Chromosome Division in Early Embryos—Is Everything under Control? And Is the Cell Size Important?
Abstract
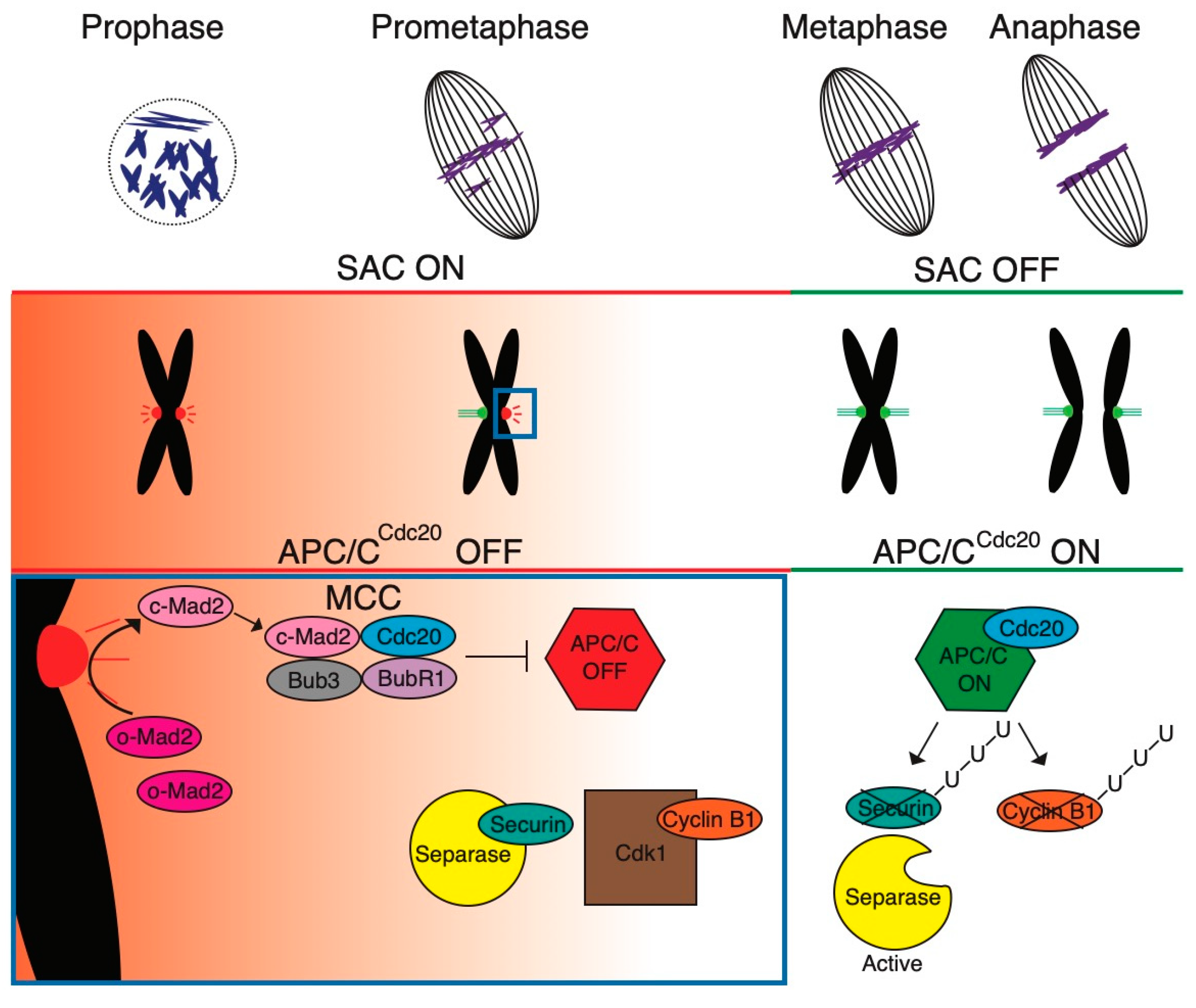
1. The Assembly of the Spindle Is under Surveillance
2. Important Aspects of Early Embryonic Development with Potential Impact on Chromosome Segregation
3. Control of Chromosome Segregation during Early Embryonic Development
4. Is Cell Size Important for the Fidelity of Chromosome Segregation?
5. Conclusions
Funding
Conflicts of Interest
Abbreviations
| APC/C | anaphase-promoting complex/cyclosome |
| Bub1 | budding uninhibited by benzimidazole 1 |
| Bub3 | budding uninhibited by benzimidazole 3 |
| BubR1 | budding uninhibited by benzimidazole-related 1 |
| c-Mad2 | closed Mad2 |
| Cdc20 | cell division cycle 20 |
| CDK | cyclin-dependent kinase |
| CDK1 | cyclin-dependent kinase 1 |
| CPC | chromosomal passenger complex |
| GVBD | germinal vesicle breakdown |
| KMN | KNL1-MIS12-NDC80 complex |
| KNL1 | kinetochore scaffold 1 |
| Mad1 | mitotic arrest deficient 1 |
| Mad2 | mitotic arrest deficient 2 |
| MCC | mitotic checkpoint complex |
| MIS12 | MIS12 kinetochore complex component |
| Mps1 | monopolar spindle 1 kinase |
| MTOC | microtubule-organising centre |
| NDC80 | NDC80 kinetochore complex component |
| NEBD | nuclear envelope breakdown |
| o-Mad2 | open Mad2 |
| PLCζ | phospholipase C zeta |
| Rec 8 | meiotic recombination protein Rec 8 |
| SAC | spindle assembly checkpoint |
| Scc1 | sister chromatid cohesion protein 1 |
| TRIP13 | thyroid hormone receptor interactor 13 |
References
- Gelens, L.; Qian, J.; Bollen, M.; Saurin, A.T. The Importance of Kinase-Phosphatase Integration: Lessons from Mitosis. Trends Cell Biol. 2018, 28, 6–21. [Google Scholar] [CrossRef]
- Basu, S.; Greenwood, J.; Jones, A.W.; Nurse, P. Core control principles of the eukaryotic cell cycle. Nature 2022, 607, 381–386. [Google Scholar] [CrossRef]
- Nasa, I.; Kettenbach, A.N. Coordination of Protein Kinase and Phosphoprotein Phosphatase Activities in Mitosis. Front. Cell Dev. Biol. 2018, 6, 30. [Google Scholar] [CrossRef] [PubMed]
- Walczak, C.E.; Heald, R. Mechanisms of mitotic spindle assembly and function. Int. Rev. Cytol. 2008, 265, 111–158. [Google Scholar] [PubMed]
- Reber, S.; Hyman, A.A. Emergent Properties of the Metaphase Spindle. Cold Spring Harb. Perspect. Biol. 2015, 7, a015784. [Google Scholar] [CrossRef] [PubMed]
- Agircan, F.G.; Schiebel, E.; Mardin, B.R. Separate to operate: Control of centrosome positioning and separation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2014, 369, 20130461. [Google Scholar] [CrossRef] [PubMed]
- Prosser, S.L.; Pelletier, L. Mitotic spindle assembly in animal cells: A fine balancing act. Nat. Rev. Mol. Cell Biol. 2017, 18, 187–201. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, I. Centrosomes in mitotic spindle assembly and orientation. Curr. Opin. Struct. Biol. 2021, 66, 193–198. [Google Scholar] [CrossRef] [PubMed]
- Dunkley, S.; Scheffler, K.; Mogessie, B. Cytoskeletal form and function in mammalian oocytes and zygotes. Curr. Opin. Cell Biol. 2022, 75, 102073. [Google Scholar] [CrossRef]
- Mullen, T.J.; Davis-Roca, A.C.; Wignall, S.M. Spindle assembly and chromosome dynamics during oocyte meiosis. Curr. Opin. Cell Biol. 2019, 60, 53–59. [Google Scholar] [CrossRef]
- Blengini, C.S.; Schindler, K. Acentriolar spindle assembly in mammalian female meiosis and the consequences of its perturbations on human reproduction†. Biol. Reprod. 2022, 106, 253–263. [Google Scholar] [CrossRef]
- Mihajlovic, A.I.; FitzHarris, G. Segregating Chromosomes in the Mammalian Oocyte. Curr. Biol. 2018, 28, R895–R907. [Google Scholar] [CrossRef]
- Mogessie, B.; Scheffler, K.; Schuh, M. Assembly and Positioning of the Oocyte Meiotic Spindle. Annu. Rev. Cell Dev. Biol. 2018, 34, 381–403. [Google Scholar] [CrossRef] [PubMed]
- Dumont, J.; Million, K.; Sunderland, K.; Rassinier, P.; Lim, H.; Leader, B.; Verlhac, M.H. Formin-2 is required for spindle migration and for the late steps of cytokinesis in mouse oocytes. Dev. Biol. 2007, 301, 254–265. [Google Scholar] [CrossRef] [PubMed]
- Azoury, J.; Lee, K.W.; Georget, V.; Rassinier, P.; Leader, B.; Verlhac, M.H. Spindle positioning in mouse oocytes relies on a dynamic meshwork of actin filaments. Curr. Biol. 2008, 18, 1514–1519. [Google Scholar] [CrossRef] [PubMed]
- Schuh, M.; Ellenberg, J. A new model for asymmetric spindle positioning in mouse oocytes. Curr. Biol. 2008, 18, 1986–1992. [Google Scholar] [CrossRef] [PubMed]
- Roeles, J.; Tsiavaliaris, G. Actin-microtubule interplay coordinates spindle assembly in human oocytes. Nat. Commun. 2019, 10, 4651. [Google Scholar] [CrossRef] [PubMed]
- Harasimov, K.; Uraji, J.; Mönnich, E.U.; Holubcová, Z.; Elder, K.; Blayney, M.; Schuh, M. Actin-driven chromosome clustering facilitates fast and complete chromosome capture in mammalian oocytes. Nat. Cell Biol. 2023, 25, 439–452. [Google Scholar] [CrossRef] [PubMed]
- Mogessie, B.; Schuh, M. Actin protects mammalian eggs against chromosome segregation errors. Science 2017, 357, eaal1647. [Google Scholar] [CrossRef] [PubMed]
- Almonacid, M.; Terret, M.É.; Verlhac, M.H. Actin-based spindle positioning: New insights from female gametes. J. Cell Sci. 2014, 127, 477–483. [Google Scholar] [CrossRef]
- Manandhar, G.; Sutovsky, P.; Joshi, H.C.; Stearns, T.; Schatten, G. Centrosome reduction during mouse spermiogenesis. Dev. Biol. 1998, 203, 424–434. [Google Scholar] [CrossRef] [PubMed]
- Khanal, S.; Jaiswal, A.; Chowdanayaka, R.; Puente, N.; Turner, K.; Assefa, K.Y.; Nawras, M.; Back, E.D.; Royfman, A.; Burkett, J.P.; et al. The evolution of centriole degradation in mouse sperm. Nat. Commun. 2024, 15, 117. [Google Scholar] [CrossRef] [PubMed]
- Gueth-Hallonet, C.; Antony, C.; Aghion, J.; Santa-Maria, A.; Lajoie-Mazenc, I.; Wright, M.; Maro, B. gamma-Tubulin is present in acentriolar MTOCs during early mouse development. J. Cell Sci. 1993, 105, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Courtois, A.; Schuh, M.; Ellenberg, J.; Hiiragi, T. The transition from meiotic to mitotic spindle assembly is gradual during early mammalian development. J. Cell Biol. 2012, 198, 357–370. [Google Scholar] [CrossRef] [PubMed]
- Clift, D.; Schuh, M. Restarting life: Fertilization and the transition from meiosis to mitosis. Nat. Rev. Mol. Cell Biol. 2013, 14, 549–562. [Google Scholar] [CrossRef] [PubMed]
- Avidor-Reiss, T.; Mazur, M.; Fishman, E.L.; Sindhwani, P. The Role of Sperm Centrioles in Human Reproduction—The Known and the Unknown. Front. Cell Dev. Biol. 2019, 7, 188. [Google Scholar] [CrossRef] [PubMed]
- Aljiboury, A.; Hehnly, H. The centrosome—Diverse functions in fertilization and development across species. J. Cell Sci. 2023, 136, jcs261387. [Google Scholar] [CrossRef] [PubMed]
- Amargant, F.; Pujol, A.; Ferrer-Vaquer, A.; Durban, M.; Martínez, M.; Vassena, R.; Vernos, I. The human sperm basal body is a complex centrosome important for embryo preimplantation development. Mol. Hum. Reprod. 2021, 27, gaab062. [Google Scholar] [CrossRef]
- Uzbekov, R.; Singina, G.N.; Shedova, E.N.; Banliat, C.; Avidor-Reiss, T.; Uzbekova, S. Centrosome Formation in the Bovine Early Embryo. Cells 2023, 12, 1335. [Google Scholar] [CrossRef]
- Kops, G.J.; Weaver, B.A.; Cleveland, D.W. On the road to cancer: Aneuploidy and the mitotic checkpoint. Nat. Rev. Cancer 2005, 5, 773–785. [Google Scholar] [CrossRef]
- Lakhani, A.A.; Thompson, S.L.; Sheltzer, J.M. Aneuploidy in human cancer: New tools and perspectives. Trends Genet. 2023, 39, 968–980. [Google Scholar] [CrossRef] [PubMed]
- Mazzagatti, A.; Engel, J.L.; Ly, P. Boveri and beyond: Chromothripsis and genomic instability from mitotic errors. Mol. Cell 2023, 84, 55–69. [Google Scholar] [CrossRef] [PubMed]
- Musacchio, A. The Molecular Biology of Spindle Assembly Checkpoint Signaling Dynamics. Curr. Biol. 2015, 25, R1002–R1018. [Google Scholar] [CrossRef] [PubMed]
- McAinsh, A.D.; Kops GJ, P.L. Principles and dynamics of spindle assembly checkpoint signalling. Nat. Rev. Mol. Cell Biol. 2023, 24, 543–559. [Google Scholar] [CrossRef] [PubMed]
- Jackman, M.; Marcozzi, C.; Barbiero, M.; Pardo, M.; Yu, L.; Tyson, A.L.; Choudhary, J.S.; Pines, J. Cyclin B1-Cdk1 facilitates MAD1 release from the nuclear pore to ensure a robust spindle checkpoint. J. Cell Biol. 2020, 219, e201907082. [Google Scholar] [CrossRef]
- Sivakumar, S.; Gorbsky, G.J. Spatiotemporal regulation of the anaphase-promoting complex in mitosis. Nat. Rev. Mol. Cell Biol. 2015, 16, 82–94. [Google Scholar] [CrossRef]
- Watson, E.R.; Brown, N.G.; Peters, J.M.; Stark, H.; Schulman, B.A. Posing the APC/C E3 Ubiquitin Ligase to Orchestrate Cell Division. Trends Cell Biol. 2019, 29, 117–134. [Google Scholar] [CrossRef]
- Hoyt, M.A.; Totis, L.; Roberts, B.T. S. cerevisiae genes required for cell cycle arrest in response to loss of microtubule function. Cell 1991, 66, 507–517. [Google Scholar] [CrossRef]
- Li, R.; Murray, A.W. Feedback control of mitosis in budding yeast. Cell 1991, 66, 519–531. [Google Scholar] [CrossRef] [PubMed]
- Foley, E.A.; Kapoor, T.M. Microtubule attachment and spindle assembly checkpoint signalling at the kinetochore. Nat. Rev. Mol. Cell Biol. 2013, 14, 25–37. [Google Scholar] [CrossRef] [PubMed]
- Lara-Gonzalez, P.; Pines, J.; Desai, A. Spindle assembly checkpoint activation and silencing at kinetochores. Semin. Cell Dev. Biol. 2021, 117, 86–98. [Google Scholar] [CrossRef]
- Santaguida, S.; Tighe, A.; D’Alise, A.M.; Taylor, S.S.; Musacchio, A. Dissecting the role of MPS1 in chromosome biorientation and the spindle checkpoint through the small molecule inhibitor reversine. J. Cell Biol. 2010, 190, 73–87. [Google Scholar] [CrossRef]
- Yamagishi, Y.; Yang, C.H.; Tanno, Y.; Watanabe, Y. MPS1/Mph1 phosphorylates the kinetochore protein KNL1/Spc7 to recruit SAC components. Nat. Cell Biol. 2012, 14, 746–752. [Google Scholar] [CrossRef]
- Raisch, T.; Ciossani, G.; d’Amico, E.; Cmentowski, V.; Carmignani, S.; Maffini, S.; Merino, F.; Wohlgemuth, S.; Vetter, I.R.; Raunser, S.; et al. Structure of the RZZ complex and molecular basis of Spindly-driven corona assembly at human kinetochores. EMBO J. 2022, 41, e110411. [Google Scholar] [CrossRef]
- Sudakin, V.; Chan, G.K.; Yen, T.J. Checkpoint inhibition of the APC/C in HeLa cells is mediated by a complex of BUBR1, BUB3, CDC20, and MAD2. J Cell Biol 2001, 154, 925–936. [Google Scholar] [CrossRef] [PubMed]
- Fischer, E.S.; Yu CW, H.; Hevler, J.F.; McLaughlin, S.H.; Maslen, S.L.; Heck AJ, R.; Freund SM, V.; Barford, D. Juxtaposition of Bub1 and Cdc20 on phosphorylated Mad1 during catalytic mitotic checkpoint complex assembly. Nat. Commun. 2022, 13, 6381. [Google Scholar] [CrossRef] [PubMed]
- Varma, D.; Salmon, E.D. The KMN protein network--chief conductors of the kinetochore orchestra. J. Cell Sci. 2012, 125, 5927–5936. [Google Scholar] [CrossRef] [PubMed]
- Mapelli, M.; Massimiliano, L.; Santaguida, S.; Musacchio, A. The Mad2 conformational dimer: Structure and implications for the spindle assembly checkpoint. Cell 2007, 131, 730–743. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Li, B.; Tomchick, D.R.; Machius, M.; Rizo, J.; Yu, H.; Luo, X. p31comet blocks Mad2 activation through structural mimicry. Cell 2007, 131, 744–755. [Google Scholar] [CrossRef]
- Luo, X.; Fang, G.; Coldiron, M.; Lin, Y.; Yu, H.; Kirschner, M.W.; Wagner, G. Structure of the Mad2 spindle assembly checkpoint protein and its interaction with Cdc20. Nat. Struct. Biol. 2000, 7, 224–229. [Google Scholar] [PubMed]
- Izawa, D.; Pines, J. The mitotic checkpoint complex binds a second CDC20 to inhibit active APC/C. Nature 2015, 517, 631–634. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, J.; Dumont, S. Mammalian kinetochores count attached microtubules in a sensitive and switch-like manner. J. Cell Biol. 2019, 218, 3583–3596. [Google Scholar] [CrossRef] [PubMed]
- Collin, P.; Nashchekina, O.; Walker, R.; Pines, J. The spindle assembly checkpoint works like a rheostat rather than a toggle switch. Nat. Cell Biol. 2013, 15, 1378–1385. [Google Scholar] [CrossRef] [PubMed]
- Dick, A.E.; Gerlich, D.W. Kinetic framework of spindle assembly checkpoint signalling. Nat. Cell Biol. 2013, 15, 1370–1377. [Google Scholar] [CrossRef] [PubMed]
- Alfieri, C.; Chang, L.; Barford, D. Mechanism for remodelling of the cell cycle checkpoint protein MAD2 by the ATPase TRIP13. Nature 2018, 559, 274–278. [Google Scholar] [CrossRef] [PubMed]
- Kelly, A.E.; Funabiki, H. Correcting aberrant kinetochore microtubule attachments: An Aurora B-centric view. Curr. Opin. Cell Biol. 2009, 21, 51–58. [Google Scholar] [CrossRef]
- Cimini, D.; Howell, B.; Maddox, P.; Khodjakov, A.; Degrassi, F.; Salmon, E.D. Merotelic kinetochore orientation is a major mechanism of aneuploidy in mitotic mammalian tissue cells. J. Cell Biol. 2001, 153, 517–528. [Google Scholar] [CrossRef]
- Gregan, J.; Polakova, S.; Zhang, L.; Tolić-Nørrelykke, I.M.; Cimini, D. Merotelic kinetochore attachment: Causes and effects. Trends Cell Biol. 2011, 21, 374–381. [Google Scholar] [CrossRef]
- Petronczki, M.; Siomos, M.F.; Nasmyth, K. Un ménage à quatre: The molecular biology of chromosome segregation in meiosis. Cell 2003, 112, 423–440. [Google Scholar] [CrossRef]
- Kitajima, T.S. Mechanisms of kinetochore-microtubule attachment errors in mammalian oocytes. Dev. Growth Differ. 2018, 60, 33–43. [Google Scholar] [CrossRef]
- Kouznetsova, A.; Lister, L.; Nordenskjöld, M.; Herbert, M.; Höög, C. Bi-orientation of achiasmatic chromosomes in meiosis I oocytes contributes to aneuploidy in mice. Nat. Genet. 2007, 39, 966–968. [Google Scholar] [CrossRef] [PubMed]
- Sakakibara, Y.; Hashimoto, S.; Nakaoka, Y.; Kouznetsova, A.; Höög, C.; Kitajima, T.S. Bivalent separation into univalents precedes age-related meiosis I errors in oocytes. Nat. Commun. 2015, 6, 7550. [Google Scholar] [CrossRef] [PubMed]
- Lampson, M.A.; Cheeseman, I.M. Sensing centromere tension: Aurora B and the regulation of kinetochore function. Trends Cell Biol. 2011, 21, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Carmena, M.; Wheelock, M.; Funabiki, H.; Earnshaw, W.C. The chromosomal passenger complex (CPC): From easy rider to the godfather of mitosis. Nat. Rev. Mol. Cell Biol. 2012, 13, 789–803. [Google Scholar] [CrossRef] [PubMed]
- Cimini, D.; Cameron, L.A.; Salmon, E.D. Anaphase spindle mechanics prevent mis-segregation of merotelically oriented chromosomes. Curr. Biol. 2004, 14, 2149–2155. [Google Scholar] [CrossRef]
- Kamenz, J.; Hauf, S. Slow checkpoint activation kinetics as a safety device in anaphase. Curr. Biol. 2014, 24, 646–651. [Google Scholar] [CrossRef]
- Maiato, H.; Silva, S. Double-checking chromosome segregation. J. Cell Biol. 2023, 222, e202301106. [Google Scholar] [CrossRef]
- Bhakta, H.H.; Refai, F.H.; Avella, M.A. The molecular mechanisms mediating mammalian fertilization. Development 2019, 146, dev176966. [Google Scholar] [CrossRef]
- Gaspa-Toneu, L.; Peters, A.H. Nucleosomes in mammalian sperm: Conveying paternal epigenetic inheritance or subject to reprogramming between generations. Curr. Opin. Genet. Dev. 2023, 79, 102034. [Google Scholar] [CrossRef]
- Saunders, C.M.; Larman, M.G.; Parrington, J.; Cox, L.J.; Royse, J.; Blayney, L.M.; Swann, K.; Lai, F.A. PLC zeta: A sperm-specific trigger of Ca(2+) oscillations in eggs and embryo development. Development 2002, 129, 3533–3544. [Google Scholar] [CrossRef] [PubMed]
- Reichmann, J.; Nijmeijer, B.; Hossain, M.J.; Eguren, M.; Schneider, I.; Politi, A.Z.; Roberti, M.J.; Hufnagel, L.; Hiiragi, T.; Ellenberg, J. Dual-spindle formation in zygotes keeps parental genomes apart in early mammalian embryos. Science 2018, 361, 189–193. [Google Scholar] [CrossRef] [PubMed]
- Ciemerych, M.A.; Maro, B.; Kubiak, J.Z. Control of duration of the first two mitoses in a mouse embryo. Zygote 1999, 7, 293–300. [Google Scholar] [CrossRef]
- Farrell, J.A.; O’Farrell, P.H. From egg to gastrula: How the cell cycle is remodeled during the Drosophila mid-blastula transition. Annu. Rev. Genet. 2014, 48, 269–294. [Google Scholar] [CrossRef] [PubMed]
- Newport, J.; Kirschner, M. A major developmental transition in early Xenopus embryos: I. characterization and timing of cellular changes at the midblastula stage. Cell 1982, 30, 675–686. [Google Scholar] [PubMed]
- Bao, Z.; Zhao, Z.; Boyle, T.J.; Murray, J.I.; Waterston, R.H. Control of cell cycle timing during C. elegans embryogenesis. Dev. Biol. 2008, 318, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Fulka, J. Checkpoint control of the G2/M phase transition during the first mitotic cycle in mammalian eggs. Human. Reprod. 1999, 14, 1582–1587. [Google Scholar] [CrossRef][Green Version]
- Ciemerych, M.A.; Sicinski, P. Cell cycle in mouse development. Oncogene 2005, 24, 2877–2898. [Google Scholar] [CrossRef]
- Schulz, K.N.; Harrison, M.M. Mechanisms regulating zygotic genome activation. Nat. Rev. Genet. 2018, 20, 221–234. [Google Scholar] [CrossRef]
- Anger, M.; Radonova, L.; Horakova, A.; Sekach, D.; Charousova, M. Impact of Global Transcriptional Silencing on Cell Cycle Regulation and Chromosome Segregation in Early Mammalian Embryos. Int. J. Mol. Sci. 2021, 22, 9073. [Google Scholar] [CrossRef]
- Bennabi, I.; Terret, M.E.; Verlhac, M.H. Meiotic spindle assembly and chromosome segregation in oocytes. J. Cell Biol. 2016, 215, 611–619. [Google Scholar] [CrossRef]
- Kovacovicova, K.; Awadova, T.; Mikel, P.; Anger, M. In Vitro Maturation of Mouse Oocytes Increases the Level of Kif11/Eg5 on Meiosis II Spindles. Biol. Reprod. 2016, 95, 18. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Diez, C.; FitzHarris, G. Causes and consequences of chromosome segregation error in preimplantation embryos. Reproduction 2018, 155, R63–R76. [Google Scholar] [CrossRef] [PubMed]
- Nagaoka, S.I.; Hassold, T.J.; Hunt, P.A. Human aneuploidy: Mechanisms and new insights into an age-old problem. Nat. Rev. Genet. 2012, 13, 493–504. [Google Scholar] [CrossRef] [PubMed]
- Mihajlovic, A.I.; Byers, C.; Reinholdt, L.; FitzHarris, G. Spindle assembly checkpoint insensitivity allows meiosis-II despite chromosomal defects in aged eggs. EMBO Rep. 2023, 24, e57227. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Mihajlović, A.I.; Yang, G.; FitzHarris, G. Kinetochore deterioration concommitant with centromere weakening during aging in mouse oocyte meiosis-I. FASEB J. 2023, 37, e22922. [Google Scholar] [CrossRef] [PubMed]
- Charalambous, C.; Webster, A.; Schuh, M. Aneuploidy in mammalian oocytes and the impact of maternal ageing. Nat. Rev. Mol. Cell Biol. 2022, 24, 27–44. [Google Scholar] [CrossRef]
- Wartosch, L.; Schindler, K.; Schuh, M.; Gruhn, J.R.; Hoffmann, E.R.; McCoy, R.C.; Xing, J. Origins and mechanisms leading to aneuploidy in human eggs. Prenat. Diagn. 2021, 41, 620–630. [Google Scholar] [CrossRef]
- Mihajlovic, A.I.; Haverfield, J.; FitzHarris, G. Distinct classes of lagging chromosome underpin age-related oocyte aneuploidy in mouse. Dev. Cell 2021, 56, 2273–2283.e3. [Google Scholar] [CrossRef]
- Hassold, T.; Hunt, P. To err (meiotically) is human: The genesis of human aneuploidy. Nat. Rev. Genet. 2001, 2, 280–291. [Google Scholar] [CrossRef]
- Shahbazi, M.N.; Wang, T.; Tao, X.; Weatherbee BA, T.; Sun, L.; Zhan, Y.; Keller, L.; Smith, G.D.; Pellicer, A.; Scott, R.T.; et al. Developmental potential of aneuploid human embryos cultured beyond implantation. Nat. Commun. 2020, 11, 3987. [Google Scholar] [CrossRef]
- Torres, E.M. Consequences of gaining an extra chromosome. Chromosome Res. 2023, 31, 24. [Google Scholar] [CrossRef]
- Krivega, M.; Stiefel, C.M.; Storchova, Z. Consequences of chromosome gain: A new view on trisomy syndromes. Am. J. Hum. Genet. 2022, 109, 2126–2140. [Google Scholar] [CrossRef]
- Tšuiko, O.; Catteeuw, M.; Zamani Esteki, M.; Destouni, A.; Bogado Pascottini, O.; Besenfelder, U.; Havlicek, V.; Smits, K.; Kurg, A.; Salumets, A.; et al. Genome stability of bovine in vivo-conceived cleavage-stage embryos is higher compared to in vitro-produced embryos. Hum. Reprod. 2017, 32, 2348–2357. [Google Scholar] [CrossRef]
- Pauerova, T.; Radonova, L.; Kovacovicova, K.; Novakova, L.; Skultety, M.; Anger, M. Aneuploidy during the onset of mouse embryo development. Reproduction 2020, 160, 773–782. [Google Scholar] [CrossRef]
- Duncan, F.E.; Chiang, T.; Schultz, R.M.; Lampson, M.A. Evidence that a defective spindle assembly checkpoint is not the primary cause of maternal age-associated aneuploidy in mouse eggs. Biol. Reprod. 2009, 81, 768–776. [Google Scholar] [CrossRef]
- Danylevska, A.; Kovacovicova, K.; Awadova, T.; Anger, M. The frequency of precocious segregation of sister chromatids in mouse female meiosis I is affected by genetic background. Chromosome Res. 2014, 22, 365–373. [Google Scholar] [CrossRef]
- Carbone, L.; Chavez, S.L. Mammalian pre-implantation chromosomal instability: Species comparison, evolutionary considerations, and pathological correlations. Syst. Biol. Reprod. Med. 2015, 61, 321–335. [Google Scholar] [CrossRef] [PubMed]
- Destouni, A.; Zamani Esteki, M.; Catteeuw, M.; Tšuiko, O.; Dimitriadou, E.; Smits, K.; Kurg, A.; Salumets, A.; Van Soom, A.; Voet, T.; et al. Zygotes segregate entire parental genomes in distinct blastomere lineages causing cleavage-stage chimerism and mixoploidy. Genome Res. 2016, 26, 567–578. [Google Scholar] [CrossRef] [PubMed]
- Hornak, M.; Oracova, E.; Hulinska, P.; Urbankova, L.; Rubes, J. Aneuploidy detection in pigs using comparative genomic hybridization: From the oocytes to blastocysts. PLoS ONE 2012, 7, e30335. [Google Scholar] [CrossRef] [PubMed]
- Daughtry, B.L.; Rosenkrantz, J.L.; Lazar, N.H.; Fei, S.S.; Redmayne, N.; Torkenczy, K.A.; Adey, A.; Yan, M.; Gao, L.; Park, B.; et al. Single-cell sequencing of primate preimplantation embryos reveals chromosome elimination via cellular fragmentation and blastomere exclusion. Genome Res. 2019, 29, 367–382. [Google Scholar] [CrossRef] [PubMed]
- Bolton, H.; Graham, S.J.; Van der Aa, N.; Kumar, P.; Theunis, K.; Fernandez Gallardo, E.; Voet, T.; Zernicka-Goetz, M. Mouse model of chromosome mosaicism reveals lineage-specific depletion of aneuploid cells and normal developmental potential. Nat. Commun. 2016, 7, 11165. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Rito, T.; Metzger, J.; Naftaly, J.; Soman, R.; Hu, J.; Albertini, D.F.; Barad, D.H.; Brivanlou, A.H.; Gleicher, N. Depletion of aneuploid cells in human embryos and gastruloids. Nat. Cell Biol. 2021, 23, 314–321. [Google Scholar] [CrossRef]
- Capalbo, A.; Poli, M.; Jalas, C.; Forman, E.J.; Treff, N.R. On the reproductive capabilities of aneuploid human preimplantation embryos. Am. J. Hum. Genet. 2022, 109, 1572–1581. [Google Scholar] [CrossRef]
- Vera-Rodriguez, M.; Chavez, S.L.; Rubio, C.; Reijo Pera, R.A.; Simon, C. Prediction model for aneuploidy in early human embryo development revealed by single-cell analysis. Nat. Commun. 2015, 6, 7601. [Google Scholar] [CrossRef]
- Eme, L.; Trilles, A.; Moreira, D.; Brochier-Armanet, C. The phylogenomic analysis of the anaphase promoting complex and its targets points to complex and modern-like control of the cell cycle in the last common ancestor of eukaryotes. BMC Evol. Biol. 2011, 11, 265. [Google Scholar] [CrossRef]
- Vleugel, M.; Hoogendoorn, E.; Snel, B.; Kops, G.J. Evolution and function of the mitotic checkpoint. Dev. Cell 2012, 23, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Kops GJ, P.L.; Snel, B.; Tromer, E.C. Evolutionary Dynamics of the Spindle Assembly Checkpoint in Eukaryotes. Curr. Biol. 2020, 30, R589–R602. [Google Scholar] [CrossRef] [PubMed]
- van Hooff, J.J.; Tromer, E.; van Wijk, L.M.; Snel, B.; Kops, G.J. Evolutionary dynamics of the kinetochore network in eukaryotes as revealed by comparative genomics. EMBO Rep. 2017, 18, 1559–1571. [Google Scholar] [CrossRef]
- Tromer, E.C.; van Hooff, J.J.; Kops, G.J.; Snel, B. Mosaic origin of the eukaryotic kinetochore. Proc. Natl. Acad. Sci. USA 2019, 116, 12873–12882. [Google Scholar] [CrossRef]
- Encalada, S.E.; Willis, J.; Lyczak, R.; Bowerman, B. A spindle checkpoint functions during mitosis in the early Caenorhabditis elegans embryo. Mol. Biol. Cell 2005, 16, 1056–1070. [Google Scholar] [CrossRef]
- Essex, A.; Dammermann, A.; Lewellyn, L.; Oegema, K.; Desai, A. Systematic analysis in Caenorhabditis elegans reveals that the spindle checkpoint is composed of two largely independent branches. Mol. Biol. Cell 2009, 20, 1252–1267. [Google Scholar] [CrossRef] [PubMed]
- Gerhold, A.R.; Ryan, J.; Vallée-Trudeau, J.N.; Dorn, J.F.; Labbé, J.C.; Maddox, P.S. Investigating the regulation of stem and progenitor cell mitotic progression by in situ imaging. Curr. Biol. 2015, 25, 1123–1134. [Google Scholar] [CrossRef] [PubMed]
- Gerhold, A.R.; Poupart, V.; Labbé, J.C.; Maddox, P.S. Spindle assembly checkpoint strength is linked to cell fate in the Caenorhabditis elegans embryo. Mol. Biol. Cell 2018, 29, 1435–1448. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Kothari, P.; Lampson, M.A. Spindle assembly checkpoint acquisition at the mid-blastula transition. PLoS ONE 2015, 10, e0119285. [Google Scholar] [CrossRef] [PubMed]
- Shao, H.; Li, R.; Ma, C.; Chen, E.; Liu, X.J. Xenopus oocyte meiosis lacks spindle assembly checkpoint control. J. Cell Biol. 2013, 201, 191–200. [Google Scholar] [CrossRef]
- Clute, P.; Masui, Y. Regulation of the appearance of division asynchrony and microtubule-dependent chromosome cycles in Xenopus laevis embryos. Dev. Biol. 1995, 171, 273–285. [Google Scholar] [CrossRef][Green Version]
- Minshull, J.; Sun, H.; Tonks, N.K.; Murray, A.W. A MAP kinase-dependent spindle assembly checkpoint in Xenopus egg extracts. Cell 1994, 79, 475–486. [Google Scholar] [CrossRef]
- Roca, M.; Besnardeau, L.; Christians, E.; McDougall, A.; Chenevert, J.; Castagnetti, S. Acquisition of the spindle assembly checkpoint and its modulation by cell fate and cell size in a chordate embryo. Development 2023, 150, dev201145. [Google Scholar] [CrossRef]
- Chenevert, J.; Roca, M.; Besnardeau, L.; Ruggiero, A.; Nabi, D.; McDougall, A.; Copley, R.R.; Christians, E.; Castagnetti, S. The Spindle Assembly Checkpoint Functions during Early Development in Non-Chordate Embryos. Cells 2020, 9, 1087. [Google Scholar] [CrossRef] [PubMed]
- Paps, J.; Rossi, M.E.; Bowles, A.; Álvarez-Presas, M. Assembling animals: Trees, genomes, cells, and contrast to plants. Front. Ecol. Evol. 2023, 11, 1185566. [Google Scholar] [CrossRef]
- McGuinness, B.E.; Anger, M.; Kouznetsova, A.; Gil-Bernabé, A.M.; Helmhart, W.; Kudo, N.R.; Wuensche, A.; Taylor, S.; Hoog, C.; Novak, B.; et al. Regulation of APC/C activity in oocytes by a Bub1-dependent spindle assembly checkpoint. Curr. Biol. 2009, 19, 369–380. [Google Scholar] [CrossRef]
- Touati, S.A.; Buffin, E.; Cladière, D.; Hached, K.; Rachez, C.; van Deursen, J.M.; Wassmann, K. Mouse oocytes depend on BubR1 for proper chromosome segregation but not for prophase I arrest. Nat. Commun. 2015, 6, 6946. [Google Scholar] [CrossRef]
- El Yakoubi, W.; Buffin, E.; Cladière, D.; Gryaznova, Y.; Berenguer, I.; Touati, S.A.; Gómez, R.; Suja, J.A.; van Deursen, J.M.; Wassmann, K. Mps1 kinase-dependent Sgo2 centromere localisation mediates cohesin protection in mouse oocyte meiosis I. Nat. Commun. 2017, 8, 694. [Google Scholar] [CrossRef]
- Nagaoka, S.I.; Hodges, C.A.; Albertini, D.F.; Hunt, P.A. Oocyte-specific differences in cell-cycle control create an innate susceptibility to meiotic errors. Curr. Biol. 2011, 21, 651–657. [Google Scholar] [CrossRef]
- Sebestova, J.; Danylevska, A.; Novakova, L.; Kubelka, M.; Anger, M. Lack of response to unaligned chromosomes in mammalian female gametes. Cell Cycle 2012, 11, 3011–3018. [Google Scholar] [CrossRef]
- Kolano, A.; Brunet, S.; Silk, A.D.; Cleveland, D.W.; Verlhac, M.H. Error-prone mammalian female meiosis from silencing the spindle assembly checkpoint without normal interkinetochore tension. Proc. Natl. Acad. Sci. USA 2012, 109, E1858–E1867. [Google Scholar] [CrossRef]
- Lane, S.I.; Yun, Y.; Jones, K.T. Timing of anaphase-promoting complex activation in mouse oocytes is predicted by microtubule-kinetochore attachment but not by bivalent alignment or tension. Development 2012, 139, 1947–1955. [Google Scholar] [CrossRef] [PubMed]
- Gui, L.; Homer, H. Spindle assembly checkpoint signalling is uncoupled from chromosomal position in mouse oocytes. Development 2012, 139, 1941–1946. [Google Scholar] [CrossRef] [PubMed]
- Holubcová, Z.; Blayney, M.; Elder, K.; Schuh, M. Human oocytes. Error-prone chromosome-mediated spindle assembly favors chromosome segregation defects in human oocytes. Science 2015, 348, 1143–1147. [Google Scholar] [CrossRef] [PubMed]
- Haverfield, J.; Dean, N.L.; Nöel, D.; Rémillard-Labrosse, G.; Paradis, V.; Kadoch, I.J.; FitzHarris, G. Tri-directional anaphases as a novel chromosome segregation defect in human oocytes. Hum. Reprod. 2017, 32, 1293–1303. [Google Scholar] [CrossRef] [PubMed]
- Macaulay, A.D.; Allais, A.; FitzHarris, G. Chromosome dynamics and spindle microtubule establishment in mouse embryos. FASEB J. 2020, 34, 8057–8067. [Google Scholar] [CrossRef] [PubMed]
- Allais, A.; FitzHarris, G. Absence of a robust mitotic timer mechanism in early preimplantation mouse embryos leads to chromosome instability. Development 2022, 149, dev200391. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Diez, C.; Yamagata, K.; Trivedi, S.; Haverfield, J.; FitzHarris, G. Micronucleus formation causes perpetual unilateral chromosome inheritance in mouse embryos. Proc. Natl. Acad. Sci. USA 2016, 113, 626–631. [Google Scholar] [CrossRef] [PubMed]
- Kort, D.H.; Chia, G.; Treff, N.R.; Tanaka, A.J.; Xing, T.; Vensand, L.B.; Micucci, S.; Prosser, R.; Lobo, R.A.; Sauer, M.V.; et al. Human embryos commonly form abnormal nuclei during development: A mechanism of DNA damage, embryonic aneuploidy, and developmental arrest. Hum. Reprod. 2016, 31, 312–323. [Google Scholar] [CrossRef] [PubMed]
- Yao, T.; Ueda, A.; Khurchabilig, A.; Mashiko, D.; Tokoro, M.; Nagai, H.; Sho, T.; Matoba, S.; Yamagata, K.; Sugimura, S. Micronucleus formation during early cleavage division is a potential hallmark of preimplantation embryonic loss in cattle. Biochem. Biophys. Res. Commun. 2022, 617, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, K.; Van de Velde, H.; De Paepe, C.; Sermon, K.; Spits, C. Mitotic spindle disruption in human preimplantation embryos activates the spindle assembly checkpoint but not apoptosis until Day 5 of development. Mol. Hum. Reprod. 2017, 23, 321–329. [Google Scholar] [CrossRef]
- Wei, Y.; Multi, S.; Yang, C.R.; Ma, J.; Zhang, Q.H.; Wang, Z.B.; Li, M.; Wei, L.; Ge, Z.J.; Zhang, C.H.; et al. Spindle assembly checkpoint regulates mitotic cell cycle progression during preimplantation embryo development. PLoS ONE 2011, 6, e21557. [Google Scholar] [CrossRef]
- Radonova, L.; Svobodova, T.; Skultety, M.; Mrkva, O.; Libichova, L.; Stein, P.; Anger, M. ProTAME Arrest in Mammalian Oocytes and Embryos Does Not Require Spindle Assembly Checkpoint Activity. Int. J. Mol. Sci. 2019, 20, 4537. [Google Scholar] [CrossRef]
- Vázquez-Diez, C.; Paim LM, G.; FitzHarris, G. Cell-Size-Independent Spindle Checkpoint Failure Underlies Chromosome Segregation Error in Mouse Embryos. Curr. Biol. 2019, 29, 865–873.e3. [Google Scholar] [CrossRef]
- Dobles, M.; Liberal, V.; Scott, M.L.; Benezra, R.; Sorger, P.K. Chromosome missegregation and apoptosis in mice lacking the mitotic checkpoint protein Mad2. Cell 2000, 101, 635–645. [Google Scholar] [CrossRef]
- Wang, Q.; Liu, T.; Fang, Y.; Xie, S.; Huang, X.; Mahmood, R.; Ramaswamy, G.; Sakamoto, K.M.; Darzynkiewicz, Z.; Xu, M.; et al. BUBR1 deficiency results in abnormal megakaryopoiesis. Blood 2004, 103, 1278–1285. [Google Scholar] [CrossRef] [PubMed]
- Iwanaga, Y.; Chi, Y.H.; Miyazato, A.; Sheleg, S.; Haller, K.; Peloponese, J.M.; Li, Y.; Ward, J.M.; Benezra, R.; Jeang, K.T. Heterozygous deletion of mitotic arrest-deficient protein 1 (MAD1) increases the incidence of tumors in mice. Cancer Res. 2007, 67, 160–166. [Google Scholar] [CrossRef] [PubMed]
- Jeganathan, K.; Malureanu, L.; Baker, D.J.; Abraham, S.C.; van Deursen, J.M. Bub1 mediates cell death in response to chromosome missegregation and acts to suppress spontaneous tumorigenesis. J. Cell Biol. 2007, 179, 255–267. [Google Scholar] [CrossRef] [PubMed]
- Tilston, V.; Taylor, S.S.; Perera, D. Inactivating the spindle checkpoint kinase Bub1 during embryonic development results in a global shutdown of proliferation. BMC Res. Notes 2009, 2, 190. [Google Scholar] [CrossRef] [PubMed]
- Galli, M.; Morgan, D.O. Cell Size Determines the Strength of the Spindle Assembly Checkpoint during Embryonic Development. Dev. Cell 2016, 36, 344–352. [Google Scholar] [CrossRef]
- Kyogoku, H.; Kitajima, T.S. Large Cytoplasm Is Linked to the Error-Prone Nature of Oocytes. Dev. Cell 2017, 41, 287–298.e4. [Google Scholar] [CrossRef]
- Lane SI, R.; Jones, K.T. Chromosome biorientation and APC activity remain uncoupled in oocytes with reduced volume. J. Cell Biol. 2017, 216, 3949–3957. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Horakova, A.; Konecna, M.; Anger, M. Chromosome Division in Early Embryos—Is Everything under Control? And Is the Cell Size Important? Int. J. Mol. Sci. 2024, 25, 2101. https://doi.org/10.3390/ijms25042101
Horakova A, Konecna M, Anger M. Chromosome Division in Early Embryos—Is Everything under Control? And Is the Cell Size Important? International Journal of Molecular Sciences. 2024; 25(4):2101. https://doi.org/10.3390/ijms25042101
Chicago/Turabian StyleHorakova, Adela, Marketa Konecna, and Martin Anger. 2024. "Chromosome Division in Early Embryos—Is Everything under Control? And Is the Cell Size Important?" International Journal of Molecular Sciences 25, no. 4: 2101. https://doi.org/10.3390/ijms25042101
APA StyleHorakova, A., Konecna, M., & Anger, M. (2024). Chromosome Division in Early Embryos—Is Everything under Control? And Is the Cell Size Important? International Journal of Molecular Sciences, 25(4), 2101. https://doi.org/10.3390/ijms25042101




