748964 | 4 transcript variants (NM_006070.6; NM_001007565.2; NM_001195478.2; NM_001195479.2) |
| SPG58 | KIF1C | 10749 | chr17:4997950-5028401 | 1 transcript variant (NM_006612.6) |
| SPG59 | USP8 | 9101 | chr15:50424405-50514421 | 3 transcript variants (NM_005154.5; NM_001128610.3; NM_001283049.2) |
| SPG76 | CAPN1 | 823 | chr11:65181928-65212006 | 4 transcript variants (NM_001198868.2; NM_005186.4; NM_001198869.2; NR_040008.2) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Damiani, D.; Baggiani, M.; Della Vecchia, S.; Naef, V.; Santorelli, F.M. Pluripotent Stem Cells as a Preclinical Cellular Model for Studying Hereditary Spastic Paraplegias. Int. J. Mol. Sci. 2024, 25, 2615. https://doi.org/10.3390/ijms25052615
Damiani D, Baggiani M, Della Vecchia S, Naef V, Santorelli FM. Pluripotent Stem Cells as a Preclinical Cellular Model for Studying Hereditary Spastic Paraplegias. International Journal of Molecular Sciences. 2024; 25(5):2615. https://doi.org/10.3390/ijms25052615
Chicago/Turabian StyleDamiani, Devid, Matteo Baggiani, Stefania Della Vecchia, Valentina Naef, and Filippo Maria Santorelli. 2024. "Pluripotent Stem Cells as a Preclinical Cellular Model for Studying Hereditary Spastic Paraplegias" International Journal of Molecular Sciences 25, no. 5: 2615. https://doi.org/10.3390/ijms25052615
APA StyleDamiani, D., Baggiani, M., Della Vecchia, S., Naef, V., & Santorelli, F. M. (2024). Pluripotent Stem Cells as a Preclinical Cellular Model for Studying Hereditary Spastic Paraplegias. International Journal of Molecular Sciences, 25(5), 2615. https://doi.org/10.3390/ijms25052615





