Applications of Genome-Editing Technologies for Type 1 Diabetes
Abstract
1. Introduction
1.1. Diabetes Etiology
1.2. Gene-Editing Technologies and CRISPR–Cas Systems
2. T1D In Vivo Models
CRISPR–Cas9-Engineered T1D Mouse Models
3. β-Cell Replacement
3.1. Insulin Production
3.2. Transplants
3.3. Stem Cells and Induced Pluripotent Stem Cell Therapy
4. Strategies for Protecting Transplanted β-Cells
5. Immunosuppressant Drugs
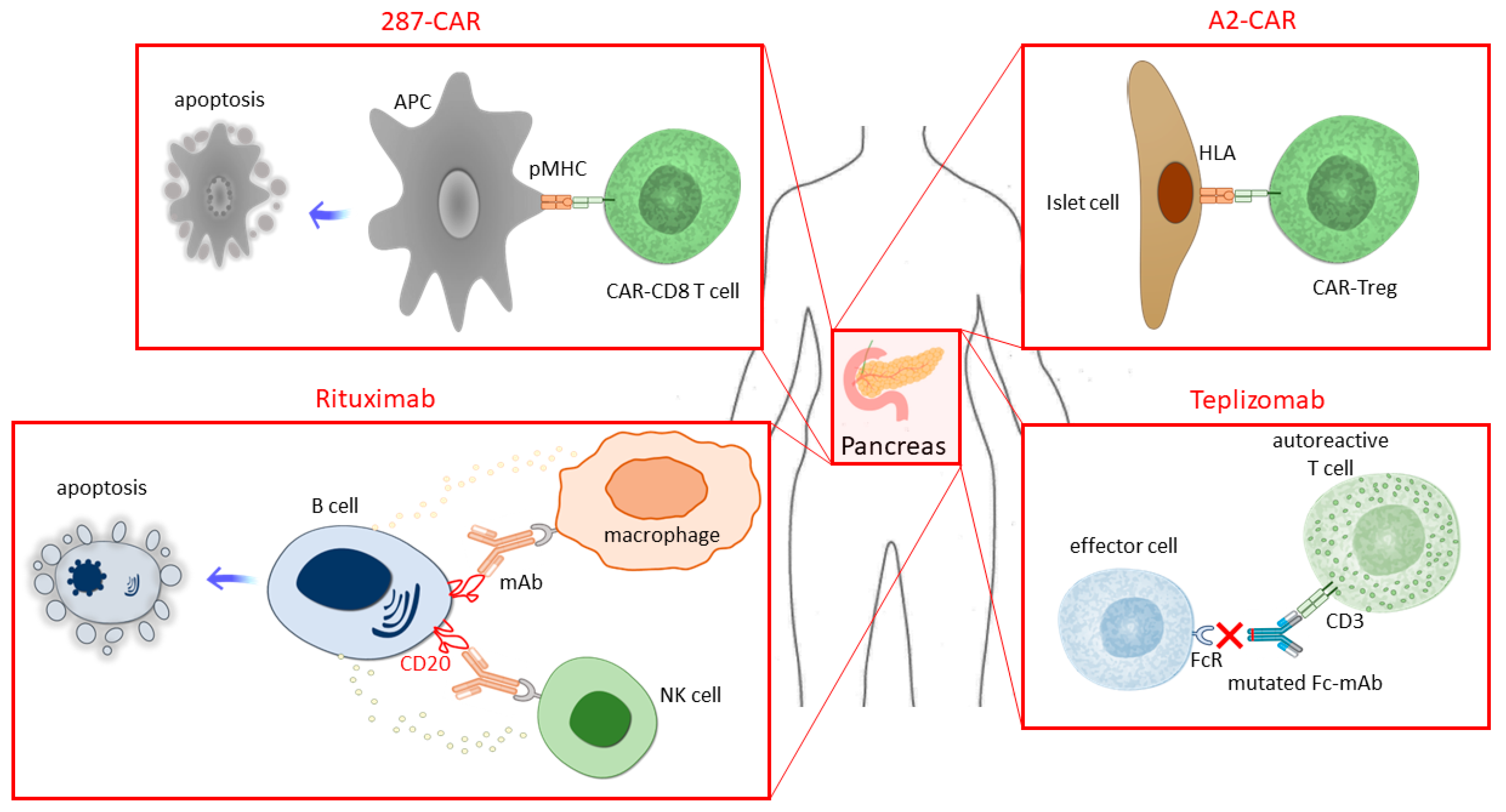
6. Engineered Immune Cells
7. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AICDA | Activation-induced cytidine deaminase gene |
| AID | Activation-induced cytidine deaminase protein |
| AMM | Automated meiotic mapping |
| c-Myc | cellular Myc |
| CD | Cluster of differentiation |
| CNS1 | Conserved non-coding sequence 1 |
| CRISPR–Cas9 | Clustered regularly interspaced palindromic repeats and Cas-associated protein 9 |
| CTLA | Cytotoxic T-lymphocyte antigen |
| DUSP10 | Dual-specificity phosphatase gene |
| ESCs | Embryonic stem cells |
| FcRn | Neonatal Fc receptor |
| FOXP3 | Forkhead box protein P3 |
| GWAS | Genome-wide association study |
| HLA | Human leukocyte antigen |
| HRD | Homology-directed repair |
| IAA | Insulin autoantibodies |
| IBMIR | Instant blood-mediated inflammatory reaction |
| Idd | Insulin-dependent diabetes |
| IFN | Interferon |
| IL-2 | Interleukin-2 |
| INS | Preproinsulin |
| IPC | Insulin-producing cells |
| IPPBC | Insulin-producing pancreatic β-cells |
| IPSCs | Induced pluripotent stem cells |
| Klf4 | Krüppel-like factor 4 |
| LV | Lentiviral vector |
| MHC | Major histocompatibility complex |
| MSC | Mesenchymal stem cells |
| NHEJ | Non-homologous end joining |
| NOD | Non-obese diabetic |
| Oct3/4 | Octamer-binding transcription factor |
| PBMCs | Peripheral blood mononuclear cells |
| PHPI | Purified human pancreatic islets |
| PTPN1 | Gene-encoding protein tyrosine phosphatases 1 |
| Ptpn22 | Protein tyrosine phosphatase nonreceptor type 22 |
| PTPN6 | Gene-encoding protein tyrosine phosphatases 6 |
| pTregs | Peripheral Treg |
| RORγt+ | Retinoic acid receptor-related orphan receptor gamma T |
| SBCs | Stem β cells |
| SC | Stem cell |
| SC-β cells | Stem cell-derived β-cells |
| sgRNA | single guide RNA |
| Sox2 | Sex-determining region Y-Box 2 |
| STING | Stimulator of IFN genes |
| T1D | Type 1 diabetes |
| TCR | T-cell receptor |
| TEC | Thymic epithelial cells |
| Teffs | Effector T cells |
| TNF | Tumor necrosis factor |
| Treg | Regulatory T cells |
| tTreg | Thymic Treg |
References
- Lucier, J.; Weinstock, R.S. Type 1 Diabetes; StatPearls Publishing: Treasure Island, FL, USA, 2023. [Google Scholar]
- Karamanou, M.; Protogerou, A.; Tsoucalas, G.; Androutsos, G.; Poulakou-Rebelakou, E. Milestones in the history of diabetes mellitus: The main contributors. World J. Diabetes 2016, 7, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Banting, F.G.; Best, C.H.; Collip, J.B.; Campbell, W.R.; Fletcher, A.A. Pancreatic Extracts in the Treatment of Diabetes Mellitus. Can Med. Assoc. J. 1922, 12, 141–146. [Google Scholar] [PubMed]
- DiMeglio, L.A.; Evans-Molina, C.; Oram, R.A. Type 1 diabetes. Lancet 2018, 391, 2449–2462. [Google Scholar] [CrossRef] [PubMed]
- Warshauer, J.T.; Bluestone, J.A.; Anderson, M.S. New Frontiers in the Treatment of Type 1 Diabetes. Cell Metab. 2020, 31, 46–61. [Google Scholar] [CrossRef] [PubMed]
- Ratiu, J.J.; Racine, J.J.; Hasham, M.G.; Wang, Q.; Branca, J.A.; Chapman, H.D.; Zhu, J.; Donghia, N.; Philip, V.; Schott, W.H.; et al. Genetic and Small Molecule Disruption of the AID/RAD51 Axis Similarly Protects Nonobese Diabetic Mice from Type 1 Diabetes through Expansion of Regulatory B Lymphocytes. J. Immunol. 2017, 198, 4255–4267. [Google Scholar] [CrossRef] [PubMed]
- Gerace, D.; Martiniello-Wilks, R.; Nassif, N.T.; Lal, S.; Steptoe, R.; Simpson, A.M. CRISPR-targeted genome editing of mesenchymal stem cell-derived therapies for type 1 diabetes: A path to clinical success? Stem. Cell Res. Ther. 2017, 8, 62. [Google Scholar] [CrossRef]
- Quattrin, T.; Mastrandrea, L.D.; Walker, L.S.K. Type 1 diabetes. Lancet 2023, 401, 2149–2162. [Google Scholar] [CrossRef]
- Todd, J.A. Etiology of type 1 diabetes. Immunity 2010, 32, 457–467. [Google Scholar] [CrossRef]
- Foray, A.P.; Candon, S.; Hildebrand, S.; Marquet, C.; Valette, F.; Pecquet, C.; Lemoine, S.; Langa-Vives, F.; Dumas, M.; Hu, P.; et al. De novo germline mutation in the dual specificity phosphatase 10 gene accelerates autoimmune diabetes. Proc. Natl. Acad. Sci. USA 2021, 118. [Google Scholar] [CrossRef]
- Thomaidou, S.; Zaldumbide, A.; Roep, B.O. Islet stress, degradation and autoimmunity. Diabetes Obes. Metab. 2018, 20 (Suppl. 2), 88–94. [Google Scholar] [CrossRef]
- Anderson, A.M.; Landry, L.G.; Alkanani, A.A.; Pyle, L.; Powers, A.C.; Atkinson, M.A.; Mathews, C.E.; Roep, B.O.; Michels, A.W.; Nakayama, M. Human islet T cells are highly reactive to preproinsulin in type 1 diabetes. Proc. Natl. Acad. Sci. USA 2021, 118, e2107208118. [Google Scholar] [CrossRef]
- Lin, X.; Pelletier, S.; Gingras, S.; Rigaud, S.; Maine, C.J.; Marquardt, K.; Dai, Y.D.; Sauer, K.; Rodriguez, A.R.; Martin, G.; et al. CRISPR-Cas9-Mediated Modification of the NOD Mouse Genome with Ptpn22R619W Mutation Increases Autoimmune Diabetes. Diabetes 2016, 65, 2134–2138. [Google Scholar] [CrossRef]
- Krogvold, L.; Edwin, B.; Buanes, T.; Frisk, G.; Skog, O.; Anagandula, M.; Korsgren, O.; Undlien, D.; Eike, C.M.; Richardson, J.S.; et al. Detection of a low-grade enteroviral infection in the islets of langerhans of living patients newly diagnosed with type 1 diabetes. Diabetes 2015, 64, 1682–1687. [Google Scholar] [CrossRef]
- Buschard, K. The etiology and pathogenesis of type 1 diabetes—A personal, non-systematic review of possible causes, and interventions. Front. Endocrinol. 2022, 13, 876470. [Google Scholar] [CrossRef]
- Brown, C.T.; Davis-Richardson, A.G.; Giongo, A.; Gano, K.A.; Crabb, D.B.; Mukherjee, N.; Casella, G.; Drew, J.C.; Ilonen, J.; Knip, M.; et al. Gut microbiome metagenomics analysis suggests a functional model for the development of autoimmunity for type 1 diabetes. PLoS ONE 2011, 6, e25792. [Google Scholar] [CrossRef]
- Pedersen, K.; Haupt-Jorgensen, M.; Krogvold, L.; Kaur, S.; Gerling, I.C.; Pociot, F.; Dahl-Jørgensen, K.; Buschard, K. Genetic predisposition in the 2′-5′A pathway in the development of type 1 diabetes: Potential contribution to dysregulation of innate antiviral immunity. Diabetologia 2021, 64, 1805–1815. [Google Scholar] [CrossRef]
- Sayed, N.; Allawadhi, P.; Khurana, A.; Singh, V.; Navik, U.; Pasumarthi, S.K.; Khurana, I.; Banothu, A.K.; Weiskirchen, R.; Bharani, K.K. Gene therapy: Comprehensive overview and therapeutic applications. Life Sci. 2022, 294, 120375. [Google Scholar] [CrossRef]
- Liu, G.; Lin, Q.; Jin, S.; Gao, C. The CRISPR-Cas toolbox and gene editing technologies. Mol. Cell. 2022, 82, 333–347. [Google Scholar] [CrossRef]
- Olivares, A.M.; Althoff, K.; Chen, G.F.; Wu, S.; Morrisson, M.A.; DeAngelis, M.M.; Haider, N. Animal Models of Diabetic Retinopathy. Curr. Diab. Rep. 2017, 17, 93. [Google Scholar] [CrossRef]
- Moshref, M.; Tangey, B.; Gilor, C.; Papas, K.K.; Williamson, P.; Loomba-Albrecht, L.; Sheehy, P.; Kol, A. Concise Review: Canine Diabetes Mellitus as a Translational Model for Innovative Regenerative Medicine Approaches. Stem. Cells Transl. Med. 2019, 8, 450–455. [Google Scholar] [CrossRef]
- van Rooijen, E.; Voest, E.E.; Logister, I.; Bussmann, J.; Korving, J.; van Eeden, F.J.; Giles, R.H.; Schulte-Merker, S. von Hippel-Lindau tumor suppressor mutants faithfully model pathological hypoxia-driven angiogenesis and vascular retinopathies in zebrafish. Dis. Model Mech. 2010, 3, 343–353. [Google Scholar] [CrossRef] [PubMed]
- Naqvi, R.A.; Naqvi, A.R.; Singh, A.; Priyadarshini, M.; Balamurugan, A.N.; Layden, B.T. The future treatment for type 1 diabetes: Pig islet- or stem cell-derived beta cells? Front. Endocrinol. 2022, 13, 1001041. [Google Scholar] [CrossRef] [PubMed]
- Kalaitzoglou, E.; Fowlkes, J.L.; Thrailkill, K.M. Mouse models of type 1 diabetes and their use in skeletal research. Curr. Opin. Endocrinol. Diabetes Obes. 2022, 29, 318–325. [Google Scholar] [CrossRef] [PubMed]
- Aoki, C.A.; Borchers, A.T.; Ridgway, W.M.; Keen, C.L.; Ansari, A.A.; Gershwin, M.E. NOD mice and autoimmunity. Autoimmun. Rev. 2005, 4, 373–379. [Google Scholar] [CrossRef] [PubMed]
- Presa, M.; Racine, J.J.; Dwyer, J.R.; Lamont, D.J.; Ratiu, J.J.; Sarsani, V.K.; Chen, Y.-G.; Geurts, A.; Schmitz, I.; Stearns, T.; et al. A Hypermorphic Nfkbid Allele Contributes to Impaired Thymic Deletion of Autoreactive Diabetogenic CD8(+) T Cells in NOD Mice. J. Immunol. 2018, 201, 1907–1917. [Google Scholar] [CrossRef]
- Wang, J.Y.; Doudna, J.A. CRISPR technology: A decade of genome editing is only the beginning. Science 2023, 379, eadd8643. [Google Scholar] [CrossRef]
- Stanley, W.J.; Trivedi, P.M.; Sutherland, A.P.; Thomas, H.E.; Gurzov, E.N. Differential regulation of pro-inflammatory cytokine signalling by protein tyrosine phosphatases in pancreatic beta-cells. J. Mol. Endocrinol. 2017, 59, 325–337. [Google Scholar] [CrossRef]
- Racine, J.J.; Stewart, I.; Ratiu, J.; Christianson, G.; Lowell, E.; Helm, K.; Allocco, J.; Maser, R.S.; Chen, Y.-G.; Lutz, C.M.; et al. Improved Murine MHC-Deficient HLA Transgenic NOD Mouse Models for Type 1 Diabetes Therapy Development. Diabetes 2018, 67, 923–935. [Google Scholar] [CrossRef]
- Holohan, D.R.; Van Gool, F.; Bluestone, J.A. Thymically-derived Foxp3+ regulatory T cells are the primary regulators of type 1 diabetes in the non-obese diabetic mouse model. PLoS ONE 2019, 14, e0217728. [Google Scholar] [CrossRef]
- Cai, E.P.; Ishikawa, Y.; Zhang, W.; Leite, N.C.; Li, J.; Hou, S.; Kiaf, B.; Hollister-Lock, J.; Yilmaz, N.K.; Schiffer, C.A.; et al. Genome-scale in vivo CRISPR screen identifies RNLS as a target for beta cell protection in type 1 diabetes. Nat. Metab. 2020, 2, 934–945. [Google Scholar] [CrossRef]
- Akazawa, S.; Mackin, L.; Jhala, G.; Fynch, S.; Catterall, T.; Selck, C.; Graham, K.L.; Krishnamurthy, B.; Pappas, E.G.; Kwong, C.-T.J.; et al. Deficiency of the innate immune adaptor STING promotes autoreactive T cell expansion in NOD mice. Diabetologia 2021, 64, 878–889. [Google Scholar] [CrossRef] [PubMed]
- Sintov, E.; Nikolskiy, I.; Barrera, V.; Kenty, J.H.-R.; Atkin, A.S.; Gerace, D.; Sui, S.J.H.; Boulanger, K.; Melton, D.A. Whole-genome CRISPR screening identifies genetic manipulations to reduce immune rejection of stem cell-derived islets. Stem. Cell Rep. 2022, 17, 1976–1990. [Google Scholar] [CrossRef] [PubMed]
- Russo, F.; Ruggiero, E.; Curto, R.; Passeri, L.; Sanvito, F.; Bortolomai, I.; Villa, A.; Gregori, S.; Annoni, A. Editing T cell repertoire by thymic epithelial cell-directed gene transfer abrogates risk of type 1 diabetes development. Mol. Ther. Methods Clin. Dev. 2022, 25, 508–519. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.H.; Shin, J.I.; Yang, J.W.; Lee, K.H.; Cha, D.H.; Hong, J.B.; Park, Y.; Choi, E.; Tizaoui, K.; Koyanagi, A.; et al. Genome Editing Using CRISPR-Cas9 and Autoimmune Diseases: A Comprehensive Review. Int. J. Mol. Sci. 2022, 23, 1337. [Google Scholar] [CrossRef] [PubMed]
- Peters, L.D.; Yeh, W.I.; Arnoletti, J.M.; Brown, M.E.; Posgai, A.L.; Mathews, C.E.; Brusko, T.M. Modeling cell-mediated immunity in human type 1 diabetes by engineering autoreactive CD8(+) T cells. Front. Immunol. 2023, 14, 1142648. [Google Scholar] [CrossRef] [PubMed]
- Demenais, F.; Margaritte-Jeannin, P.; Barnes, K.C.; Cookson, W.O.; Altmüller, J.; Ang, W.; Barr, R.G.; Beaty, T.H.; Becker, A.B.; Beilby, J.; et al. Multiancestry association study identifies new asthma risk loci that colocalize with immune-cell enhancer marks. Nat. Genet. 2018, 50, 42–53. [Google Scholar] [CrossRef] [PubMed]
- Auricchio, A.; Gao, G.P.; Yu, Q.C.; Raper, S.; Rivera, V.M.; Clackson, T.; Wilson, J.M. Constitutive and regulated expression of processed insulin following in vivo hepatic gene transfer. Gene Ther. 2002, 9, 963–971. [Google Scholar] [CrossRef][Green Version]
- Thule, P.M.; Liu, J.; Phillips, L.S. Glucose regulated production of human insulin in rat hepatocytes. Gene Ther. 2000, 7, 205–214. [Google Scholar] [CrossRef]
- Sanlioglu, A.D.; Altunbas, H.A.; Balci, M.K.; Griffith, T.S.; Sanlioglu, S. Insulin gene therapy from design to beta cell generation. Expert Rev. Mol. Med. 2012, 14, e18. [Google Scholar] [CrossRef]
- Subhashree, A.R. Red Cell Distribution Width and Serum BNP Level Correlation in Diabetic Patients with Cardiac Failure: A Cross—Sectional Study. J. Clin. Diagn. Res. 2014, 8, FC01–FC03. [Google Scholar]
- Rickels, M.R.; Robertson, R.P. Pancreatic Islet Transplantation in Humans: Recent Progress and Future Directions. Endocr. Rev. 2019, 40, 631–668. [Google Scholar] [CrossRef] [PubMed]
- Barton, F.B.; Rickels, M.R.; Alejandro, R.; Hering, B.J.; Wease, S.; Naziruddin, B.; Oberholzer, J.; Odorico, J.S.; Garfinkel, M.R.; Levy, M.; et al. Improvement in outcomes of clinical islet transplantation: 1999–2010. Diabetes Care 2012, 35, 1436–1445. [Google Scholar] [CrossRef] [PubMed]
- Vantyghem, M.C.; de Koning, E.J.; Pattou, F.; Rickels, M.R. Advances in beta-cell replacement therapy for the treatment of type 1 diabetes. Lancet 2019, 394, 1274–1285. [Google Scholar] [CrossRef] [PubMed]
- Hering, B.J.; Clarke, W.R.; Bridges, N.D.; Eggerman, T.L.; Alejandro, R.; Bellin, M.D.; Chaloner, K.; Czarniecki, C.W.; Goldstein, J.S.; Hunsicker, L.G.; et al. Phase 3 Trial of Transplantation of Human Islets in Type 1 Diabetes Complicated by Severe Hypoglycemia. Diabetes Care 2016, 39, 1230–1240. [Google Scholar] [CrossRef] [PubMed]
- Bolla, A.M.; Montefusco, L.; Pastore, I.; Lunati, M.E.; Ben Nasr, M.; Fiorina, P. Benefits and Hurdles of Pancreatic beta-Cell Replacement. Stem. Cells Transl. Med. 2022, 11, 1029–1039. [Google Scholar] [CrossRef] [PubMed]
- FDA, L. Available online: https://www.fda.gov/news-events/press-announcements/fda-approves-first-cellular-therapy-treat-patients-type-1-diabetes (accessed on 5 November 2023).
- Groth, C.G.; Tibell, A.; Tollemar, J.; Bolinder, J.; Östman, J.; Möller, E.; Reinholt, F.; Korsgren, O.; Hellerström, C.; Andersson, A. Transplantation of porcine fetal pancreas to diabetic patients. Lancet 1994, 344, 1402–1404. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, S.; Tomiya, M.; Sawamoto, O. Current status and future of clinical islet xenotransplantation. J. Diabetes 2016, 8, 483–493. [Google Scholar] [CrossRef]
- Bottino, R.; Trucco, M. Use of genetically-engineered pig donors in islet transplantation. World J. Transpl. 2015, 5, 243–250. [Google Scholar] [CrossRef]
- Thompson, P.; Badell, I.R.; Lowe, M.; Cano, J.; Song, M.; Leopardi, F.; Avila, J.; Ruhil, R.; Strobert, E.; Korbutt, G.; et al. Islet xenotransplantation using gal-deficient neonatal donors improves engraftment and function. Am. J. Transpl. 2011, 11, 2593–2602. [Google Scholar] [CrossRef]
- van der Windt, D.J.; Bottino, R.; Casu, A.; Campanile, N.; Smetanka, C.; He, J.; Murase, N.; Hara, H.; Ball, S.; Loveland, B.; et al. Long-term controlled normoglycemia in diabetic non-human primates after transplantation with hCD46 transgenic porcine islets. Am. J. Transpl. 2009, 9, 2716–2726. [Google Scholar] [CrossRef]
- Korbutt, G.S.; Elliott, J.F.; Ao, Z.; Smith, D.K.; Warnock, G.L.; Rajotte, R.V. Large scale isolation, growth, and function of porcine neonatal islet cells. J. Clin. Investig. 1996, 97, 2119–2129. [Google Scholar] [CrossRef] [PubMed]
- Koehler, N.; Buhler, L.; Egger, B.; Gonelle-Gispert, C. Multipotent Mesenchymal Stromal Cells Interact and Support Islet of Langerhans Viability and Function. Front. Endocrinol. 2022, 13, 822191. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Sordi, V.; Pellegrini, S.; Piemonti, L. Immunological Issues After Stem Cell-Based beta Cell Replacement. Curr. Diab. Rep. 2017, 17, 68. [Google Scholar] [CrossRef] [PubMed]
- Maxwell, K.G.; Millman, J.R. Applications of iPSC-derived beta cells from patients with diabetes. Cell Rep. Med. 2021, 2, 100238. [Google Scholar] [CrossRef] [PubMed]
- Stock, A.A.; Manzoli, V.; De Toni, T.; Abreu, M.M.; Poh, Y.C.; Ye, L.; Roose, A.; Pagliuca, F.W.; Thanos, C.; Ricordi, C.; et al. Conformal Coating of Stem Cell-Derived Islets for beta Cell Replacement in Type 1 Diabetes. Stem. Cell Rep. 2020, 14, 91–104. [Google Scholar] [CrossRef] [PubMed]
- Deuse, T.; Hu, X.; Agbor-Enoh, S.; Koch, M.; Spitzer, M.H.; Gravina, A.; Alawi, M.; Marishta, A.; Peters, B.; Kosaloglu-Yalcin, Z.; et al. De novo mutations in mitochondrial DNA of iPSCs produce immunogenic neoepitopes in mice and humans. Nat. Biotechnol. 2019, 37, 1137–1144. [Google Scholar] [CrossRef] [PubMed]
- Lampeter, E.F.; Homberg, M.; Gries, F.; Kolb, H.; Quabeck, K.; Schaefer, U.; Wernet, P.; Grosse-Wilde, H.; Bertrams, J. Transfer of insulin-dependent diabetes between HLA-identical siblings by bone marrow transplantation. Lancet 1993, 341, 1243–1244. [Google Scholar] [CrossRef]
- Bellin, M.D.; Moran, A.; Wilhelm, J.J.; O’Brien, T.D.; Gottlieb, P.A.; Yu, L.; Dunn, T.B. Development of Autoimmune-Mediated beta Cell Failure After Total Pancreatectomy With Autologous Islet Transplantation. Am. J. Transpl. 2015, 15, 1991–1994. [Google Scholar] [CrossRef]
- Piemonti, L.; Everly, M.J.; Maffi, P.; Scavini, M.; Poli, F.; Nano, R.; Cardillo, M.; Melzi, R.; Mercalli, A.; Sordi, V.; et al. Alloantibody and autoantibody monitoring predicts islet transplantation outcome in human type 1 diabetes. Diabetes 2013, 62, 1656–1664. [Google Scholar] [CrossRef]
- Brusko, T.M.; Russ, H.A.; Stabler, C.L. Strategies for durable beta cell replacement in type 1 diabetes. Science 2021, 373, 516–522. [Google Scholar] [CrossRef] [PubMed]
- Felton, J.L.; Conway, H.; Bonami, R.H. B Quiet: Autoantigen-Specific Strategies to Silence Raucous B Lymphocytes and Halt Cross-Talk with T Cells in Type 1 Diabetes. Biomedicines 2021, 9, 42. [Google Scholar] [CrossRef]
- Fanaropoulou, N.M.; Tsatsani, G.C.; Koufakis, T.; Kotsa, K. Teplizumab: Promises and challenges of a recently approved monoclonal antibody for the prevention of type 1 diabetes or preservation of residual beta cell function. Expert Rev. Clin. Immunol. 2023, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Bettini, M.; Bettini, M.L. Function, Failure, and the Future Potential of Tregs in Type 1 Diabetes. Diabetes 2021, 70, 1211–1219. [Google Scholar] [CrossRef] [PubMed]
- Bluestone, J.A.; Buckner, J.H.; Fitch, M.; Gitelman, S.E.; Gupta, S.; Hellerstein, M.K.; Herold, K.C.; Lares, A.; Lee, M.R.; Li, K.; et al. Type 1 diabetes immunotherapy using polyclonal regulatory T cells. Sci. Transl. Med. 2015, 7, 315ra189. [Google Scholar] [CrossRef] [PubMed]
- Raffin, C.; Vo, L.T.; Bluestone, J.A. T(reg) cell-based therapies: Challenges and perspectives. Nat. Rev. Immunol. 2020, 20, 158–172. [Google Scholar] [CrossRef]
- Muller, Y.D.; Ferreira, L.M.R.; Ronin, E.; Ho, P.; Nguyen, V.; Faleo, G.; Zhou, Y.; Lee, K.; Leung, K.K.; Skartsis, N.; et al. Precision Engineering of an Anti-HLA-A2 Chimeric Antigen Receptor in Regulatory T Cells for Transplant Immune Tolerance. Front. Immunol. 2021, 12, 686439. [Google Scholar] [CrossRef]
- Zhang, L.; Sosinowski, T.; Cox, A.R.; Cepeda, J.R.; Sekhar, N.S.; Hartig, S.M.; Miao, D.; Yu, L.; Pietropaolo, M.; Davidson, H.W. Chimeric antigen receptor (CAR) T cells targeting a pathogenic MHC class II:peptide complex modulate the progression of autoimmune diabetes. J. Autoimmun. 2019, 96, 50–58. [Google Scholar] [CrossRef]
- Crawford, F.; Stadinski, B.; Jin, N.; Michels, A.; Nakayama, M.; Pratt, P.; Marrack, P.; Eisenbarth, G.; Kappler, J.W. Specificity and detection of insulin-reactive CD4+ T cells in type 1 diabetes in the nonobese diabetic (NOD) mouse. Proc. Natl. Acad. Sci. USA 2011, 108, 16729–16734. [Google Scholar] [CrossRef]
- Lamarche, C.; Ward-Hartstonge, K.; Mi, T.; Lin, D.T.S.; Huang, Q.; Brown, A.; Edwards, K.; Novakovsky, G.E.; Qi, C.N.; Kobor, M.S.; et al. Tonic-signaling chimeric antigen receptors drive human regulatory T cell exhaustion. Proc. Natl. Acad. Sci. USA 2023, 120, e2219086120. [Google Scholar] [CrossRef]
- Chen, D.; Kakabadse, D.; Fishman, S.; Weinstein-Marom, H.; Davies, J.; Boldison, J.; Thayer, T.C.; Wen, L.; Gross, G.; Wong, F.S. Novel engineered B lymphocytes targeting islet-specific T cells inhibit the development of type 1 diabetes in non-obese diabetic Scid mice. Front. Immunol. 2023, 14, 1227133. [Google Scholar] [CrossRef] [PubMed]
- den Hollander, N.H.M.; Roep, B.O. From Disease and Patient Heterogeneity to Precision Medicine in Type 1 Diabetes. Front. Med. 2022, 9, 932086. [Google Scholar] [CrossRef] [PubMed]
- Casgevy, C. Available online: https://www.cnn.com/2023/11/16/health/uk-casgevy-approval-crispr-gene-editing-sickle-cell-scn?cid=external-feeds_iluminar_google (accessed on 28 November 2023).
| Cas13 nucleases (Type VI) | RNA-guided ribonucleases. Process pre-crRNA into mature crRNA. Collateral activity, allowing gene knockdown, base editing, and nucleic acid detection in mammalian cells. |
| Cas9 nucleases (Type II) | dsDNA cleavage with RuvC and HNH domains. Requires crRNA and tracrRNA for guide RNA. Mostly generates blunt-end cleavage. |
| Cas12 nucleases (Type V) | dsDNA cleavage with RuvC domain. Requires crRNA for guide RNA. Generates staggered-end cleavage, often with a T-rich PAM preference. |
| Cascade–Cas3 (Type I) | Type I CRISPR-associated complex for antiviral defense. Forms R-loop structure with crRNA and target DNA. Cas3 is recruited for specific dsDNA cleavage. |
| Cas7–11 nucleases (Type III) | Generally, uses several Cas proteins to target RNA. Subtype III-E uses a single-protein effector for recognizing and cleaving target RNA. Examples, such as DiCas7-11, exhibit robust RNA knockdown activity without collateral cleavage. |
| Target | Outcome | Reference |
|---|---|---|
| NOD.Ptpn22R619W | Increased IAAs | [13] |
| NOD.Aicda−/− | Delayed T1D development | [6] |
| PTPN1−/− PTPN6−/− | Protected β-cells from cytokine-induced cell death | [28] |
| NOD.HLA class II variants | Restored pathogenic CD8+ T-cell development and T1D susceptibility | [29] |
| Nfkbid−/− | Accelerated T1D onset | [26] |
| CNS1−/− | Decrease in RORγt+ Tregs | [30] |
| Rnls−/− | Protected β-cells against autoimmune destruction | [31] |
| NOD.STING−/− | Higher incidence of spontaneous diabetes | [32] |
| NOD.Dusop10−/− | Downregulation of type I interferon signature genes in islet cells | [10] |
| CXCL10−/− | Improved SC-islet survival | [33] |
| Lentivirus thymic transfer | Prevented autoimmunity | [34] |
| MND promoter | Prevented T1D development | [35] |
| TCRα | Enhanced the avidity of an antigen-specific Treg cell product | [36] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
El Nahas, R.; Al-Aghbar, M.A.; Herrero, L.; van Panhuys, N.; Espino-Guarch, M. Applications of Genome-Editing Technologies for Type 1 Diabetes. Int. J. Mol. Sci. 2024, 25, 344. https://doi.org/10.3390/ijms25010344
El Nahas R, Al-Aghbar MA, Herrero L, van Panhuys N, Espino-Guarch M. Applications of Genome-Editing Technologies for Type 1 Diabetes. International Journal of Molecular Sciences. 2024; 25(1):344. https://doi.org/10.3390/ijms25010344
Chicago/Turabian StyleEl Nahas, Rana, Mohammad Ameen Al-Aghbar, Laura Herrero, Nicholas van Panhuys, and Meritxell Espino-Guarch. 2024. "Applications of Genome-Editing Technologies for Type 1 Diabetes" International Journal of Molecular Sciences 25, no. 1: 344. https://doi.org/10.3390/ijms25010344
APA StyleEl Nahas, R., Al-Aghbar, M. A., Herrero, L., van Panhuys, N., & Espino-Guarch, M. (2024). Applications of Genome-Editing Technologies for Type 1 Diabetes. International Journal of Molecular Sciences, 25(1), 344. https://doi.org/10.3390/ijms25010344







