Transcriptome Sequencing Unveils a Molecular-Stratification-Predicting Prognosis of Sarcoma Associated with Lipid Metabolism
Abstract
1. Introduction
2. Results
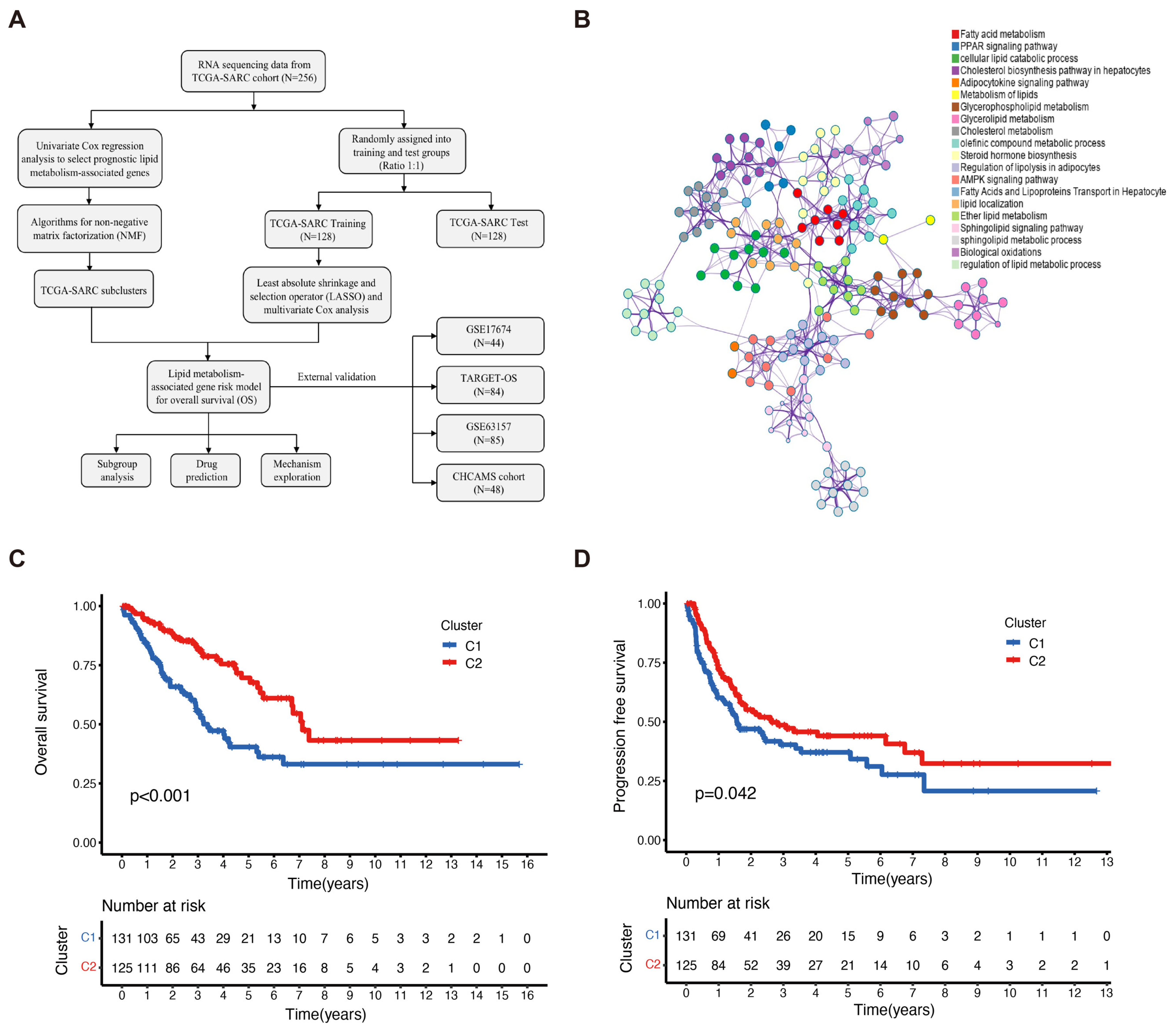
2.1. Identification of Subclusters Based on LMAGs in Sarcoma
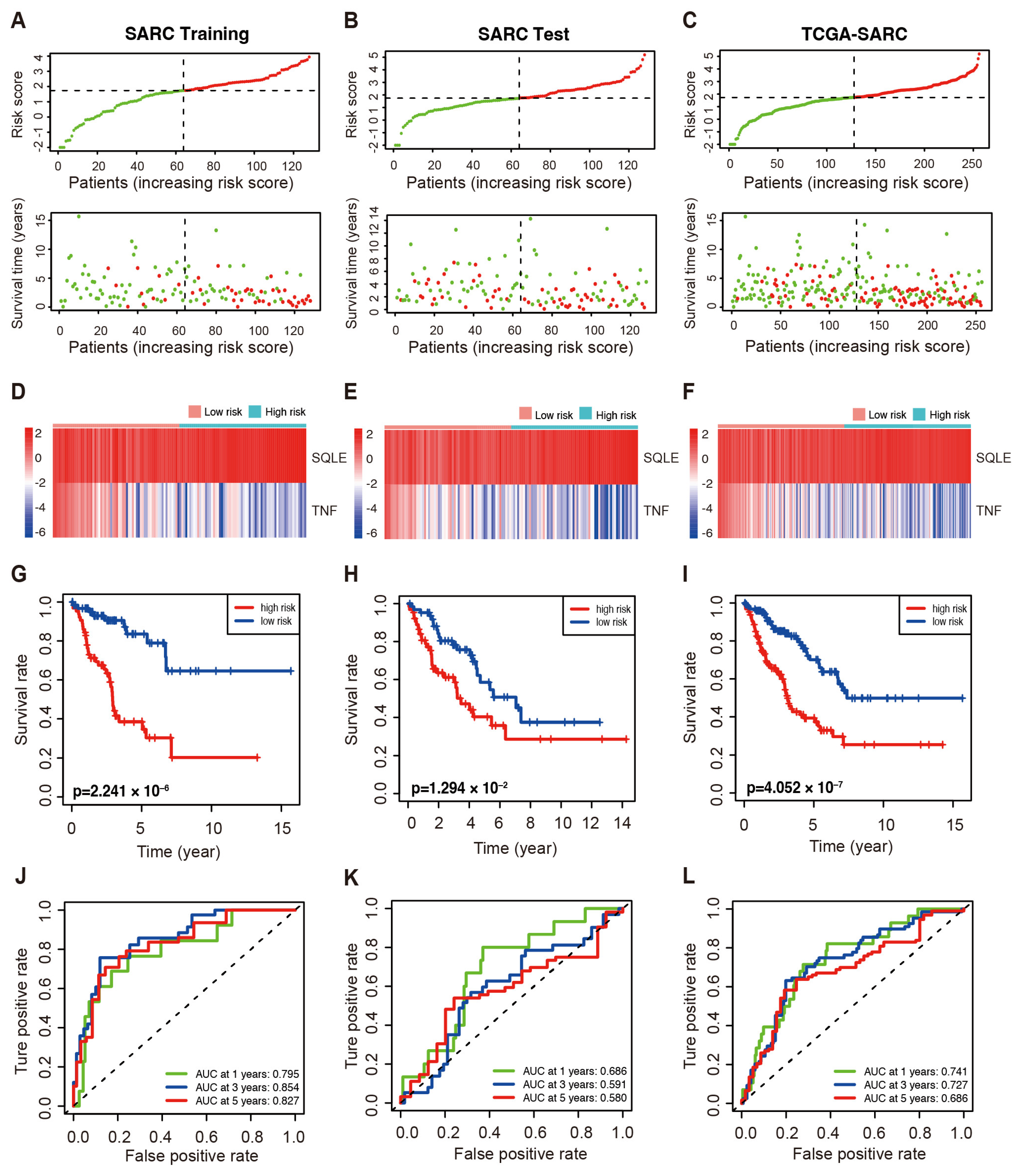
2.2. Prognostic Values of LMAGs in Sarcoma
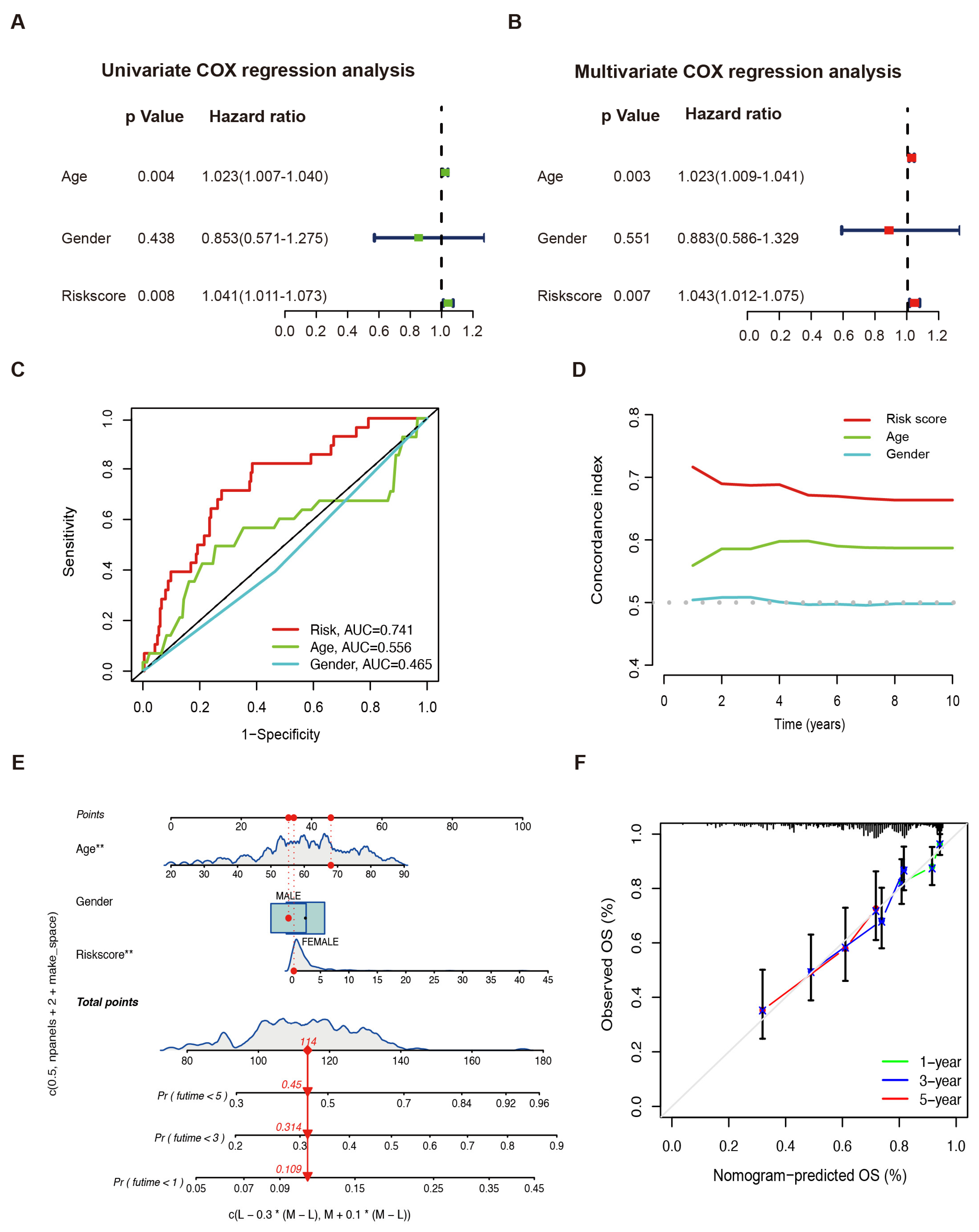
2.3. The LMAGs Risk Model as an Independent Prognostic Factor for Sarcoma
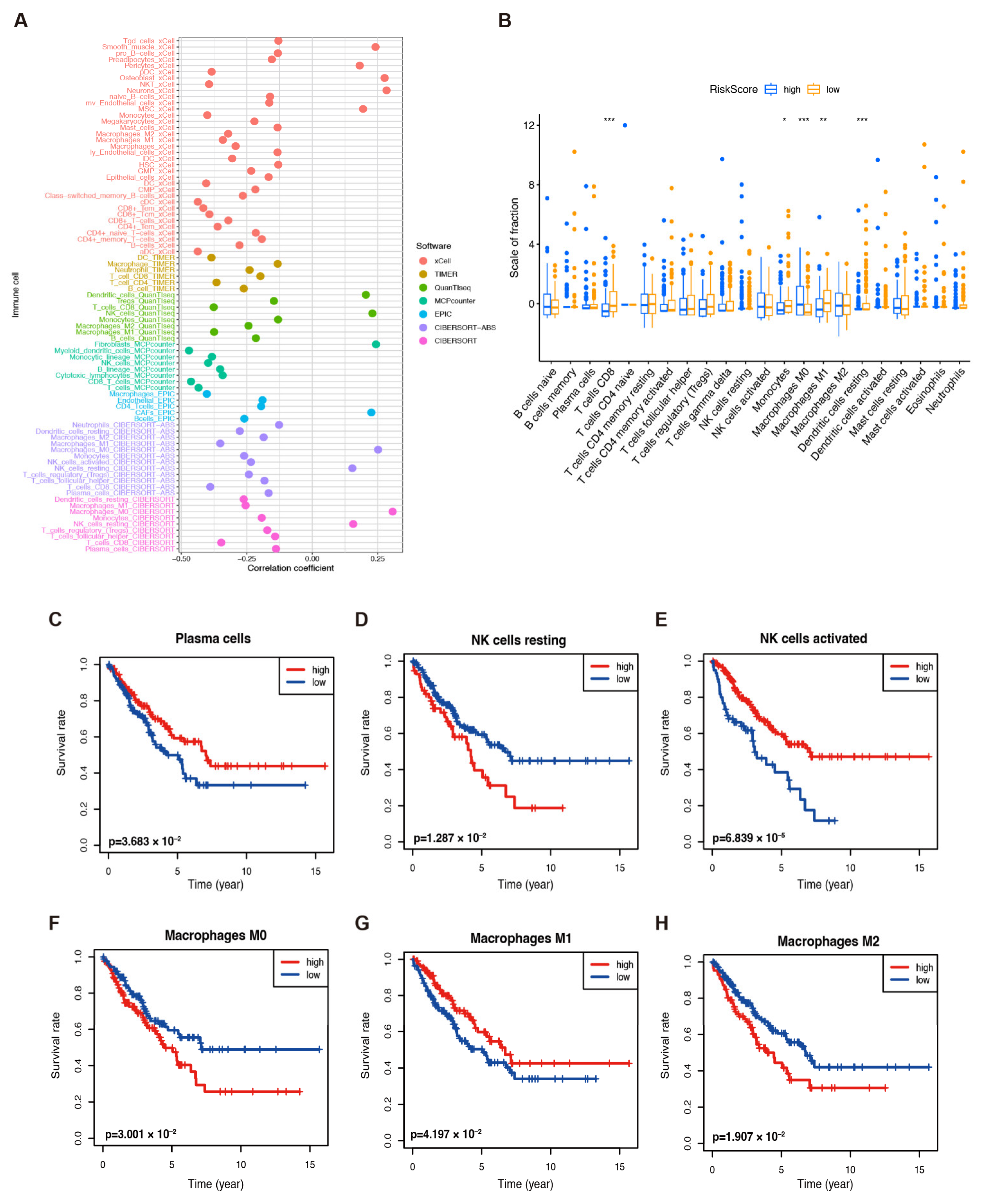
2.4. Immune Microenvironment and Infiltration in Sarcoma
2.5. Analysis of Drug Sensitivity
2.6. Validation of the LMAGs Risk Model Using the TARGET and GEO Datasets
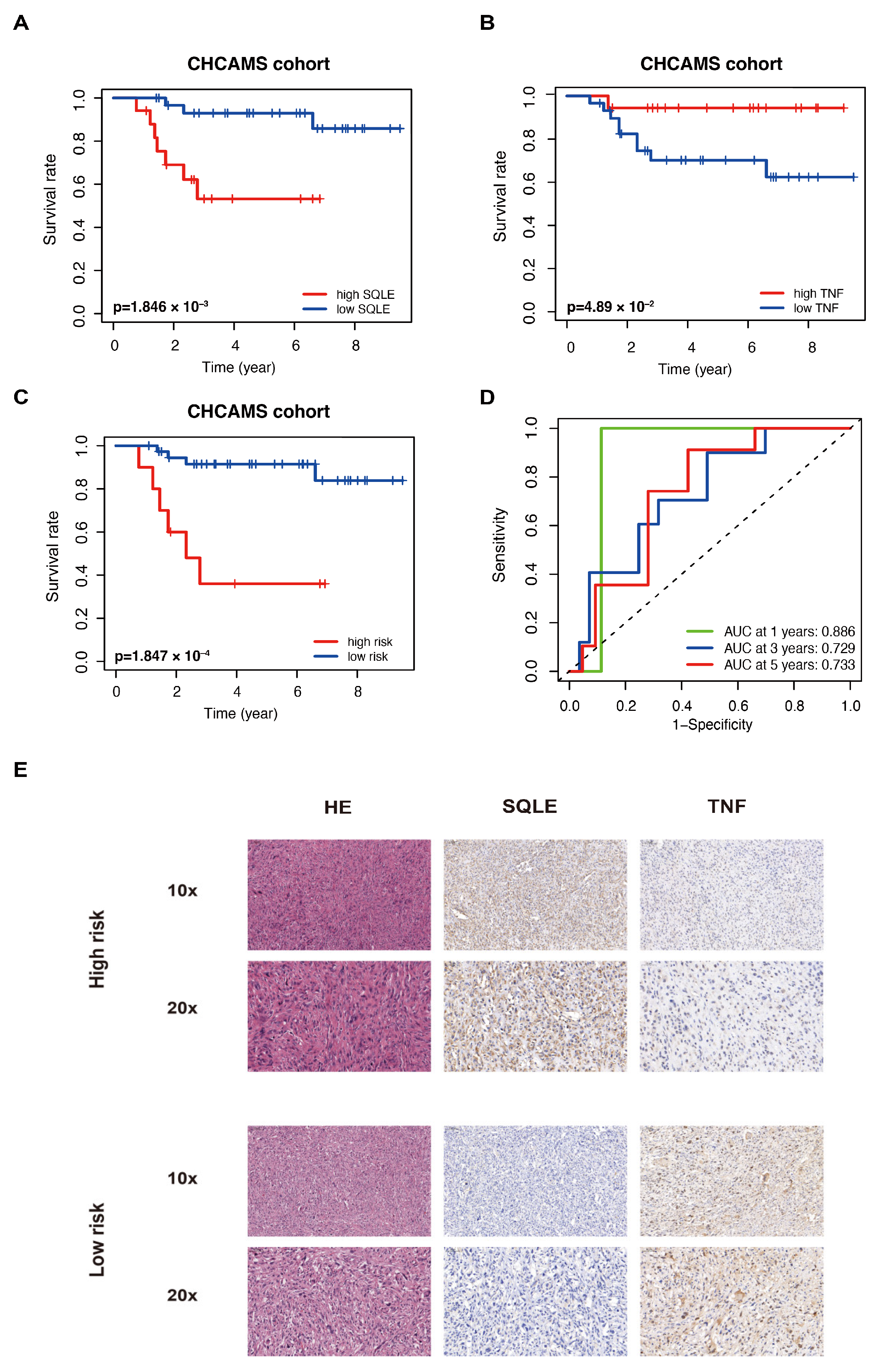
2.7. Validation of the LMAGs Risk Model by the CHCAMS Cohort
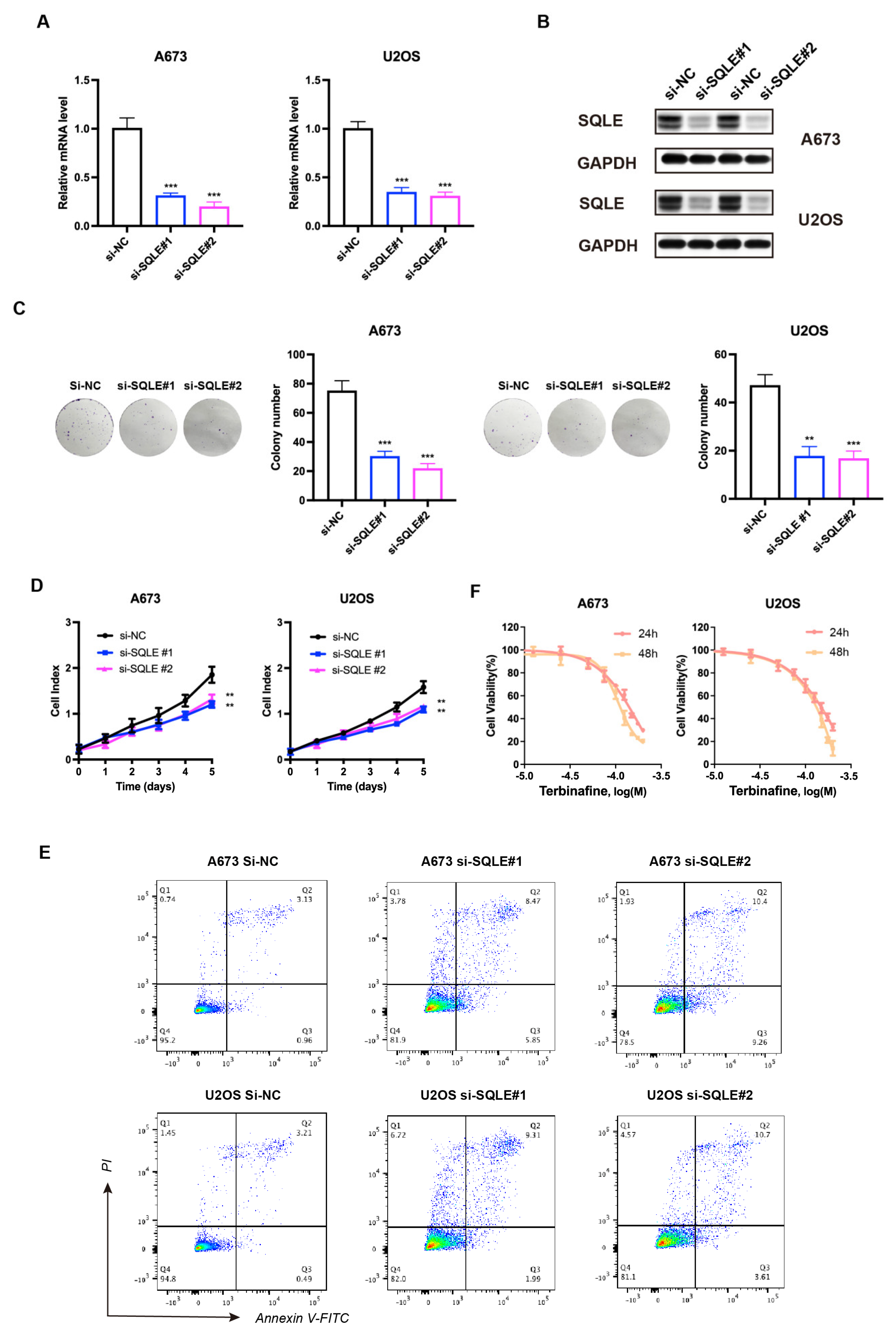
2.8. SQLE as a Potential Target for Therapy
3. Discussion
4. Materials and Methods
4.1. Data Acquisition
4.1.1. Public Datasets
4.1.2. CHCAMS Cohort
4.2. Sarcoma Subcluters Identification
4.3. The Construction and Validation of LMAGs
4.4. Analysis of Tumor Microenvironment and Clinical Treatment Response
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Choi, J.H.; Ro, J.Y. The 2020 WHO Classification of Tumors of Soft Tissue: Selected Changes and New Entities. Adv. Anat. Pathol. 2021, 28, 44–58. [Google Scholar] [CrossRef] [PubMed]
- Skubitz, K.M.; D’Adamo, D.R. Sarcoma. Mayo Clin. Proc. 2007, 82, 1409–1432. [Google Scholar] [CrossRef] [PubMed]
- Stiller, C.A.; Trama, A.; Serraino, D.; Rossi, S.; Navarro, C.; Chirlaque, M.D.; Casali, P.G. Descriptive epidemiology of sarcomas in Europe: Report from the RARECARE project. Eur. J. Cancer 2013, 49, 684–695. [Google Scholar] [CrossRef] [PubMed]
- Burgess, M.; Tawbi, H. Immunotherapeutic approaches to sarcoma. Curr. Treat. Options Oncol. 2015, 16, 26. [Google Scholar] [CrossRef]
- Chen, A.P.; Sharon, E.; O’Sullivan-Coyne, G.; Moore, N.; Foster, J.C.; Hu, J.S.; Van Tine, B.A.; Conley, A.P.; Read, W.L.; Riedel, R.F.; et al. Atezolizumab for Advanced Alveolar Soft Part Sarcoma. N. Engl. J. Med. 2023, 389, 911–921. [Google Scholar] [CrossRef] [PubMed]
- Kachare, S.D.; Brinkley, J.; Vohra, N.A.; Zervos, E.E.; Wong, J.H.; Fitzgerald, T.L. Radiotherapy associated with improved survival for high-grade sarcoma of the extremity. J. Surg. Oncol. 2015, 112, 338–343. [Google Scholar] [CrossRef]
- Ferrari, A.; Dirksen, U.; Bielack, S. Sarcomas of Soft Tissue and Bone. Prog. Tumor Res. 2016, 43, 128–141. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Röhrig, F.; Schulze, A. The multifaceted roles of fatty acid synthesis in cancer. Nat. Rev. Cancer 2016, 16, 732–749. [Google Scholar] [CrossRef]
- Mullen, P.J.; Yu, R.; Longo, J.; Archer, M.C.; Penn, L.Z. The interplay between cell signalling and the mevalonate pathway in cancer. Nat. Rev. Cancer 2016, 16, 718–731. [Google Scholar] [CrossRef]
- Ladanyi, A.; Mukherjee, A.; Kenny, H.A.; Johnson, A.; Mitra, A.K.; Sundaresan, S.; Nieman, K.M.; Pascual, G.; Benitah, S.A.; Montag, A.; et al. Adipocyte-induced CD36 expression drives ovarian cancer progression and metastasis. Oncogene 2018, 37, 2285–2301. [Google Scholar] [CrossRef] [PubMed]
- Pascual, G.; Avgustinova, A.; Mejetta, S.; Martín, M.; Castellanos, A.; Attolini, C.S.; Berenguer, A.; Prats, N.; Toll, A.; Hueto, J.A.; et al. Targeting metastasis-initiating cells through the fatty acid receptor CD36. Nature 2017, 541, 41–45. [Google Scholar] [CrossRef] [PubMed]
- Koundouros, N.; Poulogiannis, G. Reprogramming of fatty acid metabolism in cancer. Br. J. Cancer 2020, 122, 4–22. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Hu, J.W.; He, X.R.; Jin, W.L.; He, X.Y. Statins: A repurposed drug to fight cancer. J. Exp. Clin. Cancer Res. 2021, 40, 241. [Google Scholar] [CrossRef] [PubMed]
- Cardwell, C.R.; Hicks, B.M.; Hughes, C.; Murray, L.J. Statin use after colorectal cancer diagnosis and survival: A population-based cohort study. J. Clin. Oncol. 2014, 32, 3177–3183. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Rahman, O. Statin treatment and outcomes of metastatic pancreatic cancer: A pooled analysis of two phase III studies. Clin. Transl. Oncol. 2019, 21, 810–816. [Google Scholar] [CrossRef] [PubMed]
- Ho, X.D.; Phung, P.; Le, Q.V.; Nguyen, H.V.; Reimann, E.; Prans, E.; Koks, G.; Maasalu, K.; Le, N.T.; Trinh, H.L.; et al. Whole transcriptome analysis identifies differentially regulated networks between osteosarcoma and normal bone samples. Exp. Biol. Med. 2017, 242, 1802–1811. [Google Scholar] [CrossRef] [PubMed]
- Broadfield, L.A.; Pane, A.A.; Talebi, A.; Swinnen, J.V.; Fendt, S.M. Lipid metabolism in cancer: New perspectives and emerging mechanisms. Dev. Cell 2021, 56, 1363–1393. [Google Scholar] [CrossRef]
- Schade, D.S.; Shey, L.; Eaton, R.P. Cholesterol Review: A Metabolically Important Molecule. Endocr. Pract. 2020, 26, 1514–1523. [Google Scholar] [CrossRef]
- Liu, D.; Wong, C.C.; Fu, L.; Chen, H.; Zhao, L.; Li, C.; Zhou, Y.; Zhang, Y.; Xu, W.; Yang, Y.; et al. Squalene epoxidase drives NAFLD-induced hepatocellular carcinoma and is a pharmaceutical target. Sci. Transl. Med. 2018, 10, eaap9840. [Google Scholar] [CrossRef]
- Zhang, Z.; Wu, W.; Jiao, H.; Chen, Y.; Ji, X.; Cao, J.; Yin, F.; Yin, W. Squalene epoxidase promotes hepatocellular carcinoma development by activating STRAP transcription and TGF-β/SMAD signalling. Br. J. Pharmacol. 2023, 180, 1562–1581. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wang, Y.; Liu, D.; Wong, C.C.; Coker, O.O.; Zhang, X.; Liu, C.; Zhou, Y.; Liu, Y.; Kang, W.; et al. Squalene epoxidase drives cancer cell proliferation and promotes gut dysbiosis to accelerate colorectal carcinogenesis. Gut 2022, 71, 2253–2265. [Google Scholar] [CrossRef] [PubMed]
- Robichaux, J.P.; Le, X.; Vijayan, R.S.K.; Hicks, J.K.; Heeke, S.; Elamin, Y.Y.; Lin, H.Y.; Udagawa, H.; Skoulidis, F.; Tran, H.; et al. Structure-based classification predicts drug response in EGFR-mutant NSCLC. Nature 2021, 597, 732–737. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Xu, C. Immune checkpoint signaling and cancer immunotherapy. Cell Res. 2020, 30, 660–669. [Google Scholar] [CrossRef] [PubMed]
- Coleman, M.F.; Cozzo, A.J.; Pfeil, A.J.; Etigunta, S.K.; Hursting, S.D. Cell Intrinsic and Systemic Metabolism in Tumor Immunity and Immunotherapy. Cancers 2020, 12, 852. [Google Scholar] [CrossRef]
- Li, X.; Wenes, M.; Romero, P.; Huang, S.C.; Fendt, S.M.; Ho, P.C. Navigating metabolic pathways to enhance antitumour immunity and immunotherapy. Nat. Rev. Clin. Oncol. 2019, 16, 425–441. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Yang, D.; Yang, Q.; Lv, X.; Huang, W.; Zhou, Z.; Wang, Y.; Zhang, Z.; Yuan, T.; Ding, X.; et al. Single-cell RNA landscape of intratumoral heterogeneity and immunosuppressive microenvironment in advanced osteosarcoma. Nat. Commun. 2020, 11, 6322. [Google Scholar] [CrossRef]
- Zhu, M.M.T.; Shenasa, E.; Nielsen, T.O. Sarcomas: Immune biomarker expression and checkpoint inhibitor trials. Cancer Treat. Rev. 2020, 91, 102115. [Google Scholar] [CrossRef]
- Hong, Z.; Liu, T.; Wan, L.; Fa, P.; Kumar, P.; Cao, Y.; Prasad, C.B.; Qiu, Z.; Liu, J.; Wang, H.; et al. Targeting Squalene Epoxidase Interrupts Homologous Recombination via the ER Stress Response and Promotes Radiotherapy Efficacy. Cancer Res. 2022, 82, 1298–1312. [Google Scholar] [CrossRef]
- Frazier, W.T.; Santiago-Delgado, Z.M.; Stupka, K.C., 2nd. Onychomycosis: Rapid Evidence Review. Am. Fam. Physician 2021, 104, 359–367. [Google Scholar]
- Zhang, E.B.; Zhang, X.; Wang, K.; Zhang, F.; Chen, T.W.; Ma, N.; Ni, Q.Z.; Wang, Y.K.; Zheng, Q.W.; Cao, H.J.; et al. Antifungal agent Terbinafine restrains tumor growth in preclinical models of hepatocellular carcinoma via AMPK-mTOR axis. Oncogene 2021, 40, 5302–5313. [Google Scholar] [CrossRef] [PubMed]
- Tan, J.M.E.; van der Stoel, M.M.; van den Berg, M.; van Loon, N.M.; Moeton, M.; Scholl, E.; van der Wel, N.N.; Kovačević, I.; Hordijk, P.L.; Loregger, A.; et al. The MARCH6-SQLE Axis Controls Endothelial Cholesterol Homeostasis and Angiogenic Sprouting. Cell Rep. 2020, 32, 107944. [Google Scholar] [CrossRef]
- He, L.; Li, H.; Pan, C.; Hua, Y.; Peng, J.; Zhou, Z.; Zhao, Y.; Lin, M. Squalene epoxidase promotes colorectal cancer cell proliferation through accumulating calcitriol and activating CYP24A1-mediated MAPK signaling. Cancer Commun. 2021, 41, 726–746. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.P.; Huang, W.; Wang, X.; Xu, C.; Qin, W.T.; Li, D.; Tian, G.; Li, Q.; Zhou, Y.; Chen, S.; et al. Terbinafine prevents colorectal cancer growth by inducing dNTP starvation and reducing immune suppression. Mol. Ther. 2022, 30, 3284–3299. [Google Scholar] [CrossRef] [PubMed]
- Vilcek, J.; Feldmann, M. Historical review: Cytokines as therapeutics and targets of therapeutics. Trends Pharmacol. Sci. 2004, 25, 201–209. [Google Scholar] [CrossRef]
- Creaven, P.J.; Plager, J.E.; Dupere, S.; Huben, R.P.; Takita, H.; Mittelman, A.; Proefrock, A. Phase I clinical trial of recombinant human tumor necrosis factor. Cancer Chemother. Pharmacol. 1987, 20, 137–144. [Google Scholar] [CrossRef] [PubMed]
- Brown, T.D.; Goodman, P.; Fleming, T.; Macdonald, J.S.; Hersh, E.M.; Braun, T.J. A phase II trial of recombinant tumor necrosis factor in patients with adenocarcinoma of the pancreas: A Southwest Oncology Group study. J. Immunother. 1991, 10, 376–378. [Google Scholar] [CrossRef]
- Manusama, E.R.; Stavast, J.; Durante, N.M.; Marquet, R.L.; Eggermont, A.M. Isolated limb perfusion with TNF alpha and melphalan in a rat osteosarcoma model: A new anti-tumour approach. Eur. J. Surg. Oncol. 1996, 22, 152–157. [Google Scholar] [CrossRef]
- van Horssen, R.; Ten Hagen, T.L.; Eggermont, A.M. TNF-alpha in cancer treatment: Molecular insights, antitumor effects, and clinical utility. Oncologist 2006, 11, 397–408. [Google Scholar] [CrossRef]
- Munisamy, S.; Radhakrishnan, A.K.; Ramdas, P.; Samuel, P.J.; Singh, V.A. Immune Biomarkers in Blood from Sarcoma Patients: A Pilot Study. Curr. Oncol. 2022, 29, 5585–5603. [Google Scholar] [CrossRef]
- Märtson, A.; Kõks, S.; Reimann, E.; Prans, E.; Erm, T.; Maasalu, K. Transcriptome analysis of osteosarcoma identifies suppression of wnt pathway and up-regulation of adiponectin as potential biomarker. Genom. Discov. 2013, 1, 3. [Google Scholar] [CrossRef][Green Version]
- Reimann, E.; Kõks, S.; Ho, X.D.; Maasalu, K.; Märtson, A. Whole exome sequencing of a single osteosarcoma case—Integrative analysis with whole transcriptome RNA-seq data. Hum. Genom. 2014, 8, 20. [Google Scholar] [CrossRef]
- Brunet, J.P.; Tamayo, P.; Golub, T.R.; Mesirov, J.P. Metagenes and molecular pattern discovery using matrix factorization. Proc. Natl. Acad. Sci. USA 2004, 101, 4164–4169. [Google Scholar] [CrossRef] [PubMed]
- Tibshirani, R. The lasso method for variable selection in the Cox model. Stat. Med. 1997, 16, 385–395. [Google Scholar] [CrossRef]
- Bradburn, M.J.; Clark, T.G.; Love, S.B.; Altman, D.G. Survival analysis part II: Multivariate data analysis—An introduction to concepts and methods. Br. J. Cancer 2003, 89, 431–436. [Google Scholar] [CrossRef] [PubMed]
- Heagerty, P.J.; Lumley, T.; Pepe, M.S. Time-dependent ROC curves for censored survival data and a diagnostic marker. Biometrics 2000, 56, 337–344. [Google Scholar] [CrossRef] [PubMed]
- Møller-Petersen, J. Nomogram for predictive values and efficiencies of tests. Lancet 1985, 1, 348. [Google Scholar] [CrossRef]
- Mayakonda, A.; Lin, D.C.; Assenov, Y.; Plass, C.; Koeffler, H.P. Maftools: Efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018, 28, 1747–1756. [Google Scholar] [CrossRef]
- Yoshihara, K.; Shahmoradgoli, M.; Martínez, E.; Vegesna, R.; Kim, H.; Torres-Garcia, W.; Treviño, V.; Shen, H.; Laird, P.W.; Levine, D.A.; et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 2013, 4, 2612. [Google Scholar] [CrossRef]
- Maeser, D.; Gruener, R.F.; Huang, R.S. oncoPredict: An R package for predicting in vivo or cancer patient drug response and biomarkers from cell line screening data. Brief. Bioinform. 2021, 22, bbab260. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hong, Y.; Zhang, L.; Lin, W.; Yang, Y.; Cao, Z.; Feng, X.; Yu, Z.; Gao, Y. Transcriptome Sequencing Unveils a Molecular-Stratification-Predicting Prognosis of Sarcoma Associated with Lipid Metabolism. Int. J. Mol. Sci. 2024, 25, 1643. https://doi.org/10.3390/ijms25031643
Hong Y, Zhang L, Lin W, Yang Y, Cao Z, Feng X, Yu Z, Gao Y. Transcriptome Sequencing Unveils a Molecular-Stratification-Predicting Prognosis of Sarcoma Associated with Lipid Metabolism. International Journal of Molecular Sciences. 2024; 25(3):1643. https://doi.org/10.3390/ijms25031643
Chicago/Turabian StyleHong, Yuheng, Lin Zhang, Weihao Lin, Yannan Yang, Zheng Cao, Xiaoli Feng, Zhentao Yu, and Yibo Gao. 2024. "Transcriptome Sequencing Unveils a Molecular-Stratification-Predicting Prognosis of Sarcoma Associated with Lipid Metabolism" International Journal of Molecular Sciences 25, no. 3: 1643. https://doi.org/10.3390/ijms25031643
APA StyleHong, Y., Zhang, L., Lin, W., Yang, Y., Cao, Z., Feng, X., Yu, Z., & Gao, Y. (2024). Transcriptome Sequencing Unveils a Molecular-Stratification-Predicting Prognosis of Sarcoma Associated with Lipid Metabolism. International Journal of Molecular Sciences, 25(3), 1643. https://doi.org/10.3390/ijms25031643




