Comparative Transcriptome and Widely Targeted Metabolome Analysis Reveals the Molecular Mechanism of Powdery Mildew Resistance in Tomato
Abstract
1. Introduction
2. Results
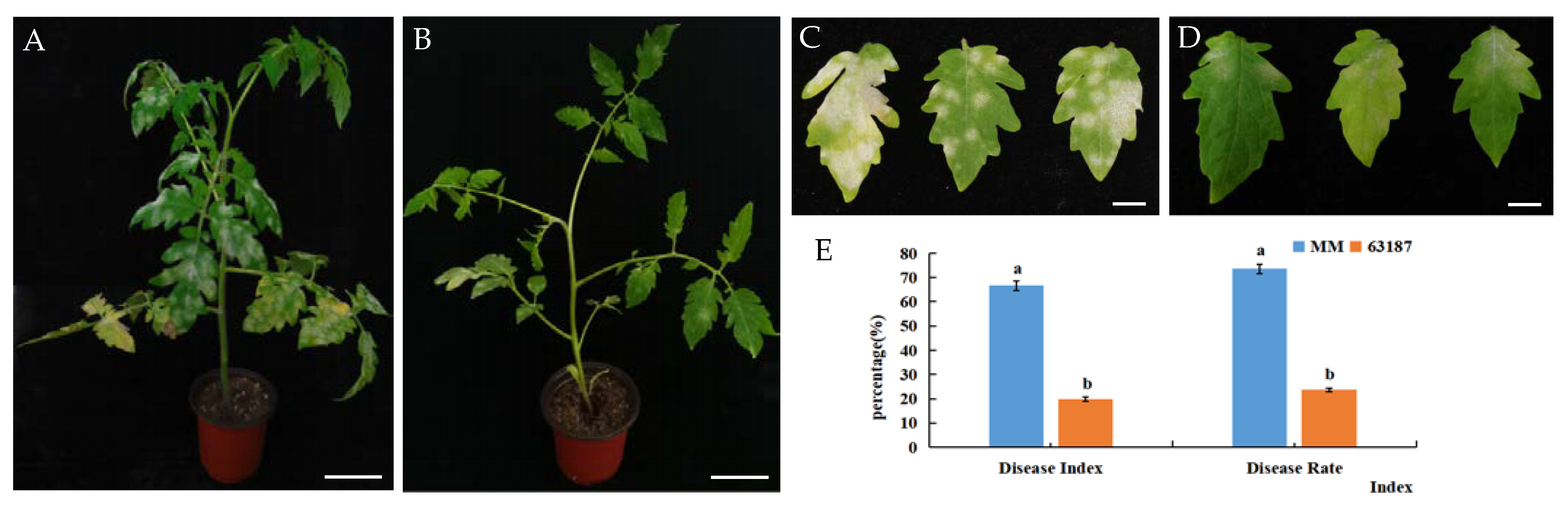
2.1. Phenotype Changes of Two Tomato Materials after Inoculation with O. neolycopersici
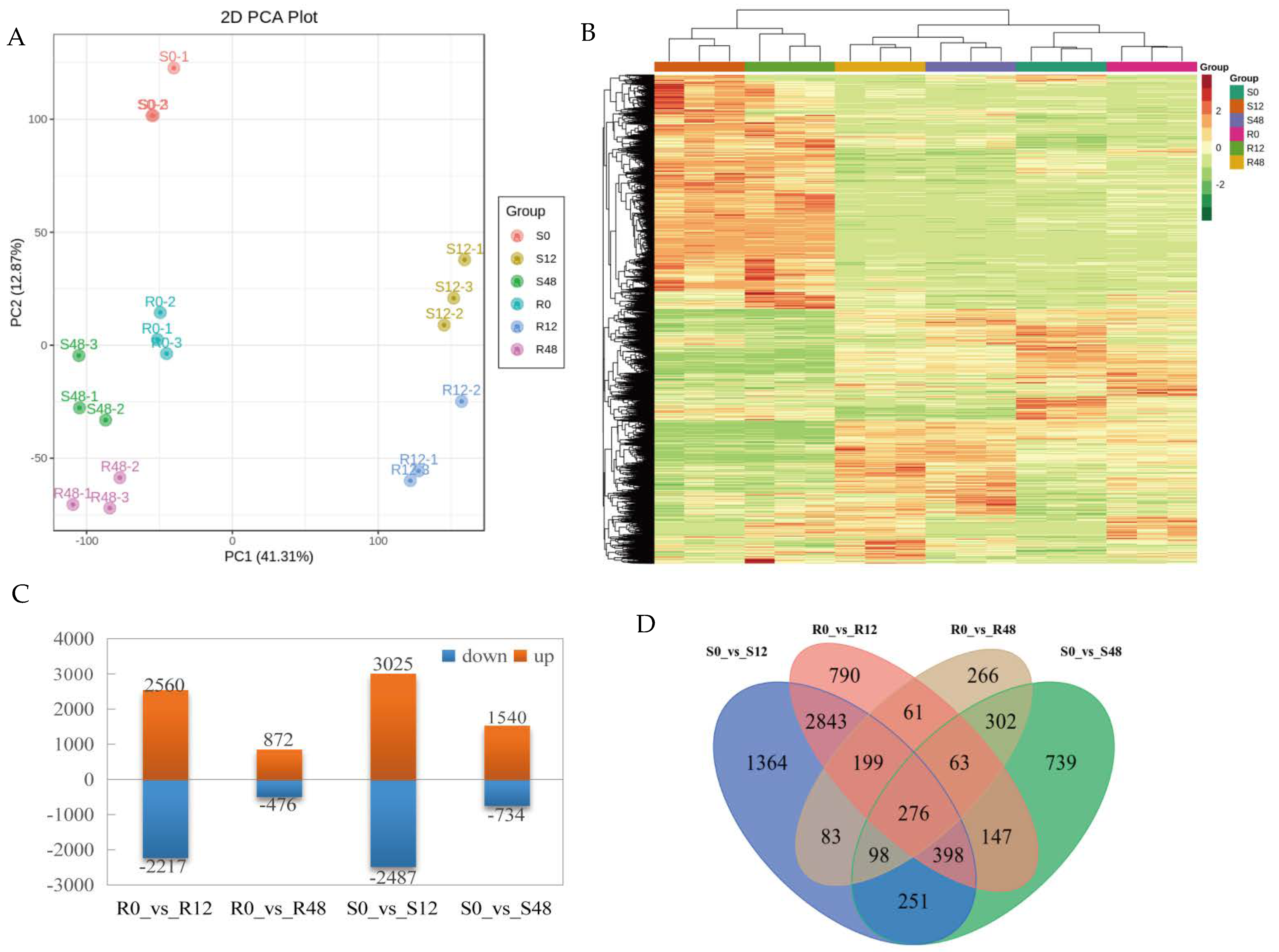
2.2. Transcriptome Analysis
2.2.1. Identification and Annotation of DEGs
2.2.2. Functional Annotation and Classification of DEGs
2.2.3. K-Means Clustering Analysis of DEGs
2.2.4. Transcription Factor Analysis of DEGs
2.2.5. Verifying DEGs by qRT-PCR
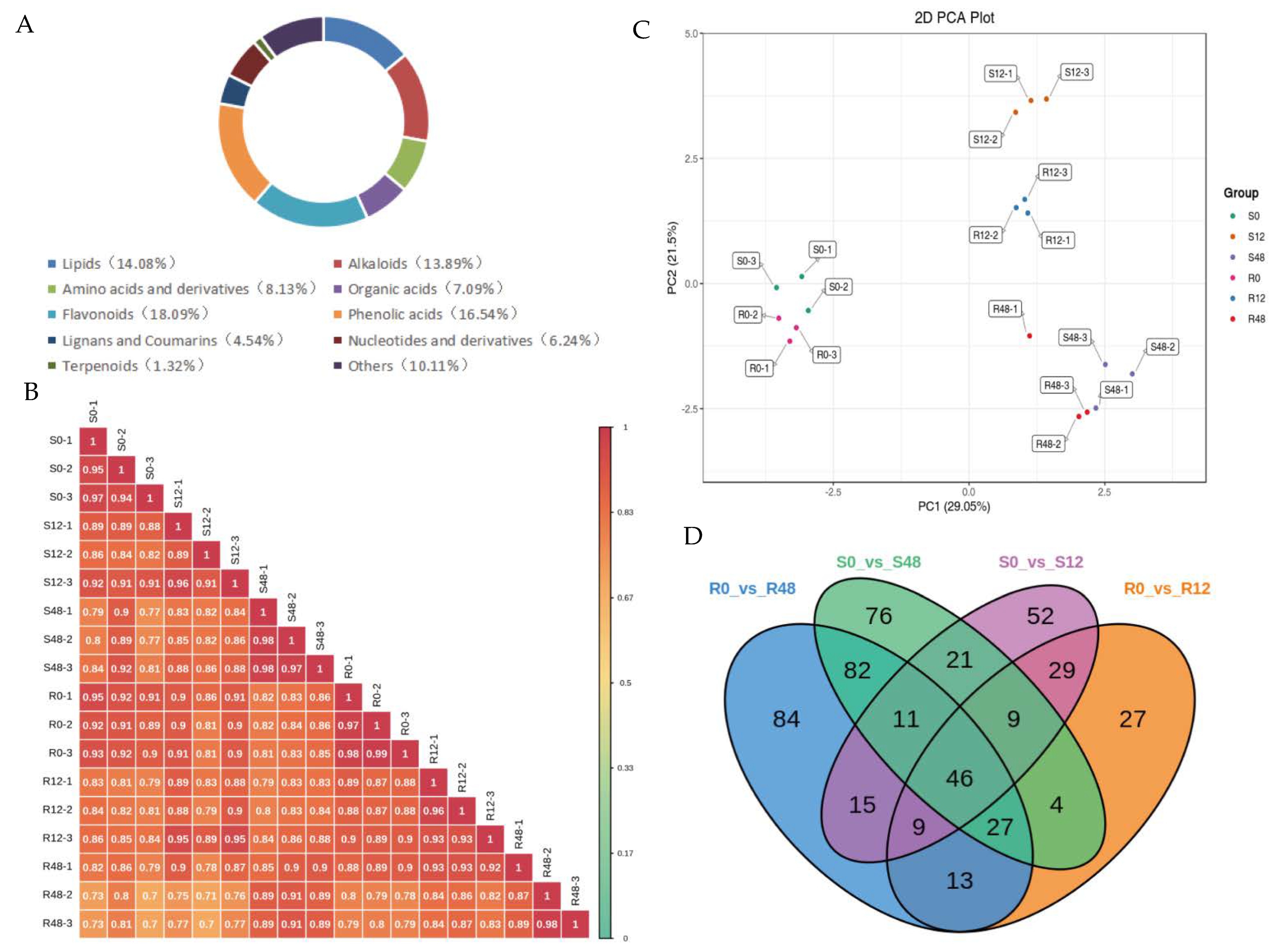
2.3. Metabolome Analysis
2.3.1. Identification of Differentially Accumulated Metabolites
2.3.2. K-Means Cluster Analysis of DAMs
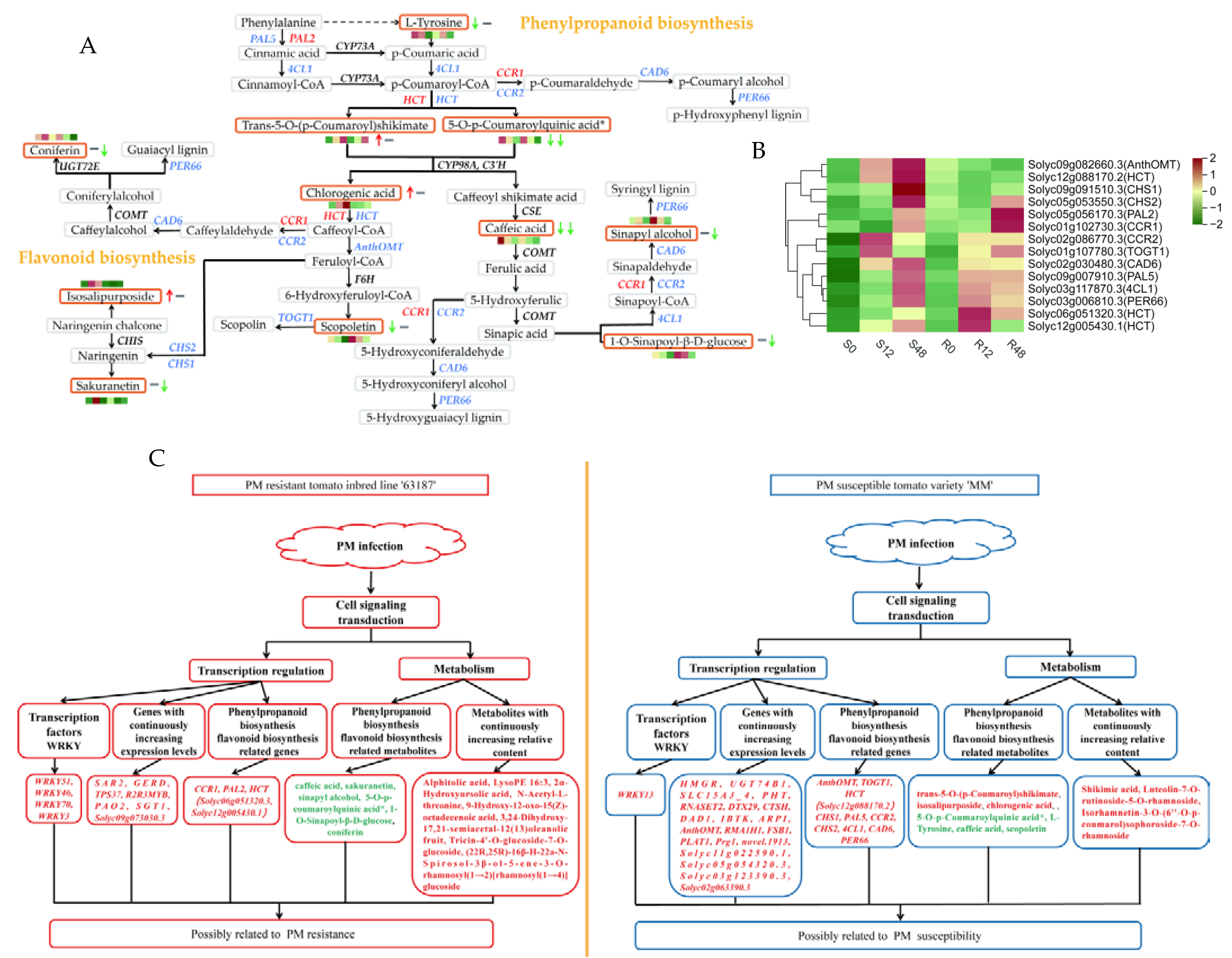
2.4. Correlation Analysis of DEGs and DAMs
2.4.1. KEGG Enrichment Analysis of Co-Expressed DEGs and DAMs
2.4.2. KEGG Enrichment Analysis of Especially Expressed DEGs and DAMs
3. Discussion
4. Materials and Methods
4.1. Plant Growth, Inoculation, and Propagation of O. neolycopersici
4.2. RNA Extraction and RNA-Seq
4.3. Read Mapping and Data Analysis
4.4. Validation of the Quantitative PCR
4.5. Metabolite Extraction
4.6. Qualitative and Quantitative Metabolite Analysis
4.7. Combined Transcriptome and Metabolome Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kurina, A.B.; Solovieva, A.E.; Khrapalova, I.A.; Artemyeva, A.M. Biochemical composition of tomato fruits of various colors. Vavilovskii Zhurnal Genet. Sel. 2021, 25, 514–527. [Google Scholar] [CrossRef] [PubMed]
- Li, C.W.; Pei, D.L.; Wang, W.J.; Ma, Y.S.; Wang, L.; Wang, F.; Liu, J.L.; Zhu, W.M. First report of powdery mildew caused by Oidium neolycopersici on tomato in China. Plant Dis. 2008, 92, 1370. [Google Scholar] [CrossRef] [PubMed]
- Dong, H. Resistance of different tomato varieties against tomato powdery mildew during different growth stages. Plant Dis. Pests 2013, 4, 15–17. [Google Scholar]
- Mieslerova, B.; Lebeda, A.; Chetelat, R.T. Variation in response of wild lycopersicon and solanum spp. against tomato powdery mildew (Oidium lycopersici). J. Phytopathol. 2000, 148, 303–311. [Google Scholar] [CrossRef]
- Kim, D.; Jin, B.; Je, B.I.; Choi, Y.; Kim, B.S.; Jung, H.J.; Nou, I.S.; Park, Y. Development of DNA markers for Slmlo1.1, a new mutant allele of the powdery mildew resistance gene SlMlo1 in tomato (Solanum lycopersicum). Genome 2018, 61, 703–712. [Google Scholar] [CrossRef]
- Bai, Y.; Huang, C.C.; van der Hulst, R.; Meijer-Dekens, F.; Bonnema, G.; Lindhout, P. QTLs for tomato powdery mildew resistance (Oidium lycopersici) in lycopersicon parviflorum G1.1601 co-localize with two qualitative powdery mildew resistance genes. Mol. Plant-Microbe Interact. 2003, 16, 169–176. [Google Scholar] [CrossRef]
- Cao, Y.Y.; Diao, Q.N.; Chen, Y.Y.; Jin, H.J.; Zhang, Y.P.; Zhang, H.M. Development of KASP markers and identification of a QTL underlying powdery mildew resistance in melon (Cucumis melo L.) by bulked segregant analysis and RNA-Seq. Front. Plant Sci. 2021, 11, 593207. [Google Scholar] [CrossRef]
- Wang, Y.H.; Wu, D.H.; Huang, J.H.; Tsao, S.J.; Hwu, K.K.; Lo, H.F. Mapping quantitative trait loci for fruit traits and powdery mildew resistance in melon (Cucumis melo L.). Bot. Stud. 2016, 57, 19. [Google Scholar] [CrossRef]
- Cheng, H.; Kun, W.P.; Liu, D.S.; Su, Y.Q.; He, Q.W. Molecular cloning and expression analysis of CmMlo1 in melon. Mol. Biol. Rep. 2012, 39, 1903–1907. [Google Scholar] [CrossRef]
- Zhao, Z.Q.; Dong, Y.M.; Wang, J.Y.; Zhang, G.L.; Zhang, Z.B.; Zhang, A.P.; Wang, Z.J.; Ma, P.P.; Li, Y.Z.; Zhang, X.Y.; et al. Comparative transcriptome analysis of melon (Cucumis melo L.) reveals candidate genes and pathways involved in powdery mildew resistance. Sci. Rep. 2022, 12, 4936. [Google Scholar] [CrossRef]
- Zheng, L.; Zhang, M.; Zhuo, Z.; Wang, Y.; Gao, X.; Li, Y.; Liu, W.; Zhang, W. Transcriptome profiling analysis reveals distinct resistance response of cucumber leaves infected with powdery mildew. Plant Biol. 2021, 23, 327–340. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.L.; Gao, P.; Wan, Y.; Cui, H.N.; Fan, C.; Liu, S.; Luan, F.S. Comparative transcriptome profiling of genes and pathways related to resistance against powdery mildew in two contrasting melon genotypes. Sci. Hortic. 2018, 227, 169–180. [Google Scholar] [CrossRef]
- Xiang, G.S.; Zhang, H.; Jian, H.Y.; Yan, H.J.; Wang, Q.G.; Zhou, N.N.; Li, S.B.; Tang, K.X.; Qiu, X.Q. De Novo assembly and characterization of the transcriptome of susceptible and resistant rose species in response to powdery mildew. Sci. Hortic. 2019, 257, 108653. [Google Scholar] [CrossRef]
- Li, P.Q.; Ruan, Z.; Fei, Z.X.; Yan, J.J.; Tang, G.H. Integrated transcriptome and metabolome analysis revealed that flavonoid biosynthesis may dominate the resistance of zanthoxylum bungeanum against stem canker. J. Agric. Food Chem. 2021, 69, 6360–6378. [Google Scholar] [CrossRef]
- Yang, C.C.; Wu, P.F.; Yao, X.H.; Sheng, Y.; Zhang, C.C.; Lin, P.; Wang, K.L. Integrated transcriptome and metabolome analysis reveals key metabolites involved in camellia oleifera defense against anthracnose. Int. J. Mol. Sci. 2022, 23, 536. [Google Scholar] [CrossRef] [PubMed]
- Zhao, F.; Niu, K.; Tian, X.; Du, W. Triticale improvement: Mining of genes related to yellow rust resistance in triticale based on transcriptome sequencing. Front. Plant Sci. 2022, 13, 883147. [Google Scholar] [CrossRef]
- Yuan, H.; Zeng, X.; Yang, Q.; Xu, Q.; Wang, Y.; Jabu, D.; Sang, Z.; Tashi, N. Gene co-expression network analysis combined with metabonomics reveals the resistance responses to powdery mildew in Tibetan hulless barley. Sci. Rep. 2018, 8, 14928. [Google Scholar] [CrossRef]
- Wen, F.; Wu, X.Z.; Li, T.J.; Jia, M.L.; Liao, L. Characterization of the WRKY gene family in akebia trifoliata and their response to colletotrichum acutatum. BMC Plant Biol. 2022, 22, 115. [Google Scholar] [CrossRef]
- Agapito, G.; Milano, M.; Cannataro, M. A python clustering analysis protocol of genes expression data sets. Genes 2022, 13, 1839. [Google Scholar] [CrossRef]
- Guo, W.L.; Chen, B.H.; Guo, Y.Y.; Yang, H.L.; Mu, J.Y.; Wang, Y.L.; Li, X.Z.; Zhou, J.G. Improved powdery mildew resistance of transgenic Nicotiana benthamiana over-expressing the cucurbita moschata CmSGT1 gene. Front. Plant Sci. 2019, 10, 955. [Google Scholar] [CrossRef]
- Xing, L.P.; Qian, C.; Cao, A.Z.; Li, Y.B.; Jiang, Z.N.; Li, M.H.; Jin, X.H.; Hu, J.M.; Zhang, Y.P.; Wang, X.; et al. The Hv-SGT1 gene from Haynaldia villosa contributes to resistances towards both biotrophic and hemi-biotrophic pathogens in common wheat (Triticum aestivum L.). PLoS ONE 2013, 8, e72571. [Google Scholar] [CrossRef] [PubMed]
- Milczarski, P.; Goralska, M.; Palatynska, K.; Wysoczanski, B.; Czyczylo-Mysza, I.; Maghuly, F.; Myskow, B. In search of the relationship between the rye polyamine oxidase (PAO) gene and resistance to powdery mildew (PM). J. Appl. Genet. 2022, 64, 65–70. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.H.; Guo, D.L.; Li, G.R.; Yang, Y.J.; Zhang, G.H.; Li, S.H.; Liang, Z.C. The grapevine R2R3-type MYB transcription factor VdMYB1 positively regulates defense responses by activating the stilbene synthase gene 2 (VdSTS2). BMC Plant Biol. 2019, 19, 478. [Google Scholar] [CrossRef]
- Thurich, J.; Meichsner, D.; Furch, A.; Pfalz, J.; Kruger, T.; Kniemeyer, O.; Brakhage, A.; Oelmuller, R. Arabidopsis thaliana responds to colonisation of Piriformospora indica by secretion of symbiosis-specific proteins. PLoS ONE 2018, 13, e209658. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.K.; Cho, S.K.; Son, O.; Xu, Z.Y.; Hwang, I.; Kim, W.T. Drought stress-induced Rma1H1, a RING membrane-anchor E3 ubiquitin ligase homolog, regulates aquaporin levels via ubiquitination in transgenic Arabidopsis plants. Plant Cell. 2009, 21, 622–641. [Google Scholar] [CrossRef] [PubMed]
- Curto, M.; Krajinski, F.; Schlereth, A.; Rubiales, D. Transcriptional profiling of medicago truncatula during Erysiphe pisi infection. Front. Plant Sci. 2015, 6, 517. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.G.; Song, Y.; Li, S.J.; Zhang, L.P.; Zou, C.S.; Yu, D.Q. The role of WRKY transcription factors in plant abiotic stresses. Biochim. Biophys. Acta 2012, 1819, 120–128. [Google Scholar] [CrossRef]
- Yin, W.; Wang, X.; Liu, H.; Wang, Y.; Nocker, S.; Tu, M.; Fang, J.; Guo, J.; Li, Z.; Wang, X. Over-expression of VqWRKY31 enhances powdery mildew resistance in grapevine by promoting salicylic acid signaling and specific metabolite synthesis. Hortic. Res.-Engl. 2022, 9, uhab064. [Google Scholar] [CrossRef]
- Bai, Y.L.; Sunarti, S.; Kissoudis, C.; Visser, R.; van der Linden, C.G. The role of tomato WRKY genes in plant responses to combined abiotic and biotic stresses. Front. Plant Sci. 2018, 9, 801. [Google Scholar] [CrossRef]
- Kissoudis, C.; Seifi, A.; Yan, Z.; Islam, A.T.; van der Schoot, H.; van de Wiel, C.C.; Visser, R.G.; van der Linden, C.G.; Bai, Y. Ethylene and abscisic acid signaling pathways differentially influence tomato resistance to combined powdery mildew and salt stress. Front. Plant Sci. 2016, 7, 2009. [Google Scholar] [CrossRef]
- Hu, Y.; Dong, Q.; Yu, D. Arabidopsis WRKY46 coordinates with WRKY70 and WRKY53 in basal resistance against pathogen Pseudomonas syringae. Plant Sci. 2012, 185–186, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.J.; Tao, F.; An, F.; Zou, Y.P.; Tian, W.; Chen, X.M.; Xu, X.M.; Hu, X.P. Wheat transcription factor TaWRKY70 is positively involved in high-temperature seedling plant resistance to Puccinia striiformis f. sp. tritici. Mol. Plant Pathol. 2016, 18, 649–661. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Huang, X.; Guo, L.; He, L.; Xiao, D.; Zhan, J.; Wang, A.; Liang, R. Comparative transcriptome analysis provides insights into the resistance in Pueraria [Pueraria lobata (Willd.) Ohwi] in response to pseudo-rust disease. Int. J. Mol. Sci. 2022, 23, 5223. [Google Scholar] [CrossRef] [PubMed]
- He, Q.Q.; Yang, L.; Zhang, J.Y.; Ma, J.N.; Ma, C.M. Chemical constituents of gold-red apple and their alpha-glucosidase inhibitory activities. J. Food Sci. 2014, 79, C1970–C1983. [Google Scholar] [CrossRef] [PubMed]
- Yeshi, K.; Turpin, G.; Jamtsho, T.; Wangchuk, P. Indigenous uses, phytochemical analysis, and anti-inflammatory properties of Australian tropical medicinal plants. Molecules 2022, 27, 3849. [Google Scholar] [CrossRef] [PubMed]
- Hu, Q.; Chen, Y.Y.; Jiao, Q.Y.; Khan, A.; Li, F.; Han, D.F.; Cao, G.D.; Lou, H.X. Triterpenoid saponins from the pulp of sapindus mukorossi and their antifungal activities. Phytochemistry 2018, 147, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Zhu, Y.Q.; Zhou, S.J. Comparative analysis of powdery mildew resistant and susceptible cultivated cucumber (Cucumis sativus L.) varieties to reveal the metabolic responses to Sphaerotheca fuliginea infection. BMC Plant Biol. 2021, 21, 24. [Google Scholar]
- Yu, H.; Li, H.Y.; Wei, R.F.; Cheng, G.; Zhou, Y.M.; Liu, J.B.; Xie, T.L.; Guo, R.R.; Zhou, S.H. Widely targeted metabolomics profiling reveals the effect of powdery mildew on wine grape varieties with different levels of tolerance to the disease. Foods 2022, 11, 2461. [Google Scholar] [CrossRef]
- Hu, X.P.; Puri, K.D.; Gurung, S.; Klosterman, S.J.; Wallis, C.M.; Britton, M.; Durbin-Johnson, B.; Phinney, B.; Salemi, M.; Short, D.; et al. Proteome and metabolome analyses reveal differential responses in tomato -verticillium dahliae-interactions. J. Proteom. 2019, 207, 103449. [Google Scholar] [CrossRef]
- Kavil, S.; Otti, G.; Bouvaine, S.; Armitage, A.; Maruthi, M.N. PAL1 gene of the phenylpropanoid pathway increases resistance to the Cassava brown streak virus in cassava. Virol. J. 2021, 18, 184. [Google Scholar] [CrossRef]
- Pant, S.R.; Irigoyen, S.; Liu, J.X.; Bedre, R.; Christensen, S.A.; Schmelz, E.A.; Sedbrook, J.C.; Scholthof, K.; Mandadi, K.K. Brachypodium phenylalanine ammonia lyase (PAL) promotes antiviral defenses against panicum mosaic virus and its satellites. mBio 2021, 12, e03518-20. [Google Scholar] [CrossRef] [PubMed]
- Kanobe, C.; McCarville, M.T.; O’Neal, M.E.; Tylka, G.L.; MacIntosh, G.C. Soybean aphid infestation induces changes in fatty acid metabolism in soybean. PLoS ONE. 2015, 10, e145660. [Google Scholar] [CrossRef] [PubMed]
- Yara, A.; Yaeno, T.; Montillet, J.L.; Hasegawa, M.; Seo, S.; Kusumi, K.; Iba, K. Enhancement of disease resistance to Magnaporthe grisea in rice by accumulation of hydroxy linoleic acid. Biochem. Biophys. Res. Commun. 2008, 370, 344–347. [Google Scholar] [CrossRef]
- Yuan, X.W.; Li, Y.X.; Liu, S.Y.; Xia, F.; Li, X.Z.; Qi, B.X. Accumulation of eicosapolyenoic acids enhances sensitivity to abscisic acid and mitigates the effects of drought in transgenic Arabidopsis thaliana. J. Exp. Bot. 2014, 65, 1637–1649. [Google Scholar] [CrossRef]
- Lee, M.W.; Padilla, C.S.; Gupta, C.; Galla, A.; Pereira, A.; Li, J.; Goggin, F.L. The fatty acid desaturase2 family in tomato contributes to primary metabolism and stress responses. Plant Physiol. 2020, 182, 1083–1099. [Google Scholar] [CrossRef]
- Zheng, K.; Jiang, J.B.; Kang, L.G. Identification method of tomato powdery mildew seedling resistance and screening of resistant germplasm resources. Plant Prot. 2012, 38, 105–107. (In Chinese) [Google Scholar]
- Jacob, D.; David, D.R.; Sztjenberg, A.; Elad, Y. Conditions for development of powdery mildew of tomato caused by Oidium neolycopersici. Phytopathology 2008, 98, 270–281. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.F.; Zhou, Y.Q.; Chen, Y.R.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018, 34, 884–890. [Google Scholar] [CrossRef]
- Kim, D.; Landmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 121–357. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. Feature counts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Hao, Q.H.; Peng, W.; Wang, J.; Tu, Y.; Li, H.; Zhu, T.M. The correlation between internet addiction and interpersonal relationship among teenagers and college students based on pearson’s correlation coefficient: A systematic review and meta-analysis. Front. Psychiatry 2022, 13, 818494. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Varet, H.; Brillet-Gueguen, L.; Coppee, J.Y.; Dillies, M.A. SARTools: A DESeq2-and EdgeR-Based R pipeline for comprehensive differential analysis of RNA-Seq data. PLoS ONE 2016, 11, e157022. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Araki, M.; Goto, S.; Hattori, M.; Hirakawa, M.; Itoh, M.; Katayama, T.; Kawashima, S.; Okuda, S.; Tokimatsu, T.; et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 2008, 36, 480–484. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. The gene ontology consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Jiao, C.; Sun, H.; Rosli, H.G.; Pombo, M.A.; Zhang, P.; Banf, M.; Dai, X.; Martin, G.B.; Giovannoni, J.J.; et al. iTAK: A program for genome-wide prediction and classification of plant transcription factors, transcriptional regulators, and protein kinases. Mol. Plant 2016, 9, 1667–1670. [Google Scholar] [CrossRef]
- Perez-Rodriguez, P.; Riano-Pachon, D.M.; Correa, L.G.; Rensing, S.A.; Kersten, B.; Mueller-Roeber, B. PlnTFDB: Updated content and new features of the plant transcription factor database. Nucleic Acids Res. 2010, 38, 822–827. [Google Scholar] [CrossRef]
- Jin, J.; Zhang, H.; Kong, L.; Gao, G.; Luo, J. PlantTFDB 3.0: A portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res. 2014, 42, 1182–1187. [Google Scholar] [CrossRef]
- Abid, M.; Gu, S.; Zhang, Y.J.; Sun, S.; Li, Z.; Bai, D.F.; Sun, L.; Qi, X.J.; Zhong, Y.P.; Fang, J.B. Comparative transcriptome and metabolome analysis reveal key regulatory defense networks and genes involved in enhanced salt tolerance of Actinidia (kiwifruit). Hortic. Res.-Engl. 2022, 9, uhac189. [Google Scholar] [CrossRef]
- Rao, X.; Huang, X.; Zhou, Z.; Lin, X. An improvement of the 2−ΔΔCt method for quantitative real-time polymerase chain reaction data analysis. Biostat. Bioinforma. Biomath. 2013, 3, 71–85. [Google Scholar]
- Yang, S.H.; Liu, Y.; Wang, Q.; Sun, Y.P.; Guan, W.; Liu, Y.; Yang, B.Y.; Kuang, H.X. UPLC-MS/MS identification and quantification of withanolides from six parts of the medicinal plant datura metel L. Molecules 2020, 25, 1260. [Google Scholar] [CrossRef]
- Zhao, S.; Liu, H.; Su, Z.; Khoo, C.; Gu, L. Identifying cranberry juice consumers with predictive OPLS-DA models of plasma metabolome and validation of cranberry juice intake biomarkers in a double-blinded, randomized, placebo-controlled, cross-over study. Mol. Nutr. Food Res. 2020, 64, e1901242. [Google Scholar] [CrossRef] [PubMed]
- Zou, S.; Wu, J.; Shahid, M.Q.; He, Y.; Lin, S.; Liu, Z.; Yang, X. Identification of key taste components in loquat using widely targeted metabolomics. Food Chem. 2020, 323, 126822. [Google Scholar] [CrossRef] [PubMed]
- Ke, M.; Ye, Y.; Zhang, Z.; Gillings, M.; Qu, Q.; Xu, N.; Xu, L.; Lu, T.; Wang, J.; Qian, H. Synergistic effects of glyphosate and multiwall carbon nanotubes on Arabidopsis thaliana physiology and metabolism. Sci. Total Environ. 2021, 769, 145156. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, W.; Wang, X.; Song, L.; Yao, W.; Guo, M.; Cheng, G.; Guo, J.; Bai, S.; Gao, Y.; Li, J.; et al. Comparative Transcriptome and Widely Targeted Metabolome Analysis Reveals the Molecular Mechanism of Powdery Mildew Resistance in Tomato. Int. J. Mol. Sci. 2023, 24, 8236. https://doi.org/10.3390/ijms24098236
Liu W, Wang X, Song L, Yao W, Guo M, Cheng G, Guo J, Bai S, Gao Y, Li J, et al. Comparative Transcriptome and Widely Targeted Metabolome Analysis Reveals the Molecular Mechanism of Powdery Mildew Resistance in Tomato. International Journal of Molecular Sciences. 2023; 24(9):8236. https://doi.org/10.3390/ijms24098236
Chicago/Turabian StyleLiu, Wenjuan, Xiaomin Wang, Lina Song, Wenkong Yao, Meng Guo, Guoxin Cheng, Jia Guo, Shengyi Bai, Yanming Gao, Jianshe Li, and et al. 2023. "Comparative Transcriptome and Widely Targeted Metabolome Analysis Reveals the Molecular Mechanism of Powdery Mildew Resistance in Tomato" International Journal of Molecular Sciences 24, no. 9: 8236. https://doi.org/10.3390/ijms24098236
APA StyleLiu, W., Wang, X., Song, L., Yao, W., Guo, M., Cheng, G., Guo, J., Bai, S., Gao, Y., Li, J., & Kang, Z. (2023). Comparative Transcriptome and Widely Targeted Metabolome Analysis Reveals the Molecular Mechanism of Powdery Mildew Resistance in Tomato. International Journal of Molecular Sciences, 24(9), 8236. https://doi.org/10.3390/ijms24098236






