De Novo Assembly of the Genome of the Sea Urchin Paracentrotus lividus (Lamarck 1816)
Abstract
1. Introduction
2. Results and Discussion
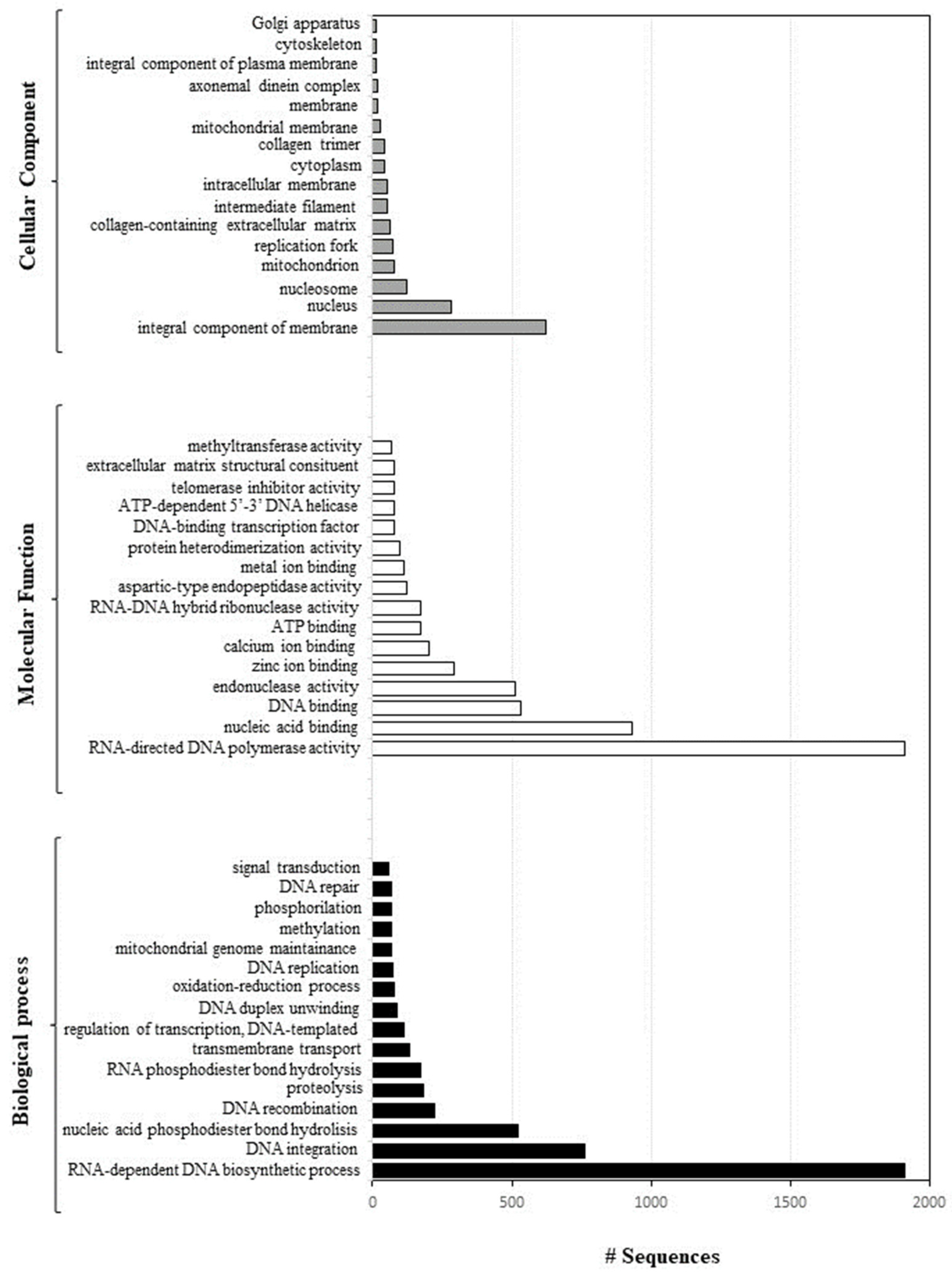
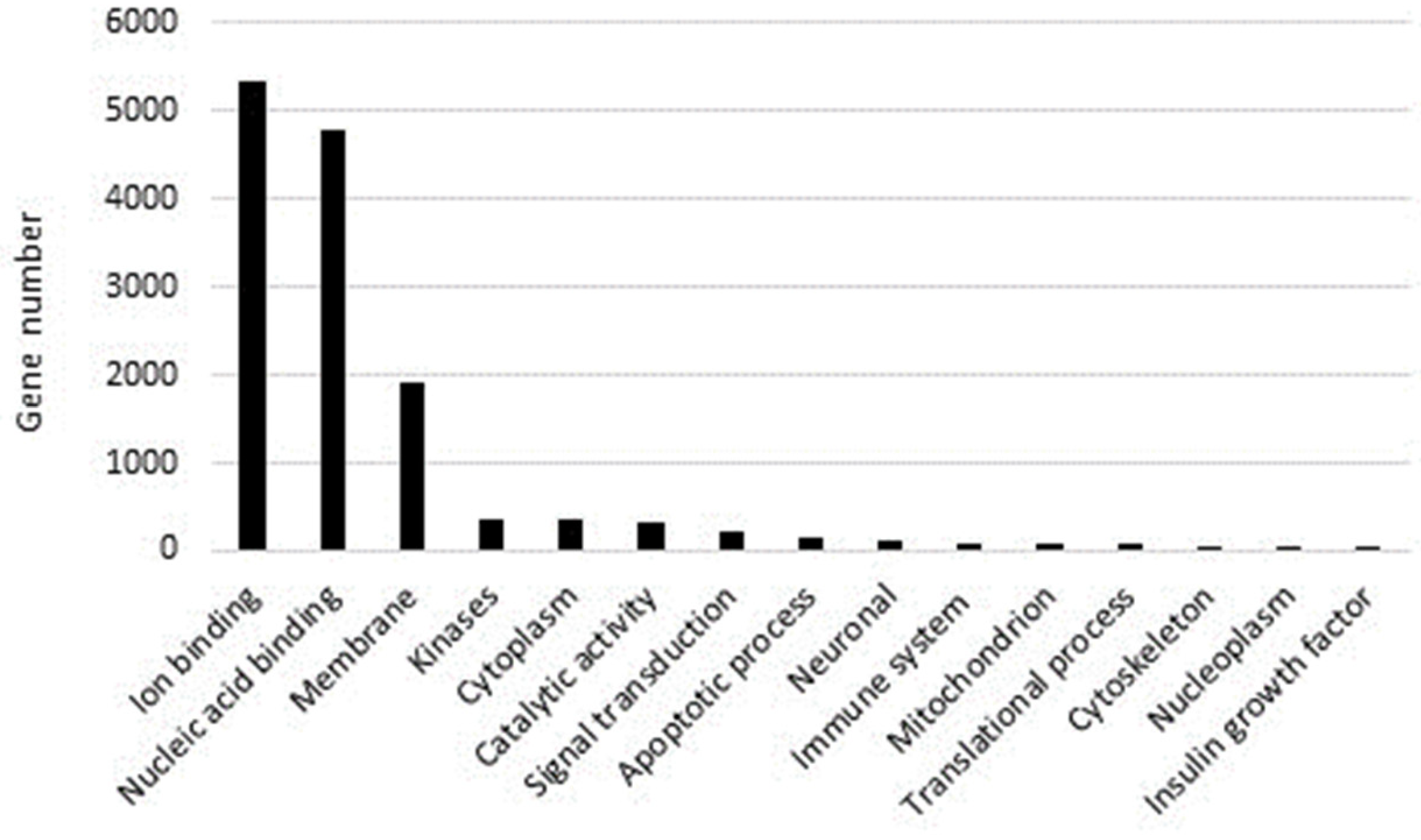
2.1. Sequencing and Annotation of Paracentrotus Lividus Genome
2.2. Key Findings and Genes
2.2.1. Complex Innate Immune Responses
2.2.2. Molecular Switches in Signal Transduction
2.2.3. Genes Regulating the Membrane Receptors
2.2.4. Nervous System and Neuronal Genes
2.2.5. The Kinome of P. lividus Resembles That of Drosophila and Human
2.2.6. Homologies with Human Oxidative Metabolism
3. Materials and Methods
3.1. Ethics Statement
3.2. Sample Collection and DNA Extraction
3.3. De Novo Genome Assembly
- Genome sequencing: the next generation sequencing experiment and bioinformatics analysis were performed using Genomix4life S.R.L. (Baronissi, Salerno, Italy). DNA concentration was assayed with a ND-1000 spectrophotometer (NanoDrop, ND-1000 UV-Vis Spectrophotometer; NanoDrop Technologies, Wilmington, DE, USA), and its quality assessed with an Agilent 4200 Tapestation (Agilent Technologies, Santa Clara, CA, USA; according manufacturer instructions). An indexed library was prepared from 1 µg of purified DNA with a Truseq DNA Nano Library Prep Kit according to the manufacturer’s instructions (Illumina, San Diego, CA, USA). The library was quantified using the Tape Station 4200 (Agilent Technologies, Santa Clara, CA, USA) and a Qubit fluorometer (Invitrogen Co., Carlsbad, CA, USA), and diluted with a final concentration of 2 nM. The sample was subject to cluster generation and sequencing using an Illumina NextSeq 500 System (Illumina) in a 2 × 150 paired-end format, according NextSeq 500 System Documentation.
- Sequencing outputs, quality control and cleaning: the most common metric was used to assess the accuracy of a sequencing platform (base calling accuracy, measured by the Phred quality score (Q score). The first step was a quality check of the raw Illumina sequencing data to remove adapter sequences and low-quality reads, using ad hoc script. The FastQC tool (available on http://www.bioinformatics.babraham.ac.uk/projects/fastqc; 1 February 2021) was used to check the quality of raw data sequencing.
- Genome assembly: to perform the de novo assembly, a KmerGenie (version 1.7044) tool was necessary to estimate the best k-mer length 66. In this case, the best k-mer predicted was 121. ABySS 2.0, an implementation of ABySS 1.0, was used to perform the de novo assembly on fastq files. The bloom filter of ABySS 2.0 was applied to avoid duplicate sequences.
- Genome assembly stats and validation: the integrity assembly was also evaluated, using several statistical tools, such as QUAST, Abyss, BBMAP, and BUSCO (Table S3).
- Genome annotation and functional analysis: Geneious software 69 was used to identify all the ORF sequences, and Blast2GO was applied to perform a blast alignment of all ORF sequences identified and to annotate everything in the Gene Ontology database.
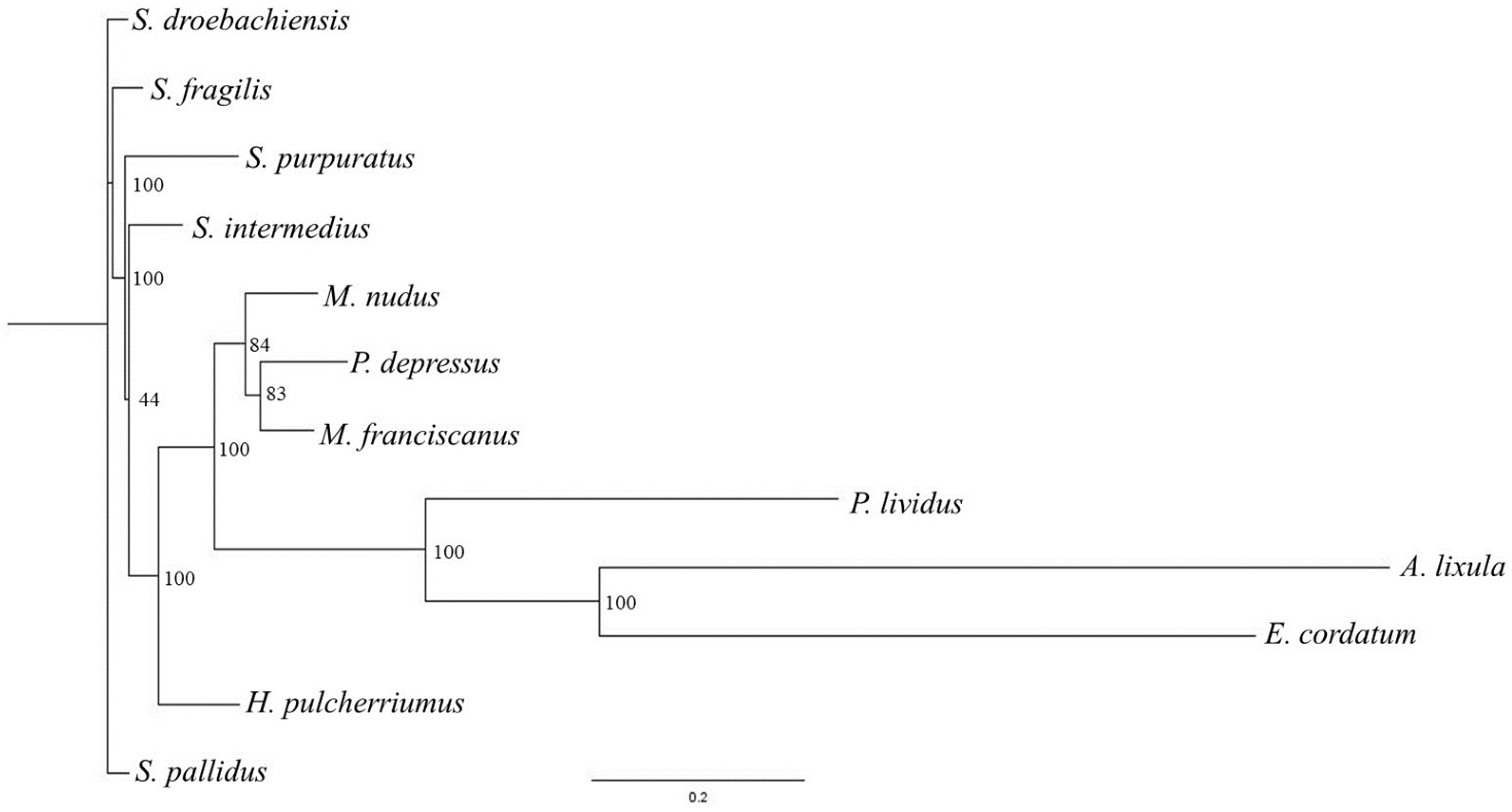
3.4. Phylogenetic Tree
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Werner, T. Next generation sequencing in functional genomics. Brief. Bioinform. 2010, 2, 499–511. [Google Scholar] [CrossRef]
- Kulski, J.K. Next-Generation Sequencing—An Overview of the History, Tools, and “Omic” Applications; Spinger: Berlin/Heidelberg, Germany, 2016; Volume 10, p. 61964. ISBN 0000957720. [Google Scholar]
- Giani, A.M.; Gallo, G.R.; Gianfranceschi, L.; Formenti, G. Long walk to genomics: History and current approaches to genome sequencing and assembly. Comput. Struct. Biotechnol. J. 2020, 18, 9–19. [Google Scholar] [CrossRef]
- Hurd, P.J.; Nelson, C.J. Advantages of next-generation sequencing versus the microarray in epigenetic research. Brief. Funct. Genom. Proteom. 2009, 8, 174–183. [Google Scholar] [CrossRef] [PubMed]
- Ekblom, R.; Galindo, J. Applications of next generation sequencing in molecular ecology of non-model organisms. Heredity 2011, 107, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Dupont, S.; Ortega-Martínez, O.; Thorndyke, M. Impact of near-future ocean acidification on echinoderms. Ecotoxicology 2010, 19, 449–462. [Google Scholar] [CrossRef] [PubMed]
- Sodergren, E.; Weinstock, G.M.; Davidson, E.H.; Cameron, R.A.; Gibbs, R.A.; Angerer, R.C.; Angerer, L.M.; Arnone, M.I.; Burgess, D.R.; Burke, R.D.; et al. The genome of the sea urchin Strongylocentrotus purpuratus. Science 2006, 314, 941–952. [Google Scholar] [CrossRef] [PubMed]
- Pawson, D.L. Narcissia ahearnae, a new species of sea star from the Western Atlantic (Echinodermata: Asteroidea: Valvatida). Zootaxa 2007, 58, 53–58. [Google Scholar] [CrossRef]
- Boudouresque, C.F.; Bernard, G.; Bonhomme, P.; Charbonnel, E.; Le Diréach, L.; Ruitton, S. Monitoring methods for Posidonia oceanica seagrass meadows in Provence and the French Riviera. Sci. Rep. 2007, 22, 17–38. [Google Scholar]
- Guidetti, P.; Terlizzi, A.; Boero, F. Effects of the edible sea urchin, Paracentrotus lividus, fishery along the Apulian rocky coast (SE Italy, Mediterranean Sea). Fish. Res. 2004, 66, 287–297. [Google Scholar] [CrossRef]
- Furesi, R.; Madau, F.; Palomba, A.; Pulina, P. Stated preferences for consumption of sea urchin: A choice experiment in Sardinia (Italy). Int. J. Food Syst. Dyn. 2014, 5, 111–119. [Google Scholar]
- Boudouresque, C.F.; Verlaque, M. Ecology of Paracentrotus lividus. In Edible Sea Urchins: Biology and Ecology; Developments in Aquaculture and Fisheries Science; Elsevier: Amsterdam, The Netherlands, 2001; pp. 177–216. [Google Scholar]
- Kober, K.M.; Bernardi, G. Phylogenomics of strongylocentrotid sea urchins. BMC Evol. Biol. 2013, 13, 88. [Google Scholar] [CrossRef]
- Marlétaz, F.; Couloux, A.; Poulain, J.; Labadie, K.; Da Silva, C.; Mangenot, S.; Noel, B.; Poustka, A.J.; Dru, P.; Pegueroles, C.; et al. Analysis of the P. lividus sea urchin genome highlights contrasting trends of genomic and regulatory evolution in deuterostomes. Cell Genom. 2023, 3, 100295. [Google Scholar] [CrossRef]
- Materna, S.C.; Berney, K.; Cameron, R.A. The S. purpuratus genome: A comparative perspective. Dev. Biol. 2006, 300, 485–495. [Google Scholar] [CrossRef]
- Cameron, R.A.; Davidson, E.H. A basal deuterostome genome viewed as a natural experiment. Gene 2007, 406, 1–7. [Google Scholar] [CrossRef]
- Lawrence, J.M. Edible sea urchins: Use and life-history strategies. In Developments in Aquaculture and Fisheries Science; Elsevier: Amsterdam, The Netherlands, 2007; pp. 1–9. [Google Scholar]
- Beane, W.S.; Gross, J.M.; McClay, D.R. RhoA regulates initiation of invagination, but not convergent extension, during sea urchin gastrulation. Dev. Biol. 2006, 292, 213–225. [Google Scholar] [CrossRef] [PubMed]
- Dehal, P.; Boore, J.L. Two rounds of whole genome duplication in the ancestral vertebrate. PLoS Biol. 2005, 3, e314. [Google Scholar] [CrossRef] [PubMed]
- Gu, X.; Wang, Y.; Gu, J. Age distribution of human gene families shows significant roles of both large- and small-scale duplications in vertebrate evolution. Nat. Genet. 2002, 31, 205–209. [Google Scholar] [CrossRef]
- Lee, T.H.; Naitoh, H.; Yamazaki, F. Chromosome studies on the mitten crabs Eriocheir japonica and E. sinensis. Fish. Sci. 2004, 70, 211–214. [Google Scholar] [CrossRef]
- Jackman, S.D.; Vandervalk, B.P.; Mohamadi, H.; Chu, J.; Yeo, S.; Hammond, S.A.; Jahesh, G.; Khan, H.; Coombe, L.; Warren, R.L.; et al. ABySS 2.0: Resource-efficient assembly of large genomes using a bloom filter effect of bloom filter false positive rate. Genome Res. 2017, 27, 768–777. [Google Scholar] [CrossRef] [PubMed]
- Fink, I.R.; Pietretti, D.; Voogdt, C.G.P.; Westphal, A.H.; Savelkoul, H.F.J.; Forlenza, M.; Wiegertjes, G.F. Molecular and functional characterization of Toll-like receptor (Tlr)1 and Tlr2 in common carp (Cyprinus carpio). Fish Shellfish Immunol. 2016, 56, 70–83. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, T.; Kawai, T. Toll-like receptor signaling pathways. Front. Immunol. 2014, 5, 461. [Google Scholar] [CrossRef] [PubMed]
- Pinsino, A.; Alijagic, A. Sea urchin Paracentrotus lividus immune cells in culture: Formulation of the appropriate harvesting and culture media and maintenance conditions. Biol. Open 2019, 8, bio039289. [Google Scholar] [CrossRef] [PubMed]
- Chiaramonte, M.; Russo, R. The echinoderm innate humoral immune response. Ital. J. Zool. 2015, 82, 300–308. [Google Scholar] [CrossRef]
- Smith, L.C.; Arriza, V.; Barela Hudgell, M.A.; Barone, G.; Bodnar, A.G.; Buckley, K.M.; Cunsolo, V.; Dheilly, N.M.; Franchi, N.; Fugmann, S.D.; et al. Echinodermata: The complex immune system in echinoderms. In Advances in Comparative Immunology; Springer: Berlin/Heidelberg, Germany, 2018; pp. 773–791. ISBN 9783319767680. [Google Scholar]
- Hibino, T.; Loza-Coll, M.; Messier, C.; Majeske, A.J.; Cohen, A.H.; Terwilliger, D.P.; Buckley, K.M.; Brockton, V.; Nair, S.V.; Berney, K.; et al. The immune gene repertoire encoded in the purple sea urchin genome. Dev. Biol. 2006, 300, 349–365. [Google Scholar] [CrossRef] [PubMed]
- Materna, S.C.; Cameron, R.A. The sea urchin genome as a window on function. Biol. Bull. 2008, 214, 266–273. [Google Scholar] [CrossRef] [PubMed]
- Chiaramonte, M.; Arizza, V.; Russo, R. Evolutionary conserved pathway of the innate immune response after a viral insult in Paracentrotus lividus sea urchin. Int. J. Immunogenet. 2019, 46, 192–202. [Google Scholar] [CrossRef]
- Adonin, L.; Drozdov, A.; Barlev, N.A. Sea urchin as a universal model for studies of gene networks. Front. Genet. 2021, 11, 1739. [Google Scholar] [CrossRef] [PubMed]
- White, D.; De Lamirande, E.; Gagnon, C. Protein kinase C is an important signaling mediator associated with motility of intact sea urchin spermatozoa. J. Exp. Biol. 2007, 210, 4053–4064. [Google Scholar] [CrossRef]
- Felix, R.C.; Trindade, M.; Pires, I.R.P.; Fonseca, V.G.; Martins, R.S.; Silveira, H.; Power, D.M.; Cardoso, J.C.R. Unravelling the evolution of the allatostatin-type A, KISS and galanin peptide-receptor gene families in bilaterians: Insights from Anopheles mosquitoes. PLoS ONE 2015, 10, e0130347. [Google Scholar] [CrossRef]
- Mirabeau, O.; Jean-Stéphane, J. Molecular evolution of peptidergic signaling systems in bilaterians. Proc. Natl. Acad. Sci. USA 2013, 110, E2028–E2037. [Google Scholar] [CrossRef]
- Lopez-Ortiz, C.; Peña-Garcia, Y.; Natarajan, P.; Bhandari, M.; Abburi, V.; Dutta, S.K.; Yadav, L.; Stommel, J.; Nimmakayala, P.; Reddy, U.K. The ankyrin repeat gene family in Capsicum spp.: Genome-wide survey, characterization and gene expression profile. Sci. Rep. 2020, 10, 4044. [Google Scholar] [CrossRef] [PubMed]
- Loza-Huerta, A.; Pacheco-Castillo, H.; Darszon, A.; Beltrán, C. Crosstalk between protein kinases A and C regulates sea urchin sperm motility. Zygote 2022, 30, 398–409. [Google Scholar] [CrossRef] [PubMed]
- Hanson, P.I.; Whiteheart, S.W. AAA+ proteins: Have engine will work. Nat. Rev. Mol. Cell Biol. 2005, 6, 519–529. [Google Scholar] [CrossRef] [PubMed]
- Erzberger, J.P.; Berger, J.M. Evolutionary relationships and structural mechanisms of AAA+ proteins. Annu. Rev. Biophys. Biomol. Struct. 2006, 35, 93–114. [Google Scholar] [CrossRef]
- Snider, J.; Thibault, G.; Houry, W.A. The AAA+ superfamily of functionally diverse proteins. Genome Biol. 2008, 9, 216. [Google Scholar] [CrossRef]
- Shorter, J.; Houry, W.A. Editorial: The role of AAA+ proteins in protein repair and degradation. Front. Mol. Biosci. 2018, 5, 85. [Google Scholar] [CrossRef]
- Galindo, B.E.; Moy, G.W.; Vacquier, V.D. A third sea urchin sperm receptor for egg jelly module protein, suREJ2, concentrates in the plasma membrane over the sperm mitochondrion. Dev. Growth Differ. 2004, 46, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Michaely, P.; Bennett, V. The ANK repeat: A ubiquitous motif involved in macromolecular recognition. Trends Cell Biol. 1992, 2, 127–129. [Google Scholar] [CrossRef]
- Breeden, L.; Nasmyth, K. Cell cycle control of the yeast HO gene: Cis- and Trans-acting regulators. Cell 1987, 48, 389–397. [Google Scholar] [CrossRef]
- Breeden, L.; Nasmyth, K. Similarity between cell-cycle genes of budding yeast and fission yeast and the Notch gene of Drosophila. Nature 1987, 329, 651–654. [Google Scholar] [CrossRef]
- Sedgwick, S.G.; Smerdon, S.J. The ankyrin repeat: A diversity of interactions on a common structural framework. Trends Biochem. Sci. 1999, 24, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Yaguchi, S.; Yaguchi, J.; Wei, Z.; Shiba, K.; Angerer, L.M.; Inaba, K. ankAT-1 is a novel gene mediating the apical tuft formation in the sea urchin embryo. Dev. Biol. 2010, 348, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Mosavi, L.K.; Cammett, T.J.; Desrosiers, D.C.; Peng, Z.Y. The ankyrin repeat as molecular architecture for protein recognition. Protein Sci. 2004, 13, 1435–1448. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Mahajan, A.; Tsai, M.D. Ankyrin repeat: A unique motif mediating protein-protein interactions. Biochemistry 2006, 45, 15168–15178. [Google Scholar] [CrossRef] [PubMed]
- Shen, G.; Kuppu, S.; Venkataramani, S.; Wang, J.; Yan, J.; Qiu, X.; Zhang, H. ANKYRIN REPEAT-CONTAINING PROTEIN 2A Is an essential molecular chaperone for peroxisomal membrane-bound ASCORBATE PEROXIDASE3 in Arabidopsis. Plant Cell 2010, 22, 811–831. [Google Scholar] [CrossRef] [PubMed]
- Collas, P.; Courvalin, J.C.; Poccia, D. Targeting of membranes to sea urchin sperm chromatin is mediated by a lamin B receptor-like integral membrane protein. J. Cell Biol. 1996, 135, 1715–1725. [Google Scholar] [CrossRef]
- Whittaker, C.A.; Bergeron, K.F.; Whittle, J.; Brandhorst, B.P.; Burke, R.D.; Hynes, R.O. The echinoderm adhesome. Dev. Biol. 2006, 300, 252–266. [Google Scholar] [CrossRef]
- Hardin, J. 4 The Cellular Basis of Sea Urchin Gastrulation. Curr. Top. Dev. Biol. 1996, 33, 159–262. [Google Scholar]
- Raible, F.; Tessmar-Raible, K.; Arboleda, E.; Kaller, T.; Bork, P.; Arendt, D.; Arnone, M.I. Opsins and clusters of sensory G-protein-coupled receptors in the sea urchin genome. Dev. Biol. 2006, 300, 461–475. [Google Scholar] [CrossRef]
- Xie, W.; Evans, R. Orphan nuclear receptors: The exotics of xenobiotics. J. Biol. Chem. 2001, 276, 37739–37742. [Google Scholar] [CrossRef]
- Klee, C.; Krinks, M. Purification of cyclic 3′5′-nucleotide phosphodiesterase inhibitory protein by affinity chromatography on activator protein coupled to sepharose. Biochemistry 1978, 17, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Wallace, R.; Lynch, T.; Tallant, E.; Cheung, W. Purification and characterization of an inhibitor protein of brain adenylate cyclase and cyclic nucleotide phosphodiesterase. J. Biol. Chem. 1979, 254, 377–382. [Google Scholar] [CrossRef] [PubMed]
- Iwasa, F.; Ishiguro, K. Calmodulin-binding protein (55K+17K) of sea urchin eggs has a Ca2+ and calmodulin-dependent phosphoprotein phosphatase activity. J. Biochem. 1986, 99, 1353–1358. [Google Scholar] [CrossRef]
- Wood, J.G.; Wallace, R.W.; Whitaker, J.N.; Cheung, W.Y. Immunocytochemical localization of calmodulin and a heat-labile calmodulin-binding protein (CaM-BP80) in basal ganglia of mouse brain. J. Cell Biol. 1980, 84, 66–76. [Google Scholar] [CrossRef] [PubMed]
- Stewart, A.; Ingebritsen, T.; Manalan, A.; Klee, C.; Cohen, P. Discovery of A Ca2+-and calmodulin-dependent protein phosphatase. FEBS Lett. 1982, 137, 80–84. [Google Scholar] [CrossRef] [PubMed]
- Washbourne, P.; Dityatev, A.; Scheiffele, P.; Biederer, T.; Weiner, J.; Christopherson, K.; El-Husseini, A. Cell adhesion molecules in synapse formation. J. Neurosci. 2004, 24, 9244–9249. [Google Scholar] [CrossRef] [PubMed]
- Lapraz, F.; Röttinger, E.; Duboc, V.; Range, R.; Duloquin, L.; Walton, K.; Wu, S.Y.; Bradham, C.; Loza, M.A.; Hibino, T.; et al. RTK and TGF-β signaling pathways genes in the sea urchin genome. Dev. Biol. 2006, 300, 132–152. [Google Scholar] [CrossRef] [PubMed]
- Sherwood, D.R.; Mcclay, D.R. Identification and localization of a sea urchin Notch homologue: Insights into vegetal plate regionalization and Notch receptor regulation. Development 1997, 124, 3363–3374. [Google Scholar] [CrossRef]
- Meyer, A.; Ku, C.; Hatleberg, W.; Telmer, C.A.; Hinman, V. New hypotheses of cell type diversity and novelty from comparative single cell and nuclei transcriptomics in echinoderms. bioRxiv 2022. [Google Scholar] [CrossRef]
- Bradham, C.A.; Foltz, K.R.; Beane, W.S.; Arnone, M.I.; Rizzo, F.; Coffman, J.A.; Mushegian, A.; Goel, M.; Morales, J.; Geneviere, A.M.; et al. The sea urchin kinome: A first look. Dev. Biol. 2006, 300, 180–193. [Google Scholar] [CrossRef]
- Rossignol, D.P.; Lennarz, W.J.; Waechter, C.J. Induction of phosphorylation of dolichol during embryonic development of the sea urchin. J. Biol. Chem. 1981, 256, 10538–10542. [Google Scholar] [CrossRef]
- Byrum, C.A.; Walton, K.D.; Robertson, A.J.; Carbonneau, S.; Thomason, R.T.; Coffman, J.A.; McClay, D.R. Protein tyrosine and serine-threonine phosphatases in the sea urchin, Strongylocentrotus purpuratus: Identification and potential functions. Dev. Biol. 2006, 300, 194–218. [Google Scholar] [CrossRef][Green Version]
- Ruocco, N.; Fedele, A.M.; Costantini, S.; Romano, G.; Ianora, A.; Costantini, M. New inter-correlated genes targeted by diatom-derived polyunsatured aldehydes in the sea urchin Paracentrotus lividus. Ecotoxicol. Environ. Saf. 2017, 142, 355–362. [Google Scholar] [CrossRef] [PubMed]
- Galtier, N.; Gouy, M.; Gautier, C. Seaview and phylo_ win: Two graphic tools for sequence alignment and molecular phylogeny. Bioinformatics 1996, 12, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 2006, 22, 2688–2690. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A. FigTree 1.4.3. Available online: https://github.com/rambaut/figtree/releases (accessed on 1 December 2023).

| Genera | Species |
|---|---|
| Strongylocentrotus | S. purpuratus |
| S. pallidus | |
| S. droebachiensis | |
| S. intermedius | |
| S. fragilis | |
| S. polyacanthus | |
| Mesocentrotus | M. franciscanus |
| M. nudus | |
| Hemicentrotus | H. pulcherrimus |
| Pseudocentrotus | P. depressus |
| Parameter | Quast | ABYSS | BUSCO |
|---|---|---|---|
| Assembly | scaffolds (min. Length = 500 bp) | scaffolds (min. Length = 500 bp) | |
| contigs (≥0 bp) | 252,952 | 252,952 | 252,952 |
| contigs (≥1000 bp) | 280 | _ | |
| contigs (≥5000 bp) | 5 | _ | |
| Total_length (≥0 bp) | 42,528,692 | _ | 42,528,692 |
| Total_length (≥1000 bp) | 515,753 | _ | |
| Total_length (≥5000 bp) | 28,242 | _ | |
| Total_length (≥10,000 bp) | 0 | _ | |
| contigs | 1757 | 1757 | |
| Largest_contig | 6806 | 6805 | |
| Total_length | 1,488,145 | 1,486,080 | |
| GC(%) | 34.77% | 31.88% | |
| N50 | 792 | 791 | 153 |
| Genes/Proteins | |
|---|---|
| Immune response | Toll-like receptor 1 |
| Toll-like receptor 2 | |
| Toll-like receptor 3 | |
| Toll-like receptor 4 | |
| Toll-like receptor 5 | |
| Toll-like receptor 6 | |
| Toll-like receptor 7 | |
| Toll-like receptor 8 | |
| Toll-like receptor 9 | |
| Toll-like receptor 10 | |
| Toll-like receptor 11 | |
| Toll-like receptor 12 | |
| Toll-like receptor 13 | |
| E3 ubiquitin-protein ligase pellino homolog 1 | |
| Signal transduction | Ras |
| Rab | |
| Ral | |
| Arf | |
| Rhodopsin | |
| 1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase beta-4 | |
| Nuclear factor kappa B | |
| Allatostatin-A receptor-like | |
| Calcium-independent protein kinase C | |
| Membrane receptors | suREJ1 |
| suREJ2 | |
| suREJ3 | |
| Ankyrin-containing gene specific for Apical Tuft | |
| Fibrillin A | |
| Rhodopsin | |
| Neuronal genes | Calcineurin |
| Neurexin | |
| Neurocan | |
| Neuroendocrine convertase 1 gene | |
| Neuron navigator 3-like | |
| Neuronal acetylcholine receptor subunit alpha-5-like | |
| Beta-adrenergic receptor kinase 2 | |
| Kinome | Adenosine kinase |
| A-kinase anchor protein 17A | |
| Bifunctional UDP-N-acetylglucosamine 2-epimerase/N-acetylmannosamine kinase isoform X1 | |
| Cell division cycle 7-related protein kinase | |
| Cyclin-dependent kinase 2-like | |
| Dolichol kinase | |
| Dual specificity mitogen-activated protein kinase kinase 7 isoform X2 | |
| Inositol hexakisphosphate | |
| Diphosphoinositol-pentakisphosphate kinase 1 isoform X1 | |
| L-fucose kinase | |
| MAP kinase | |
| Maternal embryonic leucine zipper kinase isoform X2 | |
| Membrane-associated guanylate kinase | |
| WW | |
| PDZ domain-containing protein 2-like | |
| Receptor tyrosine-protein kinase erbB-4-like | |
| Receptor-like protein kinase feronia, serine/threonine-protein kinase PAK 2 | |
| Tyrosine-protein kinase receptor Tie-1-like | |
| Wall-associated receptor kinase and wee1-like protein kinase 1-A | |
| Oxidative metabolism | CYP 1-like |
| CYP 2-like | |
| CYP 3-like | |
| CYP 4-like | |
| CYP 6-like | |
| CYP 20-like | |
| CYP 26-like | |
| CYP 27-like | |
| CYP 46-like | |
| CYP 51-like | |
| CYP 120-like |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Costantini, M.; Esposito, R.; Ruocco, N.; Caramiello, D.; Cordella, A.; Ventola, G.M.; Zupo, V. De Novo Assembly of the Genome of the Sea Urchin Paracentrotus lividus (Lamarck 1816). Int. J. Mol. Sci. 2024, 25, 1685. https://doi.org/10.3390/ijms25031685
Costantini M, Esposito R, Ruocco N, Caramiello D, Cordella A, Ventola GM, Zupo V. De Novo Assembly of the Genome of the Sea Urchin Paracentrotus lividus (Lamarck 1816). International Journal of Molecular Sciences. 2024; 25(3):1685. https://doi.org/10.3390/ijms25031685
Chicago/Turabian StyleCostantini, Maria, Roberta Esposito, Nadia Ruocco, Davide Caramiello, Angela Cordella, Giovanna Maria Ventola, and Valerio Zupo. 2024. "De Novo Assembly of the Genome of the Sea Urchin Paracentrotus lividus (Lamarck 1816)" International Journal of Molecular Sciences 25, no. 3: 1685. https://doi.org/10.3390/ijms25031685
APA StyleCostantini, M., Esposito, R., Ruocco, N., Caramiello, D., Cordella, A., Ventola, G. M., & Zupo, V. (2024). De Novo Assembly of the Genome of the Sea Urchin Paracentrotus lividus (Lamarck 1816). International Journal of Molecular Sciences, 25(3), 1685. https://doi.org/10.3390/ijms25031685









